Abstract
Hematopoietic stem cells (HSCs) reside at the top of the hematopoietic hierarchy, exhibiting a unique capacity to self-renew and differentiate into all blood cells throughout the lifetime. However, how to prevent HSC exhaustion during long-term hematopoietic output is not fully understood. Here, we show that the homeobox transcription factor Nkx2-3 is required for HSC self-renewal by preserving metabolic fitness. We found that Nkx2-3 is preferentially expressed in HSCs with excessive regenerative potential. Mice with conditional deletion of Nkx2-3 displayed a reduced HSC pool and long-term repopulating capacity as well as increased sensitivity to irradiation and 5-flurouracil treatment due to impaired HSC quiescence. In contrast, overexpression of Nkx2-3 improved HSC function both in vitro and in vivo. Furthermore, mechanistic studies revealed that Nkx2-3 can directly control the transcription of the critical mitophagy regulator ULK1, which is essential for sustaining metabolic homeostasis in HSCs by clearing activated mitochondria. More importantly, a similar regulatory role of NKX2-3 was observed in human cord blood-derived HSCs. In conclusion, our data demonstrate an important role of the Nkx2-3/ULK1/mitophagy axis in regulating the self-renewal of HSCs, therefore providing a promising strategy to improve the function of HSCs in the clinic.
Similar content being viewed by others
Introduction
Hematopoietic stem cells (HSCs) are characterized by their robust self-renewal and multilineage differentiation ability to guarantee lifelong hematopoietic output [1, 2]. In the adult mammalian bone marrow (BM), most HSCs are retained in a dormant state (G0 phase) to avoid stem cell pool exhaustion and prevent them from being insulted by various cytotoxic factors [3]. Studies have shown that HSC function is tightly regulated by many cell-intrinsic factors and BM niche-derived factors [4, 5]. However, our understanding of HSC biology remains obscure because of the heterogeneity of HSC populations and the complexity of the molecular regulatory network.
Recently, metabolic regulation of stem cell fate has become an increasing concern [6]. Under normal circumstances, HSCs mainly depend on anaerobic glycolysis rather than oxidative phosphorylation (OXPHOS) for energy supply [7, 8]. Previous studies, including our own work, have reported that abnormal activation of mitochondria will produce excessive metabolic byproducts, which disrupt HSC quiescence and function [9, 10]. Mitochondrial homeostasis is maintained by a combination of biosynthesis, fusion/fission and autophagy, also known as mitophagy [11,12,13]. Surprisingly, mitochondrial mass is higher in HSCs than their progenies [14], whereas how to maintain them in a low metabolic state is not completely understood. In addition, whether inhibiting the excessive activation of mitochondria can improve HSC function during proliferative stress is worth exploring.
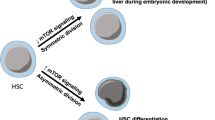
Nkx2-3 is a homeobox transcription factor family member that is involved in the differentiation and development of multiple organs by regulating tissue-specific gene expression [15]. As reported, systematic deletion of Nkx2-3 led to early postnatal lethality owing to the severe developmental defect of the small intestine [16]. In particular, Nkx2-3 knockout (Nkx2-3−/−) mice exhibited remarkable abnormalities in the hematopoietic and immune systems, which may be partly attributed to the developmental abnormity of the spleen [16, 17]. However, due to the limitations of animal models, previous studies could not clarify the role of Nkx2-3 in adult hematopoiesis. On the other hand, Nkx2-3 has been shown to facilitate tumorigenesis and progression, including lymphoma and leukemia [18, 19]. Typically, these tumors are the result of chromosomal translocations, whereby the expression of Nkx2-3 is perturbed by normally unrelated promoter activity [18, 19]. Nkx2-3 is also defined as a candidate autophagy-related gene in predicting cancer prognosis [20], while whether it participates in the autophagy process is largely unknown.
In this study, we found that Nkx2-3 is specifically enriched in HSCs with great self-renewal. Nkx2-3 deficiency resulted in gradual decreases in HSC number and long-term reconstitution ability due to the loss of HSC quiescence in both mice and humans. Mechanistically, Nkx2-3 promotes the metabolic fitness of HSCs by directly regulating the transcription of ULK1, which is required to clear activated mitochondria via mitophagy, therefore sustaining the self-renewal of HSCs. In summary, our findings not only disclose a previously unrecognized function of Nkx2-3 but also provide a deep understanding of mitophagy regulation in HSCs.
Materials and methods
Animals
Wild-type (WT) C57BL/6J mice were purchased from Vital River Lab Animal Technology Co., Ltd. (Beijing, China). Congenic CD45.1 mice were kindly gifted by Prof. Jinyong Wang [21]. Nkx2-3loxp/loxp (Nkx2-3fl/fl) mice were constructed at Cyagen Biosciences Inc (Suzhou, China). Nkx2-3EGFP/+, Mx1-Cre, Ulk1−/− and NOD-Prkdcscid Il2rgem1/Smoc (M-NSG) mice were obtained from Shanghai Model Organisms Center (China). Nkx2-3fl/fl/Mx1-Cre mice were generated by breeding Nkx2-3fl/fl mice with Mx1-Cre mice and then injected with 7 doses of polyinosinic:polycytidylic acid (pIpC; Sigma, St. Louis, MO, USA) to induce the deletion of Nkx2-3 in the hematopoietic system, as we previously reported [22]. All mice (C57BL/6J background) were used at 6–8 week (wk)-old and were sex-matched. Mice were randomly allocated and sample number was estimated based on extensive experience. No blinding was used and no sample was excluded. The animal experiments were carried out following the guidelines approved by the Animal Care Committee (Third Military Medical University, Chongqing, China).
BM transplantation (BMT) assays
For competitive BMT, 5 × 105 whole BM cells, 3 × 102 freshly sorted long-term HSCs (LT-HSCs) or 5 × 103 lentivirus-transfected Lineage- c-Kit+ Sca1- cells (LSKs) were mixed with 5 × 105 CD45.1 BM cells and then transplantated into lethally irradiated (10.0 Gy) CD45.1 mice. For serial BMT, 1 × 106 BM cells from primary recipients were transplantated into lethally irradiated (10.0 Gy) secondary CD45.1 recipient mice at 16 wk after the 1st transplantation, and a similar protocol was used for the 3rd and 4th rounds of BMT. Reciprocal transplantation assays were performed as we described previously [9, 23].
Chromatin immunoprecipitation (ChIP)
This assay was performed in LSKs after overexpression of Nkx2-3 using a NovoNGS® CUT&Tag 2.0 High-Sensitivity Kit (Novoprotein Scientific, Inc., Shanghai, China), as we reported [24].
Statistics
All experimental data were analyzed by GraphPad Prism 8.0 software (La Jolla, CA, USA). Variance was similar among the groups. Two-tailed Student’s t test and one-way analysis of variance (ANOVA) were used to compare the differences between two groups and multiple groups, respectively. Data are displayed as the means ± standard deviation (SD). All experiments were independently performed at least three times. *P < 0.05 was defined as statistically significant.
Additional methods
Other methods are provided in Supplementary Methods and antibodies or primers are listed in Supplementary Tables S1–5.
Results
Nkx2-3 is preferentially expressed in HSCs with great regenerative potential
Transcription factors are indispensable for regulating HSC self-renewal, while the underlying molecular machinery is still elusive. Here, we first performed single-cell RNA sequencing (scRNA-seq) analysis of murine hematopoietic stem progenitor cells (HSPCs) and identified 12 different cell populations (Fig. 1A, B; Supplementary Fig. S1B) according to a previous study [25]. It was found that a group of transcription factors was highly expressed in the C1 cluster, the most primitive region of HSPCs, compared with other clusters (Fig. 1C). Most of the transcription factors in the top 20 have been shown to modulate the self-renewal of HSCs, except for Nkx2-3 and Zbtb20 (Fig. 1C). Given the relative specificity of expression, we then focused on Nkx2-3. The data from public microarray, bulk RNA sequencing (RNA-seq) and scRNA-seq databases also showed that HSCs, especially LT-HSCs, expressed relatively more Nkx2-3 than committed progenitors and mature cells, which was confirmed by quantitative real-time PCR (qPCR) analysis (Fig. 1D; Supplementary Fig. S1C–F).
A–C LSKs freshly sorted form the BM of the normal WT mice were subjected to scRNA-seq analysis. Sorting strategies are provided in Supplementary Fig. S1A. Twelve different HSPC populations were identified according to reference cell clustering from a published study (GSE90742). C1-C3 unprimed (C1: the most primitive region), Mk megakaryocyte-primed, Er erythrocyte-primed, Mo1 type I macrophage-primed, Mo2 type II macrophage-primed, Ba basophilia-primed, Neu neutrophile-primed, DC dendritic cell-primed, B B cell-primed, T T cell-primed. A The workflow of scRNA-seq analysis. B The t-distributed stochastic neighbor embedding (tSNE) plots of 12 distinct clusters. C Dot plots showing the mRNA expression levels of the top 20 transcription factors in the 12 distinct clusters. D qPCR analysis of Nkx2-3 mRNA expression in the indicated cell populations purified from the BM of normal WT mice (n = 3). The relative expression of Nkx2-3 was compared with that in LT-HSCs. LT-HSC long-term HSC, ST-HSC short-term HSC, MPP multipotent progenitor, CMP common myeloid progenitor, MEP megakaryocyte erythroid progenitor, GMP granulocyte monocyte progenitor, CLP common lymphoid progenitor, Lin− Lineage− cells, Lin+ Lineage+ cells. Gating strategies are provided in Supplementary Fig. S1A. E The strategy of the generation of the Nkx2-3-EGFP reporter mouse model. F Flow cytometric analysis of the percentages of the indicated HSPCs in total, EGPF− and EGFP+ BM cells from Nkx2-3EGFP/+ mice (n = 5). Nkx2-3+/+ mice served as negative controls. G Flow cytometric analysis of EGFP expression in the indicated cell populations from Nkx2-3EGFP/+ mice (n = 5). Nkx2-3+/+ mice served as negative controls. Neg negative, MP myeloid progenitor, MFI mean fluorescence intensity. H The gating strategy for subsequent flow cytometric analysis and sorting of EGFPhi (about top 30%) and EGFPlo (about bottom 30%) LT-HSCs from the BM of Nkx2-3EGFP/+ mice. hi high expression, lo low expression. Nkx2-3+/+ mice served as negative controls. I Cell cycle analysis of EGFPhi and EGFPlo LT-HSCs from the BM of Nkx2-3EGFP/+ mice (n = 6). J Flow cytometric analysis of the percentage of LSKs after 7 days culture of 1 × 103 EGFPhi and EGFPlo LT-HSCs sorted from the BM of Nkx2-3EGFP/+ mice (n = 4). K–M A total of 3 × 102 EGFPhi or EGFPlo LT-HSCs sorted from the BM of Nkx2-3EGFP/+ mice (CD45.2) were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1). K The strategy of competitive transplantation. The percentage (L) and lineage distribution (M) of donor-derived cells in the peripheral blood (PB) of recipient mice at 16 wk after transplantation (n = 6). ***P < 0.001.
To further consolidate this finding, we generated an Nkx2-3EGFP/+ reporter mouse model, in which a 2A-EGFP cassette was knocked into the Nkx2-3 locus (Fig. 1E). EGFP knock-in did not significantly affect Nkx2-3 expression or the number and function of HSCs (Supplementary Fig. S2A–G), indicating that it is a reliable research tool. As a result, we observed a small EGFP+ population in the BM of Nkx2-3EGFP/+ mice and verified that Nkx2-3 mRNA was exclusively expressed in this subgroup (Fig. 1F; Supplementary Fig. S2H). Subsequent flow cytometric analysis showed that EGFP+ BM cells were enriched with most HSPCs (Fig. 1F). Alternatively, HSCs showed the highest expression level of Nkx2-3 in the BM (Fig. 1G). In view of the heterogeneity of HSC populations, we then divided LT-HSCs into two fractions based on EGFP expression (hereafter referred to as EGFPlo and EGFPhi HSCs) (Fig. 1H). Importantly, EGFPhi HSCs were more quiescent and displayed more potent regenerative potential both in vitro and in vivo, with no significant lineage skewing (Fig. 1I–M). Collectively, these results suggest that Nkx2-3 may play a unique role in HSC biology and can also function as a reliable marker of primitive HSCs.
Conditional knockout of Nkx2-3 results in a reduction in the HSC pool in mice
To assess whether Nkx2-3 regulates hematopoiesis, mice with conditional knockout of Nkx2-3 (Nkx2-3fl/fl/Mx1-Cre) were generated (Supplementary Fig. S3A). The deletion of Nkx2-3 was validated in LSKs only from Nkx2-3fl/fl/Mx1-Cre+ mice after pIpC injection (Supplementary Fig. S3B, C). Hereafter, Nkx2-3fl/fl/Mx1-Cre− and Nkx2-3fl/fl/Mx1-Cre+ mice are referred to as Nkx2-3fl/fl and Nkx2-3cKO mice, respectively. However, conventional hematopoietic parameters were comparable between two the groups at 4 and 20 wk after pIpC injection (Fig. 2A, B; Supplementary Fig. S3D, E).
A Nkx2-3fl/fl/Mx1-Cre+ and Nkx2-3fl/fl/Mx1-Cre− mice were intraperitoneally injected with 250 μg pIpC every two days for a total of 7 doses. At 4 wk and 20 wk after last injection, mice were subjected to further analysis. The experiment schematic is shown. B The total number of BM (left) and spleen (right) from Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). BM cells were obtained from one femur and one tibia. C Representative flow cytometric plots showing the percentages of MPs and LSKs in Nkx2-3fl/fl and Nkx2-3cKO BM. D The numbers of MPs and LSKs in the BM (one femur and one tibia) of Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). E Representative flow cytometric plots showing the percentages of LT-HSCs, ST-HSCs and MPPs in Nkx2-3fl/fl and Nkx2-3cKO BM. F The numbers of LT-HSCs, ST-HSCs and MPPs in the BM (one femur and one tibia) of Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). G Representative flow cytometric plots showing the percentage of SLAM-HSCs in Nkx2-3fl/fl and Nkx2-3cKO BM (n = 6). H The number of SLAM-HSCs in the BM (one femur and one tibia) of Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). Representative flow cytometric plots showing the percentages of MEPs, CMPs, GMPs (I) and CLPs (J) in Nkx2-3fl/fl and Nkx2-3cKO BM. K The numbers of MEPs, CMPs, GMPs and CLPs in the BM (one femur and one tibia) of Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). L The percentages of T cells (CD3e+), B cells (B220+) and myeloid cells (Gr1+ CD11b+) in the BM of Nkx2-3fl/fl and Nkx2-3cKO mice (n = 6). M A total of 1 × 102 LT-HSCs purified from the BM of Nkx2-3fl/fl and Nkx2-3cKO mice at 4 wk after pIpC injection were seeded in semi-solid medium for 7 days, and then, 1 × 104 cells were harvested for replating. Colonies were counted at 7 days after each replating (n = 4). N, O LSKs purified from the BM of WT mice were transduced with lentivirus carrying Nkx2-3 or scrambled control (Ctrl). Forty-eight hours later, transduced cells were sorted by flow cytometry. N qPCR analysis of Nkx2-3 mRNA expression in LSKs after overexpression of Nkx2-3 or Ctrl (n = 3). O Transduced cells were seeded in semi-solid medium for 7 days, and then, 1 × 104 cells were harvested for replating. Colonies were counted at 7 days after each replating (n = 4). *P < 0.05, **P < 0.01, ***P < 0.001.
Intriguingly, we found that the percentage and number of LSKs were largely unchanged at 4 wk but were decreased at 20 wk in Nkx2-3cKO mice after pIpC injection (Fig. 2C, D; Supplementary Fig. S3F). However, no such changes occurred in MPs (Fig. 2C, D; Supplementary Fig. S3F). Further analysis of LSK subpopulations showed that the percentage and number of LT-HSCs were gradually reduced in Nkx2-3cKO mice after pIpC injection, although to a lesser extent for ST-HSCs and MPPs (Fig. 2E, F; Supplementary Fig. S3G). Similar results were observed by using another set of HSC markers, signaling lymphocytic activation molecules (SLAM) (Fig. 2G, H; Supplementary Fig. S3H). Specifically, these effects were not due to increased HSC apoptosis caused by Nkx2-3 deficiency (Supplementary Fig. S3I). Among committed progenitors, only CMPs and CLPs were slightly lowered in Nkx2-3cKO mice at 20 wk after pIpC injection, along with normal distribution of mature hematopoietic cells (Fig. 2I–L; Supplementary Fig. S3J, K). These findings indicate that alterations in the HSPC phenotype following Nkx2-3 depletion might manifest over time. Consistent with this notion, the HSC serial replating experiment revealed that colony numbers were continuously diminished after a second round of replating when Nkx2-3 was deleted (Fig. 2M). However, overexpression of Nkx2-3 by lentivirus improved the replating ability of HSCs (Fig. 2N, O). Taken together, the above data demonstrate that Nkx2-3 is required for preserving the normal number of HSCs in mice.
Deletion of Nkx2-3 intrinsically compromises the long-term repopulation ability of HSCs
To test whether Nkx2-3 deletion affects HSC potential to self-renew, we first performed a competitive BMT assay (Fig. 3A). Donor chimerism levels in the peripheral blood (PB) and BM of recipients were significantly reduced when Nkx2-3 was ablated (Fig. 3B, C). The differences were gradual magnified from 4 to 16 wk after 1st and 2nd transplantation (Fig. 3B), indicating that Nkx2-3 is responsible for the maintenance of the long-term hematopoietic reconstruction ability of HSCs. However, lineage differentiation was unchanged after Nkx2-3 deletion (Fig. 3D), which is consistent with the data from untransplantated Nkx2-3cKO mice. To substantiate these findings, we conducted another BMT assay, in which Nkx2-3 knockout was induced by pIpC at 8 wk after transplantation (Fig. 3E). Similarly, we found a gradual loss of donor-derived cells in recipients’ PB and BM after Nkx2-3 ablation (Fig. 3F–H). In contrast, Nkx2-3 overexpression enforced the function of HSCs during serial transplantation (Fig. 3I–K). However, reciprocal BMT and homing assays showed that the above effects could not be attributed to microenvironment abnormities and HSC homing defects (Supplementary Fig. S4A–L). Thus, these results illustrate that Nkx2-3 sustains the self-renewal ability of HSCs in a cell-intrinsic manner.
A–D A total of 3 × 102 LT-HSCs purified from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (CD45.2) at 4 wk after pIpC injection were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1). Sixteen wk later, 1 × 106 BM cells from primary recipients were transplantated into lethally irradiated secondary recipient mice (CD45.1). A The strategy of competitive transplantation. B Donor chimerism levels in the PB of recipient mice at the indicated time after the 1st and 2nd transplantation (n = 6). Representative flow cytometric plots are shown in the left. C The chimerism levels of total BM cells, LSKs and LT-HSCs in recipient mice at 16 wk after the 1st and 2nd transplantation (n = 6). D The lineage distribution of donor-derived cells in the PB of recipient mice at 16 wk after the 1st and 2nd transplantation (n = 6). E–H A total of 5 × 105 BM cells from Nkx2-3fl/fl/Mx1-Cre− or Nkx2-3fl/fl/Mx1-Cre+ mice (CD45.2) were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1). Eight wk later, recipient mice were injected with 7 doses of pIpC to induce the deletion of Nkx2-3. At 20 wk after the 1st transplantation, 1 × 106 BM cells from primary recipients were transplantated into lethally irradiated secondary recipient mice (CD45.1). E The strategy of competitive transplantation and pIpC injection. F Donor chimerism levels in the PB recipient mice at the indicated time after the 1st and 2nd transplantation (n = 6). Representative flow cytometric plots are shown in the left. G The chimerism levels of donor-derived total BM cells, LSKs and LT-HSCs in recipient mice at 20 wk after the 1st transplantation or at 16 wk after the 2nd transplantation (n = 6). H The lineage distribution of donor-derived cells in the PB of recipient mice at 20 wk after the 1st transplantation or at 16 wk after the 2nd transplantation (n = 6). I–K A total of 5 × 103 WT LSKs (CD45.2) overexpression of Nkx2-3 or Ctrl were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1), followed by 2nd to 4th rounds of transplantation. I The strategy of Nkx2-3 overexpression and serial transplantation. The chimerism levels in the PB (J) and LT-HSC compartment (K) of recipient mice at 16 wk after the 1st to 4th rounds of transplantation (n = 6). **P < 0.01, ***P < 0.001.
Nkx2-3 deficiency significantly impairs the quiescence of HSCs
The gradual decrease in functional HSCs following Nkx2-3 ablation, combined with the finding that Nkx2-3 expression was higher in the G0 phase than in the G1 and S/G2/M phases (Fig. 4A), led us to hypothesize that Nkx2-3 may regulate the quiescence of HSCs. As anticipated, cell cycle analysis showed that HSC quiescence was modestly reduced at 4 wk but was markedly decreased at 20 wk in Nkx2-3cKO mice after pIpC injection (Fig. 4B; Supplementary Fig. S5A). Thus, the effects were intensively studied at 20 wk following pIpC injection thereafter. Further in vivo experiments displayed that Nkx2-3-null HSCs incorporated more 5-bromodeoxyuridine (BrdU) than controls (Fig. 4C). Meanwhile, the expression of several cyclins was unregulated and the expression of several cell cycle inhibitors was downregulated in HSCs after Nkx2-3 ablation (Fig. 4D). Consistent with these data, HSCs lacking Nkx2-3 displayed increased sensitivity to 5-fluorouracil (5-FU) and irradiation treatment (Fig. 4E–H; Supplementary Fig. S5B–E). Conversely, we found that overexpression of Nkx2-3 promoted the quiescence of HSCs (Supplementary Fig. S5F).
A Flow cytometric analysis of EGFP expression in the G0, G1 and S/G2/M phrases of LT-HSCs from Nkx2-3EGFP/+ mice (n = 5). B–J All mice were used at 20 wk after pIpC injection. B Cell cycle analysis of LSKs and LT-HSCs in the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 6). C Flow cytometric analysis of the percentage of BrdU+ cells in LSKs and LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 6). D qPCR analysis of mRNA expression of cell cycle-associated genes in LT-HSCs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 3). The numbers of LSKs (E) and LT-HSCs (F) from the BM (one femur and one tibia) of Nkx2-3fl/fl or Nkx2-3cKO mice after administrated with a single dose of 5-FU (150 mg/kg) via intraperitoneal injection (n = 5). G The survival rates of Nkx2-3fl/fl or Nkx2-3cKO mice after sequentially injected with 5-FU (150 mg/kg) once per wk for a total of 3 wk (n = 10). Data were analyzed by Log-rank nonparametric test and shown as Kaplan–Meier curves. H Representative immunofluorescence plots showing γ-H2AX expression in LT-HSCs freshly sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice 2 h following 2.0 Gy total body irradiation (TBI). I, J LT-HSCs purified from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice were subjected to RNA-seq analysis (n = 3). I The volcano plot showing DEGs between Nkx2-3fl/fl or Nkx2-3cKO LT-HSCs. Representative DEGs are indicated. J GSEA of gene signatures associated with HSC quiescence, stemness, long-term hematopoiesis, proliferation, cell cycle and DNA replication. NES normalized enrichment score, FDR false discovery rate. *P < 0.05, **P < 0.01, ***P < 0.001.
We then analyzed the transcriptional profile changes by bulk RNA-seq and identified 2445 differentially expressed genes (DEGs) between Nkx2-3fl/fl and Nkx2-3cKO HSCs, among which 822 genes were upregulated and 1623 genes were downregulated (Fig. 4I). In agreement with the above data, gene set enrichment analysis (GSEA) results revealed the negative enrichment of genes for HSC quiescence, stemness and long-term hematopoiesis and the positive enrichment of genes for HSC proliferation, cell cycle and DNA replication in Nkx2-3-deficient HSCs (Fig. 4J). Therefore, our findings indicate that Nkx2-3 is essential for regulating the quiescence of HSCs.
Ablation of Nkx2-3 leads to the accumulation of activated mitochondria in HSCs due to attenuated mitophagy
To gain insight into the underlying mechanism by which Nkx2-3 regulates HSC function, we further analyzed the RNA-seq data. Notably, a marked upregulation of genes involved in cell metabolism, including OXPHOS, tricarboxylic acid (TCA) cycle, NADH dehydrogenase ubiquitin activity, etc., in HSCs after Nkx2-3 deletion was observed, accompanied by strikingly enhanced expression of mitochondrial component-associated genes (Fig. 5A; Supplementary Fig. S6A). Considering that mitochondria are critical for HSC fate determination, we then examined mitochondrial properties. The results proved that mitochondrial mass and membrane potential were considerably increased in HSCs after Nkx2-3 knockout (Fig. 5B–D; Supplementary Fig. S6B). Supporting these findings, morphological analysis using transmission electron microscopy (TEM) displayed enhanced mitochondrial number and folds of cristae in Nkx2-3-deleted HSCs (Fig. 5E). In addition, glucose uptake, oxygen consumption rate (OCR) and ATP level were significantly increased, while the extracellular acidification rate (ECAR) and pyruvate and lactate content were decreased in HSCs in the absence of Nkx2-3 (Fig. 5F; Supplementary Fig. S6C–G). Moreover, ROS levels were evidently elevated in Nkx2-3-deleted HSCs, which principally originated from mitochondria (Fig. 5G; Supplementary Fig. S6H). These observations clearly demonstrate that Nkx2-3 deficiency causes the accumulation of active mitochondria in HSCs.
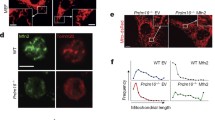
A–K All mice were used at 20 wk after pIpC injection. A GSEA results revealing upregulation of multiple metabolism processes in Nkx2-3cKO LT-HSCs. B Flow cytometric analysis of mitochondrial mass in LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice by MitoTracker Green (MTG) staining (n = 6). C Representative immunofluorescence plots showing mitochondrial mass in LT-HSCs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice by MitoTracker Deep Red (MTDR) staining. D Flow cytometric analysis of mitochondrial membrane potential in LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice by DilC1(5) staining (n = 6). E Representative TEM images showing mitochondrial changes in LSKs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice. F The OCR of LSKs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 3). Oligo, oligomycin; FCCP, carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone; R&A, rotenone and antimycin. G Flow cytometric analysis of intracellular ROS levels in LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice by DCFH-DA staining (n = 6). H The relative colocalization of TOMM20 with LAMP1 in LT-HSCs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n ≥ 15 cells). Representative immunofluorescence images are shown in the left. I qPCR analysis of mRNA expression of mitophagy-associated genes in LT-HSCs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 3). J Flow cytometric analysis of ULK1 expression in LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (n = 6). ISO, isotype control. K Representative immunofluorescence staining of ULK1 in LT-HSCs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice. *P < 0.05, **P < 0.01, ***P < 0.001.
Mitochondrial mass is regulated by biosynthesis, fusion/fission and autophagy. However, the expression of genes associated with mitochondrial biosynthesis and fusion/fission was largely unchanged in HSCs in the absence of Nkx2-3, except for PGC-1α, which was even modestly downregulated (Supplementary Fig. S6I, J). We next attempted to evaluate whether Nkx2-3 deficiency influences autophagy, which is a lysosomal-mediated process that degrades cytoplasmic components, including proteins, lipids and organelles [26]. In fact, the bulk autophagy level was only slightly but not significantly attenuated in HSCs after Nkx2-3 deletion, as determined by Cyto-ID staining (Supplementary Fig. S6K). Interestingly, immunofluorescence showed that the colocalization of the mitochondrial marker TOMM20 and lysosomal marker LAMP1 was obviously decreased in HSCs with Nkx2-3 deficiency (Fig. 5H), hinting that loss of Nkx2-3 may specifically impair mitophagy. Autophagy has been shown to contribute to the clearance of active mitochondria, which is required to keep HSC quiescence [27, 28]. However, core Atg genes were not altered, with the exception of Atg4c, which was slightly upregulated (Supplementary Fig. S6L). Of note, by analyzing mitophagy-related gene expression, we discovered that Ulk1 was significantly downregulated in Nkx2-3-deficient HSCs (Fig. 5I), which was further confirmed at the protein level (Fig. 5J, K). ULK1-dependent autophagy is identified to maintain the self-renewal of many types of stem cells, such as leukemia stem cells and embryonic stem cells [29, 30]. Indeed, we observed that Ulk1 was also relatively enriched in normal HSCs and its knockout caused substantial loss of functional HSCs due to dampened mitophagy, which is similar to that in Nkx2-3cKO mice (Supplementary Fig. S7A–K). We therefore reasonably postulated that Nkx2-3 promotes HSC maintenance probably by regulating mitophagy.
Nkx2-3 regulates mitophagy in HSCs by transcriptionally controlling the expression of Ulk1
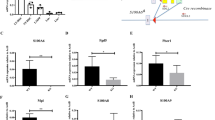
Studies have reported that AMPK and mTOR are well-recognized upstream signaling molecules that promote and inhibit ULK1 activation, respectively [31], whereas neither were altered after Nkx2-3 disruption (Supplementary Fig. S8A, B). We next sought to delineate whether Nkx2-3, as a transcription factor, can directly regulate the expression of Ulk1. By bioinformatics analysis, we searched two potential Nkx2-3 binding sites in the promoter region of the Ulk1 gene (Fig. 6A). Dual luciferase reporter assays revealed that the luciferase activity of a construct containing only WT binding site 1 was driven by Nkx2-3 overexpression (Fig. 6B). To exclude any indirect effects, we then conducted a ChIP assay and found that Nkx2-3 was able to directly bind site 1 but not site 2 (Fig. 6C). Moreover, overexpression of Nkx2-3 obviously upregulated the expression of Ulk1 in HSCs, further reinforcing this view (Fig. 6D). Overall, these results elucidate that Nkx2-3 regulates mitophagy in HSCs by directly driving Ulk1 transcription.
A Bioinformatics analysis revealing two potential Nkx2-3 binding sites in the promoter region of Ulk1 gene. B The luciferase reporter assays showing the transcriptional activity of Ulk1 upon overexpression of Nkx2-3 (n = 3). WT, wild-type; MUT, mutant. C ChIP analysis of LSKs sorted from the BM of WT mice after overexpression of Nkx2-3 or Ctrl. D qPCR analysis of Ulk1 mRNA expression in LSKs sorted from the BM of WT mice after overexpression of Nkx2-3 or Ctrl (n = 3). E–H Nkx2-3fl/fl/Mx1-Cre− or Nkx2-3fl/fl/Mx1-Cre+ mice were first injected with 7 doses of pIpC. From 5 wk to 20 wk after pIpC injection, mice were administrated with CPI-613 (12.5 mg/kg) or vehicle via intraperitoneal injection once a wk. E The strategy of CPI-613 administration. F Flow cytometric analysis of intracellular ROS levels in LT-HSCs from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice after CPI-613 or vehicle administration for 16 wk (n = 6). G The number of LT-HSCs in the BM (one femur and one tibia) of Nkx2-3fl/fl or Nkx2-3cKO mice after CPI-613 or vehicle administration for 16 wk (n = 6). H Cell cycle analysis of LT-HSCs in the BM of Nkx2-3fl/fl or Nkx2-3cKO mice after CPI-613 or vehicle administration for 16 wk (n = 6). I, J A total of 3 × 102 LT-HSCs purified from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice (CD45.2) were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1). Recipient mice were injected with CPI-613 or vehicle once a wk for a total of 16 wk. I The strategy of competitive transplantation and CPI-613 administration. J Donor chimerism levels in the PB of recipient mice after CPI-613 or vehicle administration at the indicated time after transplantation (n = 6). K–P LSKs sorted from the BM of Nkx2-3fl/fl or Nkx2-3cKO mice at 4 wk after pIpC injection were transduced with lentivirus carrying Ulk1 or Ctrl. Subsequently, transduced cells (5 × 103) were mixed with 5 × 105 BM cells (CD45.1) and then transplantated into lethally irradiated mice (CD45.1). K The strategy of Ulk1 overexpression and rescued transplantation. Flow cytometric analysis of mitochondrial mass (L), mitochondrial membrane potential (M) and intracellular ROS levels (N) in donor-derived LT-HSCs from the BM of recipients at 16 wk after transplantation (n = 5). O Cell cycle analysis of donor-derived LT-HSCs from the BM of recipients at 16 wk after transplantation (n = 5). P Donor chimerism levels in the PB of recipient mice at the indicated time after transplantation (n = 5). **P < 0.01, ***P < 0.001, ##P < 0.01, ###P < 0.001, ††P < 0.01, †††P < 0.001.
To examine whether mitochondrial activation is responsible for the impairment of HSC function after Nkx2-3 deletion, Nkx2-3cKO mice were treated with a mitochondrial metabolic inhibitor, CPI-613 [32]. Suppressing mitochondrial activity partly alleviated the loss of HSC number and quiescence in Nkx2-3cKO mice and then improved the long-term repopulation ability of HSCs with Nkx2-3 deficiency (Fig. 6E–J; Supplementary Fig. S8C–E). Consistently, scavenging intracellular ROS by N-acetylcysteine (NAC) also partly corrected the defects of HSCs in the absence of Nkx2-3 (Supplementary Fig. S8F–K), suggesting that ROS are crucial mediators of the functional damage to HSCs induced by activated mitochondria. For more direct evidence, HSCs purified from Nkx2-3cKO mice were transduced with lentivirus containing Ulk1 (Fig. 6K). Overexpression of Ulk1 substantially reduced activated mitochondria, therefore partially reversing the abnormal phenotype and function of Nkx2-3-null HSCs (Fig. 6L–P; Supplementary Fig. S8L). Altogether, Ulk1 downregulation contributes to the abnormality of HSCs with Nkx2-3 deficiency.
The NKX2-3/ULK1/mitophagy axis maintains the self-renewal of human HSCs
Finally, we wondered whether NKX2-3 plays a similar role in human HSCs. Data from two public databases showed that NKX2-3 expression was relatively higher in HSCs than in various progenitors and mature cells in human BM (Supplementary Fig. S9A, B). A similar expression pattern of NKX2-3 was confirmed in HSPCs obtained from human cord blood (CB) (Fig. 7A, B). Then, NKX2-3 was knocked down in CB-derived CD34+ HSPCs by lentivirus carrying short hairpin RNA against NKX2-3 (shNKX2-3) (Fig. 7C, D). We observed that decreasing NKX2-3 expression did not significantly change the total cultured cell number but reduced the percentage of culture-derived primitive HSCs, accompanied by decreased HSC quiescence (Fig. 7E—G). Accordingly, the replating ability of HSCs was decreased when NKX2-3 was knocked down (Fig. 7H). Subsequently, shNKX2-3-transduced CD34+ cells were transplantated into M-NSG mice, and we found that NKX2-3 knockdown impaired the hematopoietic reconstitution of HSCs, with no significant lineage skewing (Fig. 7I, J). However, the opposite effects were observed in HSCs after overexpression of NKX2-3 (Supplementary Fig. S9C-I). Further investigations showed that NKX2-3 knockdown reduced ULK1 expression and impaired mitophagy in CB-derived HSCs, as observed in mice (Fig. 7K–N; Supplementary Fig. S9J, K). Importantly, inhibition of mitochondrial metabolism or scavenging of ROS partially rescued the function of HSCs after NKX2-3 knockdown (Fig. 7O, P). Collectively, our data demonstrate that Nkx2-3 regulates mitophagy by transcriptional control of ULK1 expression, which is required to preserve HSC self-renewal (Fig. 7Q).
A The gating strategies for flow cytometric analysis or sorting of the following human CB subpopulations: hematopoietic stem cells, (HSCs; Lin− CD34+ CD38− CD90+ CD45RA−), multipotent progenitors (MPPs; Lin− CD34+ CD38− CD90− CD45RA−), multi-lymphoid progenitors (MLPs; Lin− CD34+ CD38− CD90− CD45RA+), hematopoietic stem progenitor cells (Lin− CD34+ CD38−), and hematopoietic progenitors (Lin− CD34+ CD38+). B qPCR analysis of NKX2-3 mRNA expression in the indicated cell populations purified from human CB (n = 3). The relative expression of NKX2-3 was compared with that in HSCs. C–P CD34+ cells sorted from human CB were transduced with lentivirus carrying shNKX2-3. Scrambled short hairpin RNA (shRNA) served as control and termed as shCtrl. Three days later, mCherry+ cells were purified by flow cytometry and the knockdown efficiency was tested by qPCR. Then, purified cells were used for further experiments, including culture, replating assay or xenotransplantation. C The experiment schematic is shown. D qPCR analysis of NKX2-3 mRNA expression in CD34+ cells at 3 days after shCtrl or shNKX2-3 transduction (n = 3). E The total number of cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells (n = 4). F Flow cytometric analysis of the percentage of CD34+ CD38− cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells (n = 4). G Cell cycle analysis of CD34+ CD38− cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells (n = 4). H The total number of colonies derived from CD34+ cells transducted with shCtrl or shNKX2-3 at 14 days after each replating. I, J CD34+ cells transducted with shCtrl or shNKX2-3 were transplantated into sublethally irradiated M-NSG mice. The percentage (I) and lineage distribution (J) of donor-derived cells (mCherry+ hCD45+) in the BM of recipient mice at 16 wk after transplantation were analyzed by flow cytometry (n = 8). CD19+, B cells; CD33+, myeloid cells. K qPCR analysis of ULK1 mRNA expression in CD34+ CD38− cells transducted with shCtrl or shNKX2-3 at 10 days after culture (n = 3). L Flow cytometric analysis of mitochondrial mass in CD34+ CD38− cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells by MTG staining (n = 4). M Flow cytometric analysis of mitochondrial membrane potential in CD34+ CD38− cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells by DilC1(5) staining (n = 4). N Flow cytometric analysis of intracellular ROS levels in CD34+ CD38− cells at 10 days after culture of shCtrl or shNKX2-3-transducted cells by DCFH-DA staining (n = 4). (O) CD34+ cells transducted with shCtrl or shNKX2-3 were transplantated into sublethally irradiated M-NSG mice. mice were administrated with NAC (1 mg/ml) or vehicle in drinking water for 16 wk. The percentage of donor-derived cells (mCherry+ hCD45+) in the BM of recipient mice at 16 wk after transplantation was analyzed by flow cytometry (n = 8). P CD34+ cells transducted with shCtrl or shNKX2-3 were transplantated into sublethally irradiated M-NSG mice. Recipient mice were administrated with CPI-613 (12.5 mg/kg) or vehicle via intraperitoneal injection once a wk for a total of 16 wk. The percentage of donor-derived cells (mCherry+ hCD45+) in the BM of recipient mice at 16 wk after transplantation was analyzed by flow cytometry (n = 8). Q The schematic diagram revealing how Nkx2-3 regulates the self-renewal of HSCs. *P < 0.05, **P < 0.01, ***P < 0.001.
Discussion
The maintenance of a functional HSC pool is necessary for continuous production of various blood cells throughout life and enabling hematopoietic recovery after stress injury [33]. How to elaborately regulate HSC function is still one of hot and difficult issue in the stem cell field. Recently, metabolism has emerged as a novel regulator of HSC biology, while the underlying molecular basis remains to be elucidated. In the present study, we show for the first time that Nkx2-3/ULK1/mitophagy-mediated metabolic fitness plays an important role in sustaining the self-renewal capacity of HSCs.
As a member of the Nkx gene family, Nkx2-3 was initially thought to be enriched in the spleen and small intestine but at considerably lower levels in the unfractionated BM [16]. Intriguingly, later studies using high-throughput analysis showed that Nkx2-3 is highly expressed in murine and human HSCs [34, 35], while little is known about its potential role in adult hematopoiesis. In this work, the specific expression pattern of Nkx2-3 in HSCs was corroborated by three separate methods. Subsequently, we found a gradual reduction in HSC percentage and number in Nkx2-3cKO mice after pIpC injection, revealing that Nkx2-3 is critical for the maintenance of the normal HSC pool. The discrepancy between our work and previous studies might be caused by the following reasons: (1) we used hematopoietic-specific Nkx2-3 knockout mice to rule out the possible influence of the microenvironment; (2) the deletion of Nkx2-3 was induced by pIpC injection in our study, which could exclude the developmental effect; (3) the early postnatal lethality of Nkx2-3−/− mice limited the ability of previous studies to monitor the knockdown effect over time; and (4) the phenotype of primitive HSCs was not identified in previous studies [16, 17]. However, we did not find significant differences in the conventional hematopoietic parameters in our mouse model, which provided further evidence that the perturbation of hematopoietic and immune systems in Nkx2-3−/− mice was mainly due to developmental abnormity. After all, Nkx2-3 is also abundantly present in embryonic HSCs, as shown by scRNA-seq data [36].
Next, we performed a series of BMT assays, and found that the long-term hematopoietic repopulation ability of HSCs was intrinsically impaired when Nkx2-3 was deleted. Studies have reported that Nkx2-3 governs lymphocyte homing by regulating MAdCAM-1 expression [37, 38]. However, our data showed that homing defects were not responsible for the reduced HSC function after Nkx2-3 ablation. Restriction of cell cycle activation is believed to effectively prevent the premature depletion of HSCs [4, 39]. Indeed, our data showed that loss of Nkx2-3 resulted in compromised maintenance of HSC quiescence. The above observations were further confirmed in human HSCs from CB. In contrast, enforcing Nkx2-3 expression promoted HSC quiescence and thus augmented HSC function in both mice and humans, consistent with a previous study showing that Nkx2-3 overexpression increased hemopoietic progenitor colony formation [19]. These findings indicate that Nkx2-3 may be a potential molecular target to improve HSC-related applications in the clinic. Accordingly, the above data indicate that Nkx2-3, as a transcription factor, plays a critical role in regulating the stemness of HSCs. In fact, most previously discovered stem cell regulators are transcription factors, including Egr1, Meis1, Pbx1, Nr4a1, Hlf and Mecom [40,41,42,43,44,45].
Adult HSCs prefer to utilize glycolysis instead of OXPHOS for their maintenance [7]. The conversion of the metabolic pattern from glycolysis to mitochondrial metabolism can elevate intracellular levels of ROS and intermediate metabolites, which lead to obvious oxidative stress damage and epigenetic changes, respectively, and eventually disrupt HSC quiescence and function [9, 46]. Therefore, HSC maintenance is incompatible with activated mitochondrial metabolism, while little is known about how to ensure that HSCs are in a low metabolic state. Autophagy is a lysosome-dependent pathway that acts to degrade injured or redundant intracellular constituents [47, 48]. Mitochondrial autophagy, also called mitophagy, is a special type of autophagy [49]. Studies have shown that deletion of the core Atg genes Atg5, Atg7 or Atg12 results in the accumulation of activated mitochondria and impaired HSC function, which suggests that mitophagy regulates normal hematopoiesis [27, 28]. As a consequence, disruption of PINK1-Parkin signaling, a well-recognized mitophagy-specific pathway, compromises HSC self-renewal [50, 51]. Here, we observed marked increases in mitochondrial mass and activity in HSCs after Nkx2-3 ablation, together with the downregulation of ULK1 but not PINK1 and Parkin. ULK1/ULK2 was originally defined as a core component of the autophagy initiation complex [52]. However, possibly because of the compensatory effect of ULK2, we found that knockout of Ulk1 did not significantly influence bulk autophagy, which is in accordance with previous studies [53, 54]. Particularly, ULK1, but not ULK2, has recently been shown to clear activated mitochondria by specifically mediating mitophagy, especially under hypoxic conditions [55, 56]. As expected, using Ulk1 knockout (Ulk1−/−) mice, we confirmed that Ulk1 deletion led to increased active mitochondria and defective HSC function, resembling those of Nkx2-3cKO mice in many aspects. These data prompted us to hypothesize that Nkx2-3 maintains HSC self-renewal probably by regulating ULK1-independent mitophagy.
Like other homeobox transcription factors, Nkx2-3 contains a conserved helix-turn-helix DNA-binding motif capable of binding specific DNA sequences [57]. Consequently, ChIP assays, luciferase reporter analysis and overexpression assays confirmed that Nkx2-3 can directly bind to the Ulk1 promoter region and thus regulate its expression in HSCs. It is understandable that metabolic changes can also occur following cell proliferation [6, 8]. To determine which events (metabolic and functional changes) occur earlier, we treated Nkx2-3cKO mice with a mitochondrial metabolism inhibitor or ROS scavenger. Importantly, these schemes significantly rescued the phenotype and function of Nkx2-3-deficient HSCs. Finally, an Ulk1 overexpression experiment revealed that Nkx2-3 deletion-induced Ulk1 downregulation may be a main reason for excessive mitochondrial activation in HSCs. However, we cannot entirely rule out the possibility that other ULK1-independent mechanisms may also be involved.
In summary, our data highlight the significant role of Nkx2-3 in promoting the long-term maintenance of HSCs by modulating metabolic homeostasis. Therefore, our findings unveil new insights into the metabolic regulation of HSC fate and provide a new target to improve the function of HSCs when suffering from proliferative stress.
Data availability
The raw data of scRNA-seq were deposited in the NCBI Gene Expression Omnibus (GEO) database (no. GSE221907). The raw data of bulk RNA-seq were deposited in the NCBI Sequence Read Archive (SRA) database (no. PRJNA891790).
References
Sanjuan-Pla A, Macaulay IC, Jensen CT, Woll PS, Luis TC, Mead A, et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature. 2013;502:232–6.
Cheng H, Zheng Z, Cheng T. New paradigms on hematopoietic stem cell differentiation. Protein Cell. 2020;11:34–44.
Sinha S, Dwivedi TR, Yengkhom R, Bheemsetty VA, Abe T, Kiyonari H, et al. Asrij/OCIAD1 suppresses CSN5-mediated p53 degradation and maintains mouse hematopoietic stem cell quiescence. Blood. 2019;133:2385–400.
Pinho S, Frenette PS. Haematopoietic stem cell activity and interactions with the niche. Nat Rev Mol Cell Biol. 2019;20:303–20.
Chen Z, Guo Q, Song G, Hou Y. Molecular regulation of hematopoietic stem cell quiescence. Cell Mol Life Sci. 2022;79:218.
Nakamura-Ishizu A, Ito K, Suda T. Hematopoietic stem cell metabolism during development and aging. Dev Cell. 2020;54:239–55.
Suda T, Takubo K, Semenza GL. Metabolic regulation of hematopoietic stem cells in the hypoxic niche. Cell Stem Cell. 2011;9:298–310.
Hu M, Wang J. Mitochondrial metabolism and the maintenance of hematopoietic stem cell quiescence. Curr Opin Hematol. 2019;26:228–34.
Hu M, Zeng H, Chen S, Xu Y, Wang S, Tang Y, et al. SRC-3 is involved in maintaining hematopoietic stem cell quiescence by regulation of mitochondrial metabolism in mice. Blood. 2018;132:911–23.
Zheng J, Lu Z, Kocabas F, Bottcher RT, Costell M, Kang X, et al. Profilin 1 is essential for retention and metabolism of mouse hematopoietic stem cells in bone marrow. Blood. 2014;123:992–1001.
Jin G, Xu C, Zhang X, Long J, Rezaeian AH, Liu C, et al. Atad3a suppresses Pink1-dependent mitophagy to maintain homeostasis of hematopoietic progenitor cells. Nat Immunol. 2018;19:29–40.
Luchsinger LL, de Almeida MJ, Corrigan DJ, Mumau M, Snoeck HW. Mitofusin 2 maintains haematopoietic stem cells with extensive lymphoid potential. Nature. 2016;529:528–31.
Qian P, He XC, Paulson A, Li Z, Tao F, Perry JM, et al. The Dlk1-Gtl2 locus preserves LT-HSC function by inhibiting the PI3K-mTOR pathway to restrict mitochondrial metabolism. Cell Stem Cell. 2016;18:214–28.
de Almeida MJ, Luchsinger LL, Corrigan DJ, Williams LJ, Snoeck HW. Dye-independent methods reveal elevated mitochondrial mass in hematopoietic stem cells. Cell Stem Cell. 2017;21:725–9.e4.
Holland PW. Evolution of homeobox genes. Wiley Interdiscip Rev Dev Biol. 2013;2:31–45.
Pabst O, Zweigerdt R, Arnold HH. Targeted disruption of the homeobox transcription factor Nkx2-3 in mice results in postnatal lethality and abnormal development of small intestine and spleen. Development. 1999;126:2215–25.
Tarlinton D, Light A, Metcalf D, Harvey RP, Robb L. Architectural defects in the spleens of Nkx2-3-deficient mice are intrinsic and associated with defects in both B cell maturation and T cell-dependent immune responses. J Immunol. 2003;170:4002–10.
Robles EF, Mena-Varas M, Barrio L, Merino-Cortes SV, Balogh P, Du MQ, et al. Homeobox NKX2-3 promotes marginal-zone lymphomagenesis by activating B-cell receptor signalling and shaping lymphocyte dynamics. Nat Commun. 2016;7:11889.
Dovey OM, Cooper JL, Mupo A, Grove CS, Lynn C, Conte N, et al. Molecular synergy underlies the co-occurrence patterns and phenotype of NPM1-mutant acute myeloid leukemia. Blood. 2017;130:1911–22.
Huang L, Yu X, Jiang Z, Zeng P. Novel autophagy-related gene signature investigation for patients with oral squamous cell carcinoma. Front Genet. 2021;12:673319.
Guo R, Hu F, Weng Q, Lv C, Wu H, Liu L, et al. Guiding T lymphopoiesis from pluripotent stem cells by defined transcription factors. Cell Res. 2020;30:21–33.
Hu M, Lu Y, Zeng H, Zhang Z, Chen S, Qi Y, et al. MicroRNA-21 maintains hematopoietic stem cell homeostasis through sustaining the NF-kappaB signaling pathway in mice. Haematologica. 2021;106:412–23.
Hu M, Lu Y, Wang S, Zhang Z, Qi Y, Chen N, et al. CD63 acts as a functional marker in maintaining hematopoietic stem cell quiescence through supporting TGFbeta signaling in mice. Cell Death Differ. 2022;29:178–91.
Zhang Z, Lu Y, Qi Y, Xu Y, Wang S, Chen F, et al. CDK19 regulates the proliferation of hematopoietic stem cells and acute myeloid leukemia cells by suppressing p53-mediated transcription of p21. Leukemia. 2022;36:956–69.
Rodriguez-Fraticelli AE, Wolock SL, Weinreb CS, Panero R, Patel SH, Jankovic M, et al. Clonal analysis of lineage fate in native haematopoiesis. Nature. 2018;553:212–6.
Doherty J, Baehrecke EH. Life, death and autophagy. Nat Cell Biol. 2018;20:1110–7.
Ho TT, Warr MR, Adelman ER, Lansinger OM, Flach J, Verovskaya EV, et al. Autophagy maintains the metabolism and function of young and old stem cells. Nature. 2017;543:205–10.
Mortensen M, Soilleux EJ, Djordjevic G, Tripp R, Lutteropp M, Sadighi-Akha E, et al. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J Exp Med. 2011;208:455–67.
Man N, Tan Y, Sun XJ, Liu F, Cheng G, Greenblatt SM, et al. Caspase-3 controls AML1-ETO-driven leukemogenesis via autophagy modulation in a ULK1-dependent manner. Blood. 2017;129:2782–92.
Gong J, Gu H, Zhao L, Wang L, Liu P, Wang F, et al. Phosphorylation of ULK1 by AMPK is essential for mouse embryonic stem cell self-renewal and pluripotency. Cell Death Dis. 2018;9:38.
Iorio R, Celenza G, Petricca S. Mitophagy: molecular mechanisms, new concepts on parkin activation and the emerging role of AMPK/ULK1 axis. Cells. 2021;11:30.
Bellio C, DiGloria C, Spriggs DR, Foster R, Growdon WB, Rueda BR. The metabolic inhibitor CPI-613 negates treatment enrichment of ovarian cancer stem cells. Cancers. 2019;11:1678.
Guo P, Liu Y, Geng F, Daman AW, Liu X, Zhong L, et al. Histone variant H3.3 maintains adult haematopoietic stem cell homeostasis by enforcing chromatin adaptability. Nat Cell Biol. 2022;24:99–111.
Bagger FO, Kinalis S, Rapin N. BloodSpot: a database of healthy and malignant haematopoiesis updated with purified and single cell mRNA sequencing profiles. Nucleic Acids Res. 2019;47:D881–D5.
Nagel S, Pommerenke C, Scherr M, Meyer C, Kaufmann M, Battmer K, et al. NKL homeobox gene activities in hematopoietic stem cells, T-cell development and T-cell leukemia. PLoS One. 2017;12:e0171164.
Zhou F, Li X, Wang W, Zhu P, Zhou J, He W, et al. Tracing haematopoietic stem cell formation at single-cell resolution. Nature. 2016;533:487–92.
Pabst O, Forster R, Lipp M, Engel H, Arnold HH. NKX2.3 is required for MAdCAM-1 expression and homing of lymphocytes in spleen and mucosa-associated lymphoid tissue. EMBO J. 2000;19:2015–23.
Dinh TT, Xiang M, Rajaraman A, Wang Y, Salazar N, Zhu Y, et al. An NKX-COUP-TFII morphogenetic code directs mucosal endothelial addressin expression. Nat Commun. 2022;13:7448.
Singh SK, Singh S, Gadomski S, Sun L, Pfannenstein A, Magidson V, et al. Id1 ablation protects hematopoietic stem cells from stress-induced exhaustion and aging. Cell Stem Cell. 2018;23:252–65.e8.
Kataoka K, Sato T, Yoshimi A, Goyama S, Tsuruta T, Kobayashi H, et al. Evi1 is essential for hematopoietic stem cell self-renewal, and its expression marks hematopoietic cells with long-term multilineage repopulating activity. J Exp Med. 2011;208:2403–16.
Kocabas F, Zheng J, Thet S, Copeland NG, Jenkins NA, DeBerardinis RJ, et al. Meis1 regulates the metabolic phenotype and oxidant defense of hematopoietic stem cells. Blood. 2012;120:4963–72.
Ficara F, Murphy MJ, Lin M, Cleary ML. Pbx1 regulates self-renewal of long-term hematopoietic stem cells by maintaining their quiescence. Cell Stem Cell. 2008;2:484–96.
Freire PR, Conneely OM. NR4A1 and NR4A3 restrict HSC proliferation via reciprocal regulation of C/EBPalpha and inflammatory signaling. Blood. 2018;131:1081–93.
Min IM, Pietramaggiori G, Kim FS, Passegue E, Stevenson KE, Wagers AJ. The transcription factor EGR1 controls both the proliferation and localization of hematopoietic stem cells. Cell Stem Cell. 2008;2:380–91.
Lehnertz B, Chagraoui J, MacRae T, Tomellini E, Corneau S, Mayotte N, et al. HLF expression defines the human hematopoietic stem cell state. Blood. 2021;138:2642–54.
Li C, Wu B, Li Y, Liu Y, Wang J, Xie J, et al. Loss of sphingosine kinase 2 promotes the expansion of hematopoietic stem cells by improving their metabolic fitness. Blood. 2022;140:1686–701.
Garcia-Prat L, Kaufmann KB, Schneiter F, Voisin V, Murison A, Chen J, et al. TFEB-mediated endolysosomal activity controls human hematopoietic stem cell fate. Cell Stem Cell. 2021;28:1838–50.e10.
Warr MR, Binnewies M, Flach J, Reynaud D, Garg T, Malhotra R, et al. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature. 2013;494:323–7.
Palikaras K, Lionaki E, Tavernarakis N. Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nat Cell Biol. 2018;20:1013–22.
Ito K, Turcotte R, Cui J, Zimmerman SE, Pinho S, Mizoguchi T, et al. Self-renewal of a purified Tie2+ hematopoietic stem cell population relies on mitochondrial clearance. Science. 2016;354:1156–60.
Murakami K, Kurotaki D, Kawase W, Soma S, Fukuchi Y, Kunimoto H, et al. OGT regulates hematopoietic stem cell maintenance via PINK1-dependent mitophagy. Cell Rep. 2021;34:108579.
Lee EJ, Tournier C. The requirement of uncoordinated 51-like kinase 1 (ULK1) and ULK2 in the regulation of autophagy. Autophagy. 2011;7:689–95.
Nah J, Shirakabe A, Mukai R, Zhai P, Sung EA, Ivessa A, et al. Ulk1-dependent alternative mitophagy plays a protective role during pressure overload in the heart. Cardiovasc Res. 2022;118:2638–51.
Dhingra R, Rabinovich-Nikitin I, Kirshenbaum LA. Ulk1/Rab9-mediated alternative mitophagy confers cardioprotection during energy stress. J Clin Invest. 2019;129:509–12.
Kundu M, Lindsten T, Yang CY, Wu J, Zhao F, Zhang J, et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood. 2008;112:1493–502.
Lechauve C, Keith J, Khandros E, Fowler S, Mayberry K, Freiwan A, et al. The autophagy-activating kinase ULK1 mediates clearance of free alpha-globin in beta-thalassemia. Sci Transl Med. 2019;11:eaav4881.
Han X, Yoshizaki K, Miyazaki K, Arai C, Funada K, Yuta T, et al. The transcription factor NKX2-3 mediates p21 expression and ectodysplasin-A signaling in the enamel knot for cusp formation in tooth development. J Biol Chem. 2018;293:14572–84.
Acknowledgements
We thank Yang Liu for technical support in flow cytometry, Liting Wang for technical support in immunofluorescence microscopy, and Quanfang Wei for technical assistance in transmission electron microscopy. This work was supported by grants from the National Natural Science Foundation of China (No. 81930090, 82203974, 81725019), Project funded by China Postdoctoral Science Foundation (No. 2022M723867), Postdoctoral Innovative Talent Support Program of China (No. BX20220398), Postdoctoral Innovative Talent Support Program of Chongqing (No. CQBX2021017) and the Scientific Research Project of PLA (No. BWS21J022).
Author information
Authors and Affiliations
Contributions
MH, NC and MC designed the study, performed experiments, analyzed data and wrote the paper. FC, YL and YX performed experiments and analyzed data. LY and HZ participated in some animal experiments. MS and XC participated in some in vitro experiments. SC and FW participated in the initial experimental design and discussed the manuscript. JW and SW conceived and supervised the study, and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, M., Chen, N., Chen, M. et al. Transcription factor Nkx2-3 maintains the self-renewal of hematopoietic stem cells by regulating mitophagy. Leukemia 37, 1361–1374 (2023). https://doi.org/10.1038/s41375-023-01907-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-01907-y
- Springer Nature Limited
This article is cited by
-
Trim47 prevents hematopoietic stem cell exhaustion during stress by regulating MAVS-mediated innate immune pathway
Nature Communications (2024)