Abstract
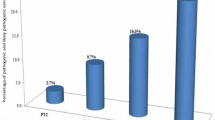
Genetic predisposition (familial risk) in the myeloproliferative neoplasms (MPNs) is more common than the risk observed in most other cancers, including breast, prostate, and colon. Up to 10% of MPNs are considered to be familial. Recent genome-wide association studies have identified genomic loci associated with an MPN diagnosis. However, the identification of variants with functional contributions to the development of MPN remains limited. In this study, we have included 630 MPN patients and whole genome sequencing was performed in 64 individuals with familial MPN to uncover recurrent germline predisposition variants. Both targeted and unbiased filtering of single nucleotide variants (SNVs) was performed, with a comparison to 218 individuals with MPN unselected for familial status. This approach identified an ATM L2307F SNV occurring in nearly 8% of individuals with familial MPN. Structural protein modeling of this variant suggested stabilization of inactive ATM dimer, and alteration of the endogenous ATM locus in a human myeloid cell line resulted in decreased phosphorylation of the downstream tumor suppressor CHEK2. These results implicate ATM, and the DNA-damage response pathway, in predisposition to MPN.
Similar content being viewed by others
Introduction
The myeloproliferative neoplasms (MPN) are characterized by increased production of mature myeloid cells due to a somatic driver mutation in either the JAK2, CALR, or MPL gene. While the majority of cases are sporadic, familial MPN is relatively common. As much as 10% of MPN cases display familial clustering, and there is a 5 to 7-fold increased risk of developing an MPN among relatives of MPN patients [1,2,3]. Identification of predisposing germline mutations in myeloid malignancies such as myelodysplastic syndrome (MDS), acute myeloid leukemia (AML), and bone marrow failure has led to a better understanding of the pathophysiology of these diseases as well as improvements in diagnosis and clinical care [4, 5]. Similar to these related myeloid diseases, familial MPN tends to display an autosomal dominant inheritance pattern with incomplete penetrance [6]. Investigation of large pedigrees with familial clustering of MPN has identified few high-risk alleles. For example, germline variants in the gene RBBP6 were found to co-segregate with the disease in three pedigrees with familial MPN with incomplete penetrance [7]. In addition, a germline duplication on chromosome 14q32 has been linked to predisposition to MPN as well as other myeloid malignancies in multiple families, though the causative gene(s) remains unclear [8, 9]. Thus, despite being relatively common, germline factors that predispose to MPN remain elusive. This has led to the hypothesis that more common, low-risk alleles account for a significant portion of the inherited risk in MPN. For example, the JAK2 46/1 haplotype is present in approximately 24% of the general population and is associated with a ~3-fold increased risk of acquiring a somatic JAK2 V617F mutation [10,11,12]. More recently, additional common germline single nucleotide variants (SNVs) that are associated with increased susceptibility for MPN have been identified via genome-wide association studies (GWAS), including low-risk alleles of TERT and TET2 [13,14,15,16]. However, the functional significance of the variants, as well as their relevance to individual patients, remains unclear. Interestingly, these same loci are also known to be associated with an increased risk for the development of clonal hematopoiesis of indeterminate potential (CHIP) [17]. Multiple studies have identified the presence of CHIP in asymptomatic individuals to be associated with an increased risk of hematologic malignancy, suggesting potential common pathophysiology in germline predisposition [18,19,20].
The lack of insight into the contribution of germline variants to familial MPN limits our ability to risk stratify MPN progression and to understand whether an MPN in an individual or a family signifies a larger cancer predisposition syndrome. To identify predisposition variants in MPN with potential functional significance, we leveraged a well-characterized cohort of MPN patients [21, 22]. Whole genome sequencing was performed on 64 unrelated patients with familial MPN and analyzed for recurrent germline variants. Both an unbiased filtering approach as well as a targeted survey of known cancer predisposition genes identified a recurrent ATM L2307F variant present in ~8% of this cohort. In silico modeling suggests this variant affects the conformational dynamics of the ATM dimer, and in vitro analysis revealed decreased phosphorylation of CHEK2, a downstream effector of ATM signaling. Together, these data implicate a germline variant in ATM as a direct mediator of germline predisposition in MPN and more broadly highlight a potential role of the germline defects in the DNA damage response (DDR) pathway in myeloid malignancies.
Materials and methods
Cohort description
The study was approved by the Johns Hopkins University institutional review board (IRB) and complied with all required ethical standards. Study participants with familial MPNs were identified from a cohort of individuals prospectively enrolled in the Johns Hopkins Center for Chronic MPN Disease Registry from 1999 to 2017 [21]. Familial MPN was defined as the diagnosis of MPN in a first- or second-degree family member of a patient with an established MPN diagnosis. The unselected cohort comprised all individuals referred for evaluation at Johns Hopkins Hospital from 2015 to 2021 who were found to have a diagnosis of MPN and underwent diagnostic next-generation sequencing (NGS) as part of their clinical evaluation. The diagnoses of MPN were based on the 2016 World Health Organization criteria [23].
Sequencing and variant calling
DNA samples for whole genome sequencing (WGS) were prepared from peripheral blood. Sequencing libraries were generated from 1 μg of sheered DNA using the Illumina TruSeq DNA PCR-Free HT Library Preparation Kit (Illumina, San Diego, CA, USA) before sequencing on an Illumina HiSeq X System (Supplementary Methods).
For germline variant calling, the GATK pipeline workflow was applied following best practices (https://software.broadinstitute.org/gatk/best-practices). FASTQ files were aligned to the human GRCh38.p12 reference genome using BWA-mem (v0.7.17). Per-patient genotyping was carried out with HaplotypeCaller and aggregated in a database for subsequent joint genotyping of the cohort. The annotation and effect prediction of the variants was performed using SnpSift/SnpEff, SIFT, Polyphen-2, and CADD (Supplementary Methods and Supplementary Fig. 1). To determine the presence of germline variants in known cancer predisposition genes we limited our analysis to 140 genes (Supplementary Table 1). The following filtering criteria were used: (1) variants reported in COSMIC v95 hg38 predicted as “pathogenic” by FATHMM, (2) variants with VAF 40–60% of >90% were included as likely germline (3) “pathogenic”, “likely pathogenic” or recurrent variants of unknown significance (VUS) as determined by VarSome automated classification based on the American College of Medical Genetics and Genomics (ACMG) guidelines were selected [24]. In the gene-agnostic approach, the variants were filtered as described in supplementary methods (Supplementary Fig. 1). Somatic variant calling for the familial cohort was performed with Mutec2 [25] and mutational signatures were compared to the COSMIC mutational signatures [26] (Supplementary Methods). Somatic analysis of the extended cohort was performed using Johns Hopkins clinical leukemia mutation panel that included 59 genes frequently mutated in hematologic malignancies (Supplementary Methods and Supplementary Table 2). Somatic structural chromosomal aberrations including copy-neutral loss of heterozygosity (CN-LOH) were assessed using VCF2CNA-Beta and Manta and annotated using AnnotSV [27,28,29].
In silico structural visualization and analysis
All structural models were analyzed using PyMOL software [30]. ATM monomer structure was predicted using ColabFold2: AlphaFold2 using MMseqs2 [31, 32] and I-TASSER: Iterative Threading ASSEmbly Refinement [33, 34].
ATM dimerization energetics
The closed ATM dimer model, 5np0 structure file [35], was retrieved from PDB [36]. The 5np0.pdb file was edited to retain only amino acids from 1995 to 3056 of ATM. Both monomers were retained in the structural file. ATM L2307F variant was introduced using the “Mutagenesis” wizard in PyMOL [30]. The ATM dimer binding energetics were predicted using SSIPe: binding affinity change estimation [37, 38] and ROSIE: The Rosetta Online Server that Includes Everyone [39, 40].
Cell line
TF1 cells were used to generate human ATM2307F mutated cell line via homology-directed repair (HDR) using the Alt-RTM CRISPR-Cas9 system with HDR donor oligo template (Integrated DNA Technologies, IDT, Coralville, IA, USA). TF1 cells were authenticated by short tandem repeat (STR) profiling and tested for Mycoplasma contamination. Single guide RNA (sgRNA) and HDR donor oligo template for ATM2307F were designed using the IDT Alt-RTM HDR design tool. The CRISPR-Cas9 ribonucleoprotein (RNP) complex and HDR donor oligo targeting ATM 2307 region were co-delivered into cells using the 4D-nucleofector system (Lonza, Bend, OR, USA). The edit location is GRCh38 chr11-108326169 and the transcript accession is NM_000051. To introduce the phenylalanine variant at ATM 2307 and a HinP1I restriction site (GCGC) for editing screening, the oligo template contained TTTGCG (Phe-Ala) in place of CTTGCC (Leu-Ala). The full length of HDR oligo template was 5′ GGAAGAAGCACAAGTATTCTGGGCAAAAAAGGAGCAGAGTTTTGCGCTGAGTATTCTCAAGCAAATGATCAAGAAGTTGGATGCCAGCTG 3′. Control cells (WT) were transfected without the HDR donor oligo template. After nucleofection, cells were cultured in media containing HDR Enhancer for 18 h and further cultured in fresh media without HDR Enhancer for 7 days for editing screening.
Screening and genotyping
To screen and isolate ATM2307F mutated cell clones, single-cell sorting was performed and clones were expanded in growth media. Genomic DNA isolation of individual clones was done following the manufacturer’s protocol (NEB, DNA isolation kit, Monarch, Ipswich, MA, USA) before PCR in batches of 10 using primers flanking the genomic region around the editing site (ATM_2307F_F: 5′-CAGACAGACAGATAGGCAGACGTGG -3′ and ATM_2307F_R: 5′- GAGTAGCTATTTCATTCATTTCCCACATACCTTTCTTG-3′). The purified PCR fragment from batched clones was digested using HinP1I (NEB), with correct digestion triggering genotyping of individual clones to identify properly targeted clones. Sanger sequencing was performed to confirm heterozygous versus homozygous targeting.
RT PCR and Western blotting
The expression of the targeted ATM was confirmed by both RT-PCR and Western blot. Cells were washed twice in PBS and protein lysates were isolated using RIPA buffer (Sigma, Burlington, MA, USA) containing protein and phosphatase inhibitors. Lysates were quantitated using BCA Protein Assay (Pierce, Thermo Fisher Scientific, Halethorpe, MD, USA) and 10 ug total lysate was run on NuPage TrisAcetate gels (Invitrogen). Transfers were performed using the iBlot System (Invitrogen), and membranes were probed overnight at 4 °C with primary antibodies: ATM (D2E2), CHEK2 (D9C6) (Cell Signaling Technology, Beverly, MA, USA), phospho-ATM, pospho-CHEK2 and anti-rabbit IgG, HRP-linked secondary antibody (DNA damage kit, Cell Signaling Technology). Antibody against GAPDH (D16H11) (Cell Signaling Technology) was used as a loading control. The visualization was performed using a chemiluminescent substrate (SuperSignal™ West Femto Maximum Sensitivity Substrate, Thermo Scientific).
Statistical analysis
Statistical analysis was performed using R 4.0.1. Categorical variables were represented as absolute numbers (percent). Contingency tables were assessed using Fisher’s exact test. The normality of the distribution was assessed using Shapiro-Wilk’s test, histogram visualization, and skewness and kurtosis assessment. Groups presenting a normal distribution were represented using mean + / − standard deviation and the differences between the two groups were assessed using Welch’s t-test. For the statistical analysis, a p-value under 0.05 was considered statistically significant.
Results
Clinical and genetic characteristics of familial MPN
The cohort of 630 patients diagnosed with MPN prospectively enrolled in the Johns Hopkins MPN registry has been previously described [21, 22]. Individuals with familial MPN were defined by the diagnosis of MPN or related myeloid malignancy (MDS, chronic myeloid leukemia, or AML) in a first- or second-degree relative. Using these criteria, 64/630 (10.2%) patients were classified as familial MPN. These included 37 (58%) individuals diagnosed with essential thrombocytosis (ET), 21 (33%) with polycythemia vera (PV), and 6 (9%) with primary myelofibrosis (PMF) (Table 1). Out of the 64 cases, 46 (72%) had a 1st degree relative with MPN, 18 (28%) had a 2nd degree relative with MPN. There were 11 cases with multiple affected relatives (5 with multiple 1st degree and 6 with 1st and 2nd degree relatives). In addition to a myeloid neoplasm, 48/64 (76.2%) patients had at least one relative with a solid tumor diagnosis. Breast, prostate, and lung cancers were the most common malignancies.
The analysis of somatic mutations revealed that JAK2 was the most commonly affected gene found in 75% (48/64) followed by CALR in 9% (6/64) and MPL in 3% (2/64). Among non-canonical mutations, TET2 was the most commonly mutated gene in the familial MPN cohort (Fig. 1A, Supplementary Table 3).
Distribution of somatic mutations and MPN phenotype in the familial (A) and unselected (B) cohorts. The right side of the graph displays individual patients, while the left side displays aggregates for each cohort. Genes are grouped by functional category. The mean age at diagnosis (+/− standard deviation) of the familial and unselected cohort is shown (C), as well as disease phenotypes (D) and relevant canonical somatic mutations and mutation categories (E).
To delineate the phenotypic and molecular aspects of familial MPN, a separate cohort of 218 consecutive patients who underwent somatic mutation profiling during their standard clinical evaluation for MPN was used for comparison (unselected cohort). The mean age at MPN diagnosis was significantly lower in familial vs. unselected cases (mean age 49.7 years vs. 54.9 years, respectively, p = 0.03; Table 1, Fig. 1C). Compared to unselected cohort, individuals with familial MPN were predominately females (41/64; 64% vs. 105/218; 48%, p = 0.03) and less likely to present as PMF (6/64; 9% vs. 50/218; 23%, p = 0.02; Fig. 1A, B, D, Table 1). Driver mutation status in both cohorts followed the pattern observed in the general MPN population with JAK2 V617F being the most common lesion, however, CALR mutations were underrepresented in the familial MPN cohort (6/64; 9.4% vs. 50/218; 22.9%, p = 0.02; Fig. 1E). Interestingly, mutations in spliceosome genes, even though frequently present in the unselected MPN cohort (6.9%), were not observed in the familial cohort (Fig. 1A, B, E, Supplementary Table 4).
Germline predisposition variants in familial MPN
Whole-genome sequencing of the 64 individuals with familial MPN was performed to identify predisposing germline variants. Sequencing data analysis was performed via two distinct approaches. First, a targeted approach was utilized focusing on 140 genes known to be involved in hereditary cancer predisposition syndromes (Supplementary Table 1). All non-coding variants were filtered out and heterozygous and homozygous variants were characterized based on their VAF (40–60% and >90%, respectively). Automated classification of germline variants via VarSome was utilized to isolate only germline variants that were deemed “pathogenic”, “likely pathogenic”, or recurrent variants of unclear significance (VUS) based on ACMG guidelines; all benign or likely benign variants were excluded. Targeted analysis of the familial MPN cohort produced 5 possibly pathogenic and/or recurrent variants in 12 individuals (Table 2). Only 1 “likely pathogenic” variant was identified- JAK2 T875N (chr9:5089726:C > A). This patient was found to have extreme thrombocytosis at birth during investigation for a heart murmur and was diagnosed with “triple negative” MPN following testing with a clinical NGS panel that was negative for somatic mutations in JAK2, CALR, and MPL. Interestingly, the patient’s mother was diagnosed with triple negative MPN at age of 5 years, and sequencing of peripheral blood DNA revealed that she was a carrier of the identical JAK2 T875N variant (Supplementary Fig. 2). ATM L2307F was the most common VUS, and it was present in 5/64 (7.8%) patients (Table 2). Three additional recurrent variants including BLM 643H, RAD50 R763H, and MET E168D, were identified in 2/64 (3%) patients each and were mutually exclusive (Supplementary Table 5). JAK2 46/1 phenotype was present in 39/64 (61%) of familial MPN cases.
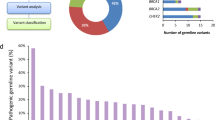
A second, unbiased, approach to the identification of predisposing variants involved the analysis of all coding variants. Via this approach, the germline variant calling pipeline identified 15,128,475 SNVs in all 64 individuals. The variant prioritization strategy limited the number of SNVs to 32,788 non-synonymous variants (Fig. 2A). Only 15 were found to occur at a significantly higher frequency than in a general population (reported in gnomAD) (Fisher Exact Test p ≤ 10−5). The in silico functional prediction revealed that only 4 variants in ATM, LLPH, IRAK3, and ZCCHC16 carried a negative functional impact (Fig. 2B, Supplementary Table 6). Of those, only ATM L2307F was designated as VUS by VarSome ACMG, while the other three were denoted either as “benign” or “likely benign”. Interestingly, the same ATM L2307F germline variant was identified in both the targeted gene screen and the gene-agnostic approach. Six variants were present in 5 individuals; patient 26 had biallelic variants due to interstitial copy-neutral loss of heterozygosity (CN-LOH) (Fig. 2C). This variant has a minor allele frequency of 0.000828 in the general population and only 2 homozygote individuals have been reported in gnomAD Genomes (both Ashkenazi Jewish). In addition to an MPN diagnosis in first-degree relatives, all 5 probands with the ATM L2307F variant had at least 1 family member with other malignancies that included lung, breast, pancreas, and non-Hodgkin lymphomas (Supplementary Table 7). The prevalence of ATM L2307F was significantly higher in familial compared to unselected MPNs (7.8% (5/64) vs 2.3% (5/218), p = 0.05). Interestingly, upon further examination, 60% (3/5) of ATM L2307F carriers in unselected MPN were diagnosed with other malignancies (2 breast and 1 renal cancer). Two individuals without a personal history of non-MPN malignancies were relatively young (17 and 19 years). Additionally, 60% (3/5) had a family history of cancer (Supplementary Table 7). Given the potential impact of ATM germline variants on structural chromosomal aberrations, we analyzed the effect of ATM L2307F on WGS-based karyotypes within the familial MPN cohort. The presence of the variant had no significant impact on structural or numerical chromosomal changes (Supplementary Fig. 3). Similarly, no significant effect was observed on mutational signature between MPN patients with and without ATM L2307 (Supplementary Fig. 4).
The filtering strategy of WGS data post variant calling for the unbiased (non-targeted) approach in the investigation of familial MPN (A). The number of variants remaining after each filtering step is provided. This strategy resulted in four recurrent variants implicated in predisposition to MPN. VarSome pathogenicity (ACMG criteria) prediction was utilized to further prioritize the candidate genes (B). Copy number alteration analysis 10 Mb upstream and 10 Mb downstream of the ATM gene in patients carrying ATM L2307F variant. Copy-neutral loss of heterozygosity (CN-LOH) was observed in Patient #26 (C).
In silico analysis of germline ATM L2307F variant
To determine the impact of L2307F variant on ATM structure, we applied ColabFold2 and I-TASSER web-based in silico prediction software (Supplementary Methods). Leucine at position 2307 falls within an alpha-helix structure and the substitution of phenylalanine for leucine did not appear to significantly change the conformation of the alpha-helix (Supplementary Fig. 5). These results were consistent between both prediction algorithms. Thus, it appeared that the L2307F variant had no significant structural impact on the ATM monomer. Since human ATM is in dynamic equilibrium between open (active) and closed (inactive) dimers, we next evaluated the impact of L2307F on homodimer function. In the closed ATM dimer L2307 on each monomer is localized in proximity to M2026 on the opposite monomer (Fig. 3A, B). Since the methionine-aromatic motif may stabilize protein structures, we assumed that the L2307F variant would introduce additional binding affinity with the opposite M2026 and stabilize ATM dimer in its closed conformation. In fact, ATM binding energetics predictions using SSIPe and ROSIE demonstrated meaningful free energy change indicating an increase in binding affinity as a result of the L2307F variant (Fig. 3C, Supplementary Table 8, and Supplementary Fig. 6). These in silico predictions suggest that even though the L2307F variant is not likely to affect ATM monomer protein structure, it may affect the homodimer binding favoring a closed (inactive) state.
ATM dimer structure in the closed conformation (A). The two ATM monomers are shown in green and yellow. The dashed box indicates the L2307 and M2026 interaction at one side of the interface. Enlargement of the L2307 and M2026 interaction between each monomer (B). Due to their proximity, a Leu to Phe alteration is likely to result in the stabilization of the ATM dimer in the closed conformation, which is confirmed by in silico binding and docking predictions (C).
In vitro effects of ATM L2307F variant
To further define the biological consequences of the ATM L2307F variant we established an in vitro model using the human myeloid TF1 cell line. The endogenous ATM locus was edited to introduce heterozygous (ATML2307F/+) and homozygous (ATML2307F/L2307) variants using the CRISPR-Cas9 system. Heterozygous and homozygous ATM L2307F variants were confirmed by single colony Sanger sequencing. While ATM expression in the heterozygous clone was similar to WT, the homozygous clone displayed decreased ATM mRNA levels likely secondary to a hemizygous deletion (Supplementary Fig. 7A, B). To confirm the in silico finding that L2307F affects ATM phosphorylation the impact of this variant on autophosphorylation of ATM at Serine 1981, which is phosphorylated in response to DNA damage, was examined. As expected, WT ATM S1981 was minimally phosphorylated at baseline and displayed a significant increase in response to ionizing radiation (Fig. 4A). The presence of ATML2307F/+ or ATML2307F/L2307 did not significantly alter S1981 phosphorylation at baseline or after ionizing radiation (IR) compared to wild-type ATM (Fig. 4A, B). To determine the impact of ATM L2307F on the ability of ATM to phosphorylate downstream mediators of the DDR pathway, expression, and phosphorylation of CHEK2 were examined. An increase in the inactive form of ATM would predict an inability to phosphorylate CHEK2, resulting in decreased degradation of total CHEK2 in response to ionizing radiation. In both the heterozygous and homozygous ATM mutant cells in the absence of ionizing radiation, a decrease in phosphorylated CHEK2 was observed with no effect on total CHEK2 expression compared to WT (Fig. 4). In the presence of ionizing radiation, total CHEK2 expression decreased in WT cells, however, this effect was impaired in ATM heterozygous and homozygous mutant cells. Further, the ratio of pCHEK2/CHEK2 was reduced in the ATM L2307F mutant cell lines. (Fig. 4A, B) Impaired phosphorylation of CHEK2 by mutant ATM was observed after IR-induced DNA damage at multiple time points. Altogether, the data indicate that ATM L2307F affects the phosphorylation of CHEK2 at baseline and after IR-induced DNA damage.
The protein level of ATM, phosphorylated ATM at S1981 (pATM S1981) and phosphorylated CHEK2 at T68 (pCHEK2 T68) are shown. The level of protein was assessed in the TF1 cell line with wild-type ATM allele (ATM+/+), heterozygous (ATML2307F/+), and homozygous (ATML2307F/L2307F) mutants at baseline and 1 h, 2 h, and 4 h after 5.2 Gy irradiation (A). The ratio of pATM/ATM and pCHEK2/CHEK2 are shown at baseline (ctrl) and after radiation (B). The level of each protein was quantified by densitometry and normalized with GAPDH.
Discussion
Investigation of a cohort of 630 MPN patients identified 64 unrelated individuals with familial MPN, consistent with the reported incidence of approximately 10% [1, 3]. Individuals with familial MPN were younger and more likely to be females than those in the unselected cohort. They were also less likely to have a diagnosis of PMF, consistent with the known associations of older age and male sex with this diagnosis [21, 41]. Analysis of somatic driver mutations was similar between both cohorts, however, mutations in spliceosome genes were more frequent in the unselected cohort, providing a potential etiology of the increased frequency of PMF observed in this cohort [42].
Eight individuals (12.5%) in the familial MPN cohort were identified as triple-negative MPN due to the lack of a canonical somatic driver mutation, with 5 of these individuals also lacking any somatic mutation in 59 additional genes associated with myeloid malignancies (Fig. 1, Supplementary Table 3). Whole genome sequencing led to the identification of a pathogenic germline JAK2 T875N mutation in one of these 5 individuals, altering the diagnosis to (non-clonal) hereditary thrombocytosis. The presence of the same germline mutation was confirmed in the patient’s mother who also carried a diagnosis of MPN. This variant was previously reported in one case of familial thrombocytosis and resulted in constitutive activation of JAK tyrosine kinase and downstream signaling [43, 44]. Given the substantial decrease in risk of thrombotic complications and exceedingly low potential for disease transformation in hereditary thrombocytosis compared to MPN, this change in diagnosis carries significant clinical implications. It also demonstrates the benefit of additional genetic testing beyond the canonical driver mutations in individuals with “triple negative” disease, particularly those with a family history of MPN.
To date, the investigation into germline predisposition of MPN via GWAS has identified multiple SNVs in linkage disequilibrium with a causal variant. However, variants exerting a direct effect on cancer predisposition have remained elusive. Our WGS analysis from 64 unrelated individuals with familial MPN identified five individuals carrying the ATM L2307F germline variant, and functional testing of this variant revealed a decrease in the ability of ATM to phosphorylate CHEK2. Distinct germline ATM VUS were identified in 2 additional individuals, as were VUS in other DDR pathway genes (Table 2 and Supplementary Table 5). Heterozygous germline variants in ATM are known to predispose to multiple cancer types including breast, prostate, pancreatic, and melanoma, and have been shown to modify disease phenotype [45,46,47]. While the role of germline ATM variants in hematologic malignancies is not well-established, SNVs in DDR genes such as ATM and CHEK2 have been found to be associated with an MPN and clonal hematopoiesis phenotype via GWAS [13, 16]. Most recently, a large GWAS study also identified a strong association between ATM, CHEK2, and PARP1 with the presence of clonal hematopoiesis [48]. In addition, germline ATM variants have been implicated in predisposition to both acute myeloid leukemia and chronic lymphoid leukemia [49,50,51]. At least one additional study identified a germline ATM variant, in addition to other DDR pathway genes, in patients with familial MPN [52].
The role of the DDR pathway in MPN is still undefined, however, there is substantial evidence to suggest its importance. The presence of the most common driver mutation in MPN, JAK2 V617F, resulted in an increase in genomic instability in a cell culture model, while primary cells from MPN patients were also shown to have abnormal DNA damage response [53,54,55]. Further, stem cells from MPN patients treated with PARP inhibitors showed reduced clonogenic growth in two independent studies [53, 56].
One potential explanation for the direct functional effect of ATM L2307F in MPN predisposition is an increase in the frequency of clonal hematopoiesis over time due to generalized DNA instability. We have demonstrated that ATM L2307F significantly affected the phosphorylation of its main downstream target CHEK2. Even though ATM/CHEK2 pathway plays a crucial role in the cellular response to DNA damage, particularly DNA double-strand breaks, we have not observed a higher frequency of structural and numerical chromosomal aberrations in ATM L2307F carriers. In addition to proper sensing of DNA double-strand breaks, ATM is involved in a plethora of cellular functions including telomere maintenance, RNA splicing, response to oxidative stress, and HSC self-renewal [57,58,59,60]. Functional studies in primary human HSCs demonstrated that CHEK2 inhibition increased the expansion of human cord blood HSC suggesting that impaired CHEK2 phosphorylation may promote HSC self-renewal and thereby increase the risk of MPN [16]. Analysis of larger cohorts of familial MPN patients as well as further study of ATM function in hematologic malignancies are necessary.
Data availability
All data generated or analyzed during this study are either included in this published paper and its supplementary materials or will be made available upon request.
References
Landgren O, Goldin LR, Kristinsson SY, Helgadottir EA, Samuelsson J, Björkholm M. Increased risks of polycythemia vera, essential thrombocythemia, and myelofibrosis among 24,577 first-degree relatives of 11,039 patients with myeloproliferative neoplasms in Sweden. Blood. 2008;112:2199–204.
Rumi E, Passamonti F, Della Porta MG, Elena C, Arcaini L, Vanelli L, et al. Familial chronic myeloproliferative disorders: clinical phenotype and evidence of disease anticipation. JCO. 2007;25:5630–5.
Sud A, Chattopadhyay S, Thomsen H, Sundquist K, Sundquist J, Houlston RS, et al. Familial risks of acute myeloid leukemia, myelodysplastic syndromes, and myeloproliferative neoplasms. Blood. 2018;132:973–6.
Godley LA. Germline mutations in MDS/AML predisposition disorders. Curr Opin Hematol. 2021;28:86–93.
Furutani E, Shimamura A. Genetic predisposition to MDS: diagnosis and management. Hematology. 2019;2019:110–9.
Jones AV, Cross NCP. Inherited predisposition to myeloproliferative neoplasms. Therapeutic Adv Hematol. 2013;4:237–53.
Harutyunyan AS, Giambruno R, Krendl C, Stukalov A, Klampfl T, Berg T, et al. Germline RBBP6 mutations in familial myeloproliferative neoplasms. Blood. 2016;127:362–5.
Babushok DV, Stanley NL, Morrissette JJD, Lieberman DB, Olson TS, Chou ST, et al. Germline duplication of ATG2B and GSKIP genes is not required for the familial myeloid malignancy syndrome associated with the duplication of chromosome 14q32. Leukemia. 2018;32:2720–3.
Saliba J, Saint-Martin C, Di Stefano A, Lenglet G, Marty C, Keren B, et al. Germline duplication of ATG2B and GSKIP predisposes to familial myeloid malignancies. Nat Genet. 2015;47:1131–40.
Jones AV, Chase A, Silver RT, Oscier D, Zoi K, Wang YL, et al. JAK2 haplotype is a major risk factor for the development of myeloproliferative neoplasms. Nat Genet. 2009;41:446–9.
Kilpivaara O, Mukherjee S, Schram AM, Wadleigh M, Mullally A, Ebert BL, et al. A germline JAK2 SNP is associated with predisposition to the development of JAK2V617F-positive myeloproliferative neoplasms. Nat Genet. 2009;41:455–9.
Olcaydu D, Harutyunyan A, Jäger R, Berg T, Gisslinger B, Pabinger I, et al. A common JAK2 haplotype confers susceptibility to myeloproliferative neoplasms. Nat Genet. 2009;41:450–4.
Hinds DA, Barnholt KE, Mesa RA, Kiefer AK, Do CB, Eriksson N, et al. Germ line variants predispose to both JAK2 V617F clonal hematopoiesis and myeloproliferative neoplasms. Blood. 2016;128:1121–8.
Oddsson A, Kristinsson SY, Helgason H, Gudbjartsson DF, Masson G, Sigurdsson A, et al. The germline sequence variant rs2736100_C in TERT associates with myeloproliferative neoplasms. Leukemia. 2014;28:1371–4.
Tapper W, Jones AV, Kralovics R, Harutyunyan AS, Zoi K, Leung W, et al. Genetic variation at MECOM, TERT, JAK2 and HBS1L-MYB predisposes to myeloproliferative neoplasms. Nat Commun. 2015;6:6691.
Bao EL, Nandakumar SK, Liao X, Bick AG, Karjalainen J, Tabaka M, et al. Inherited myeloproliferative neoplasm risk affects haematopoietic stem cells. Nature. 2020;586:769–75.
Bick AG, Weinstock JS, Nandakumar SK, Fulco CP, Bao EL, Zekavat SM, et al. Inherited causes of clonal haematopoiesis in 97,691 whole genomes. Nature. 2020;586:763–8.
Jaiswal S, Fontanillas P, Flannick J, Manning A, Grauman PV, Mar BG, et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl J Med. 2014;371:2488–98.
Xie M, Lu C, Wang J, McLellan MD, Johnson KJ, Wendl MC, et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med. 2014;20:1472–8.
Genovese G, Kähler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl J Med. 2014;371:2477–87.
Karantanos T, Chaturvedi S, Braunstein EM, Spivak J, Resar L, Karanika S, et al. Sex determines the presentation and outcomes in MPN and is related to sex-specific differences in the mutational burden. Blood Adv. 2020;4:2567–76.
Stein BL, Saraf S, Sobol U, Halpern A, Shammo J, Rondelli D, et al. Age-related differences in disease characteristics and clinical outcomes in polycythemia vera. Leuk Lymphoma. 2013;54:1989–95.
Arber DA. The 2016 WHO classification of acute myeloid leukemia: What the practicing clinician needs to know. Semin Hematol. 2019;56:90–5.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Benjamin D, Sato T, Cibulskis K, Getz G, Stewart C, Lichtenstein L. Calling somatic SNVs and indels with mutect2. Bioinformatics. 2019. https://doi.org/10.1101/861054.
PCAWG Mutational Signatures Working Group, PCAWG Consortium, Alexandrov LB, Kim J, Haradhvala NJ, Huang MN, et al. The repertoire of mutational signatures in human cancer. Nature. 2020;578:94–101.
Putnam DK, Ma X, Rice SV, Liu Y, Newman S, Zhang J, et al. VCF2CNA: a tool for efficiently detecting copy-number alterations in VCF genotype data and tumor purity. Sci Rep. 2019;9:10357.
Chen X, Schulz-Trieglaff O, Shaw R, Barnes B, Schlesinger F, Källberg M, et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32:1220–2.
Geoffroy V, Herenger Y, Kress A, Stoetzel C, Piton A, Dollfus H, et al. AnnotSV: an integrated tool for structural variations annotation. Bioinformatics. 2018;34:3572–4.
Schrödinger L. The PyMOL molecular graphics system, Version 1.3r1.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Mirdita M, Schütze K, Moriwaki Y, Heo L, Ovchinnikov S, Steinegger M. ColabFold: making protein folding accessible to all. Nat Methods. 2022;19:679–82.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2015;12:7–8.
Roy A, Kucukural A, Zhang Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc. 2010;5:725–38.
Baretić D, Pollard HK, Fisher DI, Johnson CM, Santhanam B, Truman CM, et al. Structures of closed and open conformations of dimeric human ATM. Sci Adv. 2017;3:e1700933.
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, et al. The protein data bank. Nucleic Acids Res. 2000;28:235–42.
Huang X, Zheng W, Pearce R, Zhang Y. SSIPe: accurately estimating protein-protein binding affinity change upon mutations using evolutionary profiles in combination with an optimized physical energy function. Bioinformatics. 2020;36:2429–37.
Huang X, Pearce R, Zhang Y. EvoEF2: accurate and fast energy function for computational protein design. Bioinformatics. 2020;36:1135–42.
Lyskov S, Chou F-C, Conchúir SÓ, Der BS, Drew K, Kuroda D, et al. Serverification of molecular modeling applications: The Rosetta online server that includes everyone (ROSIE). PLoS ONE. 2013;8:e63906.
Chaudhury S, Berrondo M, Weitzner BD, Muthu P, Bergman H, Gray JJ. Benchmarking and analysis of protein docking performance in Rosetta v3.2. PLoS One. 2011;6:e22477.
Komrokji RS, Verstovsek S, Padron E, List AF. Advances in the management of myelofibrosis. Cancer Control. 2012;19:4–15.
Vainchenker W, Kralovics R. Genetic basis and molecular pathophysiology of classical myeloproliferative neoplasms. Blood. 2017;129:667–79.
Mercher T, Wernig G, Moore SA, Levine RL, Gu T-L, Fröhling S, et al. JAK2T875N is a novel activating mutation that results in myeloproliferative disease with features of megakaryoblastic leukemia in a murine bone marrow transplantation model. Blood. 2006;108:2770–9.
Yoshimitsu M, Hachiman M, Uchida Y, Arima N, Arai A, Kamada Y, et al. Essential thrombocytosis attributed to JAK2-T875N germline mutation. Int J Hematol. 2019;110:584–90.
Karlsson Q, Brook MN, Dadaev T, Wakerell S, Saunders EJ, Muir K, et al. Rare germline variants in ATM predispose to prostate cancer: a PRACTICAL Consortium Study. Eur Urol Oncol. 2021;4:570–9.
Shindo K, Yu J, Suenaga M, Fesharakizadeh S, Cho C, Macgregor-Das A, et al. Deleterious germline mutations in patients with apparently sporadic pancreatic adenocarcinoma. JCO. 2017;35:3382–90.
Aoude LG, Bonazzi VF, Brosda S, Patel K, Koufariotis LT, Oey H, et al. Pathogenic germline variants are associated with poor survival in stage III/IV melanoma patients. Sci Rep. 2020;10:17687.
Kar SP, Quiros PM, Gu M, Jiang T, Mitchell J, Langdon R, et al. Genome-wide analyses of 200,453 individuals yield new insights into the causes and consequences of clonal hematopoiesis. Nat Genet. 2022;54:1155–66.
Yang F, Long N, Anekpuritanang T, Bottomly D, Savage JC, Lee T, et al. Identification and prioritization of myeloid malignancy germline variants in a large cohort of adult patients with AML. Blood. 2022;139:1208–21.
Tiao G, Improgo MR, Kasar S, Poh W, Kamburov A, Landau D-A, et al. Rare germline variants in ATM are associated with chronic lymphocytic leukemia. Leukemia. 2017;31:2244–7.
Samaraweera SE, Wang PPS, Li KL, Casolari DA, Feng J, Pinese M, et al. Childhood acute myeloid leukemia shows a high level of germline predisposition. Blood. 2021;138:2293–8.
Elbracht M, Meyer R, Kricheldorf K, Gezer D, Thomas E, Betz B, et al. Germline variants in DNA repair genes, including BRCA1/2, may cause familial myeloproliferative neoplasms. Blood Adv. 2021;5:3373–6.
Pratz KW, Koh BD, Patel AG, Flatten KS, Poh W, Herman JG, et al. Poly (ADP-Ribose) polymerase inhibitor hypersensitivity in aggressive myeloproliferative neoplasms. Clin Cancer Res. 2016;22:3894–902.
Plo I, Nakatake M, Malivert L, de Villartay J-P, Giraudier S, Villeval J-L, et al. JAK2 stimulates homologous recombination and genetic instability: potential implication in the heterogeneity of myeloproliferative disorders. Blood. 2008;112:1402–12.
Chen E, Ahn JS, Sykes DB, Breyfogle LJ, Godfrey AL, Nangalia J, et al. RECQL5 suppresses oncogenic JAK2-induced replication stress and genomic instability. Cell Rep. 2015;13:2345–52.
Patel PR, Senyuk V, Rodriguez NS, Oh AL, Bonetti E, Mahmud D, et al. Synergistic cytotoxic effect of busulfan and the PARP inhibitor veliparib in myeloproliferative neoplasms. Biol Blood Marrow Transplant. 2019;25:855–60.
Tong AS, Stern JL, Sfeir A, Kartawinata M, de Lange T, Zhu X-D, et al. ATM and ATR signaling regulate the recruitment of human telomerase to telomeres. Cell Rep. 2015;13:1633–46.
Tresini M, Warmerdam DO, Kolovos P, Snijder L, Vrouwe MG, Demmers JAA, et al. The core spliceosome as target and effector of non-canonical ATM signalling. Nature. 2015;523:53–8.
Lee J-H, Mand MR, Kao C-H, Zhou Y, Ryu SW, Richards AL, et al. ATM directs DNA damage responses and proteostasis via genetically separable pathways. Sci Signal. 2018;11:eaan5598.
Ito K, Hirao A, Arai F, Matsuoka S, Takubo K, Hamaguchi I, et al. Regulation of oxidative stress by ATM is required for self-renewal of haematopoietic stem cells. Nature. 2004;431:997–1002.
Acknowledgements
This work was supported by the National Institutes of Health (NIH), Heart, Lung and Blood Institute grant K08HL138142 (EMB), K08HL136894 (LPG), R01HL156144 (LPG), and in part by the Intramural Research Program of the National Heart, Lung, and Blood Institute of the National Institutes of Health (JG and CSH).
Author information
Authors and Affiliations
Contributions
AM, EMB and LPG identified and prepared biospecimens, and collected and annotated clinical data; CA, DH, MW, CLD, JG performed whole genome sequencing; EI and LM processed, analyzed, and interpreted WGS data; SP analyzed and interpreted clinical and sequencing data, performed genomic analysis, wrote and edited the manuscript; SW performed cell culture experiments and interpreted results; HC analyzed and interpreted sequencing data and performed protein modeling experiments; YL designed and performed protein modeling experiments; EMB and LPG supervised the experiments, interpreted the data, wrote and revised the manuscript; EMB, LPG, CH designed and funded the study. All authors critically reviewed the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Braunstein, E.M., Imada, E., Pasca, S. et al. Recurrent germline variant in ATM associated with familial myeloproliferative neoplasms. Leukemia 37, 627–635 (2023). https://doi.org/10.1038/s41375-022-01797-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-022-01797-6
- Springer Nature Limited