Abstract
Despite the success of protein kinase inhibitors as approved therapeutics, drug discovery has focused on a small subset of kinase targets. Here we provide a thorough characterization of the Published Kinase Inhibitor Set (PKIS), a set of 367 small-molecule ATP-competitive kinase inhibitors that was recently made freely available with the aim of expanding research in this field and as an experiment in open-source target validation. We screen the set in activity assays with 224 recombinant kinases and 24 G protein–coupled receptors and in cellular assays of cancer cell proliferation and angiogenesis. We identify chemical starting points for designing new chemical probes of orphan kinases and illustrate the utility of these leads by developing a selective inhibitor for the previously untargeted kinases LOK and SLK. Our cellular screens reveal compounds that modulate cancer cell growth and angiogenesis in vitro. These reagents and associated data illustrate an efficient way forward to increasing understanding of the historically untargeted kinome.
Similar content being viewed by others
Main
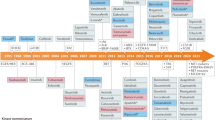
Protein kinases have been a rich source of targets for drug discovery, with more than 25 therapeutics approved by the US Food and Drug Administration (FDA) and hundreds more in clinical evaluation1,2,3. Most academic and industrial research remains focused on only a small fraction of the kinome for which evidence of therapeutic utility already exists4. This has led to a drug-development pipeline with much overlap between pharmaceutical companies and many redundant clinical studies5. To catalyze research on kinases that have received scant attention to date4, GlaxoSmithKline defined PKIS, a set of 367 kinase inhibitors that were previously published individually6.
Herein we provide a comprehensive characterization of PKIS compounds. We used target-based assays to determine inhibitor selectivity and to identify chemical starting points for previously untargeted kinases. To evaluate off-target activity outside the kinase family, we collected screening data against 24 G protein–coupled receptors (GPCRs), revealing frequent cross-reactivity of some kinase scaffolds with this diverse target family. We also found cytotoxic and cytostatic compounds in a screen of the NCI-60 cancer cell line collection, identified potential angiogenesis inhibitors in a high-content tube-formation screen and generated a selective inhibitor of the LOK and SLK kinases through structure-based modification of an inhibitor of the related kinase TIE2.
Results
Defining the set
GlaxoSmithKline and its legacy companies have published the structure and activity of >3,000 small-molecule inhibitors that cover a range of kinase targets and chemotypes. We selected compounds from this pool to form an open-source set. As an initial filter, we applied the practical requirement of a material supply of >5 mg of sample. Each compound was assigned to a specific chemotype, and in the case of heavily populated chemotypes, which have often been the focus of extensive efforts in analog synthesis, we conducted additional filtering to reduce the number of representatives to <40 while maximizing the diversity of activity profiles across kinases. Within a given chemotype, we sought to include compounds having a range of activities at the primary target and across other kinases. Compounds with less than 98% purity as measured by liquid chromatography–mass spectrometry (LC-MS) were excluded. At the end of the triage process, 31 chemotypes containing a total of 367 compounds from programs originally targeting 24 different kinases remained (chemical structures, identifying codes and literature references for each compound in the PKIS are provided in Supplementary Table 1, Supplementary Fig. 1 and Supplementary Data).
The majority of the PKIS compounds had physicochemical properties within the bounds of the commonly recognized drug-like space. Two-thirds of the set had molecular weight (MW) <500 Da and a log of the partition coefficient between octanol and water (clogP) <5. The set included some high-MW and high-clogP compounds, many of which were designed to target a conserved lipophilic 'back pocket' region. Known collectively as type II or DFG-out inhibitors because they induce a conformational change involving a loop containing conserved Asp-Phe-Gly (DFG) residues7, these inhibitors have proven useful for drug discovery despite the potential liabilities associated with their physicochemical properties.
In vitro characterization of the PKIS
Each PKIS member was previously published with a limited set of activity data. To improve annotation of the set as a whole, we screened the PKIS against a panel of 224 recombinant kinases, which included 196 unique members of the human protein kinome, 21 mutants associated with disease phenotypes and 5 lipid kinases (Fig. 1a, Supplementary Table 2 and Supplementary Data). The Nanosyn enzyme assay panel employed microfluidics capillary electrophoresis technology to measure the change in electrophoretic mobility of the substrate—usually a fluorescence-labeled peptide or lipid—upon phosphorylation. The simultaneous determination of substrate and product was employed to increase assay precision. The panel represented approximately 40% of the human protein kinome and was biased toward previously targeted kinases and close relatives. The data were collected as single measurements at two inhibitor concentrations (100 nM and 1 μM), with the concentration of ATP at the apparent Km of ATP of the particular kinase being profiled (Fig. 1b,c). Comparison of duplicate measurements for two compounds indicated that the data were concordant, affirming the utility of this screening approach (Supplementary Fig. 2).
(a) Phylogenetic tree of kinases screened against the PKIS by Caliper assay at Nanosyn, Inc. (blue circles) and by DSF (yellow circles). (b,c) Heat maps of Caliper assay screening at 0.1 μM (b) and 1 μM (c) compound. (d) Annotated heat map of Caliper assay screening at 0.1 μM compound (Supplementary Table 1). (e) Heat map of DSF screening of the PKIS against 68 kinases. Tm shifts of >4 °C are shown in yellow. The green arrow (a–e) marks the start and direction of the circular sorting order applied to the kinases sorted by circular distance around the phylogenetic tree and compounds sorted by chemotype. Inhibition from 0% to 100% is shown as a gradient of blue to yellow.
The PKIS was further screened against a set of 68 kinases by differential scanning fluorimetry (DSF), measuring the increase in melting temperature (ΔTm) of a kinase when exposed to a small-molecule inhibitor at 10 μM (ref. 8) (Fig. 1e and Supplementary Data). For the 32 kinases in common between the two panels, the correlations of assay data were r = 0.45 (Nanosyn panel screening at 0.1 μM) and r = 0.63 (Nanosyn panel screening at 1 μM) despite the difference in inhibitor concentration between assays. The degree of correlation between these two assay types might also be affected by differences in phosphorylation or activation state of the kinase. Nonetheless, the pattern of selective compounds and promiscuous chemotypes was preserved between the two panels (Fig. 1e).
Kinase screening results
Several broad trends were revealed by the screening data (Fig. 1). Visualization of the data set showed that the original kinase targets of each chemotype were often associated with the most significant cluster and that the tyrosine kinase (TK), CMGC and STE subfamilies were inhibited by a larger percentage of the inhibitors than were other subfamilies (Fig. 1d, Supplementary Table 3 and Supplementary Fig. 3). Although previous reports have associated increased inhibitor MW with promiscuity9, our data set did not show an evident relationship between MW and the number of kinases inhibited by ≥50% by a compound at either 100 nM or 1.0 μM. Indeed, the most promiscuous compounds were the relatively small pan-TK inhibitors GW559768X and GW440139A (Fig. 2a). Ten compounds inhibited more than 20 kinases by >50% at 100 nM. The original targets for these compounds were not localized in a single branch of the kinome but span the TK (GW559768X, targeting RET; GSK1173862A and GSK1326255A, targeting IGF-1R), the CMGC (GW779439X, targeting CDK) and the AGC (GSK1007102B, targeting Akt) subfamilies. Notably, the compounds showed patterns of activity that were distinct across the kinome, although each compound was associated with clusters of activity in the particular subfamily of its original kinase target (Supplementary Fig. 4). For comparison, the well-characterized nonspecific kinase inhibitor staurosporine, included as a control compound at 80 nM, inhibited 153 of the 224 kinases by >50%.
(a) Number of kinases inhibited by each compound at >50%, at 0.1 μM compound. The compounds and chemotypes are sorted as in Figure 1. Some of the most promiscuous inhibitors are depicted; selectivity profiles for these compounds are available in Supplementary Figure 3. (b) Protein kinome phylogenetic tree of PKIS original kinase targets and kinases with >50% inhibition at 0.1 μM by at least one compound. (c) Number of compounds that inhibit each kinase at >50%, at 0.1 μM compound. The kinases are sorted as in Figure 1d. (d) Selectivity profiles for example starting points toward chemical probes identified in the screening.
The profiling data also identified several compounds as highly selective across the kinase panel, including some that had been previously reported as chemical probes on the basis of fewer assays. For example, SB-686709-A and SB-698596-AC were originally reported as selective inhibitors of glycogen synthase kinase-3β (GSK3β) in a panel of 25 kinase assays10; after extensive profiling these compounds were confirmed as highly selective inhibitors of GSK3α and GSK3β (Supplementary Fig. 5). The data also verified the high selectivity of GW632580X for colony-stimulating factor-1 receptor (cFMS)11 and SB-590885-AAD for BRAF12 (Supplementary Table 4).
Only 16 compounds had a maximum inhibition of <20% at 1.0 μM across the assay panel. These structurally related inactive compounds can be useful in several contexts, such as in the interpretation of cellular profiling data and correlation with protein structural information13.
The Nanosyn panel data set allowed an assessment of the breadth of kinome activity coverage by the PKIS. Of the 224 kinase assays in the assay panel, 156 kinases were inhibited >50% by at least one compound at 100 nM (Fig. 2b), and 18 kinases were inhibited <50% at 1 μM by any compound.
Plotting the data of the unique protein kinases screened in the Nanosyn (excluding disease-related mutants) and DSF panels on a kinase phylogenetic tree showed extensive coverage with only a few small clusters of noninhibited kinases (Supplementary Fig. 6). A subset of 24 kinases were inhibited by single compounds with selectivity over other kinases (i.e., 24 kinases were inhibited by compounds having only a single kinase activity >50% at 100 nM).
This breadth of kinome coverage was observed despite the modest number of compounds in the set, the overrepresentation of inhibitors designed to target the TK and CMGC families and the composition of the assay panel that favors 'targeted' kinases. The most frequently inhibited kinases, as measured by the number of compounds with >50% inhibition at 1 μM, were the TKs LYN, PDGFRα, PDGFRβ and KDR (VEGFR2). From the more stringent data at 100 nM compound concentration, there were three CMGC kinases (GSK3α, GSK3β and p38α) and six TKs (EGFR, KDR, KIT, PDGFRα, PDGFRβ and RET) that were inhibited by more than 20 compounds (Fig. 2c).
Examples of analysis within chemotypes
A compound set composed of multiple compounds within each chemotype aids greatly in defining structure–activity relationships (SARs). Owing to space constraints it is not possible to discuss here all SARs that can be deduced from this large data set; however, we provide examples that will help to illustrate the value of this approach, both in terms of target-specific information and broader 'survey' analysis (Supplementary Figs. 7 and 8).
Example chemical probe starting points
The kinase inhibition data confirmed the utility of PKIS in the identification of chemical starting points for probes of historically understudied kinases. As an illustration of this utility, we found 89 kinases (73 wild type and 16 mutant) that were inhibited by >40% by at least one compound at 100 nM, where that compound inhibits no more than 10% of tested kinases by 50% at 1.0 μM and the kinase is in the top 5 hits for that compound (Supplementary Table 5). More than 50% of these kinases are historically understudied targets for which high-quality probes might have a substantial impact4.
Using the heat maps to compare the original target of each compound with the full range of kinase inhibition revealed numerous starting points for chemical probe development against previously untargeted kinases. We plotted a few example clusters (Figs. 1d, 2d and Supplementary Table 6). Examples of notable inhibition profiles were found in SB-739245-AC, which maintained its previously reported potency against GSK3α and GSK3β while also inhibiting cyclin-dependent kinase 6 (CDK6) and cyclin A and mitogen-activated protein kinase–interacting serine/threonine kinase 1 (MNK1); and GSK204925, from a series originally targeting polo-like kinase 1 (PLK1), which inhibited brain-specific serine/threonine kinases 1 (BRSK1) and BRSK2 and lymphocyte-oriented kinase (LOK, also known as STK10). The amino acid sequence of the two BRSK isozymes is highly conserved in the ATP binding site, and inhibitors showed no selectivity across the set (r = 0.95 at 0.1 μM; r = 0.96 at 1.0 μM, Supplementary Fig. 9). SB-633825, synthesized as a TIE2 inhibitor, showed potent inhibition of LOK and BRK (Fig. 2d) with a relatively clean inhibition profile. We follow up this observation below as an example of the development of a specific chemical probe for LOK and SLK.
Even weak hits showing inhibition only at higher concentrations may be useful starting points for further inhibitor optimization; 20% inhibition at 1 μM projects to a half-maximal inhibitory concentration (IC50) < 10 μM (Supplementary Table 7). We also observed some unexpected cross-reactivities; GSK317314A, a PLK1 inhibitor, also strongly inhibited NIMA-related kinase 9 (NEK9) at 0.1 μM (PLK1 98%, NEK9 77%) (Supplementary Table 8).
An example chemical starting point identified by DSF is SB-742864, which showed ΔTm = 7.1 °C for adapter protein 2–associated kinase 1 (AAK1). This compound had ΔTm ≥ 4 °C with only two other kinases in the panel (STK16 and CDK2, with ΔTm = 4.9 °C and 4.4 °C, respectively). Nanosyn kinase inhibition data showed that all kinases in that panel were inhibited <20% at 1 μM (except ABL T315I, at 31% inhibition). The affinity of SB-742864 for AAK1 was confirmed in a split luciferase assay system14, where it inhibited AAK1 activity with IC50 = 220 nM (data not shown). Several other members of the 3-aminoindazole series showed AAK1 ΔTm ≥ 4 °C while maintaining selectivity over the other kinases in the Nanosyn panel, increasing confidence in this series as a source of AAK1 inhibitors for optimization.
GPCR profiling
ATP-competitive kinase inhibitors can have cross-reactivity with targets beyond the kinome—for example, by interacting with the aminergic GPCRs15. Such cross-reactivity may lead to unexpected phenotypic responses but may also be used for the design of inhibitors outside the kinase family16,17. To provide additional target-based activity characterization, we profiled the PKIS against 24 GPCRs in calcium mobilization assays (Supplementary Table 9 and Supplementary Data).
No PKIS compound had any measurable agonist activity against these GPCRs. However, several compounds showed antagonist activity (Fig. 3a). We analyzed the GPCR screening results using ADDAGRA software, which represented the 24 GPCRs as nodes linked by edges18 (Fig. 3b). The presence of an edge connecting two receptor nodes indicates the similarity of the entire screening profile for all 367 compounds against the two (connected) GPCRs. The GPCRs P2Y11, V1B and, in particular, OT were clear outliers, whereas others (for example, V1A, M3D and P2Y2) clustered together.
(a) Heat map of responses with 24 clustered columns of GPCRs and 367 clustered PKIS compounds. Yellow indicates no effect; red indicates maximum potentiation effect; green indicates antagonist effect (complete inhibition of the effect of a control agonist). (b) ADDAGRA representation of GPCR antagonist profiles. (c) GPCR antagonist profile (1) and kinase inhibition profile (2) for the promiscuous compound GW406108X.
Some PKIS compounds (such as GW406108X) were characterized by complex and promiscuous GPCR activity profiles (Fig. 3c), whereas others showed highly selective profiles (Fig. 4). The concerted analysis of the kinase and GPCR screening data allowed us to select several compounds with unique profiles for confirmatory assays. For example, GSK238583A was found to be a potent epidermal growth factor receptor-4 (ERBB4) inhibitor (IC50 = 6.7 nM) but was inactive against any of the GPCRs. Another compound, GW682841X, had limited kinase inhibition potency but appeared to be a selective antagonist at P2Y receptors on the basis of the primary screening data. GW682841X was confirmed to have antagonistic activity at P2Y1, P2Y2 and P2Y11 in secondary screening assays. GW682841X reduced agonist efficacy in a concentration-dependent manner, suggesting that its interaction is based on a noncompetitive mechanism or is competitive because of hemi-equilibrium. The GW682841X antagonist activity was different at P2Y subtypes, as it reduced potency of ADP at the P2Y1 receptor and ATP at the P2Y2 receptor but enhanced ATP potency at the P2Y11 receptor. These results were consistent with an allosteric interaction with P2Y endogenous agonists. By contrast, its antagonist activity at P2Y4 and P2Y6 receptors was much weaker (data not shown).
Profiling in the NCI-60 cancer cell line panel
The NCI-60 panel of human cancer cell lines is a well-characterized platform for the cellular screening of potential anticancer agents19,20. A number of powerful online tools are available to enable data analysis, including COMPARE (https://dtp.nci.nih.gov/databases_tools/compare.htm) and CellMiner (http://discover.nci.nih.gov/cellminer/)21.
The effect on cell growth of each PKIS compound was evaluated in duplicate at five concentrations ranging from 2.5 nM to 25 μM, with tenfold dilutions, across the NCI-60 panel (Supplementary Data). Three data points were interpolated from the response data for each compound in each cell line: GI50, the concentration at which cell growth was inhibited by 50% over the 48-h time course of the assay; TGI, the concentration resulting in total cell growth inhibition; and LC50, the concentration at which 50% of the cells were killed.
The heat map of hierarchically clustered GI50 values showed a range of sensitivity and resistance for the PKIS compounds across the cell lines (Fig. 5a). The activity pattern across the panel for a given compound can provide insight into function, and the COMPARE algorithm draws on such patterns in the NCI-60 data to associate compounds with common mechanisms of action. Activity patterns of PKIS compounds that inhibit specific kinases in vitro are labeled on the heat map. In addition, a network diagram of pairwise matrix COMPARE correlation values (≥0.7) for 208 of 367 PKIS compounds that have a range of –log GI50 (also known as pGI50) values >1.0 allows for a visual representation of agents with related effects on the NCI-60 cell lines (Fig. 5b). For example, the GI50 cell-line pattern profiles of dual inhibitors of epidermal growth factor receptor (EGFR) and ERBB2 transmembrane protein kinases were, in general, highly correlated when we used the COMPARE algorithm, and one such compound (GW583373A) had a pattern highly similar to that of the FDA-approved BRAF inhibitor vemurafenib lapatinib, the FDA-approved selective dual inhibitor of EGFR and ERBB2 (r = 0.91). In another example, the activity patterns of the BRAF inhibitors also formed a tight cluster in the network diagram with the PKIS compound SB-590885 showing a high COMPARE correlation value (r = 0.91) with the structurally unrelated FDA-approved BRAF inhibitor vemurafenib.
(a) Hierarchical clustering (unweighted pair group method with arithmetic mean) heat map representation of the pGI50 NCI-60 data for PKIS compounds. (b) Network diagram of pairwise matrix COMPARE correlations for the 208 out of 367 PKIS compounds that have pGI50 values greater than 1.0. An edge connects two compounds if the pairwise correlation is greater than 0.70.
The concordance of the NCI-60 cell-line response patterns for several compounds with their known target-based profiles increased our confidence in the data set. However, the agents with cellular response patterns that are distinct from those of clinically approved, highly selective kinase inhibitors may be of more interest. One such compound is GW768505A. The GI50 values for this compound were in the approximate range of 300 nM to 3.0 μM for all cell lines except for the colon cancer-derived KM12 line, where GW768505A had GI50 = 6.3 nM. The kinase target profile of GW768505A, a compound originally developed to target KDR and TIE2 (ref. 22), showed potent inhibition (71–88% inhibition at 100 nM) of the tropomysin-related kinases TRKA, TRKB and TRKC. The KM12 cell line is associated with overexpression of TRKA and is sensitive to TRKA inhibition23. On the matrix COMPARE network diagram, the GI50 pattern for GW768505A was highly correlated (r > 0.7) with the patterns of two oxindole TRKA inhibitors (GW278681X and GW441756X). To demonstrate TRKA activity, we showed that both GW768505A and GW441756X inhibit TRKA autophosphorylation in KM12 cells (Supplementary Fig. 10), with GW441756X acting faster. GW768505A was significantly more potent against the human colon cancer cell line KM12 than was GW441756X, raising the possibility that an additional cellular target of GW768505A contributes to its potent GI50.
A second example resulted from running a standard COMPARE for the GI50 pattern derived from the KDR- and TIE2-targeting agent GW683134A24 against the NCI-60 compound collection data set. This analysis revealed a high correlation (r = 0.81) with the response pattern for the structurally unrelated multitargeted kinase inhibitor ponatinib.
Optimization of a dual chemical probe for LOK and SLK
SB-633825 profiled as a potent inhibitor of LOK in addition to its original target TIE2 and served as a starting point for generation of a selective inhibitor of the LOK and SLK kinases.
Twenty derivatives of SB-633825 were tested for activity against LOK, p38α and TIE2 (Fig. 6a). Two compounds retained inhibition of LOK while losing all activity against TIE2. GSK3037619A and SB-440719 showed high selectivity in the Nanosyn kinase assay panel (which had been expanded to 255 kinases assays compared with the original panel used for PKIS screening) with LOK and SLK as the top targets (Supplementary Table 10). LOK and SLK are close homologs and have a conserved ATP binding site (SLK was not in the earlier Nanosyn panel of 224 kinases). We co-crystallized SB-633825 with LOK and were able to obtain two X-ray structures in different crystal forms (Supplementary Table 11). The structures confirmed the ATP-competitive binding mode of SB-633825 and showed that in one crystal form, LOK was in the inactive 'DFG-out' conformation, and in the other LOK was in the active 'DFG-in' conformation (Fig. 6b). With SB-440719 it was possible to obtain a high-resolution (1.75-Å) co-crystal structure with SLK (Fig. 6c). The structure confirmed the binding mode of the inhibitor and revealed two notable features: (i) the glycine-rich loop (P-loop) formed an unusual conformation to allow Phe45 to bind the imidazole ring of the inhibitor and (ii) the activation loop packed against the inhibitor and the P-loop to reinforce this binding (Fig. 6d). The sequence of the glycine-rich loop (GELGDGAFGKVY) is well conserved only in the neurotrophic receptor tyrosine kinases 1–3, which were not inhibited by SB-633825.
(a) Identification of selective inhibitors of LOK with increased selectivity. Chemical structures of all compounds are available in the Supplementary Data. (b) Crystal structures of SB-633825 with LOK in inactive DFG-out state (left, PDB 4USD) and active DFG-in state (right, PDB 4USE). Green, N-terminal lobe; blue, C lobe; red, DFG motif. (c,d) Crystal structure of SB-440719 with SLK (PDB 4USE) (c) and showing the involvement of the activation loop (red spheres) in binding the inhibitor (yellow spheres) (d).
To validate the utility of the more potent SLK-LOK inhibitor, GSK3037619A, we first measured direct binding to LOK and SLK by biolayer interferometry. GSK3037619A bound to LOK and SLK with Kd values of 63 nM and 199 nM, respectively (Supplementary Fig. 11). These values were consistent with the enzymatic assay data. We also assessed the ability of GSK3037619A to inhibit LOK and SLK in cells. As LOK may be the dominant ezrin-radixin-moesin (ERM)-phosphorylating kinase in lymphocytes25,26,27, we assessed ERM phosphorylation in Jurkat T cells. Application of GSK3037619A had a clear effect on ERM phosphorylation in repeated experiments (Supplementary Fig. 11e).
High-content screen for regulators of angiogenesis
We also screened the PKIS for compounds targeting angiogenesis, as such targeting is a therapeutic strategy for the treatment of cancer and cardiovascular diseases28.
To characterize the activity of PKIS compounds on angiogenesis, a high-content screen was conducted using an endothelial tube-formation assay29 (Supplementary Data). This assay captured many cellular processes essential to the growing vascular network, such as establishment of new connections, branching and tubulogenesis.
The antiangiogenic activity of the PKIS compounds was correlated with the kinase inhibition data (Fig. 7a and Supplementary Fig. 12). Overall, 65 compounds from the PKIS significantly inhibited the formation of endothelial tubules (B score ≤1.5, 2× median absolute deviation (MAD)). Most of these compounds target multiple receptor tyrosine kinases such as PDGFRs, c-KIT, RET and VEGFRs, that are known to be involved in the initiation of angiogenesis. Antiangiogenic tyrosine kinase inhibitors such as sunitinib and sorafenib have been previously employed as a therapeutic strategy against cancer30,31.
(a) Kinase inhibition heat map (as in Fig. 1) with compounds sorted according to angiogenic activity (B score of total tubule length) obtained by an endothelial tube-formation assay on primary human umbilical vein endothelial cells. Red arrows mark the cutoff for angiogenesis inhibitors or enhancers (2× MAD). Kinases involved in angiogenesis are indicated on the x axis. (b) Most PKIS compounds showing >50% inhibition of KDR exhibited a significant reduction of total tubule length (angiogenesis), B score ≤1.5 (2× MAD). Orange circles represent compounds originally targeting TIE2 and KDR. (c) Left, kinase inhibition profile of GW786460X (original target TGF-βR1) showing >50% inhibition of KDR at 0.1 μM (green) and fluorescence images of endothelial capillary-like network formation produced with GW786460X (10 μM) versus DMSO (vehicle); right, kinase inhibition profile of GW795493X (original targets: TIE2 and KDR) showing inhibition of multiple receptor tyrosine kinases, including KDR, and fluorescence images showing endothelial capillary-like network structure produced with GW795493X (10 μM) versus vehicle.
The most notable target of angiogenesis inhibitors was the vascular endothelial growth factor (VEGF) signaling pathway (Fig. 7b). Among VEGF receptors, only KDR is exclusively expressed on the endothelium. Its expression level is very low in normal vasculature32, but it becomes highly activated during the growth of new blood vessels in tumors. Notably, we have identified GW786460X (originally targeting transforming growth factor-β receptor 1 (TGF-βR1), which was not in the kinase screening panel) as a potent inhibitor of KDR that is capable of substantially reducing tubule formation (Fig. 7c). It is less aggressive on the capillary-like network morphology compared to the pan-TK inhibitors such as GW795493X (originally targeting TIE2 and KDR). The use of selective KDR inhibitors may avoid undesirable side effects that result when using pan-TK inhibitors such as sunitinib33. Other PKIS compounds targeting kinases such as AKT, GSK3, p38, RAF and JNK also reduced endothelial tube formation. Compounds with cytostatic or antiproliferative properties, such as inhibitors of PLK or of EGFR and ERBB2, did not exhibit an antiangiogenic effect, an observation that may be due to the assay's duration of <12 h.
Discussion
Several large-scale kinase screens have been described previously1,9,34,35,36,37,38,39,40,41.Although they have increased understanding of many aspects of kinase pharmacology, their impact in drug discovery has been limited because they used undisclosed, unavailable and/or proprietary compounds or because the compounds studied were singletons—individual, structurally unrelated compounds that were typically clinical candidates or 'tool compounds'. In cases where structurally related compounds were included42,43, the insight into the molecular features for kinase inhibition enabled improved inhibitor design.
PKIS was designed to facilitate functional characterization of previously untargeted kinases. We provide free access to the physical compounds along with all of the chemical structures as an experiment in precompetitive compound sharing and open-access chemical biology (details are given in Online Methods). As compounds of interest are identified and testable hypotheses formed, individual solutions in larger quantity or solid samples of compounds may be made available to investigators for more detailed studies. For compounds whose supply is limited, procedures for their preparation are publicly available in the scientific literature. Compound transfer entails minimal legal requirements and no claims to intellectual property of resulting findings. However, data derived from use of the set must be placed directly into the public domain.
The uptake of PKIS in the scientific community has been encouraging. To date the set has been dispensed to over 250 collaborators, resulting in a large and growing body of data deposited at ChEMBL (https://www.ebi.ac.uk/chembl/). Applications have covered diverse biology, including characterizing the set for intrinsic luciferase inhibition activity to aid interpretation of phenotypic screens employing luciferase reporter assays44; profiling the set in an advanced model of the rare brain cancer ependymoma45; using PKIS in structural guided design of dual kinase and bromodomain inhibitors46; and progression through an in situ Western screen for inhibitors of ATM kinase47. PKIS was designed to capitalize on the chemical connectivity of ATP-competitive inhibitors that often show cross-reactivity across multiple kinases. Although medicinal chemists have developed strategies to address kinase selectivity, PKIS takes advantage of the potential for off target kinase activity to use the same chemotypes to explore the previously untargeted kinome. In the present case, chemical starting points for hitherto untargeted kinases (such as LOK, BRSK1/2, MKNK1 and AAK1) have been identified from the PKIS with modest affinity values and selectivity profiles.
In addition to the identification of chemical starting points for new kinases, the kinase activity map across a panel of hundreds of kinases can inform cellular screening efforts. Although biochemical assessments of kinase inhibitor interactions generally correlate with cellular efficacy48, the extrapolation of our in vitro results to the interpretation of cellular efficacy should proceed with a few caveats in mind. First, the Nanosyn kinase screening was conducted in the presence of ATP near the equilibrium constant for binding (Km) of the cofactor to the kinase. As such, the concentration of ATP varied across the panel by over two orders of magnitude. Cellular potency of ATP-competitive kinase inhibitors is determined not only by the intrinsic affinity of the inhibitor for the kinase but also by the Km of ATP and the cellular concentration of ATP48. Thus, the apparent potency and the rank order of kinase inhibition may differ in the cellular context. Second, many kinases in the panel are represented by constructs whose interactions with a compound could differ from that of endogenous enzyme, particularly in a cellular environment where protein scaffolding partners may be present. Third, although half of the human protein kinome is represented by the Nanosyn and DSF screening panels, there is likely additional compound activity with untested kinases or other targets.
The cellular activity of PKIS compounds was evaluated in the NCI-60 panel and in a model of angiogenesis. The results demonstrate the applicability of the set to phenotypic screening. The compounds showed a wide variety of activity profiles, potent cellular activity, and minimal nonspecific cytotoxicity. These data are expected to complement existing activity annotation and to inform future cellular screens involving PKIS.
Several recent observations suggest interactions of ATP-competitive kinase inhibitors with non-kinase targets15. ATP itself has been used to make an affinity capture reagent to identify more than 2,000 proteins that bind purine nucleotides as substrates or cofactors49. The results of screening PKIS in a panel of GPCR assays underscore the value of broad activity profiling to identify off target activity that might account for aberrant cellular activity or to increase understanding of kinase inhibitor polypharmacology.
Despite encouraging indications of the utility of the PKIS, the screening results point to opportunities to improve the set. If the figures of kinome coverage were extrapolated from the assay panels, the PKIS would provide inhibitor scaffolds for 90% of the kinome, or 467 kinases. However, many atypical kinase targets were not included in the screening panel, and the overall hit rate may ultimately be lower when these are included. It should be possible to expand coverage of the kinome through inclusion of additional compounds, whether from new chemotypes or expansion of those already present in the set. In addition, as redundant inhibition profiles or liability target activity are identified, compounds may be culled from the set.
In summary, we have presented PKIS, an open resource to stimulate research into the historically understudied kinome. It is hoped that the compounds and characterization data will help to illuminate the kinome further and identify new therapeutic targeting opportunities for pioneer medicines.
Methods
Nanosyn caliper screening.
Kinase activity was measured in vitro using an electrophoretic mobility shift assay. Kinase reactions were assembled in 384-well plates in a total volume of 25 μl. The reactions comprised: purified recombinant kinase enzyme, test compound, ATP (at apparent Km for each kinase, see Supplementary Data) FAM-labeled peptide substrates in a buffer composed of: 100 mM HEPES, pH 7.5; 10 mM MgCl2; 1 mM DTT; 0.1% BSA; 0.01% Triton X-100, and 1% DMSO (from compound). Lipid kinases PI3Kα, PI3Kβ and PI3Kγ were assayed in reaction buffer comprising 50 mM HEPES, pH 7.5, 3 mM MgCl2, 1 mM DTT, 0.012% CHAPS, 40 mM NaCl, 0.01% BSA and using BODIPY-FL-labeled phosphatidylinositol 4,5-biphosphate (PIP2) as a substrate. PI4Kβ was assayed in reaction buffer comprising: 50 mM HEPES, pH 7.5, 10 mM MgCl2, 0.1 mM DTT, 0.4% Triton X-100 and using BODIPY-FL-labeled phosphatidylinositol (PI) as a substrate. All reactions were incubated at room temperature for indicated time (Supplementary Data) and quenched by addition of 45 μL of termination buffer (100 mM HEPES, pH 7.5; 0.01% Triton X-100; 30 mM EDTA). Substrate and product peptides (or phospholipids) present in each sample were electrophoretically separated and detected using 12-channel LabChip3000 microfluidic capillary electrophoresis instrument (Caliper Life Sciences). The change in the relative fluorescence intensities of substrate and product peaks (reflecting enzyme activity) was measured. Capillary electrophoregrams were analyzed using HTS Well Analyzer software (Caliper Life Sciences). The kinase activity in each sample was determined as the product-to-sum ratio (PSR): P / (S + P), where P is the peak height of the product peptide and S is the peak height of the substrate peptide. Negative control samples (0% inhibition in the absence of inhibitor) and positive control samples (100% inhibition, in the presence of 20 mM EDTA) were assembled in replicates of four and were used to calculate percent inhibition values for each compound at each concentration. Percent inhibition (%Inhibition) was determined using the following equation:

where PSRinh is the product-sum ratio in the presence of inhibitor, PSR0% is the average product-sum ration in the absence of inhibitor and PSR100% is the average product-sum ratio in 100%-inhibition control samples. Reference inhibitors (i.e., staurosporine, SB-202190, PP2) were tested in 8-point dose-response format on each assay plate. The IC50 values of reference inhibitors were determined by fitting the inhibition curves by a four-parameter sigmoid dose-response model using XLfit 4 software (IDBS).
Differential scanning fluorimetry.
The PKIS was screened against purified protein kinases according to previously reported procedures8.
Split luciferase assay.
A coiled-coil enabled split-luciferase three-hydrid system assay was employed as previously reported14. Each test compound was screened in duplicate against AAK1 at seven different concentrations. The catalytic domain of AAK1 conjugated to the C-terminal fragment of luciferase (CFluc-kinase) was translated along with the N-terminal fragment of luciferase using a cell-free system (rabbit reticulocyte lysate) at 30 °C for 90 min. A 24-μL aliquot of this lysate containing either 1 μL of DMSO (for no-inhibitor control) or compound solution in DMSO was incubated for 30 min at room temperature and for 1 h in the presence of a kinase-specific probe. Luciferin assay reagent (80 μL) was added to each solution and luminescence was immediately measured on a luminometer. Prior to initiating IC50 determinations, the test compounds were evaluated for false positives. The percentage inhibition was calculated as

The percentage inhibition was plotted against compound concentration, and the IC50 was determined for each compound using a seven-point curve.
Biolayer interferometry.
Biotinylated LOK or SLK kinase domains were bound to tips of a FortéBio Octet RED instrument. A dilution series of each compound was used to measure a dose-response curve of association and dissocation. Kd values were calculated in GraphPad Prism using the one-site specific binding model.
GPCR screening.
Calcium mobilization assays were carried out with cells stably expressing cloned human receptors essentially as described previously50,51,52, and complete protocol details are available online (https://pdspdb.unc.edu/pdspWeb/?site=functional). In brief, cells were seeded in poly-L-lysine–coated 384-well black clear-bottom cell-culture plates using DMEM supplemented with 1% dialyzed FBS in a total of 40 μl per well and at a density of 15,000 cells per well and left overnight. On the day of assay, medium was removed and cells were loaded (20 μl/well) with Fluo-4 Direct dye (Invitrogen) prepared with assay buffer (1× HBSS, 20 mM HEPES, 2.5 mM probenecid, pH 7.40) for 1 h at 37 °C. A fluorescence imaging plate reader (FLIPRTETRA, Molecular Dynamics) was programmed to read baseline fluorescence for 10 s (1 read per second), then to add 10 μl of drug per well to assess agonist activity. The FLIPR was programmed to wash tips then add a reference agonist at a final concentration of EC80 (the concentration to elicit 80% of the maximum reference agonist, predetermined before antagonist activity assay) to assess potential antagonist activity 10 min later. The baseline for antagonist activity was the average fluorescence of 10 readings before addition of EC80 reference agonist with the fluorescence in each well corrected and normalized to corresponding baselines. The maximum signals (fold of baseline) during 60 s after drug addition were then exported. Agonist activity in initial screening assays was determined at a final concentration of 10 μM (quadruplicate determinations) and normalized with the corresponding reference agonist activity as 100% and basal as 0%. Antagonist activity in screening assays was measured at a final concentration of 8 μM and normalized with basal representing 100% inhibition and the agonist activity at EC80 concentration as 0% inhibition. For Schild plot analysis, agonist concentration-responses (16 points in triplicate) were measured in the absence and presence of increasing concentrations of test compounds. Results were analyzed using GraphPad Prism 5.0.
NCI-60 panel screening.
The set was screened across the NCI-60 panel at the National Cancer Institute using previously described methodology, protocols and cell-line verification53.
Cellular assays.
The KM12 cell line was obtained from the National Cancer Institute. The Jurkat cell line was obtained from A.P. Cribbs (Kennedy Institute of Rheumatology, Oxford University). All cell lines were regularly checked for mycoplasma contamination.
Anti-TRKA-pY496 and anti-TRKA antibodies were from Santa Cruz Biotechnology (#sc-7987 and #sc-118), anti–phospho-ERM antibody was from Pierce/Thermo Scientific (#MA5-14998) and anti-moesin was from Abcam (#ab52490) and anti-GAPDH (14C10) was from New England Biolabs (#2118S). All antibodies were validated for Western blots against the protein targets used by the antibody vendor. Antibodies were used at the following dilutions: sc-7987, 1:200; sc-118, 1:200; MA5-14998, 1:1,000; ab52490, 1:10,000; 2118S, 1:1,000. The KM12 cells were cultured in RPMI 1640 media (Sigma, R0883) supplemented with 2 mM L-glutamine (Invitrogen, 35050) and 10% heat-inactivated FBS (Life Technologies, 10500-064). Twenty-four hours before treatment cells were seeded in 6-well plates (7.5 × 105 cells per well). KM12 cells were treated with GW768505A or GW441756X positive control at a concentration of 250 nM for 2.5, 5.0, 15 or 30 min, at 37 °C and 5% CO2. DMSO was used as negative control. After treatment, cells were immediately washed and scraped using cold PBS containing 1% of PhosSTOP inhibitor cocktails (Roche, 04906845001), then collected by centrifugation at 2,500 r.p.m. at 4 °C for 3 min.
Jurkat cells were cultured in DMEM supplemented with 2 mM L-glutamine (Invitrogen, 35050) and 10% heat-inactivated FBS (Life Technologies, 10500-064). Cells were counted and aliquoted in single 2-mL centrifuge tubes at a concentration of 2 × 106 cells per tube. Jukart cells were then treated with GSK3037619A, erlotinib or staurosporine at 1 μM for 5 min at 37 °C and 5% CO2, in the presence or absence of 80 nM phosphatase inhibitor calyculin A (Cell Signaling, #9902). DMSO, plus or minus calyculin A, was used as negative control. After treatment, cells were immediately washed using cold PBS containing 1% of PhosSTOP inhibitor cocktails (Roche, 04906845001), then collected by centrifugation at 2,500 r.p.m. at 4 °C for 5 min.
Cells were lysed in RIPA buffer (Sigma-Aldrich, R0278) containing complete EDTA-free protease inhibitor cocktail (Roche, 11873580001) and PhosSTOP inhibitor cocktails. Protein concentration was determined using Pierce BCA Protein Assay Kit (Life Technologies, 23225) and 55 μg of lysate was separated by SDS-PAGE. The molecular weight marker was from LI-COR (928-40000 Odyssey Protein Molecular Weight Marker). Gels were transferred onto nitrocellulose membranes, blocked with 5% nonfat dry milk in 1× TBS containing 0.05% Tween-20 (TBST) and then incubated with the appropriate primary antibody in 5% BSA in PBST. Membranes were washed in 1× TBST and incubated for 1 h in the appropriate secondary antibody (goat anti-rabbit Alexa Fluor 680, #A21109) diluted 1:10,000 in 5% nonfat dry milk in PBST. The blots were washed and visualized using Odyssey CLx Infrared Imaging System.
High-content screening for modulators of angiogenesis.
Primary human umbilical endothelial cells (HUVECs, pooled) were purchased from Lonza. Cells were maintained in endothelial cell medium (EGM-2, Lonza). Low-passage cells were used for the screen. Cells were seeded on top of Matrigel (an extract of basement membrane proteins, BD Bioscience), in 96-well plates and incubated with PKIS compounds at concentration of 10 μM for 10 h at 37 °C and 5% CO2. The screen was performed in duplicate under rich vascular growth factor conditions (full EGM-2 medium). Endothelial cells formed capillary-like vascular structures (tubules) in the presence of the Matrigel and vascular growth factors. The endothelial cellular tubules were fixed with 4% PFA and stained with Alexa Fluor 568 phalloidin (Invitrogen) to visualize the tubules. Next, the plates were imaged using a high-content microscopy system (Operetta, PerkinElmer). The 16-bit images were processed, segmented and analyzed using fully automated Metamorph image analysis software (Molecular Devices) (Supplementary Fig. 12). Vehicle (DMSO) was used as negative control. Suramin was used as a positive control, which showed an effective inhibition of endothelial tube formation in a dose-dependent manner (pilot validation assays). The total tubule length reading values were normalized according to the B-score method with the CellHTS2 package54. The B score is a normalization method to account for plate-to-plate variances that involves the residual values calculated from median polish and the median absolute deviation (MAD) of sample to account for data variability. The selection cutoff for inhibitor hits was set at 2× MAD, which is −1.5. R2 correlation between replicates was 0.7 (Supplementary Fig. 12c) and the Z factor was >0.3 in all plates. The data were compiled and analyzed using Spotfire and GraphPad packages.
Access to PKIS.
To obtain the PKIS, prospective collaborators must agree to terms of a streamlined materials transfer agreement (MTA), the most significant requirement of which is that data produced from use of the set be made publicly available. The MTA, which can obtained via the Structural Genomics Consortium at the University of North Carolina at Chapel Hill website (https://pharmacy.unc.edu/research/sgc-unc/) or by e-mail request (SGC-UNC@unc.edu), is offered for consideration without modification. Changes requested to the template can lead to substantial delays in consideration and run the risk of failure to establish collaboration.
Accession codes.
PDB: The crystal structures of LOK (STK10) and SLK with inhibitors have been deposited in the Protein Data Bank under accession codes 4USD, 4USE and 4USF.
References
Bamborough, P. System-based drug discovery within the human kinome. Expert Opin. Drug Discov. 7, 1053–1070 (2012).
Cohen, P. & Alessi, D.R. Kinase drug discovery—what's next in the field? ACS Chem. Biol. 8, 96–104 (2013).
Wu, P., Nielsen, T.E. & Clausen, M.H. FDA-approved small-molecule kinase inhibitors. Trends Pharmacol. Sci. 36, 422–439 (2015).
Fedorov, O., Müller, S. & Knapp, S. The (un)targeted cancer kinome. Nat. Chem. Biol. 6, 166–169 (2010).
Edwards, A.M., Bountra, C., Kerr, D.J. & Willson, T.M. Open access chemical and clinical probes to support drug discovery. Nat. Chem. Biol. 5, 436–440 (2009).
Drewry, D.H., Willson, T.M. & Zuercher, W.J. Seeding collaborations to advance kinase science with the GSK Published Kinase Inhibitor Set (PKIS). Curr. Top. Med. Chem. 14, 340–342 (2014).
Zhang, J., Yang, P.L. & Gray, N.S. Targeting cancer with small molecule kinase inhibitors. Nat. Rev. Cancer 9, 28–39 (2009).
Fedorov, O., Niesen, F.H. & Knapp, S. Kinase inhibitor selectivity profiling using differential scanning fluorimetry. Methods Mol. Biol. 795, 109–118 (2012).
Bamborough, P., Drewry, D., Harper, G., Smith, G.K. & Schneider, K. Assessment of chemical coverage of kinome space and its implications for kinase drug discovery. J. Med. Chem. 51, 7898–7914 (2008).
Witherington, J. et al. 5-aryl-pyrazolo[3,4-b]pyridazines: potent inhibitors of glycogen synthase kinase-3 (GSK-3). Bioorg. Med. Chem. Lett. 13, 1581–1584 (2003).
Conway, J.G. et al. Inhibition of colony-stimulating-factor-1 signaling in vivo with the orally bioavailable cFMS kinase inhibitor GW2580. Proc. Natl. Acad. Sci. USA 102, 16078–16083 (2005).
Takle, A.K. et al. The identification of potent and selective imidazole-based inhibitors of B-Raf kinase. Bioorg. Med. Chem. Lett. 16, 378–381 (2006).
Workman, P. & Collins, I. Probing the probes: fitness factors for small molecule tools. Chem. Biol. 17, 561–577 (2010).
Jester, B.W. et al. A coiled-coil enabled split-luciferase three-hybrid system: applied toward profiling inhibitors of protein kinases. J. Am. Chem. Soc. 132, 11727–11735 (2010).
Paolini, G.V., Shapland, R.H., van Hoorn, W.P., Mason, J.S. & Hopkins, A.L. Global mapping of pharmacological space. Nat. Biotechnol. 24, 805–815 (2006).
Ciceri, P. et al. Dual kinase-bromodomain inhibitors for rationally designed polypharmacology. Nat. Chem. Biol. 10, 305–312 (2014).
Huber, K.V. et al. Stereospecific targeting of MTH1 by (S)-crizotinib as an anticancer strategy. Nature 508, 222–227 (2014).
Fourches, D. & Tropsha, A. Using graph indices for the analysis and comparison of chemical datasets. Mol. Inform. 32, 827–842 (2013).
Shoemaker, R.H. The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 6, 813–823 (2006).
Weinstein, J.N. et al. An information-intensive approach to the molecular pharmacology of cancer. Science 275, 343–349 (1997).
Reinhold, W.C. et al. CellMiner: a web-based suite of genomic and pharmacologic tools to explore transcript and drug patterns in the NCI-60 cell line set. Cancer Res. 72, 3499–3511 (2012).
Miyazaki, Y. et al. Orally active 4-amino-5-diarylurea-furo[2,3-d]pyrimidine derivatives as anti-angiogenic agent inhibiting VEGFR2 and Tie-2. Bioorg. Med. Chem. Lett. 17, 1773–1778 (2007).
Ardini, E. et al. The TPM3-NTRK1 rearrangement is a recurring event in colorectal carcinoma and is associated with tumor sensitivity to TRKA kinase inhibition. Mol. Oncol. 8, 1495–1507 (2014).
Hasegawa, M. et al. Discovery of novel benzimidazoles as potent inhibitors of TIE-2 and VEGFR-2 tyrosine kinase receptors. J. Med. Chem. 50, 4453–4470 (2007).
Belkina, N.V., Liu, Y., Hao, J.J., Karasuyama, H. & Shaw, S. LOK is a major ERM kinase in resting lymphocytes and regulates cytoskeletal rearrangement through ERM phosphorylation. Proc. Natl. Acad. Sci. USA 106, 4707–4712 (2009).
Kuramochi, S. et al. LOK is a novel mouse STE20-like protein kinase that is expressed predominantly in lymphocytes. J. Biol. Chem. 272, 22679–22684 (1997).
Viswanatha, R., Ohouo, P.Y., Smolka, M.B. & Bretscher, A. Local phosphocycling mediated by LOK/SLK restricts ezrin function to the apical aspect of epithelial cells. J. Cell Biol. 199, 969–984 (2012).
Folkman, J. Angiogenesis: an organizing principle for drug discovery? Nat. Rev. Drug Discov. 6, 273–286 (2007).
Arnaoutova, I. & Kleinman, H.K. In vitro angiogenesis: endothelial cell tube formation on gelled basement membrane extract. Nat. Protoc. 5, 628–635 (2010).
Faivre, S., Demetri, G., Sargent, W. & Raymond, E. Molecular basis for sunitinib efficacy and future clinical development. Nat. Rev. Drug Discov. 6, 734–745 (2007).
Wilhelm, S. et al. Discovery and development of sorafenib: a multikinase inhibitor for treating cancer. Nat. Rev. Drug Discov. 5, 835–844 (2006).
Olsson, A.K., Dimberg, A., Kreuger, J. & Claesson-Welsh, L. VEGF receptor signalling—in control of vascular function. Nat. Rev. Mol. Cell Biol. 7, 359–371 (2006).
Verheul, H.M. & Pinedo, H.M. Possible molecular mechanisms involved in the toxicity of angiogenesis inhibition. Nat. Rev. Cancer 7, 475–485 (2007).
Gao, Y. et al. A broad activity screen in support of a chemogenomic map for kinase signalling research and drug discovery. Biochem. J. 451, 313–328 (2013).
Anastassiadis, T., Deacon, S.W., Devarajan, K., Ma, H. & Peterson, J.R. Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1039–1045 (2011).
Davis, M.I. et al. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 29, 1046–1051 (2011).
Fedorov, O. et al. A systematic interaction map of validated kinase inhibitors with Ser/Thr kinases. Proc. Natl. Acad. Sci. USA 104, 20523–20528 (2007).
Bain, J. et al. The selectivity of protein kinase inhibitors: a further update. Biochem. J. 408, 297–315 (2007).
Davies, S.P., Reddy, H., Caivano, M. & Cohen, P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem. J. 351, 95–105 (2000).
Fabian, M.A. et al. A small molecule-kinase interaction map for clinical kinase inhibitors. Nat. Biotechnol. 23, 329–336 (2005).
Karaman, M.W. et al. A quantitative analysis of kinase inhibitor selectivity. Nat. Biotechnol. 26, 127–132 (2008).
Metz, J.T. et al. Navigating the kinome. Nat. Chem. Biol. 7, 200–202 (2011).
Posy, S.L. et al. Trends in kinase selectivity: insights for target class-focused library screening. J. Med. Chem. 54, 54–66 (2011).
Dranchak, P. et al. Profile of the GSK published protein kinase inhibitor set across ATP-dependent and-independent luciferases: implications for reporter-gene assays. PLoS One 8, e57888 (2013).
Atkinson, J.M. et al. An integrated in vitro and in vivo high-throughput screen identifies treatment leads for ependymoma. Cancer Cell 20, 384–399 (2011).
Ember, S.W. et al. Acetyl-lysine binding site of bromodomain-containing protein 4 (BRD4) interacts with diverse kinase inhibitors. ACS Chem. Biol. 9, 1160–1171 (2014).
Guo, K., Shelat, A.A., Guy, R.K. & Kastan, M.B. Development of a cell-based, high-throughput screening assay for ATM kinase inhibitors. J. Biomol. Screen. 19, 538–546 (2014).
Knight, Z.A. & Shokat, K.M. Features of selective kinase inhibitors. Chem. Biol. 12, 621–637 (2005).
Haystead, T.A. The purinome, a complex mix of drug and toxicity targets. Curr. Top. Med. Chem. 6, 1117–1127 (2006).
Besnard, J. et al. Automated design of ligands to polypharmacological profiles. Nature 492, 215–220 (2012).
Huang, X.P. et al. Parallel functional activity profiling reveals valvulopathogens are potent 5-hydroxytryptamine2B receptor agonists: implications for drug safety assessment. Mol. Pharmacol. 76, 710–722 (2009).
Wacker, D. et al. Structural features for functional selectivity at serotonin receptors. Science 340, 615–619 (2013).
Holbeck, S.L., Collins, J.M. & Doroshow, J.H. Analysis of Food and Drug Administration-approved anticancer agents in the NCI60 panel of human tumor cell lines. Mol. Cancer Ther. 9, 1451–1460 (2010).
Boutros, M., Brás, L.P. & Huber, W. Analysis of cell-based RNAi screens. Genome Biol. 7, R66 (2006).
Acknowledgements
The Structural Genomics Consortium is a registered charity (number 1097737) that receives funds from AbbVie, Bayer Pharma AG, Boehringer Ingelheim, Canada Foundation for Innovation, Eshelman Institute for Innovation, Genome Canada, Innovative Medicines Initiative (EU/EFPIA), Janssen, Merck & Co., Novartis Pharma AG, Ontario Ministry of Economic Development and Innovation, Pfizer, São Paulo Research Foundation-FAPESP, Takeda, and Wellcome Trust. A.A.H.Z. acknowledges support from the BHF Centre of Research Excellence, Oxford (RE/13/1/30181).
Author information
Authors and Affiliations
Contributions
K.E.L., J.M.E., A.A.H.Z., J. Morris, S.R., N.S., B.A.T., B.L.R., D.H.D., S.K. and W.J.Z. designed the study. M.S., V.F., K.R.A.A., J.M.E., E.S., A.A.H.Z., J. Mikolajczyk, S.R., N.S. and X.-P.H. performed experiments. J.M.E., A.A.H.Z., J. Morris, B.A.T., A.T., X.-P.H., D.F., E.M., F.L.A., J.P.O., P.B., D.H.D., S.M., S.K., E.P., M.K. and W.J.Z. analyzed data. J.M.E., B.L.R., X.-P.H., A.A.H.Z., P.B., D.H.D., D.J.P., T.M.W., S.K. and W.J.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.R. and N.S. are current employees of Nanosyn, Inc. J.P.O. is a current employee of Stratified Medical Ltd.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1-11 and Supplementary Figures 1-12 (PDF 6677 kb)
Rights and permissions
About this article
Cite this article
Elkins, J., Fedele, V., Szklarz, M. et al. Comprehensive characterization of the Published Kinase Inhibitor Set. Nat Biotechnol 34, 95–103 (2016). https://doi.org/10.1038/nbt.3374
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3374
- Springer Nature America, Inc.
This article is cited by
-
Identification of druggable regulators of cell secretion via a kinome-wide screen and high-throughput immunomagnetic cell sorting
Nature Biomedical Engineering (2023)
-
Chemogenetics for cell-type-specific modulation of signalling and neuronal activity
Nature Reviews Methods Primers (2023)
-
Opposing roles of CLK SR kinases in controlling HIV-1 gene expression and latency
Retrovirology (2022)
-
Dissecting the roles of Haspin and VRK1 in histone H3 phosphorylation during mitosis
Scientific Reports (2022)
-
A novel graph convolutional neural network for predicting interaction sites on protein kinase inhibitors in phosphorylation
Scientific Reports (2022)