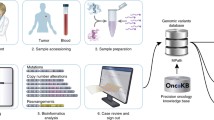
Abstract
Metastasis is the primary cause of cancer-related deaths. Although The Cancer Genome Atlas has sequenced primary tumour types obtained from surgical resections, much less comprehensive molecular analysis is available from clinically acquired metastatic cancers. Here we perform whole-exome and -transcriptome sequencing of 500 adult patients with metastatic solid tumours of diverse lineage and biopsy site. The most prevalent genes somatically altered in metastatic cancer included TP53, CDKN2A, PTEN, PIK3CA, and RB1. Putative pathogenic germline variants were present in 12.2% of cases of which 75% were related to defects in DNA repair. RNA sequencing complemented DNA sequencing to identify gene fusions, pathway activation, and immune profiling. Our results show that integrative sequence analysis provides a clinically relevant, multi-dimensional view of the complex molecular landscape and microenvironment of metastatic cancers.
Similar content being viewed by others
Main
Tumour metastasis is the process by which cancer cells disperse from a primary site and progressively colonize distant organs. In over 90% of cases, metastatic spread of tumour cells is the greatest contributor to deaths from cancer1,2. With the preponderance of cancer patients enrolled in early stage (phase I–II) clinical trials harbouring metastatic disease2, and with the advent of genomic testing of tumours, there remains the promise of matching patients to the right therapy on the basis of comprehensive molecular profiles3 of pathogenic somatic4,5 and germline6 variants, and components of the functional genome, tumour phenotype, and tumour microenvironment afforded by RNA sequencing (RNA-seq)7,8. While metastatic tumours share key driver mutations with the primary tumour from which they arise, they often develop new mutations as they evolve during metastasis and treatment9. Thus, it is preferable to match patients to potential therapies and clinical trials on the basis of a real-time analysis of their metastatic tumour, rather than archival material of their primary tumour2,10.
In 2010, we introduced the Michigan Oncology Sequencing (MI-ONCOSEQ) Program, an institutional review board (IRB)-approved protocol to perform prospective, integrative exome and transcriptome sequencing of advanced cancers in patients7, mirroring the efforts of The Cancer Genome Atlas (TCGA) project, which focuses on generating exome and transcriptome sequence11 in primary cancers. The MI-ONCOSEQ program was among the first comprehensive, clinical DNA- and RNA-seq programs offered to cancer patients8,12,13,14,15,16. Its purpose was to determine the use of genomic sequencing of tumours and germline coupled with a multidisciplinary precision medicine tumour board (PMTB) in the management of advanced cancer in patients7,15. The program transitioned into sequencing in a clinical setting (under Clinical Laboratory Improvement Amendments) as part of the Clinical Sequencing Exploratory Research consortium in 2013 (refs 14, 15, 16).
In this study, we performed clinical-grade whole-exome (tumour/normal) and -transcriptome sequencing (that is, integrative sequencing) of 500 cancer patients harbouring metastatic cancers from over 30 primary sites and biopsied from over 22 organs (abbreviated as the ‘MET500’ cohort). Sequencing-matched tumour and normal samples from patients delineated potentially pathogenic germline alterations and provided high-resolution copy-number landscapes. RNA-seq analysis provided insights into functional gene fusions, transcriptional pathway activation, and a landscape of immune infiltration.
Molecular aberrations in metastatic cancers
We successfully obtained 537 biopsies from 556 enrolled patients, and complete sequencing results on 500 patients with metastatic cancers, representing a 93% success rate. Reasons for failure included lack of tumour content on biopsy (37 cases, 6.6%) and biopsy material not available (19 cases, 3.4%; patient declined biopsy, poor physical performance, unable to image site, unsafe for biopsy, insufficient tissue, or enrolled in other clinical trials). Most patients, 468 (93.6%), were seen at the University of Michigan Comprehensive Cancer Center; however, patients from 21 other institutions were also enrolled. The patient demographics were 258 (51.6%) males, 242 females (48.4%), 460 (92%) white, and 40 (8%) non-white. The median age of the cohort was 59 years, with a range of 18–86 (Supplementary Table 1 and Extended Data Fig. 1a, b).
Figure 1a shows the cancer types (n = 20) represented in the MET500 cohort. The top three cancer types in our cohort included 93 (18.6%) metastatic prostate cancers, 91 (18.2%) metastatic breast cancers, and 42 (8.4%) soft tissue sarcomas. There were also 25 (5%) carcinomas of unknown primary origin. Figure 1b highlights the diverse metastatic sites analysed (n > 30) in the MET500 cohort. The most prevalent sites of metastases included 134 liver, 114 lymph node, 46 lung, 42 bone, and 32 abdominal mass/ascites/pleural fluid.
a, Cancer types in the MET500 cohort. Number of cases indicated for each cancer type. GIST, gastrointestinal stromal tumour. b, Site of biopsies. c, Mutational burden across tumour types from the MET500 and corresponding primary TCGA cohorts. Transparent box plots signify insignificant differences (Wilcoxon rank-sum test false discovery rate ≥ 0.1). d, e, Landscape of molecular alterations in the MET500 cohort. Each cell represents the mutation status of an individual gene for a single patient. Putative oncogenes are represented in d and putative tumour-suppressor genes in e. The percentage of mutations across the MET500 cohort is represented by vertical histograms.
For each patient, we performed paired-exome sequencing on tumour and germline DNA to identify likely pathogenic variants and resolve their somatic or germline origin. Mean target coverages for tumour and normal exomes were 180× and 120×, respectively. The average tumour content was 62%. Sequencing metrics are summarized in Supplementary Table 2. Within the targeted regions, we identified an average of 119 somatic mutations per patient. For most cancer types, the number of mutations significantly increased in metastases relative to primary tumours in TCGA (Fig. 1c). The difference was more pronounced for tumour types with low mutation burden in the primary stage, for example prostate or adrenal cancer. To identify the most recurrent, and hence likely pathogenic, targets of genetic alterations, we performed an integrative analysis of single-nucleotide variants (SNVs), copy-number variants (CNVs), and gene fusions. For each patient and gene, we classified the most recurrently mutated genes as putative tumour suppressors or oncogenes on the basis of the increase in frequency of inactivating mutations and expert knowledge (Fig. 1d, e and Supplementary Table 3). We found a long-tailed mutational spectrum for both tumour suppressors and oncogenes. TP53 (266, 53.2%), CDKN2A (80, 16%), PTEN (79, 15.8%), and RB1 (68, 13.6%) were the most frequently altered tumour suppressors, while the most frequently mutated oncogenes included PIK3CA (67, 13.4%), AR (63, 12.6%), and KRAS (51, 10.2%). Overall, tumour suppressors were altered across many cancer types (for example, TP53 and RB1), while oncogenes were more strongly associated with individual cancer types (for example, AR or GNAS) (Extended Data Fig. 1c). We further compared the alteration frequencies with those from primary tumours and found that the increase in mutation burden (Fig. 1c) was mirrored by an increase in the frequency of genetic aberrations for the most widely mutated genes (Extended Data Fig. 1d).
Germline variants in metastatic cancer
Through sequencing-matched germline DNA, we identified 63 presumed pathogenic germline mutations (PPGMs), as defined by ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/) expert curation, involving 18 genes in 61 individuals (12.2%) (Fig. 2a). These included 30 deleterious missense mutations, 8 nonsense mutations, 20 frameshift mutations, and 5 deleterious splice site mutations (Fig. 2b and Supplementary Table 4). In total, 75% of the PPGMs were in genes related to DNA repair, of these, mutations in MUTYH (n = 10, 16%), BRCA2 (n = 9, 14%), CHEK2 (n = 9, 14%), and BRCA1 (n = 5, 8%) were the most common. Outside DNA repair pathways, we observed PPGMs in APC (n = 6, 9.5%), MITF (n = 5, 8%), and HOXB13 (n = 3, 5%), among other genes. Of the 63 instances of pathogenic alleles identified, 5 were previously unreported, while the remaining alleles had existing ClinVar entries with assigned pathogenic or likely pathogenic importance. Of the 61 individuals with PPGMs identified in this study, 30 (49%) had a somatic second allele aberration within the tumour genome, including loss of heterozygosity, and exhibited molecular phenotypes consistent with pathogenicity (Supplementary Table 4). The remaining cases were of carrier status for the identified allele.
a, Pathogenic germline alterations identified in the MET500 cohort. DNA repair pathway-related variants are indicated in shades of blue, while ‘other’ alterations are indicated in shades of red. b, Gene-level schematic of pathogenic germline variants identified in the MET500 cohort. LOH, loss of heterozygosity. *, stop gain; fs, frameshift.
Next, we compared the frequencies in genes with PPGMs discovered in the MET500 cohort with the population frequencies in 52,790 individuals compiled by the Exome Aggregation Consortium (http://exac.broadinstitute.org/), excluding TCGA cancer samples17. The odds of any PPGM in metastatic cancer significantly exceeded the odds found in the populations comprising the Exome Aggregation Consortium (odds ratio = 3.00, range 2.28–3.9, P = 1 × 10−13). The genes analysed and found to be enriched in the metastatic series included BRCA1, BRCA2, APC, CHEK2, MITF, MLH1, NBN, and RB1 (Supplementary Table 5).
The gene fusion landscape of metastatic cancer
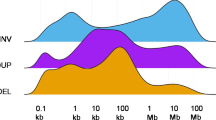
To identify both activating and inactivating gene fusions, we analysed 868 transcriptome libraries from 496 metastatic tumour RNAs. These fusion junctions involved 12,027 unique gene pairs, an average of 34 gene fusions per tumour, derived from a range of structural aberrations (Fig. 3a). Large differences in fusion burden were observed across tumour type (Extended Data Fig. 2a). One hundred and ninety-nine cases (39.8%) harboured at least one putative pathogenic fusion, with 138 activating fusions and 103 deleterious fusions (Supplementary Table 6). The activating fusions could be classified as DNA-binding (n = 88), protein kinases (n = 29), and signal transducers (n = 21) (Fig. 3b). The loss-of-function fusions segregated into canonical tumour-suppressor genes (n = 59), chromatin-modifying genes (n = 35), and genes involved in cell adhesion (n = 9). The most commonly fused tumour-suppressor genes were NF1 (n = 18), TP53 (n = 11), PTEN (n = 11), and RB1 (n = 6) (Extended Data Fig. 2b). Interestingly, we identified a series of eight novel fusion pairs in metastatic cancers that we believe are pathogenic (Fig. 3c). These include activated FGFR, BRAF, and ALK fusions with novel partners, extending the range of both fusion partners and cancer types for these clinically targetable fusions18,19,20,21,22. Novel gene fusions with functional domains include GREB1–NR4A3 in uterine leiomyosarcoma, POC5–PRKD1 in polymorphous low-grade adenocarcinoma of the tongue, and CIC–CITED1 in undifferentiated high-grade sarcoma. Notch fusions fall into two classes: those predicted to be sensitive to γ-secretase inhibition (for example, NOTCH2–SPAG17), and fusions that are independent of γ-secretase processing (for example, PARS2–NOTCH2).
a, Fusions classified by underlying structural aberrations. b, Functional gene fusions identified in the MET500 cohort. c, Molecular structure of novel, potentially activating, gene fusions in the MET500 cohort.
Transcriptional signatures of metastatic disease
To investigate the potential clinical use of metastatic expression profiles, we analysed transcriptomes of the 496 biopsy samples (868 libraries). We first evaluated to what extent tissue- and cancer-specific gene expression was maintained across metastatic lesions. We used the t-distributed stochastic neighbourhood embedding (t-SNE) projection23 to qualitatively visualize the expression of primary cancer markers across the MET500. Compared with primary tumours, metastatic samples were less well separated, more heterogeneous, and did not segregate on the basis of biopsy site, with the exception of liver biopsies (Fig. 4a and Extended Data Fig. 2c, d). We compared the expression of tissue-specific marker genes derived from 36 normal tissues24 between normal, primary, and metastatic samples, and observed significant de-differentiation with disease progression (Extended Data Fig. 2e).
a, Global gene expression patterns of the MET500 identify poorly differentiated cancers as illustrated by a t-SNE projection of the MET500 samples. Position of samples within the plot reflects the relative similarity in the expression of cancer-specific markers. Samples are colour-coded on the basis of their assigned analysis cohort. See Fig. 1c for cancer abbreviations; MISC, miscellaneous. b, Correlation heatmap and hierarchical clustering showing similarities (red) and dissimilarities (blue) in the transcriptional activity of computationally derived aggregate sets of signatures across the MET500 ‘meta-signatures’; lit., signature extracted from literature. c, Negative correlation between signatures of EMT and proliferation (S phase of the cell cycle, FANCA pathway). All MET500 samples are shown.
Next, we looked at transcriptional signatures associated with perturbed cancer-related genes25,26. Compared with normal tissues, transcriptional output was increased for most oncogenic signatures (Extended Data Fig. 3a, b), indicating a global shift towards a cancer-related transcriptional program. Unsupervised clustering of signature scores across patients revealed relevant associations between gene sets, and phenotypic similarities among patients (Extended Data Fig. 4). Inference of patient-specific activities27,28 revealed coordinated changes across curated pathways that coalesced into a small number of principal cancer hallmarks (Extended Data Fig. 5a, b): interferon response, inflammatory response, epithelial to mesenchymal transition (EMT), proliferation, and metabolism. Importantly, these associations were robust to algorithm choice. Compared with normal tissues, metastatic tumours show a global increase in proliferation, stress response, and metabolism. Conversely, hallmarks of EMT and cancer-immune responses can be either up- or downregulated (Extended Data Fig. 5a, b). Next, we computationally delineated 25 non-redundant experimental ‘meta-signatures’ (Extended Data Fig. 6). Unsupervised clustering and correlation analysis of meta-signatures revealed four of the canonical cancer hallmarks: immune response, EMT, proliferation, and metabolism (Fig. 4b). Metastatic tumours fall into two main subtypes: an EMT-like subtype associated with inflammation signatures29, and a proliferative subtype associated with increased metabolism and systemic stress. In agreement, we observed mutual exclusivity between curated proliferative and EMT gene sets (Fig. 4c). Interestingly, this trend was less prominent across primary tumours (Extended Data Fig. 7a). Importantly, meta-signature activities were found to be weakly associated with biopsy site (Extended Data Fig. 7b) and primary tissue (Extended Data Fig. 7c), and held independently for common cancer types and biopsy sites (Extended Data Fig. 7d).
The immune microenvironment of metastatic disease
To characterize the phenotype of host immune responses, we leveraged exome, RNA-seq, and a dedicated assay for T-cell repertoire profiling. On the basis of immune-cell markers proposed in ref. 30, we developed an RNA-seq-based score, MImmScore, to assess the magnitude of leukocyte infiltration. We found that MImmScore was negatively correlated with tumour content (Extended Data Fig. 8a), and positively correlated with stromal infiltration (Extended Data Fig. 8b). MImmScore was compared with canonical T-cell expression markers (RNA-seq-based) and DNA-based T-cell receptor-β (TCRβ) CDR3 sequencing: all three analyses were in good agreement (Extended Data Fig. 8c, d). We also discovered that metastatic immune infiltration was strongly determined by tumour type (Fig. 5a) and to a lesser degree by biopsy site (Extended Data Fig. 9a). Cancer types known to be infiltrated in the localized stage (Extended Data Fig. 9b), including kidney cancer31,32, lung cancer33, and melanoma34, remained infiltrated at metastatic sites. Less immunogenic types, such as breast and prostate cancers, were generally associated with a lower MImmScore in the primary and metastatic stages (Fig. 5a and Extended Data Fig. 9b). Immune infiltration was found to be heterogeneous not only across cancer types but also within individual cohorts (Extended Data Fig. 9c, d). Strikingly, individual patients with high levels of immune infiltration could be identified even within tumour types that did not thus far respond to immunotherapies.
a, Magnitude of immune (leukocyte) infiltration (MImmScore) across the MET500 analysis cohort. b, Hierarchical clustering of samples by their predicted immune infiltrates. c, d, TCR profiling by TCRβ DNA deep sequencing (TCRB–seq). Correlation of estimated T-cell numbers (templates): c, with clonal expansion (high clonality indicates that many T cells have the same TCRβ sequence) (ρ, Spearman’s rank correlation coefficient); d, with number of mutations. e, Clusters of patients based on the normalized expression levels of APC (left) or T-cell (right) surface molecules. f–h, Genomic correlates for patients grouped by their membership in immunologically active clusters: TIL-5, APC-1, Tcell-1 (silent, none; partial, some; complete, all). f, Expression of PD-L1 (t-test); RPKM, reads per kilobase per million mapped reads. g, Number of non-synonymous mutations (Wilcoxon rank-sum test). h, Response score based on a predictive gene expression signature to immunotherapy (t-test). Significance levels: *P = 0.05–0.001, **P = 0.001–10−6, ***P ≤ 10−6.
We hypothesized that metastatic tumours differ not only in the magnitude but also in composition of leukocyte cell types, tumour-infiltrating lymphocytes (TILs), and macrophages. Unsupervised clustering revealed groups of samples with significant differences in TIL composition, on the basis of bulk tumour transcriptome data35 (Fig. 5b). Cancers were most strongly typified by the different ratios of M2 to M0 (unpolarized) macrophages (clusters TIL-2, -4, -6, -7) and different CD8+ to CD4+ T-cell ratios (high CD8+ TIL-1, high CD4+ TIL-4). While immunosuppressive M2 macrophages36 were highly prevalent, pro-inflammatory, anti-tumour M1 macrophages were largely absent. A small cluster of samples (TIL-5) was characterized by a dominant ratio of cytotoxic CD8+ T cells. To assess clonal T-cell expansion, we selected index cases with a high MImmScore and CD8+ T-cell ratio or with low immune infiltration. We ascertained the identity and frequency of T-cell clones by TCRβ CDR3 deep sequencing, and found that the estimated numbers of T cells were markedly increased in the index cases (Supplementary Table 7). Most importantly, this increase was correlated with a significant expansion of T-cell clones (increased clonality) (Fig. 5c and Extended Data Fig. 9e) and a concomitant decrease in the ratio of regulatory T cells to cytotoxic T cells (Extended Data Fig. 9f). Highly mutated samples were found to be associated with a larger number of infiltrating T cells (Fig. 5d) and an increased MImmScore (Extended Data Fig. 10a).
Next, we focused on the expression of ligand/receptor pairs on the surface of T cells and antigen-presenting cells (APCs). These molecules are either co-stimulatory and required for T-cell activation, or co-inhibitory as in the case of immune checkpoints (Fig. 5e). Most patients were either immunologically silent (clusters Tcell-0, APC-0) or immunologically active (Tcell-1, APC-1), with a highly significant overlap between the independent cluster analyses (Extended Data Fig. 10b, c). Importantly, almost all samples in APC-1 expressed CD80/CD86 and almost all samples in Tcell-1 expressed CD28. CD80/CD86 are the ligands for the CD28 receptor and a critical signal for T-cell activation.
Finally, we examined the relationships between the emerging predictive biomarkers for immune therapy and the transcriptomic immune phenotypes. We stratified patients into three categories: immunologically silent, partly active, and fully active. A sample was categorized as completely or partly active if it was a member of all or at least one of the active clusters: TIL-5, APC-1, Tcell-1, respectively. Patients in the active categories exhibited increased levels of expression biomarkers PD-L1 (ref. 37) (Fig. 5f), HLA38, and granzyme39, and had higher mutational burden (Fig. 5g), which is both a prognostic and predictive marker40. Finally, leveraging a predictive signature to immunotherapy in metastatic melanoma41, we developed a clinical-response score. As expected, immunologically active patients had significantly higher clinical-response scores (Fig. 5h).
Discussion
Decreases in the cost of sequencing have led to the widespread adoption of integrative sequencing for the study of cancer and precision oncology. Accordingly, our real-time clinical sequencing program was established to explore the practical challenges of clinical translation and at the same time to characterize the genomic landscape of advanced cancer. The resulting MET500 cohort represents the first assessment of the genetic and transcriptomic heterogeneity across a wide range of metastatic cancers.
The distribution of mutation frequencies across diverse lineages of metastatic cancers is extremely long-tailed, with relatively few genes mutated at a high rate. We found that 12.2% of our cases harboured potentially pathogenic germline variants, most of which (75%) were related to DNA repair pathways. Mutations in DNA repair pathways have therapeutic implications: hypermutated tumours may respond to immune checkpoint inhibitors42, while HR deficiency could suggest sensitivity to PARP inhibitors42,43. The high prevalence of likely pathogenic germline variants suggests that metastatic patients should be considered for genetic counselling and associated germline testing.
By integrating whole-exome sequencing with RNA-seq, we were able demonstrate that transcriptome profiling provides clinically important and complementary molecular information. We have demonstrated how RNA-seq can be used in a clinical context to characterize gene fusions, outlier gene expression, transcriptional pathways, and the immune microenvironment. Across the MET500 cohort, 37% cases harboured a putative driver fusion, or an inactivating fusion in a tumour-suppressor gene. RNA-seq data played an important part in characterizing the transcriptional networks active in tumour cells as well as the metastatic tumour microenvironment, and suggest that metastatic tumours are significantly de-differentiated, but retain some tissue- and cancer-specific gene expression patterns. We were able to delineate two distinct types of metastasis: proliferative and EMT-like. Interestingly, proliferative tumours were associated with increased metabolism and stress response, while EMT-like tumours were associated with inflammation-related signatures.
Particularly valuable in the context of immunotherapy are mechanism-driven biomarkers that delineate discrete immune checkpoints or mechanisms of immune evasion. However, immune biomarkers need to characterize a complex disease state comprising the tumour genotype (for example, mutational burden), phenotype (for example, PD-L1 expression), and host response (for example, presence of CD8+ T cells). Towards comprehensive immunogenomic profiling, in this study we leveraged DNA- and RNA-seq data, which enabled us to characterize not only the tumour genotype but also the phenotype of the host immune response. Our results demonstrate the feasibility of using RNA-seq data to delineate immunologically and potentially clinically distinct subtypes of metastatic tumours, highlighting the potential of clinical RNA-seq for monitoring the tumour microenvironment and guiding immunotherapeutic approaches.
While this study has compared the molecular attributes of a metastatic cancer cohort with those of primary cancer cohorts, it did not utilize matched samples of primary and metastatic biopsies from individual cases. The sequencing of matched samples could illuminate further the processes behind tumour evolution, resistance to therapy, and immune interactions.
In summary, the metastatic solid tumour cohort represented in this study is a powerful complement to studies that have been done on primary cancers. Metastatic cancer is a highly heterogeneous disease at the genetic, transcriptomic, and microenvironment levels. Progress in the treatment of advanced cancer will therefore depend on our ability to learn the therapeutic implications of metastatic heterogeneity and to develop screening methods and clinical trial designs that match patients to the most promising therapies.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Patient enrolment
Sequencing of clinical samples was performed under our IRB-approved studies at the University of Michigan (Michigan Oncology Sequencing Protocol, MI-ONCOSEQ, IRB numbers HUM00046018, HUM00067928, HUM00056496). Medically qualified patients 18 years or older with metastatic disease (including advanced or refractory) that could be safely accessed by image-guided biopsy were eligible for inclusion. The MI-ONCOSEQ study was initiated in 2010 and is continuing as of July 2017. All patients provided written informed consent to obtain fresh tumour biopsies, and to perform comprehensive molecular profiling of tumour/germline exomes and tumour transcriptome. Four hundred and twenty-two patients (MO prefix) were enrolled under the MI-ONCOSEQ protocol, which included meeting with genetic counsellors at the time of consent. A ‘flexible default’ consent model was used with MO patients, which mandated disclosure of findings that directly impacted the current cancer management, but patients could choose whether to receive incidental results, including those with possible significance for family members or conditions unrelated to the current cancer. The remaining patients (TP prefix) were usually from external sites and were enrolled under a tissue profiling protocol without initial genetic counselling. TP patients were not given the option to decline germline findings, and germline results relevant to cancer were automatically reported. Once sequenced, a patient’s clinical course was updated quarterly to document clinical status and treatment decisions made by the physicians since the last follow-up.
Tissue acquisition and pathology review
Needle biopsies or surgically resected tissues were flash frozen in OCT and a section was cut for evaluation. Remaining portions of each specimen were retained for nucleic acid extraction. Haematoxylin and eosin (H&E)-stained frozen sections were reviewed by pathologists to identify cores or areas with the highest tumour content. In general, multiple sources of data were used to confirm the diagnosis and site of origin of the carcinomas, especially metastatic adenocarcinoma in these biopsies sent for next-generation sequencing. These included review of the electronic medical record for the clinical history, documentation of primary diagnosis and/or previously performed metastatic biopsy with confirmation of site of origin and the morphological assessment of H&E‐stained sections (frozen section slides in all cases), and review of concurrent formalin-fixed paraffin-embedded sections, as most of these cases had a previous documentation of metastasis with confirmation of site of origin. In no case were the frozen section slides used exclusively to determine site of origin. Immunohistochemistry using a panel of antibodies was performed on the formalin-fixed paraffin-embedded sections, especially if the biopsy was the first documentation of metastasis. The antibodies used for confirmation of site of primary origin were based on primary diagnosis; however, when clinically indicated, depending on the site of metastasis, the presence of multiple primaries and/or whether there was a long time gap between primary diagnosis and metastatic biopsy, to the best possible extent other sites of origin were excluded. Presentation at the PMTB included a review of morphological assessment and immunohistochemical results, and correlation of these results with expression analysis of the RNA-seq data, as well as mutation signatures. Sequencing results were used in a supportive fashion to reinforce the diagnosis of primary origin of the metastatic cancer.
Integrative clinical sequencing
Integrative clinical sequencing was performed using standard protocols in our Clinical Laboratory Improvement Amendments-compliant sequencing laboratory10,15. Tissues with the highest tumour content for each case were disrupted by 5 mm beads on a Tissuelyser II (Qiagen). Tumour genomic DNA and total RNA were purified from the same sample using an AllPrep DNA/RNA/miRNA kit (Qiagen). Matched normal genomic DNA from blood, buccal swab, or saliva was isolated using an DNeasy Blood & Tissue Kit (Qiagen). RNA integrity was measured on an Agilent 2100 Bioanalyzer using RNA Nano reagents (Agilent Technologies). RNA-seq was performed either by processed (poly(A)+) transcriptome or exome-capture transcriptome platform10,44. Both poly(A)+ and capture transcriptome libraries were prepared using 1–2 μg of total RNA. Poly(A)+ RNA was isolated using Sera-Mag oligo(dT) beads (Thermo Scientific) and fragmented with an Ambion Fragmentation Reagents kit (Ambion). cDNA synthesis, end-repair, A-base addition, and ligation of the Illumina index adapters were performed according to Illumina’s TruSeq RNA protocol (Illumina). Libraries were size-selected on 3% agarose gel. Recovered fragments were enriched by PCR using Phusion DNA polymerase (New England Biolabs) and purified using AMPure XP beads (Beckman Coulter). Capture transcriptomes were prepared as above without the upfront mRNA selection and captured by Agilent SureSelect Human All Exon v.4 probes following the manufacturer’s protocol. Library quality was measured on an Agilent 2100 Bioanalyzer for product size and concentration. Paired-end libraries were sequenced by an Illumina HiSeq 2000 or HiSeq 2500 (2 × 100-nucleotide read length), with a sequence coverage of 40–50 million paired reads. Reads that passed the chastity filter of Illumina BaseCall software were used for subsequent analysis.
Exome libraries of matched pairs of tumour/normal DNAs were prepared as described previously10,15. In brief, 1–3 μg of genomic DNA was sheared using a Covaris S2 to a peak target size of 250 base pairs (bp). Fragmented DNA was concentrated using AMPure beads, followed by end-repair, A-base addition, ligation of the Illumina indexed adapters, and size selection on 3% Nusieve agarose gels (Lonza). Fragments between 300 to 350 bp were recovered, amplified using Illumina index primers, and purified by AMPure beads. One microgram of the library was hybridized to the Agilent SureSelect Human All Exon v.4. The targeted exon fragments were captured and enriched following the manufacturer’s protocol (Agilent). Paired-end whole-exome libraries were analysed by an Agilent 2100 Bioanalyzer and DNA 1000 reagents and sequenced using an Illumina HiSeq 2000 or HiSeq 2500 (Illumina).
We used the publicly available software FastQC to assess sequencing quality. For each lane, per-base quality scores across the length of the reads were examined. Lanes were deemed passing if the per-base quality score box plot indicated that >75% of the reads had >Q20 for bases 1–80. In addition to the raw sequence quality, the alignment quality was also assessed using the Picard package. This allowed monitoring of duplication rates and chimaeric reads that may have resulted from ligation artefacts—crucial statistics for interpreting the results of copy number and structural variant analysis.
TCRβ repertoire deep sequencing (immunoSEQ)
Amplification and sequencing of (TCRB/IGH/IGKL/TCRAD/TCRG) CDR3 was performed using the immunoSEQ Platform (Adaptive Biotechnologies). The same DNA aliquot obtained from frozen tumour tissues was used as for the exome sequencing. The immunoSEQ Platform combined multiplex PCR with high-throughput sequencing and a sophisticated bioinformatics pipeline for (TCRB/IGH/IGKL/TCRAD/TCRG) CDR3 analysis, including internal PCR amplification controls. Duplicate PCR reactions were done on all samples with >1 μg of DNA. Computational analysis of sequencing data, including the estimation of the total number of templates and identification of clonotypes, was performed using the vendor-supplied analysis portal.
Mutation analysis
Whole-exome sequencing was performed on an Illumina HiSeq 2000 in paired-end mode and the primary base call files were converted into FASTQ sequence files using the bcl2fastq converter tool bcl2fastq-1.8.4 in the CASAVA 1.8 pipeline. The FASTQ sequence files generated were then processed through an in-house pipeline constructed for whole-exome sequence analyses of paired cancer genomes. The sequencing reads were aligned to the reference genome build hg19, GRCh37 using Novoalign Multithreaded (version 2.08.02) (Novocraft) and converted into BAM files using SAMtools (version 0.1.18). Sorting and indexing of BAM files used Novosort threaded (version 1.00.01) and duplicate reads were removed using Picard (version 1.74). Mutation analysis was performed with VarScan2 algorithms (version 2.3.2) using the pileup files created by SAMtools mpileup for tumour and matched normal samples, simultaneously performing the pairwise comparisons of base call and normalized sequence depth at each position. To detect SNVs, filtering parameters including coverage, variant read support, variant frequency, P value, base quality, homopolymer, and strandedness were applied. For indel analysis, Pindel (version 0.2.4) was used on tumour and matched normal samples; indels common to both samples were classified as germline, indels present in tumour but not in normal were classified as somatic. Finally, the list of candidate indels as well as somatic and/or germline mutations was generated by excluding synonymous SNVs. ANNOVAR69 was used to functionally annotate the detected genetic variants, and positions were based on Ensemble66 transcript sequences.
Tumour content for each tumour exome library was estimated from the sequence data by fitting a binomial mixture model with two components to the set of most likely SNV candidates on two-copy genomic regions. The set of candidates used for estimation consisted of coding variants that (1) exhibited at least 3 variant fragments in the tumour sample, (2) exhibited zero variant fragments in the matched benign sample with at least 16 fragments of coverage, (3) were not present in the Single Nucleotide Polymorphism database (dbSNP), (4) were within a targeted exon or within 100 bp of a targeted exon, (5) were not in homopolymer runs of four or more bases, and (6) exhibited no evidence of amplification or deletion. To filter out regions of possible amplification or deletion, we used exon coverage ratios to infer copy-number changes, as described below. Resulting SNV candidates were not used for estimation of tumour content if the segmented log-ratio exceeded 0.2 in absolute value. Candidates on the Y chromosome were also eliminated, because they were unlikely to exist in two-copy genomic regions. Using this set of candidates, we fitted a binomial mixture model with two components using the R package flexmix, version 2.3.8. One component consisted of SNV candidates with very low variant fractions, presumably resulting from recurrent sequencing errors and other artefacts. The other component, consisting of the likely set of true SNVs, was informative of tumour content in the tumour sample. Specifically, under the assumption that most or all of the observed SNV candidates in this component were heterozygous SNVs, we expected the estimated binomial proportion of this component to represent one-half of the proportion of tumour cells in the sample. Thus, the estimated binomial proportion as obtained from the mixture model was doubled to obtain an estimate of tumour content.
Recurrently mutated genes were classified as putative oncogenes and tumour suppressors. Initially, we divided them into tumour suppressors and oncogenes on the basis of the proportion of inactivating (two-hit, nonsense) aberrations (relative to all other aberrations, including missense mutations and amplifications) using a heuristic cut-off of 0.65. This initial classification was then reviewed on the basis of the distribution of somatic mutations in COSMIC (for example, presence of hotspots, ratio of inactivating mutations, prevalence of frameshifts, etc.), whether the mutations were putative gain or loss-of-function, and relevant gene-related literature. This resulted, for several oncogenes but not tumour suppressors, in the re-classification to tumour suppressor, namely FAT1, KMT2B, KMT2C, KMT2D, RAD50, RNF43, MSH2, SMC4, KEAP1, MUTYH, BRIP1, and VHL.
Copy-number aberration
Copy-number aberration was quantified and reported for each gene as the segmented normalized log2-transformed exon coverage ratios between each tumour sample and matched normal sample. To account for observed associations between coverage ratios and variation in GC content across the genome, lowess normalization was used to correct per-exon coverage ratios before segmentation analysis. Specifically, mean GC percentage was computed for each targeted region, and a lowess curve was fitted to the scatterplot of log2-coverage ratios versus mean GC content across the targeted exome using the lowess function in R (version 2.13.1) with smoothing parameter f = 0.05. Partly redundant sequencing of areas of the genome afforded the ability for cross-validation of findings. We cross-validated exome-based point mutation calls by manually examining the genomic and transcriptomic reads covering the mutation using the University of California, Santa Cruz (UCSC) Genome Browser. Likewise, gene fusion calls from the transcriptome data could be further supported by structural variant detection in the genomic sequence data, as well as copy-number information derived from the genome and exome sequencing.
Mutation burden estimation
The Varscan2 processed VCF files from 33 TCGA cohorts were downloaded from the GDC data portal and lifted-over from the GRCh38 to GRCh37 reference genome using CrossMap to compare with MET500. The mutations were filtered by coverage (at least 10×) and variant allelic fraction (at least 6%). These mutations were further narrowed down to be within 10 bp of the Agilent All Exon v.4 captured regions. The mutation burden was estimated as (total mutation/total covered bases) × 106. Finally, we identified 20 common cohorts between MET500 and TCGA.
Comparisons of gene-level aberration frequency
To compare the mutation frequency between primary and metastatic tumours, we first identified 20 analysis cohorts (tumour types) shared between the TCGA and MET500 projects. For each of those cohorts, we obtained aberration frequencies for selected most recurrently/ubiquitously mutated oncogenes (TP53, PTEN, RB1) and tumour suppressors (KRAS, PIK3CA, GNAS). We compared the aberration frequencies for each gene within each tumour type with Fisher’s exact test for a total of 120 dependent tests. To correct for multiple testing, we applied the BY method of Benjamini, Hochberg, and Yekutieli.
RNA-seq data analysis
Strand-specific RNA-seq libraries were analysed using the CRISP clinical RNA-seq pipeline, which comprises expression analysis, virus detection, and structural variant detection using a separate tool CODAC (M.C., Y.M.W., D.R.R. and A.M.C., manuscript in preparation). CRISP is composed of several tasks: pre-alignment QC, read grooming, alignment, post-align QC, quantification. Notably, fusion calling is not a part of CRISP and is done independently using multiple of CRISP output files. The components were either chosen on the basis of their performance and robustness (for example, featureCounts) or rewritten from scratch (HPSEQ, sepath, PaPy). In brief, reads that passed vendor QC thresholds were trimmed of adaptor sequences and aligned to the GRCh38 reference genome with added sequences for known oncogenic viruses and a transcript reference database on the basis of Gencode v.23. STAR_2.4.0g1 was used for alignment with the following settings (not-default): outSAMstrandField: None; alignSJoverhangMin: 8; alignSJDBoverhangMin: 3; scoreGenomicLengthLog2scale: 0; alignIntronMin: 20; alignIntronMax: 1000000; alignMatesGapMax: 1000000. For Chimeric alignment, used for structural variant detection, the following settings were applied: alignIntronMax: 400000; alignMatesGapMax: 400000; chimSegmentMin: 10; chimJunctionOverhangMin: 1; chimScoreSeparation: 0; chimScoreJunctionNonGTAG: 0; chimScoreDropMax: 1,000; chimScoreMin: 1.
The chimaeric output was analysed for chimaeric junction supported by spanning and encompassing reads, and then filtered. CRISP and CODAC were tuned to perform optimally with a custom set of reference transcripts based on Gencode v.23 (MOTR). This set lacked many questionable transcripts such as non-coding transcripts overlapping coding exons, transcripts linking two protein-coding genes, read-through transcripts, non-coding transcripts for protein-coding genes, strange isoforms with extremely long exons, intron-retention isoforms, etc. Many of those could decrease the reliability of gene-based expression estimates and, most importantly, limit the ability to detect fusions as the detection of chimaeric reads rests on the assumption that the ‘fused’ genes are not part of a known isoform. Also, by eliminating coding–non-coding overlaps, the coding (PROT) and non-coding (NONC) portions of MOTR were disjointed, simplifying many downstream analyses. In addition to standard chromosomes and unplaced contigs, we included several sequences of laboratory contaminants (for example, mycoplasma), pathogens (for example, tuberculosis), and oncogenic viruses (for example, human papilloma virus). A custom pipeline inspected the reads aligned to viral sequences (which are often problematical, for example highly repetitive). The sequencing approach used in this study did not detect HIV with reliability or sensitivity, precluding its use clinically in this regard. Paired-end reads were trimmed from adaptor sequences (in-house tool) and processed twice, once for linear alignment and downstream expression profiling, and a second time for chimaeric alignment. Before chimaeric alignment, the reads were ‘merged’: that is, if the two mates overlapped because the sequenced RNA fragment was shorter than twice the read length, the reads were combined into a synthetic single long read. This greatly improved the sensitivity by which STAR could detect a chimaeric junction. CODAC could use both alignment files to call fusions.
Fusion calling (CODAC)
Our fusion-calling pipeline allowed us to detect fusions regardless of the location of breakpoints within gene bodies, which in turn allowed us to detect a wider range of aberrations including gain-of-function fusions and truncating loss-of-function fusions. The chimaeric alignments from STAR were aggregated using custom software (M.C., Y.M.W., D.R.R. and A.M.C., manuscript in preparation) and filtered for recurrent artefacts, breakpoints within problematic repetitive regions, segmental duplications, and possible alignment errors (mismatches, pseudogenes). Variable cut-offs of supporting reads were required, depending on the breakpoint position (higher if the breakpoint was in problematic regions), with a minimum of three high-quality spanning reads defined as having a long alignment (>60 bp) on both ends of the breakpoint and a low number of mismatches (sequencing errors, SNPs, or mutations), and low repetitiveness score.
Computational fusion validation
We performed both computational and experimental validations to estimate the specificity of our fusion-calling algorithm CODAC (M.C., Y.M.W., D.R.R. and A.M.C., manuscript in preparation). First, we ran the algorithm on 50 randomly selected GTEX libraries (normal samples) and found that, for 90% of the cases, at most, two false-positive fusions were called. Next, we compared the quality of clinically reportable or pathogenic fusion calls with the remaining fusion calls, using a compound fusion-quality score taking into account several quality metrics: number of spanning reads, alignment quality, repetitiveness of the DNA, presence of splice donor–acceptor motif, etc. Likely pathogenic and non-pathogenic fusions were very similar in terms of overall quality. To further validate our algorithm, we plotted the number of fusions and number of copy-number breakpoints per sample, and observed that the number of DNA and RNA-breakpoints was highly correlated.
Experimental fusion validation
To validate the sensitivity/specificity of the fusion-calling pipeline, we randomly selected 17 private pathogenic fusions from Supplementary Table 6, and 40 fusions from 2 cases with highly rearranged genomes (20 random fusions per case, ranging from high to low supporting reads) and performed reverse transcription PCR followed by Sanger sequencing of the fusion fragments. cDNA was synthesized using a SuperScript III First-Strand Synthesis SuperMix Kit according to the manufacturer’s instructions (Thermo Fisher Scientific/Invitrogen). PCR amplification was performed using fusion-specific primers (synthesized by IDT) and a HotStarTaq Plus Master Mix Kit (Qiagen). PCR products were subjected to electrophoresis and purified using a QIAEX II Gel Extraction Kit (Qiagen) before Sanger sequencing. Sanger sequencing was performed by the University of Michigan DNA Sequencing Core. Fifty-six out of the 57 candidates (98.2%) were validated by this approach.
Adjustment
Capture and polyA expression levels could be almost perfectly adjusted across the whole dynamic range of gene expression. The adjustment was based on the (shown valid) assumption that most of the differences were due to systematic differences: for example, capture efficiency or transcript stability. These differences were estimated from data of 400 paired polyA and capture libraries using a linear model with shrinkage and variance pooling using limma with Voom precision estimates (M.C., Y.M.W., D.R.R. and A.M.C., manuscript in preparation). Correction of systematic biases through a linear model (Supplementary Methods) eliminated almost all of the apparent differences, and resulted in a good correlation between capture and polyA RNA-seq data (r = 0.97), indicating high overall reproducibility and enabling us to jointly analyse both datasets.
Selection of marker genes
We followed a multi-step procedure to identify expression markers for normal tissues and primary tumour types. First, we assembled expression compendia for both tasks. The normal tissue compendium included all the data available from GTEX and the Human Proteome Atlas, for a total of 36 different tissues/organs. The primary cancer compendium included all data from the TCGA for 33 primary tumour types. Then for each tissue/cancer type we sought to find protein-coding genes that fulfilled the following criteria: (1) highly expressed for that given tissue/cancer type; (2) upregulated in that tissue compared with other tissues/cancer types; (3) expressed in only a few tissues/cancer types; (4) not redundant with other better markers for that tissue/cancer types.
We implemented an algorithm (available at https://github.com/mcieslik-mctp/) identifying such genes using two statistical criteria: enrichment Z-score (that is, how much higher a gene was expressed in a target tissue/tumour relative to other tissues tumours) and Hoyer’s sparsity (that is, sparsity was highest if a gene was expressed only in one tissue type and lowest if it was expressed in all tissues). Next, all genes were ranked according to both analyses and an average rank computed. This ranked list of genes was traversed from the putative markers (highest rank) to worst (lowest rank) to populate a shortlist of uncorrelated markers, beginning with the top-ranking gene which was automatically included in the shortlist. For each subsequent evaluated marker, a correlation with all other previously included markers was computed and a cut-off applied (Spearman’s ρ > 0.85). This process was repeated until 50 shortlisted marker genes were identified for each tissue. We applied this algorithm to the two datasets (that is, 36 normal tissues and 33 cancer types), which resulted in two sets of marker genes, Nt36 and Tc33, respectively.
Expression signature analysis
To estimate signature/hallmark/pathway activity levels, we used two different approaches. The first relative (‘intrinsic’) approach was similar to GSVA and ssGSEA in that it estimated the activity of a pathway in one sample relative to the activity of that pathway in a cohort of other samples. In our case, the cohort was the MET500. The expression of each gene in the pathway was transformed into percentiles and the activity of each pathway was calculated as the average percentile score of all genes in a pathway minus 50 (that is, the expected median activity of a pathway). The intrinsic score has been shown to correlate very well with GSVA estimates. All GSVA analyses used default settings. The second, ‘extrinsic’ approach was analogous to the relative (intrinsic) approach, but expression percentiles for each gene were calculated, not relative within the MET500 cohort, but compared with the expression levels of that gene in over 8,000 samples from 36 normal tissues (GTEX and HUPA), each tissue represented by the same number of transcriptomes (oversampling for some of the less well studied tissues). For all pathway analyses, we used the gene sets provided by MSigDB, including the hallmark sets and the experimentally perturbed datasets.
MImmScore
The MImmScore is an aggregate analysis of immune infiltration based on the expression of multiple immune-related genes. It is derived from the ‘immune signature’ genes used in the ESTIMATE method in ref. 30. The 141 genes included in this set fulfil several criteria: (1) are common to RNA-seq and major microarray platforms, (2) are highly expressed in haematopoietic cells compared with normal tissues, (3) are upregulated in highly immune infiltrated ovarian tumours, and (4) are non-redundant. A signature derived from the expression of these genes has been shown to track with the amount of non-tumour cells on the basis of EpCam expression. We used all 141 ‘immune score’ genes, but chose a different statistical approach to transform their relative expression levels into a compound score. We used the so-called ‘inverse normal transformation’ method that is common in eQTL and other regression analyses. In brief, this rank-based method minimizes the influence of outliers, and genes with non-normally distributed expression levels, on the compound score. The first step is to transform the variable (that is, gene expression across samples) to ranks and subsequently percentiles. The percentiles are then transformed to standard normal deviates using the inverse normal (or probit function). In other words, the expression of each gene is made to follow the normal distribution. The MImmScore is simply computed by summing, for each sample, the standard normal deviates for each gene. The MImmScore is hence analogous to the ‘intrinsic’ signature expression analysis, with the added step of converting percentiles to the Z-scores associated with a given probability (using the standard qnorm function in R). Samples with high levels of immune infiltration will have many genes expressed above the 50th percentile (Z-score > 0) and hence the overall MImmScore will be positive.
Pathogenic germline variant analysis
Pathogenicities of germline variants were determined through review of the published literature, public databases including but not limited to ClinVar, the Human Genome Mutation Database, Leiden Open Variation Databases, and variant-specific databases (for example, International Agency for Research on Cancer TP53 Database, International Society for Gastrointestinal Hereditary Tumours mutation databases). Only cancer-relevant germline variants that had been previously categorized as pathogenic in ClinVar, or adjudicated at the PMTB as pathogenic, were disclosed on the clinical report. These clinically reported germline variants are shown in Supplementary Table 4. Variants with conflicting pathogenicity reports and variants not previously reported were considered to be of uncertain significance and not considered for disclosure. After disclosure, familial testing was recommended. Clinical relevance of somatic variants was investigated using an integrated approach incorporating technical considerations, (recurrence, variant allele fraction, expression levels, and predictive algorithms for pathogenicity), variant-specific information (ClinVar, published literature, and curated gene specific resources), as well as published correlations of drug/variant sensitivity profiles. Considerations of tumour heterogeneity, including clonal versus subclonal mutations, were addressed by comparing variant allele fractions and copy-number estimates for each of the mutations to post-sequencing estimates of tumour content derived from SNV and copy-number analyses.
PMTB activity
A fortnightly, multidisciplinary PMTB interpreted and deliberated on sequencing results for each patient. PMTB participants included paediatric and adult oncologists, geneticists, pathologists, biologists, bioinformaticians, bioethicists, genetic counsellors, study coordinators, and ad hoc expertise. Selected findings underwent additional independent Clinical Laboratory Improvement Amendments-validated testing, and summarized results were disclosed to treating oncologists and families by the clinical sequencing team, board-certified clinical geneticists, and/or counsellors, as appropriate. For this study, potentially actionable findings were defined as any genomic findings discovered during sequencing analysis that could (1) lead to a change in patient management by providing a targetable molecular aberration, (2) lead to a change in diagnosis or risk stratification, or (3) provide cancer-related germline findings that could inform patients/families about a potential future risk of various cancers.
Data availability
Sequencing data from the 500 patients enrolled in this study can be obtained from the Database of Genotypes and Phenotypes (dbGaP) under accession number phs000673.v2.p1. The MET500 web portal is available at http://met500.path.med.umich.edu. To make the data accessible to the broader scientific and clinical communities, we have made the somatic landscape searchable through a MAGI-based web interface45 at http://met500.path.med.umich.edu. All custom analysis software used in this study is publicly available on github at https://github.com/mcieslik-mctp/ and https://github.com/mctp/. All other data are available from the corresponding author upon reasonable request.
References
Mehlen, P. & Puisieux, A. Metastasis: a question of life or death. Nat. Rev. Cancer 6, 449–458 (2006)
Steeg, P. S. Targeting metastasis. Nat. Rev. Cancer 16, 201–218 (2016)
Friedman, A. A., Letai, A., Fisher, D. E. & Flaherty, K. T. Precision medicine for cancer with next-generation functional diagnostics. Nat. Rev. Cancer 15, 747–756 (2015)
Mauer, C. B., Pirzadeh-Miller, S. M., Robinson, L. D. & Euhus, D. M. The integration of next-generation sequencing panels in the clinical cancer genetics practice: an institutional experience. Genet. Med. 16, 407–412 (2014)
Shen, T., Pajaro-Van de Stadt, S. H., Yeat, N. C. & Lin, J. C. Clinical applications of next generation sequencing in cancer: from panels, to exomes, to genomes. Front. Genet. 6, 215 (2015)
Jones, S. et al. Personalized genomic analyses for cancer mutation discovery and interpretation. Sci. Transl. Med. 7, 283ra53 (2015)
Roychowdhury, S. et al. Personalized oncology through integrative high-throughput sequencing: a pilot study. Sci. Transl. Med. 3, 111ra121 (2011)
Byron, S. A., Van Keuren-Jensen, K. R., Engelthaler, D. M., Carpten, J. D. & Craig, D. W. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 17, 257–271 (2016)
Turajlic, S. & Swanton, C. Metastasis as an evolutionary process. Science 352, 169–175 (2016)
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015)
The Cancer Genome Atlas Research Network, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013)
Gagan, J. & Van Allen, E. M. Next-generation sequencing to guide cancer therapy. Genome Med. 7, 80 (2015)
Mardis, E. R. The translation of cancer genomics: time for a revolution in clinical cancer care. Genome Med. 6, 22 (2014)
Parsons, D. W. et al. Diagnostic yield of clinical tumor and germline whole-exome sequencing for children with solid tumors. J. Am. Med. Assoc. Oncol. 2, 616–624 (2016)
Mody, R. J. et al. Integrative clinical sequencing in the management of refractory or relapsed cancer in youth. J. Am. Med. Assoc. 314, 913–925 (2015)
Wagle, N. et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov. 2, 82–93 (2012)
Pritchard, C. C. et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N. Engl. J. Med. 375, 443–453 (2016)
Palanisamy, N. et al. Rearrangements of the RAF kinase pathway in prostate cancer, gastric cancer and melanoma. Nat. Med. 16, 793–798 (2010)
Robinson, D. R. et al. Functionally recurrent rearrangements of the MAST kinase and Notch gene families in breast cancer. Nat. Med. 17, 1646–1651 (2011)
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014)
Agaram, N. P., Zhang, L., Sung, Y. S., Singer, S. & Antonescu, C. R. Extraskeletal myxoid chondrosarcoma with non-EWSR1–NR4A3 variant fusions correlate with rhabdoid phenotype and high-grade morphology. Hum. Pathol. 45, 1084–1091 (2014)
Weinreb, I. et al. Novel PRKD gene rearrangements and variant fusions in cribriform adenocarcinoma of salivary gland origin. Genes Chromosom. Cancer 53, 845–856 (2014)
Amir, A. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013)
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013)
Liberzon, A. et al. The Molecular Signatures Database hallmark gene set collection. Cell Syst. 1, 417–425 (2015)
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011)
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013)
Vaske, C. J. et al. Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics 26, i237–i245 (2010)
López-Novoa, J. M. & Nieto, M. A. Inflammation and EMT: an alliance towards organ fibrosis and cancer progression. EMBO Mol. Med. 1, 303–314 (2009)
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013)
Attig, S. et al. Simultaneous infiltration of polyfunctional effector and suppressor T cells into renal cell carcinomas. Cancer Res. 69, 8412–8419 (2009)
Nakano, O. et al. Proliferative activity of intratumoral CD8+ T-lymphocytes as a prognostic factor in human renal cell carcinoma: clinicopathologic demonstration of antitumor immunity. Cancer Res. 61, 5132–5136 (2001)
Ruffini, E. et al. Clinical significance of tumor-infiltrating lymphocytes in lung neoplasms. Ann. Thorac. Surg. 87, 365–371 (2009)
Boon, T., Coulie, P. G., Van den Eynde, B. J. & van der Bruggen, P. Human T cell responses against melanoma. Annu. Rev. Immunol. 24, 175–208 (2006)
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015)
Sica, A., Schioppa, T., Mantovani, A. & Allavena, P. Tumour-associated macrophages are a distinct M2 polarised population promoting tumour progression: potential targets of anti-cancer therapy. Eur. J. Cancer 42, 717–727 (2006)
Patel, S. P. & Kurzrock, R. PD-L1 expression as a predictive biomarker in cancer immunotherapy. Mol. Cancer Ther. 14, 847–856 (2015)
van Houdt, I. S. et al. Favorable outcome in clinically stage II melanoma patients is associated with the presence of activated tumor infiltrating T-lymphocytes and preserved MHC class I antigen expression. Int. J. Cancer 123, 609–615 (2008)
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006)
Rizvi, N. A. et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015)
Ulloa-Montoya, F. et al. Predictive gene signature in MAGE-A3 antigen-specific cancer immunotherapy. J. Clin. Oncol. 31, 2388–2395 (2013)
Mateo, J. et al. DNA-repair defects and olaparib in metastatic prostate cancer. N. Engl. J. Med. 373, 1697–1708 (2015)
Helleday, T., Petermann, E., Lundin, C., Hodgson, B. & Sharma, R. A. DNA repair pathways as targets for cancer therapy. Nat. Rev. Cancer 8, 193–204 (2008)
Cieślik, M. et al. The use of exome capture RNA-seq for highly degraded RNA with application to clinical cancer sequencing. Genome Res. 25, 1372–1381 (2015)
Leiserson, M. D. et al. MAGI: visualization and collaborative annotation of genomic aberrations. Nat. Methods 12, 483–484 (2015)
Acknowledgements
This work was supported by a National Institutes of Health (NIH) Clinical Sequencing Exploratory Research Award NIH 1UM1HG006508. Other sources of support included the Prostate Cancer Foundation, Stand Up 2 Cancer (SU2C)-Prostate Cancer Foundation Prostate Dream Team Grant SU2C-AACR-DT0712, Early Detection Research Network grant U01 CA214170, and Prostate SPORE grant P50 CA186786. A.M.C. is a Howard Hughes Medical Institute Investigator, A. Alfred Taubman Scholar, and American Cancer Society Professor. M.C. is supported by a PCF Young Investigator Award. We acknowledge Y. Ning, R. Wang, X. Dang, M. Davis, L. Hodges, J. Griggs, J. Athanikar, C. Brennan, C. Betts, J. Chen, S. Kalyana-Sundaram, K. Giles, and R. Mehra for their contributions to this study. Over 100 physicians referred patients to this study and we acknowledge the following: K. Cooney, M. Hussain, S. Urba, N. Henry, V. Sahai, D. Simeone, C. Lao, J. Smerage, M. Caram, M. Burness, G. Kalemkerian, C. Van Poznak, M. Wicha, R. Buckanovich, J. Bufill, P. Grivas, P. Hu, A. Morikawa, P. Palmbos, B. Redman, F. Feng, G. Hammer, S. Merajver, and A. Pearson. We thank S. Roychowdhury and K. Pienta for help in protocol development for the MI-ONCOSEQ program. Most importantly, we recognize the generosity and kindness of the cancer patients and their families for participating in this study.
Author information
Authors and Affiliations
Contributions
D.R.R., Y.-M.W., and X.C. coordinated clinical sequencing. R.J.L., M.C., and P.V. developed the bioinformatics analysis. J.S. coordinated sample procurement, L.P.K., D.L., and S.A.T. led the histopathology analysis. D.C.S., S.S., M.M.Z., A.A., R.C., F.W., L.H.B., R.J.M., N.R., A.F.S., and D.F.H. coordinated patient recruitment. E.R. was the lead study coordinator. J.E., V.R., E.M.S., and J.I. provided genetic counselling and assessment of PPGMs, and J.V. and K.O. analysed relative risk assessment. J.S.R. coordinated the bioethics component. M.T. and A.M.C. coordinated IRB protocol development. D.R.R., Y.-M.W. and C.K.-S. prepared PMTBs. E.C., M.T., D.F.H., D.R.R., and Y.-M.W. implemented the clinical tiering of molecular aberrations. Y.-M.W., D.R.R., M.C., and A.M.C. developed the figures and tables. A.M.C., M.C., D.R.R., and Y.-M.W. wrote the manuscript with input from all authors. A.M.C. and M.T. designed and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Bova, P. Robbins and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Demographics of the MET500 cohort and summary of common genetic aberrations.
a, Gender distribution of the MET500 cohort. b, Age distribution of the MET500 cohort. c, Bubble plot of clinically actionable genetic aberrations. Genes have been divided by putative gain-of-function (oncogene, red) or loss-of-function (tumour suppressor, blue) status. Common aberrations are defined as those observed in five or more MET500 analysis cohorts (Fig. 1c), restricted aberrations are found in fewer than five analysis cohorts. Bubble area is proportional to the observed frequency of the aberration across the MET500 cohort. d, Comparison of genetic aberration frequencies (SNVs, indels, amplifications, predicted homozygous deletions) between primary (TCGA) and metastatic (MET500) tumours for select tumour suppressors (left) and oncogenes (right). TCGA data for the primary cancer cohorts were obtained from the cBio portal. Nominal statistical significance is based on Fisher’s exact test. Statistically significant differences in frequencies after correction for multiple dependent tests using the Benjamini–Yekutieli procedure are indicated as circles, insignificant differences are shown as triangles.
Extended Data Figure 2 Analysis of pan-cancer metastatic transcriptomes.
a, Structural rearrangements in metastatic genomes. Distribution of the number of fusions per case is plotted across the MET500 by analysis cohort (see Fig. 1c for cancer abbreviations). The y axis is truncated at 100 fusions. Error bars show the range from Q1 − 1.5 × IQR to Q3 + 1.5 × IQR, where Q1 is the first quartile, Q3 is the third quartile, and IQR is the interquartile range. b, Summary Circos diagrams of predicted inactivating fusions for select tumour-suppressor genes across the cohort. Arc end positions indicate the chimaeric junctions; colours indicate type of rearrangement. Black, tandem duplication; blue, translocation; red, inversion; grey, signifies that multiple close junctions were detected. c, The t-SNE plot for the TCGA pan-cancer meta-cohort (a random selection of cases from each primary tumour type) on the basis of the expression of tumour-type-specific marker genes (same genes as in Fig. 4a). d, The t-SNE plot for the MET500 samples coloured by biopsy site (same samples as in Fig. 4a, there coloured by cancer type). e, Average percentile expression of tissue-specific genes in normal tissues, primary cancers, and metastases. Error bars, s.d. Significance tests were done for all normal–primary and primary–metastasis pairs of samples; all comparisons were significant (P < 0.01) according to a two-tailed t-test, with the exception of those indicated with NS. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma and endocervical adenocarcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B-cell lymphoma; ESCA, oesophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LAML, acute myeloid leukaemia; LGG, brain lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, ovarian serous cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectal adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumours; THYM, thymoma; THCA, thyroid carcinoma; UCS, uterine carcinosarcoma; UCEC, uterine corpus endometrial carcinoma; UVM, uveal melanoma.
Extended Data Figure 3 Global activity of oncogenic signatures.
a, Activity of signatures is calculated relative to a normal tissue baseline: that is, activity scores are compared with a compendium of 36 normal tissues. Therefore, this plot represents a comparison of pathway activities between metastatic tissues and normal tissues. Increased activity (positive difference, red) or decreased activity (negative difference, blue) indicates that the signature genes are on average more (or less) expressed in a metastatic tumour sample relative to the baseline (in average percentile point difference labelled ‘% diff activity’). Samples (columns) are ordered from left to right by decreasing average activity difference (column averages: that is, the aggregate score in b). b, Box plots summarizing the aggregate scores (column averages of ‘% diff activity’) in a. Analysis cohorts are ordered left to right by median aggregate scores. Error bars show the range from Q1 − 1.5 × IQR to Q3 + 1.5 × IQR.
Extended Data Figure 4 Relative activity of oncogenic signatures.
Hierarchically clustered heatmap of activity scores for the most variable oncogenic signatures. In contrast to Supplementary Figure 7, here activity scores are computed intrinsically: that is, relative to other samples in the MET500 (like ssGSEA or GSVA), which represents a relative comparison between different patients/samples. Red indicates that a signature is more active (in percentile points) for a given sample relative to the median activity; blue indicates that a signature is less active for a given sample.
Extended Data Figure 5 Activity of cancer hallmarks in metastatic cancers.
Clustered heatmaps of activity scores for the 50 MSigDB cancer hallmarks are shown. a, Gene expression patterns of cancer hallmark pathways. Average increase (red) or decrease (blue) in the relative expression levels (percentiles) of transcriptional signatures associated with the hallmarks of cancers. b, Activity scores are calculated relative to a compendium of 36 normal tissues, which represent a comparison of hallmark activities between metastatic tissues and normal tissues (analogous to Supplementary Figure 7 but for a different gene set). Red indicates that a signature is more active (in percentile points) for a given sample relative to the median activity; blue indicates that a signature is less active for a given sample.
Extended Data Figure 6 Discovery of oncogenic meta-signatures.
Relative activity scores were computed for all experimental signatures in the MSigDB database across the MET500 cohort. The signatures were clustered into 25 meta-signatures on the basis of their activity profiles across the MET500. For each of the 25 meta-signature clusters, the 5 most variable signatures were selected. Red indicates that a signature is more active (in percentile points) for a given sample relative to the median activity across the MET500; blue indicates that a signature is less active for a given sample.
Extended Data Figure 7 Activity of the oncogenic meta-signatures.
a, Relative activity of EMT and proliferation signatures across the TCGA analysis meta-cohort. b, Relative activity of the 25 meta-signatures across MET500 samples from different biopsy sites. Red indicates that a signature is more active for a given biopsy site relative to the median activity; blue indicates that a signature is less active for a given biopsy site. c, Relative activity of the 25 meta-signatures across samples from different normal tissues. Red indicates that a signature is more active (in percentile points) for a given tissue relative to the median activity; blue indicates that a signature is less active for a given tissue. d, Correlations between the 25 meta-signatures. Correlation heatmap and hierarchical clustering showed similarities (red) and dissimilarities (blue) in the transcriptional activity of computationally derived aggregate sets of MSigDB signatures: that is, ‘meta-signatures’ across samples from the MET500 stratified by the most common primary tumour type (left) and biopsy site (right).
Extended Data Figure 8 Prediction of immune infiltration in cancer tissues.
a, Correlation between the MImmScore, a measurement of absolute immune infiltration in a tumour sample, with tumour content estimated from exome DNA-seq using CNVs and SNVs. b, Correlation between MImmScores and an analogous score for tumour-stromal infiltration. c, Correlation between a T-cell expression score summarizing the expression levels (RNA-seq-based) of marker genes CD3D, CD3E, CD3G, CD6, SH2D1A, TRAT1 and the estimated number of T cells based on T-cell repertoire profiling (DNA-based). d, Number of T cells based on T-cell repertoire profiling for index cases stratified into MImmScore low (<0) or MImmScore high (>0). Error bars show the range from Q1 − 1.5 × IQR to Q3 + 1.5 × IQR. Significance levels of Spearman’s rank correlation coefficient test: *P = 0.05–0.001, **P = 0.001–10−6, ***P < 10−6.
Extended Data Figure 9 Differential immune infiltration in various cancer types.
a, Distribution of MImmScores, a measurement of the magnitude of immune infiltration in a tumour sample, for MET500 samples/patients grouped by tumour biopsy site. b, Distribution of MImmScores across the TCGA meta-cohort, grouped by primary cancer designation. Haematological malignancies (DLBC, LAML) are included as positive control. Error bars show the range from Q1 − 1.5 × IQR to Q3 + 1.5 × IQR. c, Percentage of patients in each of the MET500 analysis cohorts with a high MImmScore, defined here as >80th percentile across the whole MET500. The total number of cases with high MImmScore is indicated above each bar. d, Same as c but for the TCGA meta-cohort. e, Correlation between the total number of T cells (templates) based on T-cell repertoire (DNA-seq) of the TCR CDR3 sequence and the number of expanded clones (an expanded T-cell clone is defined as having more than 30 cells with the same CDR3 sequence). f, Ratio of expression levels for markers of CD8+ T cells (CD8A, CD8B) and regulatory T cells (FOXP3) as a function of the total number of T cells. Significance levels of Spearman’s rank correlation coefficient: *P = 0.05–0.001, **P = 0.001–10−6, ***P < 10−6.
Extended Data Figure 10 Genomic correlates of immune infiltration.
a, Association between the MImmScore and mutation status (hypermutated samples have been defined here as having >250 non-synonymous mutations). Statistical significance of this association was determined using logistic regression. b, c, A χ2 test for independence is used to determine whether the clusterings of samples based on T-cell and APC markers are independent. Enrichment or depletion is calculated as the Pearson’s residual. Red indicates (positive enrichment) that the clusters overlap significantly. Blue indicates (depletion) that clusters tend to be mutually exclusive. Clustered heatmap of enrichment levels (χ2 table cell residuals) is shown in b. Enrichment levels for clusters for the active Tcell-1 and Tcell-4 clusters and all APC clusters (APC-1, -4 active) are shown in c.
Supplementary information
Supplementary Tables
This file contains Supplementary Table 1 (Demographics and clinical details), Supplementary Table 2 (Sequencing statistics), Supplementary Table 4 (Pathogenic germline variants in the MET500 cohort), Supplementary Table 5 (Germline mutations in metastatic cancer), Supplementary Table 6 (Pathogenic fusions in the MET500 cohort) and Supplementary Table 7 (Immune cell infiltration analyses).
Supplementary Table 3
This file contains Supplementary Table 3 (Recurrent molecular aberrations in the MET500 cohort).
Rights and permissions
About this article
Cite this article
Robinson, D., Wu, YM., Lonigro, R. et al. Integrative clinical genomics of metastatic cancer. Nature 548, 297–303 (2017). https://doi.org/10.1038/nature23306
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23306
- Springer Nature Limited
This article is cited by
-
Unveiling the unique role of TSPAN7 across tumors: a pan-cancer study incorporating retrospective clinical research and bioinformatic analysis
Biology Direct (2024)
-
Biomarker discovery with quantum neural networks: a case-study in CTLA4-activation pathways
BMC Bioinformatics (2024)
-
Conformational diversity and protein–protein interfaces in drug repurposing in Ras signaling pathway
Scientific Reports (2024)
-
Decoding the interplay between genetic and non-genetic drivers of metastasis
Nature (2024)
-
CRISPR/Cas9 model of prostate cancer identifies Kmt2c deficiency as a metastatic driver by Odam/Cabs1 gene cluster expression
Nature Communications (2024)