Abstract
Marine phytoplankton inhabit a dynamic environment where turbulence, together with nutrient and light availability, shapes species fitness, succession and selection1,2. Many species of phytoplankton are motile and undertake diel vertical migrations to gain access to nutrient-rich deeper layers at night and well-lit surface waters during the day3,4. Disruption of this migratory strategy by turbulence is considered to be an important cause of the succession between motile and non-motile species when conditions turn turbulent1,5,6. However, this classical view neglects the possibility that motile species may actively respond to turbulent cues to avoid layers of strong turbulence7. Here we report that phytoplankton, including raphidophytes and dinoflagellates, can actively diversify their migratory strategy in response to hydrodynamic cues characteristic of overturning by Kolmogorov-scale eddies. Upon experiencing repeated overturning with timescales and statistics representative of ocean turbulence, an upward-swimming population rapidly (5–60 min) splits into two subpopulations, one swimming upward and one swimming downward. Quantitative morphological analysis of the harmful-algal-bloom-forming raphidophyte Heterosigma akashiwo together with a model of cell mechanics revealed that this behaviour was accompanied by a modulation of the cells’ fore–aft asymmetry. The minute magnitude of the required modulation, sufficient to invert the preferential swimming direction of the cells, highlights the advanced level of control that phytoplankton can exert on their migratory behaviour. Together with observations of enhanced cellular stress after overturning and the typically deleterious effects of strong turbulence on motile phytoplankton5,8, these results point to an active adaptation of H. akashiwo to increase the chance of evading turbulent layers by diversifying the direction of migration within the population, in a manner suggestive of evolutionary bet-hedging. This migratory behaviour relaxes the boundaries between the fluid dynamic niches of motile and non-motile phytoplankton, and highlights that rapid responses to hydrodynamic cues are important survival strategies for phytoplankton in the ocean.
Similar content being viewed by others
Main
Understanding the distribution of phytoplankton in the ocean represents a longstanding challenge in oceanography9, owing to the complex, multi-scale interactions between biotic and abiotic processes10. Unfavourable conditions, such as light and nutrient limitation2, suboptimal temperature11 or strong turbulence5 can cause physiological stress and reduced growth, affecting species fitness and succession1,2,12. Phytoplankton have evolved diverse adaptive strategies to cope with environmental stressors, often by exhibiting plasticity in important functional traits13, including buoyancy14, and chlorophyll13 and lipid content, which they can modulate in response to ambient conditions.
Turbulence is a prevalent physical factor in the ecology of phytoplankton, and is implicated in the seasonal succession and bloom formation of phytoplankton species1,2,9, as first conceptualized in Margalef’s ‘mandala’1. Turbulence can be beneficial for phytoplankton—for example, by enhancing nutrient uptake15—but is often deleterious for motile species5,6, causing flagellar or body wall damage, reduced lipid production and impaired organelle functioning8. Because turbulence is often intermittent and localized7, frequently affecting only a few metres of the water column16, migration could be an effective turbulence-avoidance strategy. However, this hypothesis implies that phytoplankton can modulate migration based on turbulent cues, which has not been demonstrated until now.
Evidence for behavioural responses of phytoplankton to turbulence has remained scarce and limited primarily to drag reduction over cell-cycle timescales17. Here we demonstrate a rapid response of raphidophytes and dinoflagellates to a fundamental turbulent cue, the repeated overturning events associated with small-scale eddies, showing that approximately half of an upward-swimming population engages in a dive response within as little as 30 min. Because the vertical is the most effective direction for modulating migration to escape unfavourable conditions, we propose that this rapid behavioural diversification may be a bet-hedging strategy to increase chances of evading turbulent layers.
Many phytoplankton species migrate through the water column using gravitaxis18, a form of directed motility mediated by a stabilizing torque that biases swimming in the vertical direction. In flow, gravitactic cells can form thin layers19 and microscale patchiness20, yet whether cells can actively modulate gravitaxis and thus their direction of migration in response to flow has remained unknown.
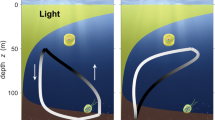
We tracked cells of the raphidophyte H. akashiwo (strain CCMP452; hereafter referred to as HA452)—a model system for gravitaxis4,18 known for its allelopathic effects21 and toxic blooms—in a millifluidic chamber (12 mm × 4 mm × 1.6 mm). Results will refer to HA452 unless otherwise indicated. The chamber could be continuously rotated around a horizontal axis by a computer-controlled motor, mimicking the overturning of cells in the ocean by the smallest (that is, Kolmogorov-scale) eddies (Fig. 1a, b). While the hydrodynamic environment in the flip chamber is not equivalent to turbulence, it captures the overturning of cells by small-scale turbulent eddies. Without rotation, cells swam strongly upwards (negative gravitaxis), accumulating at the top of the chamber (Fig. 1c).
a, The automated millifluidic chamber used to expose phytoplankton cells to turbulence-like reorientations. b, A square wave (green) comprising periodic reorientation events of amplitude π (‘flips’) and period Q = 18 s. Time series of the orientation, ψ, of a passive particle relative to the vertical in a three-dimensional isotropic turbulent flow (magenta), obtained from a direct numerical simulation. The dashed red line denotes the direction of the gravity vector, g (ψ = π/2). c, d, Equilibrium vertical distribution (each representative frame was obtained from the corresponding video after background subtraction) of HA452 over the height of the chamber, before (c) and after (d) 100 flips in the vertical plane (30 min). Hued regions (coloured layer rendering of the representative frames) extend 400 μm from the top (↑, orange) and bottom (↓, blue) of the chamber. e, f, False-colour epifluorescence micrographs of HA452 cells harvested from the top (e) and the bottom (f) of the chamber, after 30 min of flipping in the vertical plane (period Q = 18 s). g, Equilibrium vertical distributions of cells (mean of four replicates) over the chamber height upon different treatments, each lasting 30 min: still conditions (control, red, n = 1,319 cells); periodic flips in the horizontal plane (blue, n = 1,231; Q = 18 s); periodic flips in the vertical plane (green, n = 1,675; Q = 18 s); and continuous, stochastic reorientations (magenta, n = 1,932; Methods). For each case, the equilibrium vertical distribution was measured 30 min after treatment ended. Inset, the upward bias, r = (f↑ − f↓)/(f↑ + f↓), for each treatment, where f↑ and f↓ are the concentrations of cells in the top and the bottom 400 μm of the chamber (mean ± standard deviation (s.d.) of four replicates; ANOVA: P < 0.001; asterisks indicate statistically significant differences from the control, post-hoc Tukey’s honest significant difference (HSD) test reported in Supplementary Table 1).
Repeated overturning caused a striking departure from negative gravitaxis. We first imposed a series of flips, with a period (18 s) representative of the Kolmogorov timescale (0.1–30 s) (ref. 7), and specifically corresponding to a turbulent dissipation rate (ε = 3 × 10−8 W kg−1) characteristic of the ocean’s pycnocline22. When exposed to this overturning regime for 30 min (100 flips), the population split into two subpopulations: one continuing to swim upwards (negative gravitaxis; HA452(↑)) and accumulating at the top, the other swimming downwards (positive gravitaxis; HA452(↓)) and accumulating at the bottom (Fig. 1d).
We quantified the population split in terms of an upward bias index, r = (f↑ − f↓)/(f↑ + f↓), with f↑ and f↓ the concentrations of cells in the top and bottom 400 μm of the chamber, respectively (Fig. 1g). The treatment with 100 flips resulted in r = 0.13 ± 0.20 for HA452, whereas control experiments with cells held in the chamber for 30 min without flipping yielded no population split (r = 0.61 ± 0.12; Fig. 1g). A population split of the same magnitude (r = 0.22 ± 0.04; analysis of variance (ANOVA): P < 0.001; Fig. 1g and Supplementary Table 1) occurred also for non-periodic reorientations obtained from a direct numerical simulation that closely modelled the continuous, stochastic reorientation experienced by cells in isotropic turbulence (Fig. 1b; Methods), justifying the simplification of using flips in lieu of stochastic reorientations. The split occurred irrespective of continuous-light or diel-light-cycle culturing, but did not occur for a starved population (Extended Data Fig. 1a, d). Sinking was ruled out as the cause of the bottom accumulation, because HA452(↑) and HA452(↓) cells swam at comparable speeds (~74 μm s−1; Extended Data Fig. 2), indicating that the split is an active behavioural adaptation.
The population split was fast and its magnitude increased with the duration of the overturning treatment (Fig. 2a). Over the course of 30 flips (~10 min), r decreased from 0.61 to 0.30, corresponding to a 60% increase in the number of down-swimming cells and implying that even brief periods of turbulence can cause diversified migration. The response saturated after 300 flips (~2 h) at r = 0.03 ± 0.10, corresponding to a symmetric distribution of up- and down-swimmers. A similar, symmetric population split was observed in the raphidophyte Chattonella marina (CM2962) and the dinoflagellate Prorocentrum minimum (PM291) (r = −0.01 and 0.08, respectively, after 300 flips), whereas other strains tested did not split (Extended Data Fig. 3 and Extended Data Table 1).
a, Upward bias, r, as a function of the number of flips, N (period Q = 18 s, red axis denotes the equivalent time elapsed, t = NQ). For each curve, r was computed from the equilibrium vertical distribution (mean ± s.d. of four replicates) measured 30 min after treatment ended. Square symbols correspond to pre-treatment conditions (mean ± s.d. of four replicates), measured 30 min after filling the chamber for HA452 (blue, n = 960 cells) and HA3107 (green, n = 701). Flipping induces the emergence of two subpopulations with distinct migration behaviours in HA452 (blue, n = 827–1,349), as compared to the control (cells maintained in the chamber under still conditions, red, n = 1,082 cells; see top red axis for the timescale). Flipping does not affect HA3107 (green, n = 613–910). b, c, Rotation rate, ω, of HA452 cells, as a function of cell orientation to the vertical, θ, measured before (n = 2,257 cells) (b) and after (c) 30 min of flipping in the vertical plane. After flipping, the HA452(↑) (n = 1,421 cells) and HA452(↓) (n = 1,138) subpopulations have opposite stability. Solid lines (colour-coded) denote the arithmetic mean over all cell trajectories and shaded regions represent ± 1 standard error of the mean (s.e.m.). Dashed lines represent the corresponding sinusoidal fits.
The split was triggered by changes in the cell orientation relative to gravity, as revealed by the absence of splitting in HA452 (r = 0.48 ± 0.16 after 30 min; Supplementary Table 1) when flipping occurred in the horizontal plane (Fig. 1g). This result indicates that the cue for the behavioural differentiation is not fluid velocity gradients, but rather changes in the cell orientation relative to gravity, a simple yet fundamental effect of small-scale turbulent eddies that has been neglected to date.
The switch of a fraction of the cells to downward swimming corresponded to a switch in their orientational stability. This stability is quantified by the reorientation timescale, B, the characteristic time a cell takes to rotate back to its vertical equilibrium orientation once perturbed from it (Methods). A greater magnitude of B denotes less stability and the sign of B denotes upward (B > 0) or downward (B < 0) stability. For cells before flipping we found B = 10.4 ± 1.5 s (Fig. 2b). The two subpopulations that emerged after 30 min of flipping had reduced stability and, most importantly, had opposite stability, with HA452(↑) cells having B↑ = 19.3 ± 13.5 s and HA452(↓) cells having B↓ = −23.1 ± 10.2 s (Fig. 2c), confirming the active nature of the down-swimming behaviour.
The process underpinning the reversal in stability of part of the cells was a change of morphology. We harvested HA452 cells from the top and bottom of the chamber after 30 min of flipping and determined their shape through phase-contrast microscopy. We found that HA452(↑) cells were fore–aft asymmetric, narrower in the front and wider in the back (Fig. 1e and Extended Data Fig. 4). By contrast, HA452(↓) cells were fore–aft symmetric (Fig. 1f). Image analysis showed that the asymmetry was on the order of 1 μm—a minute yet robust morphological difference between the two subpopulations (Extended Data Fig. 4 and Extended Data Table 2; Methods). This fast shape change is favoured in HA452 and C. marina by the absence of a rigid cell wall, characteristic of raphidophytes23.
A cell mechanics model demonstrated that the observed shape change is sufficient to cause the stability switch. Using the cell contour, the size and position of the nucleus determined by epifluorescence microscopy, and the flow field around the cell obtained through a fluid dynamics model (Methods), we determined the position of the three points relevant for cell stability: the geometric centre, CB (where the buoyancy force acts), the centre of mass, CW (where the weight acts), and the centre of hydrodynamic stress, CH (where the resultant hydrodynamic force acts) (Fig. 3, Extended Data Fig. 4, Extended Data Table 2 and Supplementary Table 2; Methods). For HA452(↑) cells, CW was located a distance LW = 0.03 ± 0.04 μm above CB (Fig. 3a, d), making the cells top heavy. This top heaviness produces a torque, TW, which orients cells downwards. However, because of the fore–aft asymmetry, CH was located a distance LH = 1.17 ± 0.46 μm above CB (Fig. 3d and Extended Data Figs 4, 5), producing a torque TH > TW that overcame top heaviness and oriented cells upwards. By contrast, for HA452(↓) cells, LW was unchanged (0.04 ± 0.04 μm), but LH was reduced (0.24 ± 0.27 μm; Fig. 3b, e and Extended Data Figs 4, 5), because of the loss of fore–aft asymmetry. As a result, TW > TH and cells were stable downwards (Fig. 3e). In the phase space of cell stability defined by the two cellular length scales LH and LW, the loss of fore–aft asymmetry corresponds to the crossing of the line dividing upward from downward stability regimes (Extended Data Fig. 6). For comparison, for a different strain of H. akashiwo (HA3107) that had greater, upward stability (B = 4.9 ± 1.5 s; Extended Data Fig. 7b), CW was located a distance LW = 0.02 ± 0.02 μm below CB (bottom heavy; Fig. 3c, f) and CH was located a distance LH = 0.87 ± 0.23 μm above CB (Fig. 3f and Extended Data Figs 4, 5), so that both torques TW and TH stabilized cells upwards. Indeed, HA3107 did not split upon flipping (Fig. 2a).
a–c, False-colour epifluorescence micrographs of an HA452(↑) cell (representative of n = 13 cells), a HA452(↓) (n = 10) cell, and of an HA3107 cell (n = 12). White dashed lines denote the contours of the cell body and nucleus (bright orange). CW, CB, and CH are the centres of mass, buoyancy and hydrodynamic stress, respectively. d–f, Free body diagrams showing the forces acting on the cell overlaid on the numerically computed flow field around the cell (colour velocity gradient, not to scale). The swimming direction θ and the rotation rate ω are set by the competition between the gravitational and hydrodynamic torques, TW and TH, about CB. d, HA452 cells collected from the top of the chamber after 30 min of flipping are top heavy (CW above CB) and fore–aft asymmetric (CH above CB), thus TW and TH act in opposition, causing cells to orient upwards. e, HA452 cells collected from the bottom of the chamber after 30 min of flipping are fore–aft symmetric (CH = CB, TH vanishes) and top heavy, thus TW causes cells to orient downwards. f, HA3107 cells are bottom heavy (CW below CB) and fore–aft asymmetric, thus both TW and TH act to orient cells upwards.
These observations reveal that phytoplankton are capable of rapid polymorphism to diversify their migration strategy. Polymorphism is an important adaptation in phytoplankton24, yet to date has been observed mostly as large morphological changes over demographic timescales, as in the reversible loss of horns or spines to accelerate sinking17 or in chain formation5. By contrast, HA452 diversifies shape over minutes and a tiny morphological change is sufficient to drastically alter migration behaviour. These findings thus illustrate that phytoplankton are capable of precise control over movement behaviour through fine-scale control of cellular morphology. These results also indicate that small phytoplankton are capable of perceiving reorientations relative to gravity, yet the underlying mechanism remains unclear. While large (>75 μm) unicellular protists (for example, Paramecium, Tetrahymena) directly sense gravity through mechanosensitive ion channels activated by the gravitational pressure of the cytoplasm on the lower membrane25, this appears difficult in the face of thermal noise for cells in the size range of HA452 (see Supplementary Discussion).
We propose that the observed diversification of migration strategies is an escape response to avoid turbulence. A population split could benefit a migrating population when turbulence is heterogeneous over the depth of the water column, as often occurs in the ocean, where turbulent layers or patches commonly span no more than a few metres in depth7,16. When an upward-migrating phytoplankton population enters a turbulent layer, part of the population may switch to downward migration (temporarily renouncing performing photosynthesis and therefore growth) in response to the turbulent reorientations and thus avoid exposure to the stronger and potentially damaging levels of turbulence in the core of the layer. Damage from strong turbulence to motile phytoplankton has been extensively reported and includes disruption to diverse elements of their complex life cycles, including the cellular clock, mitotic cycle, nucleic acid concentration, microtubule assembly and chromosome separation, reduction in swimming velocity and loss of flagella5,8. At the same time, uncertainty about the severity of turbulence in any particular layer implies that a strategy of splitting allows some cells to attempt the ‘crossing’ of the layer, with the benefit of continued growth at the risk of turbulent damage.
This population split may be a bet-hedging strategy (in the sense described previously26) to cope with the encounter of a turbulent layer in the ocean during vertical migration. According to this scenario, by phenotypically diversifying into two subpopulations with different gravitactic behaviours, a population increases the probability that at least a fraction of the cells escape turbulence, at the cost of reduced photosynthesis because downward-swimming cells will not reach well-lit surface waters (Supplementary Discussion). This hypothesis is supported by the observation that collecting cells from the bottom of the chamber after the overturning treatment and growing them resulted in cells with the same upward stability that their ancestors had before flipping (Extended Data Fig. 1a, b). The occurrence of a split upon flipping also in a monoclonal population of HA452 (Extended Data Fig. 1c) strongly suggests that genetically identical cells can express different gravitactic phenotypes. The high intrinsic variability in the reorientation timescale B within the population—before and thus independently of the reorientation treatment—is also in line with the bet-hedging hypothesis (Extended Data Fig. 7a and Supplementary Discussion), because it allows a diversification of migration strategies through inversion of the stability of a fraction of the cells. However, other mechanisms that produce phenotypic differences cannot be ruled out, including ones that produce changes to the DNA sequence, as occurs in genetic switching27. Genetic switching may also in principle have occurred in our monoclonal population and would be difficult to distinguish from phenotypic heterogeneity28.
The escape hypothesis finds support in stress measurements, performed by quantifying the production of the free radical nitric oxide (NO) (Methods), which revealed that flipping is stressful for HA452 cells (Supplementary Discussion). After flipping, cells showed significantly higher rates of NO accumulation (3.9 ± 0.5 × 10−3 s−1 for HA452(↓); 2.9 ± 0.3 × 10−3 s−1 for HA452(↑); Fig. 4) compared with the still control (2.1 ± 0.3 × 10−3 s−1; Kruskal–Wallis test: P = 0.02; Supplementary Table 3). By demonstrating that flipping causes stress, this result supports the view of turbulence being typically deleterious for motile phytoplankton1,5,8. However, it is important to recognize that turbulence can be beneficial for some species6, in particular by enhancing mass transport to larger cells15 (Supplementary Discussion), and that considerable variability in the response to turbulence exists among species6, as also exemplified by our experiments with a range of species (Extended Data Table 1 and Extended Data Fig. 3). The extraordinary behavioural capacity of phytoplankton brought to light by our results, then, calls for an overall better understanding of the role of turbulence on these organisms.
a–c, Fluorescence intensity, measuring the production of NO in a cell collected from the top (subpopulation HA452(↑), representative of n = 4) (a) and the bottom (subpopulation HA452(↓), n = 7) (b) of the chamber, in both cases after 30 min of flipping, as well as a cell not exposed to flipping (control, n = 3) (c). d, Time series of stress accumulation, quantified as NO production (Supplementary Information), for the same three cases. Solid lines denote the arithmetic mean over all cells and the shaded regions represent ± 1 s.d. from the mean. Time represents the acquisition time under the microscope. Inset, rate of stress accumulation (mean ± 1 s.d.; Kruskal–Wallis test: P = 0.02; Dunn’s test for multiple comparison analysis reported in Supplementary Table 3).
Margalef’s mandala has remained the paradigm for the effect of turbulence on phytoplankton species succession in the ocean1,2,12. The results reported here suggest that active responses of cells to turbulence may play an important part in this paradigm, and that the spatial structure of turbulence—not just its intensity—may be a fundamental driver of phytoplankton species fitness and succession2. We thus propose that the understanding of the ocean embodied in the mandala should be augmented by a mechanistic understanding of behavioural adaptations to turbulence, to understand better how turbulence shapes phytoplankton community composition in present-day and future environmental conditions.
Methods
Cell culture
Three different strains of the raphidophyte Heterosigma akashiwo were examined: CCMP452, CCMP3374 (here referred to as HA452 and HA3374, respectively, and both obtained from the National Center for Marine Algae), and CCMP3107 (ref. 29) (courtesy of S. Menden-Deuer; here referred to as HA3107). All three strains were grown in 50 ml sterile glass tubes under continuous illumination (75 μmol m−2 s−1), in f/2 (minus silica) media. HA452 and HA3107 were cultured at 21 °C and HA3374 at 18 °C.
Additional experiments were performed (1) with cells grown under a diel light cycle (14 h light: 10 h dark) (Extended Data Fig. 1) and (2) with monoclonal cultures of HA452. The latter were grown from a single parent cell, isolated from a prior culture by means of an inoculation loop (diameter ~100 μm) developed in house. The loop was dipped into a culture to trap a thin liquid layer and microscopy was used to select the cases with only a single HA452 in the layer. Each single trapped cell was then transferred to a separate well in a 36-well plate containing fresh growth media. Monoclonal cultures were also grown under a 14:10 light cycle (Extended Data Fig. 1c).
A range of other phytoplankton species were also tested. Extended Data Table 1 summarizes the list of species, their origin, and their response to flipping. All the strains were grown in 50 ml sterile glass tubes under a 14 h light: 10 h dark cycle (75 μmol m−2 s−1), and appropriate temperature. Fibrocapsa japonica_cf (FJ2965), Chattonella sp. (CSP218), Chattonella marina_cf (CM2962), Chattonella subsalsa (CS2814), Alexandrium tamarense (AT1598 and AT2023), Karenia brevis (KB2281 and KB2229) and Prorocentrum minimum (PM696) were grown in L1 (minus silica) media at 21 °C, while Alexandrium tamarense (AT1771) and Prorocentrum minimum (PM699) were grown at 18 °C, in the same medium. Akashiwo sanguinea (AS3040), Alexandrium tamarense (AT4087) and Prorocentrum minimum (PM291) were grown at 18 °C in K/2 media (for AS3040 and AT4087 strains) and in K media (PM291 strain).
All experiments were conducted at room temperature (21 °C), between 96 h and 120 h after inoculation. This time window corresponds to the early exponential growth phase of the species, as shown for HA452 (Extended Data Fig. 8a). All experiments were carried out at a specific time of the day (between 09:00 and 12:00) to rule out any possible artefact due to the diurnal migration pattern of many phytoplankton species (as shown here for HA452; Extended Data Fig. 8b). For propagation, 2 ml of the culture was inoculated into 25 ml of fresh medium every two weeks.
Flipping chamber
All experiments were conducted in a millifluidic chamber (12 mm × 4 mm × 1.6 mm) constructed out of a transparent acrylic sheet and mounted on a supporting frame. The frame was coupled to the shaft of a stepper motor and designed to allow for full rotations from 0° to 360°. The rotation of the chamber was automated using an externally programmed controller that drove the motor, with full user control over the time series of the rotation angle. Suspensions of phytoplankton cells were gently pipetted into the chamber through one of two injection ports, which were then closed with silicone plugs. At the end of an experiment, the two ports were used to harvest cells from the top and bottom parts of the chamber using a microcapillary attached to an aspirator tube for gentle suction, for subsequent microscopic characterization. The dimension of the capillary (internal diameter = 500 μm) was chosen to avoid cell damage during sampling.
During experiments, cells in the flipping chamber were visualized using a stereoscope (Nikon SMZ1000) with a plan APO ×1 objective (0.12 NA) and a digital CMOS camera (Photron FastCam SA3). The flipping chamber was mounted on a translation stage, the position of which could be controlled using micrometre screws along all three axes. The camera was focused on a plane perpendicular to the rotation axis and halfway between the two chamber walls. The depth of focus was 750 μm, ensuring that cells were more than 400 μm (>50 cell radii) from the front and back walls of the chamber, to eliminate wall effects. Any small residual wall effects that may still have occurred would have been present for the entire duration of an experiment, and thus could not have caused the population split.
Images were acquired at 60 frames per second. The suspension was uniformly illuminated using a single 627 nm LED (0.1 W) mounted just outside of the flipping chamber. Neither of the two H. akashiwo strains tested showed any phototactic bias to wavelengths of light in the red spectrum, in agreement with literature30. All experiments were conducted under diffused room light settings, to avoid possible photo-responses.
For each treatment, a control experiment was performed in which cells were observed in the flipping chamber without rotation, for the same duration as the treatment. The vertical distribution of cells in these control experiments was quantified at regular intervals to ascertain that the upward bias of cells in the absence of overturning remained constant (Fig. 2a, red line, and Extended Data Fig. 8b).
Overturning experiments
The range of length scales characteristic of a given turbulent flow is quantified by the Taylor Reynolds number, Reλ = uRMS λ/ν, where λ = uRMS (15ν/ε)1/2 is the Taylor length scale, uRMS is the root-mean-square fluid velocity, and ε is the turbulent energy dissipation rate. Periodic flipping consisted of multiple, rapid overturnings of the chamber (180° in 3 s), each followed by 15 s at rest. The resulting period of 18 s corresponds to the Kolmogorov timescale τη = (ν/ε)1/2, associated with ε = 3 × 10−8 W kg−1. This value is typical of the ocean pycnocline22 and falls within the typical range of values for ocean turbulence7 (10−9–10−5 W kg−1).
To drive the time series of the angular orientation of the flipping chamber relative to the vertical, ψ(t), in the continuous, stochastic overturning experiments, we used the time history of the angular orientation of a small passive sphere in homogeneous isotropic turbulence, quantified from a direct numerical simulation20 at Reλ = 65 (time history courtesy of M. Cencini and G. Boffetta).
Vertical cell distribution
Histograms of normalized cell concentration in the vertical direction inside the flipping chamber, within the region captured by the camera (4 mm × 4 mm), were obtained by identifying the positions of the centroids of individual cells, imaged in the mid-chamber plane (that is, equidistant from the two side-walls perpendicular to the imaging axis). After the end of every flipping experiment, we allowed the population to reach its equilibrium distribution over the vertical by waiting 30 min. This waiting time was chosen conservatively based on the observation that the concentration profile stabilizes already after ~5 min (Extended Data Fig. 7d), and the consideration that cells swimming upward at ~50 μm s−1 (Extended Data Fig. 2) cover the depth of the flipping chamber (4 mm) in <2 min.
Upward bias index
To quantify the asymmetry in cell distribution over the vertical, we computed the upward bias r = (f↑ − f↓)/(f↑ + f↓), where f↑ and f↓ are the numbers of cells in the top 400 μm and the bottom 400 μm of the chamber, respectively. A symmetric distribution of cells corresponds to r = 0, whereas preferential upward-swimming corresponds to r > 0 and preferential downward swimming to r < 0. The two subpopulations HA452(↑) and HA452(↓) were composed of HA452 cells collected from the top/bottom of the chamber after 30 min of flipping. Control experiments consisted of cells held in the chamber for 30 min without flipping (the same duration as flipping experiments with 100 flips).
Cell tracking
For tracking, cell locations were determined by image analysis based on intensity thresholding using MATLAB (MathWorks) routines. Cell trajectories, obtained from movies recorded at 60 frames per second, were assembled by linking the locations of cells in subsequent frames, based on proximity and kinematic predictions from previous time steps, using automated software31.
Quantification of cell stability
To determine cell stability, we quantified the rotation rate ω of cells as a function of their orientation θ relative to the vertical. This is an established method18 for quantifying the reorientation timescale B, and is based on the fact that greater stability will cause faster reorientation towards the stable orientation after a cell is perturbed from it. To this end, we tracked individual cells over 15 s immediately following a single flip (which provided the perturbation from the stable orientation), and averaged their rotation rate over all cells as a function of θ. Given that the smallest dimension of the field of view is along the optical axis (750 μm, the depth of field) and that cells swam on average at 75 μm s−1, an estimated residence time in the field of view is ~10 s. Tracked trajectories in our data set are as long as 22 s, with an average of 5.5 s, which is sufficient to compute the stability parameter B. The resulting data for ω(θ) were well fitted with a sinusoidal function of the form Acos (θ + κ), where we imposed a phase shift κ equal to −π/2 for the top population and π/2 for the bottom population (simultaneously fitting both A and κ showed consistent results for this approach). We determined the reorientation timescale from the best-fit sinusoid as B = (2A)−1 cos (π/2 + κ) (ref. 18). To account for heterogeneity in the population stability (Supplementary Information), resulting in some cells reorienting faster than others, we separately quantified B for the first 5 s and the subsequent 10 s (Extended Data Fig. 7a, c). HA452 cells in the flipping chamber swam in helical patterns, characteristic of many phytoplankton species32. However, the helical component was averaged out using a 1-s moving average to reduce noise in the calculation of the stability parameter B.
Axial symmetry, fore–aft asymmetry, and centre of buoyancy
To determine the morphology of cells sampled from the millifluidic chamber, we used single-cell imaging with an inverted microscope (Nikon TE2000) equipped with a ×20 or ×40 objective and an Andor iXon Ultra 897 camera. We used phase contrast microscopy to determine the cell orientation, based on the location of the flagellum on the anterior part of the cell, and the cell contour. To assess the axial symmetry of the cell shape, we acquired images of the cross-section of ten randomly chosen HA452 cells. This was achieved by acquiring videos of each cell over tens of seconds, manually refocusing as needed to maintain the cells in focus. From each video, we identified the frames in which cells swam vertically—changing swimming plane—based on the fact that the cell boundary is in sharp focus when a cell swims vertically out of the imaging plane, but is blurry due to optical diffraction when it swims out of the imaging plane at an angle. The frame of the video in which the cell had the maximum diameter was then chosen to extract the cell’s cross-section (‘top view’, Extended Data Fig. 4d). Using these data, we quantified the ratio (R) between the major (bx) and minor (by) semi-axes of the ellipse that best fitted the top-view cross-section (Extended Data Fig. 4). This yielded R = bx/by = 1.08 ± 0.06 (mean ± s.d.), that is, an aspect ratio very close to that of a circle (R = 1). Given the small departure of R from 1, the analysis was conducted assuming axial symmetry, with b = bx = by. The cell shape was captured by the three-parameter surface33

where γ and δ are the polar and azimuthal angles measured from the major axis, a and b are the major and minor semi-axes of an original axisymmetric ellipsoid (that is, a spheroid), c measures the deviation from the symmetric shape in the fore–aft direction (that is, along the major axis), and r(γ,δ) is the distance of a point on the surface from the origin with a polar angle γ and azimuthal angle δ. To quantify a cell’s fore–aft asymmetry, the cell contour obtained from phase contrast microscopy was fitted by using equation (1) and imposing δ = 0, thus obtaining the curve ab/(a2 cos2 γ + b2 sin2 γ)1/2 + c sin γ = r(γ,δ = 0). The contours of 10 HA452 cells collected from the bottom of the flipping chamber, 13 HA452 cells collected from the top, and 12 HA3107 cells collected from the top (all after 30 min of flipping treatment) were determined through image analysis and used to determine the parameters a, b and c for each population (Extended Data Figs 4, 5). The centre of buoyancy of the cell, CB, which is the centre of mass of the (uniform) fluid displaced by the cell, was then determined by finding the geometric centre of the fitted contours.
Centre of mass
To locate a cell’s centre of mass, CW, one needs—in addition to the cell contour—the location, size and density of any heavy organelle. Of particular importance is the nucleus, because of its size and density, and we used epifluorescence microscopy to identify its size and location. The chloroplasts, having higher density than the rest of the cell, could also potentially have contributed to the location of the centre of mass, but in HA452 and HA3107 they are randomly distributed, close to the external membrane, and our analysis showed that their contribution to the offset of the centre of mass is negligible compared to the contribution of the nucleus. Chlorophyll autofluorescence imaging in HA452 cells showed that a cell typically has 15–25 chloroplasts, each 1.5 μm in diameter, in agreement with previous observations23. In the analysis, we considered 20 chloroplasts per cell, each having a diameter of 1.5 μm and a density ρchlo = 1,150 kg m−3 (higher than the density of the rest of the cell, ρcyto = 1,050 kg m−3) (ref. 34). Chloroplasts were assumed to be distributed randomly on the interior of the cell surface, such that their outmost point was in contact with the cell surface (Extended Data Fig. 5b, c), the location where their contribution to the offset would be greatest. The contribution of the chloroplasts to the offset of the centre of mass was found to be negligible compared to the contribution of the nucleus (Extended Data Fig. 5c). Prior to imaging, the nucleus was fluorescently stained by incubating cells for 20 min in the dark with 100 nM SYTO 9 (Molecular Probes, Life Technologies), a green fluorescent nucleic acid stain (excitation/emission 485/498 nm) that stains the nucleus (Extended Data Fig. 4). The intensity of the excitation light was maintained low to prevent photo-toxicity. HA452 and HA3107 cells typically have the nucleus located off-centre. By means of image analysis, we determined the size of the nucleus as well as the distance LN = (LNa2 + LNb2)1/2 between the centre of the nucleus and the centre of buoyancy (Extended Data Fig. 4, Extended Data Table 2 and Supplementary Table 2), where LNa and LNb are the offsets along the major and minor axes, respectively. By using a nucleus density of 1,300 kg m−3 (ref. 35) and an average density for the rest of the cell of 1,050 kg m−3 (ref. 34), we then also computed the distance LW between the centre of mass and the centre of buoyancy.
Centre of hydrodynamic stress
The centre of hydrodynamic stress is the point at which the resultant of all viscous stresses exerted by the fluid on the cell (in our case with swimming cells, resulting from the combination of displacement, reorientation and sedimentation) acts. Bodies with cylindrical symmetry have a centre of hydrodynamic stress located along the axis of symmetry36. To model the flow around a fore–aft asymmetric cell, we solved the three-dimensional Navier–Stokes equations around a cell with the finite element software COMSOL Multiphysics (Burlington, MA), using the characteristic size and shape determined experimentally for the upward-swimming cells (see Extended Data Table 2 and Extended Data Fig. 5) and their characteristic swimming speed (75 μm s−1). Imposing the torque-free condition (sum of all torques on the cell must vanish) allowed us to determine the position of the centre of hydrodynamic stress, CH. Specifically, the coordinates of CH were determined by finding the minimum of the surface integral of the cross product between the stress force and the surface of the cell. The length scale LH quantifies the distance between the centre of buoyancy and the centre of hydrodynamic stress.
Cell mechanics model to predict orientational stability
For a body of revolution swimming in a fluid at a low Reynolds number, there is separation of the equations for translational and rotational motion36. This allows us to write the force-free conditions along the major-axis and minor-axis directions as


where P is the propulsion force, originating from the beating of the flagellum and assumed to act along the major axis of the cell, which is at an angle φ relative to the vertical; D is the drag force, acting by definition in the centre of hydrodynamic stress CH and directed in the opposite direction to the cell swimming velocity, which is at an angle θ relative to the vertical; V is the volume of the cell; ρcell and ρfluid are the densities of the cell and the fluid, respectively. The drag D on a spheroid moving in a fluid with dynamic viscosity η at velocity v depends on the angle α = θ − φ between the body axis (oriented at an angle φ to the vertical) and the direction of motion (oriented at an angle θ to the vertical), and can be computed as D =  cosα +
cosα +  sinα (
sinα ( and
and  are the drag forces corresponding to motion along and perpendicular to the direction of the major axis of the spheroid, respectively).
are the drag forces corresponding to motion along and perpendicular to the direction of the major axis of the spheroid, respectively).
One additional equation can be derived by balancing torques around the centre of buoyancy CB, resulting in TH + TW = TV, where TH is the torque generated by the drag force D, TW is the torque generated by the weight of the cell W = V ρcell g, R ηω is the net viscous torque, R is the coefficient of resistance of the body to rotational motion, and ω is the rotation rate of the cell. The propulsion force P generates no torque about CB, because it acts along the major axis of the cell and goes through CB. The torque–balance equation reads:

where arctan(LNb/LNa) is the contribution to the gravitational torque coming from the offset LNb of the nucleus within the equatorial plane (in this case LNb = 0.25 ± 0.26 μm; Extended Data Fig. 4d). For a given swimming speed v, we numerically solved (using Mathematica) the system of three equations (2), (3) and (4) for the unknowns P, φ and ω. This yielded the rotation rate, ω(θ), as a function of the swimming orientation to the vertical, θ. To quantify the cell stability, we then extracted the reorientation timescale B from ω(θ) through the sinusoid-fitting method described earlier. We note that a stability analysis for upward swimming HA452 cells (with parameters taken from Extended Data Table 2) showed that the term arctan(LNb/LNa) does not change the stability of the cell (that is, upward versus downward; Extended Data Fig. 7e) and this term was thus neglected in the remainder of the analysis. When solving equations (2), (3) and (4) in Mathematica we made the approximation that even for fore–aft asymmetric cells the drag can be computed as the drag on the corresponding spheroid (that is, with asymmetry coefficient set to c = 0), defined as the spheroid with the same volume and same aspect ratio a/b (that is, using ‘equivalent semi-axes’, aeq and beq), because the drag for the latter is known analytically. Specifically, we have computed the drag on the most fore–aft asymmetric HA452 cell we observed (a = 7.87 μm, b = 5.60 μm, c = 1.04 μm; Extended Data Table 2) by solving the three-dimensional Navier–Stokes equations around the cell in Comsol Multiphysics. We found that the drag on the fore–aft asymmetric cell exceeds the drag on the corresponding spheroid (aeq = 7.95 μm, beq = 5.65 μm, c = 0) by only 2%, irrespective of the direction of motion (that is, for both  and
and  ). The approximation of replacing the fore–aft asymmetric shape with a spheroid for drag calculations had a negligible effect on the stability of the cell (compare red and pink curves in Extended Data Fig. 7f). For a prolate spheroid,
). The approximation of replacing the fore–aft asymmetric shape with a spheroid for drag calculations had a negligible effect on the stability of the cell (compare red and pink curves in Extended Data Fig. 7f). For a prolate spheroid,  = 6πηvreq
= 6πηvreq and
and  = 6πηvre
= 6πηvre  , where req is the radius of the sphere with volume equal to the spheroid and
, where req is the radius of the sphere with volume equal to the spheroid and  and
and  are shape factors, which depend on the ratio of minor over major axes of the spheroid, t = a/b. The shape factors are approximated36,37 as
are shape factors, which depend on the ratio of minor over major axes of the spheroid, t = a/b. The shape factors are approximated36,37 as  (t) = 4(t2 − 1)3/2/3t1/3/[(2t2 − 1)1/2 ln[t + (t2 − 1)1/2 – t (t2 − 1)1/2] and
(t) = 4(t2 − 1)3/2/3t1/3/[(2t2 − 1)1/2 ln[t + (t2 − 1)1/2 – t (t2 − 1)1/2] and  = 8(t2 − 1)3/2/3t1/3/[(2t2 − 3)1/2 ln[t + (t2 − 1)1/2 + t (t2 − 1)1/2]. The coefficient of resistance of a prolate spheroid to rotational motion is37 R(t) = 2/3t (t2 + 1) (t2 − 1)3/2/[(2t2 − 1) ln[t + (t2 − 1)1/2] – t(t2 − 1)1/2].
= 8(t2 − 1)3/2/3t1/3/[(2t2 − 3)1/2 ln[t + (t2 − 1)1/2 + t (t2 − 1)1/2]. The coefficient of resistance of a prolate spheroid to rotational motion is37 R(t) = 2/3t (t2 + 1) (t2 − 1)3/2/[(2t2 − 1) ln[t + (t2 − 1)1/2] – t(t2 − 1)1/2].
Stress quantification
To quantify the generation of nitrosative stress, cells exposed to overturning and control cells were sampled from the millifluidic chamber as described earlier, and incubated for 30 min in 500 nM DAF-FM Diacetate (Molecular Probes, Life Technologies), prepared by diluting the original aliquot in the cell culturing medium f/2. DAF-FM Diacetate is a chemical reagent that enables the detection of low concentrations of nitric oxide (NO), a common reactive nitrogen species exuded by eukaryotic cells under stress38,39 (Supplementary Information). DAF-FM by itself is not fluorescent, but forms a fluorescent benzotriazole group (excitation/emission = 495/515 nm) upon reaction with the free radical NO. After incubation, cells were illuminated using cyan light (wavelength ~495 nm) and the fluorescent readout was quantified over time using epifluorescent microscopy. For each cell, the fluorescent intensity increased over time, and reached the peak value at the onset of cell lysis. The magnitude of fluorescent intensity just before cell lysis corresponded to the maximum value of the NO accumulation in the cell. For the quantification, measurements for each cell were normalized with the value of maximum fluorescence intensity. The stress accumulation rate in each experiment was calculated using a linear fit on the portion of the data set where the stress signal was higher than 10% of its maximum intensity.
Statistical tests
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment. We performed a one-way ANOVA to test the differences in the upward bias index r between the still control and the three reorientations treatments: horizontal, vertical, and stochastic. Experiments reported in Figs 1g and 2a were carried out in four replicates. The total number of cells, n, recorded within the field of view to extract the upward bias, r, is reported in figure captions. All the replicates in our experiments were biological replicates. The Anderson–Darling test was performed to test the normality assumption in the four treatments (control: P = 0.28; horizontal: P = 0.55; vertical: P = 0.16; stochastic: P = 0.39). Bartlett’s test was performed to check for homogeneity of variance (χ3 = 4.39, P = 0.22). We made multiple comparisons using a post-hoc Tukey’s HSD test (Supplementary Table 1). A Kruskal–Wallis test was conducted (Bartlett’s test revealed variance heterogeneity) to test the stress differences between the still control and the two subpopulations, top and bottom. Dunn’s test was performed for multiple comparisons (Supplementary Table 3).
Data availability statement
The authors declare that the relevant data supporting the findings of this study are available as supplementary Source Data (Figs 1, 2, 4 and Extended Data Figs 1, 2, 5, 6, 7, 8). Any additional data that support the findings of this study are available from the corresponding author upon request.
References
Margalef, R. Life-forms of phytoplankton as survival alternatives in an unstable environment. Oceanol. Acta 1, 493–509 (1978)
Smayda, T. J. & Reynolds, C. S. Community assembly in marine phytoplankton: application of recent models to harmful dinoflagellate blooms. J. Plankton Res. 23, 447–461 (2001)
Bollens, S. M., Rollwagen-Bollens, G., Quenette, J. A. & Bochdansky, A. B. Cascading migrations and implications for vertical fluxes in pelagic ecosystems. J. Plankton Res. 33, 349–355 (2011)
Schuech, R. & Menden-Deuer, S. Going ballistic in the plankton: anisotropic swimming behavior of marine protists. Limnol. Oceanogr. Fluids Environ. 4, 1–16 (2014)
Sullivan, J. M., Swift, E., Donaghay, P. L. & Rines, J. E. B. Small-scale turbulence affects the division rate and morphology of two red-tide dinoflagellates. Harmful Algae 2, 183–199 (2003)
Berdalet, E. et al. Species-specific physiological response of dinoflagellates to quantified small-scale turbulence. J. Phycol. 43, 965–977 (2007)
Lozovatsky, I., Lee, J., Fernando, H. J. S., Kang, S. K. & Jinadasa, S. U. P. Turbulence in the East China Sea: the summertime stratification. J. Geophys. Res. Oceans. 120, 1856–1871 (2015)
Thomas, W. H. & Gibson, C. H. Effects of small-scale turbulence on microalgae. J. Appl. Phycol. 2, 71–77 (1990)
Sverdrup, H. On conditions for the vernal blooming of phytoplankton. J. Cons. Perm. Int. Explor. Mer 18, 287–295 (1953)
Barton, A. D., Dutkiewicz, S., Flierl, G., Bragg, J. & Follows, M. J. Patterns of diversity in marine phytoplankton. Science 327, 1509–1511 (2010)
Thomas, M. K., Kremer, C. T., Klausmeier, C. A. & Litchman, E. A global pattern of thermal adaptation in marine phytoplankton. Science 338, 1085–1088 (2012)
Brun, P. et al. Ecological niches of open ocean phytoplankton taxa. Limnol. Oceanogr. 60, 1020–1038 (2015)
Litchman, E. & Klausmeier, C. A. Trait-based community ecology of phytoplankton. Annu. Rev. Ecol. Evol. Syst. 39, 615–639 (2008)
Villareal, T. A. & Carpenter, E. J. Buoyancy regulation and the potential for vertical migration in the oceanic cyanobacterium Trichodesmium. Microb. Ecol. 45, 1–10 (2003)
Guasto, J. S., Rusconi, R. & Stocker, R. Fluid mechanics of planktonic microorganisms. Annu. Rev. Fluid Mech. 44, 373–400 (2012)
Franks, P. J. S. Has Sverdrup’s critical depth hypothesis been tested? Mixed layers vs. turbulence layers. J. Mar. Sci. 72, 1–11 (2015)
Zirbel, M. J., Veron, F. & Latz, M. I. The reversible effect of flow on the morphology of Ceratocorys horrida. J. Phycol. 58, 46–58 (2000)
Roberts, A. M. Geotaxis in motile micro-organisms. J. Exp. Biol. 53, 687–699 (1970)
Durham, W. M., Kessler, J. O. & Stocker, R. Disruption of vertical motility by shear triggers formation of thin phytoplankton layers. Science 323, 1067–1070 (2009)
Durham, W. M. et al. Turbulence drives microscale patches of motile phytoplankton. Nature Commun. 4, 2148 (2013)
Yamasaki, Y. et al. Extracellular polysaccharide–protein complexes of a harmful alga mediate the allelopathic control it exerts within the phytoplankton community. ISME J. 3, 808–817 (2009)
Thorpe, S. A. An Introduction to Ocean Turbulence (Cambridge Univ. Press, 2007)
Hara, Y. & Chihara, M. Morphology, ultrastructure and taxonomy of the raphidophycean alga Heterosigma akashiwo. Bot. Mag. 100, 151–163 (1987)
Smayda, T. J. Adaptations and selection of harmful and other dinoflagellate species in upwelling systems.1. Morphology and adaptive polymorphism. Prog. Oceanogr. 85, 71–91 (2010)
Hemmersbach, R., Volkmann, D. & Hader, D. P. Graviorientation in protists and plants. J. Plant Physiol. 154, 1–15 (1999)
Bulmer, M. G. Selection for iteroparity in a variable environment. Am. Nat. 126, 63–71 (1985)
Moxon, E. R., Rainey, P. B., Nowak, M. A. & Lenski, R. E. Adaptive evolution of highly mutable loci in pathogenic bacteria. Curr. Biol. 4, 24–33 (1994)
Ackermann, M. A functional perspective on phenotypic heterogeneity in microorganisms. Nature Rev. Microbiol. 13, 497–508 (2015)
Harvey, E. L., Menden-Deuer, S. & Rynearson, T. A. Persistent intra-specific variation in genetic and behavioral traits in the raphidophyte, Heterosigma akashiwo. Front. Microbiol. 6, 1277 (2015)
Foster, K. W. & Smyth, R. D. Light Antennas in phototactic algae. Microbiol. Rev. 44, 572–630 (1980)
Ouellette, N. T., Xu, H. & Bodenschatz, E. A quantitative study of three-dimensional Lagrangian particle tracking algorithms. Exp. Fluids 40, 301–313 (2006)
Gurarie, E., Grünbaum, D. & Nishizaki, M. T. Estimating 3D movements from 2D observations using a continuous model of helical swimming. Bull. Math. Biol. 73, 1358–1377 (2011)
Roberts, A. M. & Deacon, F. M. Gravitaxis in motile micro-organisms: the role of fore–aft body asymmetry. J. Fluid Mech. 452, 405–423 (2002)
Wada, M., Miyazaki, A. & Fujii, T. On the mechanisms of diurnal vertical migration behavior of Heterosigma akashiwo (Raphidophyceae). Plant Cell Physiol. 26, 431–436 (1985)
Milo, R. & Phillips, R. Cell Biology by the Numbers (Garland Science, Taylor & Francis Group, 2016)
Happel, J. & Brenner, H. Low Reynolds Number Hydrodynamics: With Special Application to Particulate Media (Mechanics of Fluids and Transport Processes) Vol. 1 (Englewood Cliffs, 1983)
Koenig, S. H. Brownian motion of an ellipsoid. A correction to Perrin’s results. Biopolymers 14, 2421–2423 (1975)
Bouchard, J. N. & Yamasaki, H. Heat stress stimulates nitric oxide production in Symbiodinium microadriaticum: a possible linkage between nitric oxide and the coral bleaching phenomenon. Plant Cell Physiol. 49, 641–652 (2008)
Vardi, A. et al. A diatom gene regulating nitric-oxide signaling and susceptibility to diatom-derived aldehydes. Curr. Biol. 18, 895–899 (2008)
Acknowledgements
We thank M. Ackermann for input on bet-hedging, G. Boffetta and M. Cencini for sharing the direct numerical simulations data, S. Menden-Deuer for providing H. akashiwo CCMP3107 and for discussions, E. Berdalet, A. Vardi, D. Anderson and D. J. McGillicuddy for suggestions, and M. Barry for initial development of the flip chamber. This work was partially supported by a Human Frontier Science Program Cross Disciplinary Fellowship (LT000993/2014-C to A.S.), a Swiss National Science Foundation Early Postdoc Mobility Fellowship (to F.C.), and a Gordon and Betty Moore Marine Microbial Initiative Investigator Award (GBMF 3783 to R.S.).
Author information
Authors and Affiliations
Contributions
A.S., F.C. and R.S. designed research. A.S. performed the flipping experiments (with assistance from F.C.) and the stress experiments. A.S. and F.C. jointly performed the image analysis and the visualization to determine cell morphology. F.C. developed the fluid dynamic simulations and the cell mechanics model (with assistance from A.S.). F.C. performed the statistical analysis. R.S. provided oversight and advice for all parts of the project. A.S., F.C. and R.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. J. Behrenfeld, S. Menden-Deuer and T. Pedley for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Vertical distribution of HA452 cells.
a–d, Vertical distribution is reported for a population grown under a diel light cycle (a), for regrown cells (b), for a monoclonal population (c) and for a starved population (d). a, Upward bias for exponential-phase HA452 cells grown under a diel light cycle (14 h light: 10 h dark), showing the characteristic split after 100 flips (blue, n = 844 cells; red curve is control, n = 910). A similar split was observed for cells cultured under constant illumination (Fig. 1g). b, Cells regrown from those collected from the bottom of the chamber after 100 flips. Although cells collected from the bottom were positively gravitactic (swimming downwards; blue curve in a), cells regrown from these are negatively gravitactic (swimming upwards; pink curve in b, n = 411). Upon exposure to 100 flips, these regrown cells again exhibited the population split (cyan curve, n = 391). a, b, The solid lines represent the mean of the equilibrium vertical distribution over time (a: mean of 92 frames; b: mean of 78 frames), and the shaded regions represent ± 1 s.d. from the mean. In the insets, the error bars of the upward bias, r, are extracted from the cells’ vertical distribution. c, The population split also occurs in a monoclonal population of HA452. The inset shows the upward bias (mean ± s.d. of three replicates) calculated from the relative distribution of the cells at equilibrium, after being exposed to 100 flips (blue, total number of cells n = 2,985) and for the control (red, n = 2,490). The star indicates statistical significance in the difference between treatment and control (two-sided t-test, t4 = 4.79, P = 0.009). d, A nutrient-starved HA452 population does not split upon flipping. Nutrient-starved cells were harvested at stationary phase (350 h after inoculation). The inset shows the upward bias (mean ± s.d. of three replicates, two-sided t-test, t4 = 0.91, P = 0.42) of the cells after being exposed to 100 flips (blue, n = 743) and for the control (red, n = 1,011). c, d, The solid lines represent the mean of the equilibrium vertical distribution over three replicates, and the shaded regions represent ± 1 s.d. from the mean.
Extended Data Figure 2 Swimming behaviour of HA452 cells after exposure to 100 flips.
a, The relative distribution of swimming speeds, obtained by image analysis of cells in the flipping chamber, showing no difference in the absolute swimming speed of the two subpopulations, HA452(↑): v = 74.5 ± 42.4 μm s−1 (n = 1,780 cells); and HA452(↓): v = 73.8 ± 46.2 μm s−1 (n = 992). b, Distribution of the vertical component of the swimming velocity in HA452(↑) and HA452(↓), showing distinct peaks in opposite directions, at approximately ±50 μm s−1, and corresponding to upward and downward swimming, respectively. c, Distribution of the horizontal component of the swimming velocity in HA452(↑) and HA452(↓), showing no appreciable difference between the two subpopulations. In all panels, speeds were obtained by tracking cells for 15 s just after a single additional flip following the 100 flips. Here, trajectories in the top 1 mm of the chamber were assigned to HA452(↑) and trajectories in the bottom 1 mm were assigned to HA452(↓). For each subpopulation, velocities were averaged over all trajectories. d, e, The joint distribution of the swimming velocity in the vertical and horizontal directions for HA452(↑) (d) and HA452(↓) (e). The colour scale indicates the relative distribution of cell trajectories counted over the 15 s movie normalized by the maximum in each subpopulation.
Extended Data Figure 3 Additional flipping experiments with a range of raphidophyte and dinoflagellate species revealed that rapid behavioural responses in phytoplankton to flipping are not restricted to HA452.
a, b, Vertical distribution of Chattonella marina_cf CM2962 (a) and Prorocentrum minimum PM291 (b), both showing a split similar to that of HA452. Solid lines represent the mean of the equilibrium vertical distribution over four and three replicates respectively. Shaded regions represent ± 1 s.d. from the mean. The insets show the upward bias, r, after 300 flips (blue) and for the control, consisting of the same time in the chamber without flipping (red, mean ± s.d.). The star indicates statistical significance (two-sided t-test) between the two treatments (CM2962: t6 = 3.66, P = 0.01; PM291: t4 = 2.85, P = 0.04). c, d, Upward bias index, r (mean ± s.d.), for seven raphidophyte strains (c) and ten dinoflagellate strains (d). The error bars of the upward bias, r, are extracted from the cells’ vertical distribution. Full names of strains are provided in Extended Data Table 1. The number of cells analysed for each case is given in Extended Data Table 1. Many of these strains showed a moderate to strong response to flipping, as shown by the change in their upward bias between treatment and control.
Extended Data Figure 4 Quantification of the shape and nucleus position of H. akashiwo cells based on single-cell microscopy, for subpopulations HA452(↑), and HA452(↓), as well as strain HA3107.
a–c, The top row in each panel shows micrographs obtained by epifluorescence microscopy (Methods), of HA452 cells harvested from the top (HA452(↑), n = 13) (a) and bottom (HA452(↓), n = 10) (b) of the millifluidic chamber after 100 flips, and of HA3107 (n = 12) (c). The cell itself was visualized using an inverted microscope (Nikon TE2000) in phase contrast, equipped with a ×20 or ×40 objective and an Andor iXon Ultra 897 camera. Prior to imaging, cells were stained with SYTO 9 (Methods) to visualize the nucleus through fluorescence microscopy (central bright spot). Image analysis was used to extract the contour of each cell and the position of its nucleus (middle rows). Experimentally obtained cell contours (black) were fitted with a three-parameter curve (equation (1); red) (bottom rows). Single-cell parameters associated with these fits are given in Extended Data Table 2 and Supplementary Table 2. d, Quantification of the axial symmetry of HA452 cells from single-cell microscopy. The top row shows micrographs obtained by epifluorescence microscopy for ten randomly chosen cells after flipping. Image analysis was used to extract the contour of each cell and the position of its nucleus (middle row). Experimentally obtained cell contours (black) were fitted with an ellipse with major and minor semi-axis bx, by (red) and a circle of radius req (blue). The degree of axial asymmetry, quantified as R = bx/by = 1.08 ± 0.06 (mean ± s.d., n = 10), was very close to that of a circle (R = 1), showing that cells were very close to axially symmetric. The offset of the position of the nucleus compared to the centre of the circle in the plane perpendicular to the major axis was LNb = 0.25 ± 0.26 μm (mean ± s.d.).
Extended Data Figure 5 Cell shapes for HA452(↑), HA452(↓) and HA3107.
a, The graph shows cell shape variation in terms of the degree of fore–aft asymmetry and minor/major axis ratio (see equation (1)). The parameter c denotes the degree of fore–aft asymmetry, a is the semi-major axis, b is the semi-minor axis. We highlighted the average contours (see Extended Data Table 2) for the subpopulation of downward swimmers (HA452(↓), blue), the subpopulation of upward swimmers (HA452(↑), orange) and HA3107 (green). Values of a, b and c are given in Extended Data Table 2 and Supplementary Table 2. b, Epifluorescence micrograph showing the chloroplasts. c, Three-dimensional schematic of an HA452 cell used to compute the contribution of the chloroplasts to the offset of the centre of mass relative to the contribution of the nucleus. The large sphere represents the nucleus (density ρN = 1,300 kg m−3, radius sN = 2.5 μm) and the 20 small spheres represent the chloroplasts (density ρchlo = 1,150 kg m−3, radius rchlo = 0.75 μm), which for the purpose of computing the contribution to the centre of mass were taken to be randomly distributed adjacent to the cell membrane. The contribution of the chloroplasts to the offset of the centre of mass from the centre of buoyancy was found to be <4% of the contribution of the nucleus and was thus neglected in the stability analysis.
Extended Data Figure 6 Regime diagram of cell stability.
Two physical features—summarized by two morphological length scales—determine cell stability: the asymmetry in shape, quantified by LH/a, and the mass distribution, quantified by LW/a, where a is the semi-major axis, LH quantifies the distance between the centre of buoyancy and the centre of hydrodynamic stress, and LW the distance between the centre of buoyancy and the centre of mass (Fig. 3). Colours denote the cell rotation rate ω following an orientational perturbation (equation (4): ω > 0 denotes negatively gravitactic cells (stable upward), ω < 0 denotes positively gravitactic cells (stable downward), and ω = 0 (white dashed line) denotes neutrally stable cells. Sample asymmetry configurations corresponding to different locations on the regime diagram are illustrated by the black-and-white schematics. Filled circles denote experimental data (see Extended Data Table 2). The morphological adaptation of HA452 cells in response to overturning causes the population stability to switch (red arrow crossing the white dashed line). The original upward swimming population splits into a subpopulation swimming downward HA452(↓) and a subpopulation swimming upward, HA452(↑).
Extended Data Figure 7 Orientational stability of H.
akashiwo . a, Rotation rate, ω, of HA452 cells before the overturning treatment, as a function of the direction, θ, of the instantaneous swimming velocity, v, relative to the vertical. The rotation rate of the cells (n = 2,257) was quantified by tracking them in the time intervals 0–5 s (grey) and 5–15 s (magenta) directly after a single flip, and averaged over all the cells as a function of θ. The difference between the two curves denotes the presence of cells that reorient more rapidly and others that reorient more slowly. Dashed lines are sinusoidal fits to the experimental data, used to obtain the reorientation timescale B. Solid lines (colour-coded) denote the arithmetic mean over all cell trajectories. The shaded region denotes ± 1 s.e.m. The reorientation timescales obtained from these data are B = 7.2 s for the first 5 s and B = 12.2 s for the following 10 s, denoting a nearly twofold higher stability for cells that were observed reorienting in the first 5 s. b, Rotation rate, ω, of HA3107 cells before the overturning treatment, as a function of the swimming direction, θ. The rotation rate was quantified by tracking cells for 15 s directly after a single flip and averaged over all cells as a function of θ (n = 1,283). The dashed line is a sinusoidal fit to the data used to obtain B. The solid line denotes the arithmetic mean over all cell trajectories. The shaded region denotes ± 1 s.e.m. The reorientation timescale obtained for HA3017 from these data was B = 4.9 s. c, Distribution of swimming orientation of HA452 cells before the overturning treatment (same data as in a). The distribution was quantified by tracking cells in the time intervals 0–5 s (black), 5–10 s (green), 10–15 s (cyan), and 15–20 s (blue) directly after a single flip, and averaged over all cells as a function of θ. Note that after 15 s the distribution does not appreciably change. d, Time series of the vertical distribution of HA452 following a 100-flip treatment (period Q = 18 s). The cell distribution inside the chamber was tracked after the end of the overturning treatment, with time zero corresponding to the termination of the treatment (between 461 and 592 cells are included in each vertical profile). At t = 1 s (blue) the cell distribution is homogeneous because the cells have been continuously flipped for 30 min and 1 s is not long enough to allow cells to reach their equilibrium profile. To traverse the chamber (4 mm), it would take 80 s for cells swimming with a vertical speed of 50 μm s−1 (Extended Data Fig. 2). In fact, it takes 90 s (orange) to establish the bimodal distribution at equilibrium, corresponding to the population split induced by overturning. The population split is then maintained for at least 7 h (black). The upward bias shown in Figs 1 and 2 and in Extended Data Figs 1 and 3 is always computed 30 min after the overturning ceases. e, Effect of the torque generated by the offset LNb of the nucleus within the equatorial plane, obtained from the cell mechanics model, shown in terms of its effect on the dependence of the rotation rate on the body axis angle for the upward-swimming subpopulation HA452(↑). The dashed red line denotes the case without offset (LNb = 0), the purple and pink lines represent the cases in which the nucleus is offset by LNb = +0.25 μm and by LNb = −0.25 μm, respectively (the average offset measured experimentally; see Extended Data Fig. 4d), and the dark green and light green lines represent the cases in which the offset corresponds to mean + s.d. of the experimentally measured values, that is, LNb = +(0.25 + 0.26) = +0.51 μm and LNb = −(0.25 + 0.26) = −0.51 μm. Note that the overall upward stability of the cells remains unchanged when one accounts for the effect of LNb, since the stable points for all the cases (coloured dots) always occur for a swimming orientation θ that is smaller than ± π/2 (dashed vertical lines), which separates upward and downward swimming (θ = ±28° for LNb = ±0.25 μm; θ = ±35° for LNb = ±0.51 μm. Note that the results are symmetric around the vertical direction, θ = 0°). f, Stability analysis demonstrating that two assumptions made in our calculations have negligible consequences, in particular the assumptions that (1) the angle α between the body axis and the direction of motion is zero (compare orange and red lines), and (2) the drag on the fore–aft asymmetric upward swimmers can be approximated by the drag on a spheroid (compare red and pink lines). Shown is the rotation rate as a function of body axis angle for three cases: a spheroidal cell in which the major axis is aligned with the direction of motion (α = 0, orange), a spheroidal cell in which the misalignment between major axis and direction of motion is accounted for (α = 4°, red), and a fore–aft asymmetric cell in which the misalignment between major axis and direction of motion is accounted for (α = 4°, pink). Parameters were taken from Extended Data Table 2 (first row), for the upward-swimming cells. The fore–aft asymmetric case was simulated with Comsol Multiphysics. Note that the cell stability is the same in all three cases, as evidenced from the fact that the curve has a stable point at a swimming angle of θ = 0 and a negative minimum at θ = π/2, which together imply upward stability. Throughout our analyses, we have thus adopted the spheroidal approximation for the calculation of the drag, and taken into account the contribution to the cell stability by the angle α.
Extended Data Figure 8 The growth curve of HA452 and the upward bias, r, of cells over the course of a day.
a, To obtain the growth curve, we sampled cells from the original culture (see Methods) at the specified time points (t = 0, 36, 68, 90, 114, 126, 140, 164, 288, 360 h). Cells were counted by imaging them inside the flip chamber, in the middle plane. Red dots represent the mean number of cells over time (between 76 and 93 frames were analysed for each data point). Error bars represent ± 1 s.d. from the mean. The growth curve of HA452 is shown in linear and semi-log scale (inset). The population density at carrying capacity was 3 × 105 cells ml−1, reached after ~2 weeks of incubation (= 360 h). The population’s intrinsic growth rate was found to be 0.4 day−1, as measured by fitting a logistic curve to the data (black line). In the inset, the shaded orange region shows the growth stage at which cells were harvested for experiments with exponential-phase cells (most experiments), while the shaded magenta region denotes the growth stage at which cells were harvested for experiments with starved cells. b, Upward bias, r, of HA452 cells over the course of a day, with time measured from midnight. For each data point the equilibrium vertical distribution was measured 30 min after termination of the overturning treatment, for both 10 flips (n = 560 cells) and 100 flips (n = 723) (for the control: 30 min after introduction of cells in the flipping chamber, n = 674). A positive upward bias denotes negatively gravitactic cells (that is, preferentially up-swimming). Gravitaxis can be seen to follow a diel cycle, even though the culture was kept under constant illumination. Flipping experiments consistently show a population split, leading to a reduction in the upward bias of the 10 flips and 100 flips treatments compared to the control treatment. The experiments were all conducted between 09:00 and 12:00, where the upward bias measured for the control cells presents the maximum stability.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and Supplementary References. (PDF 302 kb)
Supplementary Data
This file contains Supplementary Table 1. (XLSX 11 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 12 kb)
Supplementary Data
This file contains Supplementary Table 3. (XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Sengupta, A., Carrara, F. & Stocker, R. Phytoplankton can actively diversify their migration strategy in response to turbulent cues. Nature 543, 555–558 (2017). https://doi.org/10.1038/nature21415
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21415
- Springer Nature Limited
This article is cited by
-
Fluid flow drives phenotypic heterogeneity in bacterial growth and adhesion on surfaces
Nature Communications (2024)
-
Morphogenesis in space offers challenges and opportunities for soft matter and biophysics
Communications Physics (2023)
-
Heterogeneity of interaction strengths and its consequences on ecological systems
Scientific Reports (2023)
-
Aerosolization flux, bio-products, and dispersal capacities in the freshwater microalga Limnomonas gaiensis (Chlorophyceae)
Communications Biology (2023)
-
Active matter in space
npj Microgravity (2022)