Abstract
In plants and algae, light serves both as the energy source for photosynthesis and a biological signal that triggers cellular responses via specific sensory photoreceptors. Red light is perceived by bilin-containing phytochromes and blue light by the flavin-containing cryptochromes and/or phototropins (PHOTs)1, the latter containing two photosensory light, oxygen, or voltage (LOV) domains2. Photoperception spans several orders of light intensity3, ranging from far below the threshold for photosynthesis to values beyond the capacity of photosynthetic CO2 assimilation. Excess light may cause oxidative damage and cell death, processes prevented by enhanced thermal dissipation via high-energy quenching (qE), a key photoprotective response4. Here we show the existence of a molecular link between photoreception, photosynthesis, and photoprotection in the green alga Chlamydomonas reinhardtii. We show that PHOT controls qE by inducing the expression of the qE effector protein LHCSR3 (light-harvesting complex stress-related protein 3) in high light intensities. This control requires blue-light perception by LOV domains on PHOT, LHCSR3 induction through PHOT kinase, and light dissipation in photosystem II via LHCSR3. Mutants deficient in the PHOT gene display severely reduced fitness under excessive light conditions, indicating that the sensing, utilization, and dissipation of light is a concerted process that plays a vital role in microalgal acclimation to environments of variable light intensities.
Similar content being viewed by others
Main
In oxygenic photosynthesis, the absorption of light by pigments such as chlorophylls and carotenoids that are embedded in the light-harvesting complexes (LHCs) facilitates the transfer of energy to reaction centres, triggers electron flow from H2O to NADPH and generates a proton motive force across the thylakoid membranes of chloroplasts that is used to drive ATP synthesis. The ATP and NADPH molecules that are generated are utilised in CO2 fixation in the Calvin–Benson cycle5. Whenever light is absorbed beyond the CO2-assimilation capacity, over-excitation of the photosystems leads to photodamage and possibly cell death. These negative consequences are prevented by non-photochemical quenching (NPQ), an intricate photoprotective process that dissipates excess absorbed energy. The major component of NPQ is qE, which occurs at the LHCs of photosystem II (ref. 6).The process of qE is driven by acidification of the thylakoid lumen that occurs under excess light, a process that modifies the pigment composition of LHCII via the xanthophyll cycle and activates specific qE protein effectors7. In the active state, these protein effectors (PSBS in plants and LHCSR proteins in green algae8,9) increase the energy dissipation capacity of LHCII via an unresolved mechanism10,11. Whereas plants constitutively express PSBS, the green alga C. reinhardtii only accumulates LHCSR3 (the major qE effector) following environmental stresses such as excess light and nutrient starvation12. Induction of LHCSR3 requires Ca2+ signalling and active photosynthesis13,14, but the molecular mechanism behind this process is largely unknown.
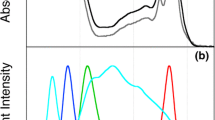
To gain insight into how LHCSR3 induction is regulated, we measured the light-colour dependency (action spectrum) of qE induction in C. reinhardtii. Low-light-acclimated cells were exposed for 4 h to highly intense monochromatic light at different wavelengths between 400 nm and 720 nm, provided by the Okazaki Large Spectrograph15, to induce LHCSR3. The kinetics of NPQ development were then followed under saturating white light for 5 min (Fig. 1a). The extent of NPQ observed at the end of illumination was used to evaluate the action spectrum of qE (Fig. 1b), corresponding to the fraction of NPQ that is rapidly reversible in the dark. This spectrum was comparable to the spectrum of LHCSR3 accumulation (Fig. 1c), confirming the close relationship between energy dissipation and LHCSR3 induction in this alga8. We found that blue light was more effective than red light in inducing the qE response (Fig. 1a, b) and LHCSR3 accumulation (Fig. 1c), although the cells absorbed both almost equally (Fig. 1b, grey area). This finding suggested that a blue-light-absorbing molecule (likely to be a blue-light receptor) is involved in the accumulation of LHCSR3 in high light intensities. To test this hypothesis we compared the kinetics of NPQ (Fig. 1a), the action spectra of qE (Fig. 1b) and the induction of LHCSR3 (Fig. 1c) in wild-type C. reinhardtii cells and in mutants lacking blue-light receptors—either the animal-like cryptochrome (aCRY)16, or PHOT17. We found that acry mutant cells (deficient in a functional ACRY gene, hereafter acry) (Extended Data Fig. 1a) showed wild-type levels of qE (Fig. 1b) and LHCSR3 accumulation (Fig. 1c) throughout the entire light spectrum, including red light, where aCRY acts16 in the absence of phytochromes in C. reinhardtii18. Conversely, phot mutant cells (deficient in a functional PHOT gene, hereafter phot; Extended Data Fig. 1b) specifically lacked blue-light induction of qE (Fig. 1b) and LHCSR3 accumulation (Fig. 1c), and therefore were more prone to photodamage (Fig. 1d). Induction of LHCSR3 and NPQ in response to high-intensity white light was also largely compromised in phot cells, in contrast to wild-type and acry cells (Extended Data Fig. 1c–e). The ability of phot cells to accumulate photoprotective carotenoids via the xanthophyll cycle was, however, similar to that of wild-type and acry cells (Extended Data Fig. 1f, g). The phenotype of phot cells was similar to that of npq4 cells ((cells deficient in functional LHCSR3.1 and LHCSR3.2 genes8); Fig. 1b, d), that we used as a qE-lacking strain. PHOT-mediated blue-light control of LHCSR3 accumulation was highly specific. Accumulation of the representative subunits of the major photosynthetic complexes was comparable in all cell lines and was wavelength-insensitive (Extended Data Fig. 2a, b). The photosynthetic electron transport rate (ETR) was reduced in phot cells, especially upon high light exposure (Extended Data Fig. 1h, i). As photosynthesis is required for proper LHCSR3 accumulation13,14, the lower electron flow capacity of phot cells could diminish LHCSR3 induction. To evaluate this hypothesis, we titrated the ETR in wild-type cells using increasing concentrations of the herbicide 3-(3,4-dichlorophenyl)-1,1-dimethylurea (DCMU), an inhibitor of electron transfer in photosystem II. The graduated addition of DCMU progressively reduced the ETR to a much lower extent in wild-type cells than in DCMU-free phot cells (Extended Data Fig. 3a). However, accumulation of LHCSR3 remained greater in DCMU-treated wild-type cells than in DCMU-untreated phot cells (Extended Data Fig. 3b). We therefore excluded the possibility that the diminished ETR observed in phot cells was the cause of the impaired LHCSR3 accumulation. We concluded instead that the impaired photoprotection in phot cells was the cause of the reduced ETR (Extended Data Fig. 1i) and enhanced photosensitivity (Fig. 1d), indicating that PHOT is a central actor of photoprotection in this alga.
a, NPQ in wild-type (137c), phot and acry cells after exposure for 4 h to different wavelengths of light (250 μmol photons m−2 s−1). Representative dataset of an experiment replicated three times on different biological samples. b, qE induction in wild-type (WT; black), acry (green) npq4 (blue) and phot (red) strains by 4 h illumination with different wavelengths of light (250 μmol photons m−2 s−1). n = 3 biological samples, mean ± s.d. Grey area, absorption spectrum of wild-type cells. RU, relative units. c, Immunoblot analysis of LHCSR3 accumulation (ATPB is the loading control). Representative dataset of an experiment replicated three times on different biological samples. d, Erlenmeyer flasks containing wild-type, acry, phot and npq4 cells after 16 h of exposure to light of 20, 200 and 750 μmol photons m−2 s−1. Representative photographs from three biological replicates.
Phototropins are ubiquitous in plants and algae. Their function in light perception involves two structurally similar LOV domains (LOV1 and LOV2) at the N terminus and downstream signalling via a C-terminal serine/threonine kinase domain2. We dissected the role of the different PHOT domains in LHCSR3-induction using existing mutants19 (Fig. 2a). Complementation of phot cells with the full-length phot cDNA (giving the PPHOT strain) restored PHOT protein expression (Extended Data Fig. 4a), expression of LHCSR3.1 and LHCSR3.2 transcripts (both of which encode LHCSR3 (ref. 8; Fig. 2b) and expression of LHCSR3 protein (Fig. 2c) to a limited extent. This partial restoration was nonetheless sufficient to largely rescue resistance to high-light-induced stress (Fig. 2d and Extended Data Fig. 4b). Moreover, introduction of the LHCSR3.1 gene into phot cells under the control of the strong, PHOT-independent PSAD promoter (the PLHCSR3 strain) also rescued qE and LHCSR3 levels (Extended Data Fig. 5). This therefore confirmed the univocal relationship between PHOT, LHCSR3 and photoprotection. Conversely, complementation of phot cells with truncated gene carrying only the photosensory domains LOV1 and LOV2 (giving the PLOV strain) did not rescue either protein expression or photosensitivity (Fig. 2c, d and Extended Data Fig. 4b). Finally, complementation with the kinase domain of PHOT (giving the PKIN strain) fully restored expression of LHCSR3 and photoprotection (Fig. 2c, d and Extended Data Fig. 4b). In this strain, however, LHCSR3 accumulation became light–colour-independent (Fig. 3a, b), in contrast to the wild-type (Fig. 1a, c) and the PPHOT (Fig. 3a, b) strain. This indicates that removal of the LOV domains leads to the loss of inhibition of the kinase activity, as has been previously reported in plants20. Deregulation of the PHOT kinase activity (as in the PKIN strain) or removal of the kinase domain (as in phot cells and in PLOV) did not alter LHCSR3 phosphorylation levels (Extended Data Fig. 6), ruling out any link between PHOT and LHCSR3 phosphorylation. We also found that LHCSR3 accumulation remained light-intensity-dependent in the PKIN strain (Fig. 3c, d), suggesting that in addition to PHOT, high-light-intensity photosynthesis was required for the accumulation of LHCSR3. In another mutant, generated by inserting an additional copy of the kinase domain into wild-type C. reinhardtii cells (giving the WTKIN strain; Extended Data Fig. 7a–d), LHCSR3 accumulation also became largely wavelength-independent but was still sensitive to light intensity. Moreover, accumulation of LHCSR3 was completely blocked upon inhibition of photosynthesis by DCMU in the PKIN and WTKIN strains (Fig. 3c, d and Extended Data Fig. 7e). Additionally, LHCSR3 was not accumulated under light from the far-red part of the spectrum (720 nm) in each cell line (Figs 1c, 3a, b and Extended Data Fig. 7c, d). This light is not absorbed by C. reinhardtii cells (Fig. 1b, grey area) and therefore does not activate photosynthesis.
a, Domains of the PHOT gene in wild-type and complemented phot lines. LOV, photosensory domain; KINASE, kinase domain; BLE, bleomycin resistance cassette. b, LHCSR3.1 and LHCSR3.2 mRNA accumulation in wild-type (cw15-302), phot, and variously phot-complemented lines exposed to low or high light intensities (LL or HL, respectively). Values relative to an endogenous control gene (GBLP) were normalized to wild-type samples under high light intensity (n = 3 biological samples, mean ± s.d.). c, Immunoblot analysis of LHCSR3 in wild-type and phot mutants (ATPB is the loading control). Representative dataset of an experiment replicated six times on different biological samples. d, Erlenmeyer flasks containing wild-type, phot, PPHOT, PKIN and PLOV strains after 20 h exposure to light of 20, 200 or 750 μmol photons m−2 s−1. Representative pictures from four biological replicates.
a, Action spectrum of LHCSR3 accumulation in PPHOT and PKIN (ATPB is the loading control). Representative dataset of an experiment replicated three times on different biological samples. b, Densitometric quantification of LHCSR3 accumulation of Fig. 3a. Values are normalized to 475 nm (n = 3 biological samples, mean ± s.d.). c, LHCSR3.1 and LHCSR3.2 mRNA accumulation in PKIN at low light intensity and at high light intensity in the presence and absence of the PSII inhibitor DCMU. Values are normalized to high light intensity (n = 3 biological samples, mean ± s.d.). d, LHCSR3 protein accumulation in PKIN at low and high light intensity in the presence or absence of DCMU for 4 h. Representative dataset of an experiment replicated three times on different biological samples.
In Arabidopsis thaliana, PHOT proteins regulate the blue-light-induced increase in cytosolic free Ca2+ (refs 21, 22). As PHOT controls LHCSR3 expression in C. reinhardtii (Fig. 1) and Ca2+ is required for the accumulation of LHCSR3 (ref. 13), a relationship between PHOT, Ca2+, and LHCSR3 can be conceived. However, a tenfold increase in extracellular Ca2+ (to 3.4 mM) did not restore LHCSR3 accumulation in the phot mutant (Extended Data Fig. 8a). This finding led us to conclude that Ca2+ signalling is not modulating LHCSR3 in the absence of PHOT. We propose that other second messengers, such as the cyclic nucleotides cAMP or cGMP, act as the signalling molecules downstream of PHOT. We tested this using a pharmacological approach and found that treatment of phot cells with 3-isobutyl-1-methylxanthine (IBMX), an inhibitor of cAMP and cGMP phosphodiesterases, rescued LHCSR3 expression (Fig. 4a and Extended Data Fig. 8b). We confirmed this effect by incubating phot cells with dibutyrylated cGMP and cAMP (DB-cGMP and DB-cAMP; Fig. 4a), suggesting that cyclic nucleotides are not only critical in mating and phototaxis18, but are also involved in photoprotection in C. reinhardtii through the regulation of LHCSR3 expression. On the other hand, induction of LHCSR3 in IBMX-treated wild-type or phot cells remained light-intensity-dependent (Fig. 4a), indicating that cAMP and/or cGMP require a high-light-photosynthesis-related signal to be effective on LHCSR3 accumulation.
a, Immunoblot analyses of LHCSR3 accumulation in wild-type (cw15-302) and phot cells treated with the phosphodiesterase inhibitor IBMX, DB-cAMP or DB-cGMP after 20 h at low or high light intensities. Representative dataset of an experiment replicated five times on different biological samples. Ctrl, control. b, Schematic representation of the relationship between photoreception, photosynthesis and -photoprotection in C. reinhardtii. cp, chloroplast; cNMP, cyclic nucleotide mono phosphate (cAMP or cGMP); ET, electron transport; eye, eyespot; nuc, nucleus; PSII, photosystem II.
Extensive research has focused on the nature and function of the molecular actors involved in the sensing (by photoreceptors) and utilization (by photosynthetic complexes) of environmental light. Our findings here uncover the molecular linkage between these two essential functions of photosynthetic organisms (Fig. 4b). Blue light perceived by PHOT mediates the photoprotection of the photosynthetic machinery (qE) in a green alga. The LOV domains of PHOT provide blue-light sensitivity while the C-terminal kinase domain performs signal transduction, possibly via the initiation of a cyclic nucleotide monophosphate signalling cascade. Downstream of PHOT, this signal integrates with another regulatory signal from the chloroplast that carries information about the amount of absorbed light that is not used for CO2 fixation. This signal relies on photosynthetic electron transfer13,14 via an as-yet-unknown mechanism. The integrated signal affects accumulation of LHCSR3.1 and LHCSR3.2 transcripts (Fig. 2b), primarily owing to the activation of transcription and partially to the stabilization of the transcripts (Extended Data Fig. 9). This contrasts with plant PHOT proteins that have been suggested to destabilize transcripts23, while little evidence exists for their role in activation of transcription2. The LHCSR3 polypeptide is then imported into the chloroplast thylakoids, where it modulates qE (Fig. 4b).
Because LHCSR3 is an ancient light-harvesting protein8 found only in lower plants including green unicellular eukaryotic algae (chlorophytes) and mosses7, as well as in some algae with secondary plastids, we propose that the molecular link between photoreception, photosynthesis, and photoprotection discovered in this study has evolved in the environment in which photosynthesis originated during evolution—namely the water columns—where blue light dominates the available spectrum. Consistent with this hypothesis, photoprotection in cyanobacteria, which are thought to share a common ancestry with chloroplasts, also relies on blue light. The cyanobacterial NPQ protein effector OCP (orange carotenoid protein) binds a single blue-light-absorbing carotenoid. NPQ in cyanobacteria is also triggered by high-intensity blue light24. Evidences also show a link between blue light and photoprotection in diatoms25 via an uncharacterized mechanism. On the other hand, the blue-light-dependent control of photoprotection was apparently lost during land-colonization. Here, a new qE process evolved reliant on the constitutively expressed protein PsbS4. Land plants, however, seem to carry a remnant of ancestral qE control by blue light, named qM, as blue light still affects NPQ in these organisms, but only via the PHOT-triggered chloroplast avoidance movement that helps chloroplasts to avoid high light intensities26.
The transition from water to land was paralleled by the emergence of a second PHOT gene. Seed plants express both PHOT1, a protein that responds to low-intensity blue light signals, and PHOT2, a protein that triggers high-intensity blue light responses. Unicellular green algae such as Chlorella variabilis, Ostreococcus tauri and C. reinhardtii only contain one PHOT gene, an orthologue of PHOT2 (ref. 27). However, in C. reinhardtii blue light triggers different biological responses depending on the photon fluence rate. Low-intensity blue light (for example, 1 μmol photons m−2 s−1) triggers gametogenesis at low nitrogen abundance28 and gene expression29 whereas blue light at higher intensities desensitizes the eyespot19 and induces LHCSR3 expression (for example, 60 μmol photon m−2 s−1, Extended Data Fig. 10a, b). Thus, we conclude that the same PHOT protein that ensures responses at low light intensities can trigger signalling at higher light intensities when acting in synergy with the photosynthetic signal. We propose that, although the transition from motility (algae) to sessility (land plants) led to an increase in specialization of the function of photoreceptors, this process occurred at the expense of some essential functions in water, probably the regulation of photoprotection that is reported here.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Chemicals
Chemicals were purchased from Sigma (DCMU) and Enzo (3-isobutyl-1-methylxanthine (IBMX), dibutyryl cyclic AMP (DB-cAMP), and dibutyryl cyclic GMP (DB-cGMP)). Stock solutions of DCMU were prepared in ethanol (40 mM) or H2O (40 μM); IBMX was dissolved in DMSO at 250 mM; DB-cAMP and DB-GMP were dissolved in H2O.
Strains and conditions
Chlamydomonas reinhardtii strains were grown under 20 μmol photons m−2 s−1 in Tris-acetate-phosphate (TAP) media30 at 23 °C. In all experiments cells were transferred to Sueoka's high salt medium31 at 2 million cells ml−1 and exposed to light intensities as described in the text and figure legends. Two C. reinhardtii wild-type strains, cw15-302 and 137c, were used as indicated in the figure legends. The phot mutant was previously generated using a homologous recombination strategy17. For the different complemented lines of phot, PsaD promoter and terminator were used for the expression of full-length PHOT cDNA (strain PPHOT; clone PPHOT3 or PPHOT4), the LOV domains (strain PLOV; clone PLOV1) and the kinase domain (strain PKIN; clone PKIN1), N-terminally fused to the zeocin resistance marker sh-Ble19. Using the same strategy, overexpression of the PHOT kinase domain in the wild-type background resulted in the strain WTKIN19. For the PHOT-independent expression of LHCSR3, the PsaD promoter and terminator were fused to a full-length genomic LHCSR3.1 gene. The generated construct with the APH7 marker gene was introduced into phot by electroporation with NEPA21 Super Electroporator (Nepa Gene). The transformants, PLHCSR3(B5) and PLHCSR3(G6), were screened for their resistance to 10 μg ml−1 hygromycin. The acry mutant, originally delivered in a SAG73.72 genetic background16, was backcrossed three times to wild-type strain 137c. Unless otherwise stated, low-light-intensity conditions corresponded to 20 μmol photons m−2 s−1 while high-light-intensity conditions corresponded to 250 μmol photons m−2 s−1 of white light. All experiments were repeated three times on different biological samples to verify their reproducibility, unless otherwise stated.
Pigment and mRNA quantification
Extraction and quantification of chlorophyll and xanthophylls, calculation of the de-epoxidation state DES, and mRNA quantification by quantitative PCR were performed as described previously14,32. A gene encoding G-protein-subunit-like protein (GBLP)33 was used as the endogenous control, and relative expression values relative to GBLP (Figs 2b and 3c) were calculated from three biological replicates, each of which contained three technical replicates. The primers used were LHCSR3.1 (5′-CACAACACCTTGATGCGAGATG-3′ and 5′-CCGTGTCTTGTCAGTCCCTG-3′), LHCSR3.2 (5′-TGTGAGGCACTCTGGTGAAG-3′ and 5′-CGCCTGTTGTCACCATCTTA-3′), GBLP (5′-CAAGTACACCATTGGCGAGC-3′ and 5′-CTTGCAGTTGGTCAGGTTCC-3′), and 18S rRNA (5′-AGCATGAGAGATGGCTACCACATC-3′ and 5′-CATTCCAATTACCAGACGCGAAGC-3′). The mRNA stability experiments were performed in the presence of the transcription inhibitor actinomycin D (Wako) at 160 μg ml−1 (ref. 34). Reverse-transcription PCR was performed using Light cycler 96 (Roche) with AptaTaqDNA GM with ROX (Roche).
Immunoblotting
Protein samples of whole cell extracts (0.5 μg chlorophyll, unless stated otherwise) were loaded on 7% or 13% SDS–PAGE gels and blotted onto nitrocellulose membranes. Antisera against D1, D2 and ATPB were obtained from Agrisera; antisera against C. reinhardtii aCRY16, PHOT (LOV1 domain)17, and all others35,36 were previously described. ATPB was used as a loading control. An anti-rabbit horseradish-peroxidase-conjugated antiserum was used for detection. The blots were developed with ECL detection reagent and images of the blots were obtained using a CCD imager (ChemiDoc MP System, Bio-Rad). For the densitometric quantification of LHCSR3, data were normalized to ATPB. LHCSR3 appears as a double band in some of the western blots (that is, Fig. 3 and Extended Data Figs 2, 4, 6, 7, 8). The upper band represents the phosphorylated form of LHCSR3 (see Extended Data Fig. 6). The extent of LHCSR3 phosphorylation was determined using a Phos-tag-based method as described37. For the dephosphorylation of LHCSR3, whole-cell extracts (1 μg total chlorophyll) were treated with either 20 U of calf intestine phosphatase (Promega) or 200 U of λ protein phosphatase (λPP) (New England Biolabs) in the presence of 0.05 (v/v) Triton X-100, at 30 °C for 1 h.
Fluorescence-based measurements
Fluorescence-based photosynthetic parameters were measured with a fluorescence imaging setup previously described38. The photosynthetic electron transfer rate was calculated as ETR = (Fm′ − F)/Fm ′× 0.84 × 0.5 × I (ref.39). qE was estimated as the fraction of NPQ that is rapidly inducible in the light and reversible in the dark, using the following equation: qE = (Fm − Fm′)/Fm′. where F and Fm′ are the fluorescence yields in steady state light and after a saturating pulse in the actinic light, respectively; Fm is the maximal fluorescence yield in dark-adapted cells; and I is the light irradiance in μmol photons m−2 s−1 (ref. 40). Before NPQ measurements, cells were exposed to high-intensity light for 4 h unless stated to induce LHCSR3 and dark acclimated for 30 min. For NPQ measurements, actinic light was set at 750 μmol photons m−2 s−1. For action spectra measurements, 5 ml of concentrated cells (2 × 107 cells ml−1) were placed in a Petri dish and exposed to intense (250 μmol photons m−2 s−1) monochromatic light (the spectral half width is 5.5 nm or less), provided by the Okazaki Large Spectrograph15 for 4 h. Samples were then collected and subjected to immunoblotting (0.1 ml) as well as qE measurement (0.2 ml) with a fluorescence video-imaging system (Fluorocam, Photon System Instruments).
References
Jiao, Y., Lau, O. S. & Deng, X. W. Light-regulated transcriptional networks in higher plants. Nat. Rev. Genet. 8, 217–230 (2007)
Christie, J. M. Phototropin blue-light receptors. Annu. Rev. Plant Biol. 58, 21–45 (2007)
Briggs, W. R. Phototropism: some history, some puzzles, and a look ahead. Plant Physiol. 164, 13–23 (2014)
Li, Z., Wakao, S., Fischer, B. B. & Niyogi, K. K. Sensing and responding to excess light. Annu. Rev. Plant Biol. 60, 239–260 (2009)
Eberhard, S., Finazzi, G. & Wollman, F. A. The dynamics of photosynthesis. Annu. Rev. Genet. 42, 463–515 (2008)
Horton, P., Ruban, A. V. & Walters, R. G. Regulation of light harvesting in green plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 655–684 (1996)
Niyogi, K. K. & Truong, T. B. Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis. Curr. Opin. Plant Biol. 16, 307–314 (2013)
Peers, G. et al. An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462, 518–521 (2009)
Tokutsu, R. & Minagawa, J. Energy-dissipative supercomplex of photosystem II associated with LHCSR3 in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 110, 10016–10021 (2013)
Ruban, A. V. et al. Identification of a mechanism of photoprotective energy dissipation in higher plants. Nature 450, 575–578 (2007)
Ahn, T. K. et al. Architecture of a charge-transfer state regulating light harvesting in a plant antenna protein. Science 320, 794–797 (2008)
Finazzi, G. & Minagawa, J. in Non-PHOTochemical Quenching and Energy Dissipation in Plants, Algae and Cyanobacteria (eds Demmig-Adams, B., Garab, G., Adams, W. III & Govindjee ) Ch 21, 445–469 (Springer Netherlands, 2014)
Petroutsos, D. et al. The chloroplast calcium sensor CAS is required for photoacclimation in Chlamydomonas reinhardtii. Plant Cell 23, 2950–2963 (2011)
Maruyama, S., Tokutsu, R. & Minagawa, J. Transcriptional regulation of the stress-responsive light harvesting complex genes in Chlamydomonas reinhardtii. Plant Cell Physiol. 55, 1304–1310 (2014)
Watanabe, M. et al. Design and performance of the Okazaki large spectrograph for photobiological research. Photochem. photobiol. 36, 491–498 (1982)
Beel, B. et al. A flavin binding cryptochrome photoreceptor responds to both blue and red light in Chlamydomonas reinhardtii. Plant Cell 24, 2992–3008 (2012)
Zorin, B., Lu, Y., Sizova, I. & Hegemann, P. Nuclear gene targeting in Chlamydomonas as exemplified by disruption of the PHOT gene. Gene 432, 91–96 (2009)
Merchant, S. S. et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245–250 (2007)
Trippens, J. et al. Phototropin influence on eyespot development and regulation of phototactic behavior in Chlamydomonas reinhardtii. Plant Cell 24, 4687–4702 (2012)
Kong, S.-G. et al. The C-terminal kinase fragment of Arabidopsis phototropin 2 triggers constitutive phototropin responses. Plant J. 51, 862–873 (2007)
Harada, A., Sakai, T. & Okada, K. phot1 and phot2 mediate blue light-induced transient increases in cytosolic Ca2+ differently in Arabidopsis leaves. Proc. Natl Acad. Sci. USA 100, 8583–8588 (2003)
Babourina, O., Newman, I. & Shabala, S. Blue light-induced kinetics of H+ and Ca2+ fluxes in etiolated wild-type and phototropin-mutant Arabidopsis seedlings. Proc. Natl Acad. Sci. USA 99, 2433–2438 (2002)
Folta, K. M. & Kaufman, L. S. Phototropin 1 is required for high-fluence blue-light-mediated mRNA destabilization. Plant Mol. Biol. 51, 609–618 (2003)
Kirilovsky, D. & Kerfeld, C. A. The orange carotenoid protein: a blue-green light photoactive protein. Photochem. photobiol. Sci. 12, 1135–1143 (2013)
Schellenberger Costa, B. et al. Blue light is essential for high light acclimation and photoprotection in the diatom Phaeodactylum tricornutum. J. Exp. Bot. 64, 483–493 (2013)
Cazzaniga, S., Dall’ Osto, L., Kong, S. G., Wada, M. & Bassi, R. Interaction between avoidance of photon absorption, excess energy dissipation and zeaxanthin synthesis against photooxidative stress in Arabidopsis. Plant J. 76, 568–579 (2013)
Galván-Ampudia, C. S. & Offringa, R. Plant evolution: AGC kinases tell the auxin tale. Trends Plant Sci. 12, 541–547 (2007)
Huang, K. & Beck, C. F. Phototropin is the blue-light receptor that controls multiple steps in the sexual life cycle of the green alga Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 100, 6269–6274 (2003)
Im, C. S., Eberhard, S., Huang, K., Beck, C. F. & Grossman, A. R. Phototropin involvement in the expression of genes encoding chlorophyll and carotenoid biosynthesis enzymes and LHC apoproteins in Chlamydomonas reinhardtii. Plant J. 48, 1–16 (2006)
Gorman, D. S. & Levine, R. P. Cytochrome f and plastocyanin: their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardi. Proc. Natl Acad. Sci. USA 54, 1665–1669 (1965)
Sueoka, N. Mitotic replication of deoxyribonucleic acid in Chlamydomonas reinhardii. Proc. Natl Acad. Sci. USA 46, 83–91 (1960)
Allorent, G. et al. A dual strategy to cope with high light in Chlamydomonas reinhardtii. Plant Cell 25, 545–557 (2013)
Schloss, J. A. A Chlamydomonas gene encodes a G protein β subunit-like polypeptide. Mol. Gen. Genet. 221, 443–452 (1990)
Gera, J. F. & Baker, E. J. Deadenylation-dependent and -independent decay pathways for α1-tubulin mRNA in Chlamydomonas reinhardtii. Mol. Cell. Biol. 18, 1498–1505 (1998)
Takahashi, H., Iwai, M., Takahashi, Y. & Minagawa, J. Identification of the mobile light-harvesting complex II polypeptides for state transitions in Chlamydomonas reinhardtii. Proc. Natl Acad. Sci. USA 103, 477–482 (2006)
Iwai, M. et al. Isolation of the elusive supercomplex that drives cyclic electron flow in photosynthesis. Nature 464, 1210–1213 (2010)
Longoni, P., Douchi, D., Cariti, F., Fucile, G. & Goldschmidt-Clermont, M. Phosphorylation of the light-harvesting complex II isoform Lhcb2 is central to state transitions. Plant Physiol. 169, 2874–2883 (2015)
Johnson, X. et al. A new setup for in vivo fluorescence imaging of photosynthetic activity. photosynth. Res. 102, 85–93 (2009)
Petroutsos, D. et al. PGRL1 participates in iron-induced remodeling of the photosynthetic apparatus and in energy metabolism in Chlamydomonas reinhardtii. J. Biol. Chem. 284, 32770–32781 (2009)
Genty, B., Briantais, J.-M. & Baker, N. R. The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochim. Biophys. Acta 990, 87–92 (1989)
Depège, N., Bellafiore, S. & Rochaix, J. D. Role of chloroplast protein kinase Stt7 in LHCII phosphorylation and state transition in Chlamydomonas. Science 299, 1572–1575 (2003)
Bergner, S. V. et al. STATE TRANSITION7-dependent phosphorylation is modulated by changing environmental conditions, and its absence triggers remodeling of photosynthetic protein complexes. Plant Physiol. 168, 615–634 (2015)
Acknowledgements
We thank K. Kamada for help with genetic crossing, Y. Yari Kamrani for help with the Okazaki Large Spectrograph experiments and K. Kosuge for help with immunoblotting experiments. We thank G. Kreimer and Y. Aihara for discussions, M. Hippler for the antibody against LHCSR3 and G. Allorent for help during the initial phase of this project. We also thank P. Longoni and M. Goldschmidt-Clermont for advice and technical assistance with Phos-tag PAGE and T. Yamasaki for assistance with quantitative PCR. This work was supported by grants from Agence Nationale de la Recherche (ANR-12-BIME-0005, GRAL Labex, ANR-10-LABX-49-01, DiaDomOil to D.P. and G.F.); the Marie Curie Initial Training Network Accliphot (FP7-PEOPLE-2012-ITN; 316427 to G.F., D.P., S.F.); the CNRS Défi (ENRS 2013 to G.F. and D.P.); the CEA Bioénergies program (to G.F. and D.P.); the HFSP (HFSP0052) to G.F. and D.P.; the NIBB Cooperative Research Program for the Okazaki Large Spectrograph (13-514 and 14-508 to G.F. and 15-609 and 16-705 to J.M.); JSPS KAKENHI (JP15H05599 to R.T., JP26251033 and JP16H06553 to J.M. and R.T.); the NEDO (P07015 to J.M.); the MEXT (through the Network of Centres of Carbon Dioxide Resource Studies in Plants to J.M.); the German Research Foundation, DFG, grants FOR1261 (to P.H., M.M. and T.K.) and Heisenberg fellowship (to T.K.). L.M. acknowledges support from the Alexander von Humboldt Stiftung/Foundation.
Author information
Authors and Affiliations
Contributions
D.P., R.T., J.M. and G.F. designed the study. D.P., R.T., S.F. and L.M performed biochemical analyses; D.P. and R.T. performed fluorescence measurements; D.P. performed pharmacological tests; D.P., R.T., J.M. and G.F. performed action spectra; R.T. performed mutant generation; R.T. and S.M. performed quantitative PCR analysis; S.F. performed pigment quantification; L.C. performed the photosensitivity tests; J.M. and G.F. performed non-photochemical quenching experiments. D.P., R.T., S.M., A.G., L.M., T.K., M.M., P.H., J.M. and G.F. provided strains and analysed the data. D.P., R.T., J.M. and G.F. wrote the manuscript, and all authors revised and approved it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks J.-D. Rochaix and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Photosynthetic properties of acry and phot mutants.
a, Immunoblot analysis of aCRY accumulation in wild-type (137c) and acry cells. The two strains were grown as previously described16. Cells were harvested at the beginning of the light phase (LD2 phase16). Whole-cell samples with 2 μg of chlorophyll were loaded on each lane. Representative dataset of an experiment replicated 3 times on different biological samples. b, Immunoblot analysis of PHOT in wild-type (cw15-302) and phot cells after 4 h exposure to high-intensity white light. Representative dataset of an experiment replicated 3 times on different biological samples. c, Immunoblot analysis of LHCSR3 and ATPB in wild-type (137c), phot and acry cells in low light intensity and after 4 h exposure to high-intensity white light (HL). ATPB was used as a loading control. Representative dataset of an experiment replicated five times on different biological samples. d, e, NPQ induction kinetics of wild-type (137c) and acry cells (d) and wild-type (cw15-302) and phot cells (e) after 4 h exposure to high-intensity white light. qE was recorded for 16 min upon illumination with 440 μmol photons m−2 s−1 (white bar) followed by 2 min of darkness (black bar), to measure qE relaxation. Representative dataset of an experiment replicated 4 (d) and 7 (e) times on different biological samples. f, g, DES indicates the xanthophyll cycle de-epoxidation state ([zeaxanthin] + 1/2[antheraxanthin])/([zeaxanthin] + [antheraxanthin] + [violaxanthin]) in wild-type (137c) and acry cells (f) and wild-type (cw15-302) and phot cells (g) after exposure to high light intensity for 4 h (n = 3 biological samples, mean ± s.d.). h, i, ETR of wild-type (137c) and acry (h) and wild-type (cw15-302) and phot (i) cells exposed at 20, 240 and 750 μmol photons m−2 s−1 for 10 h. ETR (μmol photons m−2 s−1) was measured at actinic illumination of 41 μmol photons m−2 s−1 (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 2 Accumulation of major photosynthetic complexes is unaltered in the phot cells upon exposure to different wavelengths of visible light.
a, Immunoblot analyses of ATPB, PsaA/B, D1, D2 and CP26 accumulation in wild-type (137c), phot and acry cells after 4 h exposure to 250 μmol photons m−2 s−1 of monochromatic light at the different wavelengths of the visible spectrum. b, Immunoblot analyses of major photosynthetic complexes of PSII, cyt b6f and PSII in wild-type, phot and acry cells after 4 h exposure at 250 μmol photons m−2 s−1 of white light. Representative dataset of an experiment replicated three times on different biological samples.
Extended Data Figure 3 Diminished LHCSR3 induction in phot is not caused by diminished photosynthesis.
a, Comparison of ETR in DCMU-titrated wild-type cells and DCMU-untreated phot cells exposed to high light intensity for 3 h. ETR (μmol photons m−2 s−1) was measured upon exposure to light of 170 μmol photons m−2 s−1 (n = 3 biological samples, mean ± s.d.). b, Immunoblot analyses of LHCSR3 accumulations in the wild-type and phot samples described in a. ATPB was used as a loading control. Representative dataset of an experiment replicated five times on different biological samples.
Extended Data Figure 4 PHOT protein levels pigment content in phot mutants.
a, Immunoblot analyses of PHOT accumulation in wild-type (cw15-302) and PPHOT strains. PPHOT expresses a fused PHOT–BLE protein, which has a higher molecular mass than wild-type PHOT. ATPB served as a loading control. Representative dataset of experiment replicated four times on different biological samples. b, Total cellular chlorophyll (a + b) content in wild-type, phot, PPHOT, PKIN and PLOV cells exposed to light of 20, 200 or 750 μmol photons m−2 s−1 for 20 h as in Fig. 2d (n = 3 biological samples, mean ± s.d.). Diminished chlorophyll content is a signature of pigment bleaching following photo-damage.
Extended Data Figure 5 PHOT-independent LHCSR3 expression restores photoprotection in the phot mutant.
Relative qE and LHCSR3 expression under high-light-intensity conditions in wild-type (cw15-302), phot, and two phot-transformed lines expressing an additional copy of the LHCSR3.1 gene under the control of the PsaD promoter (PLHCSR3(B5) and PLHCSR3(G6)). ATPB was used as a loading control. qE values were normalized to wild-type cells (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 6 Phosphorylation levels of LHCSR3 in phot, PPHOT, PLOV, PKIN, stt7 and wild-type cells.
Assessment of the phosphorylation levels of LHCSR3 in wild-type (cw15-302), phot, PPHOT, PLOV and PKIN cells by a mobility-shift detection of phosphorylated proteins (Phos-tag). The upper and lower bands correspond to the phosphorylated and non-phosphorylated forms of LHCSR3 (P-LHCSR3 and LHCSR3), respectively, as confirmed by treatment of the samples with either calf intestinal phosphatase (CIP) or λPP. The stt7 mutant41 was used to test the involvement of the chloroplastic serine/threonine kinase STT7 in the LHCSR3 phosphorylation. In this mutant, LHCSR3 was mostly present in the non-phosphorylated form, while an STT7-independent phosphorylation was also seen in agreement with recent findings42. Representative dataset of an experiment replicated three times on different biological samples.
Extended Data Figure 7 Phenotypic traits of the WTKIN genotype.
a, Immunoblot analyses of LHCSR3 accumulation after exposure to 240 μmol photons m−2 s−1 of white light for 20 h in wild-type (137c) and WTKIN cells. ATPB was used as a loading control. b, Erlenmeyer flasks containing wild-type and WTKIN (expressing the kinase domain of PHOT in the wild-type background) after 20 h exposure to white light of 20, 200 and 750 μmol photons m−2 s−1 and schematic drawings of the PHOT gene constructs in the two lines. Representative pictures from an experiment replicated three times on different biological samples. c, Action spectrum of LHCSR3 accumulation in wild-type and WTKIN. ATPB was used as a loading control. d, Densitometric quantification of LHCSR3 accumulation in wild-type (137c) and WTKIN (data normalized to ATPB). e, LHCSR3 protein accumulation in WTKIN at low and high light intensity in the absence and presence of the PSII inhibitor DCMU. Representative dataset of experiment replicated three times on different biological samples.
Extended Data Figure 8 Second messengers involved in the PHOT-dependent regulation of LHCSR3 expression.
a, Immunoblotting analyses of LHCSR3 accumulation after exposure to high-intensity white light for 20 h in wild-type (cw15-302) and phot cells under control conditions (0.34 mM Ca2+) or in the presence of increased Ca2+ concentration (3.4 mM Ca2+). Representative dataset of an experiment replicated three times on different biological samples. b, Statistical analyses of LHCSR3 accumulation in high-light-intensity-treated wild-type or phot cells in the absence (control) and presence of IBMX, DB-cGMP and DB-cAMP (see conditions as in Fig. 4a). Data are normalized to LHCSR3 levels of wild-type control cells (n = 5 biological samples, mean ± s.d.). Asterisks indicate statistically significant difference from wild-type control cells (t-test, P < 0.0001)
Extended Data Figure 9 LHCSR3.1 and LHCSR3.2 transcript stability in wild-type and phot cells.
Relative amounts of LHCSR3.1 and LHCSR3.2 mRNA in wild-type (cw15-302; black) and phot (red) cells were quantified by quantitative PCR. Cells exposed for 20 h to 470 nm LED light at 100 μmol photons m−2 s−1 were transfered to darkness at t = 0 and treated with actinomycin D to stop further mRNA synthesis. mRNA samples were collected at 0, 0.25, 0.5, 1, 2 and 4 h after transition to darkness. LHCSR3.1 and LHCSR3.2 transcript amounts were normalized to the amounts of 18S rRNA as endogenous control and their values were set at 100% at t = 0 (n = 3 biological samples, mean ± s.d.).
Extended Data Figure 10 qE and LHCSR3 induction requires high light in C. reinhardtii.
a, Action spectrum of qE induction in wild-type cells as a function of the light intensity. b, Immunoblot analysis of LHCSR3 accumulation in darkness (D) and under white (WL), blue (470 nm) and red (660 nm) light of low (20 μmol photons m−2 s−1) and high intensity (250 μmol photons m−2 s−1). ATPB was used as a loading control. Representative dataset of an experiment replicated two times on different biological samples.
Rights and permissions
About this article
Cite this article
Petroutsos, D., Tokutsu, R., Maruyama, S. et al. A blue-light photoreceptor mediates the feedback regulation of photosynthesis. Nature 537, 563–566 (2016). https://doi.org/10.1038/nature19358
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19358
- Springer Nature Limited
This article is cited by
-
Macroscale structural changes of thylakoid architecture during high light acclimation in Chlamydomonas reinhardtii
Photosynthesis Research (2024)
-
Small Papillae Regulated by SPD25 are Critical for Balancing Photosynthetic CO2 Assimilation and Water Loss in Rice
Rice (2023)
-
Proteomic characterization of a lutein-hyperaccumulating Chlamydomonas reinhardtii mutant reveals photoprotection-related factors as targets for increasing cellular carotenoid content
Biotechnology for Biofuels and Bioproducts (2023)
-
Light-independent regulation of algal photoprotection by CO2 availability
Nature Communications (2023)
-
Widening the landscape of transcriptional regulation of green algal photoprotection
Nature Communications (2023)