Abstract
Many hybrid devices integrate functional molecular or nanoparticle components with microstructures, as exemplified by the nanophotonic devices that couple emitters to optical resonators1 for potential use in single-molecule detection2,3, precision magnetometry4, low threshold lasing5,6 and quantum information processing7,8,9,10,11,12. These systems also illustrate a common difficulty for hybrid devices: although many proof-of-principle devices exist, practical applications face the challenge of how to incorporate large numbers of chemically diverse functional components into microfabricated resonators at precise locations. Here we show that the directed self-assembly13,14 of DNA origami15 onto lithographically patterned binding sites allows reliable and controllable coupling of molecular emitters to photonic crystal cavities (PCCs). The precision of this method is sufficient to enable us to visualize the local density of states within PCCs by simple wide-field microscopy and to resolve the antinodes of the cavity mode at a resolution of about one-tenth of a wavelength. By simply changing the number of binding sites, we program the delivery of up to seven DNA origami onto distinct antinodes within a single cavity and thereby digitally vary the intensity of the cavity emission. To demonstrate the scalability of our technique, we fabricate 65,536 independently programmed PCCs on a single chip. These features, in combination with the widely used modularity of DNA origami16,17,18,19,20, suggest that our method is well suited for the rapid prototyping of a broad array of hybrid nanophotonic devices.
Similar content being viewed by others
Main
Solution-synthesized molecules and colloidal nanoparticles offer unique optical properties, such as tunable photoluminescence (PL) spectra, which are difficult to achieve in the materials we can most easily craft into microfabricated optical cavities. Thus hybrid nanophotonic systems, in which the strongly localized optical fields within microfabricated cavities further enhance the optical properties of molecules and nanoparticles1 or create new properties such as lasing5,6 or nonlinear phenomena8, are of interest.
To reliably build such systems, it is necessary to have a fabrication method which can both introduce well defined numbers (often exactly one) of optically active components into a single cavity, and position those components relative to the antinodes of the cavity mode with a precision that is well below the wavelength of light (tens of nanometres). Further, advanced hybrid devices would organize two or more kinds of chemically diverse subcomponents—organic molecules, nitrogen–vacancy centres, quantum dots, ions, or metallic nanoparticles—in intimate contact at the length scale of the subcomponents (0.1–10 nm). Finally, whether the hybrid device is imagined to be part of a hand-held biomedical diagnostic machine, or part of a quantum circuit embedded within a classical computer which controls it, large numbers of devices must be scalably integrated on chips, using CMOS-compatible techniques.
A complete solution for hybrid device fabrication would address all of the above challenges, achieving control of component number within a cavity, precise positioning in a cavity mode, modular incorporation of diverse materials, finer subcomponent-scale arrangement, and large-scale integration. No current technique suffices. Three approaches are used to integrate heterogeneous components into microstructures: (1) fully stochastic assembly, (2) fully deterministic scanning-probe assembly, and (3) directed self-assembly. In (1), fully stochastic assembly6,7,8,9, molecules or nanoparticles are either deposited or grown at random locations on a substrate. When resonator locations are predefined8, the yield of functional devices in which a single component has landed within a resonator is limited by Poisson statistics to 37%, but yields can be lower since positioning within the cavity is uncontrolled. Alternatively, via ‘select and post-process’1, randomly arranged components can be mapped using atomic force microscopy (AFM), scanning electron microscopy (SEM), or super-resolution fluorescence microscopy and a resonator can be built around the component7,9. Scanning probe assembly (2) involves pushing components into a resonator using the scanning probe tip, one at a time10,11. Neither fully stochastic nor scanning-probe approaches can be scaled up. In contrast, directed self-assembly (3) uses lithographically defined growth sites or binding sites to localize components to microstructures with high probability, and is thus inherently scalable. But while previous directed self-assembly techniques control emitter number and position12, they do not generalize to other component types, or address subcomponent organization.
Structural DNA nanotechnology21 can create organic/inorganic hybrid components at a resolution unachievable by top-down fabrication. Particularly versatile in this regard is DNA origami15,22, in which a long single strand of DNA is folded by the concerted action of hundreds of much shorter DNA ‘staple strands’: coupling diverse subcomponents to the staple strands yields modular DNA ‘breadboards’ that carry up to 200 different elements (these can range from organic dyes16,17, metal nanoparticles16,17, nitrogen–vacancy centres23 and quantum dots17,18,19 to carbon nanotubes20 and proteins), simultaneously juxtaposed16,17,18,19 with 3–5 nm resolution. Large-scale integration of the functionalized DNA shapes can then be achieved through electrostatic immobilization onto ~100-nm binding sites that have been lithographically defined on desired substrates. When optimized, this ‘placement’ technique13,14 successfully positions single origami at 94% of binding sites with ~20-nm precision. Together, origami and placement thus meet all five of the above challenges in the context of a general hybrid nanophotonics platform: the former provides modularity and fine subcomponent-scale organization, the latter control over component number, positioning and scalability. Below we give examples in which origami and placement work together to control the emission of thousands of silicon nitride (SiN) PCCs.
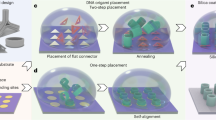
Solution-synthesized DNA origami triangles (Fig. 1a and Extended Data Fig. 1) serve both as components and adaptors, carrying either 3 or 15 cyanine dye subcomponents (Cy5, Fig. 1b) to triangular binding sites within PCCs. To form the origami, a long circular scaffold strand (the genome of bacteriophage M13mp18) is annealed (90–20 °C, 6 h) with ~200 short computer-designed staple strands, and a Cy5-labelled subcomponent strand (grey strands with a red dot) which binds to linkers (black) projecting from a subset of the staple strands. Figure 1c outlines the fabrication steps (details in Extended Data Fig. 2) used to create both binding sites and PCCs from a 275-nm-thick SiN layer (purple) on a Si wafer (grey). Binding sites are created (steps 1 and 2) via electron beam (e-beam) patterning of negatively charged carboxylate groups (red, which bind negatively charged origami strongly via positively charged Mg2+ ions from solution) within a background of hydrophobic methyl groups (blue, which bind origami poorly). Directed self-assembly proceeds by incubating a solution of purified DNA origami over the patterned substrate which results in a high yield of single origami binding events. Figure 1d depicts placement reoptimized for SiN, which achieves 98% single-origami occupancy. PCCs based on previous designs24 are fabricated after the binding sites via additional e-beam and etching (steps 3–6). Each cavity (Fig. 1e) is simply a row of three missing holes within a two-dimensional (2D) triangular lattice of air holes. In 2D, light is confined to the cavity because the spacing of the air holes creates an optical bandgap. Vertical confinement is achieved by total internal reflection at the interface of the SiN membrane and the air above and below. Cavity geometry was optimized using finite-difference time domain (FDTD) simulations to set up a high-quality mode (Fig. 1f) between 655 nm and 660 nm, near the emission peak of Cy5.
a, A single-stranded M13mp18 scaffold is annealed with staples to form origami triangles with single-stranded linkers (black squiggles) for the attachment of subcomponents. b, Fluid AFM of origami without (leftmost image) and with (rightmost image) subcomponent strands (middle, grey squiggles) labelled with Cy5 (red oval or red dots) bound to linkers (black squiggles). The inset spectrum shows Cy5’s broad emission, centred at 670 nm). c, Fabrication of a single binding site (red), within a passivation layer (blue), followed by construction of a photonic crystal cavity (PCC) around it (details in Extended Data Fig. 2). d, Diagram shows test substrate for placement fidelity (without PCCs), and the groups which mediate binding (carboxylate), and non-binding (methyl). AFM images show binding sites before (left) and after (right) placement; 24 of 25 sites have a single origami (full test substrate: 600 sites, 98% single origami). e, SEM of a PCC; a = 256 nm, r/a = 0.3, r1/a = 0.2, r2/a = 0.25, s = 0.22a. f, Low resolution PCC reflectance spectra (black) is compared to FDTD prediction (red); fundamental mode ωc agrees within 1 nm, lower Q is attributed to fabrication defects24. Inset, high resolution fit to ωc (blue). Scale bars: b, 50 nm; d, 400 nm; e, 500 nm.
A large variation in cavity emission, as a function of Cy5-origami position, highlights the importance of emitter placement in both theory and experiment (Fig. 2, Extended Data Fig. 4). Because emitters are located on the 2D cavity surface, total spontaneous emission ST from a Cy5 at location r = (x, y) for wavelength ω has three components11: direct emission into free space, indirect emission into the cavity (eventually scattered into free space by imperfect confinement), and interference between the two. The measurement set-up (Extended Data Fig. 5) sets coefficients Cdir, Ccav and Cint, which govern the relative contribution of each component to the total spontaneous emission:

where the enhancement of emission into the cavity mode, P(r, ω), has a Lorentzian line shape centred about the cavity resonance ωc, and an interference factor I(r, ω) accounts for the interference between direct and indirect emission at the collection point. Because photons are coupled multimodally through free space into the monochromator, and because Cy5-origami carry multiple emitters, we neglect Cint and assume that any observed dependence on emitter position is due to P(r, ω). In particular, P(r, ωc), the Purcell factor, is proportional to the local density of states (LDOS) at r, which exhibits strong periodic features with a spacing below the diffraction limit (116 nm from central maxima to adjacent minima; see simulations in Figs 2c and 3b). To measure these features experimentally, PL spectra (Fig. 2b) were recorded for a series of isolated cavities each with a Cy5-origami positioned at one of 21 different x-offsets along their horizontal midlines (Extended Data Fig. 4a). Oscillations were less sharp than expected, but predicted antinodes were still prominent and emission from single Cy5-origami varied up to fourfold (Fig. 2c, data are Lorentzian fits to spectra). For comparison, we modelled random placement by creating PCCs filled with tightly packed origami (Extended Data Fig. 4b). Reference patches with the same average number (26) of Cy5-origami were constructed on open SiN. After measuring spectra (Extended Data Fig. 4c) for five copies of each, we computed the ratio of enhanced to unenhanced emission; randomly placed emitters experience only a 1.5 enhancement, on average. Thus, placement of emitters is important to realize both maximal and reproducible emitter–cavity coupling.
a, Dry AFM of PCCs with a single origami (n = 15 Cy5/origami) at different x-offsets; dots marks cavity centres. Scale bars, 250 nm. b, PL spectra from PCCs with x-offsets from −450 nm to +450 nm, with ∆x of 45 nm. c, Comparison of background-subtracted and normalized PL peak emission from spectra in b with the normalized LDOS calculated via FDTD. Intensity error bars are ±1 standard deviation for 6–8 measurements; x-offset error bars are ±1 standard deviation for 2–3 post-fabrication AFM measurements, averaging ±28 nm.
a, Top, the x–y location of a Cy5-origami (left) within a single PCC (centre) translates to that PCC’s X–Y location within a 40 × 15 PCC array (right; 176 μm × 77 μm). Dashed rectangle in left image shows the area over which origami position was varied, in increments of ∆x = 26.5 nm and ∆y = 22.5 nm. Bottom, AFM of four cavities; outline colours indicate positions in array shown at top right. b, Simulated LDOS for a single PCC. c, Wide-field epifluorescence microscopy average of five copies of the PCC array. d, Comparison of b to c along their horizontal midlines; data (black) normalized to simulated LDOS (red) at central antinode. Scale bars: b, 200 nm; c, 32 μm.
The ability to spectrally characterize single cavities (Fig. 2b) is important, but for mode mapping (Fig. 2c) this approach is tedious and sensitive to alignment errors. A variety of scanning probe10,11,25,26,27 and cathodoluminescence techniques28 have been used to map microcavity modes in 1D10,11 and 2D25,26,28, but they require complex instrumentation. By fabricating a 2D array of PCCs whose microscale X–Y coordinates reflect the nanoscale x–y coordinates of a Cy5-origami within each cavity (Fig. 3a), wide-field epifluorescence microscopy (Fig. 3c) affords direct super-resolution visualization of the 2D LDOS (Fig. 3b). Although the predicted subdiffraction mode pattern is clearly visible in an image of a single array (Extended Data Fig. 6a), Fig. 3c shows an average of five arrays with better signal-to-noise. A 1D slice through the map (Fig. 3d) matches the simulated LDOS better than data in Fig. 2c (perhaps due to better e-beam alignment for close-packed PCC arrays), and emission varies by >5-fold. Resolution is limited by the spatial arrangement of emitters on the origami, the precision of placement, and the resolution with which e-beam lithography can maintain registration between the micrometre-scale PCCs. Here emitters lie equally spaced along the ~50 nm (<λ/10) inner edge of origami triangles and so emitter arrangement dominates other factors, but designs with more densely arranged emitters should allow the method to be tested down to 10-nm resolution.
We have shown that origami placement in PCCs enables super-resolution mapping of cavity modes, but the method’s implications are much broader. Our expanded-scale mode map (Fig. 3c) relies on the reproducible fabrication of 3,000 devices with precise emitter–cavity coupling, and its success implies the ability to create devices with arbitrary coupling, up to the maximum available within a single antinode. To show that we could further engineer emission through coupling to multiple antinodes, we created cavities for which a variable number of origami components, from one to seven, were positioned within the seven strongest antinodes of the cavity (Fig. 4a). Device reproducibility was assessed by creating 64 × 64 arrays of cavities, each with eight 512-cavity sub-arrays having the same number (from 0 to 7) of origami positioned within them. To demonstrate subcomponent-scale organization, we repeated the experiment for two different numbers and sublithographic arrangements of dyes (Fig. 4b, n = 3 Cy5 and n = 15 Cy5). Predicted antinode variability was small (~15% peak intensity difference between the central and six surrounding antinodes) and so a roughly linear relationship between emission and origami number was expected and observed. This allowed us to use sixteen 64 × 64 PCC arrays on the same chip to recognizably approximate a 65,536-pixel grayscale image (Extended Data Fig. 9) with eight intensity levels (Fig. 4c).
a, AFM and schematics show that up to seven origami can be placed in distinct antinodes of a single PCC. b, Epifluorescence images show test patterns of each of the seven configurations from a plus an array of zero-site controls for n = 3 and n = 15 Cy5/origami. 240 copies of each configuration are shown, see Extended Data Fig. 7 for all 512 copies studied. Plots show means from Gaussian fits to intensity histograms (Extended Data Fig. 8); error bars are ±1 standard deviation. c, Van Gogh’s The Starry Night approximated with 65,536 cavities each having from zero to seven binding sites. See Extended Data Fig. 9 for details.
More quantitatively, single-origami devices in Fig. 4b had emission variabilities of ±22.6% (that is, ±1 coefficient of variation, which is the standard deviation divided by the mean) for 3 Cy5 and ±10.9% for 15 Cy5. We modelled device emission as having a binomially distributed component, governed by the number n of Cy5 and the fraction p of functional Cy5 (simultaneously capturing the purity of Cy5-labelled strands, coupling efficiency of these strands to origami, and fraction of unbleached Cy5) multiplied by a Gaussian component, simultaneously capturing both placement and fabrication errors, which should be independent of n. Minimizing squared error between modelled and observed device variability for n = 3 and n = 15 simultaneously suggests that p = 87.2% of emitters were coupled and intact, and Gaussian noise was 4.5% (±1 standard deviation). However, the effect of Gaussian noise on total variability was small (<10%): assuming a placement and fabrication error of 0%, and solving for p in  , where np(1 − p) is the variance and np the mean of the binomial distribution, gives similar values for both 3 emitters (86.7%) and 15 emitters (85%). We did not explore n = 1 because of bleaching and instrument sensitivity, but this analysis suggests that with longer-lived emitters, 85% yield for single-photon light sources could be easily achieved.
, where np(1 − p) is the variance and np the mean of the binomial distribution, gives similar values for both 3 emitters (86.7%) and 15 emitters (85%). We did not explore n = 1 because of bleaching and instrument sensitivity, but this analysis suggests that with longer-lived emitters, 85% yield for single-photon light sources could be easily achieved.
Here we have used DNA origami as modular adaptors to quickly switch between two different tightly distributed numbers of emitting subcomponents (achieving 2.6 ± 0.6 for n = 3 and 13 ± 1.4 for n = 15; ± 1 standard deviation) simply by changing a few DNA strands. Because any material that can be attached to DNA can now be coupled to resonators, diverse applications can be explored. For label-free (emitter-less) single-molecule detection2,3, origami modularity will allow specificity to be switched quickly between small molecules, proteins and nucleic acids. For applications in quantum information or magnetometry which require emitters with better photophysics, recent advances in nanodiamond nitrogen29 and silicon30 vacancy centres may provide suitable emitters. Even without emitter improvements, hybrid nanophotonics based on origami placement will enable the cavity amplification of many phenomena, ranging from plasmonic enhancement16 and lifetime engineering19 (where origami provide metal nanoparticle and emitter integration) to superradiance and low-threshold lasing5,6 (where origami can tightly control emitter number and density). Beyond hybrid nanophotonics, our work may apply to hybrid nanoelectronics20, and to any heterogeneous fabrication wherein molecular or nanoparticulate components are integrated with microstructures—the end of low-yield methods such as ‘select and post-process’ is in sight.
Methods
DNA origami
Design. A variation of the ‘sharp triangle’ design described previously15 was chosen because the sharp triangle is rigid and it has a low tendency to aggregate. The ‘sameside sharp triangle’ used here (details of design in Extended Data Fig. 1) has the same scaffold path as the original sharp triangle, and the scaffold strand shares the same alignment as in the original structure. (For sequences and the caDNAno design, unzip Supplementary Data: specifically, files ‘ETSamSide_15Cy5_StapleList.xls’ and ‘Sameside-sharp-triangle-bridged.json’ which unzip into the directory ‘OrigamiDesignFiles’.) The difference between the original sharp triangle and the sameside sharp triangle lies in the pattern of nicks along the phosphate backbone of the staple strands: in the original sharp triangle, nick positions alternate between the two faces of the triangle, whereas the new sameside sharp triangle has all nick positions on the same face of the triangle. Additionally, 15 staples on the inner edge of the triangle occur in two versions. The first version is an unmodified staple. The second version has an 18-nucleotide poly-T extension, which serves as a linker that binds to a 21-nucleotide poly-A ‘subcomponent’ strand bearing a single Cy5 on its 5′ end. Using this basic design we prepared two version of the origami: one had n = 15 Cy5 molecules along its inner edge and the second had only n = 3 Cy5 in along its inner edge.
Synthesis. Staple strands (Integrated DNA Technologies, 100 μM each in water) and the scaffold strand (single-stranded M13mp18, Bayou Biolabs, P-107) were mixed together to target concentrations of 100 nM (each staple) and 40 nM, respectively (a 2.5:1 staple:scaffold ratio) in 10 mM Tris Base, 1 mM EDTA buffer (adjusted to pH 8.35 with HCl) with 12.5 mM magnesium chloride. (We refer to the buffer used for origami synthesis, with the above combination of Tris Base, EDTA, and Mg2+, as ‘TE/Mg2+’.) Staples with the poly-T linker and the Cy5 label were added at 5× and 150× excess respectively. 50 μl volumes of staple/scaffold mixture were heated to 90 °C for 5 min and annealed from 90 °C to 20 °C at −0.2 °C min−1 in a PCR machine. We used 0.5 ml DNA LoBind tubes (Eppendorf) to minimize loss of origami to the sides of the tubes.
Purification. Since a high concentration of excess staples interferes with origami placement, the synthesized origami were purified away from excess staples using 100 kD molecular weight cut-off spin filters (‘Amicon Ultra-0.5 Centrifugal Filter Units with Ultracel-100 membranes’). By the following protocol, recovery is generally 40%–50% and staples are not visible by agarose gel. Wet the filter by adding 500 μl TE/Mg2+. Spin filter at 2,000 r.c.f. (relative centrifugal force) for 6 min at 4 °C, until the volume in the filter is 50 μl. Discard the filtrate. Add 50 μl of unpurified origami and 400 μl TE/Mg2+. Spin at 2,000 r.c.f. for 6 min at 4 °C. Wash the origami three more times by discarding the filtrate, adding 450 μl TE/Mg2+ and spinning at 2,000 r.c.f. for 6 min at 4 °C. Invert the filter onto a clean tube and spin at 2,000 r.c.f. for 6 min at 4 °C to collect purified origami (∼50 μl). Total time for this purification is roughly 40 min.
All of the work reported in this Letter was performed with spin-column purified origami. Other protocols, which may give higher yield via PEG precipitation or magnetic bead separation, are available31,32 although we have not verified that these methods would give quantitatively similar results for placement.
Quantification. Post-purification, origami were quantified using a NanoDrop spectrophotometer (Thermo Scientific). We estimated the molar extinction coefficient of the origami triangles as that of a fully double-stranded M13mp18 molecule (ε = 123,735,380 M−1 cm−1; we did not correct for a small single-stranded loop present on one edge of the origami). The typical working concentration for origami during placement was 100 pM, which is too small to be measured with the NanoDrop, so serial dilutions were performed. For all of the experiments performed reported here, we used a single 300 μl stock of 20 nM origami solution that was diluted to 100 pM immediately before each use.
Handling origami. After purification and quantification, it is especially important to use DNA LoBind tubes (Eppendorf) for storage and dilution of low concentration DNA origami solutions. Low dilutions, for example, 100 pM, must be made fresh from more concentrated solutions and used immediately—even overnight storage can result in total loss of origami to the sides of the tube. Addition of large amounts of carrier DNA to prevent origami loss may prevent origami placement, just as excess staples do. We have not yet determined whether other blocking agents such as BSA might both prevent origami loss and preserve placement.
Photonic crystal cavity nanofabrication
We fabricated PCCs similar to previous designs24 using two slightly different approaches. For isolated PCCs occurring in widely spaced arrays, like those which appear in Fig. 2, we suspended the PCC membranes using a front etch. For large, close-packed arrays of PCCs like those which appear in Figs 3 and 4, we suspended the PCC membranes using a back etch. Schematics for both fabrication processes are given in Extended Data Fig. 2. All fabrication was carried out in Caltech’s Kavli Nanoscience Institute.
Widely spaced PCC arrays
A schematic of the fabrication process is shown in Extended Data Fig. 2a; SEM images of these arrays are given in Extended Data Fig. 3. Fabrication began with 275 nm LPCVD (low-pressure chemical vapour deposition) growth of SiN on a 100 mm single-side polished (SSP, 525 ± 25 μm thick) 〈100〉 silicon wafer as the base substrate (Rogue Valley Microdevices). The wafer was cleaned and alignment markers were defined in the SiN layer by e-beam lithography and modified Bosch ICP (inductively coupled plasma) etching. The substrate was cleaned again and silanized with a trimethylsilyl passivation layer by vapour deposition of HMDS (hexamethyldisilazane). Next, binding sites in the shape of a DNA origami were defined at specific locations using the alignment markers (defined a priori) by e-beam lithography. Binding sites were then activated with a short O2 plasma etch and the resist was stripped. Surface silanols on binding sites were converted to carboxylate groups via a second silanization. Lastly, a PCC was defined around the carboxylated binding site by e-beam lithography and modified Bosch ICP etching of the SiN layer. Finally, PCCs were suspended using a XeF2 isotropic etch of the underlying Si layer.
Here we note that all modified Bosch ICP etching of SiN, whether for definition of PCCs or alignment marks, was performed in an ICP-RIE (inductively coupled plasma-reactive ion etcher, Oxford Instruments System 100 ICP 380) at a pressure of 4 mtorr, with a flow rate of 25 sccm for SF6 and 35 sccm for C4F8, an RF generator power of 25 W, an ICP power of 1,000 W, at 4 °C. Further, we note that this modified Bosch process does not involve alternating applications of SF6 and C4F8, but rather introduces them as a mixed gas.
Definition of alignment markers. Substrates were sonicated with isopropanol (IPA) and dried in a stream of N2 to remove any particulate contaminants. Substrates were spin coated with 600 nm of ZEP 520A (10 s, 500 rpm; 10 s, 1,500 rpm; 120 s, 3,000 rpm; ZEP 520A is a high-resolution positive electron beam resist from Zeon Chemicals). Alignment marker patterns were defined by e-beam. Patterns were developed in ZED N50 (Zeon Chemicals electron beam resist developer) for 1 min. Patterns were transferred into the substrate using the modified Bosch process for about 30 min. Remaining resist was stripped with hot (50 °C) n-methyl pyrrolidone (NMP).
Surface passivation with trimethylsilyl groups. Substrates were sonicated with IPA and dried in a stream of N2, to remove any particulate contaminants. Substrates were cleaned with O2 plasma in a Plasmatherm Dual-chamber SLR-720 RIE (reactive ion etcher), with a flow rate of 50 sccm, a pressure of 50 mtorr and a power setting of 80 W (110 W cm−2), for 5 min to create surface silanols. Substrates were dehydrated on a hot plate at 150 °C for 5 min. Substrates were incubated in a 4 l chamber saturated with HMDS vapour (a 20 cm × 20 cm × 10 cm Tupperware with a 90 mm Petri dish in the corner holding 10 ml HMDS) for 20 min. This step resulted in the formation of a monolayer of trimethylsilyl groups on the surface. Substrates were heated on a hot plate at 150 °C for 30 min, to stabilize the silanized surface. It is important to bake the substrate both before and after HMDS vapour-priming to ensure high quality surface silanization and increase the robustness of the monolayer to hydrolysis. Failure to perform either of these steps leads to increased background binding of origami.
Definition of carboxylated binding sites. Binding sites were defined by e-beam lithography on a freshly prepared and silanized substrates as follows. Substrates were spin-coated with a 170–180 nm resist layer of poly(methyl methacrylate) (MicroChem Corp., 950 PMMA, A3) at 2,500 rpm for 90 s. The resist was baked at 180 °C for 30 s. Binding sites were defined in the resist using e-beam lithography with a 100 keV beam at 500 pA current. The dosage ranged from 600 to 850 μC cm−2 and patterns were proximity corrected. The resist was developed for 70 s in a 1:3 solution of methyl isobutyl ketone (MIBK) and isopropanol (IPA). Surface silanols were created in the lithographically defined regions by an O2 plasma etch (Plasmatherm Dual chamber SLR-720 RIE, a flow rate of 20 sccm, a pressure of 20 mtorr and a power setting of 50 W (92 W cm−2), for 18 s). The resist was stripped by sonicating the wafer in bath of NMP at 50 °C for 10 min. The surface silanols that were created by O2 plasma was converted to carboxylate groups by incubating the substrate in a disodium carboxyethylsilanetriol (CTES from Gelest, 25% w/v in water Catalog #SIC2263.0) silanization buffer (0.01% CTES, 10 mM Tris, pH 8.3 for 10 min) followed by sonication in milliQ water. During the last step, ensure that the silanization buffer is at pH 8.3. Higher pH leads to the trimethylsilyl background being hydrolysed and inadvertently carboxylated.
Definition of isolated PCCs. After binding site definition, the substrate was cleaned with IPA. A 600 nm thick layer of ZEP 520A was spin-coated on the substrate and baked at 180 °C for 1 min. PCCs were defined using e-beam lithography with a 100 keV beam at a current of 500 pA. The dosage ranged from 200 to 300 μC cm−2 and patterns were proximity corrected. The exposed resist was developed in ZED N50 for 1 min. PCC structures were created via modified Bosch etch for 9–12 min, depending on the etch rate observed that day. The Si under the PCC was isotropically etched with a custom-built pulsed XeF2 etcher (5 cycles of 20 s each, 500 mtorr XeF2). The resist was stripped by overnight soaking of the chip in a bath of NMP at 50 °C. It is important not to sonicate the substrate after the PCC has been suspended as it leads to the collapse of the PCC membrane.
Close-packed PCC arrays
A schematic of the fabrication process is given in Extended Data Fig. 2b. The process is a modification of procedures in ref. 33 in which extremely large SiN membranes were created (up to 2.4 mm × 2.4 mm). The fabrication process is similar to that described for isolated cavities above, with the exception that PCC arrays were created on a SiN window supported by a thin Si layer (which is removed at the very last step with an XeF2 etch). Fabrication began with double-side polished silicon wafers (DSP, 〈100〉, 380 ± 10 nm thick, University Wafers, Rogue Valley Microdevices) with 275 nm layers of LPCVD-grown SiN on both sides of each wafer.
Etching of large SiN windows. Wafers were cleaned and alignment marker were defined by e-beam lithography and ICP/modified Bosch etch (30 min) in the SiN layers on both sides, using the wafer flat for alignment. Substrates were spin-coated with a 600 nm layer of ZEP 520A on the back side of the wafer. Substrates were spin-coated with ProTEK PSB (Brewer Science) on the top side of the wafer to protect it from contamination and physical scratches. A 650 μm × 650 μm window was defined via e-beam using back-side alignment markers. The resist was developed in ZED N50 for 1 min. The SiN layer (on the back side) was etched for 1 h by the modified Bosch process, to punch through to the Si layer. Si exposed in the previous step was etched for 12 h in 50% KOH solution to a depth of approximately 300–350 μm. This created an SiN window of approximately 400 μm × 400 μm supported by an Si layer less than 80 μm thick. The ProTEK PSB was stripped using hot (50 °C) NMP.
Notes on fabrication. After Si-supported SiN windows were etched, origami binding sites and PCC arrays were defined on the top surfaces using earlier-defined top-side alignment. We note that the alignment error between patterns on the top and bottom of the wafers sometimes reached a few tens of micrometres. However the most crucial alignment errors, between the origami binding site and the PCC array, were limited to tens of nanometres, because both binding sites and PCCs were created using the same set of alignment markers on the top of the wafer. Finally, the thin Si support remaining underneath the PCC array was removed using XeF2 pulsed etching. The suspended membranes are extremely delicate so it is crucial that substrate is handled with care. It is important not to sonicate suspended photonic crystal membranes as this will break them.
Origami placement on PCCs
Here the placement DNA origami on binding sites is mediated by Mg2+ binding to surface carboxylate groups, rather than surface silanols, as in our previous work14. We have observed that the use of carboxylated binding sites allows high-quality origami placement and orientation on SiN at a much lower Mg2+ concentration (15 mM) than that required (35 mM) for binding sites activated only by an O2 plasma (and which we presume are covered with silanols). We have not measured the density of carboxylate groups or silanols in either case, but we suggest that the effect is due to the difference in pKas between these two functional groups: similar surface carboxylate groups34 have a pKa of ∼6, while silanol groups have a pKa of 8.3. Thus binding sites with carboxylate groups should carry a higher negative charge at our working pH of 8.3, they should bind more Mg2+, and they should enable the observed binding of origami at lower Mg2+ concentration.
In addition to decreasing the potential for Mg2+ salt artefacts during drying, the use of carboxylate groups has a further important added benefit. Over the course of the extensive PCC fabrication process, different areas experience many different specific series of treatments, which results in many different surface types. Some of these, for example the inside of the PCC holes or the back side of the PCC membranes, are not passivated with trimethylsilyl groups, and appear to bind some DNA origami at higher Mg2+ concentration. Thus the use of carboxylated binding sites (and hence a lower Mg2+ concentration for placement) decreases nonspecific DNA origami binding and ensures that under our buffer conditions the only locations at which origami can stably bind are the intended binding sites.
Below, we describe the placement protocol in five steps, which apart from the use of a lower Mg2+ concentration and carboxylated binding sites, is similar to that in our previous work14. Photographs in supplementary figure 3 of ref. 14 show how substrates should look at different steps of the placement process. (Note that our previous work uses carboxylation of binding sites at a different step, after origami placement, as part of a method to achieve covalent coupling of origami to substrates.)
Binding. A 50 mm Petri dish was prepared with a moistened piece of laboratory tissue paper to limit evaporation. Solution with 100 pM origami was prepared in ‘placement buffer’ (10 mM Tris, 15 mM Mg2+, pH 8.3) and a 20 μl drop was deposited in the middle of the chip, on top of the PCCs. The chip was placed in the closed, humid Petri dish and the origami solution was allowed to incubate on the chip for 1 h.
Initial wash. After the 1 h incubation, excess origami (in solution) were washed away with at least 8 buffer washes by pipetting 60 μl of fresh placement buffer onto the chip, and pipetting 60 μl off the chip. Each of the 8 washes consisted of pipetting the 60 μl volume up and down 2–3 times to mix the fresh buffer with existing buffer on the chip. This initial wash took about 2 min.
Tween wash. Next, in order to remove origami that were non-specifically bound to the passivated background, the chip was buffer-washed 5 times using a ‘Tween washing buffer’ with 0.1% Tween 20 (v/v in placement buffer). This took about 1 min. Because of the low surface tension of the Tween washing buffer, these washes were difficult to perform: they involve adding 20–40 μl of wash buffer, just enough to cover most of the chip, but not enough to spill over the chip and wet the back side of the chip (this may introduce dust contamination from the Petri dish). After the fifth wash, the chip was left to incubate for 30 min. (It is important to use Tween 20, rather than other surfactants. Tween 80 and SDS, which are two other common surfactants, lead to markedly different results—Tween 80 leads to the total removal of placed origami from the substrate, and SDS does not remove excess origami from the trimethylsilyl background. It is important that chips are not exposed to Tween 20 until after the origami have been deposited. Tween 20 applied before binding completely inactivates the binding sites.)
Final stabilizing wash. Last, the chip was buffer-washed 8 times back into a higher pH ‘stabilizing buffer’ (10 mM Tris, 15 mM Mg2+, pH 8.9) so that origami bound strongly and artefacts were minimized during subsequent drying. This took about 2 min. These washes were relatively high volume (60 μl) and were intended to completely remove the Tween 20. The amount of Tween 20 left was monitored qualitatively by the surface tension of the drop (roughly, by eye). When a 20 μl drop covered roughly the same area as the initially deposited drop, it was assumed that the Tween 20 had been sufficiently removed. In the last wash the chip was left with roughly 20 μl of stabilizing buffer, and was ready for drying.
Drying. Chips were dried by serial dilution into ethanol. The chips were dipped in 50% ethanol in water (v/v%) for 10 s, 75% ethanol in water (v/v%) for 10 s and then 120 s in 90% ethanol in water. Then the chips were air-dried. (If nanoarrays are subjected to ethanol solutions with less than 80% ethanol for more than 60–90 s, origami begin to detach from the surface. After the 90% ethanol immersion, it is necessary to let the samples air-dry rather than using N2 or compressed air, since streaking or other drying artefacts are observed in instances of forced air drying.)
Troubleshooting origami placement. If low binding, high multiple binding, or high background binding artefacts are observed, see the troubleshooting guide in Extended Data Table 1. Below we provide notes emphasizing some of the more important aspects of the placement protocol, which may be helpful in troubleshooting. Do not allow the patterned region with binding sites to dry at any point during the binding step or subsequent buffer washes. Inadvertent dewetting of the binding sites leads to distortion of the origami (causing them to ball up) as well as the formation of salt crystals on top of them. Do not use EDTA in placement, Tween washing, or imaging buffers. It is unnecessary in this context and will change the effective Mg2+ concentration for placement slightly. Make fresh buffer solutions every week. Here and elsewhere in this work, we use buffers at low strength (typically 10 mM) to minimize background binding and to make complete washing into different buffers easier. This means the buffers have low buffering capacity and the pH will decrease (and placement may cease to work, depending on conditions). For example, weak buffers made to read pH 8.35 can lose 0.05 pH units and read pH 8.3 after one week.
AFM characterization
After the cavities were fabricated and the origami were placed within the cavities, we quantified the quality of placement using dry AFM. While the presence of origami (and the fluorophores they carry) can be inferred from the optical experiments, we have used AFM because it provides the ability to unambiguously verify the presence of single origami, as well as measure the precise location of the origami (and hence fluorophores) with respect to the cavity. Unless otherwise specified (images in Fig. 1b were taken under aqueous buffer with a Bruker Fastscan AFM), all AFM images were taken in air in tapping mode with a Dimension Icon AFM/Nanoscope V Scanner (Bruker) using the short, fat cantilever from an SNL probe B (“sharp nitride lever”, 2 nm tip radius, Bruker) resonating at approximately 60 kHz. Imaging origami on suspended resonators was challenging. All images had to be acquired at the lowest possible force in order to reduce tip–sample interaction. We optimized imaging conditions on an unsuspended region near the alignment markers and then moved onto the cavity with precise stage movements. Minimization of imaging force was important: if imaging force was increased, imaging became unstable. We noticed that imaging near the edge of suspended regions was easier than at the middle; we suggest that the suspended SiN membrane is interfering with AFM measurement by acting as a mechanical resonator itself.
Each AFM image was processed using Gwyddion (http://gwyddion.net/). In assessing the quality of placement, we measured both the binding site occupancy (the percentage of sites with one or more origami), and the number of origami at a given site (0, 1, 2, or >3). In previous work14, we measured the quality of placement over dozens of repetitions of placement, where each repetition had more than 500 binding sites. Here, because of the difficulty of AFM measurements on PCC resonators, we performed a test for the quality of placement by analysing a single array of 600 binding sites on SiN (Fig. 1d), without any PCCs. Carboxylate-mediated placement on SiN resulted in no empty binding sites, and few sites with multiple bindings (∼2% of sites). Thus, we measured a single origami occupancy of 98%, consistent with that previously achieved.
For experiments in which single origami binding sites were written inside PCCs (Figs 2 and 3), AFM imaging of selected cavities showed that the fabrication steps added to create the PCCs did not degrade the quality of placement: all PCCs inspected (Fig. 2a, Fig. 3a, and Extended Data Fig. 4) had binding sites occupied by single origami, within ∼10 nm of the desired location. We attribute the low rate of multiple bindings to the low Mg2+ concentration (enabled by carboxylated binding sites), the Tween 20 washes, and the proximity correction (which enables accurate patterning of the binding sites) that we used. We attribute the low positional error to our use of multiple alignment markers within the single e-beam field, and the use of the same markers for the patterning of both the PCCs and the binding sites. As far as we can tell, the quality of the binding sites is not affected by their proximity to etched features, such as the holes in the PCCs.
For experiments in which multiple origami binding sites were written inside resonators (Fig. 4, Extended Data Figs 7 and 9), we observed that resonators with larger numbers of binding sites (that is, 5, 6 or 7) had a greater number of multiple bindings (which increase the number of origami in the resonator above that desired). We did not quantify this effect, because linear fits of emission as a function of the number of binding sites suggest that it is not a large effect (Fig. 4b). In our previous work14, we observed that crowding of binding sites over large arrays decreased site occupancy, because sites filled mostly via 2D diffusion of weakly surface-bound origami from the edges of the array. Here, crowding of binding sites within PCCs seems to have an opposite effect. The context is quite different, however. The PCC structure blocks 2D diffusion of origami to binding sites from the background, and sites are positioned closely in a different configuration which may allow extra origami to bind partially to two adjacent binding sites. The effect of the spacing between origami binding sites in different contexts thus deserves more study.
Spectroscopy
For all cross-polarization reflectance and microphotoluminescence (fluorescence) spectroscopy we used the set-up illustrated in Extended Data Fig. 5. For cross-polarization reflectance measurements, we used a Fianium supercontinuum laser as an excitation source. Broad-spectrum excitation light was polarized used a linear polarizer (polarizer 1) and focused onto the PCC sample through a 50× (0.8 NA) infinity-corrected objective. The sample was oriented at 45° to the incident polarization, and reflected signal collected by the objective passed through another polarizer (polarizer 2, orthogonal to polarizer 1) before being coupled into a monochromator (a SpectraPro-2500i with a grating of 1,800 grooves per millimetre from Princeton Instruments).
For fluorescence spectroscopy we use the same basic set-up but replaced the supercontinuum laser with a 15 mW, 638 nm laser, bandpass filtered at 635 ± 5 nm (635 BP). After being expanded to ∼500 μm in size, the laser beam was focused onto the sample through the same 50× (0.8 NA) objective. On the sample surface, the laser spot was approximately 5 μm in size, as measured by a CCD camera. Fluorescence emission was filtered through a 650-nm longpass filter (650 LP) to remove excitation light, before being coupled into the monochromator. A marked reduction in fluorescence signal was observed after illumination times of 30–45 s. Thus the potential effects of bleaching were minimized by limiting total illumination time to 10 s: final spectra were produced by averaging 10 spectral scans, each taken with an integration time of 1 s.
Wide-field epifluorescence microscopy
All fluorescence imaging was performed with an Olympus BX-61 microscope with a xenon excitation source. Excitation light was filtered with a 640-nm shortpass filter, and passed through a 645-nm dichroic to the sample. After returning through the dichroic, emission light was bandpass filtered at 655 ± 5 nm. The narrow bandpass filter was important for emission; longpass filtering of the emission light admitted too much direct Cy5 emission, rather than indirect emission through the cavity, and this made it impossible to image the 2D mode map. Fluorescence emission was imaged using a 50× (0.8 NA) objective onto a Hamamatsu EMCCD cooled to −80 °C. Complete bleaching of samples was observed at illumination times of approximately 2 min. For a particular PCC array, each final image was created by averaging 10 image acquisitions. For n = 3 Cy5, integration time for each of these acquisitions was 1 s; for n = 15 Cy5, integration time for each acquisition was 200 ms. In the case of 2D modemaps, a single final image of a particular PCC array (Extended Data Fig. 6a) was somewhat noisy, and five final images of different PCC arrays were averaged to yield the final data (Fig. 3c and Extended Data Fig. 6b). Features of the experimental images and 2D LDOS are qualitatively similar, but we did not attempt to fit experimental images to simulation, and instead matched their features by eye.
FDTD simulations for PCC design and analysis
Three dimensional (3D) finite-difference time domain (FDTD) simulation was used both for PCC design and to generate simulated LDOS for comparison with experimental maps of the resonant cavity modes. All simulations were performed using FDTD Solutions from Lumerical Solutions, Inc (https://www.lumerical.com/). Lumerical simulation files can be found by unzipping the Supplementary Data, in the directory ‘LumericalScripts’. Matlab files for creating Autocad versions of optimized resonators can also be found by unzipping the Supplementary Data, in the directory ‘AutocadScriptGenerator’.
Cavity geometry (Fig. 1e) was based on previous designs24 featuring soft confinement35. To design the photonic crystal we fixed the refractive index of SiN at 2.05, the thickness of the SiN membrane at 275 nm, and adjusted r, r/a, r1, r2 and s to maximize quality factor within the wavelength range 655–660 nm. Photonic crystal size was set to 20a in the x-direction and 34.64a in the y-direction. Boundary conditions were implemented by introducing a perfect matching layer around the structure. The simulation discretization was set to a/R in the x-direction, 0.866a/R in the y-direction, and a/R in the z-direction, where the variable R was set to 10 for PCC design (so that PCC parameters could be quickly optimized), and set to 20 to generated simulated LDOS of higher resolution for comparison with experimental mode maps. The simulation modelled emission from a single dipole with polarization P(x, y, z) = (1, 1, 0), located at a weak symmetry point close the cavity surface.
Code availability
The code used to design and simulate the PCCs as well as code to generate Autocad files for electron beam lithography defining PCCs and binding sites is available as the zip-encoded Supplementary Data file.
Online Methods
DNA origami
Design. A variation of the ‘sharp triangle’ design described previously15 was chosen because the sharp triangle is rigid and it has a low tendency to aggregate. The ‘sameside sharp triangle’ used here (details of design in Extended Data Fig. 1) has the same scaffold path as the original sharp triangle, and the scaffold strand shares the same alignment as in the original structure. (For sequences and the caDNAno design, unzip Supplementary Data: specifically, files ‘ETSamSide_15Cy5_StapleList.xls’ and ‘Sameside-sharp-triangle-bridged.json’ which unzip into the directory ‘OrigamiDesignFiles’.) The difference between the original sharp triangle and the sameside sharp triangle lies in the pattern of nicks along the phosphate backbone of the staple strands: in the original sharp triangle, nick positions alternate between the two faces of the triangle, whereas the new sameside sharp triangle has all nick positions on the same face of the triangle. Additionally, 15 staples on the inner edge of the triangle occur in two versions. The first version is an unmodified staple. The second version has an 18-nucleotide poly-T extension, which serves as a linker that binds to a 21-nucleotide poly-A ‘subcomponent’ strand bearing a single Cy5 on its 5′ end. Using this basic design we prepared two version of the origami: one had n = 15 Cy5 molecules along its inner edge and the second had only n = 3 Cy5 in along its inner edge.
Synthesis. Staple strands (Integrated DNA Technologies, 100 μM each in water) and the scaffold strand (single-stranded M13mp18, Bayou Biolabs, P-107) were mixed together to target concentrations of 100 nM (each staple) and 40 nM, respectively (a 2.5:1 staple:scaffold ratio) in 10 mM Tris Base, 1 mM EDTA buffer (adjusted to pH 8.35 with HCl) with 12.5 mM magnesium chloride. (We refer to the buffer used for origami synthesis, with the above combination of Tris Base, EDTA, and Mg2+, as ‘TE/Mg2+’.) Staples with the poly-T linker and the Cy5 label were added at 5× and 150× excess respectively. 50 μl volumes of staple/scaffold mixture were heated to 90 °C for 5 min and annealed from 90 °C to 20 °C at −0.2 °C min−1 in a PCR machine. We used 0.5 ml DNA LoBind tubes (Eppendorf) to minimize loss of origami to the sides of the tubes.
Purification. Since a high concentration of excess staples interferes with origami placement, the synthesized origami were purified away from excess staples using 100 kD molecular weight cut-off spin filters (‘Amicon Ultra-0.5 Centrifugal Filter Units with Ultracel-100 membranes’). By the following protocol, recovery is generally 40%–50% and staples are not visible by agarose gel. Wet the filter by adding 500 μl TE/Mg2+. Spin filter at 2,000 r.c.f. (relative centrifugal force) for 6 min at 4 °C, until the volume in the filter is 50 μl. Discard the filtrate. Add 50 μl of unpurified origami and 400 μl TE/Mg2+. Spin at 2,000 r.c.f. for 6 min at 4 °C. Wash the origami three more times by discarding the filtrate, adding 450 μl TE/Mg2+ and spinning at 2,000 r.c.f. for 6 min at 4 °C. Invert the filter onto a clean tube and spin at 2,000 r.c.f. for 6 min at 4 °C to collect purified origami (∼50 μl). Total time for this purification is roughly 40 min.
All of the work reported in this Letter was performed with spin-column purified origami. Other protocols, which may give higher yield via PEG precipitation or magnetic bead separation, are available31,32 although we have not verified that these methods would give quantitatively similar results for placement.
Quantification. Post-purification, origami were quantified using a NanoDrop spectrophotometer (Thermo Scientific). We estimated the molar extinction coefficient of the origami triangles as that of a fully double-stranded M13mp18 molecule (ε = 123,735,380 M−1 cm−1; we did not correct for a small single-stranded loop present on one edge of the origami). The typical working concentration for origami during placement was 100 pM, which is too small to be measured with the NanoDrop, so serial dilutions were performed. For all of the experiments performed reported here, we used a single 300 μl stock of 20 nM origami solution that was diluted to 100 pM immediately before each use.
Handling origami. After purification and quantification, it is especially important to use DNA LoBind tubes (Eppendorf) for storage and dilution of low concentration DNA origami solutions. Low dilutions, for example, 100 pM, must be made fresh from more concentrated solutions and used immediately—even overnight storage can result in total loss of origami to the sides of the tube. Addition of large amounts of carrier DNA to prevent origami loss may prevent origami placement, just as excess staples do. We have not yet determined whether other blocking agents such as BSA might both prevent origami loss and preserve placement.
Photonic crystal cavity nanofabrication
We fabricated PCCs similar to previous designs24 using two slightly different approaches. For isolated PCCs occurring in widely spaced arrays, like those which appear in Fig. 2, we suspended the PCC membranes using a front etch. For large, close-packed arrays of PCCs like those which appear in Figs 3 and 4, we suspended the PCC membranes using a back etch. Schematics for both fabrication processes are given in Extended Data Fig. 2. All fabrication was carried out in Caltech’s Kavli Nanoscience Institute.
Widely spaced PCC arrays
A schematic of the fabrication process is shown in Extended Data Fig. 2a; SEM images of these arrays are given in Extended Data Fig. 3. Fabrication began with 275 nm LPCVD (low-pressure chemical vapour deposition) growth of SiN on a 100 mm single-side polished (SSP, 525 ± 25 μm thick) 〈100〉 silicon wafer as the base substrate (Rogue Valley Microdevices). The wafer was cleaned and alignment markers were defined in the SiN layer by e-beam lithography and modified Bosch ICP (inductively coupled plasma) etching. The substrate was cleaned again and silanized with a trimethylsilyl passivation layer by vapour deposition of HMDS (hexamethyldisilazane). Next, binding sites in the shape of a DNA origami were defined at specific locations using the alignment markers (defined a priori) by e-beam lithography. Binding sites were then activated with a short O2 plasma etch and the resist was stripped. Surface silanols on binding sites were converted to carboxylate groups via a second silanization. Lastly, a PCC was defined around the carboxylated binding site by e-beam lithography and modified Bosch ICP etching of the SiN layer. Finally, PCCs were suspended using a XeF2 isotropic etch of the underlying Si layer.
Here we note that all modified Bosch ICP etching of SiN, whether for definition of PCCs or alignment marks, was performed in an ICP-RIE (inductively coupled plasma-reactive ion etcher, Oxford Instruments System 100 ICP 380) at a pressure of 4 mtorr, with a flow rate of 25 sccm for SF6 and 35 sccm for C4F8, an RF generator power of 25 W, an ICP power of 1,000 W, at 4 °C. Further, we note that this modified Bosch process does not involve alternating applications of SF6 and C4F8, but rather introduces them as a mixed gas.
Definition of alignment markers. Substrates were sonicated with isopropanol (IPA) and dried in a stream of N2 to remove any particulate contaminants. Substrates were spin coated with 600 nm of ZEP 520A (10 s, 500 rpm; 10 s, 1,500 rpm; 120 s, 3,000 rpm; ZEP 520A is a high-resolution positive electron beam resist from Zeon Chemicals). Alignment marker patterns were defined by e-beam. Patterns were developed in ZED N50 (Zeon Chemicals electron beam resist developer) for 1 min. Patterns were transferred into the substrate using the modified Bosch process for about 30 min. Remaining resist was stripped with hot (50 °C) n-methyl pyrrolidone (NMP).
Surface passivation with trimethylsilyl groups. Substrates were sonicated with IPA and dried in a stream of N2, to remove any particulate contaminants. Substrates were cleaned with O2 plasma in a Plasmatherm Dual-chamber SLR-720 RIE (reactive ion etcher), with a flow rate of 50 sccm, a pressure of 50 mtorr and a power setting of 80 W (110 W cm−2), for 5 min to create surface silanols. Substrates were dehydrated on a hot plate at 150 °C for 5 min. Substrates were incubated in a 4 l chamber saturated with HMDS vapour (a 20 cm × 20 cm × 10 cm Tupperware with a 90 mm Petri dish in the corner holding 10 ml HMDS) for 20 min. This step resulted in the formation of a monolayer of trimethylsilyl groups on the surface. Substrates were heated on a hot plate at 150 °C for 30 min, to stabilize the silanized surface. It is important to bake the substrate both before and after HMDS vapour-priming to ensure high quality surface silanization and increase the robustness of the monolayer to hydrolysis. Failure to perform either of these steps leads to increased background binding of origami.
Definition of carboxylated binding sites. Binding sites were defined by e-beam lithography on a freshly prepared and silanized substrates as follows. Substrates were spin-coated with a 170–180 nm resist layer of poly(methyl methacrylate) (MicroChem Corp., 950 PMMA, A3) at 2,500 rpm for 90 s. The resist was baked at 180 °C for 30 s. Binding sites were defined in the resist using e-beam lithography with a 100 keV beam at 500 pA current. The dosage ranged from 600 to 850 μC cm−2 and patterns were proximity corrected. The resist was developed for 70 s in a 1:3 solution of methyl isobutyl ketone (MIBK) and isopropanol (IPA). Surface silanols were created in the lithographically defined regions by an O2 plasma etch (Plasmatherm Dual chamber SLR-720 RIE, a flow rate of 20 sccm, a pressure of 20 mtorr and a power setting of 50 W (92 W cm−2), for 18 s). The resist was stripped by sonicating the wafer in bath of NMP at 50 °C for 10 min. The surface silanols that were created by O2 plasma was converted to carboxylate groups by incubating the substrate in a disodium carboxyethylsilanetriol (CTES from Gelest, 25% w/v in water Catalog #SIC2263.0) silanization buffer (0.01% CTES, 10 mM Tris, pH 8.3 for 10 min) followed by sonication in milliQ water. During the last step, ensure that the silanization buffer is at pH 8.3. Higher pH leads to the trimethylsilyl background being hydrolysed and inadvertently carboxylated.
Definition of isolated PCCs. After binding site definition, the substrate was cleaned with IPA. A 600 nm thick layer of ZEP 520A was spin-coated on the substrate and baked at 180 °C for 1 min. PCCs were defined using e-beam lithography with a 100 keV beam at a current of 500 pA. The dosage ranged from 200 to 300 μC cm−2 and patterns were proximity corrected. The exposed resist was developed in ZED N50 for 1 min. PCC structures were created via modified Bosch etch for 9–12 min, depending on the etch rate observed that day. The Si under the PCC was isotropically etched with a custom-built pulsed XeF2 etcher (5 cycles of 20 s each, 500 mtorr XeF2). The resist was stripped by overnight soaking of the chip in a bath of NMP at 50 °C. It is important not to sonicate the substrate after the PCC has been suspended as it leads to the collapse of the PCC membrane.
Close-packed PCC arrays
A schematic of the fabrication process is given in Extended Data Fig. 2b. The process is a modification of procedures in ref. 33 in which extremely large SiN membranes were created (up to 2.4 mm × 2.4 mm). The fabrication process is similar to that described for isolated cavities above, with the exception that PCC arrays were created on a SiN window supported by a thin Si layer (which is removed at the very last step with an XeF2 etch). Fabrication began with double-side polished silicon wafers (DSP, 〈100〉, 380 ± 10 nm thick, University Wafers, Rogue Valley Microdevices) with 275 nm layers of LPCVD-grown SiN on both sides of each wafer.
Etching of large SiN windows. Wafers were cleaned and alignment marker were defined by e-beam lithography and ICP/modified Bosch etch (30 min) in the SiN layers on both sides, using the wafer flat for alignment. Substrates were spin-coated with a 600 nm layer of ZEP 520A on the back side of the wafer. Substrates were spin-coated with ProTEK PSB (Brewer Science) on the top side of the wafer to protect it from contamination and physical scratches. A 650 μm × 650 μm window was defined via e-beam using back-side alignment markers. The resist was developed in ZED N50 for 1 min. The SiN layer (on the back side) was etched for 1 h by the modified Bosch process, to punch through to the Si layer. Si exposed in the previous step was etched for 12 h in 50% KOH solution to a depth of approximately 300–350 μm. This created an SiN window of approximately 400 μm × 400 μm supported by an Si layer less than 80 μm thick. The ProTEK PSB was stripped using hot (50 °C) NMP.
Notes on fabrication. After Si-supported SiN windows were etched, origami binding sites and PCC arrays were defined on the top surfaces using earlier-defined top-side alignment. We note that the alignment error between patterns on the top and bottom of the wafers sometimes reached a few tens of micrometres. However the most crucial alignment errors, between the origami binding site and the PCC array, were limited to tens of nanometres, because both binding sites and PCCs were created using the same set of alignment markers on the top of the wafer. Finally, the thin Si support remaining underneath the PCC array was removed using XeF2 pulsed etching. The suspended membranes are extremely delicate so it is crucial that substrate is handled with care. It is important not to sonicate suspended photonic crystal membranes as this will break them.
Origami placement on PCCs
Here the placement DNA origami on binding sites is mediated by Mg2+ binding to surface carboxylate groups, rather than surface silanols, as in our previous work14. We have observed that the use of carboxylated binding sites allows high-quality origami placement and orientation on SiN at a much lower Mg2+ concentration (15 mM) than that required (35 mM) for binding sites activated only by an O2 plasma (and which we presume are covered with silanols). We have not measured the density of carboxylate groups or silanols in either case, but we suggest that the effect is due to the difference in pKas between these two functional groups: similar surface carboxylate groups34 have a pKa of ∼6, while silanol groups have a pKa of 8.3. Thus binding sites with carboxylate groups should carry a higher negative charge at our working pH of 8.3, they should bind more Mg2+, and they should enable the observed binding of origami at lower Mg2+ concentration.
In addition to decreasing the potential for Mg2+ salt artefacts during drying, the use of carboxylate groups has a further important added benefit. Over the course of the extensive PCC fabrication process, different areas experience many different specific series of treatments, which results in many different surface types. Some of these, for example the inside of the PCC holes or the back side of the PCC membranes, are not passivated with trimethylsilyl groups, and appear to bind some DNA origami at higher Mg2+ concentration. Thus the use of carboxylated binding sites (and hence a lower Mg2+ concentration for placement) decreases nonspecific DNA origami binding and ensures that under our buffer conditions the only locations at which origami can stably bind are the intended binding sites.
Below, we describe the placement protocol in five steps, which apart from the use of a lower Mg2+ concentration and carboxylated binding sites, is similar to that in our previous work14. Photographs in supplementary figure 3 of ref. 14 show how substrates should look at different steps of the placement process. (Note that our previous work uses carboxylation of binding sites at a different step, after origami placement, as part of a method to achieve covalent coupling of origami to substrates.)
Binding. A 50 mm Petri dish was prepared with a moistened piece of laboratory tissue paper to limit evaporation. Solution with 100 pM origami was prepared in ‘placement buffer’ (10 mM Tris, 15 mM Mg2+, pH 8.3) and a 20 μl drop was deposited in the middle of the chip, on top of the PCCs. The chip was placed in the closed, humid Petri dish and the origami solution was allowed to incubate on the chip for 1 h.
Initial wash. After the 1 h incubation, excess origami (in solution) were washed away with at least 8 buffer washes by pipetting 60 μl of fresh placement buffer onto the chip, and pipetting 60 μl off the chip. Each of the 8 washes consisted of pipetting the 60 μl volume up and down 2–3 times to mix the fresh buffer with existing buffer on the chip. This initial wash took about 2 min.
Tween wash. Next, in order to remove origami that were non-specifically bound to the passivated background, the chip was buffer-washed 5 times using a ‘Tween washing buffer’ with 0.1% Tween 20 (v/v in placement buffer). This took about 1 min. Because of the low surface tension of the Tween washing buffer, these washes were difficult to perform: they involve adding 20–40 μl of wash buffer, just enough to cover most of the chip, but not enough to spill over the chip and wet the back side of the chip (this may introduce dust contamination from the Petri dish). After the fifth wash, the chip was left to incubate for 30 min. (It is important to use Tween 20, rather than other surfactants. Tween 80 and SDS, which are two other common surfactants, lead to markedly different results—Tween 80 leads to the total removal of placed origami from the substrate, and SDS does not remove excess origami from the trimethylsilyl background. It is important that chips are not exposed to Tween 20 until after the origami have been deposited. Tween 20 applied before binding completely inactivates the binding sites.)
Final stabilizing wash. Last, the chip was buffer-washed 8 times back into a higher pH ‘stabilizing buffer’ (10 mM Tris, 15 mM Mg2+, pH 8.9) so that origami bound strongly and artefacts were minimized during subsequent drying. This took about 2 min. These washes were relatively high volume (60 μl) and were intended to completely remove the Tween 20. The amount of Tween 20 left was monitored qualitatively by the surface tension of the drop (roughly, by eye). When a 20 μl drop covered roughly the same area as the initially deposited drop, it was assumed that the Tween 20 had been sufficiently removed. In the last wash the chip was left with roughly 20 μl of stabilizing buffer, and was ready for drying.
Drying. Chips were dried by serial dilution into ethanol. The chips were dipped in 50% ethanol in water (v/v%) for 10 s, 75% ethanol in water (v/v%) for 10 s and then 120 s in 90% ethanol in water. Then the chips were air-dried. (If nanoarrays are subjected to ethanol solutions with less than 80% ethanol for more than 60–90 s, origami begin to detach from the surface. After the 90% ethanol immersion, it is necessary to let the samples air-dry rather than using N2 or compressed air, since streaking or other drying artefacts are observed in instances of forced air drying.)
Troubleshooting origami placement. If low binding, high multiple binding, or high background binding artefacts are observed, see the troubleshooting guide in Extended Data Table 1. Below we provide notes emphasizing some of the more important aspects of the placement protocol, which may be helpful in troubleshooting. Do not allow the patterned region with binding sites to dry at any point during the binding step or subsequent buffer washes. Inadvertent dewetting of the binding sites leads to distortion of the origami (causing them to ball up) as well as the formation of salt crystals on top of them. Do not use EDTA in placement, Tween washing, or imaging buffers. It is unnecessary in this context and will change the effective Mg2+ concentration for placement slightly. Make fresh buffer solutions every week. Here and elsewhere in this work, we use buffers at low strength (typically 10 mM) to minimize background binding and to make complete washing into different buffers easier. This means the buffers have low buffering capacity and the pH will decrease (and placement may cease to work, depending on conditions). For example, weak buffers made to read pH 8.35 can lose 0.05 pH units and read pH 8.3 after one week.
AFM characterization
After the cavities were fabricated and the origami were placed within the cavities, we quantified the quality of placement using dry AFM. While the presence of origami (and the fluorophores they carry) can be inferred from the optical experiments, we have used AFM because it provides the ability to unambiguously verify the presence of single origami, as well as measure the precise location of the origami (and hence fluorophores) with respect to the cavity. Unless otherwise specified (images in Fig. 1b were taken under aqueous buffer with a Bruker Fastscan AFM), all AFM images were taken in air in tapping mode with a Dimension Icon AFM/Nanoscope V Scanner (Bruker) using the short, fat cantilever from an SNL probe B (“sharp nitride lever”, 2 nm tip radius, Bruker) resonating at approximately 60 kHz. Imaging origami on suspended resonators was challenging. All images had to be acquired at the lowest possible force in order to reduce tip–sample interaction. We optimized imaging conditions on an unsuspended region near the alignment markers and then moved onto the cavity with precise stage movements. Minimization of imaging force was important: if imaging force was increased, imaging became unstable. We noticed that imaging near the edge of suspended regions was easier than at the middle; we suggest that the suspended SiN membrane is interfering with AFM measurement by acting as a mechanical resonator itself.
Each AFM image was processed using Gwyddion (http://gwyddion.net/). In assessing the quality of placement, we measured both the binding site occupancy (the percentage of sites with one or more origami), and the number of origami at a given site (0, 1, 2, or >3). In previous work14, we measured the quality of placement over dozens of repetitions of placement, where each repetition had more than 500 binding sites. Here, because of the difficulty of AFM measurements on PCC resonators, we performed a test for the quality of placement by analysing a single array of 600 binding sites on SiN (Fig. 1d), without any PCCs. Carboxylate-mediated placement on SiN resulted in no empty binding sites, and few sites with multiple bindings (∼2% of sites). Thus, we measured a single origami occupancy of 98%, consistent with that previously achieved.
For experiments in which single origami binding sites were written inside PCCs (Figs 2 and 3), AFM imaging of selected cavities showed that the fabrication steps added to create the PCCs did not degrade the quality of placement: all PCCs inspected (Fig. 2a, Fig. 3a, and Extended Data Fig. 4) had binding sites occupied by single origami, within ∼10 nm of the desired location. We attribute the low rate of multiple bindings to the low Mg2+ concentration (enabled by carboxylated binding sites), the Tween 20 washes, and the proximity correction (which enables accurate patterning of the binding sites) that we used. We attribute the low positional error to our use of multiple alignment markers within the single e-beam field, and the use of the same markers for the patterning of both the PCCs and the binding sites. As far as we can tell, the quality of the binding sites is not affected by their proximity to etched features, such as the holes in the PCCs.
For experiments in which multiple origami binding sites were written inside resonators (Fig. 4, Extended Data Figs 7 and 9), we observed that resonators with larger numbers of binding sites (that is, 5, 6 or 7) had a greater number of multiple bindings (which increase the number of origami in the resonator above that desired). We did not quantify this effect, because linear fits of emission as a function of the number of binding sites suggest that it is not a large effect (Fig. 4b). In our previous work14, we observed that crowding of binding sites over large arrays decreased site occupancy, because sites filled mostly via 2D diffusion of weakly surface-bound origami from the edges of the array. Here, crowding of binding sites within PCCs seems to have an opposite effect. The context is quite different, however. The PCC structure blocks 2D diffusion of origami to binding sites from the background, and sites are positioned closely in a different configuration which may allow extra origami to bind partially to two adjacent binding sites. The effect of the spacing between origami binding sites in different contexts thus deserves more study.
Spectroscopy
For all cross-polarization reflectance and microphotoluminescence (fluorescence) spectroscopy we used the set-up illustrated in Extended Data Fig. 5. For cross-polarization reflectance measurements, we used a Fianium supercontinuum laser as an excitation source. Broad-spectrum excitation light was polarized used a linear polarizer (polarizer 1) and focused onto the PCC sample through a 50× (0.8 NA) infinity-corrected objective. The sample was oriented at 45° to the incident polarization, and reflected signal collected by the objective passed through another polarizer (polarizer 2, orthogonal to polarizer 1) before being coupled into a monochromator (a SpectraPro-2500i with a grating of 1,800 grooves per millimetre from Princeton Instruments).
For fluorescence spectroscopy we use the same basic set-up but replaced the supercontinuum laser with a 15 mW, 638 nm laser, bandpass filtered at 635 ± 5 nm (635 BP). After being expanded to ∼500 μm in size, the laser beam was focused onto the sample through the same 50× (0.8 NA) objective. On the sample surface, the laser spot was approximately 5 μm in size, as measured by a CCD camera. Fluorescence emission was filtered through a 650-nm longpass filter (650 LP) to remove excitation light, before being coupled into the monochromator. A marked reduction in fluorescence signal was observed after illumination times of 30–45 s. Thus the potential effects of bleaching were minimized by limiting total illumination time to 10 s: final spectra were produced by averaging 10 spectral scans, each taken with an integration time of 1 s.
Wide-field epifluorescence microscopy
All fluorescence imaging was performed with an Olympus BX-61 microscope with a xenon excitation source. Excitation light was filtered with a 640-nm shortpass filter, and passed through a 645-nm dichroic to the sample. After returning through the dichroic, emission light was bandpass filtered at 655 ± 5 nm. The narrow bandpass filter was important for emission; longpass filtering of the emission light admitted too much direct Cy5 emission, rather than indirect emission through the cavity, and this made it impossible to image the 2D mode map. Fluorescence emission was imaged using a 50× (0.8 NA) objective onto a Hamamatsu EMCCD cooled to −80 °C. Complete bleaching of samples was observed at illumination times of approximately 2 min. For a particular PCC array, each final image was created by averaging 10 image acquisitions. For n = 3 Cy5, integration time for each of these acquisitions was 1 s; for n = 15 Cy5, integration time for each acquisition was 200 ms. In the case of 2D modemaps, a single final image of a particular PCC array (Extended Data Fig. 6a) was somewhat noisy, and five final images of different PCC arrays were averaged to yield the final data (Fig. 3c and Extended Data Fig. 6b). Features of the experimental images and 2D LDOS are qualitatively similar, but we did not attempt to fit experimental images to simulation, and instead matched their features by eye.
FDTD simulations for PCC design and analysis
Three dimensional (3D) finite-difference time domain (FDTD) simulation was used both for PCC design and to generate simulated LDOS for comparison with experimental maps of the resonant cavity modes. All simulations were performed using FDTD Solutions from Lumerical Solutions, Inc (https://www.lumerical.com/). Lumerical simulation files can be found by unzipping the Supplementary Data, in the directory ‘LumericalScripts’. Matlab files for creating Autocad versions of optimized resonators can also be found by unzipping the Supplementary Data, in the directory ‘AutocadScriptGenerator’.
Cavity geometry (Fig. 1e) was based on previous designs24 featuring soft confinement35. To design the photonic crystal we fixed the refractive index of SiN at 2.05, the thickness of the SiN membrane at 275 nm, and adjusted r, r/a, r1, r2 and s to maximize quality factor within the wavelength range 655–660 nm. Photonic crystal size was set to 20a in the x-direction and 34.64a in the y-direction. Boundary conditions were implemented by introducing a perfect matching layer around the structure. The simulation discretization was set to a/R in the x-direction, 0.866a/R in the y-direction, and a/R in the z-direction, where the variable R was set to 10 for PCC design (so that PCC parameters could be quickly optimized), and set to 20 to generated simulated LDOS of higher resolution for comparison with experimental mode maps. The simulation modelled emission from a single dipole with polarization P(x, y, z) = (1, 1, 0), located at a weak symmetry point close the cavity surface.
Code availability
The code used to design and simulate the PCCs as well as code to generate Autocad files for electron beam lithography defining PCCs and binding sites is available as the zip-encoded Supplementary Data file.
References
Benson, O. Assembly of hybrid photonic architectures from nanophotonic constituents. Nature 480, 193–199 (2011)
Armani, A. M. et al. Label-free, single-molecule detection with optical microcavities. Science 317, 783–787 (2007)
Yang, D. et al. High sensitivity and high Q-factor nanoslotted parallel quadrabeam photonic crystal cavity for real-time and label-free sensing. Appl. Phys. Lett. 105, 063118 (2014)
Jensen, K. et al. Cavity-enhanced room-temperature magnetometry using absorption by nitrogen-vacancy centers in diamond. Phys. Rev. Lett. 112, 160802 (2014)
Yakunin, S. et al. Low-threshold amplified spontaneous emission and lasing from colloidal nanocrystals of caesium lead halide perovskites. Nat. Commun. 6, 8056 (2015)
Strauf, S. et al. Self-tuned quantum dot gain in photonic crystal lasers. Phys. Rev. Lett. 96, 127404 (2006)
Hennessy, K. et al. Quantum nature of a strongly coupled single quantum dot–cavity system. Nature 445, 896–899 (2007)
Englund, D. et al. Controlling cavity reflectivity with a single quantum dot. Nature 450, 857–861 (2007)
Sapienza, L., Davanço, M., Badolato, A. & Srinivasan, K. Nanoscale optical positioning of single quantum dots for bright and pure single-photon emission. Nat. Commun. 6, 7833 (2015)
Barth, M., Nüsse, N., Löchel, B. & Benson, O. Controlled coupling of a single-diamond nanocrystal to a photonic crystal cavity. Opt. Lett. 34, 1108–1110 (2009)
Englund, D. et al. Deterministic coupling of a single nitrogen vacancy center to a photonic crystal cavity. Nano Lett. 10, 3922–3926 (2010)
Lyasota, A. et al. Integration of multiple site-controlled pyramidal quantum dot systems with photonic-crystal membrane cavities. J. Cryst. Growth 414, 192–195 (2015)
Kershner, R. J. et al. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nat. Nanotechnol. 4, 557–561 (2009)
Gopinath, A. & Rothemund, P. W. K. Optimized assembly and covalent coupling of single-molecule DNA origami nanoarrays. ACS Nano 8, 12030–12040 (2014)
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Acuna, G. P. et al. Fluorescence enhancement at docking sites of DNA-directed self-assembled nanoantennas. Science 338, 506–510 (2012)
Schreiber, R. et al. Hierarchical assembly of metal nanoparticles, quantum dots and organic dyes using DNA origami scaffolds. Nat. Nanotechnol. 9, 74–78 (2014)
Bui, H. et al. Programmable periodicity of quantum dot arrays with DNA origami nanotubes. Nano Lett. 10, 3367–3372 (2010)
Ko, S. H., Du, K. & Liddle, J. A. Quantum-dot fluorescence lifetime engineering with DNA origami constructs. Angew. Chem. Int. Ed. 52, 1193–1197 (2013)
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nat. Nanotechnol. 5, 61–66 (2010)
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003)
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009)
Zhang, T. et al. DNA-based self-assembly of fluorescent nanodiamonds. J. Am. Chem. Soc. 137, 9776–9779 (2015)
Barth, M. et al. Modification of visible spontaneous emission with silicon nitride photonic crystal nanocavities. Opt. Express 15, 17231–17240 (2007)
Louvion, N. et al. Local observation and spectroscopy of optical modes in an active photonic-crystal microcavity. Phys. Rev. Lett. 94, 113907 (2005)
Mivelle, M. et al. Light funneling from a photonic crystal laser cavity to a nano-antenna: overcoming the diffraction limit in optical energy transfer down to the nanoscale. Opt. Express 22, 15075–15087 (2014)
Rotenberg, N. & Kuipers, L. Mapping nanoscale light fields. Nat. Photon. 8, 919–926 (2014)
Sapienza, R. et al. Deep-subwavelength imaging of the modal dispersion of light. Nat. Mater. 11, 781–787 (2012)
Knowles, H. S., Kara, D. M. & Atatüre, M. Observing bulk diamond spin coherence in high-purity nanodiamonds. Nat. Mater. 13, 21–25 (2014)
Neu, E. et al. Low-temperature investigations of single silicon vacancy colour centres in diamond. New J. Phys. 15, 043005 (2013)
Stahl, E., Martin, T. G., Praetorius, F. & Dietz, H. Facile and scalable preparation of pure and dense DNA origami solutions. Angew. Chem. Int. Ed. 53, 12735–12740 (2014)
Shaw, A., Benson, E. & Högberg, B. Purification of functionalized DNA origami nanostructures. ACS Nano 9, 4968–4975 (2015)
Klein, M. J. K. Wafer-Scale Fabrication of Thin SiN Membranes and Au Films and Membranes with Arrays of Sub-micron Holes Using Nanosphere Lithography . PhD thesis, École Polytechnique Fédérale de Lausanne (2010)
Noy, A. Handbook of Molecular Force Spectroscopy (Springer, 2007)
Akahane, Y., Asano, T., Song, B.-S. & Noda, S. High-Q photonic nanocavity in a two-dimensional photonic crystal. Nature 425, 944–947 (2003)
Acknowledgements
We acknowledge funding from the Army Research Office (award W911NF-11-1-0117), the Office of Naval Research (award N000141410702), the Air Force Office of Scientific Research (Young Investigator award FA9550-15-1-0252), and the US National Science Foundation (Expeditions in Computing numbers 0832824 and 1317694, Molecular Programming Project; http://molecular-programming.org). We thank J. Fakonas for discussions and B. Fultz for use of his spectrometer. Device fabrication was done at Caltech’s Kavli Nanoscience Institute.
Author information
Authors and Affiliations
Contributions
A.G. and P.W.K.R. conceived the project. A.G. performed origami synthesis, nanofabrication, AFM, SEM and fluorescence microscopy. A.G. and E.M. built the set-up for microphotoluminescence spectroscopy. All authors contributed to data interpretation and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DNA origami design showing position of Cy5 fluorophores.
a, We used a triangular DNA origami shape, ~130 nm on each edge, as an adaptor to position fluorophores within a photonic crystal cavity (PCC). The origami is composed of a 7,249-nucleotide (nt) single-stranded scaffold (the commercially available single-stranded genome of the M13mp18 bacteriophage) and approximately 200 commercially synthesized ‘staple strands’ which are typically 32 nucleotides in length. Mixed together and annealed from 90 to 20 °C over the course of about 6 h, the strands self-assemble to form equilateral triangles in high yield. Three trapezoidal domains make up this ‘sameside sharp triangle’ design. Red filled circles indicate 15 positions at which staples were extended on their 5′ end with 18-nt poly-T linkers for the synthesis of origami bearing n = 15 Cy5. Blue rings indicate the subset of positions used for origami bearing n = 3 Cy5. To these linkers, 21-nt poly-A ‘subcomponent’ strands modified with a 5′ Cy5 were hybridized. Because the Cy5 modification is on their 5′ ends, subcomponent strands put Cy5 within a few nucleotides of the origami surface, less than 1 nm from where linkers extend from the origami surface (rather than at the 6-nm-away distal end of the linker/Cy5 strand hybrid). b, Schema showing both components of the fine structure of a DNA origami (gaps between helices, and crossover positions, where helices are tangent) with linker/Cy5-label positions noted as in a. The white outer ring has a 62.4-nm diameter and the white inner ring has a 31.2-nm diameter. As in a, red dots indicate the subset of positions used for origami bearing n = 15 Cy5 and blue rings around red dots indicate positions used for origami bearing n = 3 Cy5. Rainbow colours on the origami trace the path of the scaffold as it progresses through the origami structure, from red to purple. c, Buffer AFM image showing fine structure of origami without linkers/Cy5 strands for comparison with b. d, Buffer AFM image showing origami with n = 3 linkers and Cy5 strands. In this AFM image the Cy5-labelled duplexes appear to fall on the inside of the inner triangular hole, perhaps due to adhesion to the mica substrate used in these high-resolution experiments. High-resolution imaging of origami dried onto SiN is difficult, and so the exact conformation of Cy5-labelled duplexes on resonators was not determined. However, because the carboxylated sites bind DNA strongly, it seems likely that Cy5-labelled duplexes would behave similarly and fall within the 54-nm triangular hole of the origami, between 30 and 60 nm from the centre of the origami.
Extended Data Figure 2 Process flow for fabricating PCCs.
a, Fabrication of isolated PCCs for widely spaced arrays (Fig. 2). b, Fabrication of PCCs in close-packed arrays (Figs 3, 4). Note that back-side alignment markers are omitted from b for clarity. After either fabrication process substrates are incubated in origami solution, rinsed of excess origami, subjected to an ethanol dilution series, and air-dried.
Extended Data Figure 3 SEM imaging of isolated PCCs used for taking fluorescence spectra as a function of x-position.
An 8 × 21 array was used. a, Positions 01 to 21 each indicate a row of eight copies of isolated PCCs having origami positioned at the same x-offset, as exemplified in Extended Data Fig. 4. Red squares indicate broken PCC membranes which were not used. b, Intact membrane. c, Typical broken membrane. d, Zoom-in of PCC. e, Cross-section of PCC membrane. Scale bars: a, 300 μm; b, 2 μm; d, 1 μm; e, 500 nm.
Extended Data Figure 4 AFM of isolated PCCs and control cavity.
a, Schema and representative AFM for each of 21 offsets at which spectra (Fig. 2b) were measured. b, Dry AFM of a control PCC filled with origami (n = 15 Cy5/origami). Scale bar, 250 nm. c, PL reference spectra (red) for Cy5-origami on open SiN, and for Cy5-origami filling a cavity, as in b.
Extended Data Figure 5 Optical set-up for reflectance and fluorescence spectroscopy.
The 650-nm long-pass filter (650 LP) was used only for fluorescence spectroscopy. Details in Methods.
Extended Data Figure 6 Comparison of unaveraged and averaged epifluorescence images of PCC array used for 2D mode map.
a, A single epifluorescence image of a single array. b, An average of five images taken of five different PCC arrays, as in Fig. 3c. Scale bars, 23 μm.
Extended Data Figure 7 Raw fluorescence data demonstrating digital control of cavity emission.
The central panel diagrams eight different arrangements of origami (white triangles) within PCCs, with the number of origami ranging from zero to seven. The left panel shows an epifluorescence image of eight PCC arrays, each containing 512 copies of a PCC with the origami arrangement diagrammed in the adjacent row of the central panel; each origami has n = 3 Cy5. The right panel shows an analogous epifluorescence image of eight PCC arrays for which each origami has n = 15 Cy5. Histograms summarizing these epifluorescence data are given in Extended Data Fig. 8. See Fig. 4b for plots showing the linearity of intensity as the number of origami are increased. Scale bars, 50 μm.
Extended Data Figure 8 Histograms of numerical data demonstrating digital control of cavity emission.
Eight different arrangements of origami were loaded into PCCs, with the number of origami ranging from zero to seven (see centre panel, Extended Fig. 7). 512 identical PCCs of each of the nonzero arrangements were imaged by epifluorescence microscopy (Extended Data Fig. 7, left and right panels) and their intensity was quantified. These data, for n = 3 Cy5 origami (a) and n = 15 Cy5 origami (b) are represented by coloured histograms labelled ‘one’ through ‘seven’. The number of PCCs with a given fluorescence intensity is plotted on the y-axis. c, Summary of means and standard deviations (for intensity measured as counts) from Gaussian fits in a, b. The peaks of these histograms are plotted in Fig. 4b.
Extended Data Figure 9 Schema for recreation of Vincent van Gogh’s painting The Starry Night (1889).
See Fig. 4c for recreation. a, Original image from Wikimedia Commons (https://commons.wikimedia.org/wiki/File:VanGogh-starry_night_ballance1.jpg). Because our current emitters and cavities provide only one color, with only eight levels of intensity, the original image was converted to greyscale (b) and then further converted to have three-bit colour depth (c). Because of the difficulty of handling extremely large suspended SiN membranes (which easily break) we limited our recreation to a low-resolution 256 × 256 = 65,536 pixel version, which was then further broken down into sixteen 64 × 64 pixel arrays (d). With one PCC per pixel, these arrays were small enough that we could fabricate and handle the resulting SiN membranes without too much breakage, image them separately, and then reassemble the sixteen images into a 4 × 4 array for the final recreation (Fig. 4c).
Supplementary information
Supplementary Data
This file contains Supplementary Data including (1) a DNA origami caDNAno design file and corresponding Microsoft Excel sequence list, (2) PCC design files as Matlab and Autocad scripts, and (3) FDTD simulations as Lumerical scripts. These files have been combined as a single .zip file. (ZIP 287 kb)
Rights and permissions
About this article
Cite this article
Gopinath, A., Miyazono, E., Faraon, A. et al. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 535, 401–405 (2016). https://doi.org/10.1038/nature18287
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18287
- Springer Nature Limited
This article is cited by
-
Site-directed placement of three-dimensional DNA origami
Nature Nanotechnology (2023)
-
A DNA origami rotary ratchet motor
Nature (2022)
-
Deterministic assembly of single emitters in sub-5 nanometer optical cavity formed by gold nanorod dimers on three-dimensional DNA origami
Nano Research (2022)
-
Spontaneous emission in micro- or nanophotonic structures
PhotoniX (2021)
-
A synthetic tubular molecular transport system
Nature Communications (2021)