Abstract
In salamanders, grafting of a left limb blastema onto a right limb stump yields regeneration of three limbs, the normal limb and two ‘supernumerary’ limbs1,2,3,4. This experiment and other research have shown that the juxtaposition of anterior and posterior limb tissue plus innervation are necessary and sufficient to induce complete limb regeneration in salamanders5,6,7,8,9,10. However, the cellular and molecular basis of the requirement for anterior–posterior tissue interactions were unknown. Here we have clarified the molecular basis of the requirement for both anterior and posterior tissue during limb regeneration and supernumerary limb formation in axolotls (Ambystoma mexicanum). We show that the two tissues provide complementary cross-inductive signals that are required for limb outgrowth. A blastema composed solely of anterior tissue normally regresses rather than forming a limb, but activation of hedgehog (HH) signalling was sufficient to drive regeneration of an anterior blastema to completion owing to its ability to maintain fibroblast growth factor (FGF) expression, the key signalling activity responsible for blastema outgrowth. In blastemas composed solely of posterior tissue, HH signalling was not sufficient to drive regeneration; however, ectopic expression of FGF8 together with endogenous HH signalling was sufficient. In axolotls, FGF8 is expressed only in the anterior mesenchyme and maintenance of its expression depends on sonic hedgehog (SHH) signalling from posterior tissue. Together, our findings identify key anteriorly and posteriorly localized signals that promote limb regeneration and show that these single factors are sufficient to drive non-regenerating blastemas to complete regeneration with full elaboration of skeletal elements.
Similar content being viewed by others
Main
Limb regeneration in salamanders, including axolotls, proceeds in four steps: injury, blastema formation, sustained blastema outgrowth, and differentiation. Previous studies have shown that blastema formation is dependent on the presence of nerves at the injury site, and that sustained outgrowth requires the presence of both anterior and posterior limb cells5,6,7,8,9,10. Remarkably, despite the large cadre of molecular information that has accumulated on limb patterning in the past decades, the cellular and molecular logic underlying the requirement for anterior and posterior cell juxtaposition in maintaining limb regeneration is unknown. Two models relying on different cellular principles have been proposed to explain this requirement. In the polar coordinate model, a graded spectrum of positional values around the limb circumference manifests in the form of cell surface properties. Disruption of this gradation as a result of transplantation or amputation would be recognized by cells at the border, stimulating them to proliferate and generate daughter cells that intercalate the missing circumferential identities and thereby resulting in distal outgrowth of the limb tissue11. By contrast, the boundary model subdivides the limb circumference into four domains: anterior–dorsal, anterior–ventral, posterior–dorsal, and posterior–ventral, and proposes that the intersection between the domains is required to induce the posterior domains to express a secreted morphogen that is required for growth and patterning of the limb12. Thus, complementing, cross-inductive interactions between domains rather than local detection of differences form the conceptual basis of the boundary model.
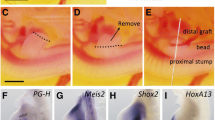
Anterior-only blastemas (ABs) composed solely of anterior tissue can be created by deviating nerve endings to an anterior-facing lateral wound in the upper limb8,9. A mid-bud blastema forms at the lateral site but eventually regresses, unless a patch of posterior limb skin (including dermal mesenchyme) is grafted to the anterior site; the graft allows the blastema to grow and form a fully patterned limb, called an accessory limb. We first searched for posterior-associated molecules that could stimulate complete limb regeneration from ABs and found that stimulation of HH signalling via the global application of smoothened agonist (SAG) was sufficient to prevent regression of an AB and to elicit regeneration of a limb with clear skeletal patterning along the proximo–distal axis and the full complement of constituent limb cell types (Fig. 1a, Extended Data Figs 1 and 2a, b, d and Supplementary Table 1). The accessory limbs displayed varying digit numbers that depended on SAG concentration (Extended Data Fig. 2a). We corroborated these findings by overexpressing human SHH in ABs via baculovirus-mediated delivery; this also led to patterned limb outgrowth (n = 8 of 32; Fig. 1b and Supplementary Table 1). These results show that activation of the HH pathway is sufficient to induce regeneration from an anterior-only blastema. Conversely, inhibition of HH signalling by global application of cyclopamine blocked accessory limb outgrowth from ABs complemented with a graft of posterior skin (Extended Data Fig. 2c), indicating that HH signalling is required for ectopic blastema outgrowth when a full complement of positional identities is present.
a, Accessory limbs develop from ABs treated with 10 nM SAG (n = 43 of 66), but not from ABs treated with water (n = 0 of 22). Alcian blue/alizarin red staining reveals the skeletal pattern with supernumerary digits and carpal elements in the accessory limb. Data from three experiments. See Supplementary Table 1 for details. b, Ectopic expression of SHH in ABs by baculovirus transduction is sufficient to drive accessory limb outgrowth (n = 8 of 32, data from two experiments). Black dashed lines demarcate the outgrowing accessory limb. See Supplementary Table 1 for details. c, Ectopic expression of FGF8 in ABs by baculovirus transduction is sufficient to drive accessory limb outgrowth (n = 14 of 57, data from four experiments). Alcian blue/alizarin red staining reveals the skeletal pattern with a single digit in the accessory limb. See Supplementary Table 4 for details. d, Expression of FGF–SHH loop components in ABs untreated or treated with 10 nM SAG and analysed with Nanostring nCounter technology. Each data point is the mean normalized count number from biological replicates (n = 3 or 4; for individual n values and complete data set, see Supplementary Table 7a; data from one experiment). Error bars represent s.d. For data points where the s.d. was low, no error bars are shown. The dashed line is for visual aid and indicates the level of mature (day 0) expression. For in situ hybridization of selected genes, see Extended Data Fig. 4. Scale bars, 2 mm. dpw, days post-wounding. For alcian blue/alizarin red staining, cartilage is blue and ossifications are red.
To gain an insight into the mechanism of SAG-induced outgrowth of ABs, we investigated whether HH signalling induced a stable change from an anterior to a posterior determination state in parts of the ectopic limb tissue, or whether SAG treatment stimulated anteriorly determined cells to grow out a limb. Reamputation of SAG-induced accessory limbs (SAG-ALs) yielded either no outgrowth (n = 2 of 5) or outgrowth of a spike (n = 3 of 5), suggesting that a stable anterior–posterior discontinuity that could induce a secondary limb had not been established (Extended Data Fig. 3a). Transplantation is the cleanest way to test determination state. Therefore we transplanted skin from SAG-ALs to innervated anterior or posterior wounds and looked for secondary accessory limb formation (Extended Data Fig. 3b). Limb outgrowth occurred when the SAG-AL skin was transplanted to posterior (n = 15 of 21) but not anterior wounds (n = 0 of 22), indicating that the SAG-AL limb tissue retains an anterior rather than posterior determination state (Extended Data Fig. 3c, d and Supplementary Table 2).
These results suggested that SAG may promote accessory limb outgrowth by upregulating a downstream growth-stimulating signalling loop that is involved in limb development; namely, posteriorly secreted SHH that stimulates upregulation of gremlin1 (grem1) and consequently fgf genes, whose protein products in turn maintain shh expression in posterior cells13,14,15,16,17,18,19,20,21,22,23,24,25. We therefore quantified the expression of key limb development factors and limb blastema genes in SAG-treated versus untreated ABs by Nanostring nCounter analysis (a method for counting RNA transcripts) and by RNA in situ hybridization. As expected, we observed upregulation of the blastema-associated transcription factors prrx1, msx1, msx2 and twist1 and of the anterior marker gli3 in ABs treated with SAG or water (Extended Data Fig. 4a, b and Supplementary Table 7a). Interestingly, a set of extracellular signalling factors that included fgf8, fgf9, fgf10 and grem1 were sustainably and highly upregulated in SAG-treated ABs but showed only an initial, unsustained upregulation in untreated ABs. Fgf17 was expressed only in ABs treated with SAG, and shh was not expressed in ABs at all (Fig. 1d, Extended Data Figs 4 and 8 and Supplementary Tables 3, 7a and 8).
These data pointed to FGF signalling as a potentially critical effector of SAG in ABs. To test this theory, SAG-treated ABs were exposed to the FGFR1/FGFR3 signalling inhibitor PD173074 starting at day 10 after wounding. Treatment of ABs with SAG alone induced accessory limb formation (n = 14 of 14), whereas addition of PD173074 blocked SAG-induced accessory limb outgrowth (n = 0 of 16) (Extended Data Fig. 5a). To test whether FGF signalling can drive accessory limb outgrowth, FGF8 was ectopically expressed in ABs by baculovirus transduction; this yielded single digit outgrowths (Fig. 1c, Extended Data Fig. 5b and Supplementary Table 4) without radius or ulna formation or anterior–posterior digit elaboration. These data show that FGF signalling is both necessary and sufficient to drive accessory limb outgrowth from ABs.
We next tested whether SAG treatment was sufficient to induce outgrowth of posterior-only blastemas (PBs). PBs also regress unless complemented with an anterior skin graft. Interestingly, we found that SAG did not induce PBs to form accessory limbs (n = 1 of 48) (Fig. 2a, Extended Data Fig. 6a–c, f and Supplementary Table 1). Nanostring analysis (Fig. 2d and Supplementary Tables 7b, 9) and RNA in situ hybridization (Extended Data Fig. 7 and Supplementary Table 3) for gli1, ptc1 and shh revealed that these factors were expressed in PBs with or without SAG, indicating that the HH signalling system is intact in PBs but is not sufficient to induce regeneration in this context. In addition, grem1 and fgf10 were also initially upregulated in PBs with or without SAG, and fgf9 was initially upregulated in untreated PBs. On the other hand, fgf8 and fgf17 were not upregulated in PBs with or without SAG (Fig. 2d, Extended Data Figs 7 and 8 and Supplementary Tables 3, 7b, 9). RNA in situ hybridization on normal blastemas revealed that the fgf8 signal localized to the anterior half of the blastema, being excluded from the posterior hand2-expressing zone. Interestingly grem1, fgf9, fgf10 and fgf17 were not excluded from the posterior hand2-expressing domain (Extended Data Fig. 9a, b). These results suggested that FGF8 is the anteriorly expressed FGF that is limiting in PB outgrowth. To test whether FGF8 expression could substitute for anterior cells in PBs to drive accessory limb outgrowth, we transduced the posterior regions of axolotl limbs with baculovirus expressing fgf8 and created PBs several days later. FGF8 expression alone was sufficient to induce accessory limb outgrowth in 54% of cases (n = 27 of 50), whereas no significant outgrowth was observed after control mCherry expression (Fig. 2b, Extended Data Fig. 6d, e, g and Supplementary Table 5). Interestingly, FGF8-mediated outgrowth from PBs required endogenous HH signalling, as co-treatment with cyclopamine blocked accessory limb outgrowth (Fig. 2c, Extended Data Fig. 10 and Supplementary Table 6). Our results show that FGF8 expression together with endogenous HH signalling is sufficient to drive accessory limb outgrowth from PBs.
a, Accessory limbs did not grow from PBs (upper row) of animals treated with 10 nM SAG (n = 1 of 32), while ABs on the contralateral forelimbs of the same animals (lower row) developed into accessory limbs (n = 23 of 32), indicating efficient deactivation of HH signalling. Data from one experiment. Yellow dashed line demarcates the AB. See Extended Data Fig. 6a–c, f for controls and Supplementary Table 1 for details. b, PBs expressing FGF8 after baculovirus transduction grow into accessory limbs (n = 27 of 50; data from three experiments). See Extended Data Fig. 6d–e, g for controls and Supplementary Table 5 for details. c, PBs expressing ectopic FGF8 and treated with 8 μM cyclopamine, an inhibitor of HH signalling, do not develop into accessory limbs (n = 3 of 48; data from four experiments). See Extended Data Fig. 10 for controls and Supplementary Table 6 for details. d, Expression of FGF–SHH loop components in PBs treated with 10 nM SAG, untreated or complemented with anterior skin, analysed with Nanostring nCounter technology. Note the lack of fgf8 induction by SAG despite the induction of the HH signalling targets gli1 and ptc1, which indicates functional HH signalling. Each data point is the mean normalized count from biological replicates (n = 2 or 3; for individual n values and complete data set, see Supplementary Table 7b; data from one experiment). Error bars represent s.d. For data points where the s.d. was low or where n = 2, no error bars are shown. The dashed lines are for visual aid and indicate the level of mature (day 0) expression. Extended Data Fig. 7 shows in situ hybridization of selected genes. Scale bars, 2 mm. dpw, days post-wounding.
We next investigated the relevance of our findings to normal regeneration. Treatment of upper arm blastemas with cyclopamine starting at day 6 after amputation blocked regeneration (Fig. 3a) and yielded changes in the expression of extracellular signalling factors (Fig. 3b and Supplementary Table 10). The limb blastema marker prrx1 showed similar profiles in blastemas treated with cyclopamine or ethanol (control) up to day 16. In contrast, fgf8, fgf9, and fgf10 showed sustained expression under control conditions, whereas in cyclopamine-treated blastemas, the initial induction of fgf8, fgf9, and fgf10 up to day 8 was followed by downregulation. Interestingly, grem1 showed a faster kinetic, peaking after 6 days before being downregulated in cyclopamine-treated blastemas, and shh expression started to be downregulated at day 10 in cyclopamine-treated blastemas. These profiles are consistent with GREM1 being upstream of fgf gene expression, and with FGFs being required for shh maintenance14,19,22,23,24. Two late-stage genes associated with distal limb bud development, fgf17 and hoxa13, were strongly expressed only in ethanol-treated blastemas, suggesting that FGF–SHH signalling is required for substantial distalization during limb regeneration. These results show that the FGF–SHH loop is active during regeneration and is required for blastema outgrowth. Additionally, the anterior localization of fgf8, but not of fgf9, fgf10 or fgf17, in the limb blastema, and the lack of expression of fgf8 in PB cells, suggest that the cross-induction between anteriorly restricted fgf8 and posteriorly restricted shh (Extended Data Fig. 9a, c) is an essential part of normal limb regeneration (Fig. 4).
a, Inhibition of HH signalling with cyclopamine blocks progression of limb regeneration. Limbs treated with cyclopamine (8 μM) starting from day 6 after amputation do not regenerate (n = 0 of 14), whereas ethanol-treated controls do (n = 6 of 6). Data from three experiments. Scale bars, 2 mm. b, Nanostring nCounter gene expression analysis of limbs treated with 8 μM cyclopamine or ethanol. The schema illustrates the experimental setup. The blastema gene prrx1 shows a similar profile in control and cyclopamine-treated samples, whereas genes involved in the SHH–FGF loop and the late genes fgf17 and hox13 diverge under the two conditions. Each data point is the mean normalized count of data from biological replicates (n = 3 or 4; for complete data set and individual n values, see Supplementary Table 10; data from one experiment). Error bars represent s.d. For data points where the s.d. was low, no error bars are shown.
Anterior cells of the blastema express fgf8, whereas shh is expressed in the most posterior part. The initial upregulation of fgf8 in anterior cells is independent of interaction with posterior cells and the initial upregulation of shh in posterior cells is independent of interaction with anterior cells. Later, the expression of fgf8 and shh becomes interdependent in a positive feedback loop, probably involving GREM1, that is expressed primarily in the anterior half of the blastema but also in some posterior hand2-expressing cells. FGFs are the factors that can drive blastema outgrowth and limb regeneration.
In summary, we have delineated the mechanism underlying the anterior–posterior juxtaposition that has long been known to be required for limb regeneration. We have identified single factors that can substitute for anterior and posterior tissue functions: SHH for posterior and FGF8 for anterior tissue. This work shows that, in axolotls, the anterior and posterior tissue provide cells that are restricted in their competence to express complementary soluble signalling factors required for the growth and patterning of a mid-bud blastema, a result consistent with the boundary model of limb regeneration12. Posteriorly localized SHH is required for the maintenance of anteriorly restricted FGF8 expression and of non-restricted expression of FGF9, FGF10 and FGF17. Sustained FGF signalling is the key limiting factor for extended proliferation of anterior and posterior cells and subsequent morphogenesis (Fig. 4).
Methods
Data reporting
Animal distribution to experimental groups was randomized within the selection criteria outlined below for different experiments. The investigators were not blinded to allocation during experiments and outcome assessment.
Animal procedures
Axolotls (Ambystoma mexicanum) used in all experiments were bred in captivity in our facility, maintained at 17–24 °C in tap water and fed with artemia. Axolotls of the leucistic (d/d) strain were used in all experiments. Non-transgenic animals were used in all experiments unless indicated otherwise. When necessary, transgenic animals of the caggs::egfp genotype were used26.
Animals of length between 2 and 4 cm from snout to cloaca (that is, between 4 and 8 cm from snout to tail tip) were used in all experiments, except for the SAG treatment experiment in which larger 13–14-cm (snout to tail tip) animals were used (Extended Data Fig. 2b). In any given experiment, the range in the length (snout to cloaca) of animals used was no more than 0.4 cm, to ensure comparable dynamics of regeneration. Among different replicate experiments, animals of similar size were used. For any given experiment, sibling animals were preferentially used. In cases when this was not possible, animals from different matings were evenly distributed among the different experimental subgroups. It was not possible to determine the sex of the animals at the stage at which the experiment was performed and therefore this was not taken into account.
All animal procedures were carried out in accordance with the laws and regulations of the State of Saxony, Germany.
Surgical procedures
For all experimental procedures, axolotls were anaesthetized in 0.03% benzocaine (Sigma) before surgery. All surgeries were performed using Olympus SZX10 microscopes.
To initiate normal limb regeneration, limbs were amputated either through the upper arm or lower arm.
To generate ABs and PBs, wounds were created by cutting out a rectangular piece of skin from the anterior or posterior side of the distal half of the upper arm. Two ventral nerves were dissected free, severed at the elbow level and deflected to the wound.
Skin transplantation to wounds was performed similarly. A slightly larger patch of skin compared to AB or PB production was removed from the host distal half of the upper arm and nerves were deflected to the wound. A square skin flap from the donor arm was placed on the distal half of the host wound, while leaving the proximal half uncovered with nerves sticking out. To allow healing of the transplant, animals were left for 15 min to 1 h in a Petri dish humidified with 0.03% benzocaine; afterwards animals were transferred into tap water.
Treatment of animals with chemical agonists or inhibitors
In all experiments that required animals to be treated with chemical compounds, the compounds were dissolved in tap water and animals were kept in that solution. Solutions were made freshly and exchanged daily. The chemical compounds used in the study were: smoothened agonist SAG (EMD Millipore), PD173074 (Tocris Bioscience), cyclopamine (LC labs), dimethyl sulfoxide DMSO (Sigma) and ethanol (99.9% VWR chemicals). Chemical solutions were prepared in glass bottles covered with aluminium foil that protected the solutions from light, and stirred for a minimum of 30 min with a magnetic stirrer. Solutions were distributed to the boxes in which animals were kept. The animals were then transferred into the boxes with solutions and kept in darkness. During treatment, animals were fed daily for 30 min to 1 h under lab light or preferentially in darkness. Feeding was started 30 min to 1 h before the chemical solution was exchanged.
SAG treatment of ABs and PBs
ABs and PBs were generated by surgical methods as described above. Treatment with SAG was performed as described above and started immediately after surgery or 3 days after surgery for some PB samples. The duration of treatment with SAG varied; the duration of treatment for each experiment is indicated in Supplementary Table 1.
Cyclopamine treatment of blastemas, ABs complemented with posterior skin, and PBs complemented with anterior skin
Limbs were amputated or injured as above. An 8 μM cyclopamine solution in tap water was prepared from a stock of 4 mM cyclopamine in ethanol as described. We found that the effect of cyclopamine was dependent not only on the final concentration but also on the volume of the solution in which animals were kept. In all experiments, the volume of solution per animal was 60 ml. A control solution of ethanol in tap water with similar dilution and volume was also prepared. Treatment with solutions was started 6 days after surgery. The duration of treatment is indicated in the experimental schemas associated with each experiment.
Exogenous expression of genes in ABs or PBs by baculovirus transduction
Baculovirus technology27,28,29 was established in our lab for use as a transient expression system in the axolotl and will be described elsewhere. Baculovirus was pseudotyped with vsv-ged gene, which was inserted into the rescue vector under the baculovirus polyhedrin promoter. Genes of interest (human SHH, axolotl fgf8, and mCherry) were cloned into the baculovirus rescue vector under the CMV promoter using standard restriction enzyme methods.
The generation of pseudotyped baculoviruses carrying the gene of interest was carried out by co-transfection of the rescue vector together with replication-incompetent baculovirus DNA into a modified Spodoptera frugiperda cell line (expresSF+, ProteinSciences Corporation). Upon culture expansion, recombinant baculoviruses were collected, concentrated and purified. Baculovirus titre was assessed by end-point dilution assay in SF-9 Easy Titer cells, a modified Spodoptera frugiperda cell line expressing a reporter egfp gene under the control of the baculovirus polyhedrin promoter30.
Expression of SHH in ABs by baculovirus transduction was achieved by injecting baculovirus–SHH into the anterior sides of mature upper arms, followed by surgery 5 days after injection. Titres of injected baculovirus suspension are shown in Supplementary Table 1.
Expression of mCherry or FGF8 was achieved by injecting the corresponding baculovirus into the posterior or anterior sides of mature upper arms. Titres of the viral suspensions injected were: mCherry, 2.7 × 1010 or 9.4 × 1010; FGF8, 2.7 × 1010 or 8.5 × 1010.
Injections were performed using a glass capillary needle pulled on a micropipette puller (Sutter Instrument Co. model P-97). Glass capillaries (1.2 mm OD, 0.9 mm ID borosilicate glass) were acquired from Harvard Apparatus. For injection into the anterior upper arm, either a tiny incision was made with surgical scissors (FineScienceTools No 15024-10) or the skin was punctured with a 30G needle (BD Bioscience) on the dorsal side of the upper arm just proximal to the elbow. The needle was inserted through the incision/puncture and the virus was injected into the biceps and surrounding connective tissue. For injection into the posterior upper arm, an incision/puncture was made posterior–dorsally just above the elbow, the needle was inserted through the incision/puncture and the virus was injected into the triceps muscles and surrounding connective tissue.
Statistical analysis of outgrowths
First, outgrowths were sorted into several categories according to the extent of outgrowth (see Supplementary Tables 4 and 6). Next, different categories were grouped into two supra-categories: substantial outgrowth versus non-substantial outgrowth, with outgrowths that have at least two cartilaginous segments being the cut-off for what marks a substantial outgrowth. We then used the Cochran–Mantel–Haenszel test with continuity correction for repeated tests of independence on categorical data to assess whether different treatments had statistically significant effects on the likelihood of substantial outgrowths. The two groups of category were: type of manipulation (for example, ABs + FGF8 versus ABs + mCherry) and type of outgrowth (for example, substantial outgrowth versus non-substantial outgrowth).
Choice of sample size in experiments
When we used SAG to investigate the role of HH signalling in accessory limb outgrowth from ABs, the sample size was chosen at random for the first experiment. For subsequent experiments it was chosen based on the rate of accessory limb induction in the first experiment. In experiments involving baculoviral delivery of human SHH or axolotl fgf8 to ABs, the sample size was also chosen based on the rate of accessory limb induction from ABs by SAG.
In experiments testing the potential of SAG to induce accessory limbs from PBs, we used four conditions: PBs treated with SAG, PBs treated with water (negative controls), PBs complemented with anterior skin (positive controls), PBs complemented with posterior skin (negative controls). The sample size for all groups was based on the rate of accessory limb induction from ABs by SAG, as described above.
In experiments involving baculoviral delivery of mCherry or fgf8 to PBs, the sample size was chosen based on the induction rate of accessory limbs from PBs complemented with anterior skin, taking into consideration the false positive rate of accessory limb induction from PBs treated with water, as identified in the PB experiments described above.
In experiments where the determination state of SAG-induced accessory limbs from ABs was assayed by transplanting accessory limb skin onto host anterior or posterior wounds, the sample size was based on preliminary experiments and resulting information about the induction rate of accessory limb outgrowth from PBs complemented with anterior skin or ABs complemented with posterior skin.
Nanostring gene expression analysis
For probe design, sequences of genes of interest were taken from the axolotl transcriptome that has been assembled from Illumina sequencing reads (to be published elsewhere). For genes whose sequence was validated by the Sanger method, the Nanostring probes were designed to the whole validated sequence. For non-validated genes, the Nanostring probes were designed only to the open reading frame of the sequence. Probes were designed by Nanostring Technologies.
RNA isolation for Nanostring analysis was done by a tandem purification using Trizol (Ambion) and Qiagen Micro Kit (Qiagen). The tissue was homogenized in Trizol using a polytron homogenizer. After chloroform addition and centrifugation, the aqueous phase was collected. An equal volume of 70% ethanol was added to the collected aqueous phase and vortexed. The solution was then loaded onto a Qiagen Micro Kit column, and RNA purification was done following the manufacturer’s protocol.
For Nanostring hybridization, each hybridized sample corresponded to one biological replicate. Each biological replicate consisted of a single blastema, AB or PB. In each hybridization we used 200 ng of RNA (analysis of blastemas) or 300 ng of RNA (analysis of ABs and PBs). Samples were hybridized according to the manufacturer’s instructions, and subsequently analysed on the Nanostring Digital Analyzer to obtain the data on the number of counts. Data were normalized based on the counts of housekeeping genes rpl4 and ef1 with Nanostring nSolver software. Statistical analysis (one-way ANOVA with false discovery rate correction, followed by Tukey post-hoc test) was performed in Genespring GX (Agilent) software. Graphs showing expression profiles of genes were generated with Prism6 (Graphpad) software.
Alcian blue/alizarin red staining
The regenerates were fixed overnight in MEMFA (1 × MEM salts, 3.7% formaldehyde), followed by three 5-min washes in PBS. The limbs were dehydrated in 25% and 50% ethanol (EtOH) for 10 min each. They were then placed in alcian blue solution and kept at 37 °C for approximately 1 h or until all cartilage elements were visibly stained. The limbs were then washed in EtOH–acetic acid mix for 1 h, 95% EtOH for 15 min and twice in 1% KOH solution for 10 min each time, until the soft tissue became clear. They were then placed in alizarin red solution, and stained at room temperature until the ossified parts of the bones became visibly stained (usually approximately 25 min). They were subsequently washed in 1% KOH–glycerol mix. Finally, the limbs were placed in 20% glycerol–PBS solution and imaged using an Olympus SZX16 microscope and Cell F software (Olympus).
Cryosectioning
Limbs with accessory limbs were collected and fixed in MEMFA overnight at 4 °C. Afterwards, the limbs were washed for 2 × 5 min and 4 × 10 min in 1× PBS and placed in 20% sucrose solution overnight at 4 °C. The next day they were embedded in optimum cutting temperature (OCT) compound and frozen in plastic blocks. Samples were sectioned into 10-μm-thick slices that were collected on Superfrost-plus microscopy slides (Menzel-Gläser). The sections were allowed to air dry for 1 h before being stored at −20 °C.
Immunostaining
The following antibodies were used for staining: anti-Meis1/2/3, clone 9.2.7 (EMD Millipore); anti-beta-3-tubulin (R&D systems MAB1195); anti-myosin heavy chain (generated in-house at MPI-CBG antibody facility). Staining was performed using protocols previously described31,32.
Paraffin embedding and sectioning
Limbs with accessory limbs or blastemas were collected and fixed in MEMFA for 3 days at 4 °C. Afterwards, the limbs were washed for 2 × 5 min in 1× PBS treated with diethylpyrocarbonate (DEPC) and sequentially dehydrated in 25%, 50%, 75% and 100% ethanol in DEPC-treated water. The samples were left overnight in 100% ethanol. The next day, the limbs were washed for 10 min in fresh 100% ethanol, then for 3× 40 min to1 h in 100% xylol at room temperature, and lastly for 3 × 1 h in melted paraffin at 70 °C. The limbs were placed on the bottom of disposable plastic base moulds filled with molten paraffin and left on a smooth surface to solidify at room temperature. Samples were cut into sections with a thickness of 7–10 μm and transferred onto superfrost-plus microscopy slides (Menzel-Gläser) covered with DEPC-treated water. The slides were placed on a heating platform at 37 °C and allowed to air dry for at least 1 h.
Isolation of genes for in situ hybridization
We used 10–12 freshly frozen 9-day or 15-day blastemas for total RNA extraction using a standard trizol-based protocol. A SuperScript III kit (Invitrogen) was used for cDNA synthesis according the manufacturer’s protocol. cDNA was used for PCR with appropriate primers to isolate the genes of interest. PCR products were run in 2% agarose gel and bands of the expected size were cut out and purified from the gel. PCR products were cloned into a PCR-II Topo vector, using TOPO TA Cloning kit (Invitrogen). Accession numbers are listed after the Acknowledgements section.
For grem1 in situ hybridization we pulled out a sequence of 937 bp by PCR using the following primers: forward TCGCCTGACACTACATAGC, reverse AGATGTGCAAAAGTTCAGAGAT. For hand2 in situ hybridization we pulled out a sequence of 632 bp by PCR using the following primers based on the newt HAND2 sequence: forward TGAGCCTGGTGGGGGGCTTCC, reverse AGCGCCCAGACGTGCTGCGGCCA. For gli3 in situ hybridization we isolated a sequence of 933 bp by PCR using the following primers: forward ATCATTAATAAAGAAGGAGATCCA, reverse ATAGGTCTCTGGGTAGGAAAAG. For hoxd13 in situ hybridization we isolated a sequence of 735 bp by PCR using the following primers: forward GGCCGAGGTTAGTTTTTATAACCGCA, reverse AAAGGAAGACTTCCAGAAGTGCGAGCTCT. For fgf8 in situ hybridization we isolated a sequence of 1,148 bp by PCR using the following primers: forward TTCACGTGCCTCACCTCCACTACCTCAGCA, reverse ATATAAATTGTTCTTCTAAAAGTCCACTGG. For fgf9 in situ hybridization we isolated a sequence of 972 bp by PCR using the following primers: forward ATTAAACCCG CTTTGATTTCTTG, reverse GTAGAGATTAGATG AGTATGTGTTA. Fgf10 and fgf17 sequences were isolated by screening an EST library of 6-day axolotl tail blastema cDNA. For ptc1 and shh in situ hybridization, we used sequences previously described33.
In situ hybridization probe synthesis
The probes were synthesized using linearized plasmids containing genes of interest, SP6 or T7 polymerase (produced in-house), and DIG-labelled nucleotide mixture (Roche). Synthesis was performed for 2–5 h at 37 °C. RNA was purified from the RNA synthesis mix using RNeasy MinElute Cleanup Kit (Qiagen) and a small amount of purified RNA was run on a 1% agarose gel to assess its quality. Concentration was measured on NanoDrop and adjusted to 100 ng μl−1 with hybridization buffer (50% formamide, 10% dextran, 5× SSC, 0.1% Tween, 1 mg ml−1 yeast RNA, 100 mg ml−1 heparin, 1× Denhardt’s solution, 0.1% CHAPS detergent and 5 mM EDTA).
In situ hybridization
Slides were deparaffinized by washing 3 times in 100% xylol and 3 times in 100% ethanol for 10 min each step, dried for 15 min and placed overnight at 70 °C in hybridization buffer containing probes at the following concentrations: fgf8 500 ng μl−1, fgf9 300 ng μl−1, fgf10 300 ng μl−1, fgf17 300 ng μl−1, gli3 500 ng μl−1, grem1 500 ng μl−1, hand2 200 ng μl−1, shh 200 ng μl−1. On the next day, slides were washed in salt solutions at 70 °C as follows: 30 min in 5 × SSC, 40 min in 2 × SSC twice, 20 min in 0.2 × SSC. To reduce background signal coming from nonspecific probe binding, slides were washed in Tris–sodium–EDTA (TNE) solution for 2 × 10 min, treated with 20 μg ml−1 RNase A (Sigma) in TNE for 15 min, and washed in TNE twice, for 2 min and 10 min. After RNase treatment, slides were washed for 2 × 5 min and 1 × 10 min in maleic acid buffer (MAB) and blocked for 1 h with blocking buffer (1% blocking reagent (Roche) dissolved in MAB). Then slides were incubated with anti-DIG-AP antibody (Roche) diluted 1:5,000 in blocking buffer for 1–2 h at room temperature or overnight at 4 °C. Slides were washed at room temperature in MAB for 2 min, 5 min and 3 × 10 min, followed by 2 × 10 min washes in AP buffer. Slides were developed in BM Purple solution (Roche) or preferentially a substrate working solution containing NBT/BCIP (Roche) and 5% polyvinyl alcohol (Sigma) at 37 °C in the dark until the signal became clearly visible. The reaction was stopped by washing in 1 × PBS several times.
In the case of the fgf9 in situ hybridization signal in untreated 12-day ABs (Extended Data Fig. 4f), the signal was localized to the base of the blastema but was not considerably stronger than the background signal that is typically seen in muscle and nerve for limb in situ hybridization samples. Combined with observed reduction in counts of fgf9 after 8 days in Nanostring data (Fig. 1d), we cannot conclusively say whether this basal fgf9 signal represents bona fide expression or background signal.
Imaging and figure assembly
Images of antibody-stained samples were acquired on a Zeiss Axioobserver Z.1 using AxioVision 4.8.1.0 software (Zeiss). Images of in situ-stained samples were acquired on an Olympus BX61VS110 system using Olympus VS-ASW software.
Images were processed with Fiji or Photoshop software. When adjusted, contrast and brightness were adjusted uniformly across the image.
Accession codes
Primary accessions
NCBI Reference Sequence
Data deposits
The following sequences have been deposited in the NCBI GenBank database with the corresponding accession numbers: fgf10, KU882013 ; fgf17, KU882014; grem1, KU882015; hand2, KU882016; gli3, KU882017; hoxd13, KU882018; fgf8, KU882019; fgf9, KU882020.
References
Iten, L. E. & Bryant, S. V. The interaction between the blastema and stump in the establishment of the anterior–posterior and proximal–distal organization of the limb regenerate. Dev. Biol. 44, 119–147 (1975)
Maden, M. Structure of supernumerary limbs. Nature 286, 803–805 (1980)
Maden, M. Supernumerary limbs in amphibians. Integr. Comp. Biol. 22, 131–142 (1982)
Stocum, D. L. Determination of axial polarity in the urodele limb regeneration blastema. J. Embryol. Exp. Morphol. 71, 193–214 (1982)
Bryant, S. V. Regenerative failure of double half limbs in Notophthalmus viridescens. Nature 263, 676–679 (1976)
Bryant, S. V. & Baca, B. A. Regenerative ability of double-half and half upper arms in the newt, Notophthalmus viridescens. J. Exp. Zool. 204, 307–323 (1978)
Stocum, D. L. Regeneration of symmetrical hindlimbs in larval salamanders. Science 200, 790–793 (1978)
Endo, T., Bryant, S. V. & Gardiner, D. M. A stepwise model system for limb regeneration. Dev. Biol. 270, 135–145 (2004)
Satoh, A., Gardiner, D. M., Bryant, S. V. & Endo, T. Nerve-induced ectopic limb blastemas in the axolotl are equivalent to amputation-induced blastemas. Dev. Biol. 312, 231–244 (2007)
Nacu, E. & Tanaka, E. M. Limb regeneration: a new development? Annu. Rev. Cell Dev. Biol. 27, 409–440 (2011)
Bryant, S. V., French, V. & Bryant, P. J. Distal regeneration and symmetry. Science 212, 993–1002 (1981)
Meinhardt, H. A boundary model for pattern formation in vertebrate limbs. J. Embryol. Exp. Morphol. 76, 115–137 (1983)
Laufer, E., Nelson, C. E., Johnson, R. L., Morgan, B. A. & Tabin, C. Sonic hedgehog and Fgf-4 act through a signaling cascade and feedback loop to integrate growth and patterning of the developing limb bud. Cell 79, 993–1003 (1994)
Chiang, C. et al. Manifestation of the limb prepattern: limb development in the absence of sonic hedgehog function. Dev. Biol. 236, 421–435 (2001)
Harfe, B. D. et al. Evidence for an expansion-based temporal Shh gradient in specifying vertebrate digit identities. Cell 118, 517–528 (2004)
Towers, M., Mahood, R., Yin, Y. & Tickle, C. Integration of growth and specification in chick wing digit-patterning. Nature 452, 882–886 (2008)
Zhu, J. et al. Uncoupling Sonic hedgehog control of pattern and expansion of the developing limb bud. Dev. Cell 14, 624–632 (2008)
Niswander, L., Jeffrey, S., Martin, G. R. & Tickle, C. A positive feedback loop coordinates growth and patterning in the vertebrate limb. Nature 371, 609–612 (1994)
Zúñiga, A., Haramis, A. P., McMahon, A. P. & Zeller, R. Signal relay by BMP antagonism controls the SHH/FGF4 feedback loop in vertebrate limb buds. Nature 401, 598–602 (1999)
Lewandoski, M., Sun, X. & Martin, G. R. Fgf8 signalling from the AER is essential for normal limb development. Nature Genet. 26, 460–463 (2000)
Moon, A. M. & Capecchi, M. R. Fgf8 is required for outgrowth and patterning of the limbs. Nature Genet. 26, 455–459 (2000)
Khokha, M. K., Hsu, D., Brunet, L. J., Dionne, M. S. & Harland, R. M. Gremlin is the BMP antagonist required for maintenance of Shh and Fgf signals during limb patterning. Nature Genet. 34, 303–307 (2003)
Panman, L. et al. Differential regulation of gene expression in the digit forming area of the mouse limb bud by SHH and gremlin 1/FGF-mediated epithelial-mesenchymal signalling. Development 133, 3419–3428 (2006)
Nissim, S., Hasso, S. M., Fallon, J. F. & Tabin, C. J. Regulation of Gremlin expression in the posterior limb bud. Dev. Biol. 299, 12–21 (2006)
Mariani, F. V., Ahn, C. P. & Martin, G. R. Genetic evidence that FGFs have an instructive role in limb proximal-distal patterning. Nature 453, 401–405 (2008)
Sobkow, L., Epperlein, H.-H., Herklotz, S., Straube, W. L. & Tanaka, E. M. A germline GFP transgenic axolotl and its use to track cell fate: dual origin of the fin mesenchyme during development and the fate of blood cells during regeneration. Dev. Biol. 290, 386–397 (2006)
Boyce, F. M. & Bucher, N. L. Baculovirus-mediated gene transfer into mammalian cells. Proc. Natl Acad. Sci. USA 93, 2348–2352 (1996)
Kost, T. A. & Condreay, J. P. Recombinant baculoviruses as mammalian cell gene-delivery vectors. Trends Biotechnol. 20, 173–180 (2002)
Kaikkonen, M. U. et al. Truncated vesicular stomatitis virus G protein improves baculovirus transduction efficiency in vitro and in vivo. Gene Ther. 13, 304–312 (2006)
Hopkins, R. & Esposito, D. A rapid method for titrating baculovirus stocks using the Sf-9 Easy Titer cell line. BioTechniques 47, 785–788 (2009)
Kragl, M. et al. Cells keep a memory of their tissue origin during axolotl limb regeneration. Nature 460, 60–65 (2009)
Nacu, E. et al. Connective tissue cells, but not muscle cells, are involved in establishing the proximo-distal outcome of limb regeneration in the axolotl. Development 140, 513–518 (2013)
Schnapp, E., Kragl, M., Rubin, L. & Tanaka, E. M. Hedgehog signaling controls dorsoventral patterning, blastema cell proliferation and cartilage induction during axolotl tail regeneration. Development 132, 3243–3253 (2005)
Acknowledgements
We thank B. Gruhl, S. Moegel, M. Armstead, H. Goers, A. Wagner and S. Kaudel for animal care, H. Q. Le and K. Goehler for assistance, C. Cannistraci for advice on statistical analysis, C. Antos, D. Knapp, W. Masselink, T. Sugiura and Y. Taniguchi for advice on the manuscript, K. Crawford for discussion, S. Eaton for providing us with the plasmid with human SHH, K. Airenne, S. Ylä-Herttuala, and M. Kaikonnen for the plasmid with VSV-GED, members of the MPI-CBG protein facility for assistance in baculovirus preparation, and members of the MPI-CBG antibody facility for the anti-MHC antibody. This work was supported by central funds from the CRTD and MPI-CBG and an ERC Advanced Investigator grant to E.M.T. C.R.O. was supported by the Portuguese Foundation for Science and Technology (FCT). Schematic illustrations of axolotl limbs were redrawn based on illustrations from ref. 10 under copyright transfer agreement to E.N. and E.M.T. as authors.
Author information
Authors and Affiliations
Contributions
E.N. and E.M.T. conceived the project, conceptualized the experiments and wrote the manuscript. E.N. designed and managed the experiments. E.N. and E.G. performed most of the limb experiments and in situ hybridizations. C.R.O. and D.D. established the baculovirus transduction system and C.R.O. performed the human SHH transductions. E.N., E.G., C.R.O. and E.M.T. analysed data. Figures and tables were prepared by E.N. and E.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 SAG-induced accessory limbs contain muscles, axons, and primarily lower arm and hand elements.
a, Immunostaining for myosin heavy chain (MHC) reveals muscle fibres in the accessory limb (AL). b, Immunostaining for β3-tubulin reveals axons in the accessory limb. c, Immunostaining for MEIS, an upper arm identity marker, reveals that the accessory limb contains primarily lower arm and hand elements, as evidenced by the absence of MEIS in the accessory limb. Expression of MEIS is seen in the host upper arm on which the accessory limb was induced. MEIS staining is also observed in the epidermis, however epidermis does not play a role in positional identity. d, Overlay of anti-MHC (green), anti-β3-tubulin (white), anti-MEIS (red) and Hoechst (marks nuclei, blue) staining. Number of accessory limbs analysed, n = 5. Scale bar, 1 mm.
Extended Data Figure 2 HH signalling is sufficient for accessory limb outgrowth from ABs and is necessary for outgrowth from ABs complemented with posterior skin.
a, Treatment of ABs with lower concentrations of SAG still results in accessory limb outgrowth, but with a decreased number of digits. Data from a single experiment. b, SAG (40 nM) also induces accessory limb outgrowth from ABs in large animals (13–14 cm snout to tail tip), indicating that this effect is not size-dependent. Data from a single experiment. c, Blocking HH signalling with cyclopamine (8 μM) in ABs complemented with posterior skin blocks regeneration. Data from a single experiment. d, Percentage of accessory limb outgrowths from ABs treated with different concentrations of SAG. See Supplementary Table 1 for details. Short blue horizontal lines represent average percentages for each condition. The dotted line is for visual aid and intersects the y axis at 0%. Numbers in upper right corners of photomicrographs indicate the number of accessory limb outgrowths out of the total number of ABs. dpw, days post-wounding. Scale bars, 2 mm.
Extended Data Figure 3 SAG treatment does not make the determination state of accessory limbs more posterior.
a, SAG-induced accessory limbs (SAG-AL) regenerate at most a spike after amputation, indicating a lack of functional positional discontinuity. SAG-ALs were amputated and imaged at 14 and 21 days post-amputation. No further growth of the spike was observed 21 days post-amputation. Red or black lines, amputation plane. No SAG-ALs regenerated a patterned accessory limb after amputation (n = 0 of 5). Experimental data from a single experiment. Alcian blue/alizarin red staining: cartilage is blue, ossifications are red. b–d, Testing determination state of SAG-ALs by skin transplantation to ABs or PBs. b, Schema illustrating the experimental setup and potential outcomes. Accessory limbs were generated by SAG treatment of ABs from transgenic animals constitutively expressing GFP. The SAG-ALs were divided into a medial and lateral side, with skin from one side transplanted to an anterior host site and skin from the opposite side transplanted to a posterior host site. Two possible outcomes were expected, depending on whether SAG posteriorized cells during initial accessory limb outgrowth. c, A host AB with a skin transplant from a SAG-AL shows no accessory limb outgrowth, indicating no determination of posterior identity during SAG-AL outgrowth (n = 0 of 22). d, A host PB with a skin transplant from the same SAG-AL as in c results in new accessory limb outgrowth, revealing the anterior identity of the transplanted skin (n = 15 of 21). Data from a single experiment. See Supplementary Table 2 for details. DF, dark field images. GFP panels show the transplanted skin. dpt, days post transplantation of skin. Scale bars, 2 mm.
Extended Data Figure 4 Gene expression analysis of ABs with and without SAG.
a, ABs express blastema markers prrx1, msx1, msx2, and twist1, and anterior marker gli3. The y axis indicates the number of normalized counts obtained by Nanostring nCounter technology. Each data point is the mean normalized count number from biological replicates (n = 3 or 4; for individual n values and complete data set, see Supplementary Table 7a; data from one experiment). Error bars represent s.d. For data points where s.d. was small, no error bars are shown. b, gli3 is expressed at 8 and 12 days in SAG-treated or untreated ABs. c–h, Expression of FGF–SHH loop components in SAG-treated versus untreated ABs. c, d, f, In ABs, fgf8, fgf9, and grem1 maintenance, but not induction, requires SAG treatment. c, d, fgf8 and grem1 are upregulated in ABs in the presence or absence of SAG at 8 days, but are downregulated at 12 days unless SAG is present. f, fgf9 is expressed at the tip of the bumps at 8 days and maintains this expression at the tip in 12-day SAG-treated samples. In untreated 12-day samples there is no more fgf9 expression at the tip, although there seems to be expression at the base of the bump. This could represent a secondary source of fgf9 that is not part of the FGF–SHH circuitry or it could be an unspecific signal; see Methods. g, fgf10 was expressed in all conditions. h, Unlike other fgf genes, fgf17 expression required SAG at all time points. fgf17 was not expressed in untreated samples at either 8 or 12 days, but was expressed at both time points when treated with SAG. e, SAG is not sufficient to upregulate shh in ABs. Shh is not expressed in ABs at 8 or 12 days in the presence or absence of SAG. For detailed information on the number of samples analysed and results, see Supplementary Table 3. Inserts show magnification of positive cells. Images of 12-day SAG-treated blastemas in panels f–h show different sections from the same limb that were hybridized separately for each of the indicated genes. Scale bars: all 8-day images, 100 μm; all 12-day images, 200 μm.
Extended Data Figure 5 FGF signalling is required for SAG-induced accessory limb outgrowth from ABs, and is sufficient to induce single-digit accessory limb outgrowth from ABs.
a, FGF signalling is required for SAG-induced accessory limb outgrowth. ABs were treated with 15 nM SAG or water starting on the day of wounding. At 10 days post-wounding, concomitant treatment with DMSO or 15 µM PD173074 (inhibitor of FGFR1 and FGFR3) was initiated. Treatment was terminated at 22 days post-wounding. Images of ABs were taken at 10, 15 and 30 days post-wounding. ABs treated with water and DMSO fail to develop accessory limbs (n = 1 of 16). ABs treated with SAG and DMSO form accessory limbs (n = 14 of 14). ABs treated with SAG and PD173074 grow until initiation of PD173074 treatment, after which growth halts and ABs fail to develop accessory limbs (n = 0 of 16), eventually regressing. This indicates that FGF signalling acts downstream of SAG and is required for accessory limb outgrowth. Dashed yellow lines demarcate ABs. The experimental data come from two experiments. dpw, days post-wounding. Scale bars, 1 mm. b, Ectopic baculovirus-induced expression of FGF8 in ABs results in single digit accessory limb outgrowths. Percentage of substantial outgrowths from ABs transduced with fgf8 or mCherry (negative control). See Supplementary Table 4 for details. Short blue horizontal lines represent average percentage for each condition. The dotted line is for visual aid and intersects the y axis at of 0%.
Extended Data Figure 6 FGF8, but not HH signalling, is sufficient to drive accessory limb outgrowth from PBs.
a–c, f, HH signalling is not sufficient to drive accessory limb outgrowth from PBs (for details see Supplementary Table 1). a, SAG is sufficient to drive accessory limb outgrowth from ABs (n = 23 of 32) but not PBs (n = 1 of 32) on the contralateral limb of the same animal. Data from one experiment. The images are the same as in Fig. 2a. b, Untreated PBs also produce no accessory limb outgrowth (n = 1 of 37; data from two experiments). c, Positive control: PBs complemented with anterior skin transplants from animals that constitutively express GFP grew into accessory limbs without SAG treatment (n = 10 of 19, data from two experiments). f, Percentage of accessory limb outgrowths from PBs without treatment (H2O), treated with SAG, or complemented with anterior skin. d, e, g, FGF8 is sufficient to drive accessory limb outgrowth from PBs (data from three experiments; for details see Supplementary Table 5). d, PBs expressing baculovirus-induced FGF8 grow into accessory limbs (n = 27 of 50). The images are the same as in Fig. 2b. e, Negative control: PBs expressing baculovirus-induced mCherry do not grow into accessory limbs (n = 3 of 49). g, Percentage of accessory limb outgrowths from PBs transduced with fgf8 or mCherry (negative control). Short blue horizontal lines in all graphs represent average percentage for each condition. Dotted lines are for visual aid and intersect the y axes at 0%. Dashed yellow lines demarcate blastemas. DF, dark field images; dpw, days post-wounding. Scale bars, 2 mm.
Extended Data Figure 7 PBs do not express fgf8 or fgf17 even though intact HH signalling is present, but do express grem1, fgf10 and fgf9.
a, g, PBs do not express fgf8 and fgf17 even upon SAG treatment. Fgf8 and fgf17 are not expressed in PBs at 8 or 12 days in the presence or absence of SAG, but are expressed in positive control PBs containing an anterior skin transplant. b–d, f, Early induction of grem1, shh, ptc1 (a HH signalling target) and fgf10 in posterior wounds is independent of SAG, and SAG is not sufficient for maintenance of grem1 and shh expression. b, c, Grem1 and shh are present in PBs at 8 days and are downregulated by 12 days independent of SAG treatment. This observation differs slightly from the Nanostring nCounter data (Fig. 2d) where grem1 is downregulated at 15 days. The discrepancy is probably due to the variation in regeneration timelines across different experiments. Grem1 and shh are present in control PBs containing an anterior skin transplant at both 8 and 12 days. d, Ptc1 is expressed at 8 days in PBs that were untreated or treated with SAG or complemented with an anterior skin graft. f, Fgf10 is present in all conditions at 8 and 12 days. e, Fgf9 is expressed in a few cells at the tip of 8-day untreated PBs, but is absent from 12-day untreated PBs and 8- or 12-day SAG-treated PBs. Fgf9 is present in control PBs complemented with anterior skin grafts. Detailed information on the number of samples analysed and results can be found in Supplementary Table 3. Inserts show magnification of positive cells. Images of 12-day PBs complemented with anterior skin (e–g) show different sections from the same limb that were hybridized separately for each of the indicated genes. Scale bars: all 8-day samples, 100 μm; all 12-day samples, 200 μm.
Extended Data Figure 8 Schematic illustration of the expression pattern of fgf8, fgf9, fgf10, fgf17, grem1 and shh in ABs and PBs treated with SAG or untreated and PBs complemented with anterior skin transplant.
ABs upregulate fgf8, fgf9, fgf10 and grem1 but not shh in the absence of SAG at 8 days post-wounding, indicating that initial upregulation of fgf8, fgf9 and fgf10 (yellow) and grem1 (blue) is independent of SAG. ABs downregulate expression of fgf8 and grem1 by 12 days, and eventually regress. ABs treated with SAG maintain fgf8 and grem1 expression at 12 days, indicating that HH signalling is necessary for their maintenance. SAG-treated ABs also express fgf17. ABs treated with SAG eventually develop accessory limbs (ALs). Untreated PBs upregulate fgf9, fgf10, grem1 and shh (red) but not fgf8 nor fgf17. SAG-treated PBs also do not upregulate fgf8 or fgf17 and do not produce accessory limb outgrowths, indicating that SHH signalling is not sufficient for accessory limb outgrowth from PBs. PBs complemented with an anterior skin transplant express fgf8, fgf9, fgf10, fgf17, grem1 and shh and form accessory limbs.
Extended Data Figure 9 Expression patterns of gli3, hand2, fgf8, fgf9, fgf10, fgf17, grem1, shh and hoxd13 in normal limb regeneration.
a, Expression of Gli3, hand2, fgf8 and grem1 was analysed on adjacent sections of a 10-day blastema. Sections are arranged from the distal tip of the blastema (top, plane A) to the proximal base of the blastema (bottom, plane E). Gli3 is expressed throughout the blastema, with stronger expression in the anterior half that lacks hand2 expression and weaker expression in the hand2-expressing zone. Fgf8 is expressed in the anterior half that lacks hand2 expression. Grem1 is expressed more proximally than the other genes (starting from plane B). It is primarily expressed in the anterior half of the blastema, but is also expressed in the posterior half, particularly in the proximal part (planes D and E). b, Unlike fgf8; fgf9, fgf10 and fgf17 are not excluded from the hand2-expressing, posterior half of a blastema (11 days). c, Shh is expressed in a small posterior domain in the blastema (12 days). The planes are sections of the same blastema along the proximo-distal axis. Scale bars, 200 μm.
Extended Data Figure 10 HH signalling is required for FGF8-induced accessory limb outgrowth from PBs.
a, Schematic outline of the experiment. Each replicate experiment consisted of transducing limbs with fgf8 or mCherry (negative control), followed by injury with nerve deviation at 10 or 14 days after transduction. Treatment of FGF8-expressing animals with 8 μM cyclopamine or an equivalent amount of ethanol (cyclopamine solvent) was started at 6 days post-wounding. Additionally a group of non-transduced PBs were complemented with anterior skin and treated with 8 μM cyclopamine (as controls for blocking HH signalling) or ethanol. b, Examples of typical outcomes in each condition. Images from the condition PB+FGF8+cyclopamine are the same as in Fig. 2c. Scale bars, 2 mm. dpw, days post-wounding. c, Percentage of substantial outgrowths in each condition. Data from four experiments. For detailed description of the number of limbs in each condition and the way they were scored see Supplementary Table 6. The dotted line is for visual aid and intersects the y axis at 0%. d, The effect of cyclopamine is specific to HH signalling. Limbs were amputated and treatment was started 6 days post-amputation with 8 μM cyclopamine alone or in combination with 600 nM SAG. Treatment with cyclopamine blocked regeneration, which was rescued by co-treatment with SAG. Numbers indicate regenerated limbs out of total amputated limbs. Data from two experiments. dpa, days post-amputation.
Supplementary information
Supplementary Data
This file contains Supplementary Tables 1-6. (PDF 284 kb)
Supplementary Data
This file contains Supplementary Table 7. (XLSX 102 kb)
Supplementary Data
This file contains Supplementary Table 8. (XLSX 60 kb)
Supplementary Data
This file contains Supplementary Table 9. (XLSX 49 kb)
Supplementary Data
This file contains Supplementary Table 10. (XLSX 68 kb)
Rights and permissions
About this article
Cite this article
Nacu, E., Gromberg, E., Oliveira, C. et al. FGF8 and SHH substitute for anterior–posterior tissue interactions to induce limb regeneration. Nature 533, 407–410 (2016). https://doi.org/10.1038/nature17972
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17972
- Springer Nature Limited
This article is cited by
-
Injury-induced cooperation of InhibinβA and JunB is essential for cell proliferation in Xenopus tadpole tail regeneration
Scientific Reports (2024)
-
Multi-species atlas resolves an axolotl limb development and regeneration paradox
Nature Communications (2023)
-
Akt/mTOR integrate energy metabolism with Wnt signal to influence wound epithelium growth in Gekko Japonicus
Communications Biology (2022)
-
The engine initiating tissue regeneration: does a common mechanism exist during evolution?
Cell Regeneration (2021)
-
Optogenetic manipulation of cellular communication using engineered myosin motors
Nature Cell Biology (2021)