Abstract
G protein-coupled receptors (GPCRs) are physiologically important transmembrane signalling proteins that trigger intracellular responses upon binding of extracellular ligands. Despite recent breakthroughs in GPCR crystallography1,2,3, the details of ligand-induced signal transduction are not well understood owing to missing dynamical information. In principle, such information can be provided by NMR4, but so far only limited data of functional relevance on few side-chain sites of eukaryotic GPCRs have been obtained5,6,7,8,9. Here we show that receptor motions can be followed at virtually any backbone site in a thermostabilized mutant of the turkey β1-adrenergic receptor (β1AR)10,11,12. Labelling with [15N]valine in a eukaryotic expression system provides over twenty resolved resonances that report on structure and dynamics in six ligand complexes and the apo form. The response to the various ligands is heterogeneous in the vicinity of the binding pocket, but gets transformed into a homogeneous readout at the intracellular side of helix 5 (TM5), which correlates linearly with ligand efficacy for the G protein pathway. The effect of several pertinent, thermostabilizing point mutations was assessed by reverting them to the native sequence. Whereas the response to ligands remains largely unchanged, binding of the G protein mimetic nanobody NB80 and G protein activation are only observed when two conserved tyrosines (Y227 and Y343) are restored. Binding of NB80 leads to very strong spectral changes throughout the receptor, including the extracellular ligand entrance pocket. This indicates that even the fully thermostabilized receptor undergoes activating motions in TM5, but that the fully active state is only reached in presence of Y227 and Y343 by stabilization with a G protein-like partner. The combined analysis of chemical shift changes from the point mutations and ligand responses identifies crucial connections in the allosteric activation pathway, and presents a general experimental method to delineate signal transmission networks at high resolution in GPCRs.
Similar content being viewed by others
Main
A thermostabilized, detergent-resistant mutant of β1AR10,11,12 (TS-β1AR; see Methods) was selectively labelled with [15N]valine and produced in insect cells without further chemical modifications. Its 28 valine residues are homogeneously distributed across the receptor (Extended Data Fig. 1) at locations suitable to sense ligand binding and receptor activation. Although resonances of main chain atoms are considerably more difficult to observe than those of mobile side chains of surface residues, they are expected to be better reporters of functional, long-range backbone motions. We succeeded to obtain well-resolved TROSY (transverse relaxation-optimized spectroscopy) spectra of the valine 1H–15N backbone resonances of detergent-solubilized TS-β1AR in its apo form and in complexes with six ligands (Extended Data Fig. 1) ranging in their efficacy from antagonists to agonists (Extended Data Table 1). Despite the absence of deuteration and very short T2 relaxation times (~4 ms for 1HN), 26 valine resonances could be observed with sufficient sensitivity and resolution. Distinct and reversible chemical shift changes were detected for many valines after ligand exchange. 16 valines were assigned unambiguously and 5 tentatively using spectra from 18 point mutants, as well as further spectral and structural information (Extended Data Table 2).
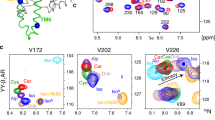
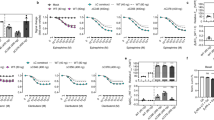
Many valine residues in the vicinity of the ligand binding pocket could be assigned, showing chemical shift changes that report on the ligand functional groups (Fig. 1). Remarkably, residue V172(4.56) (the number in parenthesis corresponds to the Ballesteros–Weinstein numbering system13), which is located close to the ligand aromatic head group, exhibits an unusual 15N chemical shift of ~105–110 p.p.m. (Extended Data Figs 1, 2 and Fig. 1c). This anomaly seems caused by a distorted backbone geometry, which is presumably conserved among adrenergic receptors and results from a missing hydrogen bond to the proline at position 176(4.60) (Extended Data Fig. 2). Instead, the carbonyl of V172(4.56) participates in a water-mediated hydrogen bond network, which connects the ligand binding site, TM3, TM4, TM5, and TM612,14. Seemingly as a result of these interactions, the V172(4.56) 1H–15N resonances cluster according to the substitution patterns of the ligand head group (Fig. 1c): one cluster is observed for the partial agonists/antagonists cyanopindolol, alprenolol and carvedilol, which have larger head groups with ortho- and/or meta-substitutions; a second cluster is observed for the agonists isoprenaline and dobutamine, which bear a meta- and para-substituted catechol ring. We attribute the distinct chemical shifts for isoprenaline or dobutamine to the loss of a coordinated water caused by specific hydrogen bond interactions between their catechol moieties and the side chain of S215(5.461) (Extended Data Fig. 2d).
a, Partial view of the β1AR–carvedilol crystal structure (4AMJ) showing valine residues (blue spheres) in the vicinity (<8.5 Å) of the ligand (magenta sticks) binding site. b, Chemical structures of the β1AR ligands used in this study. Ligand affinities derived from whole-cell binding assays on the thermostabilized β36-m23 β1AR construct17 are indicated as pKD values. Similar pK values were measured for the TS-β1AR construct (Extended Data Table 1). c, Left, ligand-induced response of V172(4.56) 1H–15N resonances. The black bar represents a scale of 0.1 p.p.m. (1H) and 1 p.p.m. (15N). The labels o, m, p indicate ligands with ortho, meta, and para substitutions at the head group, respectively. c, Right, partial view of the β1AR–carvedilol structure (4AMJ) showing the interaction network connecting V172(4.56) to S215(5.461), P219(5.50), I129(3.40), and F299(6.44). d, Left, representation as c, left, for the 1H–15N resonances of V314(6.59). Centres of resonances are indicated by circles. d, Middle, correlation of a best-fit linear combination of the V314(6.59) chemical shifts (−49.3 δ1H + 2.02 δ15N + 133) to the ligand affinity pKD (Extended Data Table 1). d, Right, partial view of the carvedilol (red, 4AMJ), dobutamine (orange, 2Y01), cyanopindolol (green, 2VT4 B), and isoprenaline (blue, 2Y03) complex structures showing the ligand-induced movement of V314(6.59). e, Left, representation as d, left, for the 1H–15N resonances of V125(3.36) and V103(2.65). e, Middle, correlations of best-fit linear combinations of chemical shifts for V125(3.36) (−0.402 δ1H + 1.17 δ15N − 132) and V103(2.65) (−336 δ1H − 634 δ15N + 7.64·104) to the ligand insertion depths and tail volumes (Extended Data Table 1), respectively. e, Right, partial view of the carvedilol, dobutamine, cyanopindolol, and isoprenaline crystal structures (representation as d, right) showing the ligand-induced movement of V125(3.36), D121(3.32), and V103(2.65).
Remarkably, the resonance positions of V172(4.56) in complex with the antagonist atenolol strongly differ from the already described complexes and the apo form: considerable 1HN (>0.4 p.p.m.) and 15N (>4 p.p.m.) upfield shifts indicate, respectively, a further weakening of the main chain hydrogen bond V172(4.56)HN···I168(4.52)O and a stronger kink of the backbone. This rearrangement seems caused by the insertion of the para-acetamide group of the ligand head between residues S215(5.461) and V172(4.56) (Extended Data Fig. 2d). This will lead to a substantial disruption of the TM3–TM4–TM5 interface, thereby precluding receptor activation, in agreement with atenolol’s inverse agonist pharmacology. Thus, the amide chemical shifts of V172(4.56) constitute a very sensitive readout for the state of this water-mediated, inter-helical activation switch.
The 1H–15N chemical shifts of further residues in the vicinity of the binding pocket report on additional characteristics of the ligands. V314(6.59) and V202(ECL2) are located at the extracellular surface of the receptor in a “vestibule” next to the entry/exit pathway of the orthosteric binding site15,16. The resonances of these residues are either severely broadened or undetectable in the absence of ligands, whereas they are observable in the presence of ligands (Fig. 1d and Extended Data Fig. 3). A line shape analysis for V314(6.59) (Extended Data Fig. 4) indicates that this extracellular part of the receptor undergoes micro- to millisecond motions in the apo form, which are quenched by ligand binding. This is consistent with results on β2AR, which suggest that high-affinity ligands stabilize the conformation of ECL2 and ECL35. The ligand-induced shifts of the V314 resonance correlate strongly (r2 = 0.95) with the reported ligand affinity17 (Fig. 1d). Interestingly, the resonances of V314(6.59) in the apo form and in high-affinity ligand complexes are very close. This may indicate that the high-affinity ligand complexes mimic the average apo conformation. Finally, the 1H–15N chemical shifts of V125(3.36) at the bottom of the binding site and of V103(2.65) close to the ligand tail reveal additional trends (Fig. 1e): the chemical shifts of V125(3.36) correlate with the depth of ligand insertion towards the central part of TM3 (r2 = 0.81), whereas those of V103 correlate with the volume of the ligand tail (r2 = 0.90).
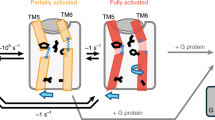
Compared to inactive β2AR, complexes of activated β2AR with either G protein2 or the G protein-mimicking nanobody NB809,18 show large movements at the intracellular sides of TM5, TM6 and their intervening loop ICL3, which form the binding site for the G protein. These conformational changes are expected to be conserved throughout the GPCR family19. Four valine residues could be assigned in this region of TS-β1AR: V226(5.57), V230(5.61), V280(6.25), and V298(6.43) (Fig. 2). In contrast to the chemical shift changes in the vicinity of the ligand binding pocket, which depend strongly on the ligand chemistry, the shifts of the TM5 residues observed in this region report on ligand efficacy. This effect is most prominent for residue V226(5.57), for which the 1H–15N resonances fall on one line from antagonists to agonists (Fig. 2a). The chemical shifts for the different ligands correlate very strongly (r2 = 0.89) with their reported17 efficacies for Gs signalling (Extended Data Table 1). This highly linear effect suggests that the receptor filters the diverse input signals from the various ligands to a unified and precise structural response on TM5, which can be read out by the chemical shifts of V226(5.57). Interestingly, the V226(5.57) atenolol peak is situated at a position corresponding to lower efficacy than for the apo receptor. This gives direct structural evidence of atenolol’s inverse agonist action, which reduces the activation relative to the basal level of the apo receptor.
a, Left, response of the V226(5.57) 1H–15N resonance to various ligands (colour coding as in Fig. 1). The centres of resonances are indicated by circles. The 1H–15N resonances fall on one line from atenolol (antagonist) over apo to alprenolol (partial agonist), carvedilol (antagonist), cyanopindolol (partial agonist), dobutamine (full agonist) and isoprenaline (full agonist). a, Right, correlation of a best-fit linear combination of the V226(5.57) chemical shifts (−515 δ1H −31.7 δ15N +8.41·103) in different ligand complexes to their efficacy for the Gs signalling pathway17. b, Left, overlay of TM5 and TM6 backbones of thermostabilized β1AR in antagonist- (blue, PDB code 4AMJ) and agonist-bound (green, PDB code 2Y03) form. The agonist does not induce detectable helix movements. b, Right, TM5 and TM6 backbone movements upon activation in human β2AR. The overlay of inactive (blue, PDB code 2RH1) and G protein-bound β2AR (magenta, PDB code 3SN6) structures shows the large bend of TM6 along with the smaller conformational change of TM5 upon activation. Hydrogen bonds 5.57-HN···5.53-O and 5.61-HN···5.57-O are indicated by dashes. According to the behaviour of the 1HN chemical shifts of V226(5.57) and V230(5.61) in TS-β1AR, these hydrogen bonds expand in an efficacy-dependent manner during agonist binding. c, Response of the 1H–15N resonances for V226(5.57), V230(5.61), V298(6.43), and V280(6.25) to various ligands. For clarity, only the centres of resonances are depicted as circles with colour coding as in Fig. 1. The black bar represents a scale of 0.1 p.p.m. and 1 p.p.m. for the 1H and 15N chemical shifts, respectively. The schematic representation of the receptor indicates the locations of the respective valine residues at the cytoplasmic sides of TM5 and TM6 within the helical bundle of β1AR.
Current high-resolution structures of β1AR do not show significant changes between antagonist- and agonist-bound forms (Fig. 2b). The decrease of the V226(5.57) 1HN chemical shift by about 0.2 p.p.m. from the agonist isoprenaline to the antagonist atenolol indicates a lengthening of the V226(5.57)-HN···I222(5.53)-O hydrogen bond by about 0.05 Å (ref. 20). This small, but clearly NMR-detectable length variation is below the resolution limit of current GPCR structures (Supplementary Text 1), but may indicate the start of TM5 bending towards the active conformation as observed in the G protein-bound form of β2AR (Fig. 2b). Remarkably, this response to agonists occurs even in the thermostabilized receptor TS-β1AR. Albeit reduced in absolute size compared to V226(5.57), V230(5.61) displays similar linear chemical shift changes as a function of ligand efficacy for the Gs pathway (Fig. 2c). As this residue is located one helical turn further towards the cytoplasm, the detected conformational change is not just local, but spans a certain length in TM5.
Compared to V226(5.57) and V230(5.61) in TM5, the chemical shift response to ligands is much less pronounced for V298(6.43) and in particular V280(6.25) at the intracellular side of TM6 (Fig. 2c). This suggests that agonist binding to the TS-β1AR does not induce the large conformational change in TM6 observed in the activated β2AR–G protein2 or β2AR–NB8018 complexes. However, G protein activation upon agonist binding has been reported for other less thermostabilized β1AR constructs17, indicating that they can still be activated, albeit at low levels. Thus we reverted the mutations most likely to interfere with the activation mechanism in TS-β1AR, that is, I129(3.40)V in the connector switch15, Y227(5.58)A in TM521,22, and Y343(7.53)L in the NPxxY motif of TM714,22 to the native residues. These reverse mutants were then tested for G protein activation and the NMR response in TM6. A summary of the results is given in Extended Data Table 3. The original TS-β1AR, the single mutants TS-β1AR(V129I), TS-β1AR(A227Y), and TS-β1AR(L343Y) as well as the double mutant TS-β1AR(V129I/A227Y) showed no detectable G protein activation upon isoprenaline binding (Extended Data Fig. 5). However, G protein activation was detectable for the least thermostable TS-β1AR(A227Y/L343Y) double mutant (Tm reduced by 11 °C relative to TS-β1AR), which recovers the conserved tyrosines in TM5 and TM7 that are known to stabilize the active state of rhodopsin22. None of the reverse mutants showed major changes in the NMR spectra of various ligand complexes compared to the original TS-β1AR (Extended Data Fig. 6). In particular, residues V298(6.43) and V280(6.25) at the intracellular side of TM6 did not show an increased response to agonists. This is in agreement with recent DEER (double electron–electron resonance) and 19F-NMR data showing that agonists alone do not fully stabilize the active state of TM6 in β2AR9. Interestingly, the V129(3.40)I and the V129(3.40)I/A227(5.58)Y mutations shifted the 1H–15N resonances of V226(5.57) towards a more active (that is, bent) state of TM5 in both the atenolol- and isoprenaline-bound forms (Extended Data Fig. 6b), thereby given direct experimental evidence for an allosteric activation pathway spanning about 13 Å from I129(3.40) on TM3 to V226(5.57) on TM5.
With the exception of rhodopsin23, the stabilization of fully active GPCR conformations seems to require binding of an agonist and an intracellular partner8,9. Indeed, when both the agonist isoprenaline and the G protein-mimicking nanobody NB8018 were added to TS-β1AR(A227Y/L343Y), very large chemical shift responses for many valine residues in TM3–TM6 were observed, whereas no change was observed for several valines in TM1, 2, and 7 (Extended Data Fig. 7). This very strong response extends even to the extracellular residue V314(6.59), providing evidence of a long-distance connection from the G protein binding site to the ligand entry site. The strong chemical shift changes are reverted when the partial agonist/antagonist cyanopindolol is added to the isoprenaline-TS-β1AR(A227Y/L343Y)–NB80 complex. The spectrum then becomes identical to that of the ‘pure’ cyanopindolol–TS-β1AR(A227Y/L343Y) complex (Extended Data Fig. 7a), indicating that cyanopindolol replaces isoprenaline and causes the release of NB80. In agreement with the G protein activation data, the isoprenaline-bound original TS-β1AR and the mutants TS-β1AR(A227Y) and TS-β1AR(L343Y) did not show binding of NB80 in the NMR spectra. Moreover, supplementing NB80 to the ultrastable TS-β1AR did not change its affinity for isoprenaline (Extended Data Fig. 7b), whereas it caused a hundred-fold affinity increase in the case of TS-β1AR(A227Y/L343Y) and the truncated native turkey β1AR receptor (tβtrunc)17. This increase is identical to data for β2AR18 and shows the energetic coupling between the NB80 and agonist binding also for β1AR.
In combination, these data prove that agonist binding, even in the absence of a G protein mimic, induces initial changes in the conformational equilibrium of TM5 towards the conformation observed in the G protein complex of β2AR. Remarkably, these rearrangements occur in all thermostabilized forms of β1AR. However, a full shift of the equilibrium towards such an active conformation, including allosteric changes at the extracellular side, occurs only when G protein or its mimetic NB80 is bound. This process requires the presence of both Y227(5.58) and Y343(7.53), which significantly reduce the thermal stability. Different active conformations may be reached for non-G protein effectors such as β-arrestin.
The possibility to detect NMR signals at many receptor sites in response to ligand binding and point mutations provides an experimental method to trace allosteric signalling paths. Figure 3 shows examples of these pathways, derived from the response to the ligands atenolol and isoprenaline and the single point mutations V129(3.40)I, A227(5.58)Y, and L343(7.53)Y. Choosing a cutoff of 0.05 p.p.m. for the resulting combined 1H, 15N chemical shift change (Fig. 3a, red line), long-range (>10 Å) connections become evident throughout the receptor (Fig. 3b, c). Whereas detected ligand signals radiate broadly to almost all helices, the point mutants give evidence of smaller interaction networks connecting TM3 to TM4/5, TM5 to TM3/4/6, as well as TM7 to TM2/3 (Fig. 3b, c). Interestingly, the TM2/TM7 network seems to be only weakly connected to the TM3–TM6 network. Together, these data provide experimental evidence at high resolution of an extensive signal transduction network that connects the ligand binding site to the intracellular sides of TM5, TM6, and TM7. Such a network of loosely coupled allosteric connections has been postulated previously for β2AR on the basis of molecular dynamics simulations24.
a, Combined 1H, 15N chemical shift deviations [Δδ = (Δδ1H2/2 + Δδ15N2/50)1/2] of valine resonances observed upon ligand binding or induced by the indicated point mutations. For ligand binding, the three pairwise deviations Δδ were calculated between the apo, atenolol-, and isoprenaline-bound forms of TS-β1AR. The maximum of these deviations is shown. For the reverse mutants, deviations Δδ are shown relative to TS-β1AR for their apo (black), atenolol- (cyan), and isoprenaline-bound (blue) bound forms. Valines within 10 Å from the Cα atom of the mutated amino acid are shown on a grey background. Distances were calculated using the coordinates of the thermostabilized β1AR (PDB code 4BVN). A red line marks a cut-off value Δδ of 0.05 p.p.m. for significant chemical shift deviations. b, Topology of the signalling network determined from point mutations (left) and ligand binding (right). Signal paths were identified by chemical shift deviations Δδ larger than 0.05 p.p.m. induced by these two perturbations (a). Signal paths to valines within 10 Å from the ligand or point mutation (that is, localized conformational changes) are indicated as dashed lines, and those beyond 10 Å (long-range conformational changes) as solid lines. The ligand signals broadly towards all helices but TM1. In contrast, the network determined by the point mutations is more localized and connects TM3 to TM4/5, TM5 to TM3/4/6, and TM7 to TM2/3. The latter network seems to be divided into two subnetworks involving TM3/4/5/6 and TM2/7. c, Long-range allosteric signal paths identified from ligand binding or point mutations (a and b) indicated on schematic β1AR representations showing the involved TMs. Helices are colour-coded according to b.
In summary, we have shown that highly resolved solution NMR backbone spectra can be obtained for a eukaryotic GPCR. The NMR observations delineate the allosteric signalling pathways and comprehensively connect many previous experimental and theoretical observations, which may ultimately allow to understand the dynamic mechanisms of GPCRs at the atomic level.
Methods
β1AR constructs
The TS-β1AR mutant was derived from the turkey β1AR44-m23 mutant used in crystallographic studies25 by adding three additional thermostabilizing mutations (I129V, D322K, and Y343L) and a neutral mutation (D200E) from the ultra-stable β1AR-JM3 mutant11. As compared to the wild type, TS-β1AR contains truncations at the amino and carboxy termini and intracellular loop (ICL3), a total of nine thermostabilizing point mutations, three further point mutations as well as a C-terminal hexahistidine tag (Extended Data Fig. 1). The final TS-β1AR sequence is MGAELLSQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGA TLVVRGTWLWGSFLCELWTSLDVLCVTASVETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAI SALVSFLPIMMHWWRDEDPQALKCYQDPGCCEFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQ IRKIDRASKRKTSRVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPKWLFVAFNWLGYA NSAMNPIILCRSPDFRKAFKRLLAFPRKADRRLHHHHHH.
Additional valine-to-alanine or isoleucine point mutations were introduced into TS-β1AR for NMR assignment purposes. All constructs were made using the QuikChange site-directed mutagenesis method (Agilent). Baculovirus for insect cell expression was generated using the Bac-to-Bac system (Invitrogen).
β1AR expression and purification
All β1AR constructs were expressed in baculovirus-infected insect cells as described26. Selective labelling by [15N]valine was achieved by growing cells on unlabelled serum-free insect cell medium (InsectXpress, Lonza) and then exchanging into custom-made serum-free medium (SF4, BioConcept) devoid of valine and yeast extract, to which 100 mg l−1 [15N]valine were supplemented. Virus was added immediately after the medium exchange. The culture was harvested at 48 or 72 h post infection.
After cell lysis, the membrane fraction was separated from the lysate via ultracentrifugation and subsequently solubilized with 2% n-decyl-β-D-maltopyranoside (DM, Anatrace). The solubilized membrane fraction was then purified by nickel ion affinity chromatography followed by alprenolol ligand affinity chromatography. The active receptor was eluted with buffer (20 mM Tris, 350 mM NaCl, 0.1% DM, pH 7.5) containing either atenolol (1 mM) or alprenolol (0.1 mM). Final yields of detergent-solubilized receptor were 1.5 mg l−1 of cell culture. The molecular weight of the receptor-detergent complex was estimated as ~100 kDa by static light scattering.
Thermal shift assays of mutant receptors
Detergent-solubilized, purified apo TS-β1AR and reverse-mutation receptors for thermal stability assays were obtained from their atenolol-bound form by washing with buffer devoid of ligand on a HiTrap SP HP (GE Healthcare) column. Their thermal stability was determined by the microscale fluorescent stability assay for binding of the thiol-specific fluorochrome N-[4-(7-diethylamino-4-methyl-3-coumarinyl)phenyl]maleimide (CPM)27 in a Rotor-Gene Q (QIAGEN) real-time PCR cycler using 1 μg of receptor in 20 mM Tris, 350 mM NaCl, 0.1% DM, pH 7.5 and a heating rate of 2 K min−1.
NMR experiments
NMR samples were prepared in Shigemi tubes as 250 μl volumes of typically 100 μM receptor, 1 mM ligand (except for apo form), 20 mM TRIS, 100 mM NaCl, 0.1% DM, 5% D2O, pH 7.5. For isoprenaline or dobutamine, 2 mM of sodium L-ascorbate were supplemented as anti-oxidant. All solution NMR measurements were carried out at 304 K on a 800 MHz or a 900 MHz Bruker Avance III spectrometer equipped with a cryogenic probe. 2D 1H,15N TROSY spectra were recorded with total acquisition periods of 16 ms (15N) and 43 ms (1H) with typical total experimental times of 24–48 h. As compared to a standard TROSY pulse sequence, the 1H–15N INEPT delays were set to 3 ms to reduce magnetization losses from relaxation.
Assignment procedure
To obtain sequence-specific assignment information, we initially attempted to detect HNCO and HNCA correlations on samples additionally labelled with 13C at specific backbone sites28. However, due to low sensitivity, only very few correlations were observable. Therefore, assignments were obtained from a combination of information from TROSY spectra recorded on 18 TS-β1AR valine point mutants with different ligands, four HN(CO) correlations and five distinct structure-based chemical shift predictions (Extended Data Table 2).
Ligand exchange experiments
Receptor complexes with different ligands were generated by sequential exchange according to increasing ligand affinity, that is, in the sequence atenolol-isoprenaline-dobutamine-alprenolol or alprenolol-carvedilol-cyanopindolol. For exchange, the sample was washed three times with buffer devoid of ligand at tenfold dilution in Amicon Ultra 50 kDa cutoff concentrators. Subsequently, the sample was washed again twice with buffer containing 100 μM new ligand, separated by a period of 1 h incubation. Final concentrations of the ligands were adjusted to 1 mM. Apo receptor was generated from the atenolol complex by six washing steps of tenfold dilution in ligand-free buffer using a 1 h incubation period for the last three steps.
NMR NB80 binding experiment
Binding of NB80 to β1AR mutants was assessed using TROSY and 1D proton NMR spectra. These spectra were recorded on the β1AR mutants (TS-β1AR: 132 μM, TS-β1AR(A227Y): 120 μM, TS-β1AR(L343Y): 110 μM, and TS-β1AR(A227Y/L343Y): 120 μM) in the presence of saturating amounts (1 mM) of the agonist isoprenaline before and immediately after addition of an equimolar (relative to the receptor) amount of NB80. For TS-β1AR(A227Y/L343Y) additional spectra were recorded after a further addition of the partial agonist cyanopindolol (1 mM) to the already present isoprenaline and NB80.
Scintillation proximity assay with 3H-dihydroalprenolol
For pharmacological binding assays membranes were prepared from SF9 insect cells as described previously26. The total protein content of the membranes was estimated by A280 measurements using an average extinction coefficient of 1.0 per mg ml−1. All assays were carried out in 96-well plates at 200 μg ml−1 total protein in membranes and 2 mg ml−1 WGA-YSi beads (Perkin-Elmer) in a 100 μl total volume per well. Samples were equilibrated at room temperature for at least 16 h. KD values for the radioactive ligand [3H]dihydroalprenolol (3H-DHA) were determined by titrating 3H-DHA from 0.032 to 100 nM. Non-specific binding was determined in presence of 1 μM S-propanolol to block the ligand binding site. Competition assays were performed in the presence of 20 nM 3H-DHA (hot ligand) and increasing concentrations of the competitor (cold ligand). Dilutions of alprenolol, atenolol, cyanopindolol, dobutamine and isoprenaline were made with phosphate buffered saline (PBS, Biochrom, Germany). Due to the limited solubility of carvedilol in water, stock dilutions of carvedilol were prepared in DMSO. The final concentration of DMSO in the samples was 5%. To test the effect of NB80 binding on the affinities (IC50) of isoprenaline for various receptor mutants, the competition assays were also carried out in the presence of saturating concentrations of NB80 (10 μM). IC50 values were determined by fitting the measured radioactive counts per minute CPM(X) at a specific concentration X of the competitor to the equation  , where CPMmax and CPMmin are maximal and minimal counts of the assay, respectively. The fits were carried in MATLAB (MathWorks, http://www.mathworks.com) with Monte-Carlo estimation of errors. Ki values were calculated from the obtained IC50 values according to the formula
, where CPMmax and CPMmin are maximal and minimal counts of the assay, respectively. The fits were carried in MATLAB (MathWorks, http://www.mathworks.com) with Monte-Carlo estimation of errors. Ki values were calculated from the obtained IC50 values according to the formula  where A is the concentration of the radioactive ligand and KD is its affinity for the receptor determined in the direct binding experiment.
where A is the concentration of the radioactive ligand and KD is its affinity for the receptor determined in the direct binding experiment.
G protein activation assay
G protein activation was measured on purified β1AR mutants reconstituted with MSP1E3D129 into POPC/POPG nanodiscs. MSP1E3D1 was expressed and purified as described29 and cleaved with TEV protease. 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC, Avanti Polar Lipids) and 1-palmitoyl-2-oleoyl-sn-glycerol-3-phospho-(1′-rac-glycerol) sodium salt (POPG, Avanti Polar Lipids) were solubilized at a ratio of 1:1.5 (w/w) POPG/POPC in ND buffer (20 mM HEPES pH 8, 100 mM NaCl, 1 mM EDTA) with 50 mM sodium cholate (Sigma-Aldrich) at 4 °C. 133.3 μM MSP1E3D1 was incubated with 8 mM solubilized POPC/POPG and 10 μM purified β1AR in ND buffer with a final concentration of 24 mM sodium cholate for 1 h at 4 °C. Nanodiscs containing the receptor were separated from empty nanodiscs using a cobalt-chelating resin. The heterotrimeric G protein was prepared by incubating 10 μM recombinant Gαi1 and 10 μM native Gβγt in activation buffer (25 mM HEPES pH 7.5, 150 mM NaCl, 2 mM MgCl2, 1 mM DTT) for 30 min at 4 °C.
G protein activation was detected by the change in tryptophan fluorescence caused by the exchange of GDP for GTPγS, associated conformational changes in the Gα subunit and its dissociation from the Gβγ subunit of the heterotrimeric G protein30. All measurements were carried out on a Varian Cary Eclipse fluorescence spectrophotometer (λex = 295 nm, λem = 340 nm, 1.5 nm excitation slit, 20 nm emission slit, 2 s averaging time, 15 s cycle time) using final sample volumes of 1 ml in 10 × 4 mm cuvettes (Hellma, CH) and magnetic stirrers at 20 °C. Prior to activation, the fluorescence intensity baseline was recorded with 100 nM heterotrimeric G protein for approximately 500 s. The activation was started by adding 6 nM β1AR and 10 μM GTPγS, and the fluorescence intensity was monitored for a further 1 h. For experiments in the presence of an agonist, the concentrated receptor stock solution (1.5 μM) was pre-incubated for 30 min at 4 °C with 40 μM isoprenaline, and the buffer during the measurements contained 2 μM of isoprenaline to maintain the saturation conditions for the receptor.
References
Rasmussen, S. G. F. et al. Crystal structure of the human β2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007)
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011)
Ghosh, E., Kumari, P., Jaiman, D. & Shukla, A. K. Methodological advances: the unsung heroes of the GPCR structural revolution. Nature Rev. Mol. Cell Biol. 16, 69–81 (2015)
Grzesiek, S. & Sass, H.-J. From biomolecular structure to functional understanding: new NMR developments narrow the gap. Curr. Opin. Struct. Biol. 19, 585–595 (2009)
Bokoch, M. P. et al. Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 463, 108–112 (2010)
Kofuku, Y. et al. Efficacy of the β2-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 3, 1045–1049 (2012)
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wüthrich, K. Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 335, 1106–1110 (2012)
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013)
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015)
Warne, T. et al. Structure of a β1-adrenergic G-protein-coupled receptor. Nature 454, 486–491 (2008)
Miller, J. L. & Tate, C. G. Engineering an ultra-thermostable β1-adrenoceptor. J. Mol. Biol. 413, 628–638 (2011)
Miller-Gallacher, J. L. et al. The 2.1 Å resolution structure of cyanopindolol-bound β1-adrenoceptor identifies an intramembrane Na+ ion that stabilises the ligand-free receptor. PLoS One 9, e92727 (2014)
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995)
Deupi, X. & Standfuss, J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Curr. Opin. Struct. Biol. 21, 541–551 (2011)
Dror, R. O. et al. Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc. Natl Acad. Sci. USA 108, 13118–13123 (2011)
González, A., Perez-Acle, T., Pardo, L. & Deupi, X. Molecular basis of ligand dissociation in β-adrenergic receptors. PLoS One 6, e23815 (2011)
Baker, J. G., Proudman, R. G. W. & Tate, C. G. The pharmacological effects of the thermostabilising (m23) mutations and intra and extracellular (β36) deletions essential for crystallisation of the turkey β-adrenoceptor. Naunyn Schmiedebergs Arch. Pharmacol. 384, 71–91 (2011)
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature 469, 175–180 (2011)
Manglik, A. & Kobilka, B. The role of protein dynamics in GPCR function: insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 27, 136–143 (2014)
Grzesiek, S., Cordier, F., Jaravine, V. & Barfield, M. Insights into biomolecular hydrogen bonds from hydrogen bond scalar couplings. Prog Nucl Mag Res Sp 45, 275–300 (2004)
Tate, C. G. & Schertler, G. F. Engineering G protein-coupled receptors to facilitate their structure determination. Curr. Opin. Struct. Biol. 19, 386–395 (2009)
Goncalves, J. A. et al. Highly conserved tyrosine stabilizes the active state of rhodopsin. Proc. Natl Acad. Sci. USA 107, 19861–19866 (2010)
Park, J. H., Scheerer, P., Hofmann, K. P., Choe, H.-W. & Ernst, O. P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 454, 183–187 (2008)
Dror, R. O. et al. Activation mechanism of the β2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011)
Warne, T. et al. The structural basis for agonist and partial agonist action on a β1-adrenergic receptor. Nature 469, 241–244 (2011)
Brueckner, F. et al. Structure of β-adrenergic receptors. Methods Enzymol. 520, 117–151 (2013)
Alexandrov, A. I., Mileni, M., Chien, E. Y. T., Hanson, M. A. & Stevens, R. C. Microscale fluorescent thermal stability assay for membrane proteins. Structure 16, 351–359 (2008)
Vajpai, N. et al. Backbone NMR resonance assignment of the Abelson kinase domain in complex with imatinib. Biomol. NMR Assign. 2, 41–42 (2008)
Bayburt, T. H., Grinkova, Y. V. & Sligar, S. G. Self-assembly of discoidal phospholipid bilayer nanoparticles with membrane scaffold proteins. Nano Lett. 2, 853–856 (2002)
Ernst, O. P., Bieri, C., Vogel, H. & Hofmann, K. P. Intrinsic biophysical monitors of transducin activation: fluorescence, UV-visible spectroscopy, light scattering, and evanescent field techniques. Methods Enzymol. 315, 471–489 (2000)
Han, B., Liu, Y., Ginzinger, S. W. & Wishart, D. S. SHIFTX2: significantly improved protein chemical shift prediction. J. Biomol. NMR 50, 43–57 (2011)
Acknowledgements
We acknowledge T. Sharpe for expert help with biophysical assays. We thank J. Steyaert for providing the NB80 plasmid and M. Rogowski for preparing wild-type β1AR baculovirus. F.B. was supported by Marie Curie and EMBO postdoctoral fellowships. This work was supported by Swiss National Science Foundation grants 31-132857 (S.G.), Sinergia 141898 (S.G., D.B.V., G.F.X.S.), 31-135754 (D.B.V.), 31-153145 (G.F.X.S.), 31-146520 (X.D.), European Union FP7 grant 242135 (S.G.), and COST Action CM1207 (GLISTEN) (X.D. and G.F.X.S.).
Author information
Authors and Affiliations
Contributions
S.G., G.F.X.S., D.B.V., X.D. and S.I. initiated and managed the project. S.I., X.D., F.B., D.B.V., G.F.X.S. and S.G. designed ligand response and selective labelling experiments. C.O. designed the initial TS-β1AR construct and established purification. S.I. designed and prepared all selectively labelled receptor mutants, performed ligand exchange, NB80 binding, and all NMR experiments. S.I. and S.G. designed NMR experiments, analysed and interpreted all data. F.M.H. and D.B.V. purified trimeric Gi protein, designed and performed radioligand affinity and Gi protein activation assays. C.-J.T. prepared NB80 nanobody. S.G., S.I., X.D., F.M.H., D.B.V. and G.F.X.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Position of valine residues in the thermostabilized mutant TS-β1AR.
a, Schematic representation of secondary structure and amino acid sequence of TS-β1AR. As compared to the wild type, the TS-β1AR has truncations at the N and C termini and the intracellular loop (ICL3) as well as nine thermostabilizing point mutations (colour coded green) and three additional point mutations (colour coded yellow). Valines labelled with 15N are indicated by bold circles for assigned (blue) and unassigned (grey) residues. b, Structure of β1AR in complex with carvedilol (PDB code 4AMJ). The protein backbone and carvedilol are shown in ribbon and red stick representation, respectively. The individual valines are depicted as spheres (blue, assigned; grey, not assigned) labelled by residue number. c, Full 1H–15N TROSY spectra of apo TS-β1AR and all investigated ligand complexes. The ligand chemical structures are shown as inserts. Resonances are marked with assignment information (black, firm; cyan, tentative).
Extended Data Figure 2 Effect of ligand head group substitution on hydrogen bond network involving V172(4.56) in β1AR.
a, Sequence alignment of turkey β1AR and human adrenoreceptors around position 4.56 in TM4 showing the conserved proline at position 4.60. b, Schematic representation of the hydrogen bond network between TM4 and TM5 involving V172(4.56) in β1AR. The hydrogen bond network (orange dotted lines) originates at the carbonyl of V172(4.56) and connects to TM5 via two water molecules (w1 and w2, red asterisks). c, Left, phi and psi dihedral angles (averages and standard deviations in case of multiple chains) in TM4 between residues 4.48 and 4.61 for different complexes of β1AR (ligand, PDB code [chain]: carvedilol, 4AMJ [A,B]; cyanopindolol, 2VT4 [B,D] and 4BVN [A]; dobutamine, 2Y00 [A,B] and 2Y01 [A,B]; isoprenaline, 2Y03 [A,B]). V165(4.49) and V172(4.56), for which 1H–15N resonances could be observed, are shown on a green and red background, respectively. c, Right, average of the 1H–15N resonance positions for V165(4.49, green) and V172(4.56, red). The phi and psi values of V172(4.56) are distorted from the normal helical angles due to the loss of the intra-helical hydrogen bond to P176(4.60). Together with the loss of the canonical hydrogen bond, this strongly shifts both the 1H and 15N chemical shifts of V172(4.56) towards smaller p.p.m. values relative to V165(4.49), which has normal, helical phi and psi angles. d, Partial views of the crystal structures of the carvedilol complex (PDB code 4AMJ), the isoprenaline complex (PDB code 2Y03) and a docking model of the atenolol complex based on the cyanopindolol complex crystal structure (PDB code 4BVN). In the carvedilol complex, one water molecule (w2) forms a hydrogen bond network between the carbonyl oxygen of V172(4.56) and the side chains of S211(5.43) and S215(5.461). In the isoprenaline complex, the side chains of S211(5.43) and S215(5.461) are rotated and form hydrogen bonds to the catechol hydroxyl groups of isoprenaline. In this structure, no water molecule is observed at the equivalent position of w2. In the atenolol complex, the para-substituted acetamide of the ligand head ring (dashed circle) inserts between S211(5.43) and S215(5.461) and disturbs the interface between TM3 and TM5 near V172(4.56) (solid circle).
Extended Data Figure 3 Response to various ligands for all assigned valine 1H–15N resonances in TS-β1AR.
Colour coding as in Extended Data Fig. 1: cyan (atenolol), blue (isoprenaline), orange (dobutamine), purple (alprenolol), red (carvedilol), green (cyanopindolol), black (apo receptor). For clarity the centres of resonances are marked by circles. Firmly (tentatively) assigned residues are marked in black (cyan).
Extended Data Figure 4 Evidence for micro- to millisecond dynamics at the ligand entry/exit pathway.
Left, region of the 1H–15N TROSYs showing the V314(6.59) 1H–15N resonance of TS-β1AR in the apo and various ligand-bound forms. The resonances of the apo and atenolol-bound forms are severely broadened in the 15N dimension indicative of backbone dynamics in the micro- to millisecond range. The broadening is not observed for the other more tightly binding ligands. The resonances for the low affinity ligands, atenolol, isoprenaline and dobutamine, and for the high affinity ligands, alprenolol, carvedilol and cyanopindolol cluster at different positions. Interestingly, the resonance of the apo form clusters with the high affinity ligands, indicative of a similar backbone conformation. Right, 15N and 1H line widths of the V314(6.59) resonance for the apo and ligand-bound forms of TS-β1AR.
Extended Data Figure 5 Isoprenaline-induced G protein activation by β1AR mutants.
Activation was measured by the change in tryptophan fluorescence upon exchange of GDP for GTPγS in the Gα subunit of the heterotrimeric Gi protein. The figure shows the time courses of tryptophan fluorescence after the addition of different mutant forms of TS-β1AR in nanodiscs and GTPγS to the heterotrimeric G protein in the presence (red) and absence (black) of the agonist isoprenaline. The increase in fluorescence intensity at time t = 0 is caused by the additional fluorescence of the receptor, the nanodisc scaffold protein and the ligand. For the active mutant TS-β1AR(A227Y/L343Y), this initial rise is followed by an exponential fluorescence increase due to the activation and dissociation of the G protein. The rate of the Gαi activation by TS-β1AR(A227Y/L343Y) (half-life ~290 s at 6 nM receptor) is approximately 4 times slower than for the activation by rhodopsin (half-life ~400 s at 1 nM receptor) using the same assay. This assumes that the Km of Gαi for TS-β1AR(A227Y/L343Y) is of the same scale as for rhodopsin (8.6 nM) and therefore significantly below the (saturating) concentration of Gαi (100 nM). Each mutant was measured three times except for TS-β1AR(A227Y/L343Y) that was measured four times. Representative fluorescence curves from single experiments are shown.
Extended Data Figure 6 Observed NMR effects of the reverse mutations of TS-β1AR towards the native β1AR sequence.
a, 1H–15N TROSY spectra of TS-β1AR and several reverse single and double mutants in complex with either atenolol (cyan) or isoprenaline (dark blue). Resonances are marked with assignment information (black, definite; cyan, tentative). b, Enlarged regions of the 1H–15N correlation spectra showing only the resonance positions of V226(5.57) and V280(6.25) for all mutants in both ligand-receptor complexes. The black bar represents 0.1 p.p.m. in 1H and 1 p.p.m. in 15N. For all mutants, the resonances for V226(5.57) show efficacy-related chemical shift changes between atenolol (diamonds) and isoprenaline (circles). For the A227Y mutants TS-β1AR(A227Y) (green), TS-β1AR(V129I/A227Y) (orange) and TS-β1AR(A227Y/L343Y) (cyan), the 1H–15N resonances of V226(5.57) also exhibit an overall shift due to a ring current effect from the introduced Y227(5.58) side chain, which has no structural significance. However, the V129I mutants TS-β1AR(V129I) (red) and TS-β1AR(V129I/A227Y) (orange) exhibit further 1H–15N shifts towards a more active (that is, bent) state of TM5 relative to the mutants that carry the V129(3.40) residue. For all mutants, the V280(6.25) resonances fall basically in identical positions and show no changes between atenolol and isoprenaline.
Extended Data Figure 7 Binding of NB80 to agonist-activated TS-β1AR(A227Y/L343Y).
a, Sequential addition of isoprenaline (1 mM, blue), NB80 (120 μM, red), and cyanopindolol (1 mM, green) to TS-β1AR(A227Y/L343Y) (120 μM) followed by 1H–15N TROSY spectra. Addition of an equimolar amount of NB80 changes the spectrum drastically. Except for few resonances (62, 89, 102, 103, 172, 280 and 326), the assignments are not transferable from the isoprenaline complex. The addition of cyanopindolol to this mixture apparently replaces isoprenaline in the receptor and drives the spectrum to the purely cyanopindolol-bound form. These results indicate that TS-β1AR(A227Y/L343Y) can be activated, but that the fully active state requires stabilization with a G protein-like partner at the intracellular site. b, Changes in isoprenaline affinity of β1AR induced by NB80 binding. The isoprenaline affinity was assayed for TS-β1AR (orange), TS-β1AR(A227Y/L343Y) (green), and the truncated native turkey β1AR receptor (tβtrunc, blue)17 by radio-ligand competition using 3H-dihydroalprenolol in the presence of saturating amounts of NB80 (10 μM, squares) or its absence (circles). For TS-β1AR(A227Y/L343Y) and tβtrunc the apparent affinity increases by two log units in the presence of NB80, whereas no change occurs for TS-β1AR. Data points are shown as mean and standard deviation of three independent experiments for each condition. c, Combined 1H and 15N chemical shift changes (Δδ = (Δδ1H2/2 + Δδ15N2/50)1/2) between the isoprenaline-TS-β1AR(A227Y/L343Y) and the NB80-isoprenaline-TS-β1AR(A227Y/L343Y) complexes as a function of residue number. Except for the few assigned peaks in the NB80-isoprenaline-TS-β1AR(A227Y/L343Y) complex, chemical shift deviations are calculated as the minimal difference from the isoprenaline peak position to the nearest peak position in the isoprenaline–NB80 complex. Note that these minimal chemical shift deviations present a lower boundary for the deviations that could be obtained from a full assignment of the isoprenaline–NB80 complex. Peak deviations are colour-coded in red (≥0.2 p.p.m.), orange (0.2–0.05 p.p.m.), blue (<0.05 p.p.m.), and grey (ambiguity from peak overlap). d, Representation of the chemical shift response to nanobody binding given in c on the structure of β1AR. Valines in TM1 and TM7 do not show a large chemical shift response. Due to overlap the response for V280(6.25) in TM6 is unclear. Valines in TM2 show a strong response around the sodium binding region near residue D87(2.50), but valines at its extracellular side are not responsive. In general, valine resonances at the extracellular sides of TM3–TM6 are severely shifted. This indicates that the NB80 binding at the intracellular side has a long-range effect on the extracellular side near the ligand binding site.
Supplementary information
Supplementary information
This file contains Supplementary Text 1. (PDF 186 kb)
Rights and permissions
About this article
Cite this article
Isogai, S., Deupi, X., Opitz, C. et al. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 530, 237–241 (2016). https://doi.org/10.1038/nature16577
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16577
- Springer Nature Limited
This article is cited by
-
Programming conformational cooperativity to regulate allosteric protein-oligonucleotide signal transduction
Nature Communications (2023)
-
E. coli “Stablelabel” S30 lysate for optimized cell-free NMR sample preparation
Journal of Biomolecular NMR (2023)
-
Filling of a water-free void explains the allosteric regulation of the β1-adrenergic receptor by cholesterol
Nature Chemistry (2022)
-
Efficient production of a functional G protein-coupled receptor in E. coli for structural studies
Journal of Biomolecular NMR (2021)
-
A high-resolution description of β1-adrenergic receptor functional dynamics and allosteric coupling from backbone NMR
Nature Communications (2020)