Abstract
Innate immunity serves as the first line of defence against invading pathogens such as bacteria and viruses1. Toll-like receptors (TLRs) are examples of innate immune receptors, which sense specific molecular patterns from pathogens and activate immune responses2. TLR9 recognizes bacterial and viral DNA containing the cytosine–phosphate–guanine (CpG) dideoxynucleotide motif3,4. The molecular basis by which CpG-containing DNA (CpG-DNA) elicits immunostimulatory activity via TLR9 remains to be elucidated. Here we show the crystal structures of three forms of TLR9: unliganded, bound to agonistic CpG-DNA, and bound to inhibitory DNA (iDNA). Agonistic-CpG-DNA-bound TLR9 formed a symmetric TLR9–CpG-DNA complex with 2:2 stoichiometry, whereas iDNA-bound TLR9 was a monomer. CpG-DNA was recognized by both protomers in the dimer, in particular by the amino-terminal fragment (LRRNT–LRR10) from one protomer and the carboxy-terminal fragment (LRR20–LRR22) from the other. The iDNA, which formed a stem-loop structure suitable for binding by intramolecular base pairing, bound to the concave surface from LRR2–LRR10. This structure serves as an important basis for improving our understanding of the functional mechanisms of TLR9.
Similar content being viewed by others
Main
The immunostimulatory activity of CpG-DNA is affected by its length, the number of CpG motifs, and the sequences flanking the CpG motif. The core CpG motif, which consists of a hexamer with a central unmethylated CpG, has the general formula RRCGYY (where R represents a purine and Y a pyrimidine)5. Unlike other TLRs, TLRs 7–9 each have a long insertion loop (Z-loop) between LRR14 and LRR15 (Extended Data Fig. 1). Several recent studies have shown that proteolytic processing at the Z-loop is necessary for the creation of functional TLR96,7,8,9,10. To reveal the molecular mechanism by which TLR9 specifically recognizes CpG-DNA and sends signals to the intracellular compartment, we performed biochemical, biophysical and crystallographic studies of the agonistic- and antagonistic-DNA-binding modes.
We screened recombinantly expressed extracellular domains of TLR9 from various species. Of the proteins we examined, the purified yields of the horse (Equus caballus; Ec), bovine (Bos Taurus; Bt) and mouse (Mus musculus; Mm) orthologues (Extended Data Fig. 1) were sufficient for crystallographic study. We used DNA1668_12mer derived from 20-mer single-stranded DNA16683 as agonistic DNA (Extended Data Fig. 2a), and iDNA408411 and iDNA_super as iDNAs (Fig. 1a). DNA1668 and iDNAs (iDNA4084 and iDNA_super) function as an agonist and antagonist, respectively, for both horse and bovine TLR9 (Extended Data Fig. 2b). Unless otherwise noted, the DNAs used in this study were single-stranded. Both DNA1668_12mer and iDNA4084 were able to bind Z-loop-processed and -unprocessed TLR9, demonstrating that the binding of both types of DNA is independent of Z-loop processing (Fig. 1b, c). Z-loop-unprocessed TLR9 remained mostly monomeric, irrespective of the presence of agonistic DNA1668_12mer, whereas the proportion of Z-loop-processed TLR9 present as a dimer increased significantly in the presence of DNA1668_12mer (Fig. 1d and Extended Data Fig. 3). These results indicate that the Z-loop is functionally significant in the ligand-dependent oligomerization of TLR9. By contrast, DNA1668_12mer_GC (DNA1668 with the CpG motif swapped for GpC) and iDNA4084 could induce the formation of only small quantities of TLR9–DNA dimers (Fig. 1d).
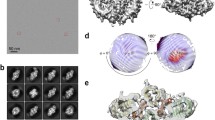
a, DNA sequences used in this study. The DNA regions involved in the intramolecular base pairing in iDNA4084 and iDNA_super are underlined. b, SDS–PAGE analysis of EcTLR9 with unprocessed and processed Z-loops. c, Gel-filtration chromatography of EcTLR9. Black line and dashed line denote absorption at 280 nm and 260 nm, respectively. The ratios of absorbances at 260 nm and 280 nm are shown in parentheses. d, The oligomerization states of EcTLR9 analysed by SV–AUC. The original c(s) distributions were normalized against the height of the main peak (see Methods). kDa, kilodalton; S, Svedberg (unit of sedimentation coefficient).
We determined the crystal structures of the unliganded, agonistic-DNA-bound, and iDNA-bound forms of Z-loop-processed TLR9 molecules (Extended Data Table 1 and Extended Data Fig. 4). In the crystals, the unliganded and iDNA-bound forms of TLR9 are monomeric (Fig. 2a, b), whereas the agonistic-DNA1668_12mer-bound forms of TLR9 are dimeric (Fig. 2c and Extended Data Fig. 5). Similar to other TLRs12,13,14,15,16,17, the agonistic-DNA-bound forms form an m-shaped 2:2 complex in which the C-termini of the two TLR9 protomers are positioned in the centre (Fig. 2c and Extended Data Fig. 5d).
a, Monomer structure of unliganded EcTLR9 showing the lateral face (left) and the convex face from the N-terminal side (right). The N-glycan residues are shown in stick representations with their O, N and C atoms coloured red, blue and grey, respectively. The N- and C-terminal halves of EcTLR9 are coloured light green and orange, respectively. b, Structure of EcTLR9 in complex with iDNA4084 showing the lateral face (left) and the concave face of the N-terminal half (right). Bound iDNA4084 is shown in stick representation and semi-transparent surface representation. The 5′ and 3′ ends of the DNA are indicated. c, Dimer structure of the EcTLR9–DNA1668_12mer complex. EcTLR9 and its dimerization partner EcTLR9* are shown in green and cyan, respectively. The second TLR9 within the dimer and its residues are indicated with asterisks. The DNA1668_12mer molecules are shown in stick representation and semi-transparent surface representation.
The agonistic DNA binds to two equivalent positions in the dimer, and each DNA1668_12mer is recognized by both TLR9 and TLR9* (The second TLR9 within the dimer and its residues are indicated with asterisks) in a bent conformation (Fig. 2c and Extended Data Fig. 6). DNA1668_12mer winds around the N-terminal fragment of TLR9 from the ascending lateral face to the concave face to interact with a region spanning from LRRNT to LRR10 (interface 1). This binding region is consistent with the results of previous reports18,19. Simultaneously, DNA1668_12mer interacts with the loop regions from LRR20–22 in the C-terminal fragment of TLR9* (interface 2) (Extended Data Fig. 6b). Thus, agonistic DNA acts as ‘molecular glue’ to bridge the two TLR9 molecules. This structural feature strongly suggests that only single-stranded DNA can act as an agonist. Accordingly, double-stranded DNAs containing the CpG motif had greatly reduced affinity (Extended Data Fig. 7 and Extended Data Table 2).
In DNA1668_12mer, the G4–T9 sequence corresponding to the consensus hexamer of GACGTT is mainly recognized by TLR9 as opposed to the other part of DNA. The bases of the CpG motif are accommodated in the groove formed by LRRNT, LRR1 and LRR2 in the ascending lateral face of TLR9 (Fig. 3a). The CpG motif and the flanking bases are recognized via interactions with multiple amino acids, as well as via water-mediated hydrogen bonds (Fig. 3a). The C6 moiety in the CpG motif forms direct hydrogen bonds: the cytosine O2 atom with Met106 N and Ser104 Oγ and the cytosine N3 atom with Ser104 Oγ. In addition, the cytosine N4 atom makes water-mediated (W1 and W2) hydrogen bonds with His76, Pro99 and Phe108, and the cytosine ring itself is wedged between Pro105, Phe108 and a neighbouring CpG guanine (G7) (Fig. 3a). Together with Trp47 and Phe49, C6 forms a three-walled cage that accommodates the neighbouring G7. The G7 N2 atom engages in hydrogen bonds with the Trp96 O and O4 atom of T9 at the +2 position, anchoring the guanine ring to the bottom of the CpG binding groove. Because the N2 atom is unique to guanine these interactions define the specificity for guanine in the CpG motif. The thymine ring at the +1 position (T8) stacks with Trp47, and the thymine ring at the +2 position (T9) is inserted into the CpG-binding groove and sandwiched between Trp47 and Trp96. T9 also forms hydrogen bonds to Ser72 and G7. The adenine ring at the –1 position (A5) stacks with Phe108, and in turn G4 (–2 position) stacks onto A5 (Fig. 3a). Because purine bases can form more extensive contacts than pyrimidine bases, the purine–purine sequence preceding the CpG motif is favoured by this three-layered stacking interaction revealed by gel-filtration analysis (Extended Data Fig. 8). The backbone phosphates of G4 and A5 are recognized electrostatically by the positively charged side chains of Lys51, Arg74, His76 and His77. The mutation of several residues (Trp47, Trp96 and Phe108) important for the interaction with DNA resulted in a protein with dramatically reduced binding affinity (Extended Data Table 2), in agreement with the results of the structural analysis.
a, Left, CpG-DNA binding groove formed by LRRNT, LRR1 and LRR2. Middle, overview of CpG-DNA recognition by TLR9 (EcTLR9_DNA1668_12mer interface 1). Right, magnified view of the CpG recognition. The simulated omit electron densities are shown at a contour level of 3.0σ. b, Magnified view of EcTLR9_DNA1668_12mer in interface 2. c, Schematic summary of CpG-DNA recognition. Dashed lines in red and black depict hydrogen bonds and van der Waals interactions, respectively. d, Magnified view of the protein–protein interface. e, NF-κB activation of mouse TLR9 mutants stimulated by agonistic DNA1668 (1 μM). Residues in parenthesis are derived from horse TLR9. ‘Ctrl’ represents the empty vector without mouse TLR9 cDNA. f, The NF-κB activation of mouse TLR9 by DNA1668, DNA1668_GC or DNA1668_met in HEK293T cells. Ctrl, no stimulation. In e and f, data from triplicate experiments are shown as mean ± s.d. and two-tailed Student’s t-test was used to evaluate the statistical significance (**P < 0.01).
Interface 2 also plays an important role in the dimerization of TLR9. Residues of interface 2 engage in four hydrogen bonds and several van der Waals interactions (Fig. 3b, c). In contrast to interface 1, which recognizes the base moieties of the CpG motif, LRR20*–LRR22* from the C-terminal fragment primarily recognize the backbone of CpG motif-containing DNA. The insertion loops in the ascending lateral surface of LRR2, LRR5, LRR8, LRR11, LRR18* and LRR20*, which is characteristic of the TLR7–9 family, are involved in protein–protein interactions with complementary shapes (Fig. 3b, d).
We mutated the residues important for the recognition and examined the ability to activate NF-κB (Fig. 3e). Most of the mutants exhibited reduced or completely abolished activation of NF-κB signalling in response to DNA1668. These results clearly demonstrate that interfaces 1 and 2 are both important for the functional integrity of TLR9. We converted the CpG motif into GC, UG, TG and CA and employed isothermal titration calorimetry (ITC) to determine the affinity of TLR9 for the resultant DNAs. The dissociation constant (Kd) values were 20 nM (CG), 569 nM (GC), 54 nM (UG), 163 nM (TG) and 883 nM (CA), respectively (Extended Data Table 2), demonstrating that the CG sequence is important for binding. The mutations of the CpG motif reduced affinities for TLR9: several interactions were disrupted by swapping CG for GC, whereas the direct interactions between G7 (N2) and Trp96 (O) and T9 (O4) were lost by the conversion of CG into CA. C6 (N4) forms a hydrogen bond with the water molecule (W1) that makes hydrogen bonds with Pro99 O and Phe108 O. The substitution of UG for CG would be unfavourable because W1 is surrounded by three hydrogen bond acceptors. The conversion of CG into TG would result in a weak affinity for the same reason as UG, and the methyl group might further weaken an affinity, possibly by disrupting water molecule clusters; the importance of water-mediated interactions for the recognition of methylated DNA in MeCP2 having been previously identified20. We also employed ITC to assess the pH-dependence of CpG-DNA binding to TLR9. Binding affinity decreased as pH increased, with Kd values ranging from 20 nM at pH 6.0 to 2500 nM at pH 8.0, revealing that the interaction was stronger under acidic than basic conditions (Extended Data Table 2). Consistent with this, the structural study revealed that His residues are concentrated around the DNA-binding region, resulting in a higher-affinity interaction under acidic conditions (Extended Data Fig. 6c). In addition, we examined the binding affinity of methylated CG (DNA1668_12mer_met) for TLR9. The Kd value for this interaction was 50 nM (Extended Data Table 2), demonstrating that methylated CG yields weaker binding. Accordingly, sedimentation velocity analytical ultracentrifugation (SV–AUC) analysis demonstrated that methylation of the CpG motif exhibited a reduced ability to dimerize (Fig. 1d). Consistent with this, DNA1668_met exhibited reduced activation (Fig. 3f).
We also determined the crystal structures of TLR9 bound to iDNAs (iDNA4084 and iDNA_super) (Fig. 2b and Extended Data Table 1). The binding site for iDNA partially overlaps with the binding site for agonistic DNA (Extended Data Fig. 6a, b), and this overlap between binding sites accounts for the antagonistic effect of iDNA. Of particular interest, both iDNA4084 and iDNA_super in complex with TLR9, interacting with LRR2–LRR11, form stem-loop structures that fit snugly into the interior of the ring structure of TLR9 (Fig. 4). The stem-loop structure of the iDNAs is formed by intramolecular base pairing between C1-C2-T3 and T7-G8-G9 (two GC pairs and one TT mismatch pair) in the iDNA4084 complex, and between C1-C2-T3-C4 and G13-A14-G15-G16 (three GC pairs and one AT pair) in the iDNA_super complex (Fig. 4 and Extended Data Fig. 4c–e). The length of the loop seems to be immaterial: iDNA_super, which has a long loop, binds similarly to iDNA4084, which has a short loop. The recognition of iDNAs is primarily mediated via the DNA backbone. Also, the base at the cohesive-end position (G10 in iDNA4084) was directly recognized (Fig. 4 and Extended Data Fig. 5f).
a, Magnified view of iDNA4084 bound to EcTLR9. b, Schematic summary of iDNA recognition. Dashed lines in red and black depict hydrogen bond and van der Waals interactions, respectively.
The structures of the unliganded, agonistic CpG-DNA-bound and iDNA-bound forms of TLR9 described in this study reveal the structural basis of CpG-DNA recognition and signalling by TLR9, as well as the inhibitory mechanism of iDNA. These results will contribute to the development of therapeutic agents that target TLR9.
Methods
Protein expression, purification and crystallization
The DNA encoding the extracellular domain of Toll-like receptor 9 (TLR9) from various species (human (Q9NR96, residues 25–818, 100%), monkey (F6UZJ0, residues 26–817, 95.7%), horse (Q2EEY0, residues 26–817, 83.6%), bovine (Q866B2, residues 25–815, 77.9%), pig (S5R6V0, residues 25–816, 80.4%), rat (M0RAA8, residues 26–818, 71.5%), mouse (Q9EQU3, residues 26–818, 73.4%) and zebrafish (B3DJW3, residues 23–844, 38.1%)), where values in parentheses corresponds to the Uniprot accession number, the region and sequence identity (versus human) of the extracellular domain, with a C-terminal thrombin cleavage site followed by protein A tag were inserted into the expression vector pMT/BiP/V5-His of the Drosophila Expression System. For the preparation of crystallization samples of MmTLR9, a total of seven mutations (N200Q, N242Q, N309Q, N495Q, N568Q, N695Q and N752Q) were introduced to produce the protein with reduced glycosylation sites. Drosophila S2 cells were co-transfected with the TLR9 and pCoHygro vectors. Stably transfected cells were selected in Sf-900 II SFM medium containing 300 μg ml−1 hygromycin. Z-loop processing of TLR9 is important for its function6,7,8,9,10. Therefore, to mimic the Z-loop processing that occurs in the cell, the purification protocol included V8-protease treatment, which yielded the Z-loop-cleaved product. After proteolytic processing, the N- and C-terminal halves of TLR9 remained associated in subsequent purification steps. Protein secreted to the supernatant was captured by IgG Sepharose 6 Fast Flow (GE healthcare) equilibrated with phosphate buffered saline (PBS), washed with ten column volumes of PBS, and eluted by 0.1 M glycine-HCl pH 3.5 and 0.15 M NaCl. Eluent was immediately neutralized by adding with 1/20 volume of 1 M Tris-HCl pH 8.0 and was concentrated to 5–10 mg ml–1 and further purified by Superdex 200 gel filtration chromatography equilibrated with 10 mM Tris-HCl pH 7.5 and 0.15 M NaCl. For BtTLR9 and EcTLR9, concentrated TLR9 was added with 1/10 volume of 1 M Na-acetate pH 5.0 and incubated overnight at room temperature with 1–2 U per mg of protein endo Hf (New England Biolabs) for saccharide trimming. Monomeric fractions from Superdex 200 were collected and was incubated with 1/20–1/50 (w/w) V8 protease (Wako) for 12 to 48 h to cleave the Z-loop and protein A tag. Z-loop-processed TLR9 was further purified by HiTrap SP (GE healthcare) cation exchange chromatography. The column was equilibrated with 10 mM Mes pH 6.0 and 0.1 M NaCl and the bound protein was eluted by a linear gradient from 0.1 to 0.7 M NaCl.
For the crystallizations, purified TLR9 was concentrated to 4.0–6.8 mg ml–1 in 10 mM Tris (pH 7.5), 150 mM NaCl. To prepare the DNA complex of TLR9, the protein solutions were combined with an approximately twofold excess of DNAs (DNA1668_12mer, iDNA4084 and iDNA_super). Crystallization experiments were performed with sitting-drop vapour-diffusion methods at 293 K. The crystallization droplets were made by mixing the equivolume of protein solution and reservoir solution, typically around 0.5–2.0 μl, except in the case of the EcTLR9–DNA1668_12mer complex where the protein solution and reservoir solution were mixed with to a 3:1 ratio. Corresponding to the observed pH dependency of TLR9, the crystals of agonistic forms of TLR9 complexed with DNA1668_12mer were obtained only in acidic conditions (pH 5.8 and pH 5.5 for horse and bovine TLR9, respectively), while the crystal of unliganded and iDNA bound forms of TLR9 were obtained in wide range of pH (4.5–8.0). The crystallization conditions are summarized in Extended Data Table 1.
Data collection and structure determination
Diffraction data sets were collected on beamlines PF-AR NE3A (Ibaraki, Japan) and SPring-8 BL41XU (Hyogo, Japan) under cryogenic conditions at 100 K. Crystals were soaked into cryoprotectant solution summarized in Extended Data Table 1 and then flash-cooled under a cold gas stream. The diffraction data sets were processed using the HKL2000 package21 or imosflm22. The initial phases for the unliganded form of MmTLR9 were determined with the molecular replacement method by using the program Molrep23 with the coordinates of the human TLR8 structure (PDB ID: 3W3J)16. The model was further refined with stepwise cycles of manual model building using the COOT program24 and restrained refinement using REFMAC25 or phenix.refine26 until the R factors converged. The EcTLR9 and BtTLR9 structures were determined by the molecular replacement method using the Molrep23 program using the refined MmTLR9 structure. Ligand molecules, N-glycans and water molecules were modelled into the electron density maps at the latter cycles of the refinement. The quality of the final structure was evaluated with MolProbity27. In the structures of EcTLR9 (unliganded), EcTLR9–DNA1668_12mer, EcTLR9–iDNA4084, BtTLR9–DNA1668_12mer, MmTLR9 (unliganded), MmTLR9–iDNA4084 (form1), MmTLR9–iDNA4084 (form2) and MmTLR9–iDNA_super, 100%, 100%, 98%, 99%, 99%, 99%, 100% and 99% of the residues were in Ramachandran favoured or allowed regions, respectively. The statistics of the data collection and refinement are summarized in Extended Data Table 1. The figures representing structures were prepared with PyMOL28.
Isothermal titration calorimetry
ITC experiments were performed at 298 K in a buffer composed of 50 mM phosphate buffer pH 6.0–8.0, 250 mM NaCl using a MicroCal iTC200 (GE Healthcare). DNAs at a concentration of 50 μM were titrated into 5 μM of wild-type or mutant horse TLR9. The titration sequence included a single 0.4 μl injection followed by 19 injections, 2 μl each, with a spacing of 120 s between the injections. OrigineLab software (GE Healthcare) was used to analyse the raw ITC data. Thermodynamic parameters were extracted from curve fitting analysis with a single-site binding model.
Analytical ultracentrifugation sedimentation velocity
SV–AUC analyses were performed in a ProteomeLab XL-I analytical ultracentrifuge (Beckman Coulter) equipped with a 4-hole An60Ti rotors at 20 °C using Beckman Coulter 12-mm double-sector charcoal-filled epon centerpieces and sapphire windows. The scanning at 42,000 r.p.m. was performed as quickly as possible between 6.0 and 7.2 cm from the axis of rotation with a radial increment of 30 μm. To analyse the dimerization induced by DNA binding, horse TLR9 Z-loop-unprocessed and Z-loop-processed samples were run at a loading concentration of 20 µM with or without equimolar concentrations of DNAs (Fig. 1d). To analyse the concentration dependence of the dimerization, the AUC measurements were performed at protein concentrations ranging from 1.5 µM to 30 µM in the presence of equimolar DNA1668_12mer (Extended Data Fig. 3). All sedimentation velocity experiments were conducted in a buffer containing 10 mM MES and 250 mM NaCl at pH 5.5.
The sedimentation coefficient distributions were obtained using the c(s) method of SEDFIT29. The sedimentation coefficients ranging from 0.1 to 50 S with a logarithmically spaced grid and resolution of 500 were used. The frictional ratio, meniscus, radial and time-invariant noise were floated during the fitting procedure, and a regularization level of 0.68 was used. The partial specific volume, the buffer density and viscosity were calculated using the program SEDNTERP 1.09 and were 0.7407 cm3 g–1, 1.00852 g ml–1 and 1.0256 cP, respectively. The percentages of monomer and dimer were calculated by dividing the corresponding peak area by the sum of the areas under two peaks.
NF-κB-dependent luciferase reporter assay
To check mouse TLR9 response, HEK293T cells were seeded in collagen-coated 6-well plates at a density of 5 × 105 cells per well, and transiently transfected with wild-type or mutant mouse TLR9 cDNAs in pMX-puro-IRES-rat CD2 (1 μg), together with wild-type mouse Unc93B1 cDNA in pMX-puro (0.5 μg) and a pELAM1-luc reporter plasmid (5 ng), using PEI (Polyethylenimine “Max”, MW40,000 ; Polysciences, Inc.) at 36 h before stimulation. To check horse and bovine TLR9 response, HEK293T cells were plated on collagen-coated 10-cm dishes at a density of 6 × 106 cells per well, and transiently transfected with wild-type horse or bovine TLR9 cDNAs in pMX-puro-IRES-rat CD2 (3 μg), together with wild-type human Unc93B1 cDNA in pMX-puro (6 μg) and a pELAM1-luc reporter plasmid (30 ng), using PEI at 30 and 24 h before stimulation. The NF-κB luciferase reporter plasmid, pELAM1-luc, was provided by T. Muta (University of Tohoku, Japan)30. Twenty-four hours after first transfection, cells were reseeded in collagen-coated flat 96-well plates (Corning) at a density of 1 × 105 cells per well. Then, after pre-culture for 4∼6 h, attached cells were stimulated with various DNAs or 100 ng ml–1 recombinant human TNF-α (Wako Pure Chemical Industries) for 6 h. Stimulated cells were lysed by 40 μl of Cell Culture Lysis Reagent (Promega) and 6 μl of lysate was subjected to a luciferase assay using the Luciferase Assay System (Promega). The relative light unit (RLU) of chemiluminescence was measured by GloMax 96 Microplate Luminometer (Promega). Since retroviral vector pMX-puro-IRES-rat CD2 allowed for indirect validation of inserted cDNA expression by checking rat CD2 expression, transfection efficiency of wild-type and mutant mouse TLR9 cDNAs in HEK293T cells was evaluated by examination of cell-surface rat CD2 expression level using a FACSCalibur flow cytometer (BD Biosciences). The activity of pELAM1-luc in each transfected HEK293T cells was also verified by stimulation of recombinant human TNF-α.
Oligonucleotide
Oligonucleotides for the gel-filtration, ITC, AUC and crystallographic analyses were single-stranded DNAs with normal phosphodiester linkage unless otherwise noted. Oligonucleotides for the luciferase reporter assay were single-stranded DNAs with phosphorothioate linkage. Phosphorothioate DNAs (DNA1668, DNA1668_GC, DNA1668_12mer and DNA1668_12mer_GC) were purchased from FASMAC (Kanagawa, Japan). Other phosphorothioate DNAs (iDNA4084, iDNA_super and DNA1668_met) and all phosphodiester DNAs were purchased from Eurofins MWG Operon (Ebersberg, Germany).
Statistical analysis
Data from triplicate samples in Fig. 3e, 3f and Extended Data Fig. 2 were shown as mean ± s.d. and subjected to statistical analysis. Statistical significance was determined by two-tailed Student’s t-tests. A P value of less than 0.01 was considered to be significant. No statistical method was used to predetermine sample size.
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The coordinates and structure-factor data of horse TLR9 (unliganded form), TLR9–DNA1668_12mer, TLR9–iDNA4084, bovine TLR9–DNA1668_12mer, mouse TLR9 (unliganded form), TLR9–iDNA4084 (form1), TLR9–iDNA4084 (form2) and TLR9–iDNA_super have been deposited in the Protein Data Bank under the accession numbers 3WPB, 3WPC, 3WPD, 3WPE, 3WPF, 3WPG, 3WPH and 3WPI, respectively.
References
Janeway, C. A., Jr & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002)
Akira, S. & Takeda, K. Toll-like receptor signalling. Nature Rev. Immunol. 4, 499–511 (2004)
Bauer, S. et al. Human TLR9 confers responsiveness to bacterial DNA via species-specific CpG motif recognition. Proc. Natl Acad. Sci. USA 98, 9237–9242 (2001)
Hemmi, H. et al. A Toll-like receptor recognizes bacterial DNA. Nature 408, 740–745 (2000)
Krieg, A. M. et al. CpG motifs in bacterial DNA trigger direct B-cell activation. Nature 374, 546–549 (1995)
Ewald, S. E. et al. Nucleic acid recognition by Toll-like receptors is coupled to stepwise processing by cathepsins and asparagine endopeptidase. J. Exp. Med. 208, 643–651 (2011)
Sepulveda, F. E. et al. Critical role for asparagine endopeptidase in endocytic Toll-like receptor signaling in dendritic cells. Immunity 31, 737–748 (2009)
Park, B. et al. Proteolytic cleavage in an endolysosomal compartment is required for activation of Toll-like receptor 9. Nature Immunol. 9, 1407–1414 (2008)
Ewald, S. E. et al. The ectodomain of Toll-like receptor 9 is cleaved to generate a functional receptor. Nature 456, 658–662 (2008)
Onji, M. et al. An essential role for the N-terminal fragment of Toll-like receptor 9 in DNA sensing. Nature Commun. 4, 1949 (2013)
Lenert, P. S. Classification, mechanisms of action, and therapeutic applications of inhibitory oligonucleotides for Toll-like receptors (TLR) 7 and 9. Mediators Inflamm. 2010, 986596 (2010)
Jin, M. S. et al. Crystal structure of the TLR1–TLR2 heterodimer induced by binding of a tri-acylated lipopeptide. Cell 130, 1071–1082 (2007)
Kang, J. Y. et al. Recognition of lipopeptide patterns by Toll-like receptor 2-Toll-like receptor 6 heterodimer. Immunity 31, 873–884 (2009)
Liu, L. et al. Structural basis of toll-like receptor 3 signaling with double-stranded RNA. Science 320, 379–381 (2008)
Park, B. S. et al. The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature 458, 1191–1195 (2009)
Tanji, H., Ohto, U., Shibata, T., Miyake, K. & Shimizu, T. Structural reorganization of the Toll-like receptor 8 dimer induced by agonistic ligands. Science 339, 1426–1429 (2013)
Yoon, S. I. et al. Structural basis of TLR5-flagellin recognition and signaling. Science 335, 859–864 (2012)
Kubarenko, A. V. et al. A naturally occurring variant in human TLR9, P99L, is associated with loss of CpG oligonucleotide responsiveness. J. Biol. Chem. 285, 36486–36494 (2010)
Peter, M. E., Kubarenko, A. V., Weber, A. N. & Dalpke, A. H. Identification of an N-terminal recognition site in TLR9 that contributes to CpG-DNA-mediated receptor activation. J. Immunol. 182, 7690–7697 (2009)
Ho, K. L. et al. MeCP2 binding to DNA depends upon hydration at methyl-CpG. Mol. Cell 29, 525–531 (2008)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
DeLano, W. L. The PyMOL Molecular Graphics System. DeLano Scientific LLC. http://www.pymol.org (2008)
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000)
Muta, T. & Takeshige, K. Essential roles of CD14 and lipopolysaccharide-binding protein for activation of toll-like receptor (TLR)2 as well as TLR4 reconstitution of TLR2- and TLR4-activation by distinguishable ligands in LPS preparations. FEBS J. 268, 4580–4589 (2001)
Xu, C., Bian, C., Lam, R., Dong, A. & Min, J. The structural basis for selective binding of non-methylated CpG islands by the CFP1 CXXC domain. Nat. Commun. 2, 227 (2011)
Arita, K., Ariyoshi, M., Tochio, H., Nakamura, Y. & Shirakawa, M. Recognition of hemi-methylated DNA by the SRA protein UHRF1 by a base-flipping mechanism. Nature 455, 818–821 (2008)
Avvakumov, G. V., Walker, J. R. & Xue, S. Li, Y. Duan, S., Bronner, C., Arrowsmith, C. H. & Dhe-Paganon, S. Structural basis for recognition of hemi-methylated DNA by the SRA domain of human UHRF1. Nature 455, 822–825 (2008)
Hashimoto, H., Horton, J. R., Zhang, X. & Cheng, X. The SRA domain of UHRF1 flips 5-methylcytosine out of the DNA helix. Nature 455, 826–829 (2008)
Acknowledgements
We thank the beamline staff members at the Photon Factory and SPring-8 for their assistance with data collection. This work was supported by a Grant-in-Aid from the Japanese Ministry of Education, Culture, Sports, Science, and Technology (U.O., S.U., K.M. and T.S.); the JSPS Japanese–German Graduate Externship (S.U.); the Senri-Life Science Foundation (S.U.); the Takeda Science Foundation (U.O. and T.S.); and the Mochida Memorial Foundation for Medical and Pharmaceutical Research (U.O.).
Author information
Authors and Affiliations
Contributions
U.O. and H.T. expressed and purified recombinant proteins. U.O. performed crystallization and structure determination. T. Shibata and K.M. performed cellular assays. E.K. and S.U. performed AUC analyses. U.O. and H.I. performed ITC experiments. U.O. and T. Shimizu directed the research and wrote the paper with assistance from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of human, horse, bovine and mouse TLR9.
Sequence alignments are displayed for each LRR module. The agonist DNA interfaces 1 and 2 deduced from the EcTLR9–DNA1668_12mer complex are indicated by blue and yellow highlighting, respectively. The antagonist DNA interface deduced from the EcTLR9–iDNA4084 complex is indicated with boxes. The protein–protein interface in the EcTLR9–DNA1668_12mer complex is indicated by bold orange bars below each LRR module. Alignments were performed using Clustal Omega software (EMBL-European Bioinformatics Institute). Residues are coloured to indicate the degree of similarity: red residues are those with the highest similarity, followed by green, blue and black (lowest similarity).
Extended Data Figure 2 The NF-κB activation experiments.
a, DNA1668_12mer retains agonistic activity to TLR9. The NF-κB activation of wild-type mouse TLR9 induced by DNA1668 (TCCATGACGTTCCTGATGCT), DNA1668_12mer (CATGACGTTCCT), DNA1668 or DNA1668_12mer with a CpG to GpC inversion (DNA1668_GC, DNA1668_12mer_GC). DNAs were all complexed with N-[1-(2,3-Dioleoyloxy)propyl]-N, N, N-trimethylammonium methyl-sulphate (DOTAP) and added at concentration of 1 μM. The activities were analysed with an NF-κB-dependent luciferase reporter assay using HEK293T cells co-expressing mouse TLR9 and mouse Unc93B1. A two-tailed t-test was used to determine the statistical significance of differences between control (Ctrl) and stimulated cells, or between each group. **P < 0.01. Data from three independent experiments are shown. b, Horse and bovine TLR9 responses against agonistic and inhibitory DNAs. The NF-κB activation of wild-type horse (left) or bovine (right) TLR9 induced by indicated DNAs. The activities were analysed with an NF-κB-dependent luciferase reporter assay using HEK293T cells co-expressing horse or bovine TLR9 and human Unc93B1. The concentration of agonistic DNAs (DNA1668, DNA1668_GC and DNA1668_met) and inhibitory DNAs (iDNA4084 and iDNA_super) were 10 μM and 1 μM, respectively. Data represent the mean fold induction of NF-κB activity +s.d. (n = 3). A two-tailed t-test was used to determine the statistical significance of differences between control (Ctrl) and stimulated cells, or between each group. **P < 0.01. Data from three independent experiments are shown.
Extended Data Figure 3 Dimerization interaction of TLR9.
a, b, The oligomerization states of EcTLR9 with an unprocessed Z-loop (a) or a processed Z-loop (b) were analysed by SV–AUC at various concentrations of TLR9–DNA1668_12mer (equimolar). The weight-average sedimentation coefficients (Sw) were plotted against TLR9–DNA1668_12mer concentration to determine the Kd value for the dimerization. The dissociation constant for the dimerization of the processed TLR9 is estimated to be 20 μM.
Extended Data Figure 4 Electron densities of DNA bound to TLR9.
a–e, The Fo–Fc omit difference electron densities of DNA1668_12mer bound to EcTLR9 (a) and BtTLR9 (b), iDNA4084 bound to EcTLR9 (c) and MmTLR9 (d), and iDNA_super bound to MmTLR9 (e) contoured at the 3.0σ level. The residues coloured blue in the sequence are not visible in the electron density map. The core hexamer could be unambiguously modelled into the continuous electron density map in the EcTLR9–DNA1668_12mer and BtTLR9–DNA1668_12mer complexes, whereas flanking bases were obscure or not visible. The A5–T12 loop connecting the base-paired region of iDNA_super was not visible in the electron density map, whereas the G4–A6 sequence of iDNA4084 was well defined.
Extended Data Figure 5 Structures of TLR9.
a, Monomer structure of EcTLR9, derived from the EcTLR9–DNA1668_12mer complex. The structure and binding mode of TLR9 are markedly different from those of other CpG-binding proteins20,31,32,33,34. b, Monomer structure of human TLR8, derived from the human TLR8–CL097 (2-(ethoxymethyl)-1H-imidazo[4,5-c]quinolin-4-amine) complex (PDB ID: 3W3J)16. The Z-loops in TLR9 and TLR8 are oriented differently with respect to the concave face of TLR and engage in different interactions with it. The latter half of the Z-loop of TLR8 extends towards the N-terminus, whereas the Z-loop of TLR9 extends towards the C-terminus to interact with LRR15–21. TLR8 has three ordered N-glycans attached to Asn293, Asn511 and Asn590 that project into the inner space of the ring structure, whereas EcTLR9 has only one N-glycan attached to Asn731 that projects inward. As a result, the ring structure of TLR9 has more unoccupied inner space than that of TLR8, an arrangement that is suitable for ligand binding on the concave interior surface. c, Superposition of the overall ligand-induced dimer structures of EcTLR9 (DNA1668_12mer complex, green) and human TLR8 (CL097 complex, purple) by PyMol28, yielding an root-mean-square deviation (r.m.s.d.) value of 2.3 Å. d, Superposition of the overall dimer structures of EcTLR9 (DNA1668_12mer complex, green) and BtTLR9 (DNA1668_12mer complex, purple) by PyMol28, yielding an r.m.s.d. value of 0.7 Å. e, Magnified view of the CpG-binding groove of EcTLR9 and BtTLR9. The amino acid at position 109 was proline for human and serine for mouse (rodents). From the structure, Pro109 made a van der Waals contact with A at the –1 position, but its contact was somewhat close. If A is changed to T, this contact is weakened. Serine at position 109 would accommodate a larger base. f, Magnified view of the G10 of iDNA4084 recognition by TLR9 (EcTLR9–iDNA4084 complex). G10 of iDNA4084 makes three hydrogen bonds with TLR9: N2, N1 and O6 atoms of G10 with the side chains of Ser205, Asp175 and Ser151, respectively. To examine the functional importance of this base, we substituted it with other bases and examined the binding affinity by ITC. The affinity of TLR9 for G10A (Kd = 6 nM) was reduced from that of the original sequence (Kd = 3 nM), but TLR9 exhibited much lower affinity for DNAs with a pyrimidine at this position (Kd = 41 nM for G10C and Kd = 76 nM for G10T), suggesting that this position favours purine over pyrimidine. g, Superposition of iDNA4084 bound to EcTLR9 (green) and MmTLR9 (purple). The binding mode of iDNA4048 is perfectly conserved between EcTLR9 and MmTLR9.
Extended Data Figure 6 Binding interfaces for agonistic and inhibitory DNA.
a, Superposition of the structures of unliganded (grey), DNA1668_12mer-bound (green), and iDNA4084-bound (blue), EcTLR9. TLR9 and DNA are shown in Cα-trace and stick representation, respectively. No significant conformational change was observed upon agonistic DNA binding, as indicated by the small r.m.s.d. value of 0.8 Å between EcTLR9 (unliganded) and the EcTLR9–DNA1668_12mer complex. Instead, the conversion of EcTLR9 into the activated form appears to involve local conformational changes in the loop regions of LRR8, LRR11 and LRR18, all of which are involved in formation of the dimer. No significant structural change was induced by binding to iDNA4084, as indicated by the small r.m.s.d. values of 0.49 Å and 0.45 Å between the unliganded and iDNA4084-bound forms of EcTLR9 and MmTLR9, respectively. b, Surface representations of EcTLR9 structures in the DNA1668_12mer (upper) and iDNA4084 complexes (lower). The protein–protein interface, TLR9_DNA1668_12mer interfaces 1 and 2, and TLR9_iDNA4084 interface are shown in orange, blue, cyan and yellow, respectively. The bound DNAs are shown in stick representation with their 5′ and 3′ ends indicated. DNA1668_12mer buries approximately 1,136 Å2 and 294 Å2 of the accessible surface area of TLR9 and TLR9*, respectively, suggesting that the N-terminal binding site of TLR9 for DNA1668_12mer makes a relatively larger contribution to binding. The binding site for iDNA partially overlaps with the binding site for agonistic DNA. Specifically, LRR4 and LRR5 are both involved in the binding sites for agonistic DNA and iDNAs, although the binding modes of DNA1668_12mer and iDNA4084 are completely different: Arg152 (LRR4), Tyr179 and Lys181 (LRR5) interact with the phosphate of C11 of DNA1668_12mer but also with G8 of iDNA4084. c, Electrostatic potential map of DNA-binding region. The map was calculated at basic and acidic conditions by PyMol28. Surface colours represent the potential from –20 kBT/e (red) to 20 kBT/e (blue), where kB is the Boltzman constant and T is the absolute temperature. The DNA molecule is shown as a stick model.
Extended Data Figure 7 ITC thermograms for ITC data (related to Extended Data Table 2).
Representative ITC thermograms for the ITC data are shown with their pH condition and Kd values for EcTLR9.
Extended Data Figure 8 Significance of the consensus sequence in the CpG-DNA.
Gel-filtration chromatography of EcTLR9 with FITC-labelled DNA. DNA binding to TLR9 was monitored by FITC-fluorescence (excitation 495 nm, emission 520 nm). The parenthesized values indicate the ratios of the fluorescence peak height of the derivative DNA to the original DNA containing the consensus sequence of GACGTT (top left). In each experiment, 0.5 μM EcTLR9 (total volume of 45 μl) with DNA (equimolar) was injected into a Superdex 200 Increase 5/150 GL (GE healthcare) gel-filtration column. The running buffer was 10 mM 2-morpholinoethanesulfonic acid and 250 mM NaCl at pH 5.5. DNAs used in the analyses are shown in each panel. Bases that are changed from the original sequence are highlighted in red. Conversion of the purine–purine sequence (GA) at the –1 and –2 positions of the CpG motif (underlined) to AA and GG resulted in DNAs with affinities similar to the wild-type DNA, but conversion to a pyrimidine–pyrimidine sequence (TT and CC) weakened the affinity, demonstrating that a purine–purine sequence is preferable at these positions. Conversion of the base at the +1 position into C, A or G led to weaker binding, suggesting that T is preferable at the +1 position, although T is not specifically recognized.
Rights and permissions
About this article
Cite this article
Ohto, U., Shibata, T., Tanji, H. et al. Structural basis of CpG and inhibitory DNA recognition by Toll-like receptor 9. Nature 520, 702–705 (2015). https://doi.org/10.1038/nature14138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14138
- Springer Nature Limited
This article is cited by
-
The architecture of transmembrane and cytoplasmic juxtamembrane regions of Toll-like receptors
Nature Communications (2023)
-
Combinatorial treatment rescues tumour-microenvironment-mediated attenuation of MALT1 inhibitors in B-cell lymphomas
Nature Materials (2023)
-
Regulation of the nucleic acid-sensing Toll-like receptors
Nature Reviews Immunology (2022)
-
Structural and functional implications of leucine-rich repeats in toll-like receptor1 subfamily
Journal of Biosciences (2022)
-
Nucleic acid binding by SAMHD1 contributes to the antiretroviral activity and is enhanced by the GpsN modification
Nature Communications (2021)