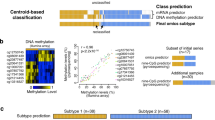
Abstract
Retinoblastoma is a childhood retinal tumour that initiates in response to biallelic RB1 inactivation and loss of functional retinoblastoma (Rb) protein. Although Rb has diverse tumour-suppressor functions and is inactivated in many cancers1,2,3,4,5, germline RB1 mutations predispose to retinoblastoma far more strongly than to other malignancies6. This tropism suggests that retinal cell-type-specific circuitry sensitizes to Rb loss, yet the nature of the circuitry and the cell type in which it operates have been unclear7,8. Here we show that post-mitotic human cone precursors are uniquely sensitive to Rb depletion. Rb knockdown induced cone precursor proliferation in prospectively isolated populations and in intact retina. Proliferation followed the induction of E2F-regulated genes, and depended on factors having strong expression in maturing cone precursors and crucial roles in retinoblastoma cell proliferation, including MYCN and MDM2. Proliferation of Rb-depleted cones and retinoblastoma cells also depended on the Rb-related protein p107, SKP2, and a p27 downregulation associated with cone precursor maturation. Moreover, Rb-depleted cone precursors formed tumours in orthotopic xenografts with histological features and protein expression typical of human retinoblastoma. These findings provide a compelling molecular rationale for a cone precursor origin of retinoblastoma. More generally, they demonstrate that cell-type-specific circuitry can collaborate with an initiating oncogenic mutation to enable tumorigenesis.
Similar content being viewed by others
Main
RB1-mutant retinoblastomas can originate from a cellular state found during retinal development in humans but not in other species9,10. Accordingly, to identify the cellular state and corresponding circuitry that sensitizes to RB1 inactivation, we examined the effects of Rb depletion on human fetal retinal cells. Samples were from post-fertilization weeks 17–19, when all retinal cell types and a range of maturation states are present.
Dissociated retinal cells were transduced with RB1-directed or control short hairpin RNAs (shRNAs), followed by co-staining for the proliferation-associated Ki67 and cell-type-specific markers. RB1 shRNAs abrogated Rb expression in long or medium wavelength (L/M)-opsin+ and thyroid hormone receptor β2+ (TRβ2+) cone precursors as well as in other cell types (Extended Data Fig. 1a). After 2 weeks, Ki67 was detected in cone-precursor-like cells co-expressing the photoreceptor marker CRX and the cone markers L/M-opsin, cone arrestin and RXRγ (Fig. 1a and Extended Data Fig. 1b–h). Ki67+ cone-marker+ cells were first detected 9 days after transduction, whereas clusters were routinely detected by day 23. Ki67 was not detected in cells expressing markers of rods (NRL), bipolar cells (strong CHX10), ganglion cells (BRN-3), or amacrine or horizontal cells (PROX1+ or PAX6+, nestin−) (Fig. 1a and Extended Data Fig. 1i, j). Ki67 was detected in cells expressing markers of retinal progenitor cells (RPCs) or Müller glia (nestin or CRALBP, SOX2), yet in similar proportions after RB1 shRNA or control shRNA (Fig. 1a and Extended Data Fig. 1j). RB1 shRNAs also induced incorporation of 5-ethynyl-2′-deoxyuridine (EdU), an indicator of S phase entry, increased expression of the mitosis marker phosphohistone H3, suppressed expression of the apoptosis marker cleaved caspase 3 (CC3), and induced proliferation in cells expressing cone but not other retinal cell markers (Fig. 1c, d and Extended Data Fig. 1k–n). By contrast, RB1 shRNAs induced CC3 and decreased the number of cells expressing markers of RPCs and glia (Fig. 1b, d and Extended Data Fig. 1n).
a, b, Responses to RB1-733 shRNA (shRB1). D, day; SCR, scrambled control. a, Percentage Ki67+ among cells expressing the indicated retinal cell-type-specific markers. b, Prevalence of DAPI+ (4′,6-diamidino-2-phenylindole+) cells expressing cone (L/M-opsin, CRX), RPC (nestin), or Müller glia (CRALBP) markers. c, d, Responses to RB1-733 and RB1-737 shRNAs (shRB1-733 and shRB1-737, respectively). c, Proliferation of cells, of which >90% were cone marker+ at day 60. d, Percentage CC3+ among cells expressing the indicated markers at day 14. Values and error bars denote mean and s.d. of triplicate assays; P values are from unpaired Student’s t-test. All results replicated at least twice.
To assess whether the Rb-deficient proliferating cone-like cells derived from post-mitotic cone precursors, we examined the effects of Rb knockdown in prospectively isolated retinal cell populations. Populations were isolated by sorting for size, for CD133, which is expressed strongly in maturing photoreceptors and weakly in RPCs11, and for a CD44 epitope expressed by Müller glia and RPCs12 (Fig. 2a). Staining for cell-type-specific markers revealed populations enriched for cone precursors, for rod plus cone precursors, for RPCs plus glia, and for a mixture of rod, ganglion, bipolar, amacrine and horizontal cells (Fig. 2b and Extended Data Fig. 2a–g). In medium and large CD133hi, CD44− populations, 96–98% of cells co-stained for CRX and cone arrestin, which is cone-specific at post-fertilization week 19 (FW19) (Extended Data Fig. 2h). A similar enrichment was observed when cone precursors were identified using CRX and RXRγ (Extended Data Fig. 2h–k).
a, Dissociated FW18 retinal cells sorted by size, CD133 and CD44, with major populations designated. FSC, forward scatter; misc, miscellaneous; SSC, side scatter. b, Percentage of cone arrestin+(cArr+), CRX+ cones, NRL+ rods, nestin+ RPCs and glia, and PAX6+, nestin− horizontal, amacrine or ganglion cells in each population. Lg, large; med, medium; sm, small. c, Responses to RB1-733 shRNA. Percentage of Ki67+ or CC3+ cells at 14 days (top), and cell numbers at days 14 and 23 (bottom). *P < 0.05, **P < 0.01 (compared to scrambled control). Results in a–c are representative of three independent experiments. d, Ki67+, L/M-opsin+ cone precursors (white arrows) in FW19 fovea 15 days after transduction with RB1-733 and RB1-737 shRNAs; and Ki67+, L/M-opsin− cells probably representing RPCs or glia (yellow arrows) after transduction with RB1 shRNA or scrambled control. Scale bar, 20 μm. Values and error bars denote mean and s.d. of triplicate assays; P values are from unpaired Student’s t-test.
RB1 shRNAs induced similar RB1 knockdown in each retinal cell population (Extended Data Fig. 3a). After 2 weeks, Ki67 was detected in 80% of cells in the cone-enriched population (Fig. 2c), probably reflecting a high ratio of shRNA-expressing lentivirus to target cells and cone precursor proliferation. After 3 weeks, cone precursor numbers increased (Fig. 2c). Rb depletion did not induce proliferation in RPCs and glia, but increased the proportion of CC3+ cells entering apoptosis (Fig. 2c). Sorted populations transduced with the scrambled control had higher CC3+ rates than unsorted cultures, potentially reflecting separation of RPCs and glia from neurons13,14. Nevertheless, Rb knockdown induced proliferation and apoptosis in cells with the same immunophenotypes as in unsorted cultures. Notably, Rb depletion induced the cell-cycle-related genes CCNE1, SKP2, E2F1, RBL1, CCNB1 and CDK1 in cone precursors, and induced p53-responsive genes in sorted RPCs and glia (Extended Data Fig. 3). Cell-cycle-related genes were induced several days before Ki67, suggesting that further reprogramming was needed for cell cycle entry.
RB1 shRNAs also induced cone precursor proliferation in intact retinas. Ki67 was detected in L/M-opsin+ cone precursors in the fovea, demarcated by cones but not rods, 15 days after transduction (Fig. 2d). Ki67 was not detected in cells expressing rod, amacrine, horizontal or ganglion cell markers (Extended Data Fig. 4a, d). Ki67 was detected in RPCs and glia marked by PAX6+, nestin+ or by CHX10+, CRX−, yet in similar proportions after transduction with RB1-directed and control shRNAs (Extended Data Fig. 4b–d). Moreover, a yellow fluorescent protein (YFP)-expressing RB1 shRNA vector selectively induced Ki67 in YFP+ cones, although all cell types were transduced (Extended Data Fig. 4e–h).
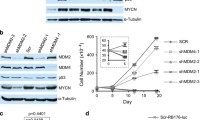
We next determined whether Rb-depleted cone precursors and retinoblastoma cells depend on similar signalling circuitry. Retinoblastoma cell proliferation requires several proteins that are prominent in cone precursors, including TRβ2, RXRγ, MYCN and MDM2 (ref. 7). Depletion of these factors suppressed Ki67 expression and cone precursor proliferation both in dissociated retinal cultures (Extended Data Fig. 5a, b) and in isolated populations (Fig. 3a). Retinoblastoma cell proliferation also requires SKP2-mediated degradation of Thr 187-phosphorylated p27 (ref. 15). Concordantly, SKP2 depletion suppressed cone precursor proliferation and increased CC3 (Fig. 3a and Extended Data Fig. 5a). Notably, maturing cone precursors had exceptionally high Thr 187-phosphorylated p27 (Extended Data Fig. 5c), coincident with a maturation-associated decrease in total p27 (ref. 16), suggesting that SKP2-mediated p27 degradation might enable cone precursor proliferation. Consistent with this view, cone precursor proliferation was suppressed by ectopic p27 and enhanced by ectopic SKP2 or p27 knockdown (Fig. 3b and Extended Data Fig. 5b), as in retinoblastoma cells15. Thus, Rb-depleted cone precursors and retinoblastoma cells had similar signalling requirements.
a, Prevalence of Ki67+ or CC3+ cells (top) and cell numbers (bottom) after shRNA transduction of isolated cone precursors. b, Isolated cone precursor response to co-transduction with RB1 shRNA and BE-Neo (BN) vector, BN-p130 or BN-SKP2. c–e, Effect of p130 overexpression or p107 knockdown on Y79 cell proliferation (c, d) and protein expression (e). *P < 0.05, **P < 0.01 (compared to scrambled and vector controls); †P < 0.05, ††P < 0.01 (compared to RB1 shRNA plus SCR (a), or to RB1 shRNA plus BN vector (b)). Results represent at least two independent experiments. Values and error bars denote mean and s.d. of triplicate assays; P values are from unpaired Student’s t-test.
We also assessed the roles of the Rb-related p107 and p130 proteins (also known as RBL1 and RBL2, respectively). In mouse models, retinal tumorigenesis required loss of Rb combined with loss of p107, p130 or p27 (refs 10, 17). However, in human retinoblastomas, p130 (also known as RBL2) losses are common, whereas p107 (also known as RBL1) losses are rare18 (Extended Data Fig. 6a). Moreover, whereas maturing cone precursors had abundant p130 and minimal p107, retinoblastomas had barely detectable p130 yet prominent p107 (Fig. 4b and Extended Data Fig. 6b), implicating p130 but not p107 in retinoblastoma suppression. Concordantly, co-knockdown of p130 with Rb increased cone precursor proliferation (Fig. 3a and Extended Data Fig. 5a, d) and p130 overexpression suppressed cone precursor and retinoblastoma cell proliferation (Fig. 3b, c, e). Meanwhile, p107 knockdown suppressed proliferation both in Rb-depleted cone precursors (Fig. 3a and Extended Data Fig. 5a, d) and in retinoblastoma cells (Fig. 3d, e and Extended Data Fig. 5e, g). In Y79 cells, p107 knockdown decreased expression of MYCN and SKP2, while it increased the SKP2 target, p27 (Fig. 3e). These effects were seen with two shRNAs and were rescued by p107 restoration (Fig. 3d, e and Extended Data Fig. 5d–i). Furthermore, p107 overexpression enhanced proliferation of retinoblastoma cells while suppressing that of neuroblastoma cells (Extended Data Fig. 5h–j). Thus, both in Rb-depleted cone precursors and in retinoblastoma cells, p130 suppressed proliferation whereas p107 had a proliferative role distinct from its function in mouse models.
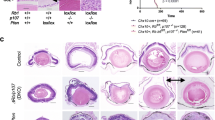
a, Haematoxylin-and-eosin-stained Rb-depleted and Rb/p130-depleted cone xenograft tumours and human retinoblastoma (n = 4). KD, knockdown. Dashed lines denote Flexner–Wintersteiner rosettes; solid lines denote fleurettes; asterisks mark rosette cavities. b, Cone- and cell-cycle-related protein expression in human retinoblastoma and cone xenografts (n = 6). Scale bars, 40 μm (a, b). c, Transmission electron microscopy of Flexner–Wintersteiner rosettes in a human retinoblastoma and a cone-derived tumour, with mitochondria (arrows) between nuclei and rosette cavity (n = 2). The ×25,000 images are from the boxed area (top) or from a rosette not shown (bottom). Results are representative of at least two experiments. d, Model of cone-precursor retinoblastoma origin highlighting proteins that suppressed (blue) or promoted (red) the proliferative response.
After several months, some cone precursor cultures depleted in Rb or in both Rb and p130 (Rb/p130-depleted) formed suspension aggregates resembling retinoblastoma cells (Extended Data Fig. 7a). Rb/p130-depleted cultures proliferated more robustly and longer than those with Rb depletion alone, consistent with p130 losses in many retinoblastoma cell lines (Extended Data Fig. 6a). The cultures had properties consistent with Rb/p130-depleted cone precursors (Extended Data Fig. 7b–h). When engrafted either 3 months or within 1 week after knockdown, Rb- or Rb/p130-depleted cone precursors formed retinoblastoma-like tumours in subretinal xenografts (Fig. 4a and Extended Data Figs 8 and 9). For cells engrafted within 1 week, tumours appeared within 6–14 months (Extended Data Fig. 8b), similar to the time needed to form tumours in children.
Cone-precursor-derived tumours had differentiated histology, little Rb or p130, many Ki67+ cells, and prominent p107 and SKP2, consistent with robust proliferation (Fig. 4a, b and Extended Data Fig. 9). They expressed the photoreceptor-related CRX, CD133 and IRBP and the cone-specific cone arrestin, L/M-opsin and RXRγ; all at levels similar to retinoblastomas and developing retinas (Fig. 4b and Extended Data Fig. 9). This is consistent with the many cone-specific proteins in retinoblastoma tumours7 (Supplementary Table 1). As in human retinoblastomas7, cone-precursor-derived tumour cells lacked numerous markers of other retinal cell types and had rare S-opsin and rhodopsin expression (Extended Data Fig. 10). Rb-depleted and Rb/p130-depleted cone precursor tumours also had structures resembling Flexner–Wintersteiner rosettes and fleurettes (Fig. 4a), which are retinoblastoma hallmarks19. Transmission electron microscopy confirmed the rosettes, with mitochondria positioned between the nuclei and rosette lumens (Fig. 4c). Dense core vesicles were not seen in two Rb-depleted cone precursor tumours nor in two retinoblastomas, consistent with the reported rarity of such structures20,21. Finally, single nucleotide polymorphism (SNP)-array analyses of two tumours revealed no megabase-size gains or losses, whereas quantitative PCR (qPCR) analyses revealed a partial RB1 loss but no other frequently reported changes (Extended Data Fig. 8c–e), consistent with the lack of DNA copy number alterations in some retinoblastomas22,23,24. Thus, cone precursor tumours resembled human retinoblastomas at the histological, ultrastructural, retinal marker and molecular cytogenetic levels.
This study examined collaboration between Rb loss and retinal cell-type-specific circuitries. We found that the circuitry of maturing L/M-cone precursors was uniquely conducive to proliferation and development of retinoblastoma-like tumours. Although we cannot exclude the possibility that Rb loss could induce a cone program and proliferation in other cell types, the robust responses of the most highly enriched cone precursor populations and of cells in an intact fovea suggest that cone precursors are the primary if not the sole responding cell type. Cone precursor features that collaborated with Rb loss included cone lineage factors (TRβ2 and RXRγ), highly expressed oncoproteins (MYCN and MDM2), and p27 downregulation probably mediated by SKP2. Some of these features may be interdependent, as RXRγ promoted MDM2 expression7, yet the larger program encompassing these features and its developmental purpose are unknown. Importantly, Rb-depleted cone precursor tumours had differentiated histology and lacked gross DNA aberrations, similar to putative early retinoblastoma elements25. These findings support a model in which Rb-deficient cone precursors form differentiated retinoblastomas, then dedifferentiate (Fig. 4d) and possibly acquire non-cone features8,22. Much of the circuitry implicated in cone precursor tumour initiation was also needed for retinoblastoma cell proliferation7,15, suggesting that tumour cells can be addicted to the cancer-predisposing circuitry of their originating cell types.
Methods
Retinoblastoma and retinal cell culture
Retinoblastoma cell lines Y79 and Weri-RB1 were obtained from the ATCC. RB177 was from an early passage culture and its identity confirmed by RB1 mutation sequencing. Retinoblastoma cells were confirmed free of mycoplasma and cultured in RB culture medium (IMDM, 10% FBS, 55 μM β-mercaptoethanol, with glutamine, penicillin, streptomycin, fungizone and 10 μg ml−1 plasmocin (Invivogen)7,26). Fetal eyes were obtained with informed consent from the Human Fetal Tissue Repository of the Albert Einstein College of Medicine and from Advanced Bioscience Resources under protocols approved by the Memorial Sloan-Kettering Cancer Center (MSKCC) Institutional Review Board, the Albert Einstein College of Medicine Institutional Review Board, and the Children’s Hospital Los Angeles Committee on Clinical Investigations. After transport in IMDM with 10% FBS on ice, eyes were rinsed in 70% ethanol for 3 s and washed in sterile PBS. Eyes were opened using a sterile scalpel and lens removed. Retinas were detached using forceps and incubated in papain solution (Worthington Tissue Dissociation Kit) for 10–30 min at 37 °C and 5% CO2, with pipette mixing every 5 min. After dissociation to ∼20-cell clusters, cells were diluted with 10 volumes of PBS and collected by centrifugation at 2,000 r.p.m. (Sorvall, Legend RT), re-washed in PBS (all centrifugations at 2,000 r.p.m. unless otherwise stated), suspended in RB culture medium as above, incubated at 37 °C with 5% CO2 overnight, and frozen in RB culture medium containing 10% dimethylsulphoxide (DMSO). Supernatant was transferred into a sterile container after each centrifugation and re-spun to prevent retinal cell losses. For lentivirus infections, cells were recovered from liquid nitrogen, cultured overnight, washed with PBS, suspended in 0.05% trypsin/EDTA (Cellgro) for 3–10 min with gentle pipetting, re-centrifuged, suspended in RB culture medium as above, and immediately infected. Cultures were maintained at high density, typically 50,000 cells per well (24-well dish) for unsorted cultures, with media changes every 3 days.
FACS
Approximately 10 million dissociated retinal cells (∼5 million per retina) were cultured for 18 h after thawing in RB culture medium, collected by centrifugation, washed with PBS, digested with 5 ml warm 0.05% trypsin/EDTA for 5–15 min while triturating in a 24-well culture plate 20–30 times per minute using a 1,000-μl tip and checking every ∼3 min, to produce 90–95% single cells, centrifuged as above (retaining the supernatant to prevent cell loss), suspended in 400 μl 5% FBS in PBS, and incubated at room temperature for 10 min. Cells (100 μl) were combined with 100 μl 4 μg ml−1 of mouse IgG (Sigma, I-8765), 300 μl of cells were combined with 300 μl of pre-mixed anti-CD133-phycoerythrin (PE) (Miltenyi Biotec, 130-080-801) at 1:6 and anti-CD44 fluorescein isothiocyanate (FITC) (clone IM7, Abcam ab19622 or BD Biosciences, BDB553133) at 1:25, to give 1:12 CD133 and 1:50 CD44 final dilutions. After 1 h at room temperature, cells were diluted with 900 μl 5% FBS in PBS, centrifuged as above, suspended in 500 μl 5% FBS/PBS with 300 ng ml−1 DAPI and held on ice until sorting. Cells were sorted using a Becton-Dickenson FACSAria SORP with 100 mW 488-nm laser, the triple bandpass filter removed in the FITC channel, FACSDiva v8.0 software, and selecting live single cells based on FSC width, SSC width, and DAPI exclusion. On FSC/SSC plots, cells were divided into small, medium, and large size groups and evaluated for CD133-PE and CD44-FITC. Eight populations collected into 500 μl complete medium as above were small, medium and large CD133hi, CD44−; small, medium, and large CD133lo, CD44+; small CD133−, CD44−; and ungated live single cells. Each population was cultured in 50% Y79-conditioned medium with fungizone (50% fresh RB culture medium combined with 50% filtered Y79-conditioned RB culture medium), and half of the volume changed with fresh 50% Y79-conditioned medium every 3 days. Sorted populations were characterized by adhering cells to poly-l-lysine-coated coverslips (1,000–2,000 cells each) for 3 h, fixing in 4% paraformaldehyde (PFA) for 5 min, washing in PBS four times, and storing at −20 °C until immunostaining. Lentivirus infection was performed within 24 h after sorting.
Lentiviral shRNA and cDNA expression constructs
pLKO lentiviral shRNA vectors from the TRC library (Open Biosystems/Thermo Scientific or MSKCC SKI High-Throughput Drug Screening and RNAi Core Facility)27 were designated by ‘sh’ followed by the name of the target gene and last 3–4 digits of the TRC or SKI identification numbers (Supplementary Table 2). shRNAs directed specifically against THRB variant 2 (also known as TRβ2) were designed using Invitrogen BLOCK-iT RNAi Designer (http://rnaidesigner.invitrogen.com/rnaiexpress/) and siDirect (http://sidirect2.rnai.jp/doc/) and cloned using the TRC cloning strategy (http://www.addgene.org/pgvec1?f=v&cmd=showfile&file=protocols) with deoxyoligonucleotides for DNA-directed RNAi (Integrated DNA Technologies). They are designated according to the position of the first shRNA target nucleotide after the translation initiation site (Supplementary Table 2). The pLKO scrambled control was Addgene plasmid 1864 (ref. 28). pLKO-YFP-shRB1-733, pLKO-YFP-shRB1-737 and pLKO-YFP-SCR control virus were produced by replacement of the puromycin resistance gene with YFP complementary DNA using In-Fusion cloning (Clontech), and provided by Z. Li. The lentiviral cDNA expression BN vector was created by replacing the EGFP gene of the BE-GFP lentiviral vector29 with the neomycin resistance gene between the EcoRI and BamHI sites (with assistance of S. Puranik). BN-p130 was produced by inserting human p130 cDNA between the BE-Neo PshAI and XbaI site. BN-SKP2 and BN-p107 were produced by inserting human SKP2 and p107 cDNA, respectively, between the BsiWI and PspXI sites of BE-Neo. Because shRBL1-2621 (shp107-1) targets the 3′ untranslated region, only the RBL1 open reading frame was cloned into BE-Neo to produce shRBL1-2621-resistant BN-p107. To produce shRBL1-2623 (shp107-2)-resistant BN-p107-2r, the shRBL1 target sequence GCAGTGAATAAGGAGTATGAA was mutated to GCAGTAAACAAAGAATATGAA without amino acid sequence changes using In-Fusion cloning (Clontech). BE-p27 was as described15.
Lentivirus production and infections
Lentiviruses were produced by reverse transfection of suspended 2 × 107 293T cells using 20 μg lentiviral vector, 10 μg pVSV-G, 20 μg pCMV-dR8.91 (ref. 30) and 100 μl Polyjet (SignaGen) or Lipofectamine 2000 (Life Technologies) in 15 cm3 dishes. The 3-ml plasmids–Polyjet complex and 1.5 ml 293T cell suspension were mixed in 50 ml centrifuge tubes and shaken for 30 min before being transferred to dishes. Virus collected 48 and 72 h after transfection was combined, concentrated 50–100-fold by centrifugation at 25,000 r.p.m. for 90 min, and suspended in RB culture medium. Concentrated virus (500–2,000 μl) was used to infect 5 × 105 Y79, Weri-1 or RB177 retinoblastoma cells, or to infect 5 × 105 total retinal cells or 1 × 105 of each sorted retinal cell population in 500 μl of filtered conditioned RB culture medium in the presence of 4 μg ml−1 polybrene (Sigma-Aldrich) followed by gentle pipetting 25 times and shaking for 10 min in the hood. After 18 h, cells were diluted in an equal volume of conditioned RB culture medium and maintained at 37 °C with 5% CO2. For co-infections, 100 μl of each concentrated virus was used to infect either 1 × 104 total retinal cells or 1 × 103 sorted retinal cells suspended in 100 μl of conditioned RB culture medium with 4 μg ml−1 polybrene in a total volume of 300 μl, and medium was replaced with 150 μl 50% Y79 and other RB cell conditioned medium 24 h after infection. Infected cells were selected starting 48 h after infection with 1.4–3 μg ml−1 puromycin for 48–72 h or with 50–100 μg ml−1 G418 for 4–7 days, and fed every 2–3 days by replacing two-thirds of the media with 50% Y79 and other RB cell-conditioned medium.
Intact FW19 retinas were either infected within the globe, by cutting a cross-section through the cornea, removing the lens and most of the vitreous, and pipetting 500 μl of concentrated pLKO versions of shRB1-733 and shRB1-737 or scrambled control lentivirus into the sub-retinal space and vitreous (causing retinal detachment) in a 24-well plate with the globe submerged in RB culture medium with 1 ml lentivirus suspension, followed after 2 days by addition of 2 ml of freshly prepared concentrated lentivirus; or infected after removal of the intact retina and residual vitreous in a 12-well plate, by addition of 1 ml of 80× concentrated pLKO-YFP-shRB1-733 or scrambled control lentivirus, reinfection with the same viruses 1 and 3 days later, and changing 50% of medium with a 1:1 mixture of fresh and ocular globe-conditioned medium daily thereafter. Displaced retinal tissue was fixed with 2% PFA in PBS for 2 h at 4 °C, and eyes with remaining tissue were fixed in 2% PFA in PBS overnight at 4 °C. Tissue samples were washed with PBS, transferred to 30% sucrose in PBS, and embedded in 30% sucrose/PBS:OCT at a 2:1 ratio, and cryosectioned at 8–10 μm.
Real-time quantitative PCR
Total RNA was isolated using StrataPrep total RNA microprep kit (Stratagene) for <1,000 cells (in FACS isolated populations) or GenElute Mammalian Total RNA Miniprep Kit (Sigma) for all other analyses. cDNA was synthesized using ImProm-II Reverse Transcription System (Promega). Primers were designed by Beacon Designer software (Premier Biosoft International) or Primer3 (http://frodo.wi.mit.edu/primer3/) (Supplementary Table 3). Relative messenger RNA levels were determined by qPCR using QuantiTect SYBR Green PCR Kit (Qiagen) or Maxima SYBR Green qPCR Master Mix (Fermentas) on an Applied Biosystems ABI 7900HT Sequence Detection System or ViiA 7 Real-Time PCR System using 95 °C 10 min followed by 40 cycles of 95 °C 20 s, 54 °C 30 s, 72 °C 30 s. Each sample was evaluated in triplicate and normalized to ACTB and GAPDH. Values represent the averages of both normalized results and error bars the standard deviation.
Immunostaining
Antibodies are described in Supplementary Table 4. Eyes were prepared and cryosectioned as described7,16. Cultured retinal cells were dissociated by gentle triturating, spread on poly-l-lysine-coated slides, incubated in a humidified incubator at 5% CO2 and 37 °C for 3 h, fixed in 4% PFA and PBS for 5 min, gently rinsed with PBS four times, vacuum-dried for 5 min, and stored at −20 °C.
The following co-staining combinations and orders were used to assess Ki67 expression in different retinal cell types. For cones: 1a. Mouse anti-cone arrestin31, anti-mouse-biotin, streptavidin-FITC, rabbit anti-Ki67, anti-rabbit-Cy3, rabbit anti-CRX and anti-rabbit-Cy5. 1b. Mouse anti-Ki67, anti-mouse-Cy3, rabbit anti-CRX, anti-rabbit-FITC, rabbit anti-human cone arrestin32 and anti-rabbit-Cy5. 2. Mouse anti-Ki67, anti-mouse-biotin, streptavidin-FITC, rabbit anti-CRX, anti-rabbit Cy3, rabbit anti-L/M-opsin and anti-rabbit-Cy5. 3. Mouse anti-RXRγ, anti-mouse-biotin, streptavidin-FITC, rabbit anti-Ki67, anti-rabbit-Cy3, rabbit anti-CRX and anti-rabbit-Cy5. For progenitors, Müller, and horizontal amacrine cells: mouse anti-Pax6, anti-mouse-biotin, streptavidin-FITC, rabbit anti-Ki67, anti-rabbit-Cy3, rabbit anti-nestin and anti-rabbit-Cy5. For other retinal cell types: mouse anti-human Ki67, anti-mouse-biotin, streptavidin-FITC, and rabbit antibodies for retinal specific markers, anti-rabbit Cy3. For BrdU labelling, 10 μM BrdU was added to medium for 2 h on day 23 after Rb knockdown and cells were stained with rat anti-BrdU, anti-rat-FITC, rabbit anti-CRX, anti-rabbit-Cy3, rabbit anti-L/M-opsin and anti-rabbit-Cy5.
For co-staining with mouse antibodies, sections or cells were treated with 1 mM EDTA/PBS for 5 min at room temperature and washed with PBS. Sections were treated with ABC kit reagent A (Vector Laboratories) in PBS for 15 min, washed in PBS, treated with ABC kit reagent B (Vector Laboratories) in PBS for 15 min, washed in PBS, blocked and permeabilized for 20 min in super block (2.5% horse serum, 2.5% donkey serum, 2.5% human serum, 1% BSA, 0.1% Triton-X-100 and 0.05% Tween-20 in PBS; filtered with 0.22 μm filter), incubated in mouse primary antibody in super block overnight at 4 °C, washed in PBS, incubated in biotinylated horse anti-mouse antibody in super block for 30 min, washed in PBS, incubated with FITC-conjugated streptavidin in PBS, and washed with PBS.
For co-staining with antibodies of other species, after completing the first staining reaction as above, sections were incubated in super block for 20 min, incubated overnight with primary antibody in super block, washed in PBS, incubated with Cy3- or Cy5-conjugated secondary antibody in super block for 30 min, and washed in PBS. Sections were then stained with 1 μg ml−1 DAPI in PBS, dried, mounted in Vectashield (Vector Labs) and imaged using an Axioplan2 (Carl Zeiss MicroImaging, LLC) or confocal DMIRE2 (Leica) microscope. Antibody specificity was confirmed by staining in parallel with control IgG or no primary antibody.
Antibody-dependent immunofluorescence signals were distinguished from autofluorescence by virtue of signal detected in only one colour channel. Cells with autofluorescence in multiple channels or with DNA condensation, fragmentation, or degradation were excluded. Cytoplasmic autofluorescence common in astrocytes, Müller glia, and ganglion cells was distinguished from authentic antigens by its detection at multiple wavelengths in cells with characteristic glial cytoplasmic and nuclear morphology. Nonspecific cytoplasmic staining by concentrated nestin antibody was distinguished from authentic nestin staining by its homogeneous rather than fibre-like structure. Nonspecific nuclear staining of L/M-opsin was avoided by using reduced antibody concentration.
EdU labelling and detection
Click-iT EdU Alexa Fluor 488 Imaging Kit was used for EdU labelling to detect proliferation. Dissociated or sorted retinal cells were infected with shRB1 or control lentivirus. After 14 days, 20 μg ml−1 of EdU was added into medium and incubated for 1 h; the cells were attached on poly-l-lysine-coated coverslips for 2 h and fixed for 5 min.
The cells were blocked and permeabilized for 20 min in super block as above and EdU was detected by addition of Click-iT reaction cocktails containing 2 μM Alexa Fluor 488 azide for 1 h. Co-staining was performed after EdU labelling with different combination of antibodies for retinal cell markers and secondary antibodies conjugated with Cy3 or Cy5, described as above. For cones, cone arrestin-Cy3 plus CRX-Cy5, RXRγ-Cy3 plus CRX-Cy5, and L/M-opsin-Cy3 plus CRX-Cy5 were used for co-staining with EdU.
Immunoblotting
Cells were washed in PBS, lysed in ELB+ (150 mM NaCl, 50 mM HEPES, pH 7.4, 0.1% NP40, 5 mM EDTA, 2 mM dithiothreitol (DTT), 1 mM phenylmethylsulfonyl fluoride, 10 mM NaF, 1 mM sodium orthovanadate, Thermo Scientific Halt phosphatase inhibitor cocktail and protease inhibitor cocktails), separated on 4–20% Ready Gel polyacrylamide gels (Jule Biotechnologies INC) or 8% polyacrylamide (for Rb western), and transferred to Hybond-ECL nitrocellulose membrane (Amersham). Membranes were probed with antibodies (Supplementary Table 4) and developed using horseradish peroxidase-conjugated anti-mouse or anti-rabbit antibodies and the ECL Advance Western Blotting Detection Kit (Amersham Biosciences) or Thermo Scientific SuperSignal-West Femto Chemiluminescent Substrate, and HyBlot CL X-Ray film (Denville Scientific).
Xenografts
All animal experiments complied with ethical regulations and were approved by the MSKCC Institutional Animal Care and Use Committee. Xenografts were performed on 6-week-old male athymic (Foxn1−/−) mice (Taconic) or 6-week-old male non-obese diabetic–severe combined immunodeficient (NOD–SCID) Il2rg−/− mice (Jackson Laboratories). Cultured cells were dissociated by pipetting, suspended in RB growth medium at 5 × 104 cells per microlitre (day 90) or 2.5 × 103 cells per microlitre (days 3 or 7), held on ice, and 2 μl injected into the subretinal space as described7. Irradiated 5053 rodent diet with amoxicillin was provided from 2 days before to 2 weeks after injection to prevent infection. Some tumour-bearing eyes were fixed and embedded as described7,16.
DNA copy number analyses
Genomic DNA from retinoblastomas, cone-derived retinoblastoma-like cells, and cone-derived xenograft tumours were isolated with QiaAMP DNA Mini kit (Qiagen). Genomic DNA of cone-derived cells was digested with XhoI to separate pLKO DNA hairpin structures. Relative DNA levels were determined in triplicate by qPCR using QuantiTect SYBR Green PCR Kit (Qiagen) on an Applied Biosystems ABI 7900HT Sequence Detection System or ViiA 7 Real-Time PCR System, using primers listed in Supplementary Table 5 and normalizing to the average of the HNF4A and BRCA1 genes. Integrated pLKO-shRB1-733, shRB-737 and shRBL2-923 copy numbers were analysed using primers corresponding to the pLKO.1 U6 promoter and RB1- or RBL2-specific shRNA sequences. High resolution SNP-array DNA copy number analyses were performed using CytoScan HD (Affymetrix, 901835). Data were analysed using Chromosome Analysis Suite 2.0 (Affymetrix).
Statistical analyses
Measurements were performed in triplicate and differences between means assessed for significance using unpaired Student’s t-tests. Sample sizes were chosen based on the maximum cell numbers that could be used for individual experiments given sample availability.
Transmission electron microscopy
Human retinoblastomas and cone-derived xenograft tumours were fixed with 4% PFA in PBS, rinsed in 0.1 M sodium cacodylate buffer, post-fixed in 2% osmium tetroxide for 1 h, rinsed in distilled water, dehydrated in a graded series of 50%, 75%, 95% and 100% ethanol, followed by two 10-min incubations in propylene oxide and overnight incubation in 1:1 propylene oxide/Poly Bed 812. The samples were embedded in Poly Bed 812 and cured at 60 °C. Ultra-thin sections were obtained with a Reichert Ultracut S microtome. Sections were stained with Uranyl Acetate and Lead Citrate and photographed using a Jeol 1200EX transmission electron microscope.
References
Weinberg, R. A. The retinoblastoma protein and cell cycle control. Cell 81, 323–330 (1995)
Cobrinik, D. Pocket proteins and cell cycle control. Oncogene 24, 2796–2809 (2005)
Gordon, G. M. & Du, W. Conserved RB functions in development and tumor suppression. Protein Cell 2, 864–878 (2011)
Viatour, P. & Sage, J. Newly identified aspects of tumor suppression by RB. Dis. Model. Mech. 4, 581–585 (2011)
Manning, A. L. & Dyson, N. J. R. B. mitotic implications of a tumour suppressor. Nature Rev. Cancer 12, 220–226 (2012)
Kleinerman, R. A. et al. Risk of new cancers after radiotherapy in long-term survivors of retinoblastoma: an extended follow-up. J. Clin. Oncol. 23, 2272–2279 (2005)
Xu, X. L. et al. Retinoblastoma has properties of a cone precursor tumor and depends upon cone-specific MDM2 signaling. Cell 137, 1018–1031 (2009)
McEvoy, J. et al. Coexpression of normally incompatible developmental pathways in retinoblastoma genesis. Cancer Cell 20, 260–275 (2011)
Gombos, D. S. Retinoblastoma in the perinatal and neonatal child. Semin. Fetal Neonatal Med. 17, 239–242 (2012)
Cobrinik, D. in Animal Models of Brain Tumors (eds Martinez-Murillo, R. & Martinez, A. ) 141–152 (Springer, 2013)
Lakowski, J. et al. Effective transplantation of photoreceptor precursor cells selected via cell surface antigen expression. Stem Cells 29, 1391–1404 (2011)
Shinoe, T. et al. Identification of CD44 as a cell surface marker for Muller glia precursor cells. J. Neurochem. 115, 1633–1642 (2010)
Hauck, S. M. et al. Identification of paracrine neuroprotective candidate proteins by a functional assay-driven proteomics approach. Mol. Cell. Proteomics 7, 1349–1361 (2008)
Xu, X. L. et al. Tumor-associated retinal astrocytes promote retinoblastoma cell proliferation through production of IGFBP-5. Am. J. Pathol. 177, 424–435 (2010)
Wang, H. et al. Skp2 is required for survival of aberrantly proliferating Rb1-deficient cells and for tumorigenesis in Rb1+/− mice. Nature Genet. 42, 83–88 (2010)
Lee, T. C., Almeida, D., Claros, N., Abramson, D. H. & Cobrinik, D. Cell cycle-specific and cell type-specific expression of Rb in the developing human retina. Invest. Ophthalmol. Vis. Sci. 47, 5590–5598 (2006)
Sangwan, M. et al. Established and new mouse models reveal E2f1 and Cdk2 dependency of retinoblastoma, and expose effective strategies to block tumor initiation. Oncogene 31, 5019–5028 (2012)
Priya, K., Jada, S. R., Quah, B. L., Quah, T. C. & Lai, P. S. High incidence of allelic loss at 16q12.2 region spanning RBL2/p130 gene in retinoblastoma. Cancer Biol. Ther. 8, 714–717 (2009)
Ts’o, M. O., Zimmerman, L. E. & Fine, B. S. The nature of retinoblastoma. I. Photoreceptor differentiation: a clinical and histopathologic study. Am. J. Ophthalmol. 69, 339–349 (1970)
Albert, D. M., Lahav, M., Lesser, R. & Craft, J. Recent observations regarding retinoblastoma. I. Ultrastructure, tissue culture growth, incidence, and animal models. Trans. Ophthalmol. Soc. U. K. 94, 909–928 (1974)
Popoff, N. A. & Ellsworth, R. M. The fine structure of retinoblastoma. In vivo and in vitro observations. Lab. Invest. 25, 389–402 (1971)
Kapatai, G. et al. Gene expression profiling identifies different sub-types of retinoblastoma. Br. J. Cancer 109, 512–525 (2013)
Corson, T. W. & Gallie, B. L. One hit, two hits, three hits, more? Genomic changes in the development of retinoblastoma. Genes Chromosom. Cancer 46, 617–634 (2007)
Zhang, J. et al. A novel retinoblastoma therapy from genomic and epigenetic analyses. Nature 481, 329–334 (2012)
Dimaras, H. et al. Loss of RB1 induces non-proliferative retinoma; increasing genomic instability correlates with progression to retinoblastoma. Hum. Mol. Genet. 17, 1363–1372 (2008)
DiCiommo, D. P., Duckett, A., Burcescu, I., Bremner, R. & Gallie, B. L. Retinoblastoma protein purification and transduction of retina and retinoblastoma cells using improved alphavirus vectors. Invest. Ophthalmol. Vis. Sci. 45, 3320–3329 (2004)
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006)
Sarbassov, D. D., Guertin, D. A., Ali, S. M. & Sabatini, D. M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005)
Cobrinik, D., Francis, R. O., Abramson, D. H. & Lee, T. C. Rb induces a proliferative arrest and curtails Brn-2 expression in retinoblastoma cells. Mol. Cancer 5, 72 (2006)
Zufferey, R., Nagy, D., Mandel, R. J., Naldini, L. & Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nature Biotechnol. 15, 871–875 (1997)
Wikler, K. C., Rakic, P., Bhattacharyya, N. & Macleish, P. R. Early emergence of photoreceptor mosaicism in the primate retina revealed by a novel cone-specific monoclonal antibody. J. Comp. Neurol. 377, 500–508 (1997)
Li, A., Zhu, X. & Craft, C. M. Retinoic acid upregulates cone arrestin expression in retinoblastoma cells through a Cis element in the distal promoter region. Invest. Ophthalmol. Vis. Sci. 43, 1375–1383 (2002)
Acknowledgements
We thank P. MacLeish, D. Forrest, C. Craft, G. Chader, C. Gregory-Evans, R. Molday, P. Hargrave, Y. Imanishi, K. Palczewsk, E. Weiss, A. Swaroop, T. Li, R. Lee and J. Saari for antibodies. We thank T. Baumgartner and P. Byrne for FACS assistance, N. Lampen for electron microscopy assistance, N. Zhou, T. Patel and J. Wang for technical assistance, S. Puranik and Z. Li for DNA constructs, and J. Aparicio for critical reading of the manuscript. Funding was received from The Gerber Foundation (X.L.X.), The Fund for Ophthalmic Knowledge (D.H.A.), the Research and Development Funds of the MSKCC Department of Pathology (S.C.J.), The Larry & Celia Moh Foundation (D.C.), and National Institutes of Health grant 1R01CA137124 (D.C.).
Author information
Authors and Affiliations
Contributions
X.L.X., S.C.J. and D.C. designed the study. X.L.X. conducted most of the experiments in S.C.J.’s laboratory, supported in part by D.C. H.P.S. and D.-L.Q. quantified Rb knockdown and confirmed effects at different time points. H.P.S. transduced retina with YFP-labelled constructs, and analysed them with X.L.X. L.W. analysed SNP arrays. D.H.A. provided retinoblastoma samples. B.K.P. provided fetal retina. D.C. wrote the manuscript with assistance from X.L.X. and review by S.C.J.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Proliferation of cone-like cells after Rb depletion in dissociated FW19 retina.
a, Decreased Rb protein in L/M-opsin+ or TRβ2+ cells (arrows) on days 5 or 23, and decreased RB1 RNA or Rb protein on day 4 after shRB1-733 transduction. b, Cone arrestin+, CRX+ cells (arrows) with or without Ki67 co-expression. c, Ki67+ and cone arrestin+ cells first detected 9 or 14 days after transduction in two experiments. d–f, Co-staining of Ki67 with RXRγ and CRX at 14 days (d), with cone arrestin and CRX at 14 days (e), or with L/M-opsin and CRX at 23 days (f) after transduction with shRB1-733 or a scrambled control. g, Percentage of cells co-expressing Ki67 with L/M-opsin and CRX, RXRγ and CRX, or cone arrestin and CRX, 23 days after transduction. h, Prevalence of cells co-staining for L/M-opsin and CRX, RXRγ and CRX, or cone arrestin and CRX, 23 days after transduction. i, Ki67 not detected in cells expressing markers of rods (NRL), ganglion cells (BRN-3), bipolar cells (strong CHX10), or horizontal cells (PROX1) 14 days after transduction. j, Co-expression of Ki67 with markers of RPCs (nestin, white arrows) or Müller glia (CRALBP or SOX2), but not in PAX6+, nestin− ganglion, amacrine or horizontal cells (yellow arrows) 14 days after transduction. k, l, EdU incorporation in cells expressing markers of cones (cone arrestin and CRX or RXRγ and CRX, yellow arrows in l) but not in cells expressing markers of rods (CNGA1, CNGB1), bipolar cells (CHX10, CRX), or ganglion, horizontal or amacrine cells (syntaxin) (white arrows in l) 14 days after transduction. Black lines above labels demarcate distinct fields. m, Co-staining of phosphohistone H3 (PH3) with cone arrestin and CRX 23 days after transduction. n, Apoptosis marker CC3 in cells expressing RPC and glial marker nestin 14 days after transduction with RB1-directed shRNAs (yellow arrow) but not with scrambled control (white arrow). Values and error bars are mean and s.d. of triplicate assays. Scale bars, 20 μm. Data are representative of at least two independent experiments.
Extended Data Figure 2 FACS isolation of retinal cell populations.
Retinal cells were isolated according to size, CD133 and CD44 staining. In study 1, cell type compositions in each fraction (a) were determined by immunostaining with cone arrestin and CRX (b, e), NRL (c), and nestin and PAX6 (d, f). In study 2, cell type compositions (i) were determined by immunostaining with RXRγ and CRX, nestin and CHX10, nestin and PAX6, and CRALBP (j, k). The percentages of the predominant cell types in each population (a, i) and marker specificities (g) are indicated. h, Cone-specific co-staining of cone arrestin and GNAT2 (top) and cone-specific co-staining of RXRγ and CRX (bottom) in FW19 retina. GCL, ganglion cell layer; INL, inner nuclear layer; ONL, outer nuclear layer. l, Co-staining of cells for EdU with cone arrestin and CRX or with RXRγ and CRX 14 days after transduction of the cone-enriched medium plus large CD133hi CD44− population isolated as in i–k with two RB1 shRNAs (yellow arrows) but not with the scrambled control (white arrows). In both studies, CD133hi CD44− medium and large size populations mainly consisted of cells expressing cone markers (CRX and cone arrestin, or CRX and RXRγ). The CD133hi CD44− small population mainly consisted of cells expressing a rod marker (NRL) with a variable proportion expressing cone markers. All CD133lo CD44+ populations mainly consisted of cells co-expressing RPC and glial markers (nestin and PAX6, nestin and CHX10, or CRALBP). The CD133lo CD44− small size population consisted of cells with diverse immunophenotypes. Values and error bars are mean and s.d. of triplicate assays. Scale bars, 30 μm.
Extended Data Figure 3 Cone precursor gene expression response to Rb depletion.
a–c, Fold change in RNA level relative to day 0 uninfected cells for RB1 (a), or the indicated E2F-responsive genes (b), or the indicated p53-regulated genes (c), 3 and 6 days after transduction of each population with a mixture of shRB1-733 and shRB1-737, or with scrambled control. *P < 0.05, **P < 0.01 (comparing shRB1 and scrambled control). Data are representative of two sets of qPCR analyses. Values and error bars are mean and s.d. of triplicate assays.
Extended Data Figure 4 Proliferation status of retinal cells other than cones 15 days after shRB1 transduction of intact FW19 retinas.
a–d, Combined transduction with pLKO-shRB1-733 and -737. a, Ki67 not detected in NRL+ or rhodopsin+ rod photoreceptors or in calbindin+ horizontal cells. b, Ki67 detected in PAX6lo, nestin+ RPCs (white arrows) but not in PAX6hi, nestin− horizontal, amacrine or ganglion cells (yellow arrows). c, Ki67 detected in CHX10+, CRX− RPCs (white arrows) but not in CHX10+, CRX+ bipolar cells (yellow arrows). d, Percentage of cells co-expressing Ki67 and retinal cell markers. e–h, Transduction with YFP-marked pLKO-YFP-shRB1-733. e, Ki67 detected in YFP+, L/M-opsin+ or YFP+, cone arrestin+ cone precursors (white arrows) and in an undefined YFP− cell (yellow arrow). f, Ki67 not detected in YFP+, calbindin+ horizontal cells, YFP+, syntaxin+ or YFP+, PAX6+ amacrine cells, or in YFP+, NRL+ rod precursors. g, Ki67 detected (white arrows) or not detected (yellow arrows) in YFP+, nestin+ RPCs or glia, or in YFP+, CHX10+ RPCs or bipolar cells. h, Proportion of Ki67+ cells co-expressing YFP and retinal markers after transduction with pLKO-YFP-shRB1-733 or scrambled control. Values and error bars are mean and s.d. of triplicate assays. Scale bars, 20 μm. Analyses in a–d and in e–h represent two independent experiments. All immunostaining was performed at least twice.
Extended Data Figure 5 Effect of cone- and Rb-related circuitry on cone precursor response to Rb depletion.
A, Percentage of Ki67+ cells among L/M-opsin+, CRX+ cells (a), among RXRγ+, CRX+ cells (b), or among cone arrestin+, CRX+cells (c); and percentage of L/M-opsin+, CRX+ cells among all cells with DAPI+ nuclei (d) after transduction of dissociated FW18 retina with shRB1-733 and shRNAs against p130, p107, TRβ2, SKP2, MDM2 and MYCN. B, Percentage of Ki67+ cells among L/M-opsin+, CRX+ cone-like cells (top) and proliferative response (bottom) after transduction of dissociated FW18 retina with shRB1-733 and with shRNAs against RXRγ and p27 (shRNAs 856+930), or with overexpression of p27 and p27-T187A. C, High-level Thr 187 phosphorylated p27 (p-p27(T187), top) coinciding with downregulation of total p27 (bottom) and prominent Rb during cone precursor maturation. C, a, Perifoveal region of FW18 retina. C, b, Enlarged view of boxed regions in C, a. Arrows, cone precursors identified by large, strongly Rb+ nuclei and lack of p27 signal in characteristic outer nuclear layer position7,16. D, Effect of two RBL1-p107 or two RBL2-p130 shRNAs on proliferation of Rb-depleted isolated cone precursors. E, Knockdown efficacy of two RBL1-p107 or two RBL2-p130 shRNAs in Y79 and RB177 retinoblastoma cells. F, Impaired proliferation of Weri-RB1 retinoblastoma cells after transduction with BN-p130 compared to vector control. G, Impaired proliferation of RB177 retinoblastoma cells following transduction with two p107 shRNAs. H, I, Impaired proliferation and MYCN expression in Y79 cells after p107 knockdown with two p107-directed shRNAs, and rescue by shRNA-resistant BN-p107 constructs. J, p27 accumulation and growth suppression following p107 knockdown with shp107-2 rescued by BN-p107-2r in RB1 wild type SKN-BE(2) neuroblastoma cells. p107 overexpression impaired SKN-BE(2) growth, contrary to its effects in Y79. *P < 0.05, **P < 0.01 (compared to SCR or vector control); †P < 0.05, ††P < 0.01 (compared to RB1-KD plus SCR or RB1-KD plus BN vector); ‡P < 0.05, ‡‡P < 0.01 (compared to shp107-2 plus BN vector) (H–J). Data are representative of more than two independent experiments except for SKN-BE(2) analyses. Values and error bars are mean and s.d. of triplicate assays.
Extended Data Figure 6 p130 copy number in retinoblastomas and cone precursor expression.
a, DNA copy number of p130, other 16q genes implicated in retinoblastoma (CDH11, CDH13), and p107 determined by qPCR (n = 6). The percentage of retinoblastomas with copy number (CN) < 1.5 was higher for p130 than for other 16q genes (summarized at right; P values relative to p130 using Fisher’s exact test). b, p130 in peripheral, lateral and central FW19 retina. Boxed region in maturing central retina (top) and enlarged view (bottom) show prominent p130 in weakly DAPI-stained cone precursor nuclei (arrows). Scale bars, 40 μm. Data are representative of at least two independent experiments. Values and error bars are mean and s.d. of triplicate assays.
Extended Data Figure 7 Characterization of Rb/p130-depleted retinoblastoma-like cells.
a, Similar appearance of Rb/p130-depleted cones and early passage retinoblastoma cells. Scale bar, 40 μm. b, c, DNA copy number of shRNA vectors (b) or selected genes (c) in cell lines derived from Rb/p130-depleted cone precursors (Cone1, Cone2, Cone5) or from Rb/p130-depleted unsorted retinal cells (All3, All4), in Rb-depleted unsorted retinal cells 4 days after transduction (All-RB1-KD-d4), in Y79 cells, or in FW21 retina (normal) (n = 6). All cell lines retained RB1 and p130 shRNA vectors and lacked RB1 or p130 copy number alterations. The Y79 MYCN copy number (∼78) is not shown (asterisk). d–g, qPCR gene expression analyses in the indicated cell lines relative to cones transduced with scrambled control or FW21 retina (n = 6). d, All cell lines had diminished RB1 and p130 expression. e–g, Altered expression of cell-cycle-related (e), cone-related (f) and apoptosis-related (g) genes. h, SNP-array analysis of two Rb/p130-depleted cone precursor cell lines (1, 2), revealing no megabase-size loss of heterozygosity (LOH) or copy number alterations (CNA). Data are representative of at least two analyses (b–g) or analyses of two cell lines (h). Values and error bars are mean and s.d. of triplicate assays.
Extended Data Figure 8 Characterization of Rb- and Rb/p130-depleted cone precursor tumours.
a, Intraocular tumour 4 months after Rb-depleted cone precursor xenograft. b, Summary of subretinal xenograft groups 1, 2 and 3. Sample size was as needed to assess tumour phenotypes. Mice were randomly assigned to different xenograft regimens and the investigator blinded to the assignment until the tumour analyses. Two mice with early death were excluded from the analyses. c, SNP-array analysis of one Rb/p130-depleted (tumour 1) or one Rb-depleted (tumour 2) cone-precursor-derived tumours from xenograft group 3, revealing no megabase-size loss of heterozygosity or copy number alterations . d, qPCR analysis of pLKO shRNA vector copy number in tumours derived from Rb/p130-depleted cone precursors (m-Cone1, m-Cone2) or from Rb/p130-depleted unsorted retinal cells (m-All3, m-All4), or in mouse ocular tissue (m-Cone-SCR), Y79 cells, or FW19 retina (normal). All tumours retained RB1 and/or p130 shRNA vector sequences, confirming their engineered cone precursor origin. e, qPCR analysis of MDM2, MDM4, RB1 and MYCN copy number in three cone-derived tumours and normal retina (n = 6). DNA copy number data (d, e) are representative of two analyses. Values and error bars are mean and s.d. of triplicate assays.
Extended Data Figure 9 Cone and cell-cycle-related proteins in Rb- or Rb/p130-depleted cone precursor tumours engrafted 3 days after transduction.
Most tumour cells expressed human nuclear antigen (HuNU), confirming their xenograft origin. They also expressed cone-related proteins (CRX, cone arrestin, L/M-opsin, RXRγ, CD133 and IRBP) and proliferation-related proteins (Ki67, SKP2, p107 and cytoplasmic p27) but lacked Rb. Tumours had elements resembling Flexner–Wintersteiner rosettes (asterisks) and fleurettes (daggers). Scale bars, 40 μm. Data are representative of three independent experiments.
Extended Data Figure 10 Analysis of non-cone cell markers in cone-precursor-derived tumours and retinoblastomas.
a, Proteins detected in normal retina but not in cone-derived tumour or human retinoblastoma cells included markers of rods (rhodopsin and CNGB1), RPCs and Müller glia (nestin, GFAP and PAX6), bipolar cells (CHX10), ganglion, amacrine and horizontal cells (calbindin and PAX6), and ganglion cells (nuclear BRN-3, thin arrows in mouse retina). PAX6+, nestin+ cells detected in human retinoblastoma were previously found to be Rb+ non-tumour cells from tumour-associated retina7. An uncharacterized cytoplasmic BRN-3 signal (bold arrows) was detected in mouse photoreceptor outer segments and in cone-derived tumour and retinoblastoma rosettes. b, L/M-opsin was detected in most cone-derived tumour cells. However, rare cells co-expressed S-opsin and L/M-opsin (arrows), as in immature L/M-cone precursors and human retinoblastomas7. c, One tumour had rare rhodopsin+, Ki67− cells but no detected rhodopsin+, Ki67+ cells, as in a previously characterized retinoma-like regions7. Scale bars, 40 μm. Data are representative of three independent xenograft experiments.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1-5. (PDF 385 kb)
Rights and permissions
About this article
Cite this article
Xu, X., Singh, H., Wang, L. et al. Rb suppresses human cone-precursor-derived retinoblastoma tumours. Nature 514, 385–388 (2014). https://doi.org/10.1038/nature13813
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13813
- Springer Nature Limited
This article is cited by
-
Inhibition of high level E2F in a RB1 proficient MYCN overexpressing chicken retinoblastoma model normalizes neoplastic behaviour
Cellular Oncology (2024)
-
Sunitinib efficacy with minimal toxicity in patient-derived retinoblastoma organoids
Journal of Experimental & Clinical Cancer Research (2023)
-
E2F1/CKS2/PTEN signaling axis regulates malignant phenotypes in pediatric retinoblastoma
Cell Death & Disease (2022)
-
Co-activation of Sonic hedgehog and Wnt signaling in murine retinal precursor cells drives ocular lesions with features of intraocular medulloepithelioma
Oncogenesis (2021)
-
Retinoblastoma tumor cell proliferation is negatively associated with an immune gene expression signature and increased immune cells
Laboratory Investigation (2021)