Abstract
For the majority of patients with advanced non-small cell lung cancer (NSCLC), the standard of care remains platinum-based chemotherapy. However, cisplatin resistance is a big obstacle to the treatment, and elucidation of its mechanism is warranted. In this study, we showed that there was no difference in intracellular uptake of cisplatin or the removal of platinum-DNA adducts between a cisplatin-resistant NSCLC cell line (A549/DR) and a cisplatin-sensitive NSCLC cell line (A549). However, the capacity to repair DNA interstrand crosslinks (ICLs) and double-strand breaks (DSBs) was significantly enhanced in the A549/DR cell line compared to 3 cisplatin-sensitive cell lines. We found that the protein and mRNA expression levels of Pol η, a Y-family translesion synthesis (TLS) polymerase, were markedly increased upon cisplatin exposure in A549/DR cells compared with A549 cells. Furthermore, intracellular co-localization of Pol η and proliferation cell nuclear antigen (PCNA) induced by cisplatin or cisplatin plus gemcitabine treatment was inhibited by depleting ataxia telangiectasia mutated and Rad-3-related (ATR). Pol η depletion by siRNA sensitized A549/DR cells to cisplatin; co-depletion of Pol η and ATR further increased A549/DR cell death induced by cisplatin or cisplatin plus gemcitabine compared to depletion of Pol η or ATR alone, concomitant with inhibition of DNA ICL and DSB repair and accumulation of DNA damage. No additional sensitization effect of co-depleting Pol η and ATR was observed in A549 cells. These results demonstrate that co-inhibition of Pol η and ATR reverses the drug resistance of cisplatin-resistant NSCLC cells by blocking the repair of DNA ICLs and DSBs induced by cisplatin or cisplatin plus gemcitabine.
Similar content being viewed by others
Introduction
Platinum drugs such as cisplatin and carboplatin are the mainstay of lung cancer chemotherapy. Although the appearance of “targeted” drugs such as erlotinib and crizotinib have led to improvement in advanced non-small cell lung cancer (NSCLC) therapy for a small population of patients1,2, the majority of patients are not candidates for treatment with targeted drugs, and for these patients, the standard of care remains platinum-based chemotherapy3. The major mechanism of action of platinum drugs is induction of the formation of crosslinked DNA adducts to block DNA replication4. A major drawback in the use of platinum, however, is the acquisition of drug resistance during the courses of therapy5. This resistance can be mediated by the DNA damage response (DDR) and DNA damage tolerance (DDT) pathways, including translesion synthesis (TLS)6,7. TLS is a mechanism naturally used by cells to prevent common DNA damage from stalling replication forks and giving rise to high levels of apoptosis, and thus, TLS is believed to contribute to the development of platinum resistance8,9,10,11,12,13,14. TLS can be either error-free or error-prone, depending on the specific lesion being bypassed and the TLS polymerases involved in inserting nucleotides opposite the lesion. For instance, the Y-family TLS polymerase η (Pol η, encoded by POLH) can bypass a cyclobutane pyrimidine dimer (CPD) induced by ultraviolet (UV) light with high fidelity8. TLS Pol η is also capable of bypassing the DNA intrastrand crosslink formed by cisplatin, accommodating it in a manner similar to the CPD15. Among the many TLS polymerases tested in vitro, Pol η is the most efficient and accurate in bypassing the platinum-GG adduct (Pt-GG adduct)15,16,17,18. Moreover, some studies showed that Pol η can also bypass the DNA interstrand crosslinks (ICLs) induced by platinum, mitomycin and psoralen in an error-prone fashion in vitro and that Pol η is involved in the repair of these drug-induced ICL lesions19,20,21,22, although other investigators reported that Pol η is dispensable for the processing of cisplatin-induced ICLs in vivo 23. In addition, Pol η can efficiently extend from gemcitabine at the 3′-termini of DNA10 and replicate across gemcitabine dFdC sites in the template DNA that was shown to block DNA polymerases24, which are thought to be associated with resistance to gemcitabine.
In addition to triggering the DNA repair pathway, the stalled replication forks produced by crosslinking agents activate the ataxia telangiectasia mutated and Rad-3-related (ATR) signaling pathway. Activated ATR phosphorylates multiple substrates, including Chk1, which help cells survive replication stress by inhibiting origin firing, inducing the checkpoint and regulating cell cycle arrest and DNA damage repair25. In the absence of ATR, stalled replication forks collapse into double-strand breaks (DSBs), which can lead to genomic rearrangements or cell death26,27. The same signal that induces the ATR checkpoint also activates the recruitment of TLS polymerases through monoubiquitination of proliferation cell nuclear antigen (PCNA)28. Although there is evidence that ATR inhibition can potentiate cisplatin cytotoxicity to cancer cells29 and that Pol η deficiency sensitizes human cells to cisplatin and gemcitabine10, whether the co-inhibition of ATR and Pol η can hypersensitize cisplatin-resistant NSCLC cells to cisplatin remains unclear. Here, we show that knockdown of Pol η synergizes with ATR inhibition to further sensitize A549/DR cells, a cisplatin-resistant NSCLS cell line, to cisplatin through the suppression of ICL and DSB repair compared with Pol η knockdown alone. This evidence indicates that the co-inhibition of Pol η and ATR can be used to improve the efficacy of NSCLC chemotherapy by reversing cisplatin resistance in drug-resistant NSCLC. Because we found that the A549/DR cell line was also resistant to gemcitabine compared to cisplatin-sensitive cell lines in initial experiments and because some studies showed that Pol η participates in gemcitabine resistance10,24, we evaluated the changes in the cytotoxicity of both cisplatin and gemcitabine to A549/DR cells, as well as the DNA damage repair response induced by cisplatin and gemcitabine after the co-inhibition of Pol η and ATR.
Materials and methods
Cell culture and materials
The NSCLC-derived cell lines A549, A549/DR, LOU-NH91 and HCC4006 were purchased from the Shanghai Institute for Biological Sciences (China). All cell lines were cultured in RPMI-1640 medium supplemented with 10% heat-inactivated fetal calf serum (FCS), L-glutamine, and 5% antibiotics (penicillin/streptomycin). A549/DR cells were routinely maintained in culture medium containing 0.5 μg/mL cisplatin and grown in drug-free medium for seven days before the experiment.
The antibodies used in this study targeted antigens including the following: FANCM, FANCJ, RAD18, RAD51, ATR, PNCA, cleaved caspase-3 and PARP from Santa Cruz; Pol η, Pol κ, Pol β, Pol μ, and Pol ν from Abcam; γ-H2AX and H2AX from Cell Signaling Technology; and p-Chk1, p-KAP1, and p-RPA2 from Calbiochem. The drugs used in this study included cisplatin from Yangtze River Pharmaceutical Group (China), carboplatin from Qilu Pharmaceutical Co, oxaliplatin from Hengrui Medicine Co, Ltd (China), gemcitabine from Eli Lilly, and VE-822 from YuduoBio.
Detection of cell viability and colony formation
Cell viability was detected by the cell counting kit-8 (CCK-8) assay according to the manufacturer's instructions, as previously described30. For the colony formation assay, cells were seeded at a density of 500 cells per well onto a 6-well culture plate in DMEM containing 10% FCS and treated with various drugs. After two weeks, the cells were fixed with 4% paraformaldehyde for 10 min and then stained with 0.05% crystal violet in ddH2O for 15 min. The colonies produced by each cell group were counted and measured using Image software.
Measurement of cisplatin-DNA adducts
Cells were plated at 6×105 cells per well, and 12 h later, the cells were exposed to 2.5–15 μmol/L cisplatin for 2 h. DNA was isolated using a Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA) according to the manufacturer's instructions, resuspended in 5% HCl, hydrolyzed for 30 min at 95 °C, and quantified spectrophotometrically based on the absorbance values at 260 nm relative to those of hydrolyzed calf thymus DNA standards. The amount of platinum adduct (Pt-adduct) in the DNA in the hydrolysate (picograms of platinum per microgram of DNA) was measured by graphite furnace atomic absorption spectrophotometry (FAAS) with Zeeman background correction (Perkin-Elmer 4100 ZL, Norwalk, CT, USA). To assess the time course of the loss of platinum from DNA, the cells were treated with 5 μmol/L cisplatin for 1 h to obtain quantifiable levels of platinum over the entire period of the experiment. DNA was isolated at 0, 6, 12, 18 or 24 h after drug exposure.
Immunofluorescence and alkaline comet assay
After transfection with various siRNAs, cells were treated with cisplatin or cisplatin plus gemcitabine for 2 h and subsequently cultured in fresh medium for 48 h. Then, immunofluorescence analysis was performed with the indicated antibodies as previously described31. The modified alkaline comet assay was performed as previously described31.
BrdU incorporation assay
Cells were cultured in RPMI medium (containing 10% FBS), and after exposure of the cells to cisplatin or cisplatin plus gemcitabine, bromodeoxyuridine (BrdU; 10 μmol/L) was added to the medium. One hour later, BrdU incorporation into DNA was analyzed with a BrdU incorporation kit (Roche Diagnostics) according to the manufacturer's instructions.
siRNA transfection
All siRNA reagents used in this study were purchased from Guangzhou RiboBio Co, Ltd (China). siRNAs were transfected using Lipofectamine 2000 (Invitrogen) according to the manufacturer's protocol, as previously described30. Three siRNAs against each gene were used for transfection of the experimental cells to minimize the possibility of off-target effects. The sequences of siRNAs targeting each gene are shown in Table S1.
Western blotting and real-time quantitative PCR
Cells were treated with the indicated drugs, and protein samples from whole cell lysates were prepared and analyzed using Western blotting as previously described30. The antibodies used to detect proteins in this study are described above.
For real-time quantitative PCR, total RNA was extracted from various cell samples using Trizol reagent (Invitrogen). Reverse transcription was conducted using Applied Biosystems Power SYBR Green PCR Master Mix, and the reactions were run on an ABI 7500 Fast Real-time PCR system, as previously described31. For RT-PCR, RNA was reverse-transcribed using a ReverTra Ace kit (Toyobo, Osaka, Japan) according to the manufacturer's instructions. The specific primer sequences of the genes detected in this study are shown in Table S2.
Sub-G1 and cell cycle analysis by flow cytometry
Cells were plated at 6×105 cells per well. After 24 h in culture, the cells were treated with the indicated drugs and incubated at 37 °C for 48 h. Then, both adherent and detached cells were harvested in cold 70% ethanol, centrifuged at 500×g, and stored at -20 °C for at least 2 h. After washing with PBS, the cells were resuspended in 200 μL of propidium iodide solution (200 μg/mL RNase A, 20 μg/mL propidium iodide, and 0.1% Triton X-100 in PBS) and incubated in the dark for 30 min. The propidium iodide fluorescence of each sample was measured on a flow cytometer (Guava, Merck Millipore). Guava Express Plus Software was used to quantify sub-diploid nuclei (sub-G1 phase), and cell cycle analysis was performed using ModFit LT software, excluding cells with a DNA content greater than 4N.
Statistical analysis
The IC50 was calculated as the cisplatin concentration that killed 50% of cells in the untreated control group. All data were expressed as the mean±SD. Statistical analysis was conducted with 2-tailed unpaired Student's t tests using SPSS 16.00 version (SPSS Inc., Chicago, IL). The differences between the compared groups were considered statistically significant at P<0.05.
Results
Response to platinum and the DNA-bound platinum levels in cisplatin resistant and sensitive NSCLC cell lines
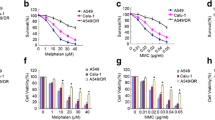
The cisplatin-resistant cell line A549/DR was generated by chronic treatment of A549 cells (human lung adenocarcinoma cell line) with low-dose cisplatin as previously described30. To determine whether the cisplatin-resistant phenotype is not specific to cisplatin but rather a phenomenon common to platinum agents, the cell viability assay was performed in A549/DR and A549 cells and two other NSCLC cell lines, LOU-NH91 (human lung squamous carcinoma cell line) and HCC4006 (human lung adenocarcinoma cell line), following treatment with cisplatin, carboplatin, oxaliplatin or gemcitabine. The results showed that A549/DR cells are also resistant to carboplatin and oxaliplatin in addition to cisplatin, even though the A549/DR cell line was derived via long-term treatment of the A549 cell line with cisplatin. The sensitivity of LOH-NH91 and HCC4006 cells to the three platinum agents was similar to that of A549 cells. Interestingly, A549/DR cells were also more resistant to gemcitabine relative to the three cisplatin-sensitive cell types despite less degree (Figure 1A, 1B, and 1C), suggesting that the mechanism of gemcitabine resistance at least partially overlaps with that of cisplatin in these NSCLC cell lines. Similar results were observed in the colony formation assay (Figure S1A–D).
Cisplatin resistance in A549/DR cells is not associated with the decrease of drug intracellular uptake and the increase of Pt-DNA adduct removal. (A) Cell viability measurement, A549, A549/DR, LOU-NH91 and HCC4006 cell lines growing in 96-well plates were treated with cisplatin, (B) carboplatin, (C) oxaliplatin, and (D) gemcitabine at indicated dose. The CCK-8 assay was used to determine cell viability. After treatment with drug as indicated for 2–4 h, cell proliferation reagent CCK-8 (DOJNDO, Japan) was added into media in each well and the cells were incubated for 2 h at 37 °C. The absorbance of each well was measured with a spectrophotometer reading at a wavelenth of 450 nm. Absorbance is assumed to be directly proportional to the number of viable cells (* P<0.05, ** P<0.01 vs A549, LOU-NH91 and HCC4006 cell lines). (E) Formation of platinum-DNA adducts in A549, A549/DR, LOU-NH91 and HCC4006 cell lines after a 2-h exposure to cisplatin as measured by the FAAS. (F) The rate of disappearance of platinum from total cellular DNA was measured in the four NSCLC cell lines after a 2-h exposure to cisplatin (10 μmol/L). Each datum represents the mean of three experiments.
One of the well-known mechanisms of cisplatin resistance is decreased drug intracellular uptake, which can result in less DNA damage and reduced cytotoxicity6. A comparison of cellular drug accumulation was carried out after the same time of exposure (2 h) to cisplatin. As shown in Figure 1E, following incubation with cisplatin, intracellular DNA-bound platinum levels were similar among all four cell lines and were increased with increasing drug concentrations. Because the rate of disappearance of platinum from the DNA accurately reflects the rate of removal of the most common platinum adduct32, the rate of disappearance of platinum from total cellular DNA was measured in the four cell lines after exposure to 10 μmol/L cisplatin. The results showed that there was no difference in the kinetics of platinum disappearance from DNA among the four NSCLC cell lines at the time point of 24 h. Although less platinum remained in the A549/DR cells at 18 and 24 h than in the other three cell lines, this difference did not reach statistical significance (Figure 1F). These data indicate that cisplatin resistance in A549/DR cells is not associated with the mechanism of the impediment of drug intracellular uptake or the enhancement of platinum DNA adduct removal.
Cisplatin induced ICLs and DSBs in cisplatin-resistant and cisplatin-sensitive NSCLC cell lines
Among the mechanisms of cisplatin resistance, the capacity of ICL and DSB repair is believed to play a main role, which is mainly carried out by the Fanconi anemia (FA) pathway in coordination with the TLS and homologous recombination (HR) pathways33. To investigate differences in DNA repair capacity, it is necessary to test the degree of damage in terms of ICLs and DSBs induced by cisplatin in different cell lines. We used fluorescence staining to determine the formation of γ-H2AX foci, a good indicator of DSBs. As shown in Figure 2A and 2B, the percentage of cells positive for γ-H2AX foci after cisplatin treatment was significantly lower in A549/DR cells than in the three cisplatin-sensitive cell lines. Moreover, the number of A549/DR cells with γ-H2AX foci declined 36 h after cisplatin treatment and returned to the untreated level by 60 h. In contrast, the number of cells with γ-H2AX foci in the three cisplatin-sensitive cell lines persisted at higher levels until 72 h post-treatment with cisplatin (Figure 2C). The DNA ICLs were determined using an alkaline comet assay at the single-cell level. Tail moment, a measurement of relative electrophoretic mobility, was markedly decreased in A549/DR cells after cisplatin treatment compared with the other three NSCLC cell lines (Figure 2D and 2E). Concomitant with the formation of DSBs and ICLs, the A549/DR cell line presented a slighter replication arrest and more rapid resumption of replication arrest compared to the three cisplatin-sensitive cell lines at 24 to 72 h after cisplatin treatment (Figure 2F). Together, these results indicate that the more effective DNA ICL and DSB repair in A549/DR cells could be responsible for their cisplatin resistance.
A549/DR cells show decreased γ-H2AX foci formation, diminished tail moment, and rapider resumption of replication arrest after cisplatin treatment. (A) A549, A549/DR, LOU-NH91 and HCC4006 cell lines were treated with 10 μmol/L cisplatin for 2 h, fixed and immunostained using anti-γ-H2AX antibody. (B) The percentage of γ-H2AX positive cells, defined as cells with a fluorescence intensity>500 units (control value) was quantified using Metafer software (* P<0.05 vs A549, LOU-NH91 and HCC4006 cell lines). (C) The four cell lines were treated with 10 μmol/L cisplatin, fixed and immunostained using anti-γ-H2A antibody. The percentages of γ-H2AX positive cells were quantified at indicated time points (* P<0.05 vs A549, LOU-NH91 and HCC4006 cell lines). (D) The four cell lines were treated with 10 μmol/L cisplatin for 2 h. The alkaline comet assay was performed to measure ICLs, and the images show detectable comet tails visualized under a fluorescent microscope. (E) Tail moments in the cells were quantified using Comet Score software version 1.5 (* P<0.05 vsA549, LOU-NH91, and HCC4006 cell lines). (F) DNA replication was measured by BrdU incorporation assay 24, 48, and 72 h after cisplatin (10 μmol/L) treatment (* P<0.05 vs A549, LOU-NH91, and HCC4006 cell lines).
TLS Pol η expression is markedly up-regulated in the A549/DR cell line compared to the cisplatin-sensitive cell lines upon cisplatin exposure
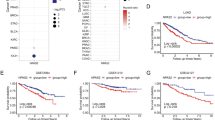
To evaluate the relationship between the expression of DNA repair pathway factors and the development of cisplatin resistance, we examined the protein and mRNA levels of several TLS polymerases and FA and HR pathway factors, including Pol η, Pol k (encoded by POLK), Pol β (encoded by POLB), Pol μ (encoded by POLM), Pol ν (encoded by POLN), FANCM, FANCJ, RAD18, RAD51, and ATR, in the A549/DR cell line and three cisplatin-sensitive NSCLC cell lines. As shown in Figure 3A–C, A549/DR cells exhibited higher protein and mRNA levels of TLS Pol η, Pol β and Pol ν as well as all FA and HR factors examined in this study than the three cisplatin-sensitive cell lines. It is noteworthy that the elevation in the levels of Pol η in A549/DR cells was the most significant among all TLS polymerases tested. Elevated FA and HR factor expression in A549/DR cells is expected because these factors are involved in cisplatin resistance32. Next, we evaluated the time-dependent expression of the TLS polymerases induced by cisplatin. Increased levels of Pol η protein and mRNA were observed in A549/DR cells upon cisplatin exposure, and the Pol η protein and mRNA levels continually increased during the 24-h post-treatment period (Figure 3D and 3F). At 16 and 24 h, the POLH mRNA levels were markedly higher than the mRNA levels of other TLS polymerases (Figure 3F). Conversely, the expression levels of all TLS polymerases tested at both the protein and mRNA levels were only slightly increased post-treatment with cisplatin in the three cisplatin-sensitive cell lines (Figures 3D–E and S1E–H). These findings suggest that Pol η may play a more important role in the cisplatin resistance of A549/DR cells.
Expressions of pol η were upregulated in A549/DR cells upon exposure to cisplatin. (A) Whole cell lysate was prepared from A549, A549/DR, LOU-NH91 and HCC4006 cell lines and subject to Western blotting with specific antibodies as indicated to measure the protein levels of various factors. (B and C) Total RNA was isolated from the four NSCLC cell lines, and subject to real-time quantitative PCR to measure the mRNA levels of various factors as indicated (* P<0.05, ** P<0.01 vs A549, LOU-NH91 and HCC4006 cell lines). (D) Protein expressions of TLS polymerases as indicated were analyzed by Western blotting using specific antibodies in whole cell lysate of A549 and A549/DR cells at different time points after cisplatin (10 μmol/L) treatment. β-actin was used as loading control. (E and F) Real-time quantitative PCR was performed to measure mRNA expression of TLS polymerases as indicated in A549 and A549/DR cells at different time points after cisplatin (10 μmol/L) treatment. The mRNA expressions of TLS polymerases were normalized to GAPDH; the untreated control was set to one (** P<0.01 vs POLK, POLB, POLM and POLN).
ATR deficiency affects the cisplatin-induced formation of Pol η foci and intracellular relocation but does not impede monoubiquitination of PCNA
The intracellular relocalization of Pol η to stalled replication forks in response to DNA-damaging agents is critical for its cellular activity34, which is dependent on the monoubiquitination of PCNA mediated by RAD18/RAD6 complexes28,35. Concomitantly, ATR is also activated by DNA-damaging agents and facilitates Pol η recruitment to stalled replication forks25,34. Western blotting showed a markedly increased level of PCNA monoubiquitination upon cisplatin exposure in the A549/DR cell line compared to the three cisplatin-sensitive cell lines (Figure 4A). Carboplatin, oxaliplatin and gemcitabine also induced PCNA monoubiquitination in A549/DR cells, although the PCNA monoubiquitination level induced by gemcitabine was lower than that induced by carboplatin or oxaliplatin (Figure 4B). To assess the impact of RAD18 and ATR inhibition on the intracellular relocation of Pol η, we depleted RAD18 and ATR using an siRNA transfection approach. The efficiency of RAD18 and ATR depletion was verified by Western blotting (Figure 4C). Parallel detection of immunofluorescence showed that the abundance of Pol η foci induced by cisplatin in the A549/DR cell line was the greatest among the four NSCLC cell lines (Figure 4D). Knockdown of RAD18 and ATR suppressed the cisplatin-mediated intracellular relocalization of Pol η foci, especially in A549/DR cells (Figure 4D). In addition, colocalization of Pol η and PCNA was observed in A549/DR cells upon exposure to cisplatin or cisplatin plus gemcitabine, and this colocalization was diminished by depleting RAD18 or ATR (Figure 4G). Although the depletion of ATR did not suppress the monoubiquitination of PCNA (Figure 4E and 4F), which is in accordance with the reports that PCNA ubiquitination is independent of ATR-mediated checkpoint activation36, the recruitment and intracellular relocalization of PCNA in ATR-deficient cells were impeded (Figure 4G), which may result from Pol η relocalization defects.
PCNA monoubiquitination and intracellular location of Pol η. (A) PCNA monoubiquitination induced by cisplatin. A549, A549/DR, LOU-NH91 and HCC4006 cell lines were treated with cisplatin (10 μmol/L), and ubiquitinated PCNA (Ub-PCNA) from the cell extracts was detected using a monoclonal anti-PCNA antibody by western blotting. (B) A549/DR cells were treated with cisplatin (Cis, 10 μmol/L), or carboplatin (Car, 10 μmol/L), or oxaliplatin (Oxa, 15 μmol/L), or gemcitabine (Gem, 10 nmol/L), and Ub-PCNA was detected as described above. (Con: negative control). (C) Western blotting was performed to verify the efficiency of transfection with siRAD18 or siATR in A549 or A549/DR cells. siCon: control siRNA. (D) After transfection with siRNAs as indicated, intracellular location of Pol η in the four NSCLC cell lines was analyzed by immunostaining using anti-Pol η body after treatment with cisplatin at indicated doses. The lower panel shows the quantification of the mean fluorescent intensity of Pol η antibody staining as calculated by Image software (** P<0.01 vs A549, LOU-NH91, and HCC4006 cell lines). (E) Intracellular colocalizations of Pol η and PCNA in A549/DR cells post-transfection with siRNAs as indicated were analyzed by immunostaining using anti-Pol η and anti-PCNA bodies after treatment with cisplatin and cisplatin plus gemcitabine. (F and G) PCNA monoubiquitinations in A549 and A549/DR cell lines post-transfection with siRNAs as indicated were analyzed by Western blotting after cisplatin (10 μmol/L) treatment.
Depletion of POLH synergizes with ATR inhibition to hypersensitize A549/DR cells to cisplatin and cisplatin plus gemcitabine
To further investigate the contribution of TLS polymerases to cisplatin sensitivity, we depleted the expression of POLH, POLB, or POLN in both the A549 and A549/DR cell lines using siRNA transfection and examined the changes in their sensitivity to cisplatin. The transfection efficiency of siRNAs against POLH, POLB, POLN, and POLH together with ATR was validated by Western blotting (Figure 5A). As expected, knockdown of these TLS factors increased the sensitivity of A549 and A549/DR cells to cisplatin or cisplatin plus gemcitabine (Figure S2A–D; Tables S3 and S4). Similar results were obtained using three different siRNA sequences each for POLH, POLB and POLN in the two cell lines, minimizing the possibility of off-target effects (Figures S2G and S2H). Importantly, the degree of drug sensitization inducted by depleting POLH was greater than that induced by depleting POLB and POLN (Figure S2A-D; Tables S3 and S4). It is notable that the sensitization effect of POLH depletion was more significant in A549/DR cells than in A549 cells. Depletion of POLH increased cisplatin sensitivity by up to 1.65-fold in A549 cells and by up to 6-fold in A549/DR cells, as indicated by analysis of the IC50 for cisplatin (Tables S3 and S4). Because A549/DR cells were also more resistant to gemcitabine in our initial experiments and because the combination of cisplatin and gemcitabine is one of the first-line chemotherapy regimens for NSCLC, we tested the impact of gemcitabine combined with cisplatin on the survival of the two NSCLC cell lines. As expected, the addition of gemcitabine increased the cytotoxicity of cisplatin, especially in A549/DR cells depleted of POLH (Figures S2B and S2D; Tables S3 and S4). Co-depletion of POLH and RAD18 did not produce additional sensitization to cisplatin or cisplatin plus gemcitabine compared to depletion of POLH alone (Figure 5B–E; Tables S3 and S4), which supports the notions that RAD18 is subordinate to Pol η and that RAD18 and Pol η function in the same pathway to confer tolerance to cisplatin-induced DNA damage37. In contrast, co-depletion of POLH and ATR further increased the sensitivity of A549/DR cells to cisplatin or cisplatin plus gemcitabine compared to depletion of POLH or ATR alone (Figures 5D–E; Table S4). In addition, we used VE-822, a selective and ATP-competitive ATR inhibitor38, to further assess the role of ATR in cisplatin resistance. The results showed that the combination of POLH knockdown with VE-822 treatment (0.5 μmol/L) significantly enhanced the cytotoxicity of cisplatin plus gemcitabine to A549/DR cells compared to POLH knockdown alone (Figure S2F), which was similar to the effect of co-depletion of POLH and ATR (Figure 5E). However, in A549 cells, depletion of both POLH and ATR as well as either POLH or ATR alone induced comparable sensitization to cisplatin or cisplatin plus gemcitabine (Figure 5B–C; Table S3). These results suggest that the sensitization produced by co-depleting POLH and ATR mainly occurs in cisplatin-resistant A549/DR cells. Taken together, these results suggest that POLH depletion can synergize with ATR inhibition to potentiate A549/DR cell death by cisplatin and cisplatin plus gemcitabine.
Co-depletion of POLH and ATR hypersensitize A549/DR cells to cisplatin, and cisplatin plus gemcitabine. (A) Western blotting was performed to verify the efficiency of the transfections with siPOLH, siPOLB, siPOLN and siATR in A549 and A549/DR cell lines siCon: control siRNA. (B and C) The viability analysis of A549 cells depleted of POLH or ATR alone, or double depleted of POLH and RAD18, or double depleted of POLH and ATR after treatment with cisplatin or cisplain plus gemcitabine as indicated doses. (D and E) The viability analysis of A549/DR cells depleted of the genes as indicated after treatment with cisplatin or cisplatin plus gemcitabine. (G and H) A549/DR cells depleted of POLH, or POLB, or POLN alone, or co-depleted of POLH and ATR were treated with cisplatin and cisplatin plus gemcitabine for 4 h and apoptotic cells was measured as sub-G1 fraction by flow cytometry (* P<0.05 vs siPOLB and siPOLN; ## P<0.01 vs siPOLH). (F) Representative imagines of cell cycle analysis show that sub-G1 peak was higher in A549/DR cells depleted of POLH alone and co-depleted of POLH and ATR than in the cells depleted of POLB or DOLN alone.
The presence of a sub-G1 cell population is indicative of DNA degradation during cell death via apoptosis. Consistent with the cell viability assay results, apoptosis analysis showed that co-depletion of POLH and ATR clearly increased the frequency of A549/DR cells in the sub-G1 phase induced by cisplatin and cisplatin plus gemcitabine compared to depletion of POLH, POLB, or POLN alone (Figure 5G and 5H). Representative images of the cell cycle analysis results showed that the A549/DR cells co-depleted of POLH and ATR displayed not only marked S/G2 phase arrest but also a higher sub-G1 peak in response to cisplatin and cisplatin plus gemcitabine (Figure 5F). Meanwhile, dramatically increased cleaved caspase-3 and cleaved PARP expression levels were observed in A549/DR cells co-depleted of POLH and ATR following cisplatin treatment (Figure S3A and S3B), further demonstrating that combined knockdown of POLH and ATR facilitates the cisplatin-induced apoptotic death of A549/DR cells.
Co-depletion of POLH and ATR impedes the repair of ICLs and DSBs induced by cisplatin plus gemcitabine
Lastly, we aimed to establish whether the cisplatin sensitization of A549/DR cells produced by depleting POLH or co-depleting POLH and ATR was derived from the impairment of ICL and DSB repair. ICLs represent a severe barrier to DNA replication, causing DSBs at blocked replication forks. To assess the impact of POLH and ATR on DNA replication and DSB formation, replication was analyzed by the BrdU incorporation assay, and DSB formation was tested by detection of γ-H2AX foci. The results revealed that depletion of POLH alone or co-depletion of POLH and ATR markedly enhanced the inhibition of replication induced by cisplatin plus gemcitabine (Figure S3C) and dramatically increased the number of cells positive for γ-H2AX foci (Figure 6A and 6B). Analysis of the kinetics of the formation of γ-H2AX foci showed that 80%–90% of A549/DR cells depleted of POLH alone or depleted of both POLH and ATR exhibited positivity for γ-H2AX foci 24 h after treatment with cisplatin plus gemcitabine, and the percentage of cells with γ-H2AX foci remained at elevated levels until 72 h post-treatment. Conversely, approximately 50% of the cells depleted of POLB or POLN were positive for γ-H2AX foci 24 h post-treatment with the same drugs, and this percentage decreased continuously and returned to the control level at 72 h (Figure 6C). Consistent with the observations regarding γ-H2AX foci, A549/DR cells depleted of POLH or co-depleted of POLH and ATR displayed a markedly prolonged tail moment in comparison with cells depleted of POLB or POLN alone (Figure 6D and 6E). These findings indicate that the impediment of ICL and DSB repair caused by depleting POLH and ATR is responsible for the drug sensitization of the cells. To further determine the effect of POLH and ATR on the DNA damage response induced by cisplatin and gemcitabine in A549/DR cells, ATR activity was analyzed by measuring P-Chk1 (S317), and DNA damage accumulation was assessed by detecting γ-H2AX (S139), P-KAP1 (S824) and P-RPA2 (S4/S8). Knockdown of POLH alone led to elevation of cisplatin-induced P-Chk1 expression and a concurrent increase in γ-H2AX, P-KAP1 and P-RPA2 expression in a concentration-dependent manner (Figure 6F), implying the activation of ATR/Chk1 signaling and the accumulation of DNA damage. Knockdown of ATR alone augmented the phosphorylation of H2AX, KAPI and RPA2, but not Chk1, which may result from ATR inhibition. Combined knockdown of POLH and ATR resulted in a decrease in P-Chk1 expression despite cisplatin exposure, but the levels of γ-H2AX, P-KAP1 and P-RPA2 were further increased under these conditions (Figure 6F), suggesting that ATR knockdown inhibited the phosphorylation of Chk1 (S317) and ATR/Chk1 signaling and augmented the accumulation of DNA damage. These results revealed that continuously increased accumulation of DNA damage (ICLs and DSBs) is a contributing mechanism responsible for the sensitization of A549/DR cells with co-depletion of POLH and ATR to cisplatin and gemcitabine.
Co-depletion of POLH and ATR in A549/DR cell resulted in marked impairment of ICL and DSB repair and increase of DNA damage accumulation. (A) After transfection with siRNAs as indicated, A549/DR cells were treated with cisplatin or cisplatin plus gemcitabine in indicated doses for 4 h, cultured in fresh medium for another 24 h, fixed and immunostained with an anti-γ-H2AX antibody. (B) The percentage of γ-H2AX foci positive cells was quantified using Metafer software (** P<0.01 vs siPOLB or siPOLN; # P<0.05 vs siPOLH). (C) The percentages of γ-H2AX positive cells were quantified at indicated time points. Each time point represents the mean of γ-H2AX positive/total cells (%) derived from five independent fields in each culture (* P<0.05 vs siPOLH).
Co-depletion of POLH and ATR in A549/DR cell resulted in marked impairment of ICL and DSB repair and increase of DNA damage accumulation. (D) A549/DR cells were transfected with siRNAs as indicated, and then treated with cisplatin or cisplatin plus gemcitabine for 4 h. An alkaline comet assay was performed to measure ICLs, and the images show detectable comet tails visualized under a fluorescent microscope. (E) Tail moments in A549/DR cells were quantified using Comet Score software version 1.5 (** P<0.01 vs siPOLB or siPOLN; # P<0.05 vs siPOLH). (F) Western blotting for phosphorylated-Chk1, -KAP1, -RPA2, and -H2AX in A549/DR cells depleted of POLH and ATR alone or co-depleted of POLH and ATR following treatment with cisplatin as indicated doses, GAPDH was used as a loading control. The intensity of protein bands was quantified by densitometry and presented in Figures S4.
Discussion
Although the mechanisms of cisplatin resistance are not fully understood, it is generally believed that decreased drug uptake, increased drug inactivation and removal of Pt-DNA adducts, and enhanced DNA repair capacity are associated with cisplatin resistance4,5,6. In this study, however, we found that the removal of Pt-DNA adducts that are mostly composed of intrastrand crosslink adducts is not responsible for the cisplatin resistance of A549/DR cells. Upon cisplatin exposure, A549/DR cells exhibited significantly enhanced ICL and DSB repair capacity compared to cisplatin-sensitive NSCLC cell lines. These results indicate that the effective repair of DNA damage is involved in the cisplatin resistance of A549/DR cells.
It is known that cisplatin-mediated ICLs and DSBs are repaired by the FA pathway in coordination with TLS and the HR pathway33. TLS DNA polymerases, such as Pol θ (encoded by POLQ), Pol ζ (consisting of the catalytic subunit REV3 and the structural subunit REV7), and Pol η, are essential for ICL repair, as cells deficient in any of these factors are especially sensitive to cross-linking agents13,14,21,33. A recent study reported by our team showed that the expression levels of Pol θ, Pol ζ and Pol η were up-regulated in A549/DR cells compared to A549 cells. The cisplatin sensitization effect of co-depleting POLQ and BRCA2, an HR factor, on A549/DR cells was more significant than that of co-depleting POLH and BRCA2, although A549/DR cells depleted of POLH alone were more sensitive to cisplatin than cells depleted of POLQ alone39. Additionally, silencing POLQ in A549/DR cells increased RAD51 expression and the formation of its foci, and inhibition of the HR pathway by depleting RAD51 or BRCA2 increased POLQ expression, suggesting that Pol θ in NSCLC cells suppresses HR activity and participates in DSB repair through an alternative pathway39. Here, we show that A549/DR cells displayed increased expression levels of TLS polymerases (such as Pol η, Pol β, and Pol ν) and FA and HR pathway factors compared with cisplatin-sensitive NSCLC cells. Moreover, the elevated level of Pol η expression was the most evident among all TLS polymerases examined in this study. Remarkably, the expression of Pol η in the A549/DR cell line was significantly up-regulated in response to cisplatin exposure, which was not found in three cisplatin-sensitive NSCLC cell lines. Furthermore, depletion of Pol η increased cisplatin sensitivity by up to 6-fold in A549/DR cells but by only 1.65-fold in A549 cells. These data suggest that Pol η may play a more important role than other TLS polymerases in mediating A549/DR cell resistance to cisplatin and that inhibition of Pol η is a potential therapeutic strategy for sensitizing cisplatin-resistant NSCLC cells to cisplatin. The correlation of high Pol η expression with cisplatin resistance in NSCLC was reported in a clinical study, which showed that high Pol η mRNA levels in tumor tissue were strongly associated with shorter survival in the group of patients with advanced NSCLC treated with platinum-based chemotherapy40. Pol η expression levels were correlated with cisplatin sensitivity in vitro in a panel of NSCLCL cell lines40, which is in accordance with our results.
The activation of TLS polymerases in response to DNA damage is mainly mediated by monoubiquitination of PCNA34,37. Monoubiquitinated PCNA triggers the translocation of Pol η to stalled DNA replication forks and forms foci in the nucleus, thereby initiating TLS and DNA damage tolerance36,37. Concomitantly, ATR is activated by DNA-damaging agents such as cisplatin to phosphorylate Pol η and facilitate Pol η recruitment to stalled DNA replication forks41. In this study, in accordance with elevated expression of Pol η mRNA and protein, the level of PCNA monoubiquitination and the abundance of Pol η foci induced by cisplatin was higher in A549/DR cells than in three cisplatin-sensitive cell lines.
Human cells deficient in Pol η are sensitive to cisplatin10,15. Knockdown of Pol η in breast cancer, glioma and melanoma cells caused marked sensitization to some ICL-inducing agents, such as fotemustine, mafosfamide and lomustine42. Given that inhibiting ATR activates origin firing under replicative stress, exhausts RPA and generates fork collapse and DNA DSBs that result in the induction of death signals25, knockdown of Pol η could synergize with ATR inhibition to hypersensitize cancer cells to cisplatin . Here, we demonstrate that depletion of Pol η combined with suppression of ATR by siRNA transfection or ATR inhibitor treatment further increases the sensitivity of A549/DR cells to cisplatin or cisplatin plus gemcitabine compared with depletion of Pol η alone. In contrast, the effect of co-inhibition of Pol η and ATR in sensitizing A549/DR cells to cisplatin or cisplatin plus gemcitabine was not observed in A549 cells, indicating that co-inhibition of Pol η and ATR can reverse the resistance of cisplatin-resistant NSCLC cells to cisplatin or cisplatin plus gemcitabine.
ICLs are the most cytotoxic and genotoxic lesions caused by cisplatin, as they block DNA replication and transcription and, if unrepaired, lead to the generation of DSBs and chromosomal rearrangements. TLS polymerases are recognized to be essential for ICL repair in both S/G2 and G1 phases by bypassing an ICL unhooked from one of the two cross-linked strands33. Pol η can bypass various structurally distinct unhooked ICLs19,20 and participate in the repair of interstrand crosslinking agent-induced ICLs21,22. Additionally, the FA pathway plays an important role in DNA damage sensing and signaling during S/G2 phase of the cell cycle in cells with ICLs33,43. The FA and ATR/Chk1 signaling pathways are activated concurrently in response to ICL-related damage. Activated ATR and its downstream kinases phosphorylate proteins upstream in the FA pathway, including the core complex proteins FANCA, FANCG, FANCE and FANCM, which are essential for efficient monoubiquitination of FANCI and FANCD2 as well as cellular tolerance to ICL-inducing agents44,45. ATR also phosphorylates HR pathway proteins such as XPCC3, BRCA1, RAD17, MCM, and RPA, which are prerequisites for HR-mediated repair of ICL-induced replication-dependent DSBs46,47. Therefore, ATR plays a crucial role in the DNA damage response by relaying and amplifying the DSB damage signal. One study reported that the ATR/Chk1 pathway is over-activated in human cells deficient in Pol η after UV exposure. Inhibition of ATR/Chk1 activity exacerbates replication fork stalling and S-phase arrest and sensitizes these cells to UV irradiation, which are not observed in wild-type cells, suggesting that in the absence of Pol η, the ATR/Chk1 pathway becomes essential for replication resumption by an alternative pathway via fork stabilization48. Thus, one explanation for our data may be that the resolution of cisplatin-induced fork blockage leading to ICLs and DSBs in cells depleted of Pol η relies on the FA and HR pathways, which are regulated by the ATR/Chk1 signaling pathway. Consequently, combined knockdown of Pol η and ATR strongly potentiates the cytotoxicity of cisplatin and gemcitabine to drug-resistant NSCLC cells.
We are aware of some limitations of the present study, including that only one cisplatin-resistant subcellular clone, A549/DR, was used for investigating sensitization to cisplatin by co-inhibition of Pol η and ATR. Further studies with several cisplatin-resistant subcellular clones from other cancer cell lines are needed.
In conclusion, we show that the elevated expression of Pol η is associated with the resistance of NSCLC cells to cisplatin. Additionally, we demonstrate that although disabling Pol η sensitizes NSCLC cells to cisplatin or cisplatin plus gemcitabine, co-inhibition of Pol η and ATR further increases cisplatin-resistant NSCLC cell death mediated by cisplatin plus gemcitabine. These findings indicate that Pol η knockdown synergizes with ATR suppression to reverse the resistance of NSCLC cells to cisplatin. Therefore, co-inhibition of Pol η and ATR may be a potential therapeutic strategy for the treatment of platinum-resistant NSCLC.
Author contribution
Xiao-qin LI and Jin REN contributed equally. Xiao-qin LI, Jin REN, and Jian LI designed research; Xiao-qin LI, Jin REN, Yu-jiao CHEN, Min WU, Yan WU, and Kang CHEN performed research; Xiao-qin LI, Jin REN, Ping CHEN and Jian LI analyzed data; Xiao-qin LI, Jin REN and Jian LI wrote paper.
References
Minguet J, Smith KH, Bramlage P. Targeted therapies for treatment of non-small cell lung cancer - Recent advances and future perspectives. Int J Cancer 2016; 138: 2549–61.
Raparia K, Villa C, DeCamp MM, Patel JD, Mehta MP. Molecular profiling in non-small cell lung cancer: a step toward personalized medicine. Arch Pathol Lab Med 2013; 137: 481–91.
Socinski MA, Evans T, Gellinger S, Hensing TA, Sequisct LVD, Ireland B, et al. Treatment of stage VI non-small cell lung cancer: diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians Evidence-Based Clinical Practice Guideline. Chest 2013; 143: e341s–e368s.
Chaney SG, Campbell SL, Bassett E, Wu Y. Recognition and processing of cisplatin - and oxaliplatin - DNA adducts. Crit Rev Oncol Hematol 2005; 53: 3–11.
Ahmad S. Platinum-DNA interactions and subsequent cellular processes controlling sensitivity to anticancer platinum complexes. Chem Biodivers 2010; 7: 543–66.
Martin LP, Hamiton TC, Schilder RJ. Platinum resistance: the rote of DNA repair pathways. Clin Cancer Res 2008; 14: 1291–5.
Lord CJ, Ashworth A. The DNA damage response and cancer therapy. Nature 2012; 481: 287–94.
Broyde S, Wang L, Rechkoblit O, Geacintov NE, Patel DJ. Lesion processing: high-fidelity versus lesion-bypass DNA polymerases. Treads Biochem Sci 2008; 33: 209–19.
Zhao Y, Biertümpfel C, Gregory MT, Hua YJ, Hanaoka F, Yang W. Structural basis of human DNA polymerase η-mediated chemoresistance to cisplatin. Proc Natl Acad Sci U S A 2012; 109: 7269–74.
Chen YW, Cheaver JE, Hanaoka F, Chang CF, Chou KM. A novel role of DNA polymerase eta in modulating cellular sensitivity to chemotherapeutic agents. Mol Cancer Res 2006; 4: 257–65.
Xie K, Doles J, Hemann MT, Walker GC. Error-prone translesion synthesis mediates acquired chemoresistance. Proc Natl Acad Sci U S A 2010; 107: 20792–7.
Lin X, Okuda T, Trang J, Howell SB. Human REV1 modulates the cytotoxicity and mutagenicity of cisplatin in human ovarian carcinoma cells. Mol Pharmacol 2006; 69: 1748–54.
Doles J, Oliver TG, Cameron ER, Hsu G, Tacks T, Walker GC, et al. Suppression of Rev3, the catalytic submit of Pol zeta, sensitize drug-resistant lung tumor to chemotherapy. Proc Natl Acad Sci U S A 2010; 107: 20786–91.
Sharma S, Shah NA, Joiner AM, Roberts KH, Canman CE. DNA polymerase ζ is a major determinant of resistance to platinum-based chemotherapeutic agents. Mol Pharmacol 2012; 81: 778–87.
Albertella MR, Green CM, Lehmann AR, O'eonor MJ. A role for polymerase η in the cellular tolerance to cisplatin-induced damage. Cancer Res 2005; 65: 9799–806.
Bassett E, King NM, Bryant MF, Hector S, Pendyala L, Chaney SG, et al. The role of DNA polymerase η in translesion synthesis past platinum-DNA adducts in human fibroblasts. Cancer Res 2004; 64: 6469–75.
Bassett E, Vaisman A, Havener JM, Masutani C, Hanaoka F, Chaney SG. Efficiency of extension of mismatched primer termini across from cisplatin and oxaliplatin adducts by human DNA polymerase beta and eta in vitro. Biochemistry 2003; 42: 14197–206.
Havener JM, Nick McElhinny SA, Bassett E, Gauger M, Ramsden DA, Chaney SG. Translesion synthesis past platinum DNA adducts by human DNA polymerase mu. Biochemistry 2003; 42: 1777–88.
Ho TV, Guainazzi A, Derkunt SB, Euoiu M, Scharer OD. Structure-dependent bypass of DNA interstrand crosslinks by translesion synthesis polymerases. Nucleic Acids Res 2011; 39: 7455–64.
Klug AR, Harbut MB, Lloyd RS, Minko IG. Replication bypass of N2-deoxyguanosine interstrand cross-links by human DNA polymerases eta and iota. Chem Res Tox 2012; 25: 755–62.
Zheng H, Wang X, Warren AJ, Legerski RJ, Nairn RS, Hamiton JW, et al. Nucleotide excision repair- and polymerase eta-mediated error-prone removal of mitomycin C interstrand cross-links. Mol Cell Biol 2003; 23: 754–61.
Mogi S, Butcher CE, Oh OH. DNA polymerase eta reduces the gamma-H2AX response to psoralen interstrand crosslinks in human cells. Exp Cell Res 2008; 314: 887–95.
Hicks JK, Chute CL, Paulsen MT, Ragland RL, Howlett NG, Guéranger Q, et al. Differential roles for DNA polymerases eta, zeta, and REV1 in lesion bypass of intrastrand versus interstrand DNA cross-links. Mol Cell Biol 2010; 30: 1217–30.
Richardson KA, Vega TP, Richardson FC, Moore CL, Rohloff JC, Tomkinson B, et al. Polymerization of the triphosphates of Arac, 2′, 2′-diffluorodeoxycytidine (dFdC) and OSI-7836 (T-araC) by human DNA polymerase alpha and DNA primase. Biochem Pharmacol 2004; 68: 2337–46.
Cimprich KA, Cortez D. ATR: an essential regulator of genome integrity. Nat Rev Mol Cell Biol 2008; 9: 616–62.
Aguilera A, Gomez-Gonzalez B. Genome instability: a mechanistic view of its cause and consequences. Nat Rev Genet 2008; 9: 204–17.
Branzei D, Foiani M. Maintaining genome stability at the replication fork. Nat Rev Mol Cell Biol 2010; 11: 208–19.
Chang DJ, Lupardus PJ, Cimprich KA. Monoubiquitination of proliferating cell nuclear antigen induced by stalled replication requires uncoupling of DNA polymerase and mini-chromosome maintenance helicase activities. J Biol Chem 2006; 281: 32081–8.
Charrier JD, Durrant SJ, Golec JM, Kay DP, Knegtel RM, MacCormik S, et al. Discovery of potent and selective inhibitors of ataxis telangiectasia mutated and Rad3 related (ATR) protein kinase as potential anticancer agents. J Med Chem 2011; 54: 2320–30.
Dai CH, Li J, Chen P, Jiang HG, Wu M, Chen YC. RNA interferences targeting the Fanconi anemia/BRCA pathway usptream genes reverse cisplatin resistance in drug-resistant lung cancer cells. J Biomed Sci 2015; 22: 77.
Chen P, Li J, Chen YC, Qian H, Chen YJ, Su JY, et al. The functional status of DNA repair pathway determines the sensitization effect to cisplatin in non-small cell lung cancer cells. Cell Oncol 2016; 39: 511–22.
Jonhnson SW, Perez RP, Godwin AK, Yeung AT, Handel LM, Ozols RF, et al. Role of platinum-DNA adduct formation and removal in cisplatin resistance in human ovarian cancer cell lines. Biochem Pharmacol 1994; 47: 689–97.
Clauson C, Schärer OD, Niedernhofer L. Advances in understanding the complex mechanisms of DNA interstrand cross-link repair. Cold Spring Harbor Perspect Biol 2013; 5: a012732.
Chem YW, Cheaver JE, Hatahet Z, Honkanen RE, Chang JY, Yen Y, et al. Human DNA polymerase η activity and translocation is regulated by phosphorylation. Proc Natl Acad Sci U S A 2008; 105: 16578–83.
Kannouche PL, Wing J, Lehmann AR. Interaction of human DNA polymerase η with monoubiquitinated PCNA: A possible mechanism for the polymerase switch in response to DNA damage. Mol Cell 2004; 14491–500.
Niimi A, Brown S, Sabbioneda S, Kannouche PL, Scott A, Yasui A, et al. Regulation of proliferating cell nuclear antigen ubiquitination in mammalian cells. Proc Natl Acad Sci U S A 2008; 105: 16125–30.
Watanabe K, Tateishi S, Kawasuji M, Tsurimoto T, Inoue H, Yamaizum M. Rad18 guides pol eta to replication stalling sites through physical interaction and PCNA monoubiquitination. EMBO J 2004; 23: 3886–96.
Fokas E, Prevo R, Pollard JR, Reaper PM, Charlton PA, Cornelissen B, et al. Targeting ATR in vivo using the novel inhibitor VE-822 results in selective sensitization of pancreatic tumors to radiation. Cell Death Dis 2012; 3: e441.
Dai CH, Chen P, Li J, Lan T, Chen YC, Qian H, et al. Co-inhibition of pol θ and HR genes efficiently synergize with cisplatin to suppress cisplatin-resistant lung cancer cells survival. Oncotarget 2016; 7:65157–70.
Ceppi P, Novello S, Cambieri A, Longo M, Monica V, Lo lacono M, et al. Polymerase η mRNA expression predicts survival of non-small cell lung cancer patients treated with platinum-based chemotherapy. Clin Cancer Res 2009; 15:1039–45.
Shechter D, Costanzo V, Gautier J. Regulation of DNA replication by ATR: signaling in response to DNA intermediates. DNA Repair (Amst) 2004; 3: 901–8.
Tomicic MT, Aasland D, Naumann SC, Meise R, Barckhausen C, Kaina B, et al. Translesion polymerase η is upregulated by cancer therapeutics and confers anticancer drug resistance. Cancer Res 2014; 74: 5585–96.
Haynes B, Saadat N, Myung B, Shekhar MPV. Crosstalk between translesion synthesis, Fanconi anemia network, and homologous recombination repair pathway in interstrand DNA crosslink repair and development of chemoresistance. Mutat Res Rev Mutat Res 2015; 763: 258–66.
Singh TR, Ali AM, Paramasivam M, Pradhan A, Wahengbam K, Seidman MM, et al. ATR-dependent phosphorylation of FANCM at serine 1045 is essential for FANCM function. Cancer Res 2013; 73: 4300–10.
Andreassen PR, D'Andrea AD, Taniguchi T. ATR couples FANCD2 monoubiquitination to the DNA-damage response. Genes Dev 2004; 18: 1958–63.
Summers KC, Shen F, Sierra Potchanant EA, Phipps EA, Hickey RJ, Malkas LH. Phosphorylation: the molecular switch of double-strand break repair. Int J Proteomics 2011; 2011: 373816.
Somyajit K, Basavaraju S, Scully R, Nagaraju G. ATM- and ATR-mediated phosphorylation of XRCC3 regulates DNA double-strand break-induced checkpoint activation and repair. Mol Cell Biol 2013; 3: 1830–44.
Despras E, Oabouss F, Hyrien O, Marheineke K, Kannouche PL. ATR/Chk1 pathway is essential for resumption of DNA synthesis and cell survival in UV-irradiated XP variant cells. Hum Mol Genet 2010; 19: 1690–701.
Acknowledgements
This research was partially supported by the National Youth Science Foundation of China (Grant No 81402485).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Supplementary Figure S1
The colony formation of four NSCLC cell lines, and cisplatin-induced the protein and mRNA expressions of TLS polymerases in LOU-NH91 and HCC4006 cell lines.
Supplementary Figure S2
The viability assay of A549 and A549/DR cell lines after different treatments.
Supplementary Figure S3
Protein expressions of cleaved caspase-3 and PARP, and DNA replication in A549/DR cells transfected with siRNAs as indicated after drug treatments.
Supplementary Information
Supplementary Tables
Rights and permissions
About this article
Cite this article
Li, Xq., Ren, J., Chen, P. et al. Co-inhibition of Pol η and ATR sensitizes cisplatin-resistant non-small cell lung cancer cells to cisplatin by impeding DNA damage repair. Acta Pharmacol Sin 39, 1359–1372 (2018). https://doi.org/10.1038/aps.2017.187
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2017.187
- Springer Nature Singapore Pte Ltd.
Keywords
This article is cited by
-
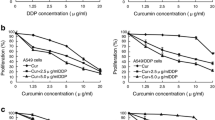
Curcumin overcome primary gefitinib resistance in non-small-cell lung cancer cells through inducing autophagy-related cell death
Journal of Experimental & Clinical Cancer Research (2019)