Abstract
Saprochaete/Magnusiomyces is among rare yeasts which might emerge as causes of breakthrough infections and nosocomial outbreaks. Identification to the species level might be a challenge in clinical laboratories. Data on virulence factors are scarce and antifungal susceptibility testing methodology is not definite. The aim of this study was to confirm species identification of clinical Saprochaete/Magnusiomyces isolates, find out their virulence factors, and obtain antifungal minimum inhibitory concentrations with two reference methods. Of the 57 isolates included, 54 were Saprochaete capitata and four were Saprochaete clavata as identified by ID32C, MALDI-TOF MS, and sequencing. When tested using phenotypic methods, all isolates were negative for coagulase, hemolysis, acid proteinase, and phospholipase, 56.1% were positive for esterase, and 19.3% had intermediate surface hydrophobicity. All isolates formed biofilms, with 40.4% of the isolates producing more biomass than biofilm-positive reference strain Candida albicans MYA-274. Antifungal susceptibility testing needed an adjusted spectrophotometric inoculum than recommended in reference methods for Candida/Cryptococcus. In conclusion, Saprochaete/Magnusiomyces species could be identified using methods available in the clinical laboratories. Despite the disadvantages of the phenotypic methods, esterase positivity was observed for the first time. A high biomass production was observed in biofilms. The need for standardization of antifungal susceptibility testing was brought to attention.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Saprochaete/Magnusiomyces species are rare causes of human infection [1]. They may cause invasive and mortal infections, especially in hemato-oncology patients and other immunocompromised hosts; nevertheless, they may be encountered in immunocompetent people [1, 2]. There are sporadic cases and nosocomial outbreaks caused by Saprochaete/Magnusiomyces species in different countries of the world [1,2,3,4]. Bloodstream infections as well as infections involving organs such as liver, lung, endocardium, brain, bone, joint, and skin were reported [1, 5, 6]. Some of these cases emerged as breakthrough infections, predominantly under echinocandins [1, 7].
Taxonomically, Saprochaete/Magnusiomyces species are classified under the Ascomycetes clade, Saccharomycetes class, order Saccharomycetales, and Dipodascaceae family [8, 9]. Different genera names, such as Geotrichum, Blastoschizomyces, and Saprochaete for anamorphs and Dipoascus and Magnusiomyces for teleomorphs, were used in the literature [1, 9]. The current nomenclature accepted for the most common clinical isolates are Magnusiomyces capitatus (An: Saprochaete capitata) and Saprochaete clavata (Tl: Magnusiomyces clavatus) [1, 8]. In order to avoid confusion, S. capitata and S. clavata were preferred throughout the text.
S. capitata and S. clavata are difficult to identify at the species level as their macroscopic and microscopic morphology is similar [1, 10]. They form white or cream yeast-like colonies which are smooth to wrinkled, irregularly spreading colonies in which the aerial hyphae become evident as they mature. Microscopically, yeast cells, true hyphae, pseudohyphae, abundant annelloconidia, and few arthroconidia can be seen on cornmeal Tween 80 medium [1, 8, 9]. Saprochaete/Magnusiomyces species are urease negative and have few different biochemical properties such as salicin and cellobiose positivity of S. clavata [1, 8]. Apart from these conventional tests, matrix-assisted laser desorption/ionization mass spectrophotometry (MALDITOF-MS) and gene sequencing methods can be used for species-level identification. Accuracy of both methods relies on the database used. MALDI-TOF MS successfully identified most fungi including Saprochaete/Magnusiomyces when larger and more reliable databases are used [1, 10]. Gene sequencing is accepted as the gold standard for microbial identification, and the ITS is accepted as the primary fungal barcode for fungal identification [11]. However, some studies could not differentiate Saprochaete/Magnusiomyces species using only ITS sequences, due to unreliable databases which contained misidentified isolates and/or insufficient length of the sequences used [10]. Analysis of additional gene sequences such as Rpb2, Tef1α, and Act, whole genome analysis, and phylogenetic trees were proposed to overcome these problems [1, 3, 8, 9]. As correctly differentiating the two species is the first step in determining the differences in epidemiology, prognosis, virulence, and management, understanding the reliability of the identification methods in clinical microbiology laboratories is fundamental.
Saprochaete/Magnusiomyces infections might have very high mortality rates, which are attributed to the vulnerability of the host and their antifungal nonsusceptibility [1, 6]. They are considered as low virulent yeasts which rarely cause disease in humans [1]. Still, as far as we know, virulence factors which they harbor are hardly investigated, except in limited studies that reported secreted acid proteinase activity [12] and biofilm production [13]. There is no data on the most common fungal virulence factors such as the production of coagulase, phospholipase, and esterase; hemolysis ability; and surface hydrophobicity. The presence and expression of variable virulence factors and their contribution to pathogenesis are yet to be explained.
Reference antifungal susceptibility testing methods were standardized for Candida and Cryptococcus, and details of testing when used for Saprochaete/Magnusiomyces are not definitive [14, 15]. Besides, neither clinical breakpoints nor epidemiological cutoff values have been determined for Saprochaete/Magnusiomyces. Still, echinocandin minimum inhibitory concentration (MIC) values are high and Saprochaete/Magnusiomyces isolates are considered resistant to echinocandins in vitro and in vivo. Except for fluconazole, which yields high MICs in Saprochaete/Magnusiomyces, azole MICs are variable in individual strains [1]. Reference methods are recommended to test the antifungal susceptibility of Saprochaete/Magnusiomyces, in order to obtain reproducible and comparable MIC values [1].
In this study, we aimed to confirm species identification, reveal virulence factors, determine antifungal susceptibility of clinical Saprochaete/Magnusiomyces isolates, and contribute to the knowledge of a rare fungal pathogen.
Materials and methods
Clinical isolates
Fungal isolates obtained from clinical specimens between 2006 and 2021 at Hacettepe University Faculty of Medicine Hospitals Microbiology Laboratory were included. The hospitals are a 1200-bed tertiary-care complex with medical, surgical, and oncological intensive care units and bone-marrow and solid-organ transplantation wards and are a center for patients with chronic pulmonary diseases, patients needing immune suppressive treatment such as malignancies, rheumatological diseases, and immune disorders. The isolates were identified by colony morphology, microscopic morphology on corn meal Tween 80 medium, negative urease test, and assimilation profile detected by ID32C (BioMérieux, France) at the Mycology Laboratory. Brain heart infusion broth with 20% glycerol was used to store isolates at −20 °C. Isolates which were identified as Saprochaete, Blastoschizomyces, or Geotrichum were subcultured on Sabouraud dextrose agar (SDA; OXOID, UK) and were checked for viability, purity, colony morphology, and morphology on Corn Meal Tween 80 medium. They were selected for further tests when the identification was compatible with Saprochaete/Magnusiomyces. If multiple isolates from a single patient were available, the first available isolate from a patient was included.
Species identification
The isolates were identified using MALDITOF-MS according to the manufacturer’s recommendations (Bruker Corporation, USA). Briefly, a sample from a colony grown overnight on Sabouraud dextrose agar was placed on target, covered by 1 μl 70% formic acid, left to dry, and then covered by 1 μl matrix solution. Matrix solution was prepared by dissolving 2.5 mg α-cyano-4-hydroxycinnamic acid (HCCA, Sigma 70990, UK), 75 μl methanol, and 150 μl organic solvent (475 μl water, 500 μl acetonitrile, 25 μl trifluoroacetic acid). Target was examined using MALDIBiotyper (Bruker Corporation, USA), and scores were noted.
For sequencing, DNA was extracted using a “heat shock” method [16]. Polymerase chain reaction for ITS and Rpb2 was performed as described elsewhere [8]. Sequences of ITS region from all isolates were analyzed for species identification. In case the ITS sequence was not sufficient to differentiate the species, Rpb2 gene sequence was evaluated additionally [8]. Products were purified (PureLink PCR Purification Kit, Thermo Fisher Scientific, USA) and sequenced (BigDye Terminator v3.1 Cycle Sequencing Kit, Thermo Fisher Scientific, USA). The sequences were checked (Bioedit 7.2.5 Sequence Alignment Editor, Ibis Therapeutics, USA) and compared to National Center for Biotechnology Information (NCBI, https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome) and International Society for Human and Animal Mycology (ISHAM) databases (https://its.mycologylab.org/page/Alignment, ITS only) to determine species identification. In order to visualize species distribution, a phylogenetic tree was constructed using ITS sequences of the isolates using Molecular Evolutionary Genetics Analysis software (MEGA 11, USA) for the maximum likelihood (ML) method (Tamura et al. 2021). The corrected Akaike information criterion (AICc) was chosen to estimate the best nucleotide substitution model by jModelTest v2.1.6 (Posada 2008). Bootstrap support value was calculated using 1000 replicates for the tree.
Screening for virulence factors
Saprochaete/Magnusiomyces isolates were tested for virulence factors observed in fungal pathogens phenotypically using conventional methods. Tests were repeated three times for each isolate.
Coagulase activity was evaluated after incubation in rabbit and human plasma supplemented with EDTA after 2, 4, 6, and 24 h of incubation at 37 °C [17,18,19].
Hemolysis was evaluated on SDA supplemented with 7% sheep blood or horse blood. Plates were incubated at 37 °C in 5% CO2 for 48 h [17].
Acid proteinase was investigated on solid media containing 1% bovine serum albumin [20, 21]. Briefly, 20 g glucose, 1 g KH2PO4, and 0.5 g MgSO4 were dissolved in 800 ml water. The solution was autoclaved, cooled to 50 °C, then mixed with filter-sterilized 2 g bovine serum albumin prepared in 200 ml water, and poured into the Petri plates. A 0.5 McFarland suspension was prepared from each isolate, and 10 μl of the suspension was placed on the medium. The plates were incubated at 37 °C for 6 days. The presence of a clear lysis zone around the growing microorganism was evaluated as positive.
Phospholipase activity was tested on media containing 8% egg yolk and citric acid disodium phosphate buffer [20, 21]. A mixture of 13 g SDA,11.7 g NaCl, and 0.111 g CaCl2 was autoclaved and cooled to 60 °C, sterile solutions of 8 ml 0.1 M citric acid (C6H8O7H2O), and 8 ml 0.2 M Na2HPO4 and 20 ml homogenized egg yolk were added. pH was adjusted to 4.2, and the final solution was transferred to sterile plates. Ten μl from a fungal suspension of 0.5 McFarland density was inoculated, and the plates were evaluated after incubation at 37 °C for 4 days. The presence of precipitation zones was sought for positive activity.
Esterase activity was observed on Tween 80 agar. A mixture of 10 g peptone, 5 g NaCl, 0.1 g CaCl2, and 1 L of distilled water was autoclaved. Separately autoclaved 5 ml Tween 80 was added to this mixture at 50 °C and adjusted to pH = 6.8. After inoculation, plates were incubated at 30 °C for 10 days [21, 22]. Precipitation around the inoculum was a positive result.
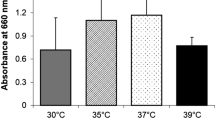
Surface hydrophobicity was tested by hydrocarbon adhesion method [21]. A solution of each isolate was prepared in PUM (phosphate, urea, MgSO4) buffer and adjusted to 1 × 107 cfu/ml, and 2 ml of the solution was transferred to 2 tubes. The first was used as a control, and 0.5 ml n-hexadecane was added to the latter. The tubes were incubated in a 37 °C water bath for 10 min, were vortexed, and were incubated for a further 30 min. A sample from the lower aqueous phase was obtained via a pipette, absorbance was measured at 660 nm, and OD values of two tubes were compared.
Biofilm formation of the isolates was investigated. Biofilm biomass was detected using crystal violet (CV), and metabolic activity was explored using 2,3-bis-(2-methoxy-4-nitro-5-sulfophenyl)-2h-tetrazolium-5-carboxanilide (XTT) [23, 24]. Biofilm-forming ability of each isolate was interpreted as negative or positive (weak, intermediate, or strong) according to OD values compared to negative controls [25]. In addition, OD values of the isolates were compared to and were given as percentage of the activity of biofilm positive reference strain (Candida albicans MYA-274) which was accepted as 100%.
In vitro antifungal susceptibility testing
Minimum inhibitory concentration (MIC) values of Saprochaete/Magnusiomyces isolates against amphotericin B, fluconazole, voriconazole, and micafungin were determined using the Clinical and Laboratory Standards Institute (CLSI) and European Committee on Antimicrobial Susceptibility Testing (EUCAST) reference microdilution methods [14, 15]. For each isolate, inoculum was prepared in 5 ml saline containing 200 μl Tween 20 (Biomatik, Canada). Cell counts were performed under the microscope for the appropriate concentration, and the percent transmission of light for the suspension was measured at 530 nm. MIC values were noted after 24 and 48 h of incubation. Quality control strains Candida krusei ATCC 6258 and Candida parapsilosis ATCC 22019 were used in each run as recommended.
Results
Identification of isolates
A total of 57 isolates were included in the study. Of these, 53 were identified as S. capitata, and four were identified as S. clavata using conventional methods and ID 32C.
Identification scores obtained by MALDITOF-MS were ≥ 2 for 54 (S. capitata n = 52, S. clavata n = 2) and between 2 and 1.7 for two (S. capitata n = 1, S. clavata n = 1) isolates. Only one S. clavata isolate yielded scores below 1.70, with no other genera/species identification options given by the system.
ITS sequences evaluated in BLAST and ISHAM databases confirmed the genus and species identification for all Saprochaete isolates. For the four S. clavata isolates, ITS gene sequences also had high scores for S. capitata, especially in the BLAST database (“percent identity” scores (> 99%). Therefore, Rpb2 sequences were also investigated for reliable differentiation of S. clavata which confirmed the identification. Moreover, the phylogenetic tree of ITS sequences showed these four S. clavata isolates clustered separately from the other isolates, which were identified as S. capitata (Supplementary Figure 1).
Virulence factors
Coagulase, hemolysis, acid proteinase, and phospholipase were negative for all Saprochaete isolates using phenotypic tests under given conditions.
Esterase activity was positive for 32 (56.1%) isolates (Supplementary Figure 2). Of these, 27 were S. capitata, and three were S. clavata.
Surface hydrophobicity was intermediate for 11 (19.3%) and weak for 46 (80.7%) isolates. All 11 isolates with intermediate surface hydrophobicity were S. capitata.
To evaluate biofilm-forming ability, CV was used to detect biomass formation, and XTT was used to understand metabolic activity. All isolates showed strong biofilm-forming ability with both methods when OD values were compared to negative controls. In order to accomplish a more detailed classification, OD values of the isolates were compared with the OD value of the biofilm-positive reference strain, C. albicans MYA-274, accepting the CV or XTT OD value of the latter as 100%. Metabolic activity was mostly lower than the reference strain, as the majority of the biofilms (n = 41, 71.9%) had a metabolic activity at 26–50%. However, biomass produced by 23 (40.4%) Saprochaete/Magnusiomyces isolates was higher than C. albicans MYA-274 (Table 1).
Antifungal susceptibility testing
Fungal suspensions consisting of 2–5 × 106 cells/ml were prepared using cell counts under the microscope, which revealed variable morphologies for Saprochaete/Magnusiomyces cells (Supplementary Figure 3). The percent transmittance of light for the solutions containing appropriate numbers of cells ranged between 62.0 and 65.8%, which was lower than Candida solutions of 0.5 McFarland (expected transmittance 80-82%) [26].
For Saprochaete/Magnusiomyces isolates included in the study, MIC values of amphotericin B, fluconazole, voriconazole, and micafungin obtained using both CLSI and EUCAST methods were noted after 24 and 48 h of incubation. For the CLSI method, 24 h incubation was insufficient, and 48 h results were used. The EUCAST method recommends an OD value of ≥ 0.200 in the growth control well for sufficient growth [15]. Only 3 of the 57 isolates reached sufficient growth after 24 h incubation. Except for these three, 48 h results were taken into account for the 54 isolates. MIC results are summarized in Table 2.
MIC values obtained by the EUCAST method were within 2 double-dilutions lower in 5 isolates (8.8%) for amphotericin B and in 1 isolate (1.8%) for micafungin, compared with those obtained by the CLSI method. For fluconazole, voriconazole, and micafungin, 56.1%, 84.2%, and 66.7% of the isolates had ≥ 2 double-dilution higher MIC values with EUCAST compared to CLSI MICs, respectively. For the rest of the isolates, the MICs obtained by the EUCAST and CLSI methods were distributed in the ±1 double-dilution range.
Discussion
Identification of Saprochaete isolates to genus level using macroscopic and microscopic morphology is applicable. However, species-level identification needs more tests, which might be a challenge for clinical laboratories. Isolates included in our study could successfully be identified to species level using a commercial kit based on biochemical reactions including carbohydrate assimilation tests (ID32C, BioMerieux, France) and MALDI-TOF MS, both of which might be available in routine clinical microbiology laboratories. Different commercial systems based on biochemical reactions are available in clinical laboratories for fungal identification. ID32C was successfully used to identify S. capitata [7] and failed to identify S. clavata [27] in previous studies, which used sequencing for confirmation of the identification. Nevertheless, all isolates included in our study were successfully identified at the species level, including four S. clavata isolates. Other commercial biochemical tests such as VITEK2 (BioMerieux, France), API20C (BioMerieux, France), and AuxaColor (BioRad, France) were also used to identify Saprochaete/Magnusiomyces with variable performances [28].
MALDI-TOF MS has been a promising method for microbiological identification and, with the expansion of the databases, might differentiate fungi to species level successfully, including Saprochaete/Magnusiomyces [3, 6, 7, 29]. The MALDI-TOF MS system used in this study (Bruker Diagnostics, USA) recommends a score of ≥ 2.0 for reliable species identification, states that a score of < 2.0–1.7 is reliable for genus identification, and considers scores < 1.7 as unreliable. Although MALDI-TOF MS generated a low score (< 1.7) for one of the S. clavata isolates, the database gave no other identification options. Kolecka et al. [29] also reported a S. clavata isolate with a MALDI-TOF MS identification score of < 1.7, for which the identification was confirmed with ITS sequencing. Improving the database using spectra from strains of confirmed identification might still be needed to eliminate doubt for Saprochaete/Magnusiomyces species.
In this study, ITS sequences were used to confirm the identification of Saprochaete/Magnusiomyces isolates, using two different databases. S. capitata isolates were identified without hesitation, as sequences yielded high scores in both databases. However, ITS sequences of four S. clavata isolates had low scores and/or high scores of both S. clavata and S. capitata. ITS sequences of these two species show high similarity (96%) which might cause misidentifications. Additional analysis of Rpb2 sequence, which was found to be the best marker for differentiation among other candidates such as LSU, Tef1α, and Act previously, was used in our study [8]. All four isolates were identified as S. clavata when Rpb2 sequences were compared in the BLAST database. In order to support the identification, a phylogenetic tree was built using ITS sequences, and the four S. clavata isolates clustered separately.
Fungal taxonomy and nomenclature are very dynamic research areas with massive current changes. Different genus and species names were proposed and used for Saprochaete/Magnusiomyces in the past decades [1, 8, 9]. Accurate identification of clinically important species in the routine microbiology laboratories is critical to reveal the real extent of the problem caused by infectious agents. S. capitata and S. clavata were described as the most common clinical isolates causing human infection, and we did not detect any other species in our culture collection.
Saprochaete/Magnusiomyces might cause infections with high mortality, even in immunocompetent hosts [1, 2]. Still, their virulence factors are yet to be described. For a first step, we looked for common fungal virulence factors via phenotypic methods. This is a limited approach as the results might vary according to test conditions and negative results do not rule out the presence of the related virulence factor. Nonetheless, when found positive, further studies might be performed to find out the related genes and understand under which conditions they are produced and how they affect pathogenesis. This study checked coagulase, haemolysis, acid proteinase, and phospholipase which were not detectable in any of the isolates with the method used. Esterase positivity was detected Saprochaete/Magnusiomyces isolates for the first time. In addition, strong biofilm biomass formation was documented.
Coagulase activity is affected by the type of plasma. Coagulase positivity was reported for Candida species, with variations in rabbit, sheep, and human plasma [17,18,19]. For Trichosporon, 5.0% and 27.5% coagulase positivity was detected using human and rabbit plasma, respectively [30]. We used both rabbit and human plasma and evaluated at different time periods but did not observe positivity in Saprochaete/Magnusiomyces isolates.
Hemolytic activity of Saprochaete/Magnusiomyces isolates was negative on SDA supplemented with 7% sheep or horse blood after 48 h. Variable rates of hemolysis were detected in Candida isolates [17]. Although Demir et al. [30] observed hemolysis after 48 of incubation in Trichosporon, Sun et al. [31] needed to extend the time period to 96 h. Therefore, extended incubation periods might alter test results for Saprochaete/Magnusiomyces, which was a limitation of our study.
Acid proteinases and phospholipases could not be detected in Saprochaete/Magnusiomyces isolates in this study. They are important virulence factors for Candida with high positivity rates [17, 18]. In another rare yeast, Trichosporon, both proteinase and phospholipase activities were negative in different studies [21, 30,31,32]. However, Pontieri et al. [12] found secreted acid protease in 3 of the 25 S. capitata isolates they tested on BSA agar and confirmed the finding by showing the hydrolysis of bovine serum hemoglobin by a spectrophotometer. Although we could not detect acid proteinase positivity, the method used was different from that of Pontieri et al.
Unlike other virulence enzymes, esterase was detected in 56.1% of the Saprochaete/Magnusiomyces isolates tested. Esterase positivity is common among Candida species and might be positive in almost all C. albicans isolates [22]. Moreover, several studies stated 100% esterase positivity in Trichosporon [30, 32]. Understanding the molecular basis of esterase production and its contribution to pathogenesis needs further studies for Saprochaete/Magnusiomyces.
Surface hydrophobicity is an important factor which affects cell-cell and cell-surface interactions and may alter virulence in medically-significant fungi. As it is directly correlated to adhesion, a shift in biofilm formation ability might be expected, but this output is not confirmed [33]. Of the Saprochaete/Magnusiomyces isolates tested, 11 S. capitata isolated showed intermediate levels of surface hydrophobicity, with remaining isolates showing low levels. Still, high biofilm-forming ability was present in all isolates, and the effect of surface hydrophobicity on biofilms of Saprochaete/Magnusiomyces is yet to be investigated.
Biofilm-forming ability could be demonstrated in all Saprochaete/Magnusiomyces isolates all of which showed strong biomass production and metabolic activity using the microplate method. Furthermore, biomass production was higher than the biofilm-positive reference strain C. albicans MYA-274 in 40.4% of the isolates. Metabolic activity was mostly slower, with 75.5% of the isolates showing an activity about 25–50% of the positive control, which might be caused by the difference between Candida and Saprochaete/Magnusiomyces. The growth rate of the latter is generally slower, which might indicate a slower metabolic activity. Biofilm production is an important virulence factor associated with adherence and persistence on surfaces, which contributes to foreign body infections and nosocomial outbreaks. ElGindi et al. [34] speculated that the site of infection is more important than the genus/species of the arthroconidial fungi to predict biofilm-forming capacity. They investigated biofilm formation of clinical and reference strains of yeasts including S. capitata CBC 197.35 under fluid flow and commented that the biofilm composition had more filamentous cells, increasing biofilm mass. Moreover, filamentous cells of S. capitata (and Trichosporon asahii) could resist shear forces due to maintained adherence. Dantonio et al. [13] observed large amounts of slime on the inner lumen of extracted catheters in six S. capitata clinical isolates obtained from catheter-related fungemia via scanning electron microscopy (SEM). Kraft et al. [35] visualized the biofilm formation of a S. clavata on a titanium-alloy device using SEM. Management strategies for rare yeast infections should be guided by the clinical characteristics and might include options for removing biofilm-harboring tissue and foreign bodies [1, 34]. Removal of all foreign bodies including central venous access devices is recommended for the management of Saprochaete/Magnusiomyces infections [1].
We determined MIC values of Saprochaete/Magnusiomyces isolates against amphotericin B, fluconazole, voriconazole, and micafungin using two reference microdilution methods. Specific instructions for antifungal susceptibility testing of Saprochaete/Magnusiomyces are not available [14, 15]. Still, reference methods developed primarily for Candida and Cryptococcus are preferred to obtain reliable and reproducible MIC results [1, 14, 15]. An inoculum of 1–5 × 106 cell/ml, prepared according to 0.5 McFarland standard, is recommended for Candida and Cryptococcus for both methods. However, when we prepared Saprochaete/Magnusiomyces 0.5 McFarland solutions of 0.08–0.13 optical density and 80–82% transmittance as recommended for Candida [26], we could not reach the recommended cell concentrations. Therefore, we ensured the solutions had the required amount of yeast cells by having cell counts and saw that these inocula ranged at 62–66% transmittance. Noster et al. [36] observed that the viable colony counts in inoculum solutions were below the recommended concentration and adjusted them to 0.75 McFarland for antifungal susceptibility testing of Saprochaete/Magnusiomyces using the EUCAST method. The need to standardize spectrophotometric measurements to obtain desired cell concentrations was addressed and applied to filamentous fungi previously [37]. As Saprochaete/Magnusiomyces cells have variable morphology and volume (Supplementary Figure 3), recommendations for Candida may not be appropriate, and further guidance might be needed.
When reading MIC values of Saprochaete/Magnusiomyces isolates, despite higher inocula were used, growth was mostly not sufficient in control wells after 24 h of incubation. For testing Candida and Cryptococcus, the CLSI method encourages reading at 24 h but extends incubation if growth in the control well is not sufficient [14]. The EUCAST method recommends reading after 24 ± 2 h of incubation but states that OD values should exceed 0.2 in the control well before determining MICs [15]. Previous studies also reported MICs after 48 h [5, 8, 38], even if a higher inoculum was used [36]. Achieving sufficient growth in the control wells might pose a problem when determining MICs for Saprochaete/Magnusiomyces.
Since clinical breakpoints and epidemiological cutoffs are not available for Saprochaete/Magnusiomyces, we only provided MIC values [39, 40]. Determination of MIC values using a reference method is recommended to decide the best management options and accumulate epidemiological data as the susceptibility of individual strains might vary [1]. CLSI and EUCAST microdilution methods mostly give compatible results. In our study, EUCAST MICs had a tendency to be higher than CLSI MICs (except amphotericin B), although variations existed. Fernandez-Ruiz et al. [38] tested three S. capitata isolates using CLSI and EUCAST, finding higher MICs with EUCAST, except amphotericin B. Esposto et al. [41] compared EUCAST, CLSI, and Sensititre and reported that MIC50 and MIC90 values were within one double-dilution. Method-specific breakpoints are recommended to avoid variations of MICs to evaluate the susceptibility of fungal isolates.
For the Saprochaete/Magnusiomyces isolates tested, amphotericin B MIC values ranged between 0.5 and 2 μg/ml. Most studies found amphotericin B MICs between 0.5 and 2 μg/ml [1]. But MICs as low as 0.06 μg/ml using CLSI [8] and EUCAST [36] and as high as 8 μg/ml using EUCAST [42] were reported using reference methods in different studies.
Fluconazole MICs were high in Saprochaete/Magnusiomyces isolates and were between 0.25 and 16 and 0.5 and 64 μg/ml with CLSI and EUCAST, respectively. High fluconazole MICs were reported in the literature for Saprochaete/Magnusiomyces, which mostly ranged between 16 and 32 μg/ml [1, 6, 8, 38, 41]. Still, high doses of fluconazole might be used to treat infections caused by isolates with low fluconazole MICs [1].
Voriconazole MICs were variable for the Saprochaete/Magnusiomyces isolates in the study. Although GM was 0.94 μg/ml with EUCAST, a MIC value of > 8 μg/ml was detected in a S. clavata isolate, and MICs of 4 μg/ml were detected in six S. capitata isolates. CLSI MICs were lower, ranging between 0.016 and 1 μg/ml. Fernandez-Ruiz et al. [38] also detected higher VOR MICs with EUCAST than CLSI, reaching 16 μg/ml in one S. capitata isolate. VOR MICs are usually between 0.03 and 0.5 μg/ml in S. capitata [1], and the detection of higher MICs in individual isolates is disturbing.
As expected, Saprochaete/Magnusiomyces isolates yielded high micafungin MICs. The lowest micafungin MIC was 0.5 μg/ml (EUCAST) at 48 h in a S. clavata isolate, which had a MIC of 0.25 μg/ml at 24 h with insufficient growth. High Saprochaete/Magnusiomyces echinocandin MICs were also reported in the literature [1, 8, 36]. Echinocandins are not an option for the treatment of Saprochaete/Magnusiomyces infections [1]. These results confirmed that this group of antifungals did not have sufficient activity against Saprochaete/Magnusiomyces, even in vitro. Individual isolates with low echinocandin (micafungin, caspofungin) MICs were detected in different studies using reference methods [1, 5, 36]. Poor growth might pose a problem when evaluating echinocandin MIC values, as reading at 24 h as recommended might not be reliable due to insufficient growth [14, 15]. Since Saprochaete/Magnusiomyces are known to have reduced susceptibility to echinocandins, if encountered, low MICs should be approached with caution and should not be considered as a potential susceptibility.
Saprochaete/Magnusiomyces species are rare human pathogens; therefore, the data on these microorganisms are limited. Recent advances in molecular taxonomy lead to nomenclature changes, and identification in routine microbiology laboratories was revisited. In this study, (1) clinical isolates included in the study might successfully be identified to species level with both MALDI-TOF MS and ID32C supported with conventional tests, as confirmed by sequencing. (2) Common fungal virulence factors were investigated via phenotypic methods as a first step to understand the pathogenicity of these opportunistic fungi. Coagulase, hemolytic activity, acid protease, and phospholipase could not be detected. However, more than half of the isolates were esterase positive. Surface hydrophobicity was weak in most isolates, but all of them formed biofilms with high biomass production. These results might change according to test conditions but might be a beginning to understand the pathogenesis of Saprochaete/Magnusiomyces. (3) Antifungal susceptibility testing methodology needs to be standardized for Saprochaete/Magnusiomyces, as spectrophotometric inoculum preparation according to 0.5 McFarland was not adequate and higher (transmittance 62–66%) was suitable. In addition, incubation for 24 h rarely yielded sufficient growth. As MIC values obtained using CLSI and EUCAST were variable, the need to establish method-based epidemiological cutoff values and clinical breakpoints stands.
Data Availability
The datasets generated during and/or analysed during the current study are available at Turkish High Education Council Thesis Center (https://tez.yok.gov.tr/UlusalTezMerkezi/giris.jsp, No: 752412) and from the corresponding author on reasonable request.
References
Chen SC, Perfect J, Colombo AL et al (2021) Global guideline for the diagnosis and management of rare yeast infections: an initiative of the ECMM in cooperation with ISHAM and ASM. Lancet Infect Dis 21:e375–e386. https://doi.org/10.1016/S1473-3099(21)00203-6
Tanabe MB, Patel SA (2018) Blastoschizomyces capitatus pulmonary infections in immunocompetent patients: case report, case series and literature review. Epidemiol Infect 146:58–64. https://doi.org/10.1017/S0950268817002643
Menu E, Criscuolo A, Desnos-Ollivier M et al (2020) Saprochaete clavata outbreak infecting cancer center through dishwasher. Emerg Infect Dis 26:2031–2038. https://doi.org/10.3201/eid2609.200341
Gurgui M, Sanchez F, March F et al (2011) Nosocomial outbreak of Blastoschizomyces capitatus associated with contaminated milk in a haematological unit. J Hosp Infect 78:274–278. https://doi.org/10.1016/j.jhin.2011.01.027
Subramanya Supram H, Gokhale S, Chakrabarti A et al (2016) Emergence of Magnusiomyces capitatus infections in Western Nepal. Med Mycol 54:103–110. https://doi.org/10.1093/mmy/myv075
Duran Graeff L, Seidel D, Vehreschild MJ et al (2017) Invasive infections due to Saprochaete and Geotrichum species: report of 23 cases from the FungiScope Registry. Mycoses 60:273–279. https://doi.org/10.1111/myc.12595
Hazırolan G, Aypak A, Aksu N (2017) An unusual case of urinary tract infection caused by Saprochaete capitata under anidulafungin treatment. J Mycol Med 27:387–390. https://doi.org/10.1016/j.mycmed.2017.04.001
Kaplan E, Al-Hatmi AMS, Ilkit M et al (2018) Molecular diagnostics of arthroconidial yeasts, frequent pulmonary opportunists. J Clin Microbiol:56. https://doi.org/10.1128/JCM.01427-17
de Hoog GS, Smith MT (2004) Ribosomal gene phylogeny and species delimitation in Geotrichum and its teleomorphs. Stud Mycol 50:489–516
Desnos-Ollivier M, Blanc C, Garcia-Hermoso D et al (2014) Misidentification of Saprochaete clavata as Magnusiomyces capitatus in clinical isolates: utility of internal transcribed spacer sequencing and matrix-assisted laser desorption ionization-time of flight mass spectrometry and importance of reliable databases. J Clin Microbiol 52:2196–2198. https://doi.org/10.1128/JCM.00039-14
Schoch CL, Seifert KA, Huhndorf S et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Pontieri E, Caracciolo C, Ceddia T et al (2005) Genetic variability among Blastoschizomyces capitatus isolates from different clinical sources. Int J Immunopathol Pharmacol 18:531–539. https://doi.org/10.1177/039463200501800313
D'Antonio D, Parruti G, Pontieri E et al (2004) Slime production by clinical isolates of Blastoschizomyces capitatus from patients with hematological malignancies and catheter-related fungemia. Eur J Clin Microbiol Infect Dis 23:787–789. https://doi.org/10.1007/s10096-004-1207-4
CLSI. CLSI Document M27A (2022) Clinical and Laboratory Standards Institute. Reference method for broth dilution antifungal susceptibility testing of yeasts, 4th edn. Clinical and Laboratory Standards Institute, Wayne, PA
EUCAST. The European Committee on Antimicrobial Susceptibility Testing. 2020. EUCAST Definitive Document E.DEF 7.3.2 Method for the determination of broth dilution minimum inhibitory concentrations of antifungal agents for yeasts. Accessed 20.09.2023; Available from: http://www.eucast.org/astoffungi/clinicalbreakpointsforantifungals/
Lockhart SR, Messer SA, Pfaller MA et al (2008) Geographic distribution and antifungal susceptibility of the newly described species Candida orthopsilosis and Candida metapsilosis in comparison to the closely related species Candida parapsilosis. J Clin Microbiol 46:2659–2664. https://doi.org/10.1128/JCM.00803-08
Al-Dabbagh AAH, Ajah HA, Salman JAS (2023) Detection of virulence factors from Candida spp. isolated from oral and vaginal candidiasis in Iraqi patients. Arch Razi Inst 78:465–474. https://doi.org/10.22092/ARI.2022.359464.2420
Jafari M, Salari S, Pakshir K et al (2017) Exoenzyme activity and possibility identification of Candida dubliniensis among Candida albicans species isolated from vaginal candidiasis. Microb Pathog 110:73–77. https://doi.org/10.1016/j.micpath.2017.06.024
Yiğit N, Aktas AE, Ayyildiz A (2008) Detection of coagulase activity in pathogenic Candida species. J Int Med Res 36:1378–1382. https://doi.org/10.1177/147323000803600627
Alp S, Arikan S (2008) Investigation of extracellular elastase, acid proteinase and phospholipase activities as putative virulence factors in clinical isolates of Aspergillus species. J Basic Microbiol 48:331–337. https://doi.org/10.1002/jobm.200700349
Dağ A, Çerikçioğlu N (2006) Investigation of some virulence factors of Trichosporon asahii strains isolated from the clinical samples of hospitalized patients. Mikrobiyol Bul 40:225–235
Slifkin M (2000) Tween 80 opacity test responses of various Candida species. J Clin Microbiol 38:4626–4628. https://doi.org/10.1128/JCM.38.12.4626-4628.2000
Melo AS, Bizerra FC, Freymuller E et al (2011) Biofilm production and evaluation of antifungal susceptibility amongst clinical Candida spp. isolates, including strains of the Candida parapsilosis complex. Med Mycol 49:253–262. https://doi.org/10.3109/13693786.2010.530032
Pierce CG, Uppuluri P, Tummala S et al (2010) A 96 well microtiter plate-based method for monitoring formation and antifungal susceptibility testing of Candida albicans biofilms. J Vis Exp. https://doi.org/10.3791/2287
Aslan H, Gulmez D (2016) Investigation of the correlation between biofilm forming ability of urinary Candida isolates with the use of urinary catheters and change of antifungal susceptibility in the presence of biofilm. Mikrobiyol Bul 50:256–265. https://doi.org/10.5578/mb.24248
CDC. OFD-500-P03 antifungal susceptibility testing yeasts using gradient diffusion strips. 2020. Accessed 10.09.2023; Available from: https://www.cdc.gov/fungal/lab-professionals/afst-yeasts.html
Aydın M, Kuştimur S, Kalkancı A et al (2019) Identification of medically important yeasts by sequence analysis of the internal transcribed spacer and D1/D2 region of the large ribosomal subunit. Rev Iberoam Micol 36:129–138. https://doi.org/10.1016/j.riam.2019.05.002
Posteraro B, Efremov L, Leoncini E et al (2015) Are the conventional commercial yeast identification methods still helpful in the era of new clinical microbiology diagnostics? A meta-analysis of their accuracy. J Clin Microbiol 53:2439–2450. https://doi.org/10.1128/JCM.00802-15
Kolecka A, Khayhan K, Groenewald M et al (2013) Identification of medically relevant species of arthroconidial yeasts by use of matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol 51:2491–2500. https://doi.org/10.1128/Jcm.00470-13
Demir F, Kuştimur S (2014) Investigation of some virulence factors in Trichosporon spp. strains. Mikrobiyol Bul 48:628–638
Sun W, Su J, Xu S et al (2012) Trichosporon asahii causing nosocomial urinary tract infections in intensive care unit patients: genotypes, virulence factors and antifungal susceptibility testing. J Med Microbiol 61:1750–1757. https://doi.org/10.1099/jmm.0.049817-0
Hazırolan G, Koçak N, Karagöz A (2018) Sequence-based identification, genotyping and virulence factors of Trichosporon asahii strains isolated from urine samples of hospitalized patients (2011-2016). J Mycol Med 28:452–456. https://doi.org/10.1016/j.mycmed.2018.06.006
Danchik C, Casadevall A (2020) Role of cell surface hydrophobicity in the pathogenesis of medically-significant fungi. Front Cell Infect Microbiol 10:594973. https://doi.org/10.3389/fcimb.2020.594973
ElGindi M, Al-Baghdadi R, Jackman AB et al (2021) Where the infection is isolated rather than the specific species correlates with adherence strength, whereas biofilm density remains static in clinically isolated Candida and arthroconidial yeasts. Can J Microbiol 67:497–505. https://doi.org/10.1139/cjm-2020-0215
Kraft L, Ribeiro VST, Petroski LP et al (2023) Saprochaete clavata invasive infection: characterization, antifungal susceptibility, and biofilm evaluation of a rare yeast isolated in Brazil. Rev Inst Med Trop Sao Paulo 65:e12. https://doi.org/10.1590/S1678-9946202365012
Noster J, Koeppel MB, Desnos-Olivier M et al (2022) Bloodstream infections caused by Magnusiomyces capitatus and Magnusiomyces clavatus: epidemiological, clinical, and microbiological features of two emerging yeast species. Antimicrob Agents Chemother 66:e0183421. https://doi.org/10.1128/AAC.01834-21
CLSI. CLSI Document M38-A (2002) Reference method for broth dilution antifungal susceptibility testing of filamentous fungi, approved standard-first edn. Clinical and Laboratory Standards Institute/National Committee for Clinical Laboratory Standards, Wayne, PA
Fernandez-Ruiz M, Guinea J, Puig-Asensio M et al (2017) Fungemia due to rare opportunistic yeasts: data from a population-based surveillance in Spain. Med Mycol 55:125–136. https://doi.org/10.1093/mmy/myw055
EUCAST. The European Committee on Antimicrobial Susceptibility Testing. 2020. Breakpoint tables for interpretation of MICs for antifungal agents Version 10.0. Accessed; Available from: http://www.eucast.org/astoffungi/clinicalbreakpointsforantifungals/
CLSI. CLSI Document M27M44S (2022) Clinical and Laboratory Standards Institute, Performance standards for antifungal susceptibility testing of yeasts, 3rd edn. Clinical and Laboratory Standards Institute, Wayne, PA
Esposto MC, Prigitano A, Lo Cascio G et al (2019) Yeast-like filamentous fungi: molecular identification and in vitro susceptibility study. Med Mycol 57:909–913. https://doi.org/10.1093/mmy/myy133
Cuenca-Estrella M, Gomez-Lopez A, Mellado E et al (2006) Head-to-head comparison of the activities of currently available antifungal agents against 3,378 Spanish clinical isolates of yeasts and filamentous fungi. Antimicrob Agents Chemother 50:917–921. https://doi.org/10.1128/AAC.50.3.917-921.2006
Acknowledgements
The authors would like to thank Prof. Dr. Sevtap Arikan-Akdagli for her constructive comments and Dr. Ceylan Polat for her help in improving Figure 1.
Funding
This study was supported by the Hacettepe University Scientific Research Unit (Project Code: THD-2022-19827).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation and data collection were performed by Beyzanur Kaşaltı, and analysis was performed by Beyzanur Kaşaltı and Dolunay Gülmez. The first draft of the manuscript was written by Beyzanur Kaşaltı and finalized by Dolunay Gülmez. All authors commented on previous and final versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Rosana Puccia
Part of this work was presented in the 37th ANKEM Rational Antibiotic Use Congress, 26–30th October 2022, Antalya, Türkiye.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kaşaltı, B., Gülmez, D. Saprochaete/Magnusiomyces: identification, virulence factors, and antifungal susceptibility of a challenging rare yeast. Braz J Microbiol 55, 41–49 (2024). https://doi.org/10.1007/s42770-024-01248-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-024-01248-7