Abstract
The objective of this study was to evaluate the antifungal activity of free methyl 3,5 dinitrobenzoate (MDNB) and its nanoemulsion (MDNB-NE) against strains of Candida albicans. Additionally, a molecular modeling study was also carried out to propose the mechanism of action and toxicity of MDNB. These results demonstrated the MDNB-NE presented a droplet size of 181.16 ± 3.20 nm and polydispersity index of 0.30 ± 0.03. MDNB and MDNB-NE inhibited the growth of all strains with minimum inhibitory concentrations of 0.27–1.10 mM. The biological results corroborated the molecular model, which pointed to a multi-target antifungal mechanism of action for MDNB in C. albicans. The study could serve as a basis for further research involving compounds with nitro groups with antifungal.
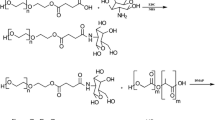
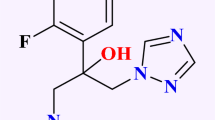
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungal infections caused by Candida species are a major health problem, resulting in high mortality and medical costs [1]. Among these strains, Candida albicans is often found in clinical contexts and is the principal cause of candidiasis, which represents one of the main causes of hospital infections [1, 2]. In immunocompromised patients, C. albicans can cause systemic invasive candidiasis, affecting the circulatory system, bones, and brain [3].
Candidiasis is treated with different classes of drugs, including echinocandins, azoles, polyenes, and 5-flucytosine, which can act as fungistatics or fungicides [4, 5]. However, the increasing number of candidiasis cases, toxicity of these drugs, and increase in the number of infections caused by resistant strains have expanded the search for new antifungal agents [6, 7]. In this context, organic synthesis represents an important strategy for the development of highly effective antifungals [8, 9]. Benzoic acid and its derivatives are biologically important compounds used as antibacterial agents and antifungal preservatives in cosmetics and medicine [10,11,12,13]. Among them, 3,5-dinitrobenzoic acid is an important intermediate in the pharmaceutical industry, with electron-withdrawing groups such as nitro groups [14]. The nitro moiety is an important pharmacophore for molecules with antimicrobial activity, and several other pharmacological activities have also been attributed to the presence of this group [15,16,17].
Previous studies have shown that some nitro compounds derived from cinnamic acid and benzoic acid have antifungal activity against strains of the genus Candida [18, 19]. Several studies have also verified the bioactivity of esters derived from benzoic acid against strains of the genus Candida, and our group has shown that esters derived from 3,5-dinitrobenzoic acid show greater antifungal activity and potency than the starting material, 3,5-dinitrobenzoic acid [11, 20,21,22].
Furthermore, nanotechnology has become a significant tool in the treatment of these infections, more efficiently exploiting the pharmacological properties of new molecules [6]. Nanoemulsions are colloidal dispersions of two immiscible liquids with the aid of a surfactant or co-surfactant mixture, with an average diameter on the nanometer scale. They have several advantages such as a controlled absorption rate, better drug stability, and improved efficacy [23,24,25].
Therefore, the objective of this study was to evaluate the antifungal activity of free methyl 3,5 dinitrobenzoate (MDNB) and its nanoemulsion (MDNB-NE) against strains of Candida albicans. Additionally, a molecular modeling study was also carried out to propose the mechanism of action and toxicity of MDNB.
Materials and methods
Chemical characterization and reagents
All reagents required for the chemical synthesis were obtained from Sigma-Aldrich. Infrared spectra were obtained using Fourier transform infrared (FTIR) spectrophotometry (Cary 630 FTIR Agilent Technologies®) and a potassium bromide (KBr) pill, with the frequency measurements in cm−1. 1H-NMR (400 MHz) and 13C-NMR (100 MHz) spectra were recorded on a BRUKER-ASCEND spectrometer. Chemical shifts (δ) were expressed in parts per million (ppm), using tetramethylsilane (TMS) as an internal standard. The spin multiplicities are given as s (singlet), d (doublet), and t (triplet). Reaction monitoring and product analysis were performed via thin layer analytical chromatography (TLC) (silica gel 60 F254) and ultraviolet light visualization using two wavelengths (254 and 366 nm).
DTA curves were obtained using a Shimadzu thermal analyzer (DSC-50) with a 5 mg sample in an alumina crucible under a nitrogen atmosphere with a flow rate of 50 mL min−1. The experiment was performed in a temperature range of 25–450 °C, at a heating rate of 10 °C min−1. The thermogravimetric curve was obtained using the Shimadzu (DSC-50) thermal analyzer with a 5 mg sample in an alumina crucible under a nitrogen atmosphere at a flow rate of 50 mL min−1. This experiment was carried out in a temperature range of 25–900 °C, at a heating rate of 10 °C min−1.
General procedure for synthesis of methyl 3,5-dinitrobenzoate (MDNB)
A mixture of 3,5-dinitrobenzoic acid (0.100 g; 0.47 mmol) and methanol (20 mL) was heated under reflux in the presence of sulfuric acid (0.025 mL) until the completion of the reaction (3 h), monitored by TLC. After the reaction, the solvent was partially removed under reduced pressure, and the solution was diluted with 20 mL of water. The obtained product was extracted using ethyl acetate (15 mL). The organic phase was treated with NaHCO3 (5% w/v) and then with water, dried with Na2SO4, filtered, and evaporated to give MDNB [26]. Spectroscopic data for the MDNB in this study are available in the Supplementary Information.
Preparation of MDNB nanoemulsions
To prepare MDNB-NE, 0.25 g of methyl 3,5-dinitrobenzoate was previously solubilized in ethanol and add to 0.5 g of Miglyol 812®. After that, the surfactants (0.12 g Tween 80® + 0.08 g Span 80®) were added to the oil phase. Finally, distilled water (9.3 g) was added to the resulting mixture. The formulations were prepared by ultrasonication, and the components were subjected to three homogenization cycles in a sonication apparatus (model QE200, Ultronique, Brazil) for 1 min at 300 W in an ice bath, followed by an ultrasonic bath for 1 min [27]. For the unloaded nanoemulsion (BNE), MDNB was replaced with Miglyol 812®.
Determination of methyl 3,5-dinitrobenzoate content in nanoemulsion
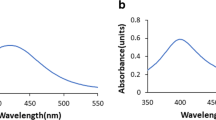
The MDNB content in the nanoemulsion was measured using ultraviolet–visible spectrophotometry (Genesys 10S UV–Vis; Thermo Fisher Scientific, Germany). First, the maximum absorption wavelength of methyl 3,5-dinitrobenzoate in dimethyl sulfoxide was determined (λmax = 260 nm). The concentration of MDNB in the nanoemulsion was calculated using a calibration curve ((n = 8) y = 0.0375x – 0.0034; R2 = 0.9991). MDNB-NE was diluted in dimethyl sulfoxide (1:2000) and filtered using a 0.2 μm membrane filter [28] before the assay. All the measurements were performed in triplicate.
Characterization of nanoemulsion
The particle size and polydispersity index (PDI) were determined with a Zetasizer instrument (Nano ZS, Malvern Instruments, UK) using the dynamic light scattering technique at 25 °C. Measurements of the mean droplet (Z-average) diameter and size distribution were performed at a fixed scattering angle of 173°. The nanoemulsions were diluted with deionized water (1:25, v/v) for characterization. All determinations were performed in triplicate, and the results are presented as means with standard deviations ( ±) [29].
Stability study
The MDNB-NEs were kept in 10 mL glass vials sealed with plastic lids and kept in a refrigerator at 4 ± 2 °C. Their stability was evaluated for 90 days. The sampling times were 1, 7, 15, 45, 60, and 90 days. The formulations were analyzed for changes in the particle size, PDI, pH, and macroscopic appearance.
The pH of the MDNB-NE was measured using a multipurpose autotitrator (model MPA-210, Tecnopon Instruments, Brazil) calibrated with buffer solutions of pH 4.0 and 7.0.
The macroscopic characteristics of the formulations were also evaluated, including the color, appearance, and phase separation.
Antifungal activity
The antifungal activity of each formulation was evaluated in relation to a reference strain obtained from the American Type Culture Collection (ATCC), C. albicans (ATCC 90028), as well as clinical strains, which were collected from the oral cavity, isolated, identified, and supplied by the Laboratory of Experimental Pharmacology and Cell Cultivation of the Federal University of Paraiba, with prior approval of the ethics committee protocol number 5.821.008. These included C. albicans PLA1, C. albicans PLA5, and C. albicans PLA15.
Determination of minimum inhibitory concentration (MIC)
The MIC was determined using the microdilution technique in 96-well plates as previously described by the Clinical and Laboratory Standards Institute (CLSI document M27-A3) [30]. The MDNB was solubilized in 5% dimethyl sulfoxide, and the MDNB-NE and BNE nanoemulsions were solubilized in distilled and sterilized water. Serial dilutions (500–31.25 μg/mL) of the tested samples (MDNB, MDNB-NE and BNE), culture medium, and fungal inoculum (2.5 × 103 CFU/mL) were added to the micro-plates. Subsequently, the plates were incubated at a temperature of 35 ± 2 °C for 24 h. A 5% dimethyl sulfoxide solution was tested to assess the possible interference of this agent in the inhibition of microbial growth. Nystatin and ketoconazole were used as positive controls [31]. The MIC was defined as the lowest concentration capable of inhibiting 100% of visible fungal growth. The viability of the strains and the culture medium sterility were assessed.
Determination of minimum fungicidal concentration (MFC)
Petri dishes containing Sabouraud dextrose agar (KASVI, Brazil) in 30 μL well aliquots corresponding to the MIC and higher concentrations were plated. The plates were incubated for 48 h at 35 ± 2 °C, and readings were performed by visual observation of fungal growth based on the counting of colony-forming units (CFU). The MFC/MIC ratio was calculated to determine whether the substance exhibited fungistatic (MFC/MIC ≥ 4) or fungicidal (MFC/MIC < 4) activity [32].
Effects of MDNB on fungal cell wall and membrane permeability
Sorbitol assay (effect on cell wall)
To determine the mode of action of the product and the nanoemulsion on the cell wall of Candida albicans ATCC 90028, the microdilution technique was performed in the presence of D-sorbitol (Sigma-Aldrich, São Paulo, Brazil). For this test, the inoculum was prepared with sorbitol at a final concentration of 0.8 M. Plates were aseptically sealed and incubated at 35 ± 2 °C for 48 h. Caspofugin was used as the positive control at an initial concentration of 1 µg/mL [33,34,35]. The MIC in the presence of sorbitol was defined as the lowest concentration of substance that inhibited visible microbial growth.
Ergosterol assay (effect on cell membrane)
The test was performed using exogenous ergosterol (Sigma-Aldrich, São Paulo, Brazil) at a concentration of 400 µg/mL using the microdilution method described above. Plates were incubated at a temperature of 35 ± 2 °C for 48 h. Nystatin was used as a positive control [33,34,35,36,37]. The assay was carried out to assess the possible effect of the product and nanoemulsion on the Candida albicans ATCC 90028 cell membrane.
Molecular modelling
Target selection
Molecular targets were selected according to homology-based methodology described in previous publications [38, 39]. For this purpose, the potential targets of MDNB were predicted using the Similarity Ensemble Approach (SEA) web server [40]. All the proteins identified in this step were used as a query for a Blast [41] search against the C. albicans (taxid 5476) proteins included in the reference protein database (refseq_protein). The proteins in C. albicans identical to at least 45% of the potential targets identified by the SEA method and covered in at least 75% of their sequences by BLAST alignment were selected as the potential targets of MDNB in the fungi.
Three sources were used for the structures of the studied targets. First, the Protein Data Bank (PDB) was searched to find the experimental structures of the proteins. For those without a solved 3D structure, homology models were generated using the SwissModel server [42]. Several different models were built with the SwissModel server, among which the one with the highest QMEAN score was selected for modeling studies. In cases where no suitable 3D model (QMEAN < -4) could be retrieved for a protein, structural models were obtained from the AlphaFold Protein Structure Data-base deposited at EMBL-EBI (https://alphafold.ebi.ac.uk/) [43].
Molecular docking
Molecular docking calculations were performed using the consensus strategy described in a previous publication [44]. One initial 3D conformation of MDNB was generated using OpenEye’s Omega [45], and partial atomic charges were added to this structure with MolCharge [46]. Docking calculations were performed with Gold software [47] using the ChemPLP scoring function for primary docking. The ligand flexibility for docking was set to be very flexible, and 30 different orientations of the ligand in the binding cavity were saved. These orientations were rescored using the Chem-Score, GoldScore, and ASP scoring functions of Gold. Next, the docking scores were transformed into Z-scores and averaged to obtain the consensus Z-score. Any ligand pose with a Z-score greater than one was selected for further analysis. If no predicted binding mode meeting the former criterion could be found, the top-scoring conformer was selected for the next modeling steps.
Molecular dynamics calculations and estimation of free energies of binding
Molecular dynamics (MD) simulations were performed using Amber 20 [48] according to the protocol described in our previous studies [44, 49]. All MD simulations were performed in an explicit solvent employing OPC water molecules. The ff19SB and gaff2 force fields were employed to parameterize the proteins and ligand, respectively. The same preparation, energy minimization, heating, and equilibration protocols were applied to all the systems, except those containing membrane receptors.
For soluble proteins, the systems were enclosed in truncated octahedral boxes, solvated, and neutralized by the addition of either Na + or Cl- ions. Two-stage energy minimization, involving the steepest descent and conjugate gradient algorithms, was implemented. All the atoms, except for the solvent and counterions, were restrained during the first energy minimization stage. Next, the systems were gradually heated from 0 to 300 K, and the heated systems were equilibrated for 100 ps.
Membrane proteins were prepared for the MD simulations using the CHARMM-GUI server [50, 51]. These were embedded in a lipid bilayer containing, on each side, 50 1-Palmotoyl-2-oleoylglycero-3-phosphocholine (POPC), 50 1-Palmitoyl-2- oleoylphosphatidylethanolamine (POPE), and 25 cholesterol (CHL) molecules. The systems containing the receptor, ligand, and lipidic bilayer were solvated and neutralized by the addition of 0.15 M KCl. The configuration files provided by the CHARMM-GUI server guided the energy minimization, heating, and equilibration processes of the membrane-containing complexes. Five short (4 ns) production runs were performed for each system. The initial velocities were randomly initialized before each production run to sample different conformational spaces of the complexes. Twenty MD snapshots, evenly distributed in a 1–4 ns time interval, were extracted from each production run to estimate the free energies of binding. These energies were computed using the MM-PBSA method implemented in Amber 20 [52]. The default implicit solvent parameters were employed for the MM-PBSA calculations, and the ionic strength was set to 150 mM. For complexes containing membrane proteins, the free energies of binding were obtained using a heterogeneous dielectric implicit membrane model (memopt = 3). The thickness and center of the implicit membrane were defined as the average distance between the N31 atoms of the lipids and the average z-coordinate for the same set of atoms, respectively.
Results and discussion
Chemistry
The MDNB synthesis method is illustrated in Fig. 1. The infrared spectrum of MDBN showed an absorption band above 3000 cm−1, corresponding to the C-H sp2 stretching of aromatics, stretch bands of aromatic C = C bonds occurring in pairs in the regions of 1630 and 1461 cm−1, stretching band at 1730 cm−1 belonging to the (C = O) of the ester, and C-O stretching assigned to the ester bond with two bands at 1301 and 1165 cm−1. The nitro substituent on the aromatic ring could be characterized by the presence of two stretching bands at 1543 and 1346 cm−1.
The 1HNMR spectrum of MDNB showed a triplet at δ 9.22 (t, J = 2.2 Hz, H-4, 1H) followed by a doublet at δ 9.16 (d, J = 2.1 Hz, H-2, H-6, 2H) attributed to the aromatic ring hydrogens. MDNB showed a signal from an aliphatic side chain at δ 4.07 (s, 3H), which was attributed to the methyl hydrogens (H-1').
The 13C NMR data showed chemical shift values, which also confirmed the structural characterization of MDNB. It is worth highlighting the signals obtained with greater displacement, including those at 163.1, which is characteristic of ester C = O; 148.8, which is assigned to carbons C-3 and C-5; 133.9 of C-1; 129.6 of C-2 and C-6; and 122.5 of C-4. In addition to these signals, the MDNB compound showed a signal at 53.7 (C-1'), corresponding to the methyl carbon. These results agreed with data from the literature [53].
Thermoanalytical techniques have been widely used in drug characterization, thermal decomposition studies, and the assessment of drug stability [54]. Table 1 lists the differential thermal analysis (DTA) data. The DTA curve of MDNB (Fig. S1) in the Supplementary Information shows the events due to phase transition as a function of temperature. The first endothermic peak occurred at a temperature of 110.68 °C, which was close to that reported by Legouin et al. [53] (109–111 °C), corresponding to the sample melt. The second endothermic event occurred at a temperature of 239.12 °C, which was attributed to the degradation of the compound.
The thermogravimetric curve of MDNB (Fig. S2) in the the Supplementary Information shows a single stage of decomposition. The decomposition process started at 231.82 °C and continued until 264.22 °C, and represented a loss of 98.98% of the sample’s mass.
Development, characterization, and stability study of methyl 3,5-dinitrobenzoate nanoemulsion
In this study, an MDNB-NE was obtained using an ultrasound technique. To assess the stability of this nanoemulsion, the droplet diameter, polydispersity index, and pH were evaluated for 90 days. The results are presented in Table 2.
The MDNB-NE had a mean droplet diameter (z-average) of 185.46 ± 4.01 nm after 90 days, close to the values of the BNE (see, Supplementary Information, Table S1), showing that the MDNB not influenced the mean droplet diameter of the nanoemulsion. A slight variation in the mean droplet size was observed compared to the mean droplet size on day 1 and 90. In addition, no cream formation or phase separation was noted on visual observation, confirming the stability of the nanosystem (Table 2). The ultrasound technique was selected to develop the nanoemulsion because of its ability to form small droplets [55]. Ultrasonication breaks emulsion droplets by applying high-frequency ultrasound waves [23, 55]. During this process, disruptive forces caused by the sonicator promote turbulence and cavitation in the sample, breaking larger droplets into smaller droplets and creating the conditions for interactions between the surfactants and the oil phase [56, 57]. Surfactants are another important component to promote stability. Two non-ionic surfactants were used in this study: Tween 80® and Span 80®. These surfactants were chosen because of their low toxicity, and because they are already used in the formulation of nanoemulsions with high stability [28, 58,59,60].
Another factor contributing to the stability of these nanoemulsions was the presence of Miglyol. The main purpose of using a medium-chain triglyceride is to carry the target molecule inside the oil droplets. In addition, medium-chain triglyceride can delay the Ostwald maturation phenomenon, which is one of the most important destabilization mechanisms of dispersed systems [61].
The PDI is used to measure the homogeneity and stability of a nanoemulsion, and the MDNB-NE did not show high PDI variation over time, with values ≤ 0.3 (see Table 2). This parameter assesses the degree of uniformity of the droplet size distribution within the system [62]. PDI values of 0 indicates a monodispersed particle. A PDI < 0.5 indicates a broad distribution, and values close to 1 indicate a highly polydisperse nanoemulsion with low stability [63, 64].
UV–Vis spectrophotometry was used to quantify the amount of MDNB in the nanoemulsion. Data were obtained using the absorbance (A = 0.37 ± 0.004) on the standard curve (y = 0.0375x + 0.0034) with a coefficient of determination of 0.9991. The formulation presented a drug recovery of 98.90% in reference to 9.89 µg/mL of MDNB. Recently, Dantas et al. [27] also used Tween 80, Span 80 and Miglyol 812 to develop a nanoemulsion through the ultrasonication method and obtained a drug recovery of 87.83%.
Antifungal activity
Although the scientific literature reports the antifungal activities of several esters derived from benzoic acid against Candida species, the bioactivity of MDNB has never been tested [11, 20,21,22]. However, in the study of Duarte et al. [65] and coworkers, esters derived from 3,5-dinitrobenzoic acid such as ethyl 3,5-dinitrobenzoate and n-propyl 3,5-dinitrobenzoate showed bioactivity against Candida strains: Candida albicans, Candida krusei and Candida glabrata that demonstrates the effect of the alkyl-substituted ester group (R) on activity against Candida strains. Furthermore, the MDNB showed greater antifungal activity (MIC = 1.10 mM) than its analogue ethyl 3,5-dinitrobenzoate (MIC = 4.71 mM) [65].
In addition, in the study of Lima et al. [22], an MDNB analog, methyl 3-methyl-4-nitrobenzoate, showed antifungal activity against different strains of the Candida genus. Thus, the present study evaluated the antifungal activity of MDNB and MDNB-NE against Candida albicans strains.
The MIC and MFC for MDB and MDB-EM are presented in Table 3. The BNE (see Supplementary Information, Table 1S) was also used as a control and showed no activity against any fungal pathogen, indicating that the formulation components did not exhibit antifungal activity at the concentrations used in this study.
The MFC/MIC ratios showed that MDNB and MDNB-NE had fungicidal effects against C. albicans ATCC 90028, C. albicans PLA5, and C. albicans PLA15. The results demonstrated that MDNB-NE did not exhibit fungicidal activity (MFC/MIC ≥ 4) against C. albicans PLA1. The MDNB-NE (MFC: 1.10 mM) was more effective than MDNB (MFC: 2.21 mM) against C. albicans ATCC 90028. Andrade et al. [66], developed a nanoformulation loaded with nitro compound which also showed an increased antimicrobial activity compared to the free compound.
All strains included in the study were sensitive to ketoconazole and nystatin. The MDNB and MDNB-NE presented MICs in the range of 0.27–1.10 mM and inhibited the growth of clinical strains, proving that the development of antifungal compounds is an essential strategy for obtaining new drugs with similar or superior efficacy to those observed for the available drugs [9].
MDNB-NE showed the same MIC (1.10 mM) as MDNB against ATCC 90028. However, for clinical strains, the MIC of MDNB-NE was higher to that of MDNB, indicating that the free compound was more effective in inhibiting the growth of these strains in vitro. This higher action on C. albicans may be related to the ready availability of the drug in spreading at the medium [67]. Furthermore, this result also may be associated to the fact that the in vitro model used in this study has limitations and cannot evaluate nanoemulsions ability to improve drug absorption and permeability [68]. Thus, complementary in vivo studies can be performed to better understand the effect of MDNB loaded in a nanoemulsion.
The mechanism by which MDNB acts against Candida albicans has not yet been described. Thus, the MDNB and its nanoemulsion were subjected to tests to elucidate the possible mechanism of action against the strains of Candida albicans ATCC 90038. Ergosterol and sorbitol were used to determine the likely targets of pharmacological action (Table 4).
Ergosterol is the main sterol in the fungal cell membrane, where it regulates fluidity and permeability, making it a target of most antifungal agents [69, 70]. The results demonstrated that the addition of ergosterol to the medium containing Candida albicans and MDNB did not affect the minimal inhibitory concentration of this compound. Likewise, MDNB-NE did not show changes in the MIC, indicating that the likely mechanism of action is not related to changes in the cell membrane function. This result corroborates with the one obtained by Andrade et al. [15], who observed that the antifungal activity of nitro compounds is not associated with the ability to bind to ergosterol. Guerra et al. [71] also reported that the addition of ergosterol does not interfere with the MIC of nitrocoumarins against Aspergillus spp. However, in the study developed by Duarte et al. [65] an MDNB analogue, ethyl 3,5-dinitrobenzoate, showed an increase in MIC (1.04 mM) in the presence of ergosterol, indicating that its antifungal mechanism of action may involve ergosterol in the fungal membrane.
Sorbitol is an essential osmotic protector for fungal growth that functions by minimizing the effects of chemical agents on the cell walls of microorganisms [72,73,74]. In our study, in the presence of sorbitol, no changes were observed in the minimum inhibitory concentrations of MDNB and MDNB-NE, suggesting that the mechanism of action of these compounds is not associated with cell wall disruption. Similarly, in a study developed by Duarte et al. [65] MDNB analogue compounds such as ethyl 3,5-dinitrobenzoate and n-propyl 3,5-dinitrobenzoate also showed no change in MIC values in the presence of sorbitol, indicating that their probable mechanism of action is not related to changes in the cell wall function.
In contrast, the MIC of the control drug caspofungin increased (from 0.000028 mM to 0.0036 mM) in the presence of sorbitol, as it is known to be involved in cell wall biosynthesis. Andrade et al. [15] also demonstrated that nitro compounds do not act on the fungal cell wall of Candida albicans ATCC 10231.
As expected, the biological results indicated that the nanoemulsion system (MDNB-NE) did not interfere with the mechanism of action. In addition, the results of in vitro tests contributed to molecular modeling data (see next section), which pointed to a multi-target antifungal mechanism of action for MDNB in C. albicans, interfering with different cellular processes. For a better understanding of the antifungal effect of MDNB-NE, we suggest that future studies evaluate the effect of this product on fungal micromorphology, allowing analysis of hyphal and pseudohyphal expression.
Molecular modeling
The molecular modeling study was carried out to identify the probable targets of MDNB in C. albicans, suggesting a possible mechanism of action for this compound. For this, potential targets of the compound were first predicted using a computational target fishing approach. Next, MDNB was docked to these predicted targets leading to several ligand-receptor binding hypotheses. Finally, the predicted complexes were subject to Molecular Dynamics (MD) simulations and the free energies of binding were predicted from conformational ensembles extracted from the MD simulations. This MD-derived energies were the criterion for selecting the most likely targets of MDNB in C. albicans.
The potential targets of MDNB were predicted following the procedure described in the Methods section and are listed in Table 5. Two of the proteins listed in the table experimental structures deposited in the Protein Data Bank. Homology models were generated for eight, and the structural models of four amino acid transporters were retrieved from the AlphaFold repository. MDNB was docked into these proteins, and the results of these calculations are provided in the Supplementary Information in Table S2. For TMP1, docking was performed separately for the substrate and cofactor sites.
Overall, the predicted docking solutions were within the receptor-binding sites, showing meaningful ligand–receptor interactions. Furthermore, for half of the studied proteins, more than one possible binding mode (consensus Z-score > 1) was predicted for MDNB. This led to 26 ligand–receptor complexes involving 14 potential receptors for analysis. According to the docking results, the top–scoring targets were DED1, GCN5, and GAP5. In contrast, the worst docking scores were obtained for MEP2, UBC2, and GNP1.
Although molecular docking is a widely used tool for studying ligand–receptor interactions, its limitations need to be considered. One such limitation is the highly simplified representation of molecular interactions in docking algorithms that is required to achieve their processing capabilities. One alternative for increasing the reliability of molecular modeling frameworks is the post-processing of molecular docking results using more accurate techniques such as MD simulations [75,76,77]. Taking this into account, the 26 complexes predicted by the molecular docking methodology were subjected to the MD simulation protocol described in the Methods section. Subsequently, the free energies for binding MDNB to its potential targets were predicted from conformational ensembles derived from MD simulations using the MM-PBSA method. This approach led to a total MD simulations time of 520 ns.
The predicted free energies for binding MDNB to its potential receptors are provided in the Supplementary Information in Table S3 and summarized in Fig. 2. These results show that the lowest (best) binding free energies were obtained for four amino acid permeases (GAP5, GAP1, GNP1, and GAP2) and the ammonium transporter MEP2. Among the non-transporter proteins, the best-ranked were GLN1 and GCN5. Based on the predicted free energies of binding, complexes of MDNB with the amino acid permeases GAP5 and GAP1, as well as with the ammonium transporter MEP2, were analyzed in detail.
The predicted binding modes of MDNB to GAP5 and GAP1 are shown in Fig. 3. The represented ligand conformation corresponds to the centroid of the most populated cluster resulting from the clustering of 100 MD snapshots employed for the MMPBSA calculations. Only the residues interacting with the receptor in at least 50% of the analyzed MD snapshots are discussed here, and the same rule is applied to label residues in the Figure. Figures containing molecular structures were produced with UCSF Chimera [78], the frequency of ligand–receptor interactions was analyzed using the software Cytoscape [79], and interaction diagrams were produced using LigPlot + [80].
Overall binding modes of MDNB to GAP1 and GAP5 receptors (top), with ligand represented as orange spheres. Detailed orientation of the ligand in the binding cavities of GAP1 (center, left) and GAP5 (bottom, left) as well as diagrams of the predicted ligand–receptor interactions. In the center and bottom representations, compound A is represented as balls and sticks. Oxygen and nitrogen atoms are red and blue, respectively
MDNB was predicted to bind to GAP1 and GAP5 in the same central region, making extensive contact with the receptors. In both cases, a hydrogen bond mediated by a carbonyl group was observed in most of the analyzed MD snapshots. These interactions were predicted to occur with the side chains of Q163 and G131 for GAP1 and GAP5, respectively. The distance from the ligand’s carbonyl oxygen to one hydrogen atom in the NH2 group of the side chain of Q163 for GAP1 was 2.04 ± 0.03 Å (median 1.98 Å), while the Donor-Hydrogen-Acceptor (DHA) angle was 152.4 ± 1.6° (median 153.4°). Likewise, these values were of 2.97 ± 0.12 Å (median 2.34 Å) and 147.3 ± 1.6° (median 148.8°) for the observed hydrogen bond with G131 from GAP5. All the distance and angle values for these atoms in the analyzed MD snapshots are provided in the Supplementary Information in Table S4. Occasional hydrogen bonds, in less than 50% of the studied snapshots, were also predicted in the MDNB-GAP5 complex with the side chain of Q202 and with G129. In addition, the ligand was predicted to orient perpendicularly to F273 and L167, with the central ring flanked by S88, F164, G276, G277, and V368 in the complex with GAP1. In the same system, one of the nitro groups pointed to G90, G92, and S274, whereas the second group interacted with S87, S88, and V368, and the hydroxymethyl moiety interacted with Q163, F273, and S436.
In addition to the previously listed hydrogen bonds, the central ring of MDNB in its complex with GAP5 was in a region lined by L206, F312, A315, and G316. In this complex, the orientation of the hydroxymethyl group was different from that observed in the ligand–GAP1 system, and it was oriented toward S127, G129, G131, and S313. Furthermore, one of the nitro groups interacted with S126, Y203, and V405 of GAP5, whereas the other group made contact mainly with Q202 and S472.
In the predicted complex of MDNB with MEP2 (Fig. 4), the compound was predicted to bind at the entrance of the ammonium transport channel. The central ring stacks in front of W167 also interacted with V111 and F119. One of the nitro groups of MDNB was exposed to the solvent, while the second was oriented parallel to the side chain of I110. Finally, the ester moiety buried the binding cavity and interacted mainly with Y122, Q123, S243, T244, and G245. A hydrogen bond contributing to the stability of the complex was predicted between the carbonyl oxygen of MDNB and the side chain of Q123. For this hydrogen bond, the carbonyl oxygen – hydrogen atom distance was 3.76 ± 0.17 Å (median 3.03 Å) and the DHA angle was 123.5 ± 2.8° (median 131.1°). These values, worse than those observed for GAP1 and GAP5, are a consequence of observing this hydrogen bond in only 50% of the analyzed MD snapshots (see Supplementary Information Table S4).
Predicted binding mode of methyl 3,5-dinitrobenzoate to MEP2. The overall orientation of the ligand is depicted in the top of the Fig., and the bottom section shows the detailed ligand–receptor interactions. The compound is depicted as orange balls and sticks, whereas oxygen and nitrogen atoms are red and blue, respectively
Overall, our results suggested a multitarget antifungal mechanism of action that interferes with different cellular processes. Among the proteins studied in C. albicans, the most probable targets of MDNB are four amino acid permeases and one ammonium transporter. Despite GAP1, GAP5 and MEP2 are all transporter proteins, they show different folding. The amino acid transporters GAP1 and GAP5 are formed by 12 transmembrane helices, while MEP2 contains 11 of these motifs. In both cases the transmembrane segments surround a central channel through which transport occurs. The predicted binding of MDNB to these transporter proteins lead us to hypothesize that the main antifungal mechanism of action of this compound is linked to the interference with the uptake of amino acids and ammonium to the cell.The uptake of amino acids is essential for fungi, and this process has been proposed as a potential target for the development of novel antifungal agents [81, 82]. MEP2 has been linked not only to ammonium transport but also to morphological changes in C. albicans, making it an attractive target for the development of antifungal compounds [83]. Although the modeling results presented herein require experimental validation, they can guide further laboratory assays focusing on the antifungal mechanism of action of MDNB. These experiments as well as the optimization of the antifungal activity of MDNB derivatives can be coupled with more specific modeling studies, for example, longer MD simulations for studying the conformational changes taking place on the receptors for a small set of complexes.
Finally, the toxicity and physicochemical properties of MDNB and the antifungal drugs nystatin and ketoconazole were predicted (Table 6). Physicochemical properties were computed with the SwissADME webserver [84], while toxicity was predicted with the pkCSM webserver [85] and show that, except for unsaturation (Fraction Csp3), the rest of the physicochemical properties of MDNB have values favorable for oral bioavailability. In contrast, nystatin exceeds the favorable values for polarity and size while ketoconazole is in the limit for almost all properties. In terms of toxicity, the three compounds are predicted with comparable toxicological profiles. For MDNB, it is predicted as producing skin sensitization and to be mutagenic. Overall, these are valuable predictions for the future optimization of the antifungal activity and pharmacological profile of MDNB. Special attention must be paid to the predicted mutagenic and skin sensitizing potentials of this compound which must be experimentally evaluated and addressed in any further hit to lead optimization campaigns. Our findings suggest that future studies should be conducted, based on both non-clinical and clinical research, to test hypotheses that MDNB exhibits an appropriate toxicological profile. Considering that the physicochemical properties of MDNB can be altered without affecting its oral bioavailability, it is possible to proceed to an optimization process that jointly considers the antifungal activity and the toxicological profile.
Conclusion
It was possible to formulate a stable MDNB-loaded nanoemulsion with an EE of 98.90% over 90 days of evaluation. The in vitro studies showed that MDNB and MDNB-NE inhibited the growth of C. albicans. However, for clinical strains, the MIC of MDNB-NE in vitro was higher than that of MDNB, indicating that the free compound was more effective in inhibiting the growth of these strains. A computational investigation of the potential mechanism of action of the MDNB compound in C. albicans revealed a possible multi-target mechanism of action involving different cellular processes and corroborated the results of in vitro tests involving ergosterol and sorbitol. Thus, the data presented in this study will serve as a basis for further research on antifungal nitro compounds.
Data availability
Data supporting the findings of this study are available within the article and can be obtained from the corresponding author upon request.
References
Pereira R, Dos Santos Fontenelle RO, de Brito EHS, de Morais SM (2021) Biofilm of Candida albicans: formation, regulation and resistance. J Appl Microbiol 131:11–22. https://doi.org/10.1111/jam.14949
Lohse MB, Gulati M, Johnson AD, Nobile CJ (2018) Development and regulation of single- and multi-species Candida albicans biofilms. Nat Rev Microbiol 16:19–31. https://doi.org/10.1038/nrmicro.2017.107
Dadar M, Tiwari R, Karthik K, Chakraborty S, Shahali Y, Dhama K (2018) Candida albicans - Biology, molecular characterization, pathogenicity, and advances in diagnosis and control - An update. Microb Pathog 117:128–138. https://doi.org/10.1016/j.micpath.2018.02.028
Garcia A, Fan YY, Vellanki S, Huh EY, Vanegas D, Wang SH, Lee SC (2020). Nanoemulsion as an effective treatment against human-pathogenic fungi. mSphere 2019, 4:e00729–19. https://doi.org/10.1128/mSphere.00729-19
Bhattacharya S, Sae-Tia S, Fries BC (2020) Candidiasis and mechanisms of antifungal resistance. Antibiotics (Basel) 9:312. https://doi.org/10.3390/antibiotics9060312
Araujo VHS, Duarte JL, Carvalho GC, Silvestre ALP, Fonseca-Santos B, Marena GD, Ribeiro TC, Dos Santos MAR, Bauab TM, Chorilli M (2020) Nanosystems against candidiasis: a review of studies performed over the last two decades. Crit Rev Microbiol 46:508–547. https://doi.org/10.1080/1040841X.2020.1803208
Van Daele R, Spriet I, Wauters J, Maertens J, Mercier T, Van Hecke S, Brüggemann R (2019) Antifungal drugs: What brings the future? Med Mycol 57:S328–S343. https://doi.org/10.1093/mmy/myz012
Lima IDO, Oliveira RDAG, Lima EDO, Farias NMP, Souza ELD (2006) Antifungal activity from essential oils on Candida species. Rev Bras Farmacogn 16:197–201
Alves DDN, Ferreira AR, Duarte ABS, Melo AKV, de Sousa DP, de Castro RD (2021) Breakpoints for the classification of anti-Candida compounds in antifungal screening. Biomed Res Int 2021:6653311. https://doi.org/10.1155/2021/6653311
Ye Y, Chen Y, Hou Y, Yu H, Zhu L, Sun Y, Zhou M, Chen Y, Dong M (2022) Two new benzoic acid derivatives from endophytic fungus Aspergillus versicolor. Nat Prod Res 36:223–228. https://doi.org/10.1080/14786419.2020.1777121
Perez-Castillo Y, Lima TC, Ferreira AR, Silva CR, Campos RS, Neto JBA, Magalhães HIF, Cavalcanti BC, Júnior HVN, de Sousa DP (2020) Bioactivity and molecular docking studies of derivatives from cinnamic and benzoic acids. Biomed Res Int 2020:6345429. https://doi.org/10.1155/2020/6345429
Del Olmo A, Calzada J, Nuñez M (2017) Benzoic acid and its derivatives as naturally occurring compounds in foods and as additives: Uses, exposure, and controversy. Cri Rev Food Sci Nutr 57:3084–3103. https://doi.org/10.1080/10408398.2015.1087964
Kalaycı M, Türkeş C, Arslan M, Demir Y, Beydemir Ş (2021) Novel benzoic acid derivatives: Synthesis and biological evaluation as multitarget acetylcholinesterase and carbonic anhydrase inhibitors. Arch Pharm (Weinheim) 354:e2000282. https://doi.org/10.1002/ardp.202000282
Li X, Du C, Zhao H (2017) Determination and modeling of binary and ternary solid-liquid phase equilibrium for the systems formed by 3, 5-dinitrobenzoic acid, m-nitrobenzoic acid and acetone. J Chem Thermodyn 105:21–29. https://doi.org/10.1016/j.jct.2016.10.004
Andrade JT, Alves SLG, Lima WG, Sousa CDF, Carmo LF, De Sá NP, Morais FB, Johann S, Villar JAFP, Ferreira JMS (2020) Pharmacologic potential of new nitro-compounds as antimicrobial agents against nosocomial pathogens: design, synthesis, and in vitro effectiveness. Folia Microbiol (Praha) 65:393–405. https://doi.org/10.1007/s12223-019-00747-7
Nascimento LG, de Morais MC, Junior JKO, Lima EO, de Sousa DMP (2022) Synthetic 2-nitrocinnamates: investigation of the antifungal action against Candida species. J Chem 2023:1–7. https://doi.org/10.1155/2023/8525145
Fries A, Bretschneider T, Winkler R, Hertweck C (2011) A ribonucleotide reductase-like electron transfer system in the nitroaryl-forming N-oxygenase AurF. ChemBioChem 12:1832–1835. https://doi.org/10.1002/cbic.201100138
Nascimento LG (2017) Preparação de ésteres nitrocinâmicos e avaliação da sua atividade antimicrobiana. Dissertation, Federal University of Paraiba
Ferreira AR (2018). Ésteres sintéticos derivados do ácido 3-metil-4-nitrobenzoico e avaliação da sua atividade antifúngica. Dissertation, Federal University of Paraiba
Silva RHN, Silva DF, da Nóbrega FR, Oliveira AJMS, Lima EO, de Sousa DP (2017) Biphenyl-4-carboxylic acid derived esters with antifungal activity. J Chem and Pharm Res 9:89–94
Lima TC, Ferreira AR, Silva DF, Lima EO, de Sousa DP (2018) Antifungal activity of cinnamic acid and benzoic acid esters against Candida albicans strains. Nat Prod Res 32:572–575. https://doi.org/10.1080/14786419.2017.1317776
Lima TC, Ferreira AR, Barboza JN (2018) Antimicrobial activity of cinnamic and benzoic methyl esters. Lat Am J Pharm 37:1011–1016
Páez-Hernández G, Mondragón-Cortez P, Espinosa-Andrews H (2019) Developing curcumin nanoemulsions by high-intensity methods: Impact of ultrasonication and microfluidization parameters. Lwt 111:291–300
Khani S, Abbasi S, Keyhanfar F, Amani A (2019) Use of artificial neural networks for analysis of the factors affecting particle size in mebudipine nanoemulsion. J Biomol Struct Dyn 37:3162–3167. https://doi.org/10.1080/07391102.2018.1510341
Pandey P, Gulati N, Makhija M, Purohit D, Dureja H (2020) Nanoemulsion: a novel drug delivery approach for enhancement of bioavailability. Recent Pat Nanotechnol 14:276–293. https://doi.org/10.2174/1872210514666200604145755
Sanderson JT, Clabault H, Patton C, Lassalle-Claux G, Jean-François J, Paré AF, Hébert MJ, Surette ME, Touaibia M (2013) Antiproliferative, antiandrogenic and cytotoxic effects of novel caffeic acid derivatives in LNCaP human androgen-dependent prostate cancer cells. Bioorg Med Chem 21:7182–7193. https://doi.org/10.1016/j.bmc.2013.08.057
Dantas AGB, de Souza RL, de Almeida AR, Xavier Júnior FH, Pitta MGDR, Rêgo MJBM, Oliveira EE (2021) Development, characterization, and immunomodulatory evaluation of carvacrol-loaded nanoemulsion. Molecules 26:3899. https://doi.org/10.3390/molecules26133899
Landa P, Kokoska L, Pribylova M, Vanek T, Marsik P (2009) In vitro anti-inflammatory activity of carvacrol: Inhibitory effect on COX-2 catalyzed prostaglandin E 2 biosynthesisb. Arch Pharm Res 32:75–78. https://doi.org/10.1007/s12272-009-1120-6
Oliveira EE, Barendji M, Vauthier C (2020) Understanding nanomedicine size and biological response dependency: what is the relevance of previous relationships established on only batch-mode DLS-measured sizes? Pharm Res 37:161. https://doi.org/10.1007/s11095-020-02869-x
CLSI (2008) Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts: Approved Standard – Third Edition. CLSI document M27-A3. Wayne, PA: Clinical and Laboratory Standards Institute
Peixoto LR, Rosalen PL, Ferreira GL, Freires IA, de Carvalho FG, Castellano LR, de Castro RD (2017) Antifungal activity, mode of action and anti-biofilm effects of Laurus nobilis Linnaeus essential oil against Candida spp. Arch Oral Biol 73:179–185. https://doi.org/10.1016/j.archoralbio.2016.10.013
Siddiqui ZN, Farooq F, Musthafa TM, Ahmad A, Khan AU (2013) Synthesis, characterization and antimicrobial evaluation of novel halopyrazole derivatives. J Soc Quím Saudita 17:237–243. https://doi.org/10.1016/j.jscs.2011.03.016
Escalante A, Gattuso M, Pérez P, Zacchino S (2008) Evidence for the mechanism of action of the antifungal phytolaccoside B isolated from Phytolacca tetramera Hauman. J Nat Prod 71:1720–1725. https://doi.org/10.1021/np070660i
Freires IA, Murata RM, Furletti VF, Sartoratto A, Alencar SM, Figueira GM, Rodrigues JAO, Duarte MC, Rosalen PL (2014) Coriandrum sativum L. (Coriander) essential oil: antifungal activity and mode of action on Candida spp., and molecular targets affected in human whole-genome expression. PLoS ONE 9:e99086. https://doi.org/10.1371/journal.pone.0099086
Lima IO, Pereira FDO, Oliveira WAD, Lima EDO, Menezes EA, Cunha FA, Diniz MDFFM (2013) Antifungal activity and mode of action of carvacrol against Candida albicans strains. J Essent Oil Res 25:138–142. https://doi.org/10.1080/10412905.2012.754728
Perlin DS (2011) Current perspectives on echinocandin class drugs. Future Microbiol 6:441–457. https://doi.org/10.2217/fmb.11.19
Pierce CG, Srinivasan A, Uppuluri P, Ramasubramanian AK, López-Ribot JL (2013) Antifungal therapy with an emphasis on biofilms. Curr Opin Pharmacol 13:726–730. https://doi.org/10.1016/j.coph.2013.08.008
Lopes SP, Yepes LM, Pérez-Castillo Y, Robledo SM, de Sousa DP (2020) Alkyl and Aryl derivatives based on p-coumaric acid modification and inhibitory action against Leishmania braziliensis and Plasmodium falciparum. Molecules 25:3178. https://doi.org/10.3390/molecules25143178
Araújo MO, Pérez-Castillo Y, Oliveira LHG, Nunes FC, de Sousa DP (2020) Larvicidal activity of cinnamic acid derivatives: investigating alternative products for Aedes aegypti L. Control Molecules 26:61. https://doi.org/10.3390/molecules26010061
Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK (2007) Relating protein pharmacology by ligand chemistry. Nat Biotechnol 25:197–206. https://doi.org/10.1038/nbt1284
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1197). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389-3402. https://doi.org/10.1093/nar/25.17.3389
Bienert S, Waterhouse A, de Beer TA, Tauriello G, Studer G, Bordoli L, Schwede T (2017) The SWISS-MODEL repository-new features and functionality. Nucleic Acids Res 45:D313–D319. https://doi.org/10.1093/nar/gkw1132
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
de Morais MC, Perez-Castillo Y, Silva VR, Santos LS, Soares MBP, Bezerra DP, de Castro RD, de Sousa DP (2021) Cytotoxic and antifungal amides derived from ferulic acid: molecular docking and mechanism of action. Biomed Res Int 2021:3598000. https://doi.org/10.1155/2021/3598000
Hawkins PC, Skillman AG, Warren GL, Ellingson BA, Stahl MT (2010) Conformer generation with OMEGA: algorithm and validation using high quality structures from the Protein Databank and Cambridge Structural Database. J Chem Inf Model 50:572–584. https://doi.org/10.1021/ci100031x
OpenEye Scientific Software. QUACPAC. Santa Fe, NM: OpenEye Scientific Software. Available online: http://www.eyesopen.com (accessed on 6 august 2021)
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748. https://doi.org/10.1006/jmbi.1996.0897
Case DA, Ben-Shalom IY, Brozell SR, Cerutti DS, Cheatham TE, Cruzeiro VWD, Darden TA, Duke RE, Ghoreishi D, Gilson MK, Gohlke H, Goetz AW, Greene D, Harris R, Homeyer N, Izadi S, Kovalenko A, Kurtzman T, Lee TS, LeGrand S, Li P, Lin C, Liu J, Luchko T, Luo R, Mermelstein DJ, Merz KM, Miao Y, Monard G, Nguyen C, Nguyen H, Omelyan I, Onufriev A, Pan F, Qi R, Roe DR, Roitberg A, Sagui C, Schott-Verdugo S, Shen J, Simmerling CL, Smith J, Salomon-Ferrer R, Swails J, Walker RC, Wang J, Wei H, Wolf RM, Wu X, Xiao L, York DM, Kollman PA (2021) AMBER 2021, University of California: San Francisco. CA, USA
Lopes SP, Castillo YP, Monteiro ML, Menezes RRPPB, Almeida RN, Martins AMC, de Sousa DP (2019) Trypanocidal mechanism of action and in silico studies of p-Coumaric acid derivatives. Int J Mol Sci 20:5916. https://doi.org/10.3390/ijms20235916
Lee J, Cheng X, Swails JM, Yeom MS, Eastman PK, Lemkul JA, Wei S, Buckner J, Jeong JC, Qi Y, Jo S, Pande VS, Case DA, Brooks CL, MacKerell AD, Klauda JB, Im W (2016) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12:405–413. https://doi.org/10.1021/acs.jctc.5b00935
Wu EL, Cheng X, Jo S, Rui H, Song KC, Dávila-Contreras EM, Qi Y, Lee J, Monje-Galvan V, Venable RM, Klauda JB, Im W (2014) CHARMM-GUI membrane builder toward realistic biological membrane simulations. J Comput Chem 35:1997–2004. https://doi.org/10.1002/jcc.23702
Miller BR, McGee TD, Swails JM, Homeyer N, Gohlke H, Roitberg AE (2012) MMPBSA.py: an efficient program for end-state free energy calculations. J Chem Theory Comput 8:3314–3321. https://doi.org/10.1021/ct300418h
Legouin B, Gayral M, Uriac P, Cupif JF, Levoin N, Toupet L, van de Weghe P (2010). Molecular Tweezers: Synthesis and Formation of Host–Guest Complexes. Eur J Org Chem 5503-5508.https://doi.org/10.1002/ejoc.201000729
Böer TM, Procópio JVV, do Nascimento TG, Macêdo RO (2013) Correlation of thermal analysis and pyrolysis coupled to GC–MS in the characterization of tacrolimus. J Pharm Biomed Anal 73:18–23. https://doi.org/10.1016/j.jpba.2012.01.040
Nirmala MJ, Durai L, Gopakumar V, Nagarajan R (2020) Preparation of celery essential oil-based nanoemulsion by ultrasonication and evaluation of its potential anticancer and antibacterial activity. Int J Nanomedicine 15:7651–7666. https://doi.org/10.2147/IJN.S252640
Wu WH, Eskin DG, Priyadarshi A, Subroto T, Tzanakis I, Zhai W (2021) New insights into the mechanisms of ultrasonic emulsification in the oil-water system and the role of gas bubbles. Ultrason Sonochem 73:105501. https://doi.org/10.1016/j.ultsonch.2021.105501
McClements DJ (2011) Edible nanoemulsions: fabrication, properties, and functional performance. Soft Matter 7:2297–2316. https://doi.org/10.1039/C0SM00549E
Felício IM, de Souza RL, Melo CO, Lima KYG, Vasconcelos U, de Moura RO, Oliveira EE (2021) Development and characterization of a carvacrol nanoemulsion and evaluation of its antimicrobial activity against selected food-related pathogens. Lett Appl Microbiol 72:299–306. https://doi.org/10.1111/lam.13411
Davis K, Rover MR, Salvachúa D, Smith RG, Beckham GT, Wen Z, Brown RC, Jarboe LR (2019) Promoting microbial utilization of phenolic substrates from bio-oil. J Ind Microbiol Biotechnol 46:1531–1545. https://doi.org/10.1007/s10295-019-02208-z
Pensado A, Fernandez-Piñeiro I, Seijo B, Sanchez A (2014) Anionic nanoparticles based on Span 80 as low-cost, simple and efficient non-viral gene-transfection systems. Int J Pharm 476:23–30. https://doi.org/10.1016/j.ijpharm.2014.09.032
Ryu V, Corradini MG, McClements DJ, McLandsborough L (2019) Impact of ripening inhibitors on molecular transport of antimicrobial components from essential oil nanoemulsions. J Colloid Interface Sci 556:568–576. https://doi.org/10.1016/j.jcis.2019.08.059
Danaei M, Dehghankhold M, Ataei S, Hasanzadeh Davarani F, Javanmard R, Dokhani A, Khorasani S, Mozafari MR (2018) Impact of particle size and polydispersity index on the clinical applications of lipidic nanocarrier systems. Pharmaceutics 10:57. https://doi.org/10.3390/pharmaceutics10020057
Mohd Narawi MM, Chiu HI, Yong YK, Mohamad Zain NN, Ramachandran MR, Tham CL, Samsurrijal SF, Lim V (2020) Biocompatible nutmeg oil-loaded nanoemulsion as phyto-repellent. Front Pharmacol 11:214. https://doi.org/10.3389/fphar.2020.00214
Deghiedy NM, Elkenawy NM, Abd El-Rehim HA (2021) Gamma radiation-assisted fabrication of bioactive-coated thyme nanoemulsion: A novel approach to improve stability, antimicrobial and antibiofilm efficacy. J Food Eng 304:110600. https://doi.org/10.1016/j.jfoodeng.2021.110600
Duarte ABS, Perez-Castillo Y, Andrade PN, de Castro RD, de Sousa DP (2022) 3,5-Dinitrobenzoate and 3,5-dinitrobenzamide derivatives: mechanistic, antifungal, and In Silico studies. J Chem 2022:1–17. https://doi.org/10.1155/2022/2336175
De Andrade LF, Apolinário AC, Rangel-Yagui CO, Stephano MA, Tavares LC (2020) Chitosan nanoparticles for the delivery of a new compound active against multidrug-resistant Staphylococcus aureus. J Drug Deliv Sci Technol 55:101363. https://doi.org/10.1016/j.jddst.2019.101363
Ribeiro RF, Motta MH, Härter APG, Flores FC, Beck RCR, Schaffazick SR, de Bona da Silva C (2016). Spray-dried powders improve the controlled release of antifungal tioconazole-loaded polymeric nanocapsules compared to with lyophilized products. Mater Sci Eng C Mater Biol Appl 59:875-884.https://doi.org/10.1016/j.msec.2015.10.035
Anuar N, Sabri AH, Bustami Effendi TJ, Abdul Hamid K (2020) Development and characterisation of ibuprofen-loaded nanoemulsion with enhanced oral bioavailability. Heliyon 6:e04570. https://doi.org/10.1016/j.heliyon.2020.e04570
Sun L, Liao K (2020) The Effect of honokiol on ergosterol biosynthesis and vacuole function in Candida albicans. J Microbiol Biotechnol 30:1835–1842. https://doi.org/10.4014/jmb.2008.08019
Rodrigues ML (2018) The multifunctional fungal ergosterol. MBio 9:e01755-e1818. https://doi.org/10.1128/mBio.01755-18
Guerra FQS, de Araújo RSA, de Sousa JP, Pereira FO, Mendonça-Junior FJB, Barbosa-Filho JM, Lima EO (2015) Evaluation of antifungal activity and mode of action of new coumarin derivative, 7-Hydroxy-6-nitro-2H-1-benzopyran-2-one, against Aspergillus spp. Evid Based Complement Alternat Med 2015:925096. https://doi.org/10.1155/2015/925096
Silva Junior IF, Raimondi M, Zacchino S, Cechinel Filho V, Noldin VF, Rao VS, Martins DT (2010) Avaliação da atividade antifúngica e modo de ação dos extratos da entrecasca, frações e ácido elágico de Lafoensia pacari A. St.-Hil., Lythraceae. Rev Bras Farmacogn 20:422–428. https://doi.org/10.1590/S0102-695X2010000300021
Łukowska-Chojnacka E, Mierzejewska J, Milner-Krawczyk M, Bondaryk M, Staniszewska M (2016) Synthesis of novel tetrazole derivatives and evaluation of their antifungal activity. Bioorg Med Chem 24:6058–6065. https://doi.org/10.1016/j.bmc.2016.09.066
Alves DN, Monteiro AFM, Andrade PN, Lazarini JG, Abílio GMF, Guerra FQS, Scotti MT, Scotti L, Rosalen PL, Castro RD (2020) Docking prediction, antifungal activity, anti-biofilm effects on candida spp., and toxicity against human cells of cinnamaldehyde. Molecules 25:5969. https://doi.org/10.3390/molecules25245969
Alonso H, Bliznyuk AA, Gready JE (2006) Combining docking and molecular dynamic simulations in drug design. Med Res Rev 26:531–568. https://doi.org/10.1002/med.20067
Liu K, Kokubo H (2017) Exploring the stability of ligand binding modes to proteins by molecular dynamics simulations: a cross-docking study. J Chem Inf Model 57:2514–2522. https://doi.org/10.1021/acs.jcim.7b00412
Guterres H, Im W (2020) Improving protein-ligand docking results with high-throughput molecular dynamics simulations. J Chem Inf Model 60:2189–2198. https://doi.org/10.1021/acs.jcim.0c00057
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786. https://doi.org/10.1021/ci200227u
McCarthy MW, Walsh TJ (2018) Amino acid metabolism and transport mechanisms as potential antifungal targets. Int J Mol Sci 19:909. https://doi.org/10.3390/ijms19030909
Garbe E, Vylkova S (2019) Role of amino acid metabolism in the virulence of human pathogenic fungi. Curr Clin Microbiol Rep 6:108–119. https://doi.org/10.1007/s40588-019-00124-5
Volkova M, Atamas A, Tsarenko A, Rogachev A, Guskov A (2021) Cation transporters of Candida albicans-new targets to fight candidiasis? Biomolecules 11:584. https://doi.org/10.3390/biom11040584
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58:4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Funding
This study was funded in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brazil (CAPES) – Process 001. The authors also thank the FAPESq-PB- Fundação de Amparo a Pesquisa do Estado da Paraíba and CM-ACQuimBio – Brazil.
Author information
Authors and Affiliations
Contributions
BSD, YPC, DNA, RDC and RLS: conceptualization, methodology, writing-original draft; DPS: conceptualization, writing review and editing, resources; EEO: conceptualization, writing-review and editing, supervision, project administration, resources.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Celia Maria de Almeida Soares
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duarte, A.B.S., Perez-Castillo, Y., da Nóbrega Alves, D. et al. Antifungal activity against Candida albicans of methyl 3,5-dinitrobenzoate loaded nanoemulsion. Braz J Microbiol 55, 25–39 (2024). https://doi.org/10.1007/s42770-023-01214-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01214-9