Abstract
The genetic diversity of the dengue virus is characterized by four circulating serotypes, several genotypes, and an increasing number of existing lineages that may have differences in the potential to cause epidemics and disease severity. Accurate identification of the genetic variability of the virus is essential to identify lineages responsible for an epidemic and understanding the processes of virus spread and virulence. Here, we characterize, using portable nanopore genomic sequencing, different lineages of dengue virus 2 (DENV-2) detected in 22 serum samples from patients with and without dengue warning signs attended at Hospital de Base of São José do Rio Preto (SJRP) in 2019, during a DENV-2 outbreak. Demographic, epidemiological, and clinical data were also analyzed. The phylogenetic reconstruction and the clinical data showed that two lineages belonging to the American/Asian genotype of DENV-2—BR3 and BR4 (BR4L1 and BR4L2)—were co-circulating in SJRP. Although preliminary, these results indicate no specific association between clinical form and phylogenetic clustering at the virus consensus sequence level. Studies with larger sample sizes and which explore single nucleotide variants are needed. Therefore, we showed that portable nanopore genome sequencing could generate quick and reliable sequences for genomic surveillance to monitor viral diversity and its association with disease severity as an epidemic unfolds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dengue virus (DENV) is an arbovirus that represents a classic example of reemerging disease, resulting in almost 400 million infections annually in the world [1], being endemic in more than 100 countries, with an incidence that increased in the last 50 years [2]. DENV belongs to the Flaviviridae family, including other clinically important arboviruses like Zika (ZIKV) and yellow fever (YFV). The flavivirus genome consists of a positive single-stranded RNA of approximately 11 kilobases (kb) in length, which encode three structural proteins (capsid (C), pre-membrane (prM), and envelope (E)) and seven nonstructural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5) [3].
The clinical manifestations of dengue are classified according to the levels of severity in (1) dengue without warning signs (patients presenting typical symptoms like nausea/vomiting, rash, headache, eye pain, muscle ache, or joint pain, leukopenia, and positive tourniquet test; (2) dengue with warning signs (patients presenting typical symptoms more other one as abdominal pain, persistent vomiting, fluid accumulation, bleeding from the mucosa, lethargy, enlarged liver, increased hematocrit with decreased platelets); and (3) severe dengue (patients that show severe plasma leakage, severe bleeding, or organ failure) [4].
DENV is classified into four serotypes (DENV-1, 2, 3, and 4) that share approximately 70% of homology at an amino acid level, and within each DENV serotype, there is some degree of genetic diversity [5]. This genetic diversity provides heterogeneity in the strain’s circulation, leading to the spread of serotypes and genotypes that differ in their potential to cause epidemics, representing a serious threat to the population. Thus, accurate identification of the genetic variability of viruses is essential for understanding and identifying a key mutation responsible for an epidemic [6].
In this study, we characterize, using a portable genomic approach [7, 8], the genetic diversity of DENV-2 detected in serum samples obtained from patients with infection classified as dengue without warning signs and dengue with warning signs attended at Hospital de Base of São José do Rio Preto, a Brazilian city located in the northeast of São Paulo State, that reported 65.7% (400,856) of all dengue cases identified in the country [9].
Material and methods
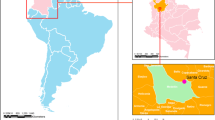
São José do Rio Preto is in the southwestern region of São Paulo State, with a population of around 469,173 inhabitants, a tropical climate, average annual rainfall of 1465 mm, and an altitude of 510 m asl [10, 11]. São José do Rio Preto is considered hyperendemic for DENV since these climatic conditions favor the proliferation of the mosquito Aedes aegypti [12] (the primary vector of DENV), whose reinfestation in São José do Rio Preto was detected in 1985 [13]. The first case of DENV in this city occurred in 1990, caused by DENV-1, and the first dengue epidemic occurred 4 years later. DENV-2, DENV-3, and DENV-4 were detected, in 1998, 2005, and 2011, respectively, causing new epidemics [14, 15]. In recent years, thousands of cases have been confirmed, highlighting 2013, 2015, and 2019 with 18,000, 22,000, and 33,000 cases, respectively [16].
To assess the genetic diversity of DENV-2 circulating in São José do Rio Preto, we randomly selected 22 RT-qPCR positive samples of DENV-2 from the 2019 bank, collected from 1 to 25 days after the onset of symptoms. Samples were provided by the Hospital de Base of São José do Rio Preto in a study submitted and approved by the Ethical Review Board (process number 28309320.3.0000.5415) from Faculdade de Medicina de São José do Rio Preto (FAMERP), São Paulo, Brazil.
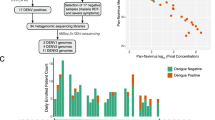
The samples collected during the 2019 dengue epidemic, between February and May, were submitted to total viral RNA extraction using QIAmp Viral RNA Kit (Qiagen®), following the manufacturer’s recommendations. Subsequently, the total RNA was submitted to RT-qPCR Singleplex-DENV [17] to check the CT (cycle threshold) values. DENV-2 whole-genome sequencing was performed using a multiplex PCR primer scheme to amplify the DENV-2 genome [18]. The extracted RNA was transcribed to cDNA using Superscript IV First-Strand Synthesis System (Thermo Fisher Scientific, MA, USA) and random hexamer oligonucleotide (Invitrogen®). Then, multiplex PCR was performed to generate overlapping amplicons from the entire viral genome. The amplification of the DENV-2 genome consisted of 35 PCR cycles according to the reaction and thermocycling mixture described by Quick and collaborators [19]. Magnetic beads (AmpureXP (Beckman Coulter, High Wycombe, UK)) were used to purify the PCR products, which were then quantified with Qubit dsDNA High Sensitivity on a Qubit 3.0 device (Life Technologies).
Sequencing libraries were generated using the Genomic DNA Sequencing Kit SQK-LSK108 (Oxford Nanopore Technologies), combining, in equimolar proportions, a total of 250 ng of PCR products using the Native Barcoding Kit (NBD103, Oxford Nanopore Technologies, Oxford, UK). The libraries were loaded into an Oxford Nanopore R9.4 Flow cell (FLO-MIN106). The generated reads were analyzed using the Guppy Basecalling Software version 3.0.3.2 [20], in which the electrical signal of the MinION run is analyzed and converted into nucleotide bases. Basecalled reads were then analyzed in Porechop version 0.2.3 [21] software which removes the adapters and recognizes the sequences of the two barcodes in each final portion of the reads, thus demultiplexing the reads. Subsequently, demultiplexed reads were aligned with the reference sequence of the DENV-2 complete genome (EU920833.1) using bwa-bwa (Alignment via Burrows-Wheeler transformation) version 0.7.17-r1188 [22]. Consensus sequence generation and variant calling were performed using the Nanopolish version 0.11.1 [23], which also checks the original sequencing data generated using Variant Calling. Finally, a consensus sequence was generated for all genomic sites with a depth of at least 20 reads for each site. In cases below this value, “N” bases were added to these positions in the consensus sequence.
We constructed a maximum likelihood (ML) phylogeny to identify the genotypes of the samples. The ML phylogenetic analysis was based on comparing the study sequences with the sequences stored in a database belonging to the different DENV-2 genotypes and lineages originating in different regions of the world in different years of circulation. Sequences were aligned using MAFFT v 7.450 [23] and edited on AliView version 1.23 [24]. Phylogenetic reconstruction was performed using IQ-TREE multicore software version 2.0-rc2 [25], using TIM3 + F + I + G4 as the nucleotide substitution model. The reliability of the branches was tested through ultrafast bootstrapping and approximate likelihood-ratio test (SH-aLRT), 1000 runs each. After that, we inferred time-scaled trees by using TreeTime [26]. The final tree generated was edited and visualized using FigTree v.1.4.4 [27]. Demographic, epidemiological, and clinical data (symptoms, radiologic, and laboratory findings) were obtained from the Hospital de Base of São José do Rio Preto database. Confidentiality was ensured by de-identifying all questionnaires and samples before data entry and analyses. The data collected were entered into a spreadsheet using Excel® software (Microsoft, Redmond, WA, USA) and imported to SPSS software for IOS (version 25, SPSS, Inc; Chicago, IL, USA). Pearson’s chi-square test was used to determine whether the expected frequency in the groups was met. When the chi-square criteria were not achieved, Fisher’s test was chosen. A significance level of 0.05 (5%) was adopted for all tests employed.
Results
To explore the genetic diversity of DENV in patients with different clinical forms of dengue fever, we sequenced 22 samples from São José do Rio Preto with collection dates from February to May 2019. The 22 DENV-2 sequences are available in GenBank (https://www.ncbi.nlm.nih.gov; GenBank accession numbers can be found in Table S1 in Supplementary Material).
Epidemiological data of DENV-2 samples from patients at São José do Rio Preto are presented in Table 1. About the patients, the age range was 24 − 82 years, and the median age was 51 years. The frequency of males and females was 36.7% and 63.3%, respectively. All samples were positive for DENV-2 by RT-qPCR, with CT values ranging from 18.82 to 30.72, presenting an average of 25.51 and a median of 25.58. All samples were positive for DENV-2 by RT-qPCR, and CT values with an average of 25.51 (Table 1). These samples were submitted to the amplicon-based sequencing protocol adapted to MinION nanopore sequencing [19]. The generated consensus genomes had a mean coverage of 82% of the genome at 20 × minimum sequencing depth.
The 22 newly generated genomes from São José do Rio Preto were subtyped using the Dengue Virus Typing tool [28] and identified as belonging to Genotype III. The consensus sequences were analyzed and aligned with 595 other DENV-2 sequences from the Americas from 1986 to 2019 (Accession numbers can be found in Table S2 in Supplementary Material).
During the sequence analysis, we detected that the genome coverage is not complete. For this reason, we decided to apply the phylogenetic reconstruction only to the envelope, a region that presented complete coverage and that has already been widely used for the detection of DENV strains [29]. Genotyping analyses (Fig. 1) showed that all Sao José do Rio Preto sequences were classified as DENV-2 Genotype III (DENV-2-III, Asian-American genotype).
Most samples were classified as DENV-2-III BR4 lineage (n = 20, 91%), and only two samples (9%) belonged to the DENV-2-III BR3 lineage. DENV-2-III BR4 lineage can be further subdivided into DENV-2-III BR4L1 and DENV-2-III BR4L2, according to their clustering patterns. In our dataset, we found that 75% of the samples (n = 15/20) were identified as DENV-2-III BR4L1 and the remaining 25% (n = 5/20) as DENV-2-III BR4L2.
The nucleotide composition of all dengue sequences obtained in this work was analyzed by structural (capsid, pre-membrane/membrane, and envelope) and non-structural (NS1, NS2A, NS2B, NS3, NS4A, NS4B, NS5) proteins (Fig. 2). We observed different proportions of nucleotides when comparing the three lineages. In the membrane, for example, we found that 44% of BR3 genome samples correspond to G, but 29% and 28% of BR4L1 and BR4L2, respectively, correspond to this mentioned nucleotide. However, these differences did not result in many amino acid changes, except for the presence of a Serine in the BR3 lineage and an Alanine in both BR4 lineages, in 697 positions. Some amino acid differences were noted in some samples of each lineage, including D23N in the BR3 samples, A121T in six BR4L1 samples, A32T in two BR4L1 samples, E558K in eleven BR4L1 samples, A120T in three BR4L2 samples, E558K in four BR4L2 samples, and I109L in two BR4L2 samples.
Concerning the dengue clinical classification defined by WHO 2009 [4], eight cases were classified as dengue without warning signs, 14 as dengue with warning signs and/or severe dengue (with neurologic impairment). We did not find a relation between the clinical outcome and the lineage and sublineage classification, as the phylogenetic analyses showed clustering of both clinical forms within different lineages and sublineages. In addition to clinical forms, the frequency of signs and symptoms was analyzed separately. Fever was the most common symptom reported at hospital admission (21/22; 95.45%), followed by myalgia (17/22; 77.27%), headache (14/22; 63.63%), exanthema (7/22; 31.81%), ocular pain (7/22; 31.81%), arthralgia (6/22; 27.27%), and vomits (4/22; 18.18%). The most frequent warning signs were abdominal pain (6/12; 50%), hemoconcentration (6/12; 50%), fluid accumulation (4/12; 33.33%), and bleeding (3/12; 25%). The severe form case evolved with seizures, hemodynamic dysfunction, fluid accumulation, and brain bleeding. The frequency of signs and symptoms according to lineage classification is presented in Tables 2 and 3.
Discussion
Globalization, climate change, and increased human mobility have facilitated the increased incidence and co-circulation of multiple DENV serotypes, genotypes, and lineages in the same region [30]. In endemic and hyperendemic regions, the epidemiological dynamics of the DENVs are typically characterized by clade replacement from time to time [31,32,33,34]. The replacement of circulating DENV lineages is usually associated with an increase in the number of total cases and the number of severe cases. Thus, the early identification of new lineages must be carried out to rapidly track virus diversity during an outbreak. Genomic surveillance can help guide measures to contain the emergence and spread of new DENV outbreaks and decrease the number of severe dengue cases, as well as deaths [28, 29]. Here, we sequenced DENV-2 clinical samples and demonstrate that the use of Oxford Nanopore technology in the diagnosis of DENV could be helpful to identify and describe the circulating lineages.
In 2019, the predominant serotype in São José do Rio Preto was DENV-2 [35]. This serotype can be classified into six genotypes, named I-VI. Until 2019, three genetic lineages of the DENV-2 Genotype III (DENV2-III) had been reported in Brazil. These three circulating lineages were named lineages 1–3 [31] or BR1-BR3 [29]. However, Jesus et al. [36] demonstrated, through phylogenetic analysis, the introduction of another lineage of DENV-2-III in São Paulo State, called DENV-2-III BR-4.
In this study, we sequenced 22 DENV-2 samples, obtained for routine diagnosis in a Hospital of São José do Rio Preto, and showed the co-circulation of two lineages belonging to the DENV-2 Genotype III: DENV-2-III BR3 and DENV-2-III BR4 (BR4L1 and BR4L2). The findings suggest no evidence of phylogenetic clustering of cases with warning signs, indicating that it may be challenging to distinguish patients who will evolve to more severe clinical forms using only information at the consensus sequence level. It is important to highlight that sample size is a limitation of our study; thus, our results should be interpreted with caution and as a preliminary analysis. Studies with larger sample sizes and which explore single nucleotide variants are needed.
Chin-inmanu et al. sequenced the variants of 40 samples from patients diagnosed with DENV 1, 2, 3, or 4. They showed that the comparison between the data of the dengue fever (dengue without warning signs) and dengue hemorrhagic fever (dengue with warning signs) specimens was conserved between their major populations with the consensus sequences sharing 99% similarity [37]. Although our study and that of Chin-inmanu et al. have different objectives, both studies were unable to differentiate the three clinical dengue classifications (dengue without warning signs, dengue with warning signs, and severe dengue) by analyzing only sequence variability and phylogenetic relationships.
In 2009, the WHO reclassified dengue disease, differing significantly from the previous clinical classification [4] newest classification correlated clinical findings associated with the natural history of plasma leakage and organ failure and aimed to identify severe dengue [38]. Since then, dengue has been classified as dengue without warning signs, dengue with warning signs, and severe dengue, reflecting its intended use as a case-clinical management tool [4]. Severe dengue may occur by all four DENV serotypes; however, each genotype presents different virulence, which may be noted historically (reviewed by Bhatt et al. [39]). DENV-2 has previously been associated with dengue epidemics, especially when variants emerge subsequently [40]. The model of antibody-dependent enhancement (ADE) has been the most accepted to justify the severe forms of the disease, which occur mainly in heterotypic secondary DENV infection (a distinct DENV serotype from the primary infecting serotype) [39]. However, several other factors play a role in the pathogenesis of severe forms, including non-structural protein 1 (NS1) viral antigen [41], cytokine levels [42], memory cross-reactive T cells [43], age [44], viral serotypes [45, 46], genotypes [47], and lineages [44].
In conclusion, our phylogenetic reconstruction showed that in São José do Rio Preto, two lineages belonging to the American/Asian genotype of DENV-2: BR3 and BR4 (BR4L1 and BR4L2) are co-circulating. Our preliminary results suggest no specific association between DENV-2 clinical form and phylogenetic clustering at the virus consensus sequence level. In this work, we demonstrate that a protocol of genome sequencing using the portable nanopore MinION platform can be used as a methodology for genomic surveillance to monitor viral lineages during an epidemic and that this approach was able to produce quick and reliable sequences. Given our sample size limitation, larger studies are needed to confirm our findings, but the data does encourage us to follow this approach.
Data Availability
All sequences generated and analyzed in this study are available in the GenBank database under the accession numbers ON123816-ON123833.
References
Bhatt S, Gething PW, Brady OJ et al (2013) The global distribution and burden of dengue. Nature 496(7446):504–507. https://doi.org/10.1038/nature12060
World Health Organization (2021) Global strategy for dengue prevention and control 2012–2020. Accessed April 24, 2023. https://apps.who.int/iris/bitstream/handle/10665/75303/9789241504034_eng.pdf;jsessionid=5FFB71EB2A8FE6C940F83C7F7B57CB28?sequence=1
Lindenbach BD, Murray CL, Thiel HJ, Rice C (2013) Flaviviridae. In: Fields Virology (6 ed). Philadelphia, USA, pp 712–746
World Health Organization (2021) Dengue: guidelines for diagnosis, treatment, prevention and control available. Accessed April 24, 2023. https://apps.who.int/iris/bitstream/handle/10665/44188/9789241547871_eng.pdf?sequence=1&isAllowed=y
Chen R, Vasilakis N (2011) Dengue-Quo tu et quo vadis? Viruses 3(9):1562–1608. https://doi.org/10.3390/v3091562
Weaver SC, Vasilakis N (2009) Molecular evolution of dengue viruses: contributions of phylogenetics to understanding the history and epidemiology of the preeminent arboviral disease. Infect Genet Evol 9(4):523–540. https://doi.org/10.1016/j.meegid.2009.02.003
Faria NR, Kraemer MUG, Hill SC et al (2018) Genomic and epidemiological monitoring of yellow fever virus transmission potential. Science 361(6405):894–899. https://doi.org/10.1126/science.aat7115
Faria NR, Quick J, Claro IM et al (2017) Establishment and cryptic transmission of Zika virus in Brazil and the Americas. Nature 546(7658):406–410. https://doi.org/10.1038/nature22401
Secretaria de Vigilância em Saúde - Ministério da Saúde (2020) Monitoramento Dos Casos de Arboviroses Urbanas Transmitidas Pelo Aedes Aegypti (Dengue, Chikungunya e Zika), Semanas Epidemiológicas 1 a 46, 2020. Accessed April 24, 2023. https://www.gov.br/saude/pt-br/centrais-de-conteudo/publicacoes/boletins/epidemiologicos/edicoes/2020/boletim_epidemiologico_svs_48.pdf
Cidade Brasil (2021) Município de São José do Rio Preto. Published. Accessed April 24, 2023. https://www.cidade-brasil.com.br/municipio-sao-jose-do-rio-preto.html
Instituto Brasileiro de Geografia e Estatística. São José do Rio Preto. Published 2022. Accessed April 24, 2023. https://www.ibge.gov.br/cidades-e-estados/sp/sao-jose-do-rio-preto.html
Manandhar KD, McCauley M, Gupta BP et al (2021) Whole genome sequencing of dengue virus serotype 2 from two clinical isolates and serological profile of dengue in the 2015–2016 Nepal outbreak. Amer Trop Med Hyg 104(1):115–120. https://doi.org/10.4269/AJTMH.20-0163
Chiaravalloti NF (1997) Description of Aedes aegypti colonization in the region of São José Do Rio Preto. Published online, São Paulo
Mondini A, de Moraes Bronzoni RV, Nunes SHP et al (2009) Spatio-temporal tracking and phylodynamics of an urban dengue 3 outbreak in São Paulo, Brazil. PLoS Negl Trop Dis 3(5). https://doi.org/10.1371/journal.pntd.0000448
Rocco IM, Silveira VR, Maeda AY et al (2012) Primeiro isolamento de dengue 4 no estado de são paulo, Brasil, 2011. Rev Inst Med Trop Sao Paulo 54(1):49–51. https://doi.org/10.1590/S0036-46652012000100009
Secretaria de Saúde - Prefeitura de São José do Rio Preto. Boletim de Dengue. Accessed April 24, 2023. https://saude.riopreto.sp.gov.br/transparencia/boletim_dengue_saude_riopreto.php
Johnson BW, Russell BJ, Lanciotti RS (2005) Serotype-specific detection of dengue viruses in a fourplex real-time reverse transcriptase PCR assay. J Clin Microbiol 43(10):4977–4983. https://doi.org/10.1128/JCM.43.10.4977-4983.2005
Hill SC, de Vasconcelos JN, Granja BG et al (2019) Early genomic detection of cosmopolitan genotype of dengue virus serotype 2, Angola, 2018. Emerg Infect Dis 25(4):784–787. https://doi.org/10.3201/eid2504.180958
Quick J, Grubaugh ND, Pullan ST et al (2017) Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc 12(6):1261–1266. https://doi.org/10.1038/nprot.2017.066
Wick RR, Judd LM, Holt KE (2019) Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol 20(1). https://doi.org/10.1186/s13059-019-1727-y
Hill SC, de Souza R, Thézé J et al (2020) Genomic surveillance of yellow fever virus epizootic in São Paulo, Brazil, 2016 – 2018. PLoS Pathog 16(8). https://doi.org/10.1371/JOURNAL.PPAT.1008699
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Loman NJ, Quick J, Simpson JT (2015) A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat Methods 12(8):733–735. https://doi.org/10.1038/nmeth.3444
Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30(22):3276–3278. https://doi.org/10.1093/bioinformatics/btu531
Trifinopoulos J, Nguyen LT, von Haeseler A, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44(W1):W232–W235. https://doi.org/10.1093/NAR/GKW256
Sagulenko P, Puller V, Neher RA (2018) Treetime: maximum-likelihood phylodynamic analysis. Virus Evol 4(1). https://doi.org/10.1093/ve/vex042
Yenamandra SP, Koo C, Chiang S et al (2021) Evolution, heterogeneity and global dispersal of cosmopolitan genotype of dengue virus type 2. Sci Rep 11(1). https://doi.org/10.1038/s41598-021-92783-y
Fonseca V, Libin PJK, Theys K et al (2019) A computational method for the identification of dengue, zika and chikungunya virus species and genotypes. PLoS Negl Trop Dis 13(5). https://doi.org/10.1371/journal.pntd.0007231
Drumond BP, Mondini A, Schmidt DJ, de Bronzoni RVM, Bosch I, Nogueira ML (2013) Circulation of different lineages of dengue virus 2, genotype American/Asian in Brazil: dynamics and molecular and phylogenetic characterization. PLoS One 8(3). https://doi.org/10.1371/journal.pone.0059422
Tian H, Sun Z, Faria NR et al (2017) Increasing airline travel may facilitate co-circulation of multiple dengue virus serotypes in Asia. PLoS Negl Trop Dis 11(8). https://doi.org/10.1371/journal.pntd.0005694
Nunes MRT, Palacios G, Faria NR et al (2014) Air travel is associated with intracontinental spread of dengue virus serotypes 1–3 in Brazil. PLoS Negl Trop Dis 8(4). https://doi.org/10.1371/journal.pntd.0002769
Carneiro AR, Cruz ACR, Vallinoto M (2012) Molecular characterisation of dengue virus type 1 reveals lineage replacement during circulation in Brazilian territory. https://doi.org/10.1590/S0074-02762012000600016
de Bruycker-Nogueira F, Mir D, dos Santos FB, Bello G (2016) Evolutionary history and spatiotemporal dynamics of DENV-1 genotype V in the Americas. Infect Genet Evol 45:454–460. https://doi.org/10.1016/j.meegid.2016.09.025
Mir D, Romero H, Fagundes De Carvalho LM, Bello G (2014) Spatiotemporal dynamics of DENV-2 Asian-American genotype lineages in the Americas. PLoS One 9(6). https://doi.org/10.1371/journal.pone.0098519
Secretaria da Saúde - Governo do Estado de São Paulo (2019) Distribuição Dos Casos de Dengue Notificados e Confirmados (Autóctones e Importados) Segundo Município de Residência, Por Mês de Início de Sintomas, ESP, 2019. Accessed April 24, 2023. https://www.saude.sp.gov.br/resources/cve-centro-de-vigilancia-epidemiologica/areas-de-vigilancia/doencas-de-transmissao-por-vetores-e-zoonoses/dados/dengue/2019/dengue19_import_autoc_res.htm
de Jesus JG, Dutra KR, Sales FC da S et al (2020) Genomic detection of a virus lineage replacement event of dengue virus serotype 2 in Brazil, 2019. Mem Inst Oswaldo Cruz 115(4). https://doi.org/10.1590/0074-02760190423
Chin-Inmanu K, Mairiang D, Khongthon P et al (2022) Genetic diversity of the dengue virus population in dengue fever and dengue hemorrhagic fever patients. Asian Pac J Allergy Immunol. https://doi.org/10.12932/ap-230620-0887
Srikiatkhachorn A, Rothman AL, Gibbons RV et al (2011) Dengue-how best to classify it. Clin Infect Dis 53(6):563–567. https://doi.org/10.1093/cid/cir451
Bhatt P, Sabeena SP, Varma M, Arunkumar G (2021) Current understanding of the pathogenesis of dengue virus infection. Curr Microbiol 78(1):17–32. https://doi.org/10.1007/s00284-020-02284-w
Oliveira MF, Araújo JMG, Ferreira OC et al (2010) Two lineages of dengue virus type 2. Brazil Emerg Infect Dis 16(3):576–578. https://doi.org/10.3201/eid1603.090996
Modhiran N, Watterson D, Muller DA et al (2015) Dengue virus NS1 protein activates cells via toll-like receptor 4 and disrupts endothelial cell monolayer integrity. Sci Trans Med 7(304):304ra142. https://doi.org/10.1126/scitranslmed.aaa3863.
Raj Kumar Patro A, Mohanty S, Prusty BK et al (2019) Cytokine signature associated with disease severity in dengue. Viruses 11(1). https://doi.org/10.3390/v11010034
Kurane I, Matsutani T, Suzuki R et al (2011) T-cell responses to dengue virus in humans. Trop Med Health 39(4 SUPPL.):45–51. https://doi.org/10.2149/tmh.2011-S09
Nunes PCG, Sampaio SAF, Rodrigues da Costa N et al (2016) Dengue severity associated with age and a new lineage of dengue virus-type 2 during an outbreak in Rio De Janeiro. Brazil. J Med Virol 88(7):1130–1136. https://doi.org/10.1002/jmv.24464
Vicente CR, Herbinger KH, Fröschl G, Romano CM, Cabidelle A de SA, Junior CC (2016) Serotype influences on dengue severity: a cross-sectional study on 485 confirmed dengue cases in Vitória, Brazil. BMC Infect Dis 16(1). https://doi.org/10.1186/s12879-016-1668-y
Rocha BAM, Guilarde AO, Argolo AFLT et al (2017) Dengue-specific serotype related to clinical severity during the 2012/2013 epidemic in centre of Brazil. Infect Dis Poverty 6(1). https://doi.org/10.1186/s40249-017-0328-9
Williams M, Mayer SV, Johnson WL et al (2014) Lineage II of southeast Asian/American DENV-2 is associated with a severe dengue outbreak in the Peruvian Amazon. Am J Trop Med Hyg 91(3):611–620. https://doi.org/10.4269/ajtmh.13-0600
Funding
Funding support is acknowledged from the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP 2019/21711–9) and the Brazil-UK Centre for Arbovirus Discovery, Diagnosis, Genomics and Epidemiology (CADDE).
Author information
Authors and Affiliations
Contributions
NRF and MLN conceived the study and its design; BCM, DSC, KRD, FCSS, and JGJ conducted the molecular investigation, genome sequencing, and genome assembling; BCM and LS performed the phylogenetic analysis; BCM, CFE, and FSD carried out clinical data analyses. BCM, LS, CAB, CFE, DSC, and MLN wrote and revised the manuscript. NRF and MLN provided the resources. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This data is available as an abstract in the annual book of 2021 Annual Meeting of The American Society of Tropical Medicine and Hygiene. The abstract can be accessed at: https://www.astmh.org/getmedia/59a95de8-1a06-49ca-9fd0-286454cc241a/ASTMH-2021-Annual-Meeting-Abstract-Book.pdf.
Responsible Editor: Jônatas Abrahão
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
de Carvalho Marques, B., Sacchetto, L., Banho, C.A. et al. Genetic differences of dengue virus 2 in patients with distinct clinical outcome. Braz J Microbiol 54, 1411–1419 (2023). https://doi.org/10.1007/s42770-023-01006-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-023-01006-1