Abstract
The Xanthomonadaceae family comprises the genera Xanthomonas and Xylella, which include plant pathogenic species that affect economically important crops. The family also includes the plant growth-promoting bacteria Pseudomonas geniculata and Stenotrophomonas rhizophila, and some other species with biotechnological, medical, and environmental relevance. Previous work identified molecular signatures that helped to understand the evolutionary placement of this family within gamma-proteobacteria. In the present study, we investigated whether insertions identified in highly conserved proteins may also be used as molecular markers for taxonomic classification and identification of members within the Xanthomonadaceae family. Four housekeeping proteins (DNA repair and replication-related and protein translation enzymes) were selected. The insertions allowed discriminating phytopathogenic and plant growth-promoting groups within this family, and also amino acid sequences of these insertions allowed distinguishing different genera and, eventually, species as well as pathovars. Moreover, insertions in the proteins MutS and DNA polymerase III (subunit alpha) are conserved in Xylella fastidiosa, but signatures in DNA ligase NAD-dependent and Valyl tRNA synthetase distinguish particular subspecies within the genus. The genus Stenotrophomonas and Pseudomonas geniculata could be distinguishable based on the insertions in MutS, DNA polymerase III (subunit alpha), and Valyl tRNA synthetase, although insertion in DNA ligase NAD-dependent discriminates these bacteria at the species level. All these insertions differentiate species and pathovars within Xanthomonas. Thus, the insertions presented support evolutionary demarcation within Xanthomonadaceae and provide tools for the fast identification in the field of these bacteria with agricultural, environmental, and economic relevance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xanthomonadales is an order of bacteria currently classified in the class Gamma-proteobacteria. The species of this order comprise bacteria that differ in lifestyles and inhabit a variety of environments, including some that live in extreme conditions [1]. In previous studies, molecular signatures have been identified in highly conserved DNA repair and replication-related proteins that provide evidence for the evolutionary placement of Xanthomonadales. This order is one of the deepest branching lineages within the Gamma-proteobacteria clade, and some signatures exclusive to certain genera included in Xanthomonadales have been identified [2]. Based on the study of Conserved Signature Indels (insertions and deletions—CSIs) and the result of phylogenetic analysis performed on a concatenated sequence alignment of 15 highly conserved housekeeping proteins, Naushad and coworkers proposed the division of Xanthomonadales (synonym for the order Lysobacterales) into two families, Xanthomonadaceae and Rhodanobacteraceae. These authors described 10 CSIs specific for Xanthomonadaceae and 11 specific for Rhodanobacteraceae [3].
The family Xanthomonadaceae, proposed by Naushad and colleagues as emended description of the family Lysobacteraceae, includes the genera Arenimonas, Luteimonas, Lysobacter, Metallibacterium, Panacagrimonas, Pseudoxanthomonas, Silanimonas, Stenotrophomonas, Thermomonas, Xanthomonas, and Xylella [3]. This family comprises phytopathogenic bacteria that produce serious agricultural and economic impacts, mainly grouped within the genera Xanthomonas and Xylella [4]. For example, Xanthomonas harbors a vast number of species in which all reported strains are plant-associated and most are recognized as being pathogenic to particular plant hosts such as rice [5], citrus and crucifers [6, 7], tomatoes and pepper [8], onion and garlic [9], sugarcane [10], and aroids [11]. Similarly, Xylella fastidiosa causes Pierces’s disease of grapevines [12], variegated chlorosis in citrus [13], olive quick decline syndrome [14], and leaf scorch diseases of almonds and oleanders [15].
In contrast, members of the genus Lysobacter represent a source of biocontrol agents against a broad range of phytopathogenic fungi [16]. The species L. enzymogenes strain 3.1T8 inhibits mycelial growth of Phytophthora aphanidermatum, which causes damping-off in many plants, including the tobacco (Nicotiana tabacum L.) [17]. Moreover, L. enzymogenes strain B25 has been reported to promote plant growth and protect plants against parasitic nematodes [18].
In addition to these genera, the family also includes the genus Stenotrophomonas that harbors S. maltophilia, a multidrug-resistant opportunist human-pathogen species responsible for important nosocomial infections [19]. However, three S. maltophilia strains isolated from healthy tomato plants have been identified as plant growth-promoting bacteria able to produce indole-3-acetic acid; two of these strains had phosphate solubilization ability [20]. Moreover, S. maltophilia P9 was recovered from algal biomass and identified as capable of producing pectinase with biotechnological relevance [21]. Some other genera are ecologically important and may be used for soil and water biodegradation. Members of the genus Pseudoxanthomonas, for example, have been isolated from biofilters, anaerobic digesters, organic fertilizers, human urine, and urban riverside soil [22].
Proteomics studies and the availability of completely sequenced genomes for a large number of bacteria constitute an opportunity for taxonomic and phylogenetic studies. The alignment of homologous protein sequences from different species makes possible the identification of CSIs. These insertions and deletions can be specific for taxonomic clades placed at different branching depths and represent molecular synapomorphies useful to identify and demarcate specific bacterial groups in clear molecular terms [23, 24].
Because Xanthomonadaceae includes not only important plant and human pathogens but also species with biotechnological and environmental relevance, studies on this important family are necessary. The order Xanthomonadales has been extensively studied, but few studies have been conducted to identify CSIs for taxonomy purposes specifically within Xanthomonadaceae [2, 3]. These CSIs could be used as molecular markers to distinguish members of this family from other groups of bacteria, considering that the 16S rRNA gene sequence has shown limited ability to resolve the relationships among members from Xanthomonadales [2, 3].
Insertions in highly conserved proteins have been used to assess the evolutionary placement of Xanthomonadales [2]. In the present study, we investigated whether these insertions may be used as molecular markers to delineate subgroups within Xanthomonadaceae and distinguish genera with agricultural, environmental, and economic relevance within this diverse family, based on variations in amino acid residues.
Materials and methods
Identification of conserved signature sequences
In this work, we carried out comparative proteomic analyses on members of Xanthomonadaceae, including species from Rhodanobacteraceae family and other proteobacteria as outgroup. Four housekeeping proteins were analyzed: DNA polymerase III (subunit alpha), NAD-dependent DNA ligase, Valyl tRNA synthetase, and the mismatch repair protein MutS, which evidenced useful signatures in previous work [2, 3]. The evolutionary rate for these proteins in Xanthomonadaceae varies from 0.85 for MutS to 1.08 for DNA ligase NAD-dependent (https://www.orthodb.org) [25].
The ortholog sequences were mainly obtained from the UniprotKB/Swiss-Prot database available at the Universal Protein Resource (UniProt) and increased from the GenBank non-redundant databases by similarity BLASTp searches [26, 27], using the default parameters on each protein and the sequences of Xylella fastidiosa 9a5c as query. The result of blast searches was analyzed in order to select high scoring similar proteins from Xanthomonadaceae, those with E value < 0.001, identity > 35%, and Bit score > 50 [28]. To assess the specificity of the insertions identified, recursive BLASTp searches against the NCBI database were performed, using as query the sequence region containing the insertion and the flanking regions [2, 3].
Multiple sequence alignments were performed using ClustalX2 [29] by progressive alignment strategies, and the alignment parameters considered were pairwise alignment (gap opening, 35.0 and gap extension, 0.75) and multiple alignment (gap opening, 15.0 and gap extension, 0.30) [30]. The results of the sequence alignments were analyzed by visual inspection in order to identify CSIs. Only insertions were considered for this work and those flanked on both sides by at least 5 conserved residues in the neighboring 30–40 amino acids were selected [31].
Phylogenetic analysis
Phylogenetic relations were inferred through Maximum Likelihood (ML) and Bayesian inference (BI). Protein trees were constructed for a concatenated sequence alignment of the four proteins and for each protein alignment sequence independent. Escherichia coli, Pseudomonas aeruginosa, and several Rhodanobacteraceae were used as outgroups in all analyses.
Models of molecular evolution for each independent protein alignment sequence were selected through Bayesian information criterion (BIC) with ProtTest. In the case of concatenated sequences, best-fit partitioning schemes and models of molecular evolution for each partition were selected through the BIC with PartitionFinderProtein v1.1.0 [32].
The ML and BI analysis were conducted with RaxML-HPC2 and MrBayes v.3.2.6 respectively through the web portal Cyberinfrastructure for Phylogenetic Research: CIPRES (http://www.phylo.org) [33, 34]. ML trees were inferred by heuristic searches with the Randomized Axelerated Maximum Likelihood algorithm. The robustness of ML inferences was assessed with a bootstrap analysis. One thousand bootstrap replicates were set automatically with the “autoMRE” criterion (auto Majority Rule Criterion) [35]. The BI analyses were conducted with default priors in two independent runs of 2 million generations (sampled each 1000 generations) and four Markov-chain Monte Carlo (MCMC). Convergence of the two runs onto a stationary distribution was assessed by examining the average standard deviation of the split frequencies, in MrBayes v.3.2.6 [34] and trace plots and effective sample size (ESS) of the parameters (ESS > 200 was considered acceptable) in Tracer v.1.7 [36]. Burn-in samples (25%) were discarded, and the remaining samples were combined in a majority rule consensus tree with node posterior probabilities equal to bipartition frequencies.
Results
The insertions identified in the proteins NAD-dependent DNA ligase and MutS were reported in a previous work, but only species from the genera Xanthomonas, Xylella, and Stenotrophomonas were included [2]. The presence of these signatures, in addition to the insertion evaluated in the protein Valyl tRNA synthetase, was later analyzed for a larger number of genera in the order [37].
In the present work, the signatures were evaluated specifically for members of the family Xanthomonadaceae (emended description of Lysobacteraceae) including some genera that were not analyzed before such as Arenimonas, Luteimonas, Lysobacter, Silanimonas, and Thermomonas and the species Pseudomonas geniculata [2, 37]. We investigated whether these insertions may be used as molecular markers useful to distinguish subgroups within Xanthomonadaceae, as a continuation of the study in which the signatures were analyzed only to uncover taxonomic relationships of the Xanthomonadales order within the proteobacteria based on the presence or absent of the markers [2].
The analysis of the proteins evidenced four signature insertions in important housekeeping proteins, which are summarized in Table 1. The first one is a five amino acid insertion (variable in sequence) found in the mismatch repair protein MutS (Fig. 1). This signature is present in the genera Xanthomonas, Xylella, and Stenotrophomonas, and curiously is shared with the species Pseudomonas geniculata, Pseudoxanthomonas spadix, Lysobacter arseniciresistens, and Luteimonas abyssi. Most of the members from the genera Lysobacter and Pseudoxanthomonas, in addition to Luteimonas, Arenimonas, Thermomonas, and Silanimonas, lack the insertion which is also absent in members from the family Rhodanobacteraceae and in other proteobacteria.
Partial sequence alignment of the protein MutS, showing a five amino acid length insertion present in some genera of Xanthomonadaceae family. Dots (.) in this and other alignments denote identity with the amino acid on the top line and gaps are indicated by empty spaces; the insertion is highlighted by a box. Accession numbers are given after each species name. CorelDRAW Graphics Suite 2019 was used to create the artwork
All the subspecies of Xylella fastidiosa included share the amino acid sequence TYEGG in this insertion. Despite most of the species belonging to Xanthomonas share the sequences SHAGG, this marker has some variation within the genus, allowing some species discrimination; the sequence for X. albilineans is PVEGG and X. translucens has valine and glutamine at the positions 2 and 3, respectively. Moreover X. translucens pv. translucens has valine at the second position and glutamic acid at the third position and X. retroflexus has glutamine at the first position and glutamic acid at the position 3.
The genus Stenotrophomonas shares the amino acid sequence QHEGG with the species Pseudomonas geniculata, except for S. acidaminiphila, which has proline instead of glutamine at the first position. Otherwise, Pseudoxanthomonas spadix and Luteimonas abyssi share the sequence SFEGG, whereas Lysobacter arseniciresistens has the sequence PNRSG.
Another signature, not reported in previous works, is a 14–15 amino acid insertion identified in the protein DNA polymerase III, subunit alpha. This marker is presents in the genera Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, Thermomonas, and Pseudomonas geniculata, but absent in homologs from the genera Luteimonas, Lysobacter, Arenimonas, and Silanimonas, as well as the family Rhodanobacteraceae and the other proteobacteria (Fig. 2). The amino acid sequence of this insertion distinguishes the different genera sharing it. For example, the homolog sequences from Xylella fastidiosa are characterized by a conserved insertion that, considering the amino acid residues (EKPEQAHPKKAKQA), distinguishes these bacteria from the rest of members of Xanthomonadaceae. The insertion present in Xanthomonas is also distinctive in amino acid sequences for this genus (EKPDQADPKKAKQA), although the species X. fragariae has asparagine at the position 4 and Xanthomonas citri is differentiated by a threonine at the second position.
A similar distinction is possible to observe for Stenotrophomonas maltophilia, S. rhizophila, the three strains of Stenotrophomonas sp. and Pseudomonas geniculata based on the shared conserved sequences of their insertion (AKPDQADPKKAKQA) which, as the signature described in MutS, support a close relation for these species. In contrast, the insertion in Pseudoxanthomonas has variations at the positions 5, 6, 7, 13, and 14 differentiating members of this genus. Whereas the species Thermomonas fusca has a very particular sequence in its one amino acid longer signature.
The protein DNA ligase NAD-dependent is characterized by a 39–49 amino acid insertion common to the genera Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, and Luteimonas and the species Pseudomonas geniculata, but is not found in homologs from Arenimonas, Lysobacter, Thermomonas, and Silanimonas, neither in homologs from the family Rhodanobacteraceae and other proteobacteria (Fig. 3). The fact that Luteimonas mephitis and Pseudoxanthomonas suwonensis also lack the insertion points to an old divergence of these species within their specific groups. This signature has some variation in length, indicating that the region of the protein affected has some structural flexibility due to low-selective pressure [2]. The most parsimonious explanation is that the insertion probably arose in the branch including the genera Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, Luteimonas, and Pseudomonas geniculata and the event was followed by deletions in specific lineages.
The insertion in Xanthomonas and Xylella is long (48–49 aa) and conserved, whereas the other genera have a smaller insert that differs in sequences. Based on the insertion size, this marker is useful to differentiate these two genera from other members of the family. However, based on the amino acid sequence, we can distinguish Xanthomonas and Xylella; although the insertion present in Xylella fastidiosa has some variation at the 39th position for X. fastidiosa 9a5c, X. fastidiosa Mul-MD, and X. fastidiosa Temecula, which share a lysine at this position, the rest of the sequence is conserved (PTMLLREARDHVTGMRYQQLEEILRTVGVDLSGEGDVPEHWQIDVLRA). This signature has variations in sequence for Xanthomonas that allows for the distinguishing of species within this genus. The species Stenotrophomonas maltophilia, Stenotrophomonas sp. SKA14, S. rhizophila, and P. geniculata share a 40 aa length insertion which has variations that distinguish these bacteria with each other, especially for S. rhizophila. The signature was also previously reported, but some genera belonging to the family and the species Pseudomonas geniculata were not included and the amino acid sequence was not analyzed [37].
On the other hand, the alignment of the protein Valyl tRNA synthetase evidenced a 19 amino acid length insertion in a highly conserved region, common to the genera Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, Luteimonas, Thermomonas, and Silanimonas and the species Pseudomonas geniculata, Arenimonas metalli and A. composti. The signature is absent in the genus Lysobacter, most of the species that belong to the genus Arenimonas and also in the family Rhodanobacteraceae and the outgroup species Neisseria meningitidis, Geobacter metallireducens, and Geobacter sulfurreducens (Fig. 4).
The amino acid sequence has some variations at the positions 5 and 14 in Xylella fastidiosa. The subspecies Xylella fastidiosa 9a5c has valine instead of isoleucine at the 5th position, and also shares an isoleucine with X. fastidiosa Dixon at the position 14 which contrasts with the threonine shared by X. fastidiosa Ann-1 and X. fastidiosa Temecula. This signature has also variations within Xanthomonas at the positions 1, 5, 7, 12, and 13 that distinguish species within this genus (Xanthomonas translucens pv. translucens, X. axonopodis, X. campestris, X. fragariae, and X. albilineans). On the other hand, Stenotrophomonas maltophilia, Stenotrophomonas sp. RIT309, S. rhizophila, and Pseudomonas geniculata have the same amino acid residues in the insertion shared by these bacteria (SYEHVERDADGVETLRETR). This signature distinguished species within the genera Pseudoxanthomonas and it is very variable in Luteimonas, Thermomonas, and Silanimonas and even in the two species of Arenimonas that share it.
This insertion has also been reported before for the genera Pseudoxanthomonas, Stenotrophomonas, Xanthomonas, and Xylella, and curiously partial insertion (6 amino acids) was shared by several species belonging to the class Alphaproteobacteria but having different amino acid sequence. The authors concluded that the shared presence of the CSIs in these two groups was the result of independent events [37]. In the present study, smaller insertions that differ in sequences from those observed in the in-group species were also observed in Pseudomonas aeruginosa, Escherichia coli, Vibrio vulnificus, Ralstonia metallidurans, Bradyrhizobium japonicum, and Mesorhizobium loti that were included as outgroup species.
Discussion
The identification of CSIs constitutes a reliable and consistent method for taxonomic and diagnosis studies and has been applied before to determine evolutionary relationships [38, 39] and specifically in Xanthomonadales [2, 3, 37]. We selected DNA repair and replication-related and protein translation enzymes which display slow rates of sequence evolution allowing them to be used for phylogenetic analysis. These evolutionarily conserved proteins participate in informational processes and their function is dependent upon interaction with other proteins [2, 3, 37].
Figure 5 shows schematic evolutionary flow of Xanthomonadaceae based on the insertions identified. The signatures found separate two main groups, the first one includes those genera that share all or most of the insertions analyzed, and the second group is formed by the genera lacking all these signatures. The insertions are likely molecular synapomorphies that distinguish the lineages Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, Luteimonas, Thermomonas, and Pseudomonas geniculata, from Arenimonas and Lysobacter which most of their species lack the insertions.
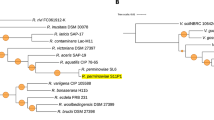
This inference is congruent with the result of the phylogenetic analysis. The two concatenated protein sequence–based phylogenetic trees obtained by performing different methods (BI and ML) exhibited similar branching order for each branch (Fig. 6). Members of the family Xanthomonadaceae are divided into two subgroups. One group consisting of species that belong to the genera Xanthomonas, Xylella, Stenotrophomonas, Pseudoxanthomonas, and Thermomonas and the species Pseudomonas geniculata formed a well-supported monophyletic group, confirming that these members share a common ancestor. The second group is not forming a monophyletic group but branched out of the main group. This last group includes the genera Arenimonas and Lysobacter, which lack all the insertions identified (except for A. composti and A. metalli, which share the insertion found in the protein Valyl tRNA synthetase), and based on the phylogenetic analyses are confirmed as early-branching lineages within the family.
The monophyletic grouping of Xanthomonas, Xylella, Stenotrophomonas, and Pseudomonas geniculata based on the shared presence of the insertion in MutS (Fig. 1) is congruent with the phylogenetic analysis in which this clade is supported by a high bootstrap and posterior probability values (Fig. 6). Considering that the ecological niche of this group includes species with agricultural relevance, the ancestor could be plant-associated bacteria. These phylogenetic inferences are in agreement with the phylogenomic analyses obtained in previous studies in which those genera are placed in the same branch [3].
Moreover, Xanthomonas and Xylella are forming a well-supported monophyletic group in the concatenated phylogenetic tree (Fig. 6). These two genera are differentiated from other members of Xanthomonadaceae based on their characteristic longer insertion in the protein NAD-dependent DNA ligase. One important point is that the family harbors many plant pathogens that affect economically important crops and plants specifically included in Xanthomonas and Xylella. Thus, the insertion could be considered a useful tool for molecular identification of these important phytopathogenic bacteria.
The species Pseudoxanthomonas spadix, Lysobacter arseniciresistens, and Luteimonas abyssi also present the insertion in the protein MutS. Curiously, Pseudoxanthomonas spadix and Luteimonas abyssi share the same amino acid sequence, which differs from the in-group homologs. On the contrary, the specific amino acid sequence of Lysobacter arseniciresistens is not shared with any other genera. The relationship of Pseudoxanthomona spadix with the clade formed by Xylella, Xanthomonas, Stenotrophomonas, and Pseudomonas geniculata is supported by a high bootstrap and posterior probability values in the concatenated phylogenetic tree (Fig. 6), confirming the proximity of these groups of bacteria as already demonstrated in previous studies [3]. However, the trees do not support a monophyletic grouping of Lysobacter arseniciresistens with this clade. Instead, this species branched with other members of its genus, confirming that the shared presence of the insertion in the protein MutS constitutes a homoplasy for L. arseniciresistens due to independent genetic events.
Molecular approaches are being used increasingly not only in evolutionary and taxonomic studies, but also in epidemiological studies. The insertions identified are well-defined in size, except for NAD-dependent DNA ligase, which has variations in length, and most of these have a strong degree of conservation (in their amino acid sequences) within specific genera or species, and are also flanked by highly conserved regions in the proteins. Because of their specificity and conservation, these are useful as molecular markers for detection and identification of both known as well as unknown bacteria belonging to specific genera or species in different environments by means of PCR-based (polymerase chain reaction) and immunological methods, as well as by in silico similarity BLASTp searches.
The insertions identified in the proteins MutS and DNA polymerase III (subunit alpha) are highly conserved in all the subspecies of Xylella fastidiosa analyzed (Figs. 1 and 2). This characteristic is very important and may be of practical significance, as PCR-based protocols could possibly contribute to pathogen detection and identification at the species level. However, the signatures in DNA ligase NAD-dependent and Valyl tRNA synthetase have some variations in sequence within this species, which distinguish particular subspecies (Figs. 3 and 4). The 39th position of the insertion present in DNA ligase NAD-dependent distinguishes Xylella fastidiosa 9a5c, X. fastidiosa Mul-MD, and X. fastidiosa Temecula from Xylella fastidiosa M12, X. fastidiosa Ann 1, and X. fastidiosa Dixon. On the other hand, the positions 5 and 14 of the insertion in Valyl tRNA synthetase differentiate Xylella fastidiosa 9a5c and X. fastidiosa Dixon from X. fastidiosa Ann-1 and X. fastidiosa Temecula. The diagnosis of X. fastidiosa is very important in terms of phytosanitary regulations because this bacterium constitutes a regulated phytopathogen in many parts of the world [40].
Moreover, the control of Xanthomonas is very difficult and requires the use of economically and environmentally unsatisfactory strategies. The bacterium X. axonopodis pv. allii, for example, is associated with outbreaks on Allium cepa L., A. fistulosum L., A. sativum L., A. porrum L., A. schoenoprasum L., and A. cepa var. ascalonicum. The lesions produced by the pathogen in onion result in a reduction of bulb size and, consequently, in yield losses [9]. On the other hand, X. campestris pv. vesicatoria causes bacterial leaf spot (BLS) in tomato (Solanum lycopersicum L.) and capsicum or chili (Capsicum annuum L.) which results in extensive damage to crops. The identification of these bacteria at the genus level can be achieved through sequencing of 16S rDNA or characterization of xanthomonadin pigments, but for identification at the pathovar level, sequence-based PCR is used. However, the biochemical tests and the species-specific PCR protocol currently used may fail to detect X. campestris pv. vesicatoria isolates [41].
These signatures could be used as molecular markers to design PCR-based protocols for identification at the species level in Xanthomonas. The insertion in MutS discriminates species and pathovars considering that X. albilineans, X. translucens, X. translucens pv. translucens, and X. retroflexus have specific variations in sequences that distinguish them from other members of the genus. Similarly, amino acid sequence of the insertion in DNA polymerase III, subunit alpha, differentiates X. fragariae and X. citri in addition to the insertion in Valyl tRNA synthetase which distinguishes the species X. axonopodis, X. campestris, X. fragariae, X. translucens pv. translucens, and X. albilineans. Furthermore, the variability in sequence of the long insertion in NAD-dependent DNA ligase differentiates species within this genus.
However, the most important challenges in agriculture are not only the identification of pathogenic bacterial isolates but also the isolation and identification of potential bacterial candidates with plant growth-promoting traits. Based on the amino acid sequence of the insertion shared by the species belonging to the genus Stenotrophomonas and Pseudomonas geniculata in the proteins DNA polymerase III (subunit alpha) and Valyl tRNA synthetase, these bacteria could be distinguishable from the rest of genera included in the family (Figs. 2 and 4). In the concatenated tree (Fig. 6), P. geniculata is also included in the well-supported monophyletic group of Stenotrophomonas, as previously reported [4]. This affiliation of P. geniculata to the cluster of the genus Stenotrophomonas was analyzed in earlier studies based on 16S rRNA sequence; this species does not group with other pseudomonas and is misclassified as Pseudomonas [42]. Thus, the current nomenclature of this species requires substantial revision.
Based on the insertion estimated in the protein MutS, the genus Stenotrophomonas shares the same amino acid sequence with the species Pseudomonas geniculata, but the homolog sequence of the specie S. acidaminiphila varies at the first position. The 40 aa length insertion characteristic of these groups in DNA ligase NAD-dependent has also variations in sequence that allow the discrimination of these bacteria at the species level.
These findings have practical use considering the agricultural importance of these species as plant growth-promoting bacteria. P. geniculata has been reported as endophytic bacteria in stress-resistant plants, and also can be found in the rhizosphere of rice, tobacco, or maize [43]. Similarly, Stenotrophomonas maltophilia and S. rhizophila are associated to the rhizosphere of some crops, especially under saline conditions, and some strains have biotechnological importance [44]. Based on their characteristic and specific insertion in the protein NAD-dependent DNA ligase, the species S. rhizophila and P. geniculata can be distinguished from other groups of bacteria.
Conclusions
The insertions evaluated are useful as molecular signatures for systematics and diagnosis studies. These confirm the current taxonomy of Xanthomonadaceae, support the demarcation of two subgroups within this family, and provide tools for molecular identification of bacteria with agricultural, environmental, and economic relevance.
References
Saddler GS, Bradbury J (2005) Xanthomonadales ord. In: Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual® of systematic bacteriology, Vol. 2, 2nd. Springer, New York, pp 63. https://doi.org/10.1007/0-387-28022-7_3,
Cutiño-Jimenez AM, Martins-Pinheiro M, Lima WC, Martin-Tornet A, Morales O, Martins Menck CF (2010) Evolutionary placement of Xanthomonadales based on conserved protein signature sequences. Mol Phylogenet Evol 54:524–534. https://doi.org/10.1016/j.ympev.2009.09.026
Naushad S, Adeolu M, Wong S, Sohail M, Schellhorn HE, Gupta RS (2015) A phylogenomic and molecular marker based taxonomic framework for the order Xanthomonadales: proposal to transfer the families Algiphilaceae and Solimonadaceae to the order Nevskiales ord. nov. and to create a new family within the order Xanthomonadales, the family Rhodanobacteraceae fam. nov., containing the genus Rhodanobacter and its closest relatives. Antonie Van Leeuwenhoek 107(2):467–485. https://doi.org/10.1007/s10482-014-0344-8
Parte AC (2018) LPSN-list of prokaryotic names with standing in nomenclature (bacterio.net), 20 years on. Int J Syst Evol Microbiol 68(6):1825–1829. https://doi.org/10.1099/ijsem.0.002786
Zhou L, Huang TW, Wang JY, Sun S, Chen G, Poplawsky A, He YW (2013) The Rice bacterial pathogen Xanthomonas oryzae pv. oryzae produces 3-Hydroxybenzoic acid and 4-Hydroxybenzoic acid via XanB2 for use in Xanthomonadin, ubiquinone, and exopolysaccharide biosynthesis. Mol Plant-Microbe Interact 26:1239–1248. https://doi.org/10.1094/MPMI-04-13-0112-R
da Silva AR, Ferro JA, Reinach F, Farah C, Furlan L, Quaggio R et al (2002) Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature 417:459–463. https://doi.org/10.1038/417459a
Palmieri ACB, Amaral AMD, Homem RA, Machado MA (2010) Differential expression of pathogenicity- and virulence-related genes of Xanthomonas axonopodis pv. citri under copper stress. Genet Mol Biol 33:348–353. https://doi.org/10.1590/S1415-47572010005000030
Giovanardi D, Biondi E, Ignjatov M, Jevtić R, Stefani E (2018) Impact of bacterial spot outbreaks on the phytosanitary quality of tomato and pepper seeds. Plant Pathol 67(5):1168–1176. https://doi.org/10.1111/ppa.12839
European and Mediterranean Plant Protection Organization (2016) Data sheets on pests recommended for regulation. Xanthomonas axonopodis pv. allii. Bull OEPP/EPPO 46(1):4–7. https://doi.org/10.1111/epp.12273
Pieretti I, Cociancich S, Bolot S, Carrère S, Morisset A, Rott P, Royer M (2015) Full genome sequence analysis of two isolates reveals a novel Xanthomonas species close to the sugarcane pathogen Xanthomonas albilineans. Genes 6(3):714–733. https://doi.org/10.3390/genes6030714
Constantin EC, Haegeman A, Van-Vaerenbergh J, Baeyen S, Van-Malderghe C et al (2017) Pathogenicity and virulence gene content of Xanthomonas strains infecting Araceae, formerly known as Xanthomonas axonopodis pv. dieffenbachiae. Plant Pathol 66:1539–1554. https://doi.org/10.1111/ppa.12694
Su CC, Chang CJ, Chang CM, Shih HT, Tzeng KC, Jan FJ, Kao CW, Deng WL (2013) Pierce’s disease of grapevines in Taiwan: isolation, cultivation and pathogenicity of Xylella fastidiosa. J Phytopathol 161:389–396. https://doi.org/10.1111/jph.12075
Niza B, Coletta-Filho H, Merfa MV, Takita MA, De Souza AA (2015) Differential colonization patterns of Xylella fastidiosa infecting citrus genotypes. Plant Pathol 64(6):1259–1269. https://doi.org/10.1111/ppa.12381
Almeida RPP, Nunney L (2015) How do plant diseases caused by Xylella fastidiosa emerge? Plant Dis 99:1457–1467. https://doi.org/10.1094/PDIS-02-15-0159-FE
Amanifar N, Taghavi M, Izadpanah K, Babaei G (2014) Isolation and pathogenicity of Xylella fastidiosa from grapevine and almond in Iran. Phytopathol Mediterr 53:318–327. https://doi.org/10.14601/Phytopathol_Mediterr-12647
Puopolo G, Giovannini O, Pertot I (2014) Lysobacter capsici AZ78 can be combined with copper to effectively control Plasmopara viticola on grapevine. Microbiol Res 169:633–642. https://doi.org/10.1016/j.micres.2013.09.013
Olson JD, Damicone JP, Kahn BA (2016) Identification and characterization of isolates of Pythium and Phytophthora spp. from snap beans with cottony leak. Plant Dis 100:1446–1453. https://doi.org/10.1094/PDIS-06-15-0662-RE
Hernández I, Fernández C (2017) Draft genome sequence and assembly of a Lysobacter enzymogenes strain with biological control activity against root knot nematodes. Genome Announc 5:e00271–e00217. https://doi.org/10.1128/genomeA.00214-17
Sánchez MB (2015) Antibiotic resistance in the opportunistic pathogen Stenotrophomonas maltophilia. Front Microbiol 6:658. https://doi.org/10.3389/fmicb.2015.00658
Aydi AR, Jabnoun-Khiareddine H, Nefzi A, Daami-Remadi M (2018) Evaluation of the growth-promoting potential of endophytic bacteria recovered from healthy tomato plants. J Hortic 05:5(2). https://doi.org/10.4172/2376-0354.1000234
Sharma P, Sharma N (2018) Molecular identification, production and optimization of pectinase by using Stenotrophomonas maltophilia P9 isolated from algal biomass of Himachal Pradesh, India. Int J Curr Microbiol App Sci 7(1):670–680. https://doi.org/10.20546/ijcmas.2018.701.082
Nayak AS, Vijaykumar MH, Karegoudar TB (2009) Characterization of biosurfactant produced by Pseudoxanthomonas sp. PNK-04 and its application in bioremediation. Int Biodeterior Biodegradation 63:73–79. https://doi.org/10.1016/j.ibiod.2008.07.003
Gupta RS (2014) Identification of conserved indels that are useful for classification and evolutionary studies. In: Goodfellow M, Sutcliffe L, Jongsik C (eds) Methods in microbiology, vol 41. Academic Press, New York, pp 153–182. https://doi.org/10.1016/bs.mim.2014.05.003
Gupta R (2016) Impact of genomics on the understanding of microbial evolution and classification: the importance of Darwin’s views on classification. FEMS Microbiol Rev 40:520–553. https://doi.org/10.1093/femsre/fuw011
Kriventseva EV, Tegenfeldt F, Petty TJ, Waterhouse RM, Simao FA, Pozdnyakov IA, Zdobnov EM (2015) OrthoDB v8: update of the hierarchical catalog of orthologs and the underlying free software. Nucleic Acids Res 43(D1):D250–D256. https://doi.org/10.1093/nar/gku1220
UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. https://doi.org/10.1093/nar/gky1049
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Pearson WR (2013) An introduction to sequence similarity (“homology”) searching. Curr Protoc Bioinformatics 42(1):3–1. https://doi.org/10.1002/0471250953.bi0301s42
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins D (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882. https://doi.org/10.1093/nar/25.24.4876
Hall BG (2005) Comparison of the accuracies of several phylogenetic methods using protein and DNA sequences. Mol Biol Evol 22(3):792–802. https://doi.org/10.1093/molbev/msi066
Khadka B, Persaud D, Gupta RS (2020) Novel sequence feature of SecA translocase protein unique to the thermophilic bacteria: bioinformatics analyses to investigate their potential roles. Microorganisms 8(1):59. https://doi.org/10.3390/microorganisms8010059
Lanfear R, Calcott B, Ho SYW, Guindon S (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Ronquist F, Teslenko M, Van-Der-Mark P, Ayres DL, Darling A et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A (2010) How many bootstrap replicates are necessary. J Comput Biol 17:337–354. https://doi.org/10.1089/cmb.2009.0179
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst Biol 67(5):901–904. https://doi.org/10.1093/sysbio/syy032
Naushad HS, Gupta RS (2013) Phylogenomics and molecular signatures for species from the plant pathogen-containing order Xanthomonadales. PLoS One 8:e55216. https://doi.org/10.1371/journal.pone.0055216
Gupta RS, Naushad S, Baker S (2015) Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbales ord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov. Int J Syst Evol Microbiol 65:1050–1069. https://doi.org/10.1099/ijs.0.070136-0
Ho J, Adeolu M, Khadka B, Gupta RS (2016) Identification of distinctive molecular traits that are characteristic of the phylum “Deinococcus-Thermus” and distinguish its main constituent groups. Syst Appl Microbiol 39:453–463. https://doi.org/10.1016/j.syapm.2016.07.003
Harper SJ, Ward LI, Clover GRG (2010) Development of LAMP and real-time PCR methods for the rapid detection of Xylella fastidiosa for quarantine and field applications. Phytopathology 100(12):1282–1288. https://doi.org/10.1094/PHYTO-06-10-0168
Roach R, Mann R, Gambley CG (2018) Identification of Xanthomonas species associated with bacterial leaf spot of tomato, capsicum and chilli crops in eastern Australia. Eur J Plant Pathol 150:595–608. https://doi.org/10.1007/s10658-017-1303-9
Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (2000) Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol 50(4):1563–1589. https://doi.org/10.1099/00207713-50-4-1563
Gopalakrishnan S, Srinivas V, Prakash B, Sathya A, Vijayabharathi R (2015) Plant growth-promoting traits of Pseudomonas geniculata isolated from chickpea nodules. 3 Biotech 5:653–661. https://doi.org/10.1007/s13205-014-0263-4
Schmidt CS, Alavi M, Cardinale M, Müller H, Berg G (2012) Stenotrophomonas rhizophila DSM14405T promotes plant growth probably by altering fungal communities in the rhizosphere. Biol Fertil Soils 48:947–960. https://doi.org/10.1007/s00374-012-0688-z
Funding
The work of CFMM is supported by FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo, Grant no. 2019/19435-3), CAPES (Coordenação de Aperfeiçoamento de Pessoal de Ensino Superior, Funding Code 001), and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico, Grant no. 308868/2018-8), Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Derlene Attili Agellis
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cutiño-Jiménez, A.M., Menck, C.F.M., Cambas, Y.T. et al. Protein signatures to identify the different genera within the Xanthomonadaceae family. Braz J Microbiol 51, 1515–1526 (2020). https://doi.org/10.1007/s42770-020-00304-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-020-00304-2