Abstract
Chagas disease is a serious health problem in Central and South America for which effective treatment is not currently available. This illness is caused by the protozoa Trypanosoma cruzi, a species that relies on a thiol-based metabolism to regulate oxidative stress. Trypanothione reductase enzyme plays a central role in the metabolic pathway of the parasite. In this work, a virtual screening of a library of novel thiadiazine derivatives against trypanothione reductase using molecular docking was performed. Four different series of hybrid ligands having in the structure one or two peptoid moieties (series I and II) or the tetrazole ring (series III and IV) were considered. An ad hoc numerical index called poses ratio was introduced to interpret the results of the docking analysis and to establish relevant structure-interaction relationships. In addition, six binding modes were found for the ligands with the highest populated conformational clusters after applying contact-based analysis. The most regular and relevant were binding modes I and II, found mainly for ligands from series I. A subsequent molecular docking on human glutathione reductase enzyme allowed to assess the possible cytotoxicity of the ligands towards human cells. A selective binding profile was found for ligands with interactions in the Hydrophobic cleft, the spermidine and the Z subsites inside the active site of trypanothione reductase. At the end of the study, new thiadiazine-based compounds were identified as plausible candidates to selectively inhibit the parasitic enzyme.
Graphic abstract

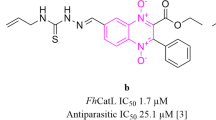
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Chagas disease or American trypanosomiasis is still considered a serious health problem in Central and South America, according the Worldwide Health Organization (WHO), despite it was discovered more than a century ago. Furthermore, imported Chagas disease is increasingly viewed as a global emergent health issue, due to the migration of infected individuals to other continents, mainly North America and Europe. Around six to seven millions of people worldwide are affected with this illness, with 12 000 fatalities reported each year and around 65 million of human beings are in risk of contracting it. Chagas disease is caused by the protozoa agent Trypanosoma cruzi (T. cruzi) and has vector-borne transmission and non-vectorial transmission mechanisms [1,2,3]. In the absence of effective anti-trypanosomal treatment the sickness has acute and chronic phases. The acute phase, that only lasts 8–12 weeks, is characterized by circulating trypomastigotes in blood; while in the chronic phase parasitemia levels become undetected by microscopy. Around 30% of the chronic patients develop, over a period of years to decades, a clinically evident cardiac and/or gastrointestinal disease, probably related to the presence of T. cruzi in smooth muscle and autonomic ganglia in heart, esophagus or colon [1,2,3].
The treatment currently available is based mainly on the drugs benznidazole and nifurtimox, which were introduced around fifty years ago. Unfortunately, both medications are only effective in the acute phase of the disease, with around 80% of effectiveness. In addition, several adverse effects have been reported; including dermatologic side effects, gastrointestinal tracks complaints and affections to the central nervous system in around 40% of the patients. Because of this, benznidazole is only commercially available in the US since 2018, while nifurtimox has just been recently reapproved by the FDA. [3, 4]
The parasitic agent T. cruzi presents a thiol-based metabolism that has been a regular target in experimental and theoretical studies. The structures of some natural molecules that have been in the center of such research projects are depicted in Fig. 1. Trypanothione dithiol (N1,N8-bis-(glutathionyl)-spermidine or T(SH)2) is a crucial intermediate that guarantees protection against oxidative stress. T(SH)2 participates in several enzymatic and non-enzymatic reactions in which it is oxidized to trypanothione-disulfide (TS2), which is then reduced back by the action of the enzyme trypanothione reductase (TR) (Fig. 1a) [5,6,7]. Trypanothione—both the reduced T(SH)2 and the oxidized TS2 forms—is a modified peptide that comprises seven building blocks (Fig. 1b). The structure can be pictured as the union of two molecules of the tripeptide L-γ-glutamyl-L-cysteinyl-glicine linked at the C-terminus by a basic spermidine (Spm) chain. Because the spermidine component is not symmetrical, the molecule is not really a dimer and the two tripeptide portions are distinguishably named I or II. Hence, trypanothione structure is often abbreviated γ-GluI-CysI-GlyI-Spm-GlyII-CysII-γ-GluII. Mammals have an equivalent enzyme designated as glutathione reductase (GR), which exerts a similar catalytic function than TR (Fig. 1a). GR reduces glutathione dimer (GSSG) to its thiol monomeric form L-γ-glutamyl-L-cysteinyl-glycine (GSH). GSSG and GSH from mammals are the molecular analogs of TS2 and T(SH)2 from T. cruzi, respectively (Fig. 1b). GSH scavenges potentially dangerous oxidizing reagents which are the result of aerobic metabolism within the cell (Fig. 1c) [8]. Because GSSG lacks the Spm residue, its overall charge is negative, in contrast with the larger and positively charged TS2.
Natural molecules relevant in the context of this study. a Enzymes trypanothione reductase (left; PDB:1BZL) [9] from T. cruzi and glutathione reductase (right; PDB:1GRA) [10] from H. sapiens. Both proteins are depicted in the same scale and the surface of the corresponding active sites are highlighted. b Structures of trypanothione disulfide (TS2) and glutathione dimer (GSSG), the natural substrates for TR and GR, espectively. The positively charged spermidine (Spm) residue, distinctive of TS2 is colored red. c Biochemical reactions catalyzed by TR and GR. NADP+: nicotinamide adenine dinucleotide phosphate and its reduced form NADPH
Enzymes GR and TR belong to the family of flavin-containing disulfide oxidoreductases. They are active as homo-dimers of about 52 kDa per subunit and have an overall 30% of sequence identity, with 14 conserved residues in the substrate binding site [8, 10]. In both molecules, the catalytic site is rather complex, comprising two separate regions, i.e., the NADP site (N-site) and the active site (G-site), connected to each other by the flavin ring of flavin adenine dinucleotide (FAD) and a redox active disulfide bridge [8, 11]. Despite all these similarities, the specific amino acid alterations among both enzymes affect the size and the chemistry of the disulfide-substrate-binding sites. In the literature, the difference in the substrate specificity of TR and GR has been explained in terms of steric and electrostatic factors [11]. The active site of TR is rich in glutamic acid and non-polar residues; while in GR basic residues (such as arginine) prevail. The orientations of the domains in the enzymes are also different, with the consequence that the disulfide-substrate-binding sites don’t have the same size. Therefore, the parasitic TR binds the larger positively charged TS2, while human GR binds smaller double negatively charged GSSG (Fig. 1b). The fact that, in general, the enzymes bind different substrates, has been the cornerstone of many studies pursuing selective compounds against Chagas disease [11].
Thiadiazines, more specifically 3,5-disubstituted thiadiazine-2-thione (THTT), have been studied by the Laboratory of Organic Synthesis (LSO) for more than 20 years [12,13,14,15,16]. Several biological studies in vitro have demonstrated the antiparasitic activity of THTT, which is related to a non-specific high cytotoxicity [12, 13]. In the last years, our group has focused on obtaining molecules containing the thiadiazine core with higher activity and/or lower cytotoxicity. Although the exact mechanism of action of thiadiazines remains obscure, the possibility of a selective inhibition of TR by some of thiadiazine-based compounds might reduce their inherent cytotoxicity. With that purpose, multicomponent procedures such as the Ugi-four-components reaction have been used for diversity-oriented synthesis of hybrid molecules with improved properties [15]. This versatile reaction enables the modification of molecules by introducing a dipeptoid residue [16] or a tetrazole ring and additional substituents—the diversity points—in the molecular scaffolds, which ease the design of libraries of new compounds. Similar synthetic approaches has been extensively used for the design of thousands of biologically active molecules [17]. In particular, tetrazoles have found widespread use in the context of medicinal chemistry because of their simple preparation, bioisosteric properties and high chemo and enzymatic stability [18].
As a continuation of our previous studies [15, 16], we designed a virtual library of 2094 hybrid THTT (Fig. 2) to assess their potential role as TR inhibitors through an in silico approach. The chemical structures of all the substituents (see Supplementary Material SA1) within the ligands were chosen so that any compound in this analysis may be easily synthesized from readily available commercial chemicals, including natural or synthetic mimetics of amino acids [15, 16, 18,19,20]. In order to select the best candidates for a future experimental study, the compounds were screened using a filtering protocol based on molecular docking. It is widely known that this computational tool effectively predicts the affinity and binding mode of a given ligand towards a particular macromolecular target [21, 22]. Molecular docking has been previously reported as a useful tool in the context of searching potential drugs against T. cruzi using TR, GAPDH, trans-sialidase, polyamine transport and cruzain enzymes as receptors [6, 7, 9, 11, 23,24,25,26,27,28,29,30,31,32,33,34,35]. In the present theoretical study, TR was selected as target taking into account the decisive role of this enzyme in T. cruzi metabolism. Therefore, the ligands were designed to be structurally similar to known inhibitors of TR—having a heterocyclic and peptidic nature–. Ligands with the highest potential to act as TR inhibitors were included in an additional docking simulation towards GR to analyze their potential cytotoxicity towards human cells and therefore their profile as selective inhibitors. As the examination of molecular docking results is often cumbersome because of the large amount of ligands considered and different enzyme-ligand interaction patterns, we have introduced a novel parameter called Pose ratio (Pr). This is a numerical index that eases the interpretation of the clustering by assessing the propensity of an individual ligand towards a specific binding mode and the effect of a particular substituent in the overall binding with the enzyme.
Chemical structure of the four thiadiazine-based series of ligands designed for molecular docking. The distinguishing molecular scaffolds appear colored: the thiadizine ring is burgundy, the N-substituted dipeptide is green and the tetrazole is ultramarine.*In this position, both possible stereoisomers were considered
2 Computational methods
2.1 Preparation of the ligands and TR protein
The designed library of potential ligands comprises four series or families of new hybrid thiadiazine-based structures. Series I and series II contain one or two peptoid moieties, respectively; while series III and IV have a tetrazole ring and differ mainly in the position of this heterocycle. Polar, charged, aliphatic and aromatic substituents were included in the structures. Neutral media (pH = 7.0) was considered to set the protonation states of the ligands, i.e. protonated—positively charged—amines, guanidine and imidazoles; and deprotonated—negatively charged—carboxylic acids and 1H-tetrazoles. The three-dimensional structure coordinates were generated using Avogadro 1.1.1 [36] software and were pre-optimized by molecular mechanics with the Steepest Descent Algorithm and force-field MMFF94 [37]. Further optimization with PM6-DH2 semi-empirical Hamiltonian [38, 39] implemented in MOPAC package [40] followed. All ligand PDB files were converted into PDBQT format, the input of AutoDock Vina program [41], in the guest user interphase AutoDockTools [42]. Charges on the ligand atoms were generated using the Gasteiger model, non-polar hydrogens were merged and default active torsions (rotatable bonds) were retained using the TORSDOF utility (Supplementary Material SA0).
The crystal structure of the protein TR (PDB: 1BZL) [9] and GR (PDB: 1GRA) [10] resolved at 2.40 Å and 2.00 Å, respectively, were retrieved from the Protein Data Bank (PDB, http://www.rcsb.org). BIOMT transformations were applied to generate the coordinates for the multimer representing known biologically significant oligomerization state of the enzyme 1GRA. These structures were optimized using pdb2pqr.py (Version 2.1.0) online server [43] with AMBER force field [44] and the protonation states of ionizable groups at pH = 7.0 were assigned by using PROPKA [45]. The ligands and waters were eliminated and the PDBQT files were generated with AutoDockTools [42]. The size of the simulation grid was selected at 36 \(\times\) 44 \(\times\) 44 Å3 and 26 \(\times\) 30 \(\times\) 28 Å3 for TR and GR, respectively. The center of the simulation box was set at the geometrical center of the active site.
2.2 Docking procedure and analysis of enzyme-ligand complexes
For the validation of the docking run, previously randomized structures of the natural substrates of the enzymes (TS2 and GSSG) were submitted to re-docking experiments. The obtained poses were compared with the experimental geometries in the enzyme-ligand complexes through the total Root-Mean-Square-Deviation (RMSD). A contact-based analysis allowed to match the enzyme-ligand interactions present in the obtained theoretical poses with those present in the crystal pose.
Docking simulations were performed using AutoDock Vina 1.1.2 program [41], which will be referred as Vina in the upcoming explanations. The active sites of TR and GR were kept rigid during the simulation. The docking parameters were set to default with the exception of the following: exhaustiveness = 32 and num_modes = 2. The Vina code predicts the adopted conformation of an enzyme-ligand complex and organizes the solutions based on a scoring function, which correlates the obtained conformations with the binding affinity (kcal/mol). The best 2 docked conformations or poses (num_modes = 2) from 10 independent runs were analyzed, to produce 20 final docked poses per ligand. These poses were then clustered based upon RMSD using a cutoff of 2.5 Å with an ad hoc Python (Version 3.7) [46] script that implements Daura’s clustering algorithm [47].
The binding energy of each cluster was set as the average binding energy of all the poses. For a particular ligand, the three clusters with the highest number of poses were selected as its binding modes; the remaining poses were considered as non-clustered. A binding mode described by a cluster with 10 or more poses was considered as a representative binding mode. This threshold, that includes at least half of the poses generated by the program for each ligand, was selected to be in agreement with the criterion of Rosenfeld, Olson and co-workers [48]. For several ligands, two representative binding modes were found, i.e. two clusters with 10 poses each.
Having the clusters for each binding mode, the parameter Pr (as from Poses ratio) was defined as the ratio between the number of representative poses (those forming a representative binding mode) and the number of non-representative poses (any other pose, whether clustered or not) for ligands having a common substituent in their structure. Pr values were calculated for all the substituents, while the mean Pr and the corresponding standard deviation were calculated for each position (see Supplementary Material SA1) within each series. Supplementary Material SD contains an Excel spreadsheet exemplifying the calculation of Pr for the substituents in compounds from Series IV.
Further analysis of interatomic contacts between the ligands with representative binding modes and the enzymes TR and GR was performed with the freely available Python-implemented computer algorithm BINANA [49]. Key binding characteristics such as hydrogen bonds, hydrophobic contacts, salt bridges (electrostatic attractions among opposed charged fragments) and π interactions were identified. All adjustable parameters by BINANA were set as default (see Supplementary Material SA0). When a ligand showed contacts with at least 60% of the amino acid residues forming a subsite, that interaction was considered as important. Ligands were regarded as potential inhibitors of TR and GR enzymes when they showed interactions with at least six (out of ten) or four (out of six) subsites, respectively. Ligands with two representative binding modes were considered as potential inhibitors of an enzyme if any of the two analyzed poses fulfilled the previous requirement. Figures depicting the binding modes of the ligands were prepared using PyMOL™ Version 2.3.1 [50].
3 Results and discussion
In order to evaluate the accuracy of the selected docking parameters, the natural substrates TS2 and GSSG were docked in the active sites of TR and GR, respectively. Two clusters with favorable energy were obtained for both natural substrates (Table 1). The poses showed significant differences in position (RMSD > 2 Å) when compared to the experimental conformations of the co-crystallized ligands. However, contact-based analysis using BINANA showed that more than half of the interactions found in the crystallized protein complexes obtained from the PDB were reproduced. These results are in agreement with similar studies in the literature [25, 31,32,33,34,35]. The fairly good coincidence between the binding mode of TS2 and GSSG in the two clusters encouraged us to retain the same docking parameters for the subsequent simulation runs.
3.1 Unravelling the role of the substituents on the binding
According to Rosenfeld-Olson, binders may be distinguished from non-binders, to some extent, using a combination of two trends, namely, predicted binding energy and level of clustering [48]. Conversely, it is known that the energy results derived from docking programs must be treated with caution because values rely on the size (number and type of atoms) of each particular molecule. Thus, absolute energy values are somewhat meaningless, unless they are supported by experimental data, which is not the case at this time [33].
The designed hybrid thiadiazine-based ligands were docked into the active site of TR using Vina program [41]. The binding energy of each cluster was calculated and negative values were obtained in all cases. Insofar the energy results are concerned, all the ligands from the library were able to dock at TR active site with comparable negative values of binding energies (between − 3.32 and − 9.90 kcal/mol) (See Supplementary Material SB1.1–4). Hence, the distinction among potential inhibitors and non-inhibitors was determined taking into account the poses population of the clusters and the enzyme-ligand interactions as described below.
Table 2 summarizes the results of the clustering following the docking towards TR enzyme. Increasing the number of active torsions in the ligands scaffolds rises the amount of non-clustered poses and less populated (non-representative) binding modes are achieved. Thus, the percentage of clustered poses and amount of representative binding modes followed the order Series II < < Series I < Series III < Series IV. Evidently, ligands from series I, III and IV have higher potentialities as TR binders. In the case of ligands with many active torsions (Series II), non-representative binding modes were mostly obtained, except for two cases. In total, 858 ligands from the whole library showed non-representative binding modes, thus having less potential for any inhibitory activity. Consequently, these compounds were ignored in the following examination.
Furthermore, aiming to get insights into important structure-interaction relationships, the parameter Poses ratio (Pr) was introduced (see the section Material and Methods for the relevant descriptions). Figure 3 illustrates the Pr values calculated for all the substituents of the ligands from series I, III and IV, as well as the mean and standard deviation found in each position. Similar Pr mean values were obtained for different positions in the same series. Nevertheless, high dispersion (standard deviation) of the values was found in all cases, which it is associated to the different nature of the substituents and their particular structural requirements to bind in a well-defined (representative) binding mode to the enzyme target TR. As a general trend, positive statistical significance on Pr values were found for voluminous substituents with no active torsions, where the steric factor (volume) is determinant. These substituents increase the enzyme-ligand interactions while only adds few active torsions to the structure. Negative statistical significance was obtained for long chains, groups with several routable bonds.
The influence of the substituents on cluster formation. The structures of all the substituents (from R1 to R11 in X axis) can be found in Supplementary Material SA1. Substituents with a positive ( +) or negative ( −) statistically significant difference with mean values are highlighted. *For the substituent in this position, the values of Pr encompass the two possible stereoisomers
In Series I, the largest Pr was observed for compounds with a tert-butyl (4a) in position R4. Large Pr were also encountered for compounds with the substituents 2e, 2m, 2o and 3g which correspond to rather diverse amino acid side chains of Glu, Trp, His and D-Phe, respectively. Low Pr were obtained for 1b (isobutyl), 3b (isobutyl) and 3f (CH2CO2CH3); whereas the lowest values corresponded to the highly flexible pentylene and propylene aliphatic chains of 2b and 2c, respectively. In the case of ligands from Series III, only those with very bulky substituents were favored. The largest Pr were obtained for substituents 7a (cyclohexyl), 8b (benzyl) and 9e (benzyl). In contrast with the previous case, the presence of the tert-butyl (7b) at the R7 position did not exert a statistically significant difference. Noteworthy, the absence of a substituent at positions R8 and R9 (8a, 9a = H) substantially diminished the formation of representative binding modes, even when there is no variation in the overall amount of active torsions of the structures. This last particular example illustrates the diversity of factors influencing Pr, such as the strength of the enzyme-ligand interactions. Regarding compounds from series IV, ligands with a cyclohexyl substituent (10a) in position R10 were favored over other possibilities. Reasonably, this ring has the ability to pack and interact strongly with the residues within the catalytic site. In position R11 the most suitable substituent was the isopropyl (11c) group.
3.2 Identifying the potential inhibitors via contact-based analysis
Crystallographic and molecular modelling data has allowed to get insights into the interaction between TR enzyme and its natural substrate TS2 [9, 27, 28, 30]. Table 3 contains the amino acid residues from TR implicated in the binding with TS2—that is, the contacts—via hydrogen bonding, van der Waals and other non-covalent interactions that have been identified [4]. These amino acid residues have been grouped in seven subsites, based on their interaction distance restriction of 4.0 Å with each building block from the natural substrate trypanothione, namely, γ-GluI, CysI, GlyI, Spm, γ-GluII, CysII and GlyII (Fig. 4) [9]. Other subsites (Z, Y and Hydrophobic cleft) relevant for the interaction have also been identified within TR active site (Fig. 4b) [9, 11, 26]. These subsites have been previously used as a template for contact-based analysis in the rational design of potential TR compounds by molecular docking. For example, analogs of clomipramine bearing a tricyclic dibenzosuberyl moiety that showed antitrypanocidal activities in vitro were recognized as competitive inhibitors of TR [24]. Docking studies identified five interaction subsites (γ-GluI, Z, CysI, GlyI and Spm sites) as the binding locations of these tricyclic inhibitors. Likewise, a comprehensive analysis of the binding modes of novel triterpenic chemotypes from the Natural Products Database (NatProDB) from Bahia State allowed the identification of new active molecules, which interacted with the subsites Hydrophobic cleft, Z, γ-GluI and Y [27].
Interaction subsites within the active site of TR enzyme. a Active site of TR in complex with trypanothione disulfide (PDB:1BZL). The structure of TS2 is depicted with colored building blocks. b Different subsites of interaction within the disulfide-binding site of TR. Each subsite-building block couple are colored the same
Investigation that relies exclusively on the predictions of the scoring function provided by the docking software is not able to accurately find all active compounds. Therefore, to discriminate potentially active from non-active ligands, one model pose from the 1355 representative binding modes of the 1236 ligands (see Table 2) obtained after the clustering were considered for a contact-based examination. The purposes of this step were: (1) to make a further filtering of the ligands, (2) to understand the kind of interactions present in the enzyme-ligand complexes and (3) to identify potential inhibitors of TR. First, the contacts of the ligands with the 28 amino acid residues constituting the 10 subsites within TR active site were obtained using BINANA (see Supplementary Material SA2 and SA4). Interactions consisted mainly in van der Waals contacts, though several hydrogen bonds, π-interactions and salt bridges were also obtained (Supplementary Material SA4). Then, the potentially active ligands were identified (see Computational Methods section for the criteria used). It was found that 277 ligands fairly mimic the interaction pattern of the natural substrate TS2, and also include contacts in the distinctive binding subsites of known inhibitors (Hydrophobic cleft, Z and Y). Taking a general picture, compounds from series I and III proved the more suitable binders. Though ligands from Series IV rendered highly populated clusters, their interaction profile was mostly poor, which could be due to their small size. It was also reinforced the idea that neutral bulky substituents with few active torsions in the ligands scaffold contributed more effectively to achieve favorable enzyme-ligand interactions.
For a better understanding of the binding of the novel THTT scaffolds to TR enzyme, the 1355 poses—from the 1236 ligands obtained from the clustering—were sorted into six average binding modes (BMTR), based on similarities in the interaction pattern found in enzyme-ligands complexes. Table 4 summarizes the important interactions of the BMTR with the subsites and the most recurrent amino acid contacts. Figure 5 depicts the orientation of the ligands forming each of the BMTR inside the active site. Binding mode I (BMTR I) comprises ligands that showed interactions in the Hydrophobic cleft and Spm subsites of TR, the vast majority from Series I. Hence, BM I is located in the left corner of the TR active site and involves also \(\gamma\)-GluI, GlyI and CysI subsites, where most ligands also showed contacts. Ligands with binding mode II (BMTR II) showed interactions in the Z subsite of TR. This subsite has proven of great importance in the design of selective inhibitors over GR human enzyme [26, 51]. Contacts in the \(\gamma\)-GluII, \(\gamma\)-GluI, CysI and GlyI subsites were also achieved. Therefore, BMTR II encompass a large region in the middle and bottom sections of TR active site. Binding mode III (BMTR III) is composed by ligands from series I and series III. It is located in the middle top of TR active site. These ligands displayed contacts in the region I of TR natural substrate site (encompassing \(\gamma\)-GluI, CysI and GlyI), as well as in the Spm and Y subsites. Binding mode IV (BMTR IV) encloses ligands from series I and series III and is located in the middle bottom of TR active site. These ligands show interactions with the region II of natural substrate sites (GlyII, CysII and \(\gamma\)-GluII) as well as with the Z subsite. Binding mode V (BMTR V) is located in the top corner of TR active site, with interactions in the subsites \(\gamma\)-Glu I and Y. Binding mode VI (BMTR VI) contains ligands from series I, III and IV with interactions in the \(\gamma\)-Glu I subsite.
3.3 Inspecting selectivity against human glutathione reductase enzyme
The design of selective inhibitors of TR over GR has been main topic of several molecular docking-based studies aiming to identify potential non-cytotoxic lead compounds for the treatment of Chagas disease. For example, nitrofurans and phenothiazine derivatives have been previously identified as non-selective inhibitors of TR based mostly on the binding energy obtained from molecular docking simulations for the ligands complexes with both enzymes [32,33,34]. It has also been found in enzymatic studies that the selective inhibitors bind at Z and Hydrophobic cleft subsites from TR active site [11, 26, 29]. As in the analogous TS2-TR system, crystallographic and molecular modelling data has allowed to identify the amino acid residues involved in the binding of GSSG with the active site of GR (Fig. 6) [32,33,34]. This information was applied in the present study for the identification of potential selective inhibitors.
Interaction subsites within the active site of GR enzyme. a Active site of GR in complex with GSSG (PDB:1GRA). The structure of GSSG is depicted with colored building blocks. b Different subsites of interaction within the disulfide-binding site of GR. Each site-building block interacting couple are colored the same
A further docking study towards GR was carried out in order to recognize selectivity in the 277 potential inhibitors of TR (Supplementary Material SB2). By this approach, less cytotoxic compounds with a superior profile as chemotherapeutics for Chagas disease might be found. In a similar fashion to the preceding study, Rosenfeld-Olson criterion was applied to discriminate binders from non-binders [48]. As shown in Table 5, 74 ligands presented non-representative binding modes and were considered automatically non-binders of GR. On the contrary, 203 ligands had representative binding modes, which makes them binders. As binding does not necessary implies inhibition, a second contact-based analysis followed, applying the same approach used previously. The contacts of the ligands with the amino acid residues constituting GR active site were obtained using BINANA (see Supplementary Material SA3 and SA5).Then, the potentially active ligands were identified (see Computational Methods section for the criteria used). On the other hand, the poses from 29 ligands showed interactions with less than three (out of six) subsites. Consequently, these compounds were also regarded as potentially non-inhibitors of GR. Summarizing, at the end of the study, a total of 103 ligands were selected as potential selective inhibitors of TR enzyme (see Supplementary Material SC). These compounds bind strongly to TR active site through representative binding modes, whereas anchor at GR active site with non-representative binding modes (74 examples) or by interacting in a minor amount of the natural substrate subsites (29 examples).
We were interested in the interaction pattern of the THTT ligands with the six natural subsites in GR enzyme (see Supplementary Material SA3 and SA5). Two average binding modes (BMGR) were identified (Table 6 and Fig. 7). BMGR I included ligands with contacts in CysII and \(\gamma\)-GluI subsites, while ligands from BMGR II displayed contacts in the CysII and \(\gamma\)-GluII subsites. Regardless, both BMGR had additional recurrent interactions with other subsites. Many hydrogen bonds were found, commonly with R37(A), Y114(A) and R347(A) residues; and a few salt bridges were achieved, almost exclusively with E472(B) residues (see Supplementary Material SA5). Additionally, 24 ligands could not be clustered in any defined binding mode and were labeled as BMGR III.
Among the 103 selective TR inhibitors found, 95 ligands belonged to series I, 1 ligand belonged to series II and 7 ligands belonged series III. Their more important binding modes were BMTR I (28 ligands) and BMTR II (59 ligands), which have interactions in \(\gamma\)-GluI, CysI, GlyI and \(\gamma\)-GluII subsites corresponding to the natural substrate. This binding modes also display important contacts in the Hydrophobic cleft, the spermidine and Z subsites (see Table 4). These later subsites do not have analogous in the active site of GR and therefore are essential for the selectivity in the binding of the ligands [11, 26, 29]. Table 7 contains some example of ligands from Series I that do not form any clusters with GR. The substituents present in their structure are neutral or positive charged, which accounts for the selectivity towards the negatively charged binding site of TR, rich in non-polar and acidic residues, as mentioned above. Positions R1 and R4 are occupied by a variety of substituents, which are all neutral and bulky. Position R2 admits a neutral voluminous or a basic positively charged group (guanidino or amino), with the notable exception of I-1521, that carries a hydrogen atoms in this place.
Interaction pattern of a I-0103, b I-0332 and c I-1001 within the active site of TR enzyme. The interaction surface of each ligand is highlighted in the active site according to their binding mode. Hydrogen bonds (green), cation-\(\pi\) (magenta), \(\pi\)-\(\pi\) stacked (violet) and salt bridge (yellow) contacts are also highlighted. Residues with van der Waals interactions are displayed as cyan sticks. All distances are given in Å
The specific contacts of three ligands (I-0103, I-0332 and I-1001) with the active site of trypnothione reductase are represented in Fig. 8. Though the compounds are bound to TR mostly by van der Waals forces, hydrogen bonds, \(\pi\)-\(\pi\) stackings, cation-\(\pi\) interactions and salt bridges are also present. Compounds I-0103 and I-0332 accommodate in a very similar manner, as expected from their high structural similarity. The benzyl ring in position R1 of these ligands protrudes towards the Z subsite, affording contacts with P462(B) and L399(B) in the case of I-0103, or with P398(B) and F396(B) in the case of I-0332. This conformation is further stabilized by non-classic interactions involving the benzene ring: I-0103 has a cation-\(\pi\) contact with H461(B), whereas I-0332 has a \(\pi\)-\(\pi\)-stacking with F396(B). The thiadiazine ring lays out in the middle, just above the \(\gamma\)-Glu II subsite and close to V54(A) and H461(B). The basic substituent in position R2 (3-guanidinopropyl in I-0103 and 4-aminobutyl in I-0332) penetrates to the Gly I subsite, the more flexible 4-aminobutyl chain reaching deeper, down to L18(A) and Y111(A). The basic guanidine and amine groups form hydrogen bonds with G50(A) and S15(A), respectively. The substituent in position R3 reinforces the interaction with the Gly I subsite; while the tert-butyl at R4 position occupies the \(\gamma\)-Glu II subsite. In contrast, ligand I-1001 arranges in a very different way. The benzyl group at position R1 and the thiadizine ring interact with the Hydrophobic cleft, Gly I and Spm subsites. Interestingly, the positively charged tertiary amine of the heterocycle forms a salt bridge with residue E19(A). The indole group in R2 position projects to the Y subsite, interacting with the polar residue S470(B) and making a cation-\(\pi\) contact with R472(B). The substituent in position R3 and the ligand backbone are surrounded by residues H461(B) and E466(B) from the \(\gamma\)-Glu II subsite, while the very long aliphatic n-octyl chain extends up to the highly hydrophobic residues F396(B), P398(B), L399(B) in the Z-subsite.
4 Conclusions
Summarizing, in this study novel potential selective inhibitors of TR enzyme from T. cruzi, the pathogen of Chagas disease, were identified in silico. With that purpose, a library containing 2094 hybrid molecules based on the THTT heterocyclic core was designed. Such thiadiazines are molecules with a broad spectrum of biological activities that has been neglected in the last years due to their general cytotoxicity; however, we believe that is plausible to modulate their biological action by chemical modification. Having this hypothesis in mind, we considered ligands modified with peptoid or tetrazole moieties, which can be easily introduced with multicomponent reactions. The molecules were docked towards TR using the freely available software AutoDock Vina. A further molecular docking on human enzyme GR was carried out to explore the possible cytoxicity of the most promising candidates. The parameters chosen to analyze the results and select the most promising ligands were the population of conformational clusters and contact-based analysis of the enzyme-ligand complexes. Based on the Rosenfeld-Olson criterion, the novel numerical index called Poses ratio (Pr) was proposed to analyze the clustering results in a straightforward manner. When Pr was calculated for the different substituents within the structures of interest, fragments with a tendency to form highly populated binding modes were identified, and hence, valuable substituent-interaction relationships could be determined. We envision that this new parameter may be particularly useful as a support of other established tools in high throughput virtual screenings where thousands of compounds are considered for lead identification. In the present particular case, it was found that medium size ligands with a few rotatable bonds and large substituents are the most promising inhibitors of TR. A contact-based analysis carried out with BINANA software, permitted to sort the ligands into six binding modes, according to their interaction pattern with ten relevant subsites within the active site of TR. These subsites are described in the literature to play a crucial role in the binding of the natural substrate and known inhibitors of TR. The most recurrent amino acid contacts were found, encompassing hydrogen bonds, polar and other non-covalent interactions. To the best of our knowledge, no other docking study filtering the ligands according to their interaction profile has been previously performed in the quest of active compounds against T. cruzi. At the end of the study, 103 ligands that have a plausible selective profile in their inhibitory action were recognized. The filtering docking-based protocol presented here may be extended to other enzymes of interest.
Availability of data and material
The online version of this article contains supplementary material, which is available to authorized users.
References
Santos S, de Araújo RV, Giarolla J, El Seoud O, Ferreira EI (2020) Searching drugs for chagas disease, leishmaniasis and schistosomiasis: a brief review. Int J Antimicrob Agents 55(4):105906. https://doi.org/10.1016/j.ijantimicag.2020.105906
Pérez-Molina JA, Molina I (2018) Chagas disease. Lancet 391(10115):82–94. https://doi.org/10.1016/S0140-6736(17)31612-4
López-Vélez R, Norman FF, Bern C (2013) American trypanosomiasis (Chagas Disease). In: Magill AJ, Hill DR, Solomon T, Ryan ET (eds) Hunter’s tropical medicine and emerging infectious disease. Elsevier Inc., Amsterdam, pp 725–738. https://doi.org/10.1016/B978-1-4160-4390-4.00098-9
Bayer HealthCare Pharmaceuticals Inc., Whippany, NJ 07981 USA (2020) LAMPIT (nifurtimox) tablets, for oral use (PDF). Instructions for use approved by U.S. Food and Drug Administration (FDA). https://www.lampit.com/hcp/about-lampit. Accessed Jan 2021
Leroux AE, Krauth-Siegel RL (2016) Thiol redox biology of trypanosomatids and potential targets for chemotherapy. Mol Biochem Parasitol 206(1–2):67–74. https://doi.org/10.1016/j.molbiopara.2015.11.003
Saha D, Sharma A (2015) Docking-based screening of natural product database in quest for dual site inhibitors of Trypanosoma cruzi Trypanothione Reductase (TcTR). Med Chem Res 24(1):316–333. https://doi.org/10.1007/s00044-014-1122-x
da Rocha Pita SS, Albuquerque MG, Rodrigues CR, Castro HC, Hopfinger AJ (2012) Receptor-dependent 4D-QSAR analysis of peptidomimetic inhibitors of trypanosoma cruzi trypanothione reductase with receptor-based alignment. Chem Biol Drug Des 79(5):740–748. https://doi.org/10.1111/j.1747-0285.2012.01338.x
Couto N, Wood JL, Barber J (2016) The role of glutathione reductase and related enzymes on cellular redox homoeostasis network. Free Radic Biol Med 95:27–42. https://doi.org/10.1016/j.freeradbiomed.2016.02.028
Bond CS, Zhang Y, Berriman M, Cunningham ML, Fairlamb AH, Hunter WN (1999) Crystal structure of Trypanosoma cruzi trypanothione reductase in complex with trypanothione, and the structure-based discovery of new natural product inhibitors. Structure 7(1):81–89. https://doi.org/10.1016/S0969-2126(99)80011-2
Karplus PA, Schulz GE (1989) Substrate binding and catalysis by glutathione reductase as derived from refined enzyme: substrate crystal structures at 2Å resolution. J Mol Biol 210(1):163–180. https://doi.org/10.1016/0022-2836(89)90298-2
Tiwari N, Tanwar N, Munde M (2018) Molecular insights into trypanothione reductase-inhibitor interaction: a structure-based review. Arch Pharm (Weinheim) 351(6):e1700373. https://doi.org/10.1002/ardp.201700373
Bermello JC, Piñeiro RP, Fidalgo LM, Cabrera HR, Navarro MS (2011) Thiadiazine derivatives as antiprotozoal new drugs. Open Med Chem J 5:51–60. https://doi.org/10.2174/1874104501105010051
Rodríguez H, Suárez M, Albericio F (2012) Thiadiazines, N, N-Heterocycles of biological relevance. Molecules 17(7):7612–7628. https://doi.org/10.2174/1874104501105010051
Rodríguez H, Coro J, Suárez M, Martínez-Álvarez R, Martín N, Albericio F (2012) Liquid phase organic synthesis of 3, 5-disubstituted 1, 3, 5-thia-diazinane-2-thione derivatives on polyethylene glycol (PEG) support. Arkivoc 8:326–338. https://doi.org/10.3998/ark.5550190.0013.828
Echemendía R, Fernández O, Coro J, Suárez M, Rivera DG (2017) A versatile approach to hybrid thiadiazine-based molecules by the Ugi four-component reaction. Tetrahedron Lett 58(18):1784–1787. https://doi.org/10.1016/j.tetlet.2017.03.075
Echemendía R, Rabêlo WF, López ER, Coro J, Suárez M, Weber M, Rivera DG (2018) A bidirectional access to novel thiadiazine hybrid molecules by double multicomponent reactions. Tetrahedron Lett 59(7):4050–4053. https://doi.org/10.1016/j.tetlet.2018.09.070
Dömling A, Wang W, Wang K (2012) Chemistry and biology of multicomponent reactions. Chem Rev 112(6):3083–3135. https://doi.org/10.1021/cr100233r
Ojeda-Carralero GM, Coro J, Valdés-Palacios A (2020) Green alternatives for the synthesis of tetrazolic acids. Chem Heterocycl Comp 56(4):408–421. https://doi.org/10.1007/s10593-020-02676-7
Zhao T, Kurpiewska K, Kalinowska-Tłuścik J, Herdtweck E, Dömling A (2016) α-Amino acid-isosteric α-amino tetrazoles. Chem Eur J 22(9):3009–3018. https://doi.org/10.1002/chem.201504520
Neochoritis CG, Zhao T, Dömling A (2019) Tetrazoles via multicomponent reactions. Chem Rev 119(3):1970–2042. https://doi.org/10.1021/acs.chemrev.8b00564
Fradera X, Babaoglu K (2017) Overview of methods and strategies for conducting virtual small molecule. Curr Protoc Chem Biol 9(3):196–212. https://doi.org/10.1002/cpch.27
Gimeno A, Ojeda-Montes MJ, Tomás-Hernández S, Cereto-Massagué A, Beltrán-Debón R, Mulero M, Pujadas G, Garcia-Vallvé S (2019) The light and dark sides of virtual screening: what is there to know? Int J Mol Sci 20(6):1375–1399. https://doi.org/10.3390/ijms20061375
Cortés-Ruiz EM, Palomino-Hernández O, Rodríguez-Hernández KD, Espinoza B, Medina-Franco JL (2018) Computational methods to discover compounds for the treatment of chagas disease. Adv Protein Chem Struct Biol 113:119–142. https://doi.org/10.1016/bs.apcsb.2018.03.005
O’Sullivan MC, Durham TB, Valdes HE, Dauer KL, Karney NJ, Forrestel AC, Bacchi CJ, Baker JF (2015) Dibenzosuberyl substituted polyamines and analogs of clomipramine as effective inhibitors of trypanothione reductase; molecular docking, and assessment of trypanocidal activities. Bioorg Med Chem 23(5):996–1010. https://doi.org/10.1016/j.bmc.2015.01.018
Din ZU, Kaplum V, Lazarin-Bidóia D, Garcia FP, Nakamura CV, Rodrigues-Filho E (2019) The structure design of biotransformed unsymmetrical nitro-contained 1,5-diaryl-3-oxo-1,4-pentadienyls for the anti-parasitic activities. Arab J Chem 12(8):4006–4016. https://doi.org/10.1016/j.arabjc.2016.03.005
Chan C, Yin H, Garforth J, Mckie JH, Jaouhari R, Speers P, Douglas KT, Rock PJ, Yardley V, Croft SL, Fairlamb AH (1998) Phenothiazine inhibitors of trypanothione reductase as potential antitrypanosomal and antileishmanial drugs. J Med Chem 41(2):148–156. https://doi.org/10.1021/jm960814j
da Paixão VG, Pita SSDR (2019) In silico identification and evaluation of new Trypanosoma cruzi trypanothione Reductase (TcTR) inhibitors obtained from Natural Products Database of the Bahia semi-arid region (NatProDB). Comput Biol Chem 79:36–47. https://doi.org/10.1016/j.compbiolchem.2019.01.009
Arias DG, Herrera FE, Garay AS, Rodrigues D, Forastieri PS, Luna LE, Bürgi MDLM, Prieto C, Iglesias AA, Cravero RM, Guerrero SA (2017) Rational design of nitrofuran derivatives: synthesis and valuation as inhibitors of Trypanosoma cruzi trypanothione reductase. Eur J Med Chem 125:1088–1097. https://doi.org/10.1016/j.ejmech.2016.10.055
Santos NC, da Paixão VG, da Rocha Pita SS (2019) New trypanosoma cruzi trypanothione reductase inhibitors identification using the virtual screening in database of Nucleus Bioassay, Biosynthesis and Ecophysiology (NuBBE). Anti-Infective Agents 17(2):138–149. https://doi.org/10.2174/2211352516666180928130031
Rodríguez-Becerra J, Cáceres-Jensen L, Hernández-Ramos J, Barrientos L (2017) Identification of potential trypanothione reductase inhibitors among commercially available β-carboline derivatives using chemical space, lead-like and drug-like filters, pharmacophore models and molecular docking. Mol Divers 21(3):697–711. https://doi.org/10.1007/s11030-017-9747-6
Romero AH, López SE (2017) In silico molecular docking studies of new potential 4-phthalazinyl-hydrazones on selected trypanosoma cruzi and Leishmania enzyme targets. J Mol Gr Model 76:313–329. https://doi.org/10.1016/j.jmgm.2017.07.013
Iribarne F, González M, Cerecetto H, Aguilera S, Tapia O, Paulino M (2007) Interaction energies of nitrofurans with trypanothione reductase and glutathione reductase studied by molecular docking. J Mol Struct THEOCHEM 818(1–3):7–22. https://doi.org/10.1016/j.theochem.2007.04.035
Iribarne F, Paulino M, Aguilera S, Tapia O (2009) Assaying phenothiazine derivatives as trypanothione reductase and glutathione reductase inhibitors by theoretical docking and molecular dynamics studies. J Mol Gr Model 28(4):371–381. https://doi.org/10.1016/j.jmgm.2009.09.003
Vega-Teijido M, Caracelli I, Zukerman-Schpector J (2006) Conformational analyses and docking studies of a series of 5-nitrofuran- and 5-nitrothiophen-semicarbazone derivatives in three possible binding sites of trypanothione and glutathione reductases. J Mol Gr Model 24(5):349–355. https://doi.org/10.1016/j.jmgm.2005.09.008
da Roche Pita SS, Cirino JJ, de Alencastro RB, Castro HC, Rodriguez CR, Albuquerque MG (2009) Molecular docking of a series of peptidomimetics in the trypanothione binding site of T. cruzi trypanothione reductase. J Mol Gr Model 28(4):330–335. https://doi.org/10.1016/j.jmgm.2009.08.011
Hanwell MD, Curtis DE, Lonie DC, Vadermeersh T, Zurek E, Hutchison GR (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform 4(1):17. https://doi.org/10.1186/1758-2946-4-17
Halgren TA (1996) Merck molecular force field. V. Extension of MMFF94 using experimental data, additional computational data, and empirical rules. J Comput Chem 17(5–6):616–641. https://doi.org/10.1002/(SICI)1096-987X(199604)17:5/6%3c616::AID-JCC5%3e3.0.CO;2-X
Řezáč J, Fanfrlík J, Salahub D, Hobza P (2009) Semiempirical quantum chemical PM6 method augmented by dispersion and H-Bonding correction terms reliably describes various types of noncovalent complexes. J Chem Theory Comput 5(7):1749–1760. https://doi.org/10.1021/ct9000922
Korth M, Pitoňák M, Řezáč J, Hobza P (2010) A transferable H-bonding correction for semiempirical quantum-chemical methods. J Chem Theory Comput 6(1):344–352. https://doi.org/10.1021/ct900541n
Stewart JJP (2016) MOPAC2016. Stewart Computational chemistry, Colorado Springs, CO, USA. http://OpenMOPAC.net. Accessed Aug 2020
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461. https://doi.org/10.1002/jcc.21334
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) Autodock4 and AutoDockTools4: automated docking with selective receptor flexiblity. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256.43
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA (2004) PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucl Acids Res 32:W665–W667. https://doi.org/10.1093/nar/gkh381
Ponder JW, Case DA (2003) Force fields for protein simulations. Adv Protein Chem 66:27–85. https://doi.org/10.1016/S0065-3233(03)66002-X
Rostkowski M, Olsson MHM, Søndergard CR, Jensen JH (2011) Graphical analysis pH-dependent properties of proteins predicted using PROPKA. BMC Struct Biol 11:1–6. https://doi.org/10.1186/1472-6807-11-6
Python Software Foundation. Python language reference, version 3.7. http://www.python.org. Accessed Aug 2020
Daura X, Gademann K, Jaun B, Seebach D, van Gunsteren WF, Mark AE (1999) Peptide folding: when simulation meets experiment. Angew Chem Int Ed 38(1–2):236–240. https://doi.org/10.1002/(SICI)1521-3773(19990115)38:1/2%3c236::AID-ANIE236%3e3.0.CO;2-M
Rosenfeld RJ, Goodsell DS, Musah RA, Morris GM, Goodin DB, Olson AJ (2003) Automated docking of ligands to an artificial active site: aumenting crystallographic analysis with computer modeling. J Comput Aided Mol Des 17(8):525–536. https://doi.org/10.1023/B:JCAM.0000004604.87558.02
Durrant JD, McCammon JA (2011) BINANA: a novel algorithm for ligand-binding characterization. J Mol Gr Model 29(6):888–893. https://doi.org/10.1016/j.jmgm.2011.01.004
DeLano WL (2010) The PyMOL molecular graphics system, version 1.3r1. Schrödinger, LLC. http://www.pymol.org. Accesed Sept 2020
da Rocha Pita SS, Batista PR, Albuquerque MG, Pascutti PG (2012) Molecular dynamics simulations of peptide inhibitors complexed with trypanosoma cruzi trypanothione reductase. Chem Biol Drug Des 80(4):561–571. https://doi.org/10.1111/j.1747-0285.2012.01429.x
Acknowledgements
We thank High Performance Computing Cluster of the Central University “Marta Abreu” of Las Villas (HPC UCLV) for providing the computational resources. GMOC is grateful to VLIR-UOS (project code CU2018TEA458A101) and KU Leuven, Belgium for ongoing financial support of his PhD. EMC is thankful to DAAD and IPB Halle for providing a doctoral position.
Author information
Authors and Affiliations
Contributions
The project was conceived and headed by JC-B, whereas the computational work was supervised by EM-C. JEA-R performed the calculations, optimizations and the preparation of the supplementary material. All the authors collected, organized and analyzed the data. ERL-R and GMO-C prepared the first draft of the manuscript, while all the authors revised and contributed to following versions. DA made the artwork. All the authors read and approved the final manuscript prior to submission.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
42452_2021_4375_MOESM1_ESM.xlsx
Materials and Methods criterion (SA0). Pr values obtained for the substituents in the study (SA1). Ligands of each binding mode distribution into its amino acid contacts with TR as target (SA2). Ligands of each binding mode distribution into its amino acid contacts with GR as target (SA3). Contact-based analysis using TR as target (SA4). Contact-based analysis using GR as target (SA5). Series I ligands clustering results after the docking simulation with TR as target (SB1.1). Series II ligands clustering results after the docking simulation with TR as target (SB1.2). Series III ligands clustering results after the docking simulation with TR as target (SB1.3). Series IV ligands clustering results after the docking simulation with TR as target (SB1.4). Clustering results after the docking simulation with GR as target (SB2). Potential selective TR inhibitors (SC). Example of calculation of Pr (SD) (XLSX 796 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Coro-Bermello, J., López-Rodríguez, E.R., Alfonso-Ramos, J.E. et al. Identification of novel thiadiazin derivatives as potentially selective inhibitors towards trypanothione reductase from Trypanosoma cruzi by molecular docking using the numerical index poses ratio Pr and the binding mode analysis. SN Appl. Sci. 3, 376 (2021). https://doi.org/10.1007/s42452-021-04375-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-021-04375-0