Abstract
The asthma pandemic imposes a huge burden on patients and health systems in both developed and developing countries. Despite available treatments, symptom control is generally suboptimal, and hospitalizations and deaths remain at unacceptably high levels. A pivotal aspect of asthma that warrants further exploration is the influence of the respiratory microbiome and virome in modulating disease activity. A plethora of studies report that the respiratory microbiome is characteristically dysbiotic in asthma. In addition, our data suggest that dysbiosis is also observed on the respiratory virome, partly characterized by the reduced abundance of bacteriophages (phages). Even though phages can naturally infect and control their bacterial prey, phage therapy has been grossly neglected in the Western world, although more recently it is more widely used as a novel tool against bacterial infections. However, it has never been used for tackling microbiome dysbiosis in human non-communicable diseases. This review provides an up-to-date understanding of the microbiome and virome's role within the airways in relation to asthma morbidity. It also advances the rationale and hypothesis for the CURE project. Specifically, the CURE project suggests that managing the respiratory microbiome through phage therapy is viable and may result in restoring eubiosis within the asthmatic airway. This entails controlling immune dysregulation and the clinical manifestation of the disease. To accomplish this goal, it is crucial to predict the effects of introducing specific phage mixtures into the intricate ecology of the airways and devise suitable interventions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Asthma imposes a significant burden globally, with microbial communities, particularly viral and bacterial components, playing roles in its initiation, persistence, and exacerbation. |
Early intestinal and nasopharyngeal microbiomes, including the infant virome, influence asthma development, indicating the importance of microbiome composition in disease progression. |
Dysbiosis in the upper respiratory virome, characterized by reduced bacteriophage abundance and increased eukaryotic viruses, correlates with asthma severity, control, and lung function. |
The CURE project explores phage therapy as a novel intervention for asthma, aiming to manipulate the respiratory microbiome to reinstate a healthy microbial ecology and improve clinical outcomes. |
Through comprehensive characterization of the respiratory microbiome and its interactions with the immune system, the CURE project seeks to identify deviations from health, understand virome dynamics in asthma, and develop phage-based interventions, which is potentially revolutionizing. |
Introduction
During the last decades, the asthma pandemic has imposed a huge morbidity burden on patients and health systems in developed and developing countries in both children and adults [1]. Despite available treatments, symptom control is generally suboptimal, while hospitalizations and deaths remain at unacceptably high levels [2]. One aspect of asthma that requires further attention is its association with microbial communities. Viral and bacterial infections have been associated with both protection and initiation or persistence and exacerbation of asthma [3]. Moreover, in atopic asthmatic patients, a compromised immune response affects the host's capability to manage infections effectively [4]. The bacterial component of the nasopharyngeal microbiome in infants has been identified as a determinant of infection spread to the lower airways, enhancing the severity of inflammation, and, most importantly, the risk of future asthma development [5]. Birth cohort studies confirmed the role of certain early intestinal microbiome, i.e., Christensenellaceae, taxa family as a significant risk factor for later development of asthma [6]. Moreover, evaluation of bacterial communities in the upper and lower airways of people with asthma demonstrated abnormal microbial community structures in respect to functional or local composition or to their metabolic activities (dysbiosis), suggesting a potential underlying mechanism for disease activity [7]. Recently, there has been a growing focus on researching the role of the virome in the onset and persistence of asthma. Specifically, virome represents a methodology rather than a biological entity, including the collection of complete and/or partial genome sequences of viruses identified within an environmental site, including the human body, utilizing high-throughput metagenomic sequencing [8]. Recent research has confirmed the significant impact of the relatively unexplored infant virome in the gut on subsequent asthma development and host immune responses [9]. Specifically, temperate gut phage taxa have been identified as being associated with the later development of asthma. Furthermore, interactions between phages and host immune responses have been shown to play a role in modulating the associated risk of asthma. Data from the PreDicta study (7th Framework Program for Research and Technological Development (FP7) European Commission-funded project_260895 PREDICTA) indicate that a newly identified feature of dysbiosis in asthma involves the upper respiratory virome, i.e., the collection of viruses with different hosts and pathogenic potential for humans [10]. Bacteriophages, also known as phages, are viruses that infect and replicate within bacterial cells and are important determinants due to their ability to infect other bacteria, while they serve as mediators between pathogenic and non-pathogenic bacteria [11]. Specifically, during an asymptomatic/non-infectious state, we observed reduced abundance of bacterial viruses (bacteriophages or phages for short), and increased presence of eukaryotic viruses in preschool children, predominantly in those with uncontrolled and severe asthma [10]. Similar findings have been reported in the lower respiratory tract, where bacteriophage abundance in induced sputum was severely reduced in patients with asthma, predominantly in severe asthmatic individuals, while such reduction was significantly correlated with increased asthma activity and worse lung function, as assessed by the asthma control test, and forced expiratory volume (1st sec)/forced vital capacity, respectively [12]. These reports suggest that microbiome dysregulation in asthma also involving the viral component, is detected throughout the respiratory trac and is linked to disease severity, control, and lung function.
It is well known that phages can naturally control bacterial populations [13]. It is thus plausible that different virome elements may play a direct or indirect role in the control of inflammation, disease expression, and/or susceptibility to asthma attacks. Phage therapy has been largely overlooked in the Western world, although it has been recently used as an effective tool against severe infections [14]; however, it has never been used for rebalancing microbiome dysbiosis in humans with chronic respiratory diseases. Within the context of “Future and Emerging Technologies open call on novel ideas for radically new technologies (Horizon-2016–2017)”, the “Constructing a Eubiosis Reinstatement Therapy for Asthma” (CURE) proposal aims to develop respiratory phage therapies capable of improving clinical outcomes in asthma (https://www.cureasthma.eu/).
The core hypothesis underpinning CURE is that the appropriate management of the respiratory microbiome, achievable through phage intervention, may reinstate a healthy microbial ecology in the airways, termed eubiosis, which corresponds to the interspecies balance of microbiota community, impacting significantly the immunological balance and clinical expression in asthma. Herein, we present the evolution of the CURE hypothesis, which has been formulated based on the latest understanding of the airway microbiome and virome's influence on asthma. We elucidate the objectives of the project and its intended evaluations.
This article is based on previously conducted studies and does not contain any new studies with human participants or animals performed by any of the authors.
The Respiratory Microbiome in Health and Asthma
In the last decade, the view of the human microbiome as a major component of homeostasis has been well established, mostly based on studies of the bacterial communities in the gut [15]. The respiratory tract, however, has been much less studied and it was not until 2010 when it was shown that lungs are not sterile [16]. The human respiratory tract, from the nostrils to the lung alveoli, is inhabited by niche-specific communities of bacteria [17]. The level of similarity between the upper and the lower respiratory microbiomes fluctuates with age, health, and disease type, and is influenced by the direct mucosal flow and micro-aspiration of microbes to the lungs [17]. A high similarity exists in bacterial microbiota across the nasal airway while the within-individual diversity is reduced from the upper to the lower airways [18]. Microbial dispersal across the respiratory tract is dependent on immigration and extinction factors described up to date by the “Adapted Island Model of Lung Biogeography” [19]. Moreover, the gut microbiota plays a crucial role in the formation of lung microbiota, further affecting pulmonary health and disease, such as asthma, tuberculosis, and lung cancer, through cross-talk between the two sites [20]. It is well accepted that the lung microbiome is habituated by microbes and gastric-related substances that colonize the gut, forming a bidirectional gut–lung axis [7].
The composition of the nasopharyngeal microbiota is closely associated with upper respiratory infections and asthma [21]. Using 16S gene-targeted sequencing, dysbiotic bacterial communities both in the upper and lower airways have been associated with asthma activity in both adults and children [22]. The relative abundance of specific bacteria, including members of the Comamonadaceae, Sphingomonadaceae, Oxalobacteraceae families, has been associated with bronchial hyper-responsiveness [23, 24], while changes in bacterial diversity are reflected in the lung function [16]. More recently, the use of inhaled corticosteroids in asthmatic patients has been shown to alter airway fungal, but not bacterial composition, as this was assessed by 16S rRNA and internal transcribed spacer (ITS) sequencing in induced sputum samples [25]. Of interest, the well-described “seasonal asthma epidemic” has been strongly associated with seasonal enrichment of the upper airways with increased abundance of Moraxella and Haemophilus members, enhancing virus-induced asthma exacerbations in childhood [26]. More recently, positive associations of certain Bacteroides abundances in the gut with increased risk of asthma occurrence were confirmed in food-allergic children, more so in African-American and Black children [27]. In the same context, obesity per se, has been associated with higher relative abundance of Prevotella, Gemella, and Streptococcus species in sputum, potentially contributing to increased asthma morbidity [28].
Despite the above progress, knowledge on the viral component of the respiratory microbiome remains rudimentary [29]. This is primarily attributed to the technical challenges, the complexity of the analysis, and the significant cost associated with shotgun sequencing for both the DNA and RNA components of the virome. The respiratory virome is comprised of viruses with diverse genetic makeup (DNA, RNA, retrotranscribed), varying hosts, and infection kinetics (including eukaryotic viruses and bacterial viruses or phages). Moreover, virus-derived genetic elements can integrate within the host's chromosomes, exhibiting immunomodulatory activity, as recently demonstrated for anelloviruses, which are recognized as a significant viral component in the lungs [30]. It is well established that phages are key controllers of the microbial ecology [31, 32] and can even provide ‘non-host immunity’, either through direct interaction with the mucosal surfaces or by enhancing host innate immunity that induces a long-lasting adaptive immune response [33]. Information about the virome of healthy individuals comes from only one direct study [34] and indirectly from control participants [35]. Insufficient data on common virome profiles exist in large cohorts of patients with diverse disease states due to the limited number of studies, small sample sizes, challenges in accessing the lower airways, and high phenotypic variability among asthma patients. Moreover, how these virome profiles correlate with clinical, epidemiological, and socioeconomic characteristics remains unclear. The association of asthma exacerbations with acute viral-induced infections by specific agents such as respiratory syncytial virus or common cold viruses such as rhinoviruses, is well documented [36]. Nevertheless, such cases have been traditionally interrogated using targeted molecular approaches, therefore overlooking the underlying viral diversity. To date, only a handful of cross-sectional studies have specifically examined the respiratory virome during periods of exacerbated asthma activity. Remarkably, very little is understood about virome profiles during periods of stable or controlled disease activity [10, 22]. The most identified viruses in the respiratory tract are the Anelloviridae family, which represents one of the stable components of the human respiratory virome [37]. Limited data on the nasopharyngeal virome composition of patients with asthma have shown that certain viruses such as rhinovirus-C, and respiratory syncytial virus are the most represented viruses during asthma exacerbations in hospitalized children [38]. More recently, studies have revealed significant differences in virome diversity and composition, particularly noting a significant reduction in phages among individuals with asthma compared to controls [12]. Within the Copenhagen birth cohort study, increased abundance of Caudoviral families and specific temperate phage species in the gut, significantly increased the risk of later asthma occurrence, potentially though alterations in host immune responses and interactions with specific bacteriome signatures [7]. Importantly, there is currently no information available regarding the temporal covariation of the respiratory virome and the microbiome. Initial data from the CURE cohort, which evaluated the nasopharyngeal microbiome using metagenomic sequencing, indicate increased diversity, inter-sample variability, and intra-individual temporal dynamics. These findings suggest a dysbiotic microbiome in children with asthma, as this was noted during a 1-month period of stable disease activity [39].
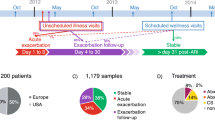
Using a metagenomic approach, we recently analyzed both the DNA and RNA components of the upper respiratory virome (nasopharynx/nasal cavity) in preschool-aged asthmatic and healthy children, deriving from the PreDicta cohort [40]. During a period of steady disease state characterized by mild-to-moderate disease severity, lasting at least 4 weeks without an asthma exacerbation or respiratory infection, we have identified shared virome profiles across individuals residing in different geographic locations across Europe. In most children, viromes were dominated by eukaryotic viruses such as Anelloviridae and common cold viruses. Interestingly, rhinovirus B species consistently dominated the virome of asthma patients. The analysis of the within individual virome ecology revealed that Anelloviridae richness and diversity were increased in asthma, along with the co-occurrence of different Anellovirus genera. Of note, bacteriophages were richer and more diverse in healthy individuals, in contrast to asthma patients who had an overall richer and more diverse eukaryotic virome [39]. An unsupervised clustering model based on virome ecological and compositional features identified three virome profile groups, characterized by gradients of increased richness and diversity of phages, eukaryotic viruses, and Anelloviridae, respectively. These virome profiles correlated to asthma severity and control and were independent of treatment. Investigation of the virus cross-species ecological interactions with the bacterial component of the microbiome (virus interactome) revealed that bacteriophages in health formed interactions of different importance for the structure of the local interactome and occupied more central positions (keystone species) within it. On the contrary, an expanded eukaryotic virus interactome was observed in asthma [39]. Overall, the above differences in the ecological structure of the virome (richness, diversity, and interactions) in the upper respiratory tract suggests that virome “dysbiosis” is a novel feature of pre-school asthma pathophysiology during asymptomatic/non-infectious states.
Host–microbiome crosstalk
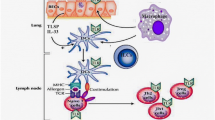
Recently, considerable attention has been directed towards exploring the interactions between bacterial communities and the immune system in both health and disease [41]. However, little is known about virome-immune connections and even less about mechanisms by which the immune system affects or is affected by the viral-bacterial interactions in an ecological context. Data from the PreDicta study indicate that preschool-aged children with asthma with low systemic (blood cells) innate interferon responses have higher abundance of picornaviruses (mostly rhinoviruses) and phages in the nasopharynx. In addition, phages, particularly Siphoviridae, appear to be sensitive sensors of host antimicrobial capacity, while Anelloviruses are not affected by TLR-induced immune response [42]. The data above suggest a probable interplay between viral stimulation and immune responses, which can subsequently alter microbial composition. Consequently, targeted manipulation of the airway virome could potentially impact asthma management. While extensive research has focused on signaling pathways in response to eukaryotic viruses, it has been revealed that prokaryotic viruses (phages) exert a profound influence on the genetic diversity, composition, and functionality of bacterial microbiota. This impact occurs through two primary mechanisms: acting as vehicles for the horizontal transfer of virulence, antibiotic resistance, and metabolic determinants, and predating susceptible bacterial strains [43]. Moreover, phages can trigger or modulate humoral immune responses, inhibit activation and proliferation of human T cells in vitro, or alter the expression of innate immune genes in mouse tissues. They can also stimulate macrophages to inhibit phagocytosis and production of inflammatory cytokines, and activate dendritic and innate lymphoid cells to produce antiviral cytokines and chemokines IFNs [44]. It is plausible that phages primarily interact with specific types of myeloid cells in the human airways. Consequently, such interactions may influence immunological homeostasis in asthma [45].
The asthmatic epithelium presents structural and functional abnormalities, while prolonged inflammation results in remodeling of its structural characteristics. Thus allergens and viruses, mostly Rhinoviruses, can further disrupt the tight junctions of the epithelium, resulting in impairment of the epithelial barrier integrity [46, 47]. The direct association of phages with the mucosal surfaces, specifically the airway epithelium, is a recently proposed concept where phages can promote synergistic and/or antagonistic relationships with bacteria on airway epithelium, potentially regulating chronic airway inflammation [48]. The triptych of airway epithelial cells, bacterial symbionts, and phages should be considered as functional and interdependent unit with direct implications on the respiratory and overall homeostasis [33]. In this context, the beneficial effect of commercial phage cocktail on epithelial cells exposed to Staphylococcus aureus was documented in an in vitro homeostasis model by down-regulating bacterial-induced inflammation, cell death, and epithelial barrier disruption [49].
The Bacterial Airway Composition in Asthmatic Patients
Microbial composition of the asthmatic airways across the spectrum of disease severity differ significantly compared with healthy individuals [50, 51]. The healthy lung microbial composition is characterized by the prevalence of bacteria belonging to the phyla Bacteroidetes, Actinobacteria, and Firmicutes, with Haemophilus being more frequent in asthmatics and Prevotella in controls, although it is well accepted that alterations are noted in relation to age, geographic area, exposure to environmental exposures etc. [16]. Using terminal restriction fragment-length polymorphism profiling for analyzing bacterial communities, it was shown that in the sputum of treatment-resistant severe asthmatics Moraxella (M) catarrhalis or members of the Haemophilus or Streptococcus genera were mostly identified, and such colonization was significantly associated with markers of asthma morbidity, such as longer asthma duration and worse lung function, as assessed by spirometry [52]. Increased abundance of M. catarrhalis, Staphylococcus species-dominated microbiota was associated with increased risk of asthma exacerbation and eosinophil activation in children with asthma [26]. Increased bacterial diversity was observed in patients with sub-optimally controlled asthma, even though diversity was inversely correlated with asthma hyperresponsiveness; of note, increased inter-individual microbiome similarity was positively correlated with bronchial hyperresponsiveness [23].
It is well accepted that bacterial diversity has been implicated in asthma but without a clear direction (higher or lower) or mechanism of action [48, 53]. Although no difference in bacterial diversity among patients with atopic asthma, atopy per se (without asthma), and healthy controls is noted, a trend towards higher bacterial phylogenetic burden/diversity has been shown in subjects with mild asthma, while an inverse relationship is observed in T2-high subjects [54]. Proteobacteria prevalence has been dominated in patients with lung diseases such as asthma, suggesting that certain bacterial colonization results in inflammatory signals may in turn induce persistent inflammation, bronchoconstriction, and bronchial hyperresponsiveness, although the possibility of the tissue microenvironment selecting specific microbes cannot be excluded [55]. It should be noted, however, that mechanistic studies investigating the influence of airway microbiomes on asthma development in murine models [56] indicate that the dynamic changes in microbiota profiles can result in enhanced airway inflammatory responses and airway remodeling, thus facilitating asthma development.
Bacteriophages as Novel Therapeutic Agents
The current paradigm for asthma management focuses on disease control through patient education, trigger avoidance, and pharmacotherapy with anti-inflammatory and bronchodilator drugs [2]. In the last few years, several monoclonal antibodies targeting key asthma mediators involved in the asthma pathophysiology have been developed including anti-IgE, anti-interleukin (IL)5, anti-IL5r, anti-IL4/13, and anti-TSLP have been developed for severe asthma [57]. The new paradigm of precision medicine is envisioned although not yet achieved. Despite advanced unsupervised clustering into diverse sub-phenotypes, treatment response prediction is still based on a small number of ‘traditional’ biomarkers (IgE, eosinophils, periostin, fractional exhaled nitric oxide_FeNO), with suboptimal specificity [58].
Treatment of infection with phages was proposed soon after their discovery in 1915. Despite initial popularity, the idea was abandoned in the ‘Western’ world after the advent of antibiotics [59]. Currently, they are only available for ‘compassionate use’, in instances where conventional antibiotics have failed [60]. However, in the former Soviet Union, phage intervention has developed into a robust clinical tool, where it is still used for the treatment of severe skin, lung, or systemic infections, albeit in an empirical way [61]. No severe side effects have ever been registered [62]. Currently, the interest is reemerging in Europe, the UK, US, and Australia [63], while workshops on the therapeutic use of bacteriophages have been organized by the European Medical Agency [64]. The Agency declared its openness for discussion and suggested that fixed phage cocktails may be evaluated within the existing regulatory framework [65]. There is, however, an ongoing debate, especially around the complexity of preparations and the scarcity of safety and efficacy data on phage use [66]. The first EU-funded clinical trial following international standards looking into the efficacy of phage therapy in burn victims has shown that phage preparation is able to decrease bacterial burden in burn wounds at a slower pace than standard of care [67]. Nevertheless, the science of phage therapy is rapidly developing around the world; studies have confirmed the significant clinical improvement in patients with resistant-to-treatment infections in different systems including lungs, urinary tract, musculoskeletal, heat and skin infections, and, most importantly, patients with sepsis in intensive care units, suggesting that such a therapy could be a key component in the battle against antimicrobial resistance [68]. Moreover, data from 52 studies have confirmed that phage therapy is well tolerated in most cases, with minimal adverse reactions, which resolved after termination of the therapy. New agents are regularly isolated and characterized [69] and phages with wider specificities are being sought [70]. Novel experimental formulations are being developed, including oral and inhaled ones for respiratory infections [71]. Phages have been used in the context of genome engineering to deliver CRISPR genes and RNA guides for sequence-specific antimicrobials.
The ‘CURE’ Concept and Hypothesis
The hypothesis underpinning CURE is that appropriate manipulation of the respiratory microbiome, achievable through phage intervention, may reinstate a healthy microbial ecology in the airways, significantly impacting the immunological balance and clinical expression in asthma (eubiosis) (Fig. 1). The project aims to advance our knowledge of the characteristics and dynamics of the human respiratory microbiome in health and asthma by describing, in detail, the currently unexplored virome using the deep analytical power of metagenomics in healthy and asthmatic individuals. The important cross-species interactions’ network including host-specificity relationships, as an anticipated driver of ecological balance, are worthy of exploring. By using longitudinal approaches within the project, we will assess the important ecological processes shaping the microbiome and its interaction to disease.
We consider that the in-depth characterization of tissue-specific and systemic immune homeostasis, as a function of microbial pressure by investigating the interactions of immune cell populations and signaling modules (cell-specific transcriptomics) with the respiratory microbiome, is an important aspect. To supplement the clinical data, the effect of phage families previously identified as part of the ‘core’ respiratory virome (Siphoviridae, Myoviridae), on macrophage, dendritic cell, and epithelial cell populations, will be assessed using in vitro co-culture models. In addition to our understanding of ecological dynamics, the development of tangible intervention agents is a key requirement. By synergizing the technologies described above, CURE aims to establish a completely new paradigm for asthma phenotyping and introduce a radically novel class of intervention. It is possible that using features describing characteristics of the microbial community, precision medicine “meta-biomarkers” can be generated. CURE assesses the upper respiratory tract (URT) microbiome, knowing that the nasal cavity and nasopharynx act as a reservoir for lower respiratory microbes, mainly through subclinical micro-aspiration, in a continuous way [72, 73]. Although the oral microbiome has been considered as the primary source of lung microbiota in a healthy state, there is substantial evidence that enhanced similarity between nasal and lung bacteria exists during disease [74]. The URT is easily accessible, which is crucial in potentially large-scale medical interventions, therefore the URT microbiome will be targeted in asthma and CURE will establish and optimize a novel collection of bacteriophages, selected for the purposes of the intervention. In parallel, phages related to the most frequently identified microbes in asthmatic airways, such as M. catarrhalis, Neisseria meningitis, Haemophilus influenza, and Prevotella, for which there is very limited information, especially on their lytic versions, which are usually selected for construction of therapeutic and prophylactic phage-based preparations, will be explored. Finally, based on the recent data from Jover and colleagues who used simulated data to show that systems with both bacteria and viruses can also be parameterized from time-series data [75]. In CURE, we aim to achieve the first empirical parameterization of a virus–bacteria interaction network in the human airways. Additionally, we will develop computational tools to inform therapeutic interventions aimed at restoring healthy symbiotic communities. The project investigates the temporal variability of the respiratory microbiome to identify microbiome states that deviate from healthy states. Furthermore, it aims to characterize the microbiome components that are linked to disease activity. The study has obtained approvals from the regional ethics committees of the clinical study centers in Greece and Poland.
Overall, the CURE hypothesis presents an innovative and ambitious proposal that requires evaluation across diverse populations and disease states. Clinical interventions are envisioned as the goal, although the realization of its full potential also depends on the development of relevant regulations. The assembled CURE dataset and biobank represent invaluable resources that are currently undergoing analysis.
References
Papadopoulos NG, Miligkos M, Xepapadaki P. A current perspective of allergic asthma: From mechanisms to management. Handb Exp Pharmacol. 2022;268:69–93.
Global Initiative for Asthma (GINA) report. Global strategy for asthma management and prevention 2023 update.
Papadopoulos NG, Christodoulou I, Rohde G, et al. Viruses and bacteria in acute asthma exacerbations—a GA(2) LEN-DARE systematic review. Allergy. 2011;66(4):458–68.
Contoli M, Message SD, Laza-Stanca V, et al. Role of deficient type III interferon-lambda production in asthma exacerbations. Nat Med. 2006;12(9):1023–6.
Teo SM, Mok D, Pham K, et al. The infant nasopharyngeal microbiome impacts severity of lower respiratory infection and risk of asthma development. Cell Host Microbe. 2015;17(5):704–15.
Lee-Sarwar KA, Kelly RS, Lasky-Su J, et al. Integrative analysis of the intestinal metabolome of childhood asthma. J Allergy Clin Immunol. 2019;144(2):442–54.
Castro-Nallar E, Shen Y, Freishtat RJ, et al. Integrating microbial and host transcriptomics to characterize asthma-associated microbial communities. BMC Med Genom. 2015;8:50.
Bai GH, Lin SC, Hsu YH, Chen SY. The human virome: viral metagenomics, relations with human diseases, and therapeutic applications. Viruses. 2022;14(2):278.
Leal Rodriguez C, Shah SA, Rasmussen MA, et al. The infant gut virome is associated with preschool asthma risk independently of bacteria. Nat Med. 2024;30(1):138–48.
Spyridon M, Bede C, Paraskevi X, et al. Bacteriophage deficiency characterizes respiratory virome dysbiosis in childhood asthma. BioRxiv. 2020.
Sweere JM, Van Belleghem JD, Ishak H, et al. Bacteriophage trigger antiviral immunity and prevent clearance of bacterial infection. Science. 2019. https://doi.org/10.1126/science.aat9691.
Choi S, Sohn KH, Jung JW, et al. Lung virome: new potential biomarkers for asthma severity and exacerbation. J Allergy Clin Immunol. 2021;148(4):1007–15 (e9).
Chevallereau A, Pons BJ, van Houte S, Westra ER. Interactions between bacterial and phage communities in natural environments. Nat Rev Microbiol. 2022;20(1):49–62.
Strathdee SA, Hatfull GF, Mutalik VK, Schooley RT. Phage therapy: From biological mechanisms to future directions. Cell. 2023;186(1):17–31.
Valdes AM, Walter J, Segal E, Spector TD. Role of the gut microbiota in nutrition and health. BMJ. 2018;361: k2179.
Hilty M, Burke C, Pedro H, et al. Disordered microbial communities in asthmatic airways. PLoS One. 2010;5(1): e8578.
Man WH, de Steenhuijsen Piters WA, Bogaert D. The microbiota of the respiratory tract: gatekeeper to respiratory health. Nat Rev Microbiol. 2017;15(5):259–70.
Mason RD, Welles HC, Adams C, et al. Targeted isolation of antibodies directed against major sites of SIV Env vulnerability. PLoS Pathog. 2016;12(4): e1005537.
Dickson RP, Erb-Downward JR, Freeman CM, et al. Spatial variation in the healthy human lung microbiome and the adapted island model of lung biogeography. Ann Am Thorac Soc. 2015;12(6):821–30.
Zhou A, Lei Y, Tang L, et al. Gut microbiota: the emerging link to lung homeostasis and disease. J Bacteriol. 2021;203(4):10–1128.
Kang HM, Kang JH. Effects of nasopharyngeal microbiota in respiratory infections and allergies. Clin Exp Pediatr. 2021;64(11):543–51.
Liu T, Lin CH, Chen YL, et al. Nasal microbiome change during and after exacerbation in asthmatic children. Front Microbiol. 2021;12: 833726.
Huang YJ, Nelson CE, Brodie EL, et al. Airway microbiota and bronchial hyperresponsiveness in patients with suboptimally controlled asthma. J Allergy Clin Immunol. 2011;127(2):372–81 (e1-3).
Lynch SV. The lung microbiome and airway disease. Ann Am Thorac Soc. 2016;13(Suppl 2):S462–5.
Huang C, Ni Y, Du W, Shi G. Effect of inhaled corticosteroids on microbiome and microbial correlations in asthma over a 9-month period. Clin Transl Sci. 2022;15:1723–36.
McCauley K, Durack J, Valladares R, et al. Distinct nasal airway bacterial microbiotas differentially relate to exacerbation in pediatric patients with asthma. J Allergy Clin Immunol. 2019;144(5):1187–97.
Mahdavinia M, Fyolek JP, Jiang J, et al. Gut microbiome is associated with asthma and race in children with food allergy. J Allergy Clin Immunol. 2023;152(6):1541–9 (e1).
Kozik AJ, Begley LA, Lugogo N, et al. Airway microbiota and immune mediator relationships differ in obesity and asthma. J Allergy Clin Immunol. 2023;151(4):931–42.
Mageiros L, Megremis S, Papadopoulos NG. The virome in allergy and asthma: a nascent, ineffable player. J Allergy Clin Immunol. 2023;152(6):1347–51.
Dodi G, Attanasi M, Di Filippo P, Di Pillo S, Chiarelli F. Virome in the lungs: the role of anelloviruses in childhood respiratory diseases. Microorganisms. 2021;9(7):1357.
Weinbauer MG, Rassoulzadegan F. Are viruses driving microbial diversification and diversity? Environ Microbiol. 2004;6(1):1–11.
Koskella B, Brockhurst MA. Bacteria-phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol Rev. 2014;38(5):916–31.
Tzani-Tzanopoulou P, Skliros D, Megremis S, et al. Interactions of bacteriophages and bacteria at the airway mucosa: new insights into the pathophysiology of asthma. Front Allergy. 2020;1: 617240.
Wylie KM, Mihindukulasuriya KA, Zhou Y, Sodergren E, Storch GA, Weinstock GM. Metagenomic analysis of double-stranded DNA viruses in healthy adults. BMC Biol. 2014;12:71.
Mitchell AB, Oliver BG, Glanville AR. Translational aspects of the human respiratory virome. Am J Respir Crit Care Med. 2016;194(12):1458–64.
Xepapadaki P, Papadopoulos NG. Childhood asthma and infection: virus-induced exacerbations as determinants and modifiers. Eur Respir J. 2010;36(2):438–45.
Freer G, Maggi F, Pifferi M, Di Cicco ME, Peroni DG, Pistello M. The virome and its major component, anellovirus, a convoluted system molding human immune defenses and possibly affecting the development of asthma and respiratory diseases in childhood. Front Microbiol. 2018;9:686.
Romero-Espinoza JA, Moreno-Valencia Y, Coronel-Tellez RH, et al. Virome and bacteriome characterization of children with pneumonia and asthma in Mexico City during winter seasons 2014 and 2015. PLoS One. 2018;13(2): e0192878.
Megremis S, Constantinides B, Xepapadaki P, et al. Respiratory eukaryotic virome expansion and bacteriophage deficiency characterize childhood asthma. Sci Rep. 2023;13(1):8319.
Xepapadaki P, Bachert C, Finotto S, et al. Contribution of repeated infections in asthma persistence from preschool to school age: Design and characteristics of the PreDicta cohort. Pediatr Allergy Immunol. 2018;29(4):383–93.
Belkaid Y, Naik S. Compartmentalized and systemic control of tissue immunity by commensals. Nat Immunol. 2013;14(7):646–53.
Rovira Rubio J, Megremis S, Pasioti M, et al. Respiratory virome profiles reflect antiviral immune responses. Allergy. 2023;78(5):1258–68.
Duerkop BA, Hooper LV. Resident viruses and their interactions with the immune system. Nat Immunol. 2013;14(7):654–9.
Federici S, Nobs SP, Elinav E. Phages and their potential to modulate the microbiome and immunity. Cell Mol Immunol. 2021;18(4):889–904.
Jonczyk-Matysiak E, Weber-Dabrowska B, Owczarek B, et al. Phage-phagocyte interactions and their implications for phage application as therapeutics. Viruses. 2017;9(6):150.
Looi K, Buckley AG, Rigby PJ, et al. Effects of human rhinovirus on epithelial barrier integrity and function in children with asthma. Clin Exp Allergy. 2018;48(5):513–24.
Celebi Sozener Z, Cevhertas L, Nadeau K, Akdis M, Akdis CA. Environmental factors in epithelial barrier dysfunction. J Allergy Clin Immunol. 2020;145(6):1517–28.
Leung CYJ, Weitz JS. Modeling the synergistic elimination of bacteria by phage and the innate immune system. J Theor Biol. 2017;429:241–52.
Tzani-Tzanopoulou P, Rozumbetov R, Taka S, et al. Development of an in vitro homeostasis model between airway epithelial cells, bacteria and bacteriophages: a time-lapsed observation of cell viability and inflammatory response. J Gen Virol. 2022. https://doi.org/10.1099/jgv.0.001819.
Carr TF, Alkatib R, Kraft M. Microbiome in mechanisms of asthma. Clin Chest Med. 2019;40(1):87–96.
Venkataraman A, Bassis CM, Beck JM, et al. Application of a neutral community model to assess structuring of the human lung microbiome. MBio. 2015. https://doi.org/10.1128/mBio.02284-14.
Green BJ, Wiriyachaiporn S, Grainge C, et al. Potentially pathogenic airway bacteria and neutrophilic inflammation in treatment resistant severe asthma. PLoS One. 2014;9(6): e100645.
Shade A. Diversity is the question, not the answer. ISME J. 2017;11(1):1–6.
Durack J, Lynch SV, Nariya S, et al. Features of the bronchial bacterial microbiome associated with atopy, asthma, and responsiveness to inhaled corticosteroid treatment. J Allergy Clin Immunol. 2017;140(1):63–75.
Hufnagl K, Pali-Scholl I, Roth-Walter F, Jensen-Jarolim E. Dysbiosis of the gut and lung microbiome has a role in asthma. Semin Immunopathol. 2020;42(1):75–93.
Zheng J, Wu Q, Zou Y, Wang M, He L, Guo S. Respiratory microbiota profiles associated with the progression from airway inflammation to remodeling in mice with OVA-induced asthma. Front Microbiol. 2021;12: 723152.
Papadopoulos NG, Barnes P, Canonica GW, et al. The evolving algorithm of biological selection in severe asthma. Allergy. 2020;75(7):1555–63.
Xepapadaki P, Adachi Y, Pozo Beltran CF, et al. Utility of biomarkers in the diagnosis and monitoring of asthmatic children. World Allergy Organ J. 2023;16(1): 100727.
Kutter E, Sulakvelidze A. Bacteriophages. Biology and Applications; 2004.
Onsea J, Uyttebroek S, Chen B, et al. Bacteriophage therapy for difficult-to-treat infections: the implementation of a Multidisciplinary Phage Task Force (The PHAGEFORCE Study Protocol). Viruses. 2021;13(8):1543.
Chanishvili N. Bacteriophages as therapeutic and prophylactic means: summary of the Soviet and post-Soviet experiences. Curr Drug Deliv. 2016;13(3):309–23.
Pirnay JP, Blasdel BG, Bretaudeau L, et al. Quality and safety requirements for sustainable phage therapy products. Pharm Res. 2015;32(7):2173–9.
Dicks LMT, Vermeulen W. Bacteriophage-host interactions and the therapeutic potential of bacteriophages. Viruses. 2024;16(3):478.
Pelfrene E, Willebrand E, Cavaleiro Sanches A, Sebris Z, Cavaleri M. Bacteriophage therapy: a regulatory perspective. J Antimicrob Chemother. 2016;71(8):2071–4.
Verbeken G, Pirnay JP, Lavigne R, et al. Call for a dedicated European legal framework for bacteriophage therapy. Arch Immunol Ther Exp (Warsz). 2014;62(2):117–29.
Jault P, Leclerc T, Jennes S, et al. Efficacy and tolerability of a cocktail of bacteriophages to treat burn wounds infected by Pseudomonas aeruginosa (PhagoBurn): a randomised, controlled, double-blind phase 1/2 trial. Lancet Infect Dis. 2019;19(1):35–45.
Uyttebroek S, Chen B, Onsea J, et al. Safety and efficacy of phage therapy in difficult-to-treat infections: a systematic review. Lancet Infect Dis. 2022;22(8):e208–20.
Weber-Dabrowska B, Jonczyk-Matysiak E, Zaczek M, Lobocka M, Lusiak-Szelachowska M, Gorski A. Bacteriophage procurement for therapeutic purposes. Front Microbiol. 2016;7:1177.
Ross A, Ward S, Hyman P. More is better: selecting for broad host range bacteriophages. Front Microbiol. 2016;7:1352.
Zelasko S, Gorski A, Dabrowska K. Delivering phage therapy per os: benefits and barriers. Expert Rev Anti Infect Ther. 2017;15(2):167–79.
Charlson ES, Bittinger K, Haas AR, et al. Topographical continuity of bacterial populations in the healthy human respiratory tract. Am J Respir Crit Care Med. 2011;184(8):957–63.
Boutin S, Graeber SY, Weitnauer M, et al. Comparison of microbiomes from different niches of upper and lower airways in children and adolescents with cystic fibrosis. PLoS One. 2015;10(1): e0116029.
Huffnagle GB, Dickson RP, Lukacs NW. The respiratory tract microbiome and lung inflammation: a two-way street. Mucosal Immunol. 2017;10(2):299–306.
Jover LF, Romberg J, Weitz JS. Inferring phage-bacteria infection networks from time-series data. R Soc Open Sci. 2016;3(11): 160654.
Acknowledgements
The members of the CURE consortium are: Grigoris Kaltsas, Evangelia Lebessi, Anastassios Doudoulakakis, Stella Taka, Panagiota Tzani Tzanopoulou, Evangelia Legaki, Rena Stergiou, David Robertson, Tucker Gilman, Mark Muldoon, Avraam Tapinos, Chuan Fu Yap, George Gkimpas, Joe Busby, Mubeccel Akdis, Cezmi Akdis, Anna Globinska, Ramazan Rozumbetov, Vangelis Andreakos, Ioanna Galani, Mikaela Koutrouli, Vaso Triantafullia, Hannah Wanstall, Maria Papadaki, Marek† Kowalski, Aleksandra Wardzyńska, Maciej Chałubiński, Nina Chanishvili, Elene Kakabadze, Marina Goderdzishvili, Valeria Ramiconi, Isabel Proano, Sofia Romagosa, Christos Ilioudis, Athina Thanopoulou, Dimitris Raptis.
Funding
The study has received funding from the Horizon 2020 research and innovation program of the European Union, under contract number no 767015.
Author information
Authors and Affiliations
Consortia
Contributions
Paraskevi Xepapadaki: draft and final editing of the manuscript. Maria Pasioti, Maria Kritikou Nikoletta Rovina, and Aleksandra Wardzyńska: inclusion and follow-up of the patients; Spyridon Megremis: metagenomic analysis, reviewed and approved the paper; Nikolaos G. Papadopoulos: concept, edited, and approved the paper. NGP is the coordinator of the CURE project.
Corresponding author
Ethics declarations
Conflict of interest
Paraskevi Xepapadaki reports outside the submitted work payment or honoraria for lectures, presentations, speakers’ bureaus, Galenica, GlaxoSmithKline, Menarini, Novartis, Uriach, Nestle, Nutricia. Nikoloas G. Papadopoulos reports grants or contracts from any entity Capricare, Nestle, Numil, Vianex, consulting fees from Abbott, AbbVie, Astra Zeneca, GSK, HAL, Medscape, Menarini/Faes Farma, Mylan, Novartis, Nutricia, OM Pharma, Regeneron/Sanofi, outside the submitted work. NGP is an Editorial Board member of Pulmonary Therapy. Nikos Papadopoulos was not involved in the selection of peer reviewers for the manuscript nor any of the subsequent editorial decisions. Spyridon Megremis, Nikoletta Rovina, Aleksandra Wardzynska, Maria Pasioti, Maria Kritikou have nothing to declare.
Ethics approval
This article is based on previously conducted studies and does not contain any new studies with human participants or animals performed by any of the authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Xepapadaki, P., Megremis, S., Rovina, N. et al. Exploring the Impact of Airway Microbiome on Asthma Morbidity: A Focus on the “Constructing a ‘Eubiosis Reinstatement Therapy’ for Asthma—CURE” Project. Pulm Ther 10, 171–182 (2024). https://doi.org/10.1007/s41030-024-00261-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41030-024-00261-3