Abstract
One of the biggest challenges of wheat production worldwide is to overcome fungal diseases, and among these is Fusarium head blight (FHB) caused mainly by the Fusarium graminearum species complex (FGSC). The occurrence of FHB can lead to contamination of grains with mycotoxins, which are harmful to human and animal health. In South America, in addition to F. graminearum sensu stricto (F. graminearum), which is the most common causal agent of FHB worldwide, F. meridionale and F. austroamericanum have also been found in wheat fields. Special attention should be paid to F. meridionale because it produces the trichothecene mycotoxin nivalenol, which has potentially higher levels of toxicity when compared with that of other trichothecene produced by the FGSC. Knowledge of the diversity and genetic structure of populations of pathogens can assist in understanding pathogen survival in the face of host genetic resistance, crop rotation, and management with fungicides. In this study, we used 10 inter-simple sequence repeat (ISSR) primers to estimate the variability, diversity, and genetic structure of populations of F. graminearum, F. meridionale, and F. austroamericanum co-occurring in wheat fields in Paraná State, Brazil. The population of F. graminearum showed the greatest genetic variability according to its polymorphism (P = 86.61%), Nei’s genetic diversity (h = 0.28), and Shannon diversity (I = 0.43) indices as compared with that of the populations of F. meridionale (P = 75.89%, h = 0.23, I = 0.36) and F. austroamericanum (P = 67.86%, h = 0.22, I = 0.33). The dendrogram, principal coordinate analysis, Bayesian analysis (K = 4), and the differentiation index (GST = 0.27) showed very high structure and, consequently, high species differentiation. The overall analysis of our data indicates high variability and genetic structure of the studied populations. This genetic condition guarantees high resilience of these species and could make it difficult to obtain wheat cultivars with good resistance to FHB, as well as may cause chemical control to be less effective.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat (Triticum aestivum L.) is the source of 20% of the total calories consumed by humanity and is the second crop in grain production worldwide, surpassed only by corn (Chaves et al. 2013). Maintaining and increasing the production of this cereal is fundamental for world food security (Chaves et al. 2013; Boff et al. 2019). Among the fungal diseases that affect wheat Fusarium head blight (FHB), caused mostly by the members of Fusarium graminearum species complex (FGSC), is one of the most relevant (Van Der Lee et al. 2015; Wang et al. 2019; Xia et al. 2020). Losses caused by the occurrence of FHB in wheat in the 1990s were estimated at US $ 2.5 billion and US $ 220 million in the USA and Canada, respectively (Windels 2000), and from 2008 to 2015, there was a significant increase in FHB epidemics in China (Wang et al. 2019). In South America, several FHB epidemics have been recorded in recent decades (Del Ponte et al. 2009; Palazzini et al. 2017). The importance of FHB in world agriculture is substantial enough that F. graminearum, the species most commonly causing FHB, was ranked as the fourth most important fungus in the world (Dean et al. 2012). The level of concern regarding FHB results not only from its devastating impact in the field but also from the frequent contamination of the harvested grains with mycotoxins, mainly trichothecenes, making unsafe their use as human food or animal feed unsafe (Schothorst and Van Egmond 2004).
Among the more than 150 trichothecene mycotoxin types, deoxynivalenol (DON) is a common contaminant of wheat grains, and wheat-based products worldwide (Sobrova et al. 2010). Deoxynivalenol, its acetylated derivatives 3-acetyldeoxynivalenol (3-ADON) and 15-acetyldeoxynivalenol (15-ADON), and nivalenol (NIV) are the most common trichothecenes produced by the members of FGSC that cause FHB (Bertero et al. 2018). Even at low concentrations, these trichothecenes are harmful to human and animal health (Varga et al. 2015; Bertero et al. 2018).
Since 2004, 15 phylogenetic species have been identified within the FGSC (O’Donnell et al. 2000; O’Donnell et al. 2004; Starkey et al. 2007; O’Donnell et al. 2008; Yli-Mattila et al. 2009; Sarver et al. 2011). The most widely geographically distributed member of FGSC causing FHB in wheat is F. graminearum sensu stricto. This species is predominant in wheat fields in the Americas (North and South), Europe, and Asia (Laday et al. 2004; Zeller et al. 2003; Zeller et al. 2004; Ramirez et al. 2007; Scoz et al. 2009; Van Der Lee et al. 2015). Despite the great prevalence of F. graminearum in wheat fields around the world, other species predominate in specific regions. For example, in both China and Japan, in the northern wheat fields, there is a predominance of the species F. graminearum, whereas to the south F. asiaticum prevails (Suga et al. 2008; Zhang et al. 2012). Likewise, in Ethiopia, the most common species in wheat is F. aethiopicum (O’Donnell et al. 2008). Another FHB aspect that deserves to be highlighted is the frequent co-occurrence of more than one FGSC member with differing trichothecene genotypes.
In South America, in Argentina, Brazil, and Uruguay, where the areas of larger wheat production are located, F. graminearum genotype 15-ADON is predominant, but several other species and genotypes have been identified as co-occurring. In Uruguay, Umpiérrez-Failache et al. (2013) reported the presence of five FGSC members of three genotypes in wheat fields [F. graminearum (15-ADON and 3-ADON), F. asiaticum (NIV), F. cortaderiae (NIV), F. brasilicum (NIV), and F. austroamericanum (3-ADON)], with. F. graminearum (15-ADON) being the most common. In Argentina, F. graminearum 15-ADON producer also prevail (Ramírez et al. 2006; Fernández Pinto et al. 2008; Alvarez et al. 2009; Reynoso et al. 2011; Malbrán et al. 2014; Ortega et al. 2016). Recently, Palacios et al. (2017) evaluated isolates sampled from Triticum turgidum L. var. durum and identified, in addition to F. graminearum with the 15-ADON genotype, F. cerealis with the NIV genotype.
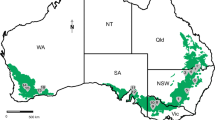
In Brazil, the evaluation of 671 isolates sampled from 200 wheat fields over a 3-year period (2009 to 2011) identified the occurrence of five FGSC members of three trichothecene genotypes [(F. graminearum 15-ADON (83%), F. meridionale NIV (12.8%), F. asiaticum NIV (0.4%), F. cortaderiae NIV or 3-ADON (2.5%), and F. austroamericanum NIV or 3-ADON (0.9%)] (Del Ponte et al. 2015). Among these species, F. asiaticum and F. cortaderiae were isolated only from wheat fields located in the extreme southern area (state of Rio Grande do Sul) of the region under study. It is interesting to note that in South America, F. meridionale (genotype NIV) was observed in wheat fields only in Brazil, whereas F. austroamericanum (genotype NIV or 3-ADON) was found in Brazil and Uruguay. In the state of Paraná, Brazil, the largest wheat producer in the country, F. meridionale (NIV producer) is of great importance as it is responsible for approximately 30% of cases of FHB (Del Ponte et al. 2015). Fusarium meridionale has been described as endemic to Latin America (Aoki et al. 2012), but its presence has also been reported in wheat fields in Australia (Amarasinghe et al. 2019), China (Zhang et al. 2014), Nepal (Desjardins and Proctor 2011), South Africa (Boutigny et al. 2011), and Taiwan (Wang and Cheng 2017). Contrarily, F. austroamericanum has been detected in wheat only in South America (O’Donnell et al. 2008; Umpiérrez-Failache et al. 2013; Del Ponte et al. 2015).
Knowledge of variability, diversity, and genetic structure of populations of pathogens can aid in the prediction of their behavior against resistant cultivars, application of fungicides, and climatic changes. This prediction is based on the premise that the higher the genetic diversity and variability of a species, the higher will be its resilience in the face of situations that affect its survival (Nevo 1978; Barrett and Schluter 2008). For a few members of the FGSC, population-genetic data are available, with F. graminearum being the most studied. For this species, high levels of genetic diversity have been observed (Mishra et al. 2004; Astolfi et al. 2011; Albayrak et al. 2016; Pan et al. 2016; Palacios et al. 2017; Malbrán et al. 2019; Yerkovich et al. 2020) suggesting that, although the species is homothallic and self-fertile, sexual reproduction is probably occurring. For F. meridionale and F. austroamericanum, there are no data available on genetic variability, diversity, or genetic structure of populations. Population-genetic data is an important step in understanding the dynamics of F. meridionale and F. austroamericanum in the region where these occur. The use of inter-simple sequence repeat (ISSR) DNA markers (Zietkiewicz et al. 1994) has already been shown to be effective to reveal population-genetic data in species of Fusarium (Arif et al. 2008; Dinolfo et al. 2010; Divakara et al. 2013; Albayrak et al. 2016; Santillán-Mendoza et al. 2019). Thus, the goals of this study were to estimate genetic variability, diversity, and structure of populations among isolates of F. graminearum sensu stricto, F. meridionale, and F. austroamericanum sampled in wheat fields in southern Paraná State, Brazil.
Material and methods
Populations, DNA extraction, and quantification
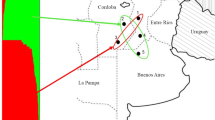
In this study, 49 fungal isolates belonging to three populations of F. graminearum (n = 19), F. meridionale (n = 16), and F. austroamericanum (n = 14) were evaluated (Fig. 1). These isolates were sampled between the years 2012 and 2016 in several wheat fields of southern Paraná State, Brazil, and belong to the fungal collection of the Plant Pathology Laboratory of the Universidade Estadual de Maringá (UEM), Paraná, Brazil. They were previously identified at the Mycotoxin Prevention and Applied Microbiology Research Unit, National Center for Agricultural Utilization Research, Agricultural Research Service, USDA, Peoria, Illinois, USA, using the multilocus genotyping approach developed by Ward et al. (2008). The population molecular analysis was done at the Central Laboratory of the Cooperativa Agrária Agroindustrial and in the Plant Molecular Biology Laboratory of the Universidade Estadual do Centro-Oeste, Guarapuava, PR, Brazil.
For DNA extraction, each isolate was plated in potato dextrose agar medium and kept for 7 days in a growth-chamber room (25 ± 2 °C). After this period, the mycelium was scraped with a platinum loop, dipped in liquid nitrogen, and crushed until a fine powder was obtained. The crushed fungal tissue was stored in a microtube at − 20 °C until DNA extraction. The DNA was extracted according to the protocol proposed by Doyle and Doyle (1987). For each isolate, 1 mL of extraction buffer (20 mM EDTA, 100 mM Tris-HCl pH 8.0, 1.4 M NaCl, 2% CTAB, 0.4% β-mercaptoethanol) was added to 100 mg of crushed tissue and incubated in a water bath at 65 °C for 45 min. The DNA was separated from the solution by precipitation using chloroform: isoamyl alcohol (24:1 v/v) followed by centrifugation. To obtain genetic material of high purity, successive washes with ethanol were performed. After extraction, the DNA was resuspended in TE (10 mM Tris-HCl; pH 8.0; 1 mM EDTA), treated with RNAse at 37 °C for 1 h and stored at − 20 °C until use. The quantification of DNA and confirmation of its integrity was conducted using electrophoresis on 0.9% agarose gel stained with ethidium bromide and visualized using UV light. To determine the DNA concentration, after electrophoresis, each sample was compared with known amounts (100, 200, and 400 ng) of intact DNA from the Lambda phage (Thermo Fisher Scientific) run in different wells of the same gel as the samples.
PCR, electrophoresis, and gel analysis
The DNA of each isolate was amplified using 10 ISSR primers developed by the University of British Columbia (UBC), Vancouver, Canada (Table 1). The DNA amplification reactions were conducted in a final volume of 12.5 μL containing 20 ng of DNA, 0.2 μM of primer, 200 μM of each dNTP, 1.5 mM of MgCl2, 1 U of Taq DNA polymerase, and 1X PCR buffer [200 mM Tris-HCl (pH 8.4), 500 mM KCl]. All reagents and enzymes used in PCR were from Invitrogen. For amplification, the thermocycler (Veriti, Applied Biosystems) was programmed with an initial denaturation step at 94 °C for 5 min, followed by 35 cycles at 90 °C for 45 s, and annealing temperature of the primer for 30 s (Table 1) and 72 °C for 60 s. At the end of the cycles, a step of 10 min at 72 °C was added for the final extension of the fragments. The amplification products were separated by electrophoresis on 1.8% agarose gel stained with ethidium bromide (0.5 μg mL−1), with a 4 h run at 110 V. To determine the size of the amplified fragments, a molecular weight marker 100 bp DNA ladder (Invitrogen) was used. The electrophoresis result was visualized in UV light and photo-documented with a digital system (L-PIX EX, Loccus do Brasil).
Statistical analysis
The gels were visually evaluated for the presence (1) or absence (0) of bands in each amplified locus. Only polymorphic bands with good definition were used to construct the binary matrix used in the statistical analysis. The genetic relationship among the genotypes was estimated by Nei’s genetic distance (Nei 1972) using the NTSYS 2.2 software (Rohlf 2000) and a dendrogram was obtained by the neighbor-joining method also using the NTSYS. Fusarium proliferatum was used as an outgroup and the tree obtained was not rooted so as not to lose the genetic distance scale. The reliability of branches in the tree was assessed by bootstrapping with 1000 replicates using the FreeTree software version 0.9.1.50 (Hampl et al. 2001). The percentage of polymorphism (P), Shannon diversity index (I), Nei’s genetic diversity (h), and the coefficient of genetic differentiation (GST) were calculated using the POPGENE software version 1.31 (Yeh et al. 1999).
A principal coordinate analysis (PCoA) was performed using GenAlEx software version 6.501 (Peakall and Smouse 2012). The Bayesian analysis was conducted to determine the number of clusters (genetic groups) that best explained the observed genetic variability, using STRUCTURE program version 2.3 (Pritchard et al. 2010). To determine the best-supported number of clusters (K), simulations were made based on the principle that it is possible to obtain any number of clusters between 1 and 10, with 10 repetitions for each simulation. For this analysis, the ancestry admixture model was used, and the allelic frequencies were correlated using 500,000 burn-in and 1,000,000 MCMC (Markov Chain Monte Carlo) steps after the burn-in. To define the K (number of clusters - genetic groups) most likely in relation to those proposed by the analysis, the criteria suggested by Evanno et al. (2005) were used with the help of the Structure Harvester program version 0.9.94 (Earl and VonHold 2012). For all analyses, each group of isolates belonging to the same species was considered a population.
Results
The 10 ISSR primers amplified 128 bands and of them, 112 (87%) were polymorphic and were used in population analysis (Table 1).
The isolates of F. graminearum showed the highest polymorphism and were more genetically diverse according to both Nei’s genetic diversity and the Shannon diversity index (Table 2). The percentage of polymorphism was above 67% when considering each species individually and 100% considering all species (Table 2). The genetic differentiation index (GST = 0.27) showed that populations were highly structured. The dendrogram obtained based on Nei’s genetic distance (Fig. 2) showed that F. meridionale and F. austroamericanum were more genetically related and that the F. graminearum was the most basal. One isolate of F. austroamericanum and three of F. meridionale were genetically more similar to F. meridionale and F. austroamericanum, respectively (Fig. 2). The PCoA reduced the dimensionality of the data and allowed the observation of the spatial structure of the isolates and their contribution to the diversity of the species (Fig. 3). Coordinates 1, 2, and 3 explained 21.31%, 13.07%, and 10.62% of the observed genetic variability, respectively. The cumulative explanation of the observed genetic diversity by the three coordinates was 45.00% (Fig. 3). Additionally, the PCoA allowed us to observe that most isolates were structured according to the species they belonged to, in accordance with the clusters observed in the dendrogram (Fig. 2).
Dendrogram based on Nei’s genetic distance of the Fusarium isolates sampled in southern Brazil. The dendrogram was obtained using 112 ISSR (inter-simple sequence repeat) loci and by the neighbor-joining grouping method. The numbers on the branches indicate the bootstrapping values obtained with 1000 replicates. Only a bootstrapping value above 50 was shown
In the simulations made by the STRUCTURE program using Bayesian inference, the highest probabilities were for the number of Ks (clusters, genetic groups) of two and four (Fig. 4). In K = 2, the species F. meridionale and F. austroamericanum were predominant in the genetic group represented by green, whereas F. graminearum was predominant in the genetic group represented by red (Fig. 5). At K = 4, each species exhibited predominance in a separate genetic group. The magenta-colored genetic group is the one distributed among the three species studied (Fig. 5).
Discussion
Genetic variability is the driving factor for evolution because it allows for adaptation (Nevo 1978; Barrett and Schluter 2008). In this sense, the knowledge of the genetic variability in populations of plant pathogens allows the prediction of their adaptive behavior in the face of the genetic resistance of the host and chemical control. The ability to predict a probable evolutionary behavior of a pathogen can assist in the development of management strategies to reduce losses or even prevent epidemics (Burdon et al. 2006). The species F. graminearum, F. meridionale, and F. austroamericanum evaluated in our study showed high molecular variability and genetic diversity, from which a high capacity for adaptation can be inferred. High genetic structuring was also observed in populations, corroborating the concept of phylogenetic species proposed by O’Donnell et al. (2004).
High genetic variability in the species included in this study is evidenced by the percentage of polymorphism (Table 2) and no polytomy in the dendrogram (Fig. 1). For the F. austroamericanum and F. meridionale species, there was no previous literature information on the percentage of polymorphism. Thus, in comparison with studies conducted with F. graminearum can help us understand the variability in these species. For F. graminearum, polymorphism has been variable using dominant markers [ISSR and AFLP (amplified fragment length polymorphism)]. Albayrak et al. (2016) evaluated 405 ISSR loci in 43 F. graminearum isolates from Turkey and Iran and only 5.62% were polymorphic. Mishra et al. (2004), evaluating eight ISSR primers in 70 F. graminearum isolates from Canada, reported polymorphism ranging from 56 to 85%. Malbrán et al. (2019) obtained 71% polymorphism by evaluating 80 ISSR loci in 109 F. graminearum isolates sampled in Argentina, whereas Palacios et al. (2017), using 72 ISSR loci in 62 isolates from the same country, obtained 92% polymorphism. Pan et al. (2016), evaluating 172 AFLP loci in 103 F. graminearum isolates from Uruguay, observed 89.5% of polymorphism. In southern Brazil, Astolfi et al. (2011) evaluating populations (111 isolates) from three counties in Rio Grande do Sul using 89 AFLP loci identified polymorphism ranging from 63 to 67%. Cunha et al. (2015) evaluating 54 ISSR loci in 152 isolates of F. oxysporum f. sp. cubense, sampled in 21 counties in southern Brazil, obtained 81% polymorphism. In this sense, comparing our data with the polymorphism data for F. graminearum at a global level and for the Fusarium genus at a regional level leads to the conclusion that F. austroamericanum and F. meridionale populations have high polymorphism. Nonetheless, it is important to emphasize that the isolates of the two species we studied were from a small geographical area compared with that of other studies in the literature, which further strengthens the finding of high polymorphism in these populations. The same comparison above applied to the F. graminearum population evaluated in our study also showed the high polymorphism of this species in the studied region. Polymorphism levels are important information when working with dominant markers because they are one of the parameters that can be used to indicate genetic variability (Fedrigo et al. 2016; Rosa et al. 2017; Mariano et al. 2019). The high polymorphism observed in the populations of F. graminearum, F. meridionale, and F. austroamericanum in southern Brazil implies high genetic variability in these species; thus, presenting a high adaptative potential.
When comparing the values of genetic diversity in the three species studied (Table 2) with data from the literature, we observed that the values we obtained were similar. Mishra et al. (2004) reported Nei’s genetic diversity ranging from 0.18 to 0.23 in isolates of F. graminearum sampled in Canada. Talas et al. (2011) evaluating F. graminearum isolates from Germany using microsatellite markers obtained Nei’s genetic diversity ranging from 0.30 to 0.54 and Shannon diversity from 0.53 to 0.96. For the southern region of Brazil, Astolfi et al. (2011), using AFLP markers, obtained Nei’s genetic diversity ranging from 0.12 to 0.13 in populations of F. graminearum when all polymorphic loci were used. Among the three populations studied by us, F. graminearum presented the highest genetic diversity, corroborating the data on genetic variability. For F. meridionale and F. austroamericanum species, there is no data on diversity in the literature; however, the comparison with data from other species of the genus Fusarium and with consideration for the limited geographical origin and number of our isolates, we can conclude that besides the high genetic variability, these species also have high genetic diversity, which could enhance their adaptive capacity.
The dendrogram (Fig. 2) corroborated the high genetic variability and diversity and indicated no polytomy. Additionally, it allows observing genetic proximity between the F. meridionale and F. austroamericanum species (Fig. 4). Also, in the dendrogram considering isolates (Fig. 2), it was possible to observe the high structuring of the F. graminearum species. The positioning in the dendrogram of three isolates of F. meridionale and three of F. austroamericanum among the main groups of F. austroamericanum and F. meridionale may be due to the low number of isolates used or due to the resolving power of the marker used. Albayrak et al. (2016) evaluating 43 isolates of F. graminearum and 20 of F. culmorum from Turkey also obtained groups in the dendrogram according to the species and with no polytomy; however, the isolates within each group were more genetically similar as compared with those in our study. Dinolfo et al. (2010) evaluating F. poae isolates from Argentina and England were unable to separate the isolates according to their geographical origin and also observed polytomy in the dendrogram. The results of these studies show that there are differences in genetic diversity and genetic relationship between isolates both when comparing species and when considering geographic regions. These results justify the need for studies of populations in less geographic coverage to better understand the dynamics of the variability in Fusarium species at the regional scale. This information can assist in the development of more effective strategies for managing the pathogen in regions with a history of FHB epidemics in wheat.
The PCoA results (Fig. 3) corroborated those of the dendrogram and showed that the genetic variation is well distributed among the species and also illustrated genetic proximity between the F. meridionale and F. austroamericanum species. Fusarium graminearum was the species that most contributed to the variation in the group, which was expected because it had greater variability and diversity and also by the number of isolates used. Bayesian inference analysis showed that F. meridionale and F. austroamericanum were the species with the greatest genetic similarity when considering K = 2, corroborating the data from the dendrogram and PCoA. Furthermore, when considering K = 4 (which also presented strong support in explaining the genetic variation of populations), the species were more distinguishable, with each one presenting a separate genetic group. One of the genetic groups (represented by the magenta color) is not predominant in any species and seems to be responsible for the genetic relationships among the species (Fig. 5). The number of genetic groups that best explains the genetic variation of populations or species is dependent on the type of data analyzed (species or populations of the same species) and the sample size. For example, Altinok et al. (2018) evaluating 200 isolates from Turkey observed five genetic groups (K = 5) in populations of F. oxysporum f. sp. melongenae. Yerkovich et al. (2020) evaluating 75 F. graminearum isolates from Argentina observed two genetic groups. In our study, the limited number of isolates used of each species may be the cause of not obtaining K = 3 which would be expected since the number of species evaluated.
In this study, the fixation index (GST = 0.27) indicates a very high genetic structure in the populations, corroborating the proposal by O’Donnell et al. (2004) of phylogenetic species in the FGSC. Additionally, the dendrogram obtained by Nei’s genetic distance (Fig. 2) and the structuring data obtained by Bayesian inference (Fig. 5) showed high genetic differentiation among populations, which was expected based on populations of distinct species. Nonetheless, the genetic relationship observed between populations was in agreement with that proposed by O’Donnell et al. (2004), who identified a greater genetic relationship between the species F. meridionale and F. austroamericanum when compared with F. graminearum.
The occurrence of more than one species capable of causing FHB in wheat fields in southern Brazil and the high genetic variability and diversity of the populations of these species may be an aggravating factor in the control of those pathogens. High genetic variability and diversity of populations can result in a failure in using fungicides for the chemical control of the pathogen (Georgopoulos and Skylakakis 1986), as well as of the ineffectiveness of resistant cultivars (McDonald and Linde 2002). This is because pathogen populations with high variability and genetic diversity have vastly variable genetic constitutions that allow for the rapid selection of genotypes that override chemical control or resistance of cultivars. Feksa et al. (2019) carried out evaluations using a series of fungicides with various active ingredients to prevent or control FHB pathogens in wheat inoculated with a mix of 10 isolates with genotypes 15-ADON, 3-ADON, and NIV. The results suggested that a better strategy to control FHB pathogens and decrease DON production in the humid regions of the state of Paraná should involve multiple applications of fungicides with various modes of action. That study included some isolates used in our work. The high genetic variability observed in the FGSC members causing FHB in the state of Paraná in our study might explain the variation in the FHB severity and DON production found by Feksa et al. (2019). This observation strengthens the hypothesis regarding the difficulty in controlling the pathogens when are present a series of related species with high genetic variability.
The overall analysis of our data together with the literature showed that the high variability found in the populations of F. graminearum, F. austroamericanum, and F. meridionale that cause FHB in southern Brazil is an important factor to be considered in the chemical control of the pathogen with fungicides and through breeding programs aimed at obtaining resistant wheat cultivars. Certainly, selection strategies in breeding programs can be more effective when lineages are selected considering more than one isolate of the causal agent of FHB.
References
Albayrak G, Yörük E, Gazdağli A, Sharifnabi B (2016) Genetic diversity among Fusarium graminearum and F. culmorum isolates based on ISSR markers. Archives of Biological Sciences 68:333–343
Altinok HH, Can C, Altinok MA (2018) Characterization of Fusarium oxysporum f. sp. melongenae isolates from Turkey with ISSR markers and DNA sequence analyses. European Journal of Plant Pathology 150:609–621
Alvarez CL, Azcarate MP, Pinto VF (2009) Toxigenic potential of Fusarium graminearum sensu stricto isolates from wheat in Argentina. International Journal of Food Microbiology 135:131–135
Amarasinghe C, Sharanowski B, Fernando WGD (2019) Molecular phylogenetic relationships trichothecene chemotype diversity and aggressiveness of isolates in a global collection of Fusarium graminearum species. Toxins 11:e263
Aoki T, Ward TJ, Kistler HC, O’Donnell K (2012) Systematics phylogeny and trichothecene mycotoxin potential of Fusarium head blight cereal pathogens. Mycotoxins 62:91–102
Arif M, Zaidi NW, Haq QMR, Singh US (2008) Genetic variability within Fusarium solani as revealed by PCR-fingerprinting based on ISSR markers. Indian Phytopathology 61:305–310
Astolfi P, dos Santos J, Schneider L, Gomes LB, Silva CN, Tessmann DJ, Del Ponte EM (2011) Molecular survey of trichothecene genotypes of Fusarium graminearum species complex from barley in southern Brazil. International Journal of Food Microbiology 148:197–201
Barrett RD, Schluter D (2008) Adaptation from standing genetic variation. Trends in Ecology & Evolution 23:38–44
Bertero A, Moretti A, Spicer LJ, Caloni F (2018) Fusarium molds and mycotoxins: potential species-specific effects. Toxins 10:e244
Boff T, Espindula LF, Bücker-Neto L, Minella E, Milacha SCK, Da-Silva PR (2019) Inheritance of aluminum tolerance in the wheat cultivar Toropi and new findings about the introduction of this trait into the Brazilian wheat germplasm. Environmental and Experimental Botany 157:91–99
Boutigny A-L, Ward TJ, Van Coller GJ, Flett B, Lamprecht SC, O’Donnell K, Viljoen A (2011) Analysis of the Fusarium graminearum species complex from wheat, barley and maize in South Africa provides evidence of species-specific differences in host preference. Fungal Genetics and Biology 48:914–920
Burdon JJ, Thrall PH, Ericson L (2006) The current and future dynamics of disease in plant communities. Annual Review of Phytopathology 44:19–39
Chaves MS, Martinelli JA, Wesp-Guterres C, Graichen FAS, Brammer SP, Scagliusi SM, Da-Silva PR, Wietholter P, Torres GAM, Lau EY, Consoli L, Chaves ALS (2013) The importance for food security of maintaining rust resistance in wheat. Food Security 5:157–176
Cunha CMS, Hinz RH, Pereira A, Tcacenco FA, Stadnik MJ (2015) Aggressiveness and genetic diversity of Fusarium oxysporum f. sp. cubense from Santa Catarina, southern Brazil. Tropical Plant Pathology 40:326–334
Dean R, Van Kan JAL, Pretorius ZA, Hammond-Kosack KE, Di Pietro A, Spanu PD, Rudd JJ, Dickman M, Kahmann R, Ellis J, Foster GD (2012) The top 10 fungal pathogens in molecular plant pathology. Molecular Plant Pathology 13:414–430
Del Ponte EM, Fernandes JMC, Pavan W, Baethgen WE (2009) A model-based assessment of the impacts of climate variability on Fusarium head blight seasonal risk in southern Brazil. Journal of Phytopathology 157:675–681
Del Ponte EM, Spolti P, Ward TJ, Gomes LB, Nicolli CP, Kuhnem PR, Siva CN, Tessmann DJ (2015) Regional and field-specific factors affect the composition of Fusarium head blight pathogens in subtropical no-till wheat agroecosystem of Brazil. Phytopathology 105:246–254
Desjardins AE, Proctor RH (2011) Genetic diversity and trichothecene chemotypes of the Fusarium graminearum clade isolated from maize in Nepal and identification of a putative new lineage. Fungal Biology 115:38–48
Dinolfo MI, Stenglein SA, Moreno MV, Nicholson P, Jennings P, Salerno GL (2010) ISSR markers detect high genetic variation among Fusarium poae isolates from Argentina and England. European Journal of Plant Pathology 127:483–491
Divakara ST, Santosh P, Aiyaz M, Ramana MV, Hariprasad P, Nayaka SC, Niranjana SR (2013) Molecular identification and characterization of Fusarium spp. associated with sorghum seeds. Journal of the Science of Food and Agriculture 94:1132–1139
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 9:11–15
Earl DA, VonHold BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14:2611–2620
Fedrigo K, Giacomin RM, Faria CMDR, Da-Silva PR (2016) ISSR primers for analysis of genetic variability of Stenocarpella maydis. Tropical Plant Pathology 41:1–6
Feksa HR, Couto HTZ, Garozi R, Almeida JL, Gardiano CG, Tessmann DJ (2019) Pre- and postinfection application of strobilurin-triazole premixes and single fungicides for control of fusarium head blight and deoxynivalenol mycotoxin in wheat. Crop Protection 117:128–134
Fernández Pinto VE, Terminiello L, Basilico JC, Rittieni A (2008) Natural occurrence of Nivalenol and mycotoxigenic potential of Fusarium graminearum isolates in wheat affected by head blight in Argentina. Brazilian Journal of Microbiology 39:157–162
Georgopoulos SG, Skylakakis G (1986) Genetic variability in the fungi and the problem of fungicide resistance. Crop Protection 5:299–305
Hampl V, Pavlícek A, Flegr J (2001) Construction and bootstrap analysis of DNA fingerprinting-based phylogenetic trees with a freeware program FreeTree: application to trichomonad parasites. International Journal of Systematic and Evolutionary Microbiology 51:731–735
Laday M, Juhàsz A, Mulè G, Moretti A, Szécsi A, Logrieco A (2004) Mitochondrial DNA diversity and lineage determination of European isolates of Fusarium graminearum (Gibberella zeae). European Journal of Plant Pathology 110:545–550
Malbrán I, Mourelos CA, Girotti JR, Balatti PA, Lori GA (2014) Toxigenic capacity and trichothecene production by Fusarium graminearum isolates from Argentina and their relationship with aggressiveness and fungal expansion in the wheat spike. Phytopathology 104:357–364
Malbrán I, Mourelos CA, Balatti PA, Lori GA (2019) Aggressiveness and genetic variability of Fusarium graminearum populations from the main wheat production area of Argentina. Journal of King Saud University - Science 31:1215–1219
Mariano LC, Zchonski FL, da Silva CM, Da-Silva PR (2019) Genetic variability in a Brazilian apple germplasm collection with low chilling requirements. PeerJ 6:e6265
McDonald BA, Linde C (2002) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
Mishra PK, Tewari JP, Clear RM, Turkington TK (2004) Molecular genetic variation and geographical structuring in Fusarium graminearum. The Annals of Applied Biology 145:299–307
Nei M (1972) Genetic distance between populations. The American Naturalist 106:283–291
Nevo E (1978) Genetic variation in natural populations: patterns and theory. Theoretical Population Biology 13:121–177
O’Donnell K, Kistler HC, Tacke BK, Casper HH (2000) Gene genealogies reveal global phylogeographic structure and reproductive isolation among lineages of Fusarium graminearum, the fungus causing wheat scab. Proceedings of the National Academy of Sciences of the United States of America 97:7905–7910
O’Donnell K, Ward TJ, Geiser DM, Kistler HC, Aoki T (2004) Genealogical concordance between the mating type locus and seven other nuclear genes supports formal recognition of nine phylogenetically distinct species within the Fusarium graminearum clade. Fungal Genetics and Biology 41:600–623
O’Donnell K, Ward TJ, Aberra D, Kistler HC, Aoki T, Orwig N, Kimura M, Bjørnstad Å, Klemsdal SS (2008) Multilocus genotyping and molecular phylogenetics resolve a novel head blight pathogen within the Fusarium graminearum species complex from Ethiopia. Fungal Genetics and Biology 45:1514–1522
Ortega LM, Dinolfo MI, Astoreca AL, Alberione EJ, Stenglein SA, Alconada TM (2016) Molecular and mycotoxin characterization of Fusarium graminearum isolates obtained from wheat at a single field in Argentina. Mycological Progress 15:e1
Palacios SA, Merlera GG, Erazo J, Reynoso MM, Farnochi MC, Torres AM (2017) Trichothecene genotype and genetic variability of Fusarium graminearum and F. cerealis isolated from durum wheat in Argentina. European Journal of Plant Pathology 149:969–981
Palazzini JM, Yerkovich N, Alberione E, Chiotta M, Chulze SN (2017) An integrated dual strategy to control Fusarium graminearum sensu stricto by the biocontrol agent Streptomyces sp. RC 87B under field conditions. Plant Gene 9:13–18
Pan D, Mionetto A, Calero N, Reynoso MM, Torres A, Bettucci L (2016) Population genetic analysis and trichothecene profiling of Fusarium graminearum from wheat in Uruguay. Genetics and Molecular Research 15:1–11
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research - an update. Bioinformatics 28:2537–2539
Pritchard JK, Wen X, Falush D (2010) Documentation for STRUCTURE software: version 2.3. University of Chicago, Chicago, USA
Ramírez ML, Reynoso MM, Farnochi MC, Chulze S (2006) Vegetative compatibility and mycotoxin chemotypes among Fusarium graminearum (Gibberella zeae) isolates from wheat in Argentina. European Journal of Plant Pathology 115:139–148
Ramirez ML, Reynoso MM, Farnochi MC, Torres AM, Leslie JF, Chulze SN (2007) Population genetic structure of Gibberella zeae isolated from wheat in Argentina. Food Additives and Contaminants 24:1115–1120
Reynoso MM, Ramirez ML, Torres AM, Chulze SN (2011) Trichothecene genotypes and chemotypes in Fusarium graminearum strains isolated from wheat in Argentina. International Journal of Food Microbiology 145:444–448
Rohlf FJ (2000) NTSYS-pc numerical taxonomy and multivariate analysis system version 22. Exeter Software, Setauket, New York
Rosa J, Weber GG, Cardoso R, Górski F, Da-Silva PR (2017) Variability and population genetic structure in Achyrocline flaccida (Weinm.) DC., a species with high value in folk medicine in South America. PLoS ONE 12:e0183533–e0183531
Santillán-Mendoza R, Pineda-Vaca D, Fernández-Pavía SP, Montero-Castro JC, Goss EM, Benítez-Malvido J, Rodríguez-Alvarado G (2019) Genetic diversity of Fusarium mexicanum, causal agent of mango and big-leaf mahogany malformation in Mexico. Molecular Biology Reports 46:3887–3897
Sarver BAJ, Ward TJ, Gale LR, Broz K, Kistler HC, Aoki T, Nicholson P, Carter J, O’Donnell K (2011) Novel Fusarium head blight pathogens from Nepal and Louisiana revealed by multilocus genealogical concordance. Fungal Genetics and Biology 48:1096–1107
Schothorst RC, van Egmond HP (2004) Report from SCOOP task 3.2.10 “collection of occurrence data of Fusarium toxins in food and assessment of dietary intake by the population of EU member states”: subtask: trichothecenes. Toxicology Letters 153:133–143
Scoz LB, Astolfi P, Reartes DS, Schmale DG III, Moraes MG, Del Ponte EM (2009) Trichothecene mycotoxin genotypes of Fusarium graminearum sensu stricto and Fusarium meridionale in wheat from southern Brazil. Plant Pathology 58:344–351
Sobrova P, Adam V, Vasatkova A, Beklova M, Zeman L, Kizek R (2010) Deoxynivalenol and its toxicity. Interdisciplinary Toxicology 3:94–99
Starkey DE, Ward TJ, Aoki T, Gale LR, Kistler HC, Geiser DM, Suga H, Tóth B, Varga J, O’Donnell K (2007) Global molecular surveillance reveals novel Fusarium head blight species and trichothecene toxin diversity. Fungal Genetics and Biology 44:1191–1204
Suga H, Karugia GW, Ward T, Gale LR, Tomimura K, Nakajima T, Miyasaka A, Koizumi S, Kageyama K, Hyakumachi M (2008) Molecular characterization of the Fusarium graminearum species complex in Japan. Phytopathology 98:159–166
Talas F, Parzies HK, Miedaner T (2011) Diversity in genetic structure and chemotype composition of Fusarium graminearum sensu stricto populations causing wheat head blight in individual fields in Germany. European Journal of Plant Pathology 131:39–48
Umpiérrez-Failache M, Garmendia G, Pereyra S, Rodríguez-Haralambides A, Ward TJ, Vero S (2013) Regional differences in species composition and toxigenic potential among Fusarium head blight isolates from Uruguay indicate a risk of nivalenol contamination in new wheat production areas. International Journal of Food Microbiology 166:135–140
Van der Lee T, Zhang H, van Diepeningen A, Waalwijk C (2015) Biogeography of Fusarium graminearum species complex and chemotypes: a review. Food Additives and Contaminants 32:453–460
Varga E, Wiesenberger G, Hametner C, Ward TJ, Dong Y, Schöfbeck D, McCormick S, Broz K, Stückler R, Schuhmacher R, Krska R, Kistler HC, Berthiller F, Adam G (2015) New tricks of an old enemy: isolates of Fusarium graminearum produce a type a trichothecene mycotoxin. Environmental Microbiology 17:2588–2600
Wang C, Cheng Y (2017) Identification and trichothecene genotypes of Fusarium graminearum species complex from wheat in Taiwan. Botanical Studies 58:e4
Wang J, Zhao Z, Yang X, Yang J, Gong A, Zhang J, Chen L, Zhou C (2019) Fusarium graminearum species complex and trichothecene genotype. In: JW DV, Trucksess MW, Jackson LS (eds) Mycotoxins and food safety, vol 2019, 1st edn. IntechOpen, London UK, pp 1–19
Ward TJ, Clear RM, Rooney AP, O’Donnell (2008) An adaptive evolutionary shift in Fusarium head blight pathogen populations is driving the rapid spread of more toxigenic Fusarium graminearum in North America. Fungal Genetics and Biology 45:473–484
Windels CE (2000) Economic and social impacts of Fusarium head blight: changing farms and rural communities in the Northern Great Plains. Phytopathology 90:17–21
Xia R, Schaafsma AW, Wu F, Hooker DC (2020) Impact of the improvements in Fusarium head blight and agronomic management on economics of winter wheat. World Mycotoxin Journal (online first):1–18. https://doi.org/10.3920/WMJ2019.2518
Yeh FC, Yang RC, Boyle T (1999) POPGENE: Microsoft Windows-based freeware for population genetic analysis Version 1.31. University of Alberta Canada and Centre for International Forestry Research, Canada
Yerkovich N, Fumero MV, Cantoro R, Palazzini JM, Chulze SN (2020) Population structure and genetic diversity of Fusarium graminearum sensu stricto, the main wheat pathogen producing Fusarium head blight in Argentina. European Journal of Plant Pathology 156:635–646
Yli-Mattila T, Gagkaeva T, Ward TJ, Aoki T, Kistler HC, O’Donnell K (2009) A novel Asian clade within the Fusarium graminearum species complex includes a newly discovered cereal head blight pathogen from the Russian Far East. Mycologia 101:841–852
Zeller KA, Bowden RL, Leslie JF (2003) Diversity of epidemic populations of Gibberella zeae from small quadrats in Kansas and North Dakota. Phytopathology 93:874–880
Zeller KA, Bowden RL, Leslie JF (2004) Population differentiation and recombination in wheat scab populations of Gibberella zeae from the United States. Molecular Ecology 13:563–571
Zhang H, Van der Lee T, Waalwijk C, Chen W, Xu J, Xu J, Zhang Y, Feng J (2012) Population analysis of the Fusarium graminearum species complex from wheat in China show a shift to more aggressive isolates. PLoS ONE 7:e31722
Zhang H, Luo W, Pan Y, Xu J, Xu JS, Chen WQ, Feng J (2014) First report of Fusarium maize ear rot caused by Fusarium meridionale in China. Plant Disease 98:e1156
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeats (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Funding
The authors thank Fundacão Araucária de Apoio ao Desenvolvimento Científico e Tecnológico do Estado do Paraná (grant number 229/2010), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, CAPES (grant number 001), and Cooperativa Agrária Agroindutrial for financial support to this research.
Author information
Authors and Affiliations
Contributions
Conceptualization: PRS, DJT, MHMA; methodology: PRS, MHMA, FLZ, YKS, DLL; formal analysis and investigation: PRS, MHMA, FLZ, YKS, DLL; writing—original draft preparation: MHMA, FLZ; writing—review and editing: Paulo PRS, DJT, MHMA; funding acquisition: PRS, DJT, MHMA; supervision: PRS. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(TXT 12 kb)
Rights and permissions
About this article
Cite this article
de Arruda, M.H.M., Zchosnki, F.L., Silva, Y.K. et al. Genetic diversity of Fusarium meridionale, F. austroamericanum, and F. graminearum isolates associated with Fusarium head blight of wheat in Brazil. Trop. plant pathol. 46, 98–108 (2021). https://doi.org/10.1007/s40858-020-00403-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-020-00403-3