Abstract
Purpose of Review
Ecological models can provide critical guidance to conservation programs both as problem-solving tools and by projecting future outcomes, specifically when time and resources limit directly testing alternative management approaches and scenarios. Due to the complexity of aquatic systems, environmental and climatic factors co-vary, multiple risk factors interact, and driving ecological and evolutionary processes are characterized by non-linear, higher-order interactions. Recent modeling advancements allow for better accounting of variation across time and space in ecological and genetic processes, but more progress is needed to inform conservation and address biodiversity decline. Modeling approaches that can explicitly incorporate the ongoing, rapid transformation of climate and landscapes and demogenetic and eco-evo consequences are useful for supporting and informing conservation planning strategies. In this narrative perspective, we present the history and role of individual-based models (IBMs) in aquatic systems to guide management.
Recent Findings
We present exemplary cases that cover (1) the conservation and management of native species in systems impacted by invasive species, (2) life history evolution impacts on the management of fisheries, (3) predictions of the interaction between changing environments and management decisions, and (4) testing factors that drive system dynamics in order to prioritize management decisions. We summarize potential platforms and software available to researchers and managers and discuss future opportunities and challenges.
Summary
While this review focuses on the use of IBMs in aquatic systems, we assert that this foundational knowledge is applicable across systems and encourages researchers and managers to consider incorporating individual-based modeling perspectives to inform conservation as appropriate.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Individual-based models (IBMs)—also known as agent-based models (ABMs)—have a long history across a wide array of systems for simulating bottom-up mechanistic processes and producing emergent top-down patterns (see Box 1 for definitions of italicized terminology). IBMs are often used when considering individually variable traits, such as behavior and physiology, necessary to produce mechanistic simulations to understand population-level patterns and processes and have been broadly applied in aquatic ecology and management [1]. While IBMs have been widely adopted in academic research focused on aquatic systems, their implementation into plans for conservation and management has lagged (i.e., the research-implementation gap [2]). IBMs have at times been criticized for being a “black box” due to a lack of transparency and approachability. Our primary goals with this review/prospectus are to highlight the utility of IBMs for practical applications while opening the IBM “black box” for practitioners in aquatic systems. First, we introduce the general concepts, history, and terminology related to IBMs and describe their potential for research and management such as those related to aquatic systems, specifically for eco-evolutionary (i.e., “eco-evo”) simulation modeling incorporating demogenetic processes. Second, we provide case studies to highlight the utility and practicality of using IBMs to answer a wide variety of eco-evo questions important for the management and conservation of aquatic ecosystems. We focus on the current utility and future potential of IBMs for eco-evo processes and highlight studies that have used this approach, including (1) the conservation and management of native species in systems impacted by invasive species, (2) life history evolution and implications for fisheries management, (3) predictions of the interaction between changing environments and management decisions, and (4) testing factors that drive system dynamics in order to evaluate consequences of management actions. Third, we discuss the future of IBMs, including ongoing challenges, opportunities, and future considerations. In sum, we hope to provide encouragement and foundational knowledge for researchers to consider supplementing current projects with IBMs where appropriate.
IBM Concepts, History, and Usage
What is an IBM? Definition and History in Aquatic Systems
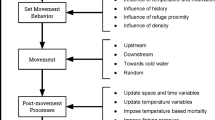
At their core, IBMs differ from analytical or equation-based models in that they simulate individual organisms, heterogeneity among individuals, and interactions among individuals and between individuals and their environment to consider changes through time and space (modified from [3] and [4]). This definition that we present represents an inclusive definition of both IBMs and ABMs, with ABMs being explicit about resource availability, adaptive behavior, and variation within age classes (for IBM versus ABM, see Box 1). However, the overall process for an IBM is similar to other modeling frameworks and generally may follow an iterative framework or “modeling cycle” that includes the development of research questions, software selection or model creation, and sensitivity analyses among other steps (Section 2.3 of [3]; Fig. 1).
Iterative model process, starting with the co-development of research questions among researchers and managers which can be applied to individual-based models to co-develop research and build toward making management decisions. After determining questions which need to be answered with simulations of individuals, factors within the environment will need to be selected, with an understanding of simplifying assumptions and pieces not captured by reality. Then, researchers should decide either the software or, if developing the model from scratch, the coding language (Table 1). After that, empirical data will need to be collected and the literature reviewed to parameterize the model. Once the model is created, it can be validated and evaluated, and then, researchers should complete a sensitivity analysis of model outcomes to different parameters. This can lead to inferences about model structure and the system, which may result in the need to collect more data and better parameterize the models. Conversely, these inferences may lead to sharing the results with collaborators and then publishing. This process may be implemented within structured decision-making and adaptive management but may also be pursued independently
Similar to various other models, IBMs can be used to forecast or hindcast and are generally appropriate for evaluating diverse scenarios. IBMs may be particularly useful when average values describing population characteristics are insufficient, and variation among individuals’ attributes (e.g., location, size) is critical for the effective modeling of social and ecological processes [5]. The process-based nature of IBMs also makes them useful when considering modeling when future or past conditions are outside of the values initially used to create a model. Modeling outside of the bounds of creation is inappropriate for several other model types. They also are useful for exploring emergent properties of complex systems [6], which arise from the fine-scale interactions occurring at the individual level. Many studies have used IBMs to address ecological questions, with several syntheses available on behavior, such as habitat selection [7, 8] and social interactions [9]. By coupling with geographic information systems (GIS), IBMs can directly incorporate real, spatially explicit environmental factors to answer questions related to spatial ecology and animal movements [5, 10]. This ability to incorporate spatially explicit data points to the particular utility of IBMs for landscape-level research. Evolutionary processes, however, have received less attention in the IBM literature, although the incorporation of evolution into ecological IBMs has been growing including for management purposes [11, 12], driven by research questions requiring the inclusion of individual genotypes (e.g., thermal tolerance and gene flow [13]) and increasing model complexity coupled with faster processing speeds and greater computational power.
Why use IBMs?
The reasons for the current growth of IBM use are similar to the three reasons previously outlined by Judson [14]: the difficulties of generality and modeling in ecology, ecological interactions of individuals are important, and computational resources, including computer power and software, have increased accessibility of modeling. As we learn more about systems, their complexity, or the number of independent and interacting components [15], makes generalized modeling efforts difficult [14]. Because individual organism characteristics and interactions between organisms may drive system patterns, understanding these processes has gained greater interest. Moreover, improvements in computational resources have only continued to expand since the publication of Judson [14].
IBMs are prevalent across scientific disciplines, including many subdisciplines of biology and social science, and as the field has grown and computational resources and methodologies have improved, researchers have been able to simulate individuals with both ecological and evolutionary processes occurring in tandem, moving into the realm of eco-evo simulations. For many research questions and management goals, the interaction of ecological and evolutionary processes is important for understanding overall system dynamics. Furthermore, these eco-evo processes and the emergent patterns of the system are often best addressed by simulating individuals, including population viability [16, 17], hybridization [18], invasive species management [19], and impacts of barriers to movement [20]. These eco-evo models expand the realm of areas where IBMs are able to help explore research questions related to management goals that cannot be effectively examined using empirical approaches alone or other modeling techniques. Because of the utility of IBMs to management where eco-evo models are beneficial, there are many examples in the literatures of their uses. This includes software that is ready to be used immediately, as well as modeling efforts where coding is done from the bottom up. Across these approaches, there are numerous examples in aquatic systems, often within a spatially explicit context (Table 1).
IBM Case Studies and Management/Conservation Applications for Aquatic Systems
Because IBMs are mechanistic and have the capability of being both broad- and fine-scale, they are particularly useful in the context of management programs. As compared to population-based and stage-structured models of differential equations, they also provide the ability to be more easily transferred to different systems, future scenarios (e.g., climate change), and novel management practices as they are developed due to the ability to better understand the emergent properties of the system while simulating the various mechanistic pieces at the individual level [25]. In addition, the emergence and transferability of mechanistic models make them attractive for their ability to be more robust to novel scenarios and environments than classical modeling approaches [25]. To illustrate these characteristics of IBMs, in each subsection below, we introduce the topic being covered, examples of IBM applications, and some of the reasons why IBMs provide a benefit over other modeling approaches. Although not all examples from the literature provided are eco-evo IBMs in aquatic systems, the topics being covered lend themselves to understanding potential future dynamics in aquatic systems. In addition, many of these aquatic examples are interested in questions at the landscape extent and are spatially explicit because of the necessity to include space when addressing management goals and research questions. However, in some cases, IBMs may be spatially implicit or have a pseudo-geography because they are based on relative distances or positions [26]. However, these may not be directly tied to environmental heterogeneity, and the position of features (landscape configuration) of the environment and may not be appropriate for certain research questions and fields (e.g., landscape genetics).
Models for Understanding and Mitigating Invasive Species Impacts
Management and conservation of native species in systems affected by invasive species present unique challenges to resource managers. Often, goals involve the evaluation of mechanisms underlying successful invasions and strategies to control or eradicate invasive species while simultaneously minimizing additional impacts on native species. IBMs can help model such systems by incorporating individual interactions which may be critical for determining population patterns and informing management decisions. An example of an IBM-informed management program that considers both native and non-native aquatic species is the invasive brook trout (Salvelinus fontinalis) suppression program in Sullivan Creek in northeastern Washington State, USA [27]. This program is focused on eliminating brook trout in Sullivan Creek to benefit the native cutthroat trout (Oncorhynchus clarkii lewisi). In concert with continued mechanical suppression activities in the watershed, a novel approach was introduced in which brook trout with two Y chromosomes (YY-males) are bred in captivity and then released in Sullivan Creek to reproduce with the naturalized brook trout population [28]. The goal of the YY-male program is to shift the sex ratio of the naturalized brook trout population until no females are left and the population is extirpated, because no long-term data sets exist to guide the best management actions, a spatially explicit IBM of the brook trout population to model the performance of released YY-males, which simulated the movement, reproduction, inheritance of chromosomes, and mortality of individual fish [29]. Using an IBM was an appropriate modeling approach, because there were no broad-scale patterns with which to consider potential outcomes. Instead, fine-scale, individual-based interactions parameterized with empirical data were used to ultimately scale up to emergent model outcomes (Fig. 2 [30]). This IBM has been critical for informing the best approach for implementing the YY-male program in several ways. First, simulation outcomes provided annual benchmarks against which the ongoing management was compared and progress toward brook trout extirpation was reevaluated (Fig. 2). Second, the model identified key drivers of brook trout extirpation, informing the optimal use of limited resources and reducing time to achieving management goals. For example, simulations demonstrated a strong effect on the reproductive status of stocked YY fish, supporting the management decision to produce a higher ratio of mature to immature YY fish in the hatchery [30]. Finally, the modeling work identified key gaps in information, guiding empirical work that focused on factors such as YY-male dispersal and fitness because of the significant effect that these factors had on model outcomes [29]. In turn, ongoing data collection informed the accuracy of the model, which was then recalibrated to produce results that were more closely aligned with reality. Thus, management actions and the IBM worked iteratively to understand how to optimize outcomes for this novel management method.
Including invasive species impacts. Emergent simulation outcomes of brook trout abundance before and after introduction of YY males (top) and the proportion of ~ 200 m patches occupied in the stream network of the Sullivan Creek Watershed in Washington State (bottom) where brook trout are invasive. Lines represent averages across 10 simulation replicates and polygons represent 95% confidence bands. Population declines occur due to skewing the sex ratio through the release of captive-raised individuals. Figure reproduced from Day et al. [29]
Considering Life History Evolution Impacts on the Management of Fisheries
Evolutionary impacts have important implications for fisheries management [31, 32]. The potential evolutionary effects of fisheries harvest have been explored through a variety of study approaches including controlled experiments [33], genetic analyses [34], and statistical inquiries of multi-year assessment data [35]. However, taken as a whole, these approaches do not adequately account for variation across time, space, or ecogenetic processes driven by individual interactions, providing a unique opportunity for IBMs. IBMs are useful when considering fisheries-induced evolution given the interaction between ecological and evolutionary dynamics, the multi-generational effects of selection, and the ability to evaluate multiple harvest scenarios, as well as accounting for potential short-term ecological and long-term evolutionary consequences of environmental stressors and targeted management actions, such as harvest [36, 37]. Historically, IBMs allowed for individual phenotypic variation (e.g., in foraging, growth, and reproductive rates) to arise through chance encounters which would be exacerbated through interactions and feedbacks. Ecogenetic IBMs (or demo-genetic agent-based models [16]) incorporate genetic differences among individuals with phenotypic differences arising through a combination of genetic differences, stochastic events, and ecological feedbacks. These have been developed and applied to model fisheries-induced evolution in both generalized theoretical frameworks and to assess effects on specific freshwater and marine fish stocks. Ecogenetic IBMs represent a subset of potential eco-evo IBMs, although many models incorporating evolution may require genetic differences among individuals to be explicitly simulated.
As a specific example, fisheries harvest based on individual size or some other desirable characteristic can eliminate individuals with the highest reproductive output. Such discriminatory harvest exerts an additional selective force that can dramatically alter the combinations of life history traits that are appropriate for maximizing lifetime reproductive success. Due to this increased selection favoring certain sizes or characteristics, harvest can lead to changes in average behaviors, growth rates, maturation schedules, and ultimately the productive capacity of fish populations [38]. Several studies indicate that targeted harvesting of older, larger fish selects for maturation at an earlier age and smaller size [39]. Earlier maturation likely comes at a cost to growth and may exacerbate recruitment [40]. Evolution induced by harvest could decrease the productive capacity of fish stocks and the ability of these stocks to recover during fishery closures [41]. At the same time, detecting evolutionary effects from harvest is not straightforward. In particular, the ecological and evolutionary responses to harvest may actually mask each other. These models allow for fisheries harvest to act simultaneously on ecological and evolutionary dynamics, and patterns can be related to empirical data [42].
Collectively, such IBMs have evaluated the response to the harvest of traits such as maturation schedules, growth, behavior, and reproductive investment [16, 43, 44]. In turn, at a population level, IBMs have helped reveal effects on stock productive potential [45], the ability of stocks to recover during fishery closures [46], and potential economic repercussions of fisheries-induced evolution [47]. As a specific example, Ayllón et al. [48•] used IBMs to simulate the interaction of harvest and life history evolution with climate change. They looked at the response of brown trout (Salmo trutta) across > 1500 combinations of climate changes and harvest scenarios by using artificial neural networks and found that any size selection during the harvest process in conjunction with climate changes could lead to population declines. Even without climate change, conservation goals often were not met except under limited size-selection scenarios. Ultimately, they suggested that banning harvest may be necessary with climatic changes to conserve existing populations.
Incorporating Climate Change and Adaptive Capacity to Guide Management
The application of IBMs in understanding species response to climate change was identified by Shugart et al. [49] 30 years ago. Climate change is an inherently spatial process, including heterogeneous rates of warming across the globe [50]. The adaptive capacity of a species is its ability to accommodate, cope with, or respond to changing conditions, via dispersal, acclimation, or phenotypic plasticity, and evolution [51,52,53,54]. Overall, complex feedbacks exist between the three components of adaptive capacity, and IBMs provide a mechanism for controlling for a single factor while varying the others [11]. For example, evolution may reduce extinction risk through high genetic variance [50]. Because these processes span the interface of ecology and evolution which directly are impacted by how individuals interact with each other and the environment, complex IBMs can be useful for estimating range shifts and persistence. In contrast, simpler modeling approaches—including early species distribution modeling techniques—may not effectively capture important processes [55]. In addition, climate change impacts may have synergistic with landscape change in aquatic systems [56], increasing overall management challenges. The incorporation of multiple processes is challenging, and the ability to model some processes independently, such as plasticity, in a spatially explicit manner is still needing further development [11, 57]. These challenges have limited implementation of IBMs which cover the eco-evo processes influencing adaptive capacity; for example, in marine systems, there are a limited number of studies using IBMs to evaluate adaptive capacity and instead have focused only on genetic adaptation [11, 13]. However challenging, by modeling with the eco-evo processes present across a spatially heterogeneous environment, researchers may over- or underestimate adaptive capacity. For example, a model which only includes local adaptation may underestimate the probability of population persistence in species with high levels of phenotypic plasticity. One important aspect of modeling is that some processes may need to act independently, because local adaptation and plasticity may vary spatially (e.g., the size of response with latitude [58]). However, not all aspects are independent, such that plasticity itself may evolve, or that local adaptation is interconnected with gene flow [51, 59].
Multiple examples of forecasting models covering eco-evo processes and the impacts of climate change across varied taxa exist, with model complexity varying with a priori knowledge, computational needs, and specific aims of the project (see Table 1). Depending on the specific research question and system, these forecasting simulations can be population instead of individual-based [60]. Early IBMs related to climate change were focused on dispersal or other ecological processes [10]. Models now routinely track not just the individual location, phenotype, and movement, but also the underlying genotype. This allows for the investigation of eco-evo process relationships, such as dispersal and local adaptation [61]. From a management standpoint, these simulations may also highlight the impact of the translocation of individuals under an assisted gene flow framework [60, 62, 63].
Testing Factors that Drive System Dynamics in Order to Inform Management Decisions
A crucial step in all model development is to demonstrate that it accomplishes its intended purpose to answer the desired research question and produces outputs that match expectations under controlled test conditions. IBMs are developed using literature and field estimates. However, decision-makers need to know whether a model is a sufficiently accurate representation of a natural system and can answer their research question. In some cases, models may not need to perfectly represent reality if they are able to address the specific question [64]. The “evaludation” framework, which combines validated and evaluation, provides guidelines for understanding the effects of parameters on model processes and outcomes [65]. Without such an understanding, model results are vulnerable to spurious or biased outcomes, whereby model results are not a function of any tested mechanism but of random chance as a result of some particular combination of parameters. Spurious results can be limited by developing and testing mechanistic hypotheses within the model but also by conducting sensitivity analysis to understand relationships among model parameters and model outcomes [66, 67]. Sensitivity analysis becomes more important as uncertainty in model parameterization increases, and the range of uncertainty can be explored. Furthermore, sensitivity analysis should be conducted not only on key parameters of interest, but within the range of uncertainty on all model parameters to ensure that the model functions as intended and continues to behave realistically under extreme parameter values. It is also important to note that the sensitivity of aquatic systems to different factors may vary through time [68]. Which parameters to conduct the sensitivity analysis on and the breadth of parameter space explored, using methods such as Latin hypercube sampling or a fully factorial design, is an important consideration and may be limited based on available computational resources [69, 70]. Conversely, for some researchers, the ability to test an increasing range of parameter values and the overall number of models continues to become easier as access to computing clusters and the resources within those clusters continues to improve. Overall, sensitivity analysis highlights those parameters that have the greatest influence on simulation outcomes.
In IBMs, sensitivity analysis is particularly important because model outcomes and relationships among variables can be difficult to predict due to the emergence of patterns that result from many non-linear functions and fine-scale, including higher-order, interactions [71]. Multiple approaches and methods are available [66], but they can be broadly broken into two categories: global and local. Global sensitivity analysis is useful for parameters of low interest to the researcher and is conducted by simultaneously varying parameter values across hyper-dimensional parameter space and extracting the effects of individual parameters on model outcomes [29]. Local sensitivity analysis is most useful for key parameters of interest to model hypotheses and usually involves holding values for all but one parameter constant and exploring the parameter of interest [72]. In addition, sensitivity to model structure should be considered, such as changes in inference if certain processes are included or not, and the form of the process, such as whether the process is stochastic or not. Structural uncertainty and determining the necessary processes to answer the research question can be challenging [73].
To help ground this process, it is worth considering the example of Baggio et al. [74] which explored the genetic differentiation of fish populations separated by dams. Using a demographic-genetic model, researchers assessed movement within metapopulations before and after dams, with tributary spawning groups acting as subpopulations. Sensitivity analysis was conducted to determine which parameters most influenced population structure. They considered the straying rate, carrying capacity, mutation rates, mortality rate, and permeability of the system through the addition of a dam (see Table 2 of the manuscript). Sensitivity revealed that population structure stabilized around 200 generations, and it was not sensitive to mutation rate but was sensitive to straying rate, carrying capacity, and mortality rates. These relationships were not always linear, and these sensitivity analyses helped to highlight the need to monitor spawning sites and consider the impacts of different connectivity scenarios.
Current Challenges and Future Opportunities
Incorporating Multi-species Interactions
Many IBMs are developed for a single system, where a single species of interest is simulated to understand how individuals interact with one another and the abiotic environment. While valuable for understanding eco-evo mechanisms, single-species IBMs have a narrower scope of research questions they can answer, potentially limiting their ability to contribute to ecological theory or to conserve broader landscapes and functional communities [75]. Furthermore, these models do not capture the complex behaviors or demographics that are expressed as a result of multi-species interactions (e.g., competition, symbiosis).
Research paradigms are moving toward incorporating the effects of multi-species interactions, especially in conservation and connectivity modeling [76,77,78]. These approaches can incorporate the effects of species interactions (e.g., predation, competition, parasitism, mutualism) and community formation on ecological and evolutionary processes. They can also model scenarios with implications for many species or whole communities for broader application for research into conservation, community ecology, and ecological theory. These models can be difficult to generate, given they require parameterization of the generalized effects of landscapes and their environments with interactions across multiple species; however, they show great promise in providing an understanding of ecosystem properties and species demographics [79, 80].
For multi-species IBMs in aquatic systems, there is a robust body of research describing organismal and community ecology, including IBM software targeted toward multi-species models of fish and marine organisms (e.g., PISCATOR [81]). Many of these are focused on community structure [82, 83], food webs [84, 85], and population dynamics [86, 87]. While there are plentiful examples of ecological multi-species models in aquatic systems, those that explore evolutionary processes remain rare. An exciting opportunity is available to explore more evolutionary mechanisms [16], including hybridization, landscape connectivity, local and community adaptation, and how multi-species interactions can lead to co-expressed phenotypes. This expansion of IBMs to incorporate a greater range of research questions then leads to a greater range of applications for resource managers.
Genomics and Large-Scale Data Challenges
The field of evolutionary biology has experienced unprecedented advancements in high-throughput sequencing technologies over the past two decades increasing available genomic data and resources [88]. With these approaches, researchers can now explore molecular diversity across whole genomes to better understand demographics [89] and mechanisms that underpin phenotypes of interest [90] in populations of non-model organisms, with implications for informing an array of management actions. For example, a recent study by Yoshida and Yáñez [91] used dense genotyping to reveal five polymorphisms that explain most of the variation in body weight in rainbow trout (Oncorhynchus mykiss). To better understand ecological and evolutionary processes in wild systems, there is an opportunity to incorporate genomic information into IBMs. The integration of genetics into demographic models was a useful step in understanding structure and predictions for management interventions, such as translocations at the population level [12, 92, 93]. By shifting to simulating genomic data integrated into IBM demographic models, researchers are able to answer questions both about neutral processes determined with increased statistical power compared to genetics, such as population structure and migration rates, as well as adaptive processes, like responses to environmental change or local adaptation [94]. Without considering genomic data, the response to certain management goals, such as climate change adaptation, would not be possible.
We anticipate that this integration can be done in one of two ways. First, genomic-based demographic information (e.g., population structure, gene flow rates, hybridization zones) can be used to guide the development of more ecologically relevant models. Second, evolutionary mechanisms (e.g., trait heritability, plasticity, or the inheritance of genomic variants) can be incorporated into IBMs to simulate the inheritance and expression of genomic-based traits from one generation to the next. The latter approach is of interest to both ecological and evolutionary biologists alike, as it provides the ability to replicate evolutionarily relevant mechanisms (e.g., allele frequencies) and forecast the ecological and evolutionary ramifications of functional variation that are discovered. While there is an appetite in the research community to develop eco-evo IBMs [12, 95, 96], few researchers have incorporated genomic data into IBMs to date (but see also [95, 97]). This may be the result of a research gap between evolutionary biologists and ecological modelers or because it is difficult to model complex and large genomic datasets. The decision between selecting software and models that may be better at simulating genomics than ecology and landscape data is important to acknowledge; however, this division may be reduced either through software advances or the decision of researchers to employ multiple modeling methods [20]. This second approach is particularly useful, because a different modeling structure presents different assumptions. Hence, if two models of the same system give a different inference, the assumptions themselves may provide an important understanding of the system.
The combination of increased genomic, phenotypic, and environmental data availability, size, and complexity all act in conjunction to create data challenges in multiple forms when simulating individuals. For example, depending on the scale, it is computationally expensive to model the complexities of the genome across thousands of individuals. Coupling with the genomic data is the relationship of individual genotypes and phenotypes with the environment. Gene-by-environment interactions are one of the hallmarks of eco-evo IBMs. The growing challenges of big data with geographic information systems (GISs) and other remote sensing technologies have been reviewed previously [98], in addition to the ability of these technologies to enhance understanding of the landscape for the purposes of ecology resources [99]. For some research questions, the environment is also dynamic, and spatiotemporal data can cause additional challenges, such as attribute information, algorithms, and conceptual frameworks [99]. In addition, many ecologically relevant traits may be polygenic, which makes modeling in a mechanistic manner difficult, although this challenge is becoming easier to address due to advances in sequencing and modeling [100•]. If models include many genome-wide markers, high resolution or large extent environment data, and many individuals with long periods of time, it may be too computationally intensive to perform or for future researchers to replicate. There are potential solutions to these big data GIS challenges through cloud computing resources or machine learning algorithms, with particular benefits to parallel computing [101,102,103].
Continued Stakeholder Engagement and Integration with the Social-Ecological System
Previous studies have highlighted the importance of co-developing research questions with researchers and practitioners to bridge the research-implementation gap [2]. The ability to tie spatially explicit, customized IBMs to management questions helps provide context and can increase the sense of understanding and ownership of the data and results. Due to the extensive impact of humans on natural systems, understanding eco-evo processes and patterns is key to effective management [104•]. Early engagement and co-development, even with limited data, can help identify research priorities and data needs for further evaluation [105]. Engaging stakeholders and managers in the development of IBMs can ensure that the most relevant questions are being asked and therefore aid in effective decision-making [106]. Ideally, this is accomplished through a formal framework such as adaptive management [107], where IBMs can model a suite of alternative management actions to evaluate predicted outcomes, describe associated uncertainty, identify critical knowledge gaps, and optimize the allocation of limited resources.
Specifically, adaptive management and structured decision-making frameworks have often relied on modeling efforts to aid in decision-making [108,109,110]. IBMs, when used in conjunction with an iterative modeling framework and the co-production of research (Fig. 2), can be easily integrated within adaptive management and structured decision-making, although they are not obligate processes. IBMs can be particularly useful for adaptive management because management actions can be simulated and evaluated before any decisions are made, thus reducing the chances of surprises [111]. When multiple management decisions surrounding management, such as restoration, occur, it is possible that the different decisions may produce opposing outcomes [112]. In the scientific literature, sometimes the application of IBMs is limited to the discussion and potential for future expansion of the model to create a direct connection to management decisions, instead of the model creation and simulation experiments being intertwined with direct management decisions from the beginning. There are a number of examples where modeling was conducted with the management objective explicitly in mind [29]. Besides IBMs and other bottom-up approaches being better at exploring future novel conditions as previously discussed, they may also lend themselves well from the standpoint of communication among stakeholders and researchers. Although IBMs may encompass many parameters, often these parameters are more directly linked to the empirical data being collected (e.g., individual lengths), and individuals are an intuitive unit for simulating, drawing inference from, and communicating, as opposed to other population approaches (e.g., individual fecundity compared to intrinsic population growth rates) [113]. These two components can help with building trust in model outcomes and integration within management decisions and actions.
Placing model outcomes in a social-ecological system (SES) context by testing scenarios informed by a wide range of stakeholders can provide more relevant insight into the management applications of IBMs. The SES framework considers natural resources within the interactions and interdependence of the users and their governance [114], and integrating social and ecological IBMs can identify important dynamics between social and ecological systems that might otherwise be missed [111, 115]. As one example of the process of integrating IBMs with social-ecological research, Jossie et al. [116] used a spatially explicit IBM approach to explore how stakeholder-informed stream connectivity scenarios determined by semi-structured interviews analyzed with mental modeling software can influence hybridization dynamics of native Yellowstone cutthroat trout (Oncorhynchus clarkii bouvieri) and invasive rainbow trout (Oncorhynchus mykiss). In this example, humans are not directly simulated within the system to create a holistic IBM of the social-ecological system, but their actions defined the parameters and parameter values being explored to understand the tradeoff of managing for large body size, migratory life history variation, and exposure to invasive species. The connectivity scenarios were informed by a diverse array of stakeholders with a variety of interests including fisheries, agriculture, conservation, housing, and commercial development. Using IBMs in this study worked because we were able to simulate the contexts of real streams as described by the participants of the SES. Interestingly, the most important factors for achieving desirable outcomes for Yellowstone cutthroat trout (i.e., increased abundance) were the traits of the fish themselves and their behavior (i.e., large body size, migratory life history, and self-preference mating), rather than different scales and approaches to barrier removal identified by stakeholders. However, there were some differences among connectivity scenarios that may provide some insight into the possible outcomes of the different management approaches associated with each scenario. For example, scenarios with relatively high connectivity resulted in the complete loss of genetically “pure” Yellowstone cutthroat trout when there was no reproductive separation modeled, even when large, migratory Yellowstone cutthroat trout were modeled. Using a spatially explicit IBM offered the ability to further understand how fish behavior (i.e., expressing a migratory life history, reproductive overlap with an invasive species) might interact with physical changes to the landscape (i.e., barrier removal in streams) to achieve desirable outcomes for populations of conservation concern.
Integration with the overall social-ecological system can happen in four ways. First, research on the social components, such as collecting stakeholder information, can be used to inform the IBM of the ecological system, such as determining parameter space [105, 106]. For example, integrating qualitative stakeholder interview data into IBM parameterization can capture the richness of individual decision-making and complex social-ecological dynamics [116, 117]. This allows managers and researchers to understand if model outputs are responsive to conservation needs to inform conservation implications and decisions [105, 106]. Second, IBMs (or more often referenced as ABMs) of the social system can occur to inform the ecological system patterns or responses to management decisions [118]. The third option is to complete IBMs of both the social and ecological components in tandem. These are relatively rare (but see Rammer and Seidl [119] for an example of forest management and succession dynamics) but may be the area for the greatest expansion and future development. Last, IBMs of the ecological system could be done in tandem with non-ABM models of human managers. In the future, developing stakeholder-informed, social-ecological IBMs across all social-ecological system components, including humans, will provide a useful tool for conservation management.
Balancing System Complexity, Development, and Interpretation
Ecologists expect to find principles rather than mere rules of thumb in their study systems, even if these principles are not as concrete as those found in other fields, such as thermodynamics [14]. There has been a long history of specifically pointing to IBMs as one means to illuminate a unifying theory of ecology [120, 121]. Complexity and limited generality are often quoted as the main limitations of individual-based models [122]. This may be exacerbated by increasing knowledge of a system and the potential parameters and their values. The exploration of increasing parameter numbers and space will continue to grow with computational power but may result in increased amounts of time in exploratory steps. For example, the complexity of inheritance processes and models of landscape change may make generality more difficult as research and knowledge growth continues. However, in a modeled system, researchers have explicit knowledge of the rules, as opposed to natural systems. Because IBMs may consist of several thousand lines of code, it can often be just as complicated to understand the model as it is to understand the real system. In general, as the system grows in complexity, the number of parameters that must be estimated also increases. Parameter estimation introduces uncertainty into the system due to experimental error, differences in experimental assays, or error from data fitting techniques. Moreover, when the number of agents is large and their interactions numerous and complicated, it can become difficult to extract or isolate the key processes responsible for a given outcome [123]. IBMs may not always be rigorously described or mathematically represented as equations. Instead, IBM descriptions may be somewhat “fuzzy,” defined by a series of algorithms, and many descriptions of IBMs fail to state their assumptions in depth, making the comparison of models even more difficult [4, 124]. One additional consideration of any model must be how well it can account for data from natural systems [14] and the process of verifying, calibrating, and validating. This can be challenging with the complexity of IBMs.
These challenges related to complexity, development, and interpretation do not have a single solution. The solutions may also be systems specific and difficult to generalize. For example, a system and IBM focused on population structure may have unique complexity, development, and interpretation challenges when compared to an IBM focused on environmental change and evolution. Some projects may look to unify these two research lines, which would add another layer to work through to address the challenges.
Despite these challenges, IBMs still present many advantages. As discussed previously, IBMs may allow researchers to answer novel questions not possible with other modeling approaches, such as when individual mate choice may be important. In addition, they may be more appropriate to address questions and management goals when individual-level processes and the diversity of traits among individuals are important and are likely the drivers of the population- and landscape-level patterns. For example, it may be better to use an IBM approach to understand population dynamics if individual movement may be essential as opposed to modeling in a classic metapopulation approach.
Recommendations
Successful implementation of IBMs to inform specific management decisions and broader adaptive management strategies is easier with early engagement between modelers and managers (Fig. 2). By reaching out to modelers to co-develop potential research questions, the selection process of software, the gathering of existing data, and the planning for new empirical data generation, projects are more likely to be successful and efficient. Implementing IBMs can be challenging, and for managers to use software without guidance from experienced modelers, modelers will need to improve the usability of software and modeling code. One exciting development for those interested in IBMs is the growing availability of generalized modeling software which can facilitate addressing many eco-evo questions, as well as opportunities for researchers to write their own modeling software (Table 1). Out-of-the-box software reduces the barrier to entry to the field and allows researchers to run simulations of their own systems without the need to develop novel model code. In addition, standards of reporting, validating, and evaluating models continue to be developed and help standardize the field and improve accessibility [125••]. It is also important for modelers to create user interfaces and make it easier for managers to implement the software they have developed.
Conclusions
Eco-evo IBMs provide an opportunity to explore the complexities of natural systems and apply ecological and evolutionary processes across systems. While this review has focused primarily on aquatic systems, it is important to note that these approaches are applicable across a diversity of systems. We hope that by providing a conceptual framework, citing a wide variety of examples, and highlighting future challenges and opportunities, researchers interested in questions focused on eco-evo processes will find this paper as a building block from which to develop system-specific IBMs. In addition, although this review has focused on accessibility for practitioners and stakeholders, it is also important that IBM researchers and academics make their models approachable and practical for managers for continued and improved application of IBMs to happen. The complex nature of many of these processes and the complexity of the data were often prohibitive in the past. Because of this, modeling efforts are occurring in an exciting time where IBMs have never been more accessible across research questions and disciplines.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
DeAngelis DL, Mooij WM. Individual-based modeling of ecological and evolutionary processes. Annu Rev Ecol Evol Syst. 2005;36(1):147–68.
Knight AT, Cowling RM, Rouget M, Balmford A, Lombard AT, Campbell BM. Knowing but not doing: selecting priority conservation areas and the research–implementation gap. Conserv Biol. 2008;22(3):610–7.
Grimm V, Railsback SF. Individual-based modeling and ecology. Individual-based Modeling and Ecology: Princeton University Press; 2005.
DeAngelis DL. Individual-based models and approaches in ecology: populations, communities and ecosystems. CRC Press. 2018;663.
Bithell M, Brasington J, Richards K. Discrete-element, individual-based and agent-based models: tools for interdisciplinary enquiry in geography? Geoforum. 2008;39(2):625–42.
Funtowicz S, Ravetz JR. Emergent complex systems. Futures. 1994;26(6):568–82.
Day CC, McCann NP, Zollner PA, Gilbert JH, MacFarland DM. Temporal plasticity in habitat selection criteria explains patterns of animal dispersal. Behav Ecol. 2019;30(2):528–40.
Railsback SF, Harvey BC. Analysis of habitat-selection rules using an individual-based model. Ecology. 2002;83(7):1817–30.
Pitt WC, Box PW, Knowlton Frederick F. An individual-based model of canid populations: modelling territoriality and social structure. Ecol Model. 2003;166(1):109–21.
Clark ME, Rose KA, Levine DA, Hargrove WW. Predicting climate change effects on Appalachian trout: combining GIS and individual-based modeling. Ecol Appl. 2001;11(1):161–78.
Romero-Mujalli D, Jeltsch F, Tiedemann R. Individual-based modeling of eco-evolutionary dynamics: state of the art and future directions. Reg Environ Change. 2019;19(1):1–12.
Seaborn T, Andrews KR, Applestein CV, Breech TM, Garrett MJ, Zaiats A, et al. Integrating genomics in population models to forecast translocation success. Restor Ecol. 2021;29(4): e13395.
Xuereb A, Rougemont Q, Tiffin P, Xue H, Phifer-Rixey M. Individual-based eco-evolutionary models for understanding adaptation in changing seas. Proc R Soc B Biol Sci. 2021;288(1962):20212006.
Judson OP. The rise of the individual-based model in ecology. Trends Ecol Evol. 1994;9(1):9–14.
Popovics G, Monostori L. An approach to determine simulation model complexity. Procedia CIRP. 2016;1(52):257–61.
Lamarins A, Fririon V, Folio D, Vernier C, Daupagne L, Labonne J, et al. Importance of interindividual interactions in eco-evolutionary population dynamics: the rise of demo-genetic agent-based models. Evol Appl. 2022;15(12):1988–2001.
Pierson JC, Beissinger SR, Bragg JG, Coates DJ, Oostermeijer JGB, Sunnucks P, et al. Incorporating evolutionary processes into population viability models. Conserv Biol. 2015;29(3):755–64.
Nathan LR, Mamoozadeh N, Tumas HR, Gunselman S, Klass K, Metcalfe A, et al. A spatially-explicit, individual-based demogenetic simulation framework for evaluating hybridization dynamics. Ecol Model. 2019;1(401):40–51.
Messager ML, Olden JD. Individual-based models forecast the spread and inform the management of an emerging riverine invader. Divers Distrib. 2018;24(12):1816–29.
Araya-Donoso R, Baty SM, Alonso-Alonso P, Sanín MJ, Wilder BT, Munguia-Vega A, et al. Implications of barrier ephemerality in geogenomic research. J Biogeogr. 2022;49(11):2050–63.
DeAngelis DL, Grimm V. Individual-based models in ecology after four decades. F1000prime Rep. 2014;6.
Almeida LZ, Sesterhenn TM, Rucinski DK, Höök TO. Nutrient loading effects on fish habitat quality: trade-offs between enhanced production and hypoxia in Lake Erie. North America Freshw Biol. 2022;67(5):784–800.
Bonabeau E. Agent-based modeling: methods and techniques for simulating human systems. PNAS. 2002;99:7280–7.
Botkin DB, Janak JF, Wallis JR. Some ecological consequences of a computer model of forest growth. J Ecol. 1972;60:849–72.
Radchuk V, Kramer-Schadt S, Grimm V. Transferability of mechanistic ecological models is about emergence. Trends Ecol Evol. 2019;34(6):487–8.
DeAngelis DL, Yurek S. Spatially explicit modeling in ecology: a review. Ecosystems. 2017;20(2):284–300.
Day CC, Landguth EL, Bearlin A, Holden ZA, Whiteley AR. Using simulation modeling to inform management of invasive species: a case study of eastern brook trout suppression and eradication. Biol Conserv. 2018;1(221):10–22.
Schill DJ, Heindel JA, Campbell MR, Meyer KA, Mamer ERJM. Production of a YY male brook trout broodstock for potential eradication of undesired brook trout populations. North Am J Aquac. 2016;78(1):72–83.
Day CC, Landguth EL, Simmons RK, Baker WP, Whiteley AR, Lukacs PM, et al. Simulating effects of fitness and dispersal on the use of Trojan sex chromosomes for the management of invasive species. J Appl Ecol. 2020;57(7):1413–25.
Day CC, Landguth EL, Simmons RK, Baker WP, Whiteley AR, Lukacs PM, et al. Evaluation of management factors affecting the relative success of a brook trout eradication program using YY male fish and electrofishing suppression. Can J Fish Aquat Sci. 2021;78(8):1109–19.
Dunlop ES, Enberg K, Jørgensen C, Heino M. Toward Darwinian fisheries management. Evol Appl. 2009;2(3):245–59.
Hard JJ, Gross MR, Heino M, Hilborn R, Kope RG, Law R, et al. Evolutionary consequences of fishing and their implications for salmon. Evol Appl. 2008;1(2):388–408.
Conover DO, Munch SB. Sustaining fisheries yields over evolutionary time scales. Science. 2002;297(5578):94–6.
Pinsky ML, Eikeset AM, Helmerson C, Bradbury IR, Bentzen P, Morris C, et al. Genomic stability through time despite decades of exploitation in cod on both sides of the Atlantic. Proc Natl Acad Sci. 2021;118(15): e2025453118.
Wang HY, Höök TO, Ebener MP, Mohr LC, Schneeberger PJ. Spatial and temporal variation of maturation schedules of lake whitefish (Coregonus clupeaformis) in the Great Lakes. Can J Fish Aquat Sci. 2008;65(10):2157–69.
Laikre L, Ryman N. Effects on intraspecific biodiversity from harvesting and enhancing natural populations. Ambio. 1996;25(8):504–9.
Stokes TK, McGlade JM, Law R. The exploitation of evolving resources: proceedings of an international conference, held at Jülich, Germany, September 3–5, 1991. Springer Science & Business Media. 2013;270.
Dunlop ES, Feiner ZS, Höök TO. Potential for fisheries-induced evolution in the Laurentian Great Lakes. J Gt Lakes Res. 2018;44(4):735–47.
Olsen EM, Heino M, Lilly GR, Morgan MJ, Brattey J, Ernande B, et al. Maturation trends indicative of rapid evolution preceded the collapse of northern cod. Nature. 2004;428(6986):932–5.
Hsieh CH, Reiss CS, Hunter JR, Beddington JR, May RM, Sugihara G. Fishing elevates variability in the abundance of exploited species. Nature. 2006;443(7113):859–62.
Feiner ZS, Chong SC, Knight CT, Lauer TE, Thomas MV, Tyson JT, et al. Rapidly shifting maturation schedules following reduced commercial harvest in a freshwater fish. Evol Appl. 2015;8(7):724–37.
Dunlop ES, Heino M, Dieckmann U. Eco-genetic modeling of contemporary life-history evolution. Ecol Appl. 2009;19(7):1815–34.
Dunlop ES, Shuter BJ, Dieckmann U. Demographic and evolutionary consequences of selective mortality: predictions from an eco-genetic model for smallmouth bass. Trans Am Fish Soc. 2007;136(3):749–65.
Wang HY, Höök TO. Eco-genetic model to explore fishing-induced ecological and evolutionary effects on growth and maturation schedules. Evol Appl. 2009;2(3):438–55.
Dunlop ES, Eikeset AM, Stenseth NC. From genes to populations: how fisheries-induced evolution alters stock productivity. Ecol Appl. 2015;25(7):1860–8.
Enberg K, Jørgensen C, Dunlop ES, Heino M, Dieckmann U. Implications of fisheries-induced evolution for stock rebuilding and recovery. Evol Appl. 2009;2(3):394–414.
Eikeset AM, Richter A, Dunlop ES, Dieckmann U, Stenseth NChr. Economic repercussions of fisheries-induced evolution. Proc Natl Acad Sci. 2013 Jul 23;110(30):12259–64.
• Ayllón D, Nicola GG, Elvira B, Almodóvar A. Climate change will render size-selective harvest of cold-water fish species unsustainable in Mediterranean freshwaters. J Appl Ecol. 2021;58(3):562–75. This paper highlights using individual-based models for understanding body size evolution.
Shugart HH, Smith TM, Post WM. The potential for application of individual-based simulation models for assessing the effects of global change. Annu Rev Ecol Syst. 1992;23(1):15–38.
Rantanen M, Karpechko AY, Lipponen A, Nordling K, Hyvärinen O, Ruosteenoja K, et al. The Arctic has warmed nearly four times faster than the globe since 1979. Commun Earth Environ. 2022;3(1):1–10.
Norberg J, Urban MC, Vellend M, Klausmeier CA, Loeuille N. Eco-evolutionary responses of biodiversity to climate change. Nat Clim Change. 2012;2(10):747–51.
Beever EA, O’Leary J, Mengelt C, West JM, Julius S, Green N, et al. Improving conservation outcomes with a new paradigm for understanding species’ fundamental and realized adaptive capacity. Conserv Lett. 2016;9(2):131–7.
Seaborn T, Griffith D, Kliskey A, Caudill CC. Building a bridge between adaptive capacity and adaptive potential to understand responses to environmental change. Glob Change Biol. 2021;27(12):2656–68.
Forester BR, Murphy M, Mellison C, Petersen J, Pilliod DS, Van Horne R, et al. Genomics-informed delineation of conservation units in a desert amphibian. Mol Ecol. 2022;31(20):5249–69.
DeMarche ML, Doak DF, Morris WF. Incorporating local adaptation into forecasts of species’ distribution and abundance under climate change. Glob Change Biol. 2019;25(3):775–93.
Murdoch A, Yip DA, Cooke SJ, Mantyka-Pringle C. Evidence for the combined impacts of climate and landscape change on freshwater biodiversity in real-world environments: state of knowledge, research gaps and field study design recommendations. Curr Landsc Ecol Rep. 2022;7(4):68–82.
Scheiner SM, Barfield M, Holt RD. The genetics of phenotypic plasticity. XVII. Response to climate change. Evol Appl. 2020;13(2):388–99.
Sasaki MC, Dam HG. Integrating patterns of thermal tolerance and phenotypic plasticity with population genetics to improve understanding of vulnerability to warming in a widespread copepod. Glob Change Biol. 2019;25(12):4147–64.
Carja O, Plotkin JB. The evolutionary advantage of heritable phenotypic heterogeneity. Sci Rep. 2017;7(1):5090.
Bay RA, Rose NH, Logan CA, Palumbi SR. Genomic models predict successful coral adaptation if future ocean warming rates are reduced. Sci Adv. 2017;3(11): e1701413.
Kubisch A, Degen T, Hovestadt T, Poethke HJ. Predicting range shifts under global change: the balance between local adaptation and dispersal. Ecography. 2013;36(8):873–82.
Aitken SN, Whitlock MC. Assisted gene flow to facilitate local adaptation to climate change. Annu Rev Ecol Evol Syst. 2013;44(1):367–88.
Chen Z, Grossfurthner L, Loxterman JL, Masingale J, Richardson BA, Seaborn T, et al. Applying genomics in assisted migration under climate change: framework, empirical applications, and case studies. Evol Appl. 2022;15(1):3–21.
Rykiel EJ Jr. Testing ecological models: the meaning of validation. Ecol Model. 1996;90(3):229–44.
Augusiak J, Van den Brink PJ, Grimm V. Merging validation and evaluation of ecological models to ‘evaludation’: a review of terminology and a practical approach. Ecol Model. 2014;24(280):117–28.
Hamby DM. A review of techniques for parameter sensitivity analysis of environmental models. Environ Monit Assess. 1994;32(2):135–54.
Cariboni J, Gatelli D, Liska R, Saltelli A. The role of sensitivity analysis in ecological modelling. Ecol Model. 2007;203(1):167–82.
Railsback SF, Harvey BC, Kupferberg SJ, Lang MM, McBain S, Welsh HH. Modeling potential river management conflicts between frogs and salmonids. Can J Fish Aquat Sci. 2016;73(5):773–84.
Beaudouin R, Monod G, Ginot V. Selecting parameters for calibration via sensitivity analysis: an individual-based model of mosquitofish population dynamics. Ecol Model. 2008;218(1):29–48.
Rubin IN, Ellner SP, Kessler A, Morrell KA. Informed herbivore movement and interplant communication determine the effects of induced resistance in an individual-based model. J Anim Ecol. 2015;84(5):1273–85.
Breckling B, Middelhoff U, Reuter H. Individual-based models as tools for ecological theory and application: understanding the emergence of organisational properties in ecological systems. Ecol Model. 2006;194(1):102–13.
Trapp SE, Day CC, Flaherty EA, Zollner PA, Smith WP. Modeling impacts of landscape connectivity on dispersal movements of northern flying squirrels (Glaucomys sabrinus griseifrons). Ecol Model. 2019;24(394):44–52.
Lindenschmidt KE, Fleischbein K, Baborowski M. Structural uncertainty in a river water quality modelling system. Ecol Model. 2004;204(3–4):289–300.
Baggio RA, Araujo SBL, Ayllón D, Boeger WA. Dams cause genetic homogenization in populations of fish that present homing behavior: evidence from a demogenetic individual-based model. Ecol Model. 2018;24(384):209–20.
Grimm V, Ayllón D, Railsback SF. Next-generation individual-based models integrate biodiversity and ecosystems: yes we can, and yes we must. Ecosystems. 2017;20(2):229–36.
Balkenhol N, Cushman SA, Waits LP, Storfer A. Current status, future opportunities, and remaining challenges in landscape genetics. In: Landscape genetics. John Wiley & Sons, Ltd; 2015 p. 247–56. Available from: https://onlinelibrary.wiley.com/doi/abs/10.1002/9781118525258.ch14.
Keller D, Holderegger R, van Strien MJ, Bolliger J. How to make landscape genetics beneficial for conservation management? Conserv Genet. 2015;16(3):503–12.
Day CC, Landguth EL, Simmons RK. CDMetaPOP 2: a multispecies, eco-evolutionary simulation framework for landscape genetics and connectivity. Ecography. 2023;e06566.
Hollowed AB, Bax N, Beamish R, Collie J, Fogarty M, Livingston P, et al. Are multispecies models an improvement on single-species models for measuring fishing impacts on marine ecosystems? ICES J Mar Sci. 2000;57(3):707–19.
Latour RJ, Brush MJ, Bonzek CF. Toward ecosystem-based fisheries management. Fisheries. 2003;28(9):10–22.
van Nes EH, Lammens EHRR, Scheffer M. PISCATOR, an individual-based model to analyze the dynamics of lake fish communities. Ecol Model. 2002;152(2):261–78.
Shin YJ, Cury P. Exploring fish community dynamics through size-dependent trophic interactions using a spatialized individual-based model. Aquat Living Resour. 2001;14(2):65–80.
Campbell MD, Rose K, Boswell K, Cowan J. Individual-based modeling of an artificial reef fish community: effects of habitat quantity and degree of refuge. Ecol Model. 2011;222(23):3895–909.
Giacomini HC, De Marco P, Petrere M. Exploring community assembly through an individual-based model for trophic interactions. Ecol Model. 2009;220(1):23–39.
Rose KA, Curchitser E, Fiechter J, Hedstrom K, Haynie A, Bernal M, et al. Simulating the effects of zooplankton food web dynamics on the sardine fishery using an individual-based multi-species model of the California Current.
Baveco H, Focks A, Laender FD. Individual-based simulation models of multi-species systems under the impact of chemicals: complex dynamics can lead to.
Sable SE, Rose KA. A comparison of individual-based and matrix projection models for simulating yellow perch population dynamics in Oneida Lake, New York, USA. Ecol Model. 2008;215(1):105–21.
Hohenlohe PA, Funk WC, Rajora OP. Population genomics for wildlife conservation and management. Mol Ecol. 2021;30(1):62–82.
Carreras C, Ordóñez V, Zane L, Kruschel C, Nasto I, Macpherson E, et al. Population genomics of an endemic Mediterranean fish: differentiation by fine scale dispersal and adaptation. Sci Rep. 2017;7(1):43417.
Wu Y, Zhou Z, Pan Y, Zhao J, Bai H, Chen B, et al. GWAS identified candidate variants and genes associated with acute heat tolerance of large yellow croaker. Aquaculture. 2021;15(540): 736696.
Yoshida GM, Yáñez JM. Increased accuracy of genomic predictions for growth under chronic thermal stress in rainbow trout by prioritizing variants from GWAS using imputed sequence data. Evol Appl. 2022;15(4):537–52.
Frank BM, Baret PV. Simulating brown trout demogenetics in a river/nursery brook system: the individual-based model DemGenTrout. Ecol Model. 2013;10(248):184–202.
Seaborn T, Goldberg CS. Integrating genetics and metapopulation viability analysis to inform translocation efforts for the last northern leopard frog population in Washington State, USA. J Herpetol. 2020;54(4):465–75.
Benestan LM, Ferchaud AL, Hohenlohe PA, Garner BA, Naylor GJP, Baums IB, et al. Conservation genomics of natural and managed populations: building a conceptual and practical framework. Mol Ecol. 2016;25(13):2967–77.
Pertoldi C, Topping C. Impact assessment predicted by means of genetic agent-based modeling. Crit Rev Toxicol. 2004;34(6):487–98.
Moustakas A, Evans MR. Integrating evolution into ecological modelling: accommodating phenotypic changes in agent based models. PLoS One. 2013;8(8): e71125.
Pracana R, Burns R, Hammond RL, Haller BC, Wurm Y. Individual-based modeling of genome evolution in haplodiploid organisms. Genome Biol Evol. 2022;14(5):evac062.
Ma Y, Wu H, Wang L, Huang B, Ranjan R, Zomaya A, et al. Remote sensing big data computing: challenges and opportunities. Future Gener Comput Syst. 2015;1(51):47–60.
Yang C, Clarke K, Shekhar S, Tao CV. Big spatiotemporal data analytics: a research and innovation frontier. Int J Geogr Inf Sci. 2020;34(6):1075–88.
• Oomen RA, Kuparinen A, Hutchings JA. Consequences of single-locus and tightly linked genomic architectures for evolutionary responses to environmental change. J Hered. 2020;111(4):319–32. This paper includes evolution in projections of climate change.
Haefner JW. Parallel computers and individual-based models: an overview. In: Individual-based models and approaches in ecology. Chapman and Hall/CRC; 1992.
Yao X, Li G, Xia J, Ben J, Cao Q, Zhao L, et al. Enabling the big Earth observation data via cloud computing and DGGS: opportunities and challenges. Remote Sens. 2020;12(1):62.
Heppenstall A, Crooks A, Malleson N, Manley E, Ge J, Batty M. Future Developments in Geographical Agent-Based Models: Challenges and Opportunities. Geogr Anal. 2021;53(1):76–91.
• Wood ZT, Palkovacs EP, Olsen BJ, Kinnison MT. The importance of eco-evolutionary potential in the Anthropocene. Bioscience. 2021;71(8):805–19. This review paper covers eco-evolutionary feedbacks, which may be one of the areas best explored with individual-based models.
Samson E, Hirsch PE, Palmer SCF, Behrens JW, Brodin T, Travis JMJ. Early engagement of stakeholders with individual-based modeling can inform research for improving invasive species management: the round goby as a case study. Front Ecol Evol. 2017;5.
Wood KA, Stillman RA, Goss-Custard JD. Co-creation of individual-based models by practitioners and modellers to inform environmental decision-making. J Appl Ecol. 2015;52(4):810–5.
Walters CJ. Adaptive management of renewable resources [Internet]. Basingstoke: Macmillan Publishers Ltd; 1986. 384 p. Available from: https://iiasa.dev.local/.
Walters CJ, Holling CS. Large-scale management experiments and learning by doing. Ecology. 1990;71(6):2060–8.
Williams BK. Adaptive management of natural resources—framework and issues. J Environ Manage. 2011;92(5):1346–53.
Gregory R, Failing L, Harstone M, Long G, McDaniels T, Ohlson D. Structured decision making: a practical guide to environmental management choices. John Wiley & Sons; 2012.
Bach A, Minderman J, Bunnefeld N, Mill AC, Duthie AB. Intervene or wait? A model evaluating the timing of intervention in conservation conflicts adaptive management under uncertainty. Ecol Soc. 2022; Available from: https://eprints.ncl.ac.uk
Railsback SF, Gard M, Harvey BC, White JL, Zimmerman JKH. Contrast of degraded and restored stream habitat using an individual-based salmon model. North Am J Fish Manag. 2013;33(2):384–99.
Bart J. Acceptance criteria for using individual-based models to make management decisions. Ecol Appl. 1995;5(2):411–20.
Ostrom E. A general framework for analyzing sustainability of social-ecological systems. Science. 2009;325(5939):419–22.
Synes NW, Brown C, Palmer SCF, Bocedi G, Osborne PE, Watts K, et al. Coupled land use and ecological models reveal emergence and feedbacks in socio-ecological systems. Ecography. 2019;42(4):814–25.
Jossie E, Seaborn T, Baxter CV, Morey B. Using social-ecological models to explore stream connectivity outcomes for stakeholders and Yellowstone cutthroat trout. Ecol Appl. 2023;In Review.
Elsawah S, Guillaume JHA, Filatova T, Rook J, Jakeman AJ. A methodology for eliciting, representing, and analysing stakeholder knowledge for decision making on complex socio-ecological systems: from cognitive maps to agent-based models. J Environ Manage. 2015;15(151):500–16.
Lindkvist E, Wijermans N, Daw TM, Gonzalez-Mon B, Giron-Nava A, Johnson AF, et al. Navigating complexities: agent-based modeling to support research, governance, and management in small-scale fisheries. Front Mar Sci. 202;6.
Rammer W, Seidl R. Coupling human and natural systems: simulating adaptive management agents in dynamically changing forest landscapes. Glob Environ Change. 2015;1(35):475–85.
Huston M, DeAngelis D, Post W. New computer models unify ecological theory: computer simulations show that many ecological patterns can be explained by interactions among individual organisms. Bioscience. 1988;38(10):682–91.
Grimm V. Ten years of individual-based modelling in ecology: what have we learned and what could we learn in the future? Ecol Model. 1999;115(2):129–48.
Uchmański J, Grimm V. Individual-based modelling in ecology: what makes the difference? Trends Ecol Evol. 1996;11(10):437–41.
Bauer AL, Beauchemin CAA, Perelson AS. Agent-based modeling of host–pathogen systems: the successes and challenges. Inf Sci. 2009;179(10):1379–89.
Robertson D, editor. Eco-logic: logic-based approaches to ecological modelling. Cambridge, Mass: MIT Press; 1991. 243 p. (Logic programming).
•• Grimm V, Railsback SF, Vincenot CE, Berger U, Gallagher C, Deangelis DL, et al. The ODD protocol for describing agent-based and other simulation models: a second update to improve clarity, replication, and structural realism. J Artif Soc Soc Simul. 2020;23(2). Available from: http://eprints.bournemouth.ac.uk/33918/. This paper summarizes key steps to reproducible modeling research.
Macdonald EA, Cushman SA, Landguth EL, Hearn AJ, Malhi Y, Macdonald DW. Simulating impacts of rapid forest loss on population size, connectivity and genetic diversity of Sunda clouded leopards (Neofelis diardi) in Borneo. PLoS One. 2018;13(9): e0196974.
Mims MC, Day CC, Burkhart JJ, Fuller MR, Hinkle J, Bearlin A, et al. Simulating demography, genetics, and spatially explicit processes to inform reintroduction of a threatened char. Ecosphere. 2019;10(2): e02589.
Brauer CJ, Beheregaray LB. Recent and rapid anthropogenic habitat fragmentation increases extinction risk for freshwater biodiversity. Evol Appl. 2020;13(10):2857–69.
Galloway J, Cresko WA, Ralph P. A few stickleback suffice for the transport of alleles to new lakes. G3-Genes Genom Genet. 2020;10(2):505–14.
• Dominguez Almela V, Palmer SCF, Andreou D, Gillingham PK, Travis JMJ, Britton JR. Predicting the outcomes of management strategies for controlling invasive river fishes using individual-based models. J Appl Ecol. 2021;58(11):2427–40. This paper highlights using individual-based models to understand invasive species under different management strategies.
Bocedi G, Palmer SCF, Pe’er G, Heikkinen RK, Matsinos YG, Watts K, et al. RangeShifter: a platform for modelling spatial eco-evolutionary dynamics and species’ responses to environmental changes. Methods Ecol Evol. 2014;5(4):388–96.
• Snyder MN, Schumaker NH, Dunham JB, Ebersole JL, Keefer ML, Halama J, et al. Tough places and safe spaces: can refuges save salmon from a warming climate? Ecosphere. 2022;13(11):e4265. This paper highlights addressing climate change with individual-based model that was custom built for the system.
Piou C, Prévost E. A demo-genetic individual-based model for Atlantic salmon populations: model structure, parameterization and sensitivity. Ecol Model. 2012;24(231):37–52.
Landguth EL, Muhlfeld CC, Luikart G. CDFISH: an individual-based, spatially-explicit, landscape genetics simulator for aquatic species in complex riverscapes. Conserv Genet Resour. 2012;4(1):133–6.
Landguth EL, Muhlfeld CC, Waples RS, Jones L, Lowe WH, Whited D, et al. Combining demographic and genetic factors to assess population vulnerability in stream species. Ecol Appl. 2014;24(6):1505–24.
Acknowledgements
We would like to thank the invitation to submit this review for consideration. In addition, Steve Railsback provided very helpful feedback prior to the peer review process. We would also like to thank anonymous reviewers.
Funding
This publication was made possible by the NSF Idaho and National EPSCoR Programs and by the National Science Foundation under award numbers OIA-1757324 and OIA-1826801.
Ethics declarations
Conflict of Interest
Travis Seaborn and Stephanie Galla received financial support from the National Science Foundation grants listed under funding. All other authors declare no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Seaborn, T., Day, C.C., Galla, S.J. et al. Individual-Based Models for Incorporating Landscape Processes in the Conservation and Management of Aquatic Systems. Curr Landscape Ecol Rep 8, 119–135 (2023). https://doi.org/10.1007/s40823-023-00089-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40823-023-00089-8