Abstract
Purpose of review
In this review, we describe recent research on: (1) the occurrence of antibiotic resistant genes (ARGs) in waste materials and in soils receiving land application of ARG-contaminated waste materials; (2) the environmental behavior of ARGs in soils; (3) approaches to waste material management in terms of the elimination and reduction of ARGs. Land application of organic waste materials provides valuable nutrients to plant growth, but it can also introduce emerging organic contaminants, for example, antibiotics, into the environment. The widespread use of antibiotics and their entry into the environment are believed to result in the development and proliferation of antibiotic-resistant bacteria and ARGs in humans and animals, as well as in the environment. For this reason, ARGs are classified as emerging organic contaminants.
Recent findings
Land application of manure, biosolids from wastewater treatment plants, and wastewater can potentially load elevated levels of antibiotics, antibiotic-resistant bacteria, and ARGs into soils. A host of environmental and biological factors may influence the environmental behavior of ARGs, including persistence and mobility, as well as their transmission among microorganisms in soil matrixes. Traditional on-site waste material management approaches may not be able to completely remove ARGs from the waste material before it is applied to the soils. However, recent research has suggested certain waste material land application management approaches may be able to further reduce the input of ARGs and mobility in the environment.
Summary
Knowledge of the environmental behavior of ARGs is essential for better understanding and predicting the development and spread of antibiotic resistance in the environment. Information on the ecological consequences of ARG contamination and the possible entry of ARGs into the human food chain is also needed for better risk assessment of land application of ARG-contaminated waste materials. There is an urgent need to develop effective management and land application approaches to minimize/eliminate the input and mobility of ARGs in the soil environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Land application of organic waste materials, such as manure and wastewater treatment plant (WWTP) biosolids, is a common practice in agriculture to provide additional nutrients to plants [1]. Wastewater irrigation is also a common practice to enrich soil nutrients and overcome water shortages in many countries. However, these land application practices can introduce undesirable pollutants into the environment, of which those commonly addressed are heavy metals, organic compounds, and salts [2]. These practices could also introduce a broad spectrum of emerging organic contaminants into the environment [3]. Emerging organic contaminants, also or better termed as contaminants of emerging concern, are defined by the U.S. Environmental Protection Agency as “chemicals and other substances that have no regulatory standard, have been recently “discovered” in natural streams, often because of improved analytical chemistry detection levels, and potentially cause deleterious effects in aquatic life at environmentally relevant concentrations” [4].
One environmentally significant class of emerging organic contaminants is antibiotics. Antibiotics are widely used for treating human illness and animal diseases and promoting animal growth, as well as for a multitude of agricultural purposes [5]. For example, about 80% of the more than 17 million kilograms of antibiotics sold yearly in the USA is used in 60–80% of food-producing animal production for therapeutic and sub-therapeutic purposes [6–8]. Up to 95% of the antibiotics administered to animals and humans can eventually be excreted in urine or feces as parent compounds or metabolites [9–11], resulting in their frequent detection at significant levels in manure from a variety of farm animals [12–19] and in wastewater and biosolids [20–24]. Research has demonstrated that the overuse and/or misuse of antibiotics can facilitate the development and proliferation of antibiotic resistance in the microbiome of animal and human digestive systems [25, 26], resulting in increased levels of antibiotic-resistant bacterial populations and elevated levels of antibiotic resistance genes (ARGs) in animal manure [27–31] and human waste [32]. Therefore, land application of animal manure, wastewater, and biosolids can be sources of antibiotics, antibiotic-resistant bacteria (ARB), and ARGs in the environment.
The concentrations of antibiotics in biosolids and biosolids-amended soils have been reported to reach levels of up to 58 mg kg−1 [dry weight (d.w.)] and 900 μg kg−1 (d.w.), respectively [3], and the concentrations of antibiotics in wastewater treatment plant influents and effluents to reach levels of up to 122 μg L−1 and 6 μg L−1, respectively [33]. In soils irrigated with wastewater, antibiotics at concentrations ranging from 1.4 to 66.7 μg kg−1 have been detected [34]. In manure and wastewater collected from a swine feedlot in China, the highest total tetracycline and total sulfonamide concentrations were 167 mg kg−1 and 65 μg kg−1, respectively, in the manure and 389 and 7.6 μg L−1, respectively, in the wastewater [35]. In another study, 17 antibiotics were detected in approximately 39% of the 219 dried manure-based fertilizer samples tested, with the mean concentrations ranging from 20 μg kg−1 to 2 mg kg−1; one sample contained the highest concentration of antibiotic, namely, 73 mg kg−1 doxycycline [23]. The highest concentrations of tetracycline and chlortetracycline reported in pig manure-amended soils were 199 μg kg−1 and 7.3 μg kg−1, respectively [36].
Research has shown that elevated levels of antibiotics in the soil environment can potentially further enhance the populations of ARB and levels of ARGs in soils in addition to those that are already associated with the land-applied waste materials [5, 37–39]. Elevated ARGs in the environment have been linked to increasing occurrence of antibiotic resistance in human populations and reduced/compromised effectiveness of antibiotic therapy for humans [40, 41]. In the European Union alone, ARB are estimated to cause 25,000 deaths and cost more than U.S. $1.5 billion in healthcare expenses and productivity losses every year [42]. Antibiotic resistance has become a global concern and was recently identified as the “major health security challenge of the 21st century” by the World Health Organization [43, 44]. It should be noted that while most ARB-induced infections result from the overuse of antibiotics by humans in hospitals [45], studies have revealed that ARB have been isolated from humans and a wide range of animals not subject to significant antibiotic exposure [46], implying a role of environmental microbiomes in the rise of antibiotic resistance [47].
The development of antibiotic resistance among bacteria arises from ARG expression. ARGs are DNA sequences encoding proteins responsible for the resistance [48]. Because of their essential role in the development of antibiotic resistance, ARGs were recently included as an important class of emerging organic contaminants that can greatly impact environmental and human health [49]. Extensive reviews on the occurrence, environmental behaviors, and ecological and human health impacts of antibiotics and ARB in soils subject to land applications with waste materials have been published in recent years [5, 50]. However, similar systematic information on ARGs in the soil environment is lacking. Therefore, the main objective of this review is to synthesize the most recent literature on: (1) the occurrence and distribution of ARGs in land applications of waste materials; (2) the environmental behaviors of ARGs in soils receiving land applications of waste materials; (3) the effectiveness of current on-site management approaches in reducing ARGs in waste materials; (4)) current knowledge gaps on the factors contributing to the enrichment of ARGs in soils, the potential risk to ecological and human health, and the best approaches to land application management with waste materials.
Occurrence of ARGs in Land-Applied Waste Materials and the Affected Soils
Land-Applied Waste Materials
The reported occurrence and distribution of a diverse ARGs in a wide range of waste materials is summarized in Table 1. Culture-independent methods, such as quantitative PCR assays and metagenomics sequence analysis, have been commonly used to identify, detect, and quantify ARGs [72–74]. As shown in Table 1, the detected ARGs conferred resistance to all major classes of antibiotics and encompassed all of the main resistant mechanisms.
Tetracycline resistance (tet) genes have been found to have the highest diversity in a variety of waste materials, with a total of 19 reported tet genes that include 11 genes encoding efflux pumps, seven genes encoding ribosome protection, and one gene encoding enzymatic inactivation (Table 1). The most frequently reported types of ARGs are those which encode macrolide (e.g., ermB), sulfonamide (e.g., sul1), and tetracycline (e.g., tetA) resistance genes (Table 1). While absolute abundance (gene copies/unit matrix) has been used in many studies, total extracted DNA or copies of 16 s rRNA copies from sample matrices have been used in some studies to normalize the total number of ARG copies to be expressed as relative abundance [51, 69].
The absolute abundance of ARGs has been reported to range from 105 to 1011 gene copies g−1 (d.w.) and from 102 to 1011 copies mL−1 in solid waste materials and wastewater samples, respectively (Table 1). The absolute abundance of species–specific-ARG also varies largely case by case. For example, the sulfonamide resistance (sul) gene sul1 was the most abundant ARG gene detected in WWTP effluents in northern China, with an absolute abundance of 105 copies mL−1 [52]; however, its absolute abundance was four orders of magnitude lower than that detected in the WWTP effluents in eastern China (109 copies mL−1) [53]. The relative abundances of four tet genes in activated biosolids from a Hong Kong WWTP ranged from 100 to 104 gene copies ng−1 DNA [51]. The relative abundances of chloramphenicol resistance genes and two sul genes (sul1 and sul2) in wastewater from a number of pig, cattle, and chicken farms in China ranged from 10−3 to 10−2 and from 101 to 102 copies/16 s rRNA copies, respectively [54, 67]. Clearly, comparing studies with ARG results expressed as absolute abundance with those expressed as relative abundance is difficult without knowing the levels of total DNA or 16sRNA in a sample. Therefore, a standardized culture-independent ARG analysis is essential for meaningful comparisons of ARG abundance from different investigations.
The observed statistically significant positive correlations between antibiotics and resistance genes in different waste materials suggest that antibiotics exert a selection effect on resistant bacteria and the corresponding ARGs [35, 67, 68]. For example, the relative abundances (10−6 to 100 copies/16 s rRNA copies) of 14 tet and three sul genes in manure and wastewater samples from a swine facility were found to be positively correlated with the concentrations of the antibiotic [35].
Soils Amended With Waste Materials
Among all of the environmental compartments, soil is the major receiver and reservoir of ARB and ARGs. First, soil is a major reservoir of antibiotics naturally produced by the indigenous microorganisms [75]. Furthermore, extensive use of antibiotics can result in their eventual release into the soil environment due to land application of waste materials that often have elevated antibiotic concentrations [75]. Elevated antibiotic concentrations in soils can potentially create a selective pressure on both ARB and ARGs in those soils [75]. Land application of waste materials with elevated levels of ARB and ARGs may further enhance their absolute abundance in soils.
Recent research has provided ample evidence that land application of waste material can significantly elevate the abundance of ARGs in soils. For example, the relative abundances (10−6 to 10−3 copies/16 s rRNA copies) of six tet genes increased by one to two orders of magnitude in paddy surface soils (0–5 cm) with long-term (9–33 years) manure application compared to soil without manure application (10−7 to 10−3 copies/16 s rRNA copies) [76]. The high relative abundance of ARGs (10−3 to 10−1 copies/16 s rRNA copies) in the manure led to significantly higher relative abundance of ARGs in manure-amended soils compared to the control soils. The authors of this study also reported that the relative abundances of four tet genes (tetA, tetM, tetQ, tetW) positively correlated with the concentrations of tetracyclines in soils (R 2 ranged from 0.45 to 0.52, p < 0.01) , as well as with soil pH (R 2 ranged from 0.54 to 0.62, p < 0.05) and soil organic matter contents (R 2 ranged from 0.47 to 0.58, p < 0.01). The results of this study indicate that both antibiotics and soil properties play an important role in the selection of ARGs in soils [76]. As such they are consistent with those of another study in which the relative abundances of three tet genes, tetG, tetL, and tetB(P), were found to be one to two orders of magnitude higher in soils with 6-year-long land applications of fresh or composted swine manure than in the control soils without manure application (10−5 to 10−4 copies/16 s rRNA copies) [55]. In another study a total of 149 unique ARGs were detected in swine manure-amended soils, and the diversities of the ARGs were threefold higher in the manure-amended soils than in the control soils [77]. The abundances of the top 63 unique ARGs were significantly enriched in the manure-amended soils, at an overall median enrichment of 192-fold, compared to control soils [77]. In Finland, which has the fourth lowest use of veterinary antibiotics per production animal among the 25 European Union countries, only two- to sevenfold increases in the relative abundances of tetM, sul1, and blaOXA-58 were detected in soils receiving manure from animals treated with lower levels of antibiotics compared to those in the investigations discussed in the preceding text [64]. Antibiotics are only allowed in Finland for therapeutic purposes, so antibiotic concentrations in manure are low. Nevertheless, the results of this study demonstrate that although the enrichment of ARGs is much lower in soils amended with manure from animals which only received antibiotics for therapeutic purposes, reducing the use of antibiotics in animal production does not appear to completely prevent the ARG enrichment in the manure-amended soils [64].
Wastewater irrigation and the application of biosolids may also elevate the abundance of ARGs in soils. For example, the relative abundances of three sul and ten tet genes in long-term (>30years) wastewater-irrigated soils in China were one to two orders of magnitude higher than those in the non-irrigated soils (not determined to 10−3 copies/16 s rRNA copies) [78]. The authors of this study also found that the relative abundance of ARGs were positively correlated with the concentrations of oxytetracyclines, sulfadiazine, and sulfamethoxazole in the soils. These findings are consistent with those of another study which showed that the relative abundances of sul1 and sul2 in soils irrigated for 100 years with wastewater were one to two orders of magnitude higher than those in the non-irrigated soils (10−5 copies/16 s rRNA copies) [79]. In a laboratory study, the absolute abundance of sul2 increased by fivefold in soils following repeated irrigation (60 days) with secondary wastewater effluent compared to the control soils [56]. To the contrary, in a study examining soils with high absolute abundance of ARGs (108 to 109 gene copies g−1 soil) in the natural soil microbiome, irrigation with wastewater for 6–18 years did not result in a significant enrichment of relative abundance of ARGs compared to the soils irrigated with fresh water [80]. The authors of this latter study suggested that the ARB in wastewater were not able to compete with the abundant native soil microbiome or survive in the soil environment and, therefore, did not significantly contribute ARGs to the native soil bacteria.
A total of 108 unique ARGs were found to be significantly enriched in soils receiving applications of biosolids for >10 years compared to those which did not [81]. Among the 108 ARGs detected in this study, mexF [encoding FCA (fluoroquinolone, quinolone, florfenicol, chloramphenicol, and amphenicol) resistance] showed the highest enrichment (up to 3845-fold), while some unique ARGs, such as floR, aadE (encoding aminoglycoside resistance), pikR2 [encoding MSLB (macrolide–lincosamide–streptogramin B resistance) resistance], ermB, and ermD were enriched by more than 100-fold. In a different field study, the relative abundances of ten ARGs were found to be significantly elevated up to levels of 10−1 copies/16 s rRNA copies in the biosolids-treated plots compared to the control plots, in many of which the ARGs were below the detection limit (absolute abundance = 5 copies g−1 soil) [82]. Even 530 days after application of the biosolids, the relative abundance of one macrolide resistance gene (mphA) in the biosolids-applied soil was still detectable—at a level sixfold higher than that in the control plot. Because the relative abundance of gene targets was highly variable from year to year, the authors concluded that it was difficult to distinguish whether the enriched gene originated from the contents of the applied biosolids or from native soil bacteria [82].
Environmental Behaviors of ARGs in Soils Amended With Waste Materials
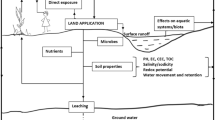
Antibiotic resistance genes, carried by either the bacterial chromosome or mobile genetic elements (e.g., plasmids), can be present either in the host cells of resistant bacteria or in the environmental media [83]. Free forms of ARGs can be released into environmental media from the host cells, either by the active secretion of resistant bacteria or by the rupture of dead cells [84]. Following their release into the soils, multiple factors, including soil physicochemical properties and biological conditions, could affect and/or determine the multitude of potential environmental behaviors of the ARGs, such as sorption/desorption, transformation, transport, and gene transmission. The possible fate of ARGs in soils receiving land application of waste materials is illustrated in Fig. 1.
Sorption and Persistence of ARGs Introduced to Soils Receiving Land Applications of Waste Materials
The sorption of DNA to soil can be significantly influenced by many factors, such as soil organic matter content, soil solution chemistry, and surface properties of the organic and inorganic components of the soil [85–87]. A batch experiment showed that the sorption of DNA (solid mass 10 mg, solution volume 1 mL, DNA initial concentration 100 μg mL−1) in a natural soil (andosols) gradually decreased from 100 to 25% when the pH of the soil slurry increased from 2 to 5 and that it was relatively constant at 20–25% between pH 5 and 9 [88]. Because the isoelectric point of DNA is around pH 5, the relative amount of negative charges on the DNA molecule would increase at higher pH, resulting in decreased sorption of the DNA onto the andosols. The authors of the same study also showed that DNA sorption to soil particles was 75% when the ionic strength (for MgCl2) was 640 mM at pH 6.7–7.1 [88]. One explanation for this effect is that the high levels of Mg2+ in the soil solution may have reduced the repulsive force between the negatively charged DNA molecule and the negative charges on soil particles by acting as bridging cations [85]. These results indicate that environmental relevant conditions, such as normal soil solution pH range (6–8) and soil solution ionic strength (<10 mM), are unfavorable for DNA sorption onto soil particles.
ARGs can persistent in soils for extended periods. A field study has shown that the relative abundances of sul1 and sul2 decreased one order of magnitude (from 10−6 to 10−5 and from 10−5 to 10−4 copies/16 s rRNA copies, respectively) in a soil during the first 10 days after swine manure application [89]. However, after 10 days, the relative abundance of sul1 (10−6 copies/16 s rRNA copies) remained at a stable level until 175 days, while that of sul2 further declined by another one order of magnitude (from 10−6 to 10−5 copies/16 s rRNA copies). These results are consistent with those in another field study in which the relative abundance increased by 40 and 58% for sul1 and by 67% and 107% for sul2, respectively, in loamy sand and silt loam that had been repeatedly amended with pig manure [90]. Similarly, the half-lives of five ARGs, namely, ermB, sul1, tetA, tetW, and tetX, ranged from 13 to 150 days and from 31 to 440 days in a biosolids-applied sandy loam and a silty loam, respectively [57]. Therefore, separate from the long-term application of ARG-containing waste materials that are able to continuously introduce ARGs into soils, the persistence of ARGs may also contribute to the accumulation of ARGs in soils.
Transmission of ARGs Among Microorganisms in Soils Receiving Waste Material Applications
There are two pathways of genetic transmission of ARGs: vertical inheritance and horizontal gene transfer [91]. In vertical inheritance (as known as vertical gene transfer), ARGs can be transmitted during the growth of resistant bacteria when they are passed on from parental resistant bacteria to its own progeny (offspring). In horizontal gene transfer (known as lateral gene transfer), susceptible bacteria may acquire ARGs via the intake and recombination of ARGs from other microorganisms that are already resistant or from the environmental media through three major pathways: conjugation, transformation and transduction [92]. The ARGs of parental bacteria may be the result of gene mutation (e.g., following exposure to antibiotics), horizontal gene transfer (from an unrelated resistant microorganism), or vertical inheritance (from its own parent).
Compared to the number of studies on gene transfer by vertical inheritance among microorganisms in waste materials-applied soils, relatively more studies have examined the possibility of horizontal gene transfer by monitoring the abundance of mobile genetic elements (e.g., transposases and introns) in soils [77, 81]. Since these mobile genetic elements play an important role in bacterial genetic exchange, their enrichment in soils may increase the likelihood of horizontal ARG transfer in soils. Land application of waste materials may elevate the abundance of mobile genetic elements in soils. For example, nine mobile genetic elements were enriched by as much as 1000-fold in ARG-polluted soils from farmland adjacent to a swine feedlot in China, and there was a positive correlation (R 2 = 0.97) between the relative abundance of tet genes and transposase genes [77]. These results suggest that manure application may accelerate the dissemination of ARGs in soils through horizontal gene transfer. Similarly, a positive correlation (R 2 = 0. 47, p < 0.01) between the relative abundance of mobile genetic elements and ARGs was observed in soils to which biosolids were applied for >10 years [81].
Extracellular DNA can be adsorbed by surface-reactive particles in soils and sediments while still retaining protection against degradation by nucleases and maintaining the capacity to transform competent bacterial cells [93]. Consequently, the extended persistence of ARGs via sorption can increase the incidence of potential horizontal gene transfer. The concentration of DNase required to inhibit transformation by bound DNA has been shown to be higher than that required to inhibit transformation by comparable amounts of free DNA [94]. In that same study, considerably more bound DNAase than free DNase was required to inhibit transformation by the same amount of free DNA [94].
Under certain selection pressures (e.g., treatment with antibiotics), ARB can survive while other bacteria are killed, resulting in the proportion of the population of resistant bacteria increasing in a microbial community [91, 95, 96]. As the resistant bacteria in this scenario would be able to pass their ARGs to subsequent generations via vertical inheritance, the selection pressure would lead to an elevated level of relative abundance of ARGs. Importantly, antibiotic exposure is not the only factor that can exert a selective pressure; selective pressure can also be exerted by other factors, such as heavy metals (e.g., As, Cu, Zn, Mn, Co, Ag, Hg, and Ni) and biocides. Co-selection of antibiotic and metal resistance has been widely reported [97–100]. The authors of one study reported that 17% of bacterial genomes (a total of 2522 genomes located on chromosomes or plasmids) carried both ARGs and biocide/metal resistance genes [101] and also suggested that heavy metal or biocides exposure has the potential to select ARB through chromosomal biocide/metal resistance genes. Some studies have reported higher concentrations of heavy metals and ARGs in land to which waste materials have been applied compared to those in control soils [77, 102]. For example, a positive correlation between Cu, Zn, and As and the relative abundance of total ARGs (R 2 ranged from 0.38 to 0.52, p <0.05) was observed in the manure-amended soils in one study [77]. A long-term (4–5 years) field experiment showed that Cu contamination could significantly affect the abundance, diversity, and mobility potential of a broad spectrum of ARGs in two contrasting agricultural soils (red and fluvo-aquic soil) [103].
The relative abundances of a β-lactamase resistance gene (blaCEP04) encoding AmpC increased in soils to which manure from animals not treated with antibiotics was applied [104]. The authors of this study observed that nutrients in the manure induced a bloom of Pseudomonas spp. which may be an important biological reservoir of β-lactamase resistance genes in the soil, leading them to highlight the role(s) of unexpected selection pressures (due to manure application) that could increase the abundance of resident ARB in soils and ARGs. On the other hand, the nutrients in manure may also promote the growth and production of β-lactam producers (e.g., a certain group of fungi, actinomycetes, unicellular bacteria) in soils, resulting in a selection pressure [104]. Unfortunately, this study did not examine the antibiotic concentrations in the manure and the manure-amended soil [104]. Therefore, the possibility that trace levels of antibiotics can be generated from the native microbiome of manure or manure-amended soils cannot be completely ruled out, even when the manure is from animals not treated with antibiotics.
Transport of ARGs Into the Aquatic Environment
Numerous studies have examined surface runoff losses and the leaching of contaminants, such as nitrate [105], phosphate [106], pesticides [107], and antibiotics [108], from agricultural soils amended with waste materials. For example, in one study the relative surface runoff losses of chlortetracycline, monensin, and tylosin were <5% of the total amount applied to the soil in manure [108], suggesting that the antibiotics were retained in the soil profile with the potential to exhibit continued pressure on the soil microbiome. Likewise, ARGs applied to agricultural fields through land application of waste materials can move into groundwater through leaching or into surface water via surface runoff. The transport behavior of ARGs can be mitigated by sorption, but under unfavorable sorption conditions, ARGs can be transported over a considerably long distance [109, 110]. For example, the transport of antibiotic resistance plasmids in quartz sand-packed columns was found to be significantly inhibited at high ionic strength (0.05 M for CaCl2), a condition which enhances ARG sorption [110]. The same study also showed that following a reduction in ionic strength, the resistance plasmids were desorbed, demonstrating that ARG sorption by soils is a reversible process. These results suggest that water irrigation and rainfall, which can reduce the ionic strength of the soil solution, can potentially enhance the transport of ARGs. In a rainfall simulation study [111], the absolute abundances of tetQ and ermB were one to two orders of magnitude higher in surface runoff from manure-amended plots than in runoff from the control plots. Thus, surface runoff can act as a nonpoint source of ARGs pollution in the environment.
The absolute abundances of tetO, tetX, and tetM in soil samples collected from farmlands adjacent to swine feedlots were detected at up to 107 copies g−1 soil at a depth of 60–80 cm [58]. In this study, the abundance of ARGs at this soil depth were one to two orders of magnitude lower than those in top soil (0–20 cm). The presence of ARGs in deep soil layers may result from vertical transport, but this study did not report the abundance of ARGs in the same layer of a soil without manure application, making it difficult to draw a definite conclusion on the potential for downward ARG movement along soil profiles [58]. To the contrary, another study showed that the absolute abundance of ARGs in both soils at a depth of 5–20 cm and in the receiving water of the stream immediately down gradient of a field amended with manure, sawdust, and straw mixtures did not increase compared to those before land application [112]. The authors suggested that ARG transport to the aquatic systems may depend on characteristics of the waste materials applied and the soils, as well as the application approach [112].
If ARGs reach the aquifer, the groundwater flow may further carry them to a larger area. One important consequence of elevated levels of ARGs in groundwater could be their impact on well water, which is a very important drinking water source in many countries. Up to 1.5 × 104 copies ng−1 DNA of tetZ and tetQ were detected in groundwater wells adjacent to a manure lagoon in a pig production farm, while no ARGs were detected in a background monitoring well [113]. Similarly, many communities also rely on surface water as drinking water sources. The authors of one study reported a positive correlation between the relative abundance of riverine sul1 and both the size of animal feeding operations upstream (R 2 = 0.35, p < 0.001) and the treatment capacity of wastewater treatment (R 2 = 0.34, p < 0.001); these results suggest that surface water may represent a key dissemination pathway of ARGs [114]. The levels of ARGs in South Platte River samples collected from sites impacted by animal feeding operations or/and waste water treatment plants were two to three orders of magnitude higher than those in samples from pristine sites [114].
Association of ARGs With Plants Grown in Soils Amended With Waste Materials
Plant surfaces provide a natural habitat for a variety of bacteria, yeast, and fungi as phyllosphere microorganisms [115]. Plants are also hosts to a variety of endophytic microorganisms [116]. Microorganisms associated with plants can be potential hosts of ARGs and through the consumption of ARB-contaminated plant products, humans can ultimately be exposed to these ARGs. Reports of elevated levels of ARGs in soils amended with waste materials have led to concern about the association between ARGs and the edible portions of plants and that this association will result in a potential negative impact on human health [117, 118]. One possible pathway leading to an association between plants and ARGs is through plant surface contamination. As plants are close to the ground, contamination may occur by soil resuspension, rain splash, and animal disturbance [119]. The other possible pathway is through plant root uptake. Research has shown that DNA fragments of 120–3000 nucleotides in length could be actively absorbed by Arabidopsis thaliana root cells [120]. If an ARG were to be carried by DNA fragments within this length range, it could potentially be taken up by plants. However, this pathway is only speculation and has to be further tested by research on plant uptake of ARGs. The passive uptake of antibiotics with water by plants grown in manure-amended soils and soils irrigated with wastewater has also been reported [121–123]. It is therefore possible that antibiotics taken up by plants may introduce a selective pressure on antibiotic-resistant endophytic bacteria and result in a yet further increased abundance of ARGs associated with plants. More studies are needed to investigate this notion.
The limited number of reports published to date suggests the possibility of an elevated association of ARGs with plants grown in soils with elevated levels of antibiotics and ARGs. In one comparative study, the relative abundances of sul1, tetG, tetC, tetA, and tetM detected in leaves of lettuce grown in soils with the higher levels of antibiotics (100–200% higher) were three to four levels of magnitude higher than those with the relatively lower levels of antibiotics; the abundance of ARGs in the former was also three to four orders of magnitude higher than that in the latter [124]. A separate study also detected three or four orders of magnitude higher relative abundances of ten ARGs (tetW, tetM, tetB, tetQ, sul1, sul2, ermA, ermB, catI, and catII) in the stem, leaf, fruit, and seed of bell peppers grown in cow manure-applied soils compared to soils with reduced levels of ARGs [125]. More investigation is needed to enable more definitive conclusions to be drawn on the impacts of the land application of waste materials on the association of ARGs with plants, the extent of ARGs entering the food chains, and the subsequent effects on human health.
ARGs in the Air Near Soils Amended with Waste Materials
Antibiotic resistance genes can also reach humans through the inhalation of aerated solids (aerosols) containing ARB or ARGs originating from contaminated soils after land application. For example, the absolute abundances of sul and blaSHV were higher (up to 750 and 870 copies m−3, respectively, in air samples) in air samples collected near three conventional farms than in those collected at an organic beef cattle farm [126]. However, the authors of this study did not compare the ARG abundance in air samples collected upwind and downwind. The same study reported that the bacteria isolated from the air samples near the three conventional farms showed more resistance to certain antibiotics compared to those collected at organic locations. A number of microorganisms can survive by forming resistant spores under harsh environmental conditions [127]. In addition, non-spore-forming species may enter into a viable but not culturable state. One study showed that antibiotic pressure can induce a viable but not culturable state in Staphylococcus aureus growing in biofilms [128]. These strategies allow the bacteria to survive for a much longer time, possibly enabling these ARB to be transported for long distances in a windblown environment. Thus, more studies are needed to gain a better understanding of this subpopulation of ARB and the fate of their corresponding ARGs.
On-site Waste Material Treatment for the Elimination/Reduction of ARGs
Before a waste material is applied to land, it is common practice to pre-treat it to destroy pathogens and remove other organic contaminants [129]. Since antibiotics can create a selective pressure on resistant bacteria and ARGs, management approaches that can reduce the release and spread of antibiotics in the environment may not be efficacious in terms of reducing the abundance of ARGs in the waste materials because the environmental behaviors of ARGs could be very different from that of antibiotics due to DNA being a macromolecule (biopolymers) and antibiotics being small molecules. In host cells, ARGs can amplify via gene replication that accompanies the proliferation of resistant bacteria, which might occur even in the absence of antibiotics, resulting in the hereditary and horizontal transfer of the ARGs to a greater population of bacteria. Efficient waste material management approaches for the removal of ARGs have yet to be identified.
Pretreatment of manure
Some manure management approaches can remove resistant bacteria. For example, multidrug-resistant bacteria were reduced by >90% by co-digestion of dairy manure and waste milk under anaerobic conditions [130]. According to the U.S. Food and Drug Administration’s (FDA) Food Safety Modernization Act, manure management is acceptable for land application if the numbers of pathogens (e.g., Salmonella sp. and Listeria sp.) are reduced to below standardized levels. Of all the manure management approaches, composting is a common practice to prevent and minimize pathogens in manure and may also reduce the levels of antibiotics [131]. However, since ARGs can be released from the cells upon rupturing, any reduction in the abundance of resistant bacteria during composting does not necessarily indicate a reduced abundance of ARGs in manure. The change in ARG abundance during composting has also been studied directly [65, 132]. In one of these studies [65], the relative abundances of tetM, tetO, tetQ, and tetW in swine manure decreased by 38.5 to 93.5% after 6 days of composting, and these decreases correlated with a reduced concentration of tetracycline; in contrast,, the relative abundances of sul1 and sul2 increased by approximately 100 and 350%, respectively, during the same period. These results indicate that proliferations of unique ARGs could occur during composting [65]. A separate study found that the relative abundances of five tet genes (tetA, tetC, tetG, tetL, and tetX) and two sul genes (sul1 and sul2) in swine manure significantly increased by one to two orders of magnitude from 10−5 to 10−3 copies/16 s rRNA copies to 10−3 to 10−1 copies/16 s rRNA copies after 32 days of composting [133]. Another study showed that of the 158 ARGs investigated, the abundances of 94 ARGs significantly reduced while 23 were significantly enriched during composting of a swine manure for 6 days [134]. The contradictions in outcomes regarding the changes in the abundance of ARGs during manure composting suggest that the efficiency of composting in the removal of ARGs prior to land application depends on the species of ARGs. Composting conditions such as pH and temperature in both the thermophilic and mesophilic stages affect the levels that antibiotics are reduced, and perhaps ARG levels also [135]. According to U.S. federal guidelines, two composting approaches, namely, static and tuned, are recommended. Therefore, assessments of U.S. FDA-approved manure management approaches are warranted.
Pretreatment of WWTP wastewater and biosolids
Wastewater or biosolids from WWTP are hotspots for ARGs [72], and the treatment of these hotspots is another opportunity to control ARGs before they are introduced into the environment [136]. Traditional treatments in WWTP are effective for removing resistant bacteria during the disinfection process using UV and chlorination treatments. However, the efficiency of the disinfection process to remove ARGs is variable. In one study, the absolute abundances of four erythromycin resistance genes (ermA, ermB, ereA, and ereB) and two tet genes (tetA and tetO) was reduced by three orders of magnitude (initial concentrations ranged from 100 to 102 gene copies mL−1) during UV treatment of wastewater, indicating the effectiveness of the UV treatment in reducing ARG abundance in municipal wastewater; in contrast, chlorination could not eliminate or reduce the levels of ARGs in the wastewater [137]. In another study, the absolute abundances of tetA, ereA, and ermB fell by 24, 87, and 40%, respectively, with chlorination treatment while there was no significant change in the absolute abundances of tetB, tetO, ereB, and ermA; in fact, even an increased dose of chlorination did not improve the reduction rate of ARGs [137]. A recent study at a drinking water treatment plant demonstrated that chlorination can even cause an enrichment of the total abundance of ARGs [139]. There are also some studies suggesting that disinfection processes (UV and chlorination) do not contribute to the reduction of ARGs given that no significant difference in ARGs abundance between pre- and post-disinfected wastewater samples was found, indicating that ARG removal from wastewater is system-specific [140, 141].
The effect of anaerobic digestion of wastewater is ARG-specific. For example, although anaerobic digestion reduced the absolute abundances of tetG, tetM, tetO, tetW, tetX, sul1, sul2, ereA, ermB, ermF, and mefA in swine wastewater by 37–95% [63], when ARGs were normalized to 16 s rRNA gene, the relative abundances of tetG, tetM, sul1, sul2, ereA were found to be enriched by 2–530%. The reason why the relative abundance of unique ARGs increased is yet to be identified. It may be that the ARB carrying these genes proliferated under selective pressure during anaerobic digestion or that these ARGs horizontally transferred to other species. Regarding anaerobic digestion of biosolids, the absolute abundances of tetA, tetW, sul1, sul2, and blaTEM decreased by 30–98% during anaerobic digestion [142]. In another study, Metagenomics analysis revealed that 21 of the ARGs in biosolids were removed at a >90% efficiency during anaerobic digestion. However, aadA, macB, and sul1 were found to be enriched by 26–33% during the thermophilic anaerobic digestion process, while tetM, sul1 and genes encoding ereA were enriched by 40–60% during the mesophilic anaerobic digestion process [143]. Whether anaerobic digestion could reduce the abundance of ARGs in wastewater and biosolids is still inconclusive, but the process appears to be species-specific. Further research is needed on this subject.
Knowledge gap
Environmental risk assessment of ARGs associated with waste material
To date, there have been few documented approaches to assess the risk of ARGs. A previous review summarized the research needed to enable human health risk assessment for environmental development and the transfer of antibiotic resistance [144]. ARGs are ubiquitous in soils, however, the long-term and repeated land application of waste material contaminated with ARGs can significantly increase the levels of ARGs, as described in preceding sections of the current review. Pertinent to any assessment of the environmental risk of ARGs associated with waste material is the key question of whether the elevated levels of ARGs in the environment can increase the likelihood of those soil bacteria acquiring resistance and, therefore, contributing to the development and spread of ARB that could become potential human pathogens.
There are natural barriers which may prevent the spread of ARGs. With respect to bacteria, potential human pathogens may acquire ARGs from soils containing ARB [145]. However, ARB introduced by land application of waste material may not able to compete with indigenous organisms [146]. One study showed that a 15-month period of land application of biosolids did not increase the percentage of antibiotic-resistant culturable bacteria above the background levels [147]. Another study showed that 20 years of applying Class B biosolids to soil did not increase either the overall microbial population in the soil or bacterial or viral pathogens [148]. With regards to DNA, soil through sorption can mitigate ARG transport, but as the process is reversible subsequent mobilization would be a concern. Soil also is abundant in DNase enzymes, which could degrade desorbed DNA, thus limiting its persistence in soils and its long-range transport.
The frequency of horizontal gene transfer in soils receiving applications of waste materials needs to be further evaluated and validated. In one study, the transferability of sul1 was analyzed in soil amended with manure from sulfadiazine-treated pigs, by capturing resistance plasmids from soil communities into Escherichia coli [149]. Manure from pigs not receiving sulfadiazine treatment was used as a control treatment. During a 6-month experimental period, the transfer frequency of resistant plasmids was up to two orders of magnitude higher in the soil amended with sulfadiazine-treated manure than in the control soil, indicating that the application of waste materials may potentially encourage horizontal gene transfer in soils through the excessive loading of resistant bacteria and ARGs [149]. On the contrary, some studies have shown a lack of evidence of horizontal gene transfer in soils [150, 151]. For example, in a greenhouse study, horizontal gene transfer from a model strain (Bacillus subtili) to soil bacteria was not observed during 6 months of cultivation [151]. Competent cells are essential for DNA to across their cell membranes, however, it is still questionable if there is an adequate number of natural competent cells in soils [152, 153]. Without a selective pressure, resistant bacteria are at a competitive disadvantage to sensitive bacteria [154]. One potential benefit of natural competence is that starving bacteria can take up exogenous DNA as a nutrient for competitive survival in environments containing limited nutrients [155, 156]. Conjugative plasmid transfer between bacteria was observed under an oligotrophy environment [157]. However, if an oligotrophy condition is important, the incidence of horizontal gene transfer would be low in the rhizosphere where vast amount of nutrients is excreted by plants. In addition, spatial separation of donor and recipient cells in the soil may also limit the possibility of horizontal gene transfer [158]. Further research is needed to identify the factors influencing the competency of indigenous soil bacteria, and improved quantitative models for horizontal gene transfer are needed.
On the other hand, some studies have suggested that it is the shift in the bacteria community—rather than horizontal gene transfer—that plays the dominant role in the enrichment of ARGs in soils. In one study, mobile genetic elements only explained 4.1% variation of ARGs in soils after the application of biosolids [81] and, similarly, in another study mobile genetic elements only contributed to 2.6% of the variation in ARGs during biosolids composting [159]. Five bacterial phyla, including Chloroflexi, Planctomycetes, Firmicutes, Gemmatimonadetes, and Bacteroidetes, were found to be significantly correlated with the relative abundance of ARGs in soil. In another study, six genera, including Flavobacterium, Poriferibacter, Bacteroides, Acinetobacter, Actinobaculu, and Streptococcus, were found to be correlated with ARGs in a WWTP, indicating that these genera played important roles in shaping the ARG profiles [160]. More data are required to understand the biological reservoir of ARGs in both waste material and waste material-applied soils.
Best Land Application Management Approaches for Waste Materials
Since common waste material pretreatment approaches are not efficient in reducing ARGs in waste materials, improved practices in waste materials land application strategies might be another option to control the dissemination of ARGs in soils. The transport of ARGs in soils can largely depend on a farmer’s approach(es) to the land application of waste material. For example, a recent study has shown that the injection of animal manure into the soil subsurface, a best land application management approach originally designed for reducing surface runoff of N and P [161], can reduce the surface runoff of antibiotics applied with dairy manure [162]. A comparison of the abundance of ARGs in runoff and manure land applied via broadcast, incorporation and injection methods after rainfall simulation revealed that the broadcast method resulted in significantly higher erm genes in runoff than did the incorporation and injection methods. However, the authors found no clear trend in ARG levels in the soil, likely because different host cells may respond differently to the soil environments created by various land application methods [111]. More information is needed to evaluate whether the manure subsurface injection approach is also an effective approach for minimizing and preventing the spread of ARB and ARGs in a wide range of soils amended with different types of waste materials.
It has also been shown that narrow grass hedges as buffer strips along waste material-applied fields are effective in reducing the levels of tylosin and erm(B) in runoff, suggesting that the implementation of buffer strips could be another management practice to control the transport of antibiotics, ARB, and ARGs in surface runoff of waste material-amended soils [163]. More investigation is needed to test whether this practice would be universally effective for different ecosystems. Also Adding other amendments when waste materials are applied to soils may also help reduce ARG abundance in waste material-applied soils. For example, when calcined eggshell waste was added to soils, the relative abundance of ten ARGs (tetW, tetM, tetB, tetQ, sulI, sulII, ermA, ermB, catI, and catII) fell by two to four orders of magnitude compared to the untreated soils [125]. Other materials that can be used as co-amendments with waste materials for controlling enrichment of ARGs in the waste material-affected soils need to be further investigated.
Conclusions
Waste materials contain high levels of antibiotics, ARB, and ARGs. Soil application of waste materials may introduce these emerging organic contaminants into the environment. Although the majority of antibiotic resistant bacterial infection is due to the overuse of antibiotics in the clinical environment, the contribution of elevated levels of antibiotics, ARB, and ARGs in the environment to the development of antibiotic resistance in potential pathogens should not be ignored. A better understanding of the environmental behaviors of ARGs in soils is critical to the control of ARG pollution. There is the potential risk that human health-related pathogens are becoming more resistant due to elevated levels of ARGs in the environment. Current strategies incorporating traditional waste material management approaches are not promising in reducing ARGs in waste materials. More importantly, agricultural activities may introduce selective pressure on natural soil-borne ARGs. It is unrealistic to expect to be able to eliminate completely ARGs in waste material-amended soils since ARGs could also be naturally produced. The question to be asked is at which levels should ARGs in soils be considered as safe for the ecosystem and humans. At the present time there are no environmental regulation standards for ARGs in the environmental matrixes. More studies are needed for environmental risk assessment. Information on the biological reservoirs of ARGs, the transferability of ARGs, and the possibility of ARGs entering food chain are also essential. Effective land application management approaches for waste materials are needed to prevent and minimize the accumulation and dispersion of ARGs in waste material-amended soils.
References
Centner TJ, Newton GL. Meeting environmental requirements for the land application of manure. J Anim Sci. 2008;86(11):3228–34. doi:10.2527/jas.2008-1161.
Hung CV, Cam BD, Mai PT, Dzung BQ. Heavy metals and polycyclic aromatic hydrocarbons in municipal sewage sludge from a river in highly urbanized metropolitan area in Hanoi, Vietnam: levels, accumulation pattern and assessment of land application. Environ Geochem Health. 2015;37(1):133–46. doi:10.1007/s10653-014-9635-2.
Clarke BO, Smith SR. Review of ‘emerging’ organic contaminants in biosolids and assessment of international research priorities for the agricultural use of biosolids. Environ Int. 2011;37(1):226–47. doi:10.1016/j.envint.2010.06.004.
Environmental Protection Agency. Aquatic life criteria for contaminants of emerging concern. 2008. https://www.epa.gov/sites/production/files/2015-08/documents/white_paper_aquatic_life_criteria_for_contaminants_of_emerging_concern_part_i_general_challenges_and_recommendations_1.pdf. Accessed 01 Sept 2016.
Chee-Sanford JC, Mackie RI, Koike S, Krapac IG, Lin YF, Yannarell AC, et al. Fate and transport of antibiotic residues and antibiotic resistance genes following land application of manure waste. J Environ Qual. 2009;38(3):1086–108. doi:10.2134/jeq2008.0128.
Hollis A, Ahmed Z. Preserving antibiotics, rationally. N Engl J Med. 2013;369(26):2474–6.
U.S. Environmental Protection Agency (EPA). Literature review of contaminants in livestock and poultry manure and implications for water quality. EPA 820-R-13-002. U.S. EPA, Washington, DC. 2013.
U.S. Food and Drug Administration (FDA). CVM reports on antimicrobials sold or distributed for food-producing animals for 2014. 2015. http://www.fda.gov/downloads/ForIndustry/UserFees/AnimalDrugUserFeeActADUFA/UCM476258.pdf. Accessed 1 April 2016.
Kim K-R, Owens G, Kwon S-I, So K-H, Lee D-B, Ok Y. Occurrence and environmental fate of veterinary antibiotics in the terrestrial environment. Water Air Soil Pollut. 2011;214(1–4):163–74. doi:10.1007/s11270-010-0412-2.
Kumar KC, Gupta S, Chander Y, Singh AK. Antibiotic use in agriculture and its impact on the terrestrial environment. In: Donald LS, editor. Advances in agronomy. Amsterdam: Academic Press; 2005. p. 1–54.
Hidaka K, Yamamoto H, Harada S, Katano M, Fujiwara H, Mizoguchi T, et al. Excretion of antibiotics into bile and urine in patients with external cholecystostomy done in order to treat obstructive jaundice. Jpn J Antibiot. 1988;41(9):1295–303.
Johnson AC, Williams RJ, Matthiessen P. The potential steroid hormone contribution of farm animals to freshwaters, the United Kingdom as a case study. Sci Total Environ. 2006;362(1):166–78. doi:10.1016/j.scitotenv.2005.06.014.
Sarmah AK, Meyer MT, Boxall ABA. A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere. 2006;65(5):725–59. doi:10.1016/j.chemosphere.2006.03.026.
Martínez-Carballo E, González-Barreiro C, Scharf S, Gans O. Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ Pollut. 2007;148(2):570–9. doi:10.1016/j.envpol.2006.11.035.
Huang CH, Renew JE, Smeby KL, Pinkston K, Sedlak DL. Assessment of potential antibiotic contaminants in water and preliminary occurrence analysis. J Contemp Water Res Educ. 2001;120:30–40.
Campagnolo ER, Johnson KR, Karpati A, Rubin CS, Kolpin DW, Meyer MT, et al. Antimicrobial residues in animal waste and water resources proximal to large-scale swine and poultry feeding operations. Sci Total Environ. 2002;299(1–3):89–95. doi:10.1016/S0048-9697(02)00233-4.
Boxall ABA, Kolpin DW, Halling-Sørensen B, Tolls J. Peer reviewed: are veterinary medicines causing environmental risks? Environ Sci Technol. 2003;37(15):286A–94. doi:10.1021/es032519b.
Meyer MT. Use and environmental occurrence of veterinary pharmaceuticals in the United States. In: Kummerer K, editor. Pharmaceuticals in the environment: sources, fate, effects, and risks. New York: Springer; 2004. p. 155–63.
Zhao L, Dong YH, Wang H. Residues of veterinary antibiotics in manures from feedlot livestock in eight provinces of China. Sci Total Environ. 2010;408(5):1069–75. doi:10.1016/j.scitotenv.2009.11.014.
Hu X, Zhou Q, Luo Y. Occurrence and source analysis of typical veterinary antibiotics in manure, soil, vegetables and groundwater from organic vegetable bases, northern China. Environ Pollut. 2010;158(9):2992–8. doi:10.1016/j.envpol.2010.05.023.
Ho YB, Zakaria MP, Latif PA, Saari N. Occurrence of veterinary antibiotics and progesterone in broiler manure and agricultural soil in Malaysia. Sci Total Environ. 2014;488–489:261–7. doi:10.1016/j.scitotenv.2014.04.109.
Li W, Shi Y, Gao L, Liu J, Cai Y. Occurrence, distribution and potential affecting factors of antibiotics in sewage sludge of wastewater treatment plants in China. Sci Total Environ. 2013;445–446:306–13. doi:10.1016/j.scitotenv.2012.12.050.
Qian M, Wu H, Wang J, Zhang H, Zhang Z, Zhang Y, et al. Occurrence of trace elements and antibiotics in manure-based fertilizers from the Zhejiang Province of China. Sci Total Environ. 2016;559:174–81. doi:10.1016/j.scitotenv.2016.03.123.
Brown KD, Kulis J, Thomson B, Chapman TH, Mawhinney DB. Occurrence of antibiotics in hospital, residential, and dairy effluent, municipal wastewater, and the Rio Grande in New Mexico. Sci Total Environ. 2006;366(2–3):772–83. doi:10.1016/j.scitotenv.2005.10.007.
Looft T, Allen HK, Cantarel BL, Levine UY, Bayles DO, Alt DP, et al. Bacteria, phages and pigs: the effects of in-feed antibiotics on the microbiome at different gut locations. ISME J. 2014;8(8):1566–76. doi:10.1038/ismej.2014.12.
Forslund K, Sunagawa S, Kultima JR, Mende DR, Arumugam M, Typas A, et al. Country-specific antibiotic use practices impact the human gut resistome. Genome Res. 2013;23(7):1163–9. doi:10.1101/gr.155465.113.
Yang Q, Ren S, Niu T, Guo Y, Qi S, Han X, et al. Distribution of antibiotic-resistant bacteria in chicken manure and manure-fertilized vegetables. Environ Sci Pollut Res. 2014;21(2):1231–41. doi:10.1007/s11356-013-1994-1.
Walczak JJ, Xu S. Manure as a source of antibiotic-resistant escherichia coli and enterococci: a case study of a Wisconsin, USA Family Dairy Farm. Water Air Soil Pollut. 2011;219(1):579–89. doi:10.1007/s11270-010-0729-x.
Zhou W, Wang Y, Lin J. Functional cloning and characterization of antibiotic resistance genes from the chicken gut microbiome. Appl Environ Microbiol. 2012;78(8):3028–32. doi:10.1128/AEM.06920-11.
Looft T, Johnson TA, Allen HK, Bayles DO, Alt DP, Stedtfeld RD, et al. In-feed antibiotic effects on the swine intestinal microbiome. Proc Natl Acad Sci USA. 2012;109(5):1691–6. doi:10.1073/pnas.1120238109.
Hoelzel CS, Harms KS, Kuechenhoff H, Kunz A, Mueller C, Meyer K, et al. Phenotypic and genotypic bacterial antimicrobial resistance in liquid pig manure is variously associated with contents of tetracyclines and sulfonamides. J Appl Microbiol. 2010;108(5):1642–56. doi:10.1111/j.1365-2672.2009.04570.x.
Agga GE, Arthur TM, Durso LM, Harhay DM, Schmidt JW. Antimicrobial-resistant bacterial populations and antimicrobial resistance genes obtained from environments impacted by livestock and municipal waste. PLoS One. 2015;10(7):e0132586. doi:10.1371/journal.pone.0132586.
Michael I, Rizzo L, McArdell CS, Manaia CM, Merlin C, Schwartz T, et al. Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: a review. Water Res. 2013;47(3):957–95. doi:10.1016/j.watres.2012.11.027.
Pan M, Wong CK, Chu LM. Distribution of antibiotics in wastewater-irrigated soils and their accumulation in vegetable crops in the Pearl River Delta, southern China. J Agric Food Chem. 2014;62(46):11062–9. doi:10.1021/jf503850v.
Wang J, Ben W, Yang M, Zhang Y, Qiang Z. Dissemination of veterinary antibiotics and corresponding resistance genes from a concentrated swine feedlot along the waste treatment paths. Environ Int. 2016;92–93:317–23. doi:10.1016/j.envint.2016.04.020.
Hamscher G, Sczesny S, Hoper H, Nau H. Determination of persistent tetracycline residues in soil fertilized with liquid manure by high-performance liquid chromatography with electrospray ionization tandem mass spectrometry. Anal Chem. 2002;74(7):1509–18. doi:10.1021/ac015588m.
Knapp CW, Dolfing J, Ehlert PA, Graham DW. Evidence of increasing antibiotic resistance gene abundances in archived soils since 1940. Environ Sci Technol. 2010;44(2):580–7. doi:10.1021/es901221x.
Heuer H, Smalla K. Manure and sulfadiazine synergistically increased bacterial antibiotic resistance in soil over at least two months. Environ Microbiol. 2007;9(3):657–66. doi:10.1111/j.1462-2920.2006.01185.x.
Liu J, Zhao Z, Orfe L, Subbiah M, Call DR. Soil-borne reservoirs of antibiotic-resistant bacteria are established following therapeutic treatment of dairy calves. Environ Microbiol. 2016;18(2):557–64. doi:10.1111/1462-2920.13097.
Sipahi OR. Economics of antibiotic resistance. Expert Rev Anti-Infect Ther. 2008;6(4):523–39. doi:10.1586/14787210.6.4.523.
Martinez JL. Antibiotics and antibiotic resistance genes in natural environments. Science. 2008;321(5887):365–7. doi:10.1126/science.1159483.
World Health Organization. Fact sheet on antibiotic resistance. 2015. http://www.who.int/mediacentre/factsheets/antibiotic-resistance/en/. Accessed 15 July 2016.
Huttner A, Harbarth S, Carlet J, Cosgrove S, Goossens H, Holmes A, et al. Antimicrobial resistance: a global view from the 2013 World Healthcare-Associated Infections Forum. Antimicrob Resist Infect Control. 2013;2:31–43. doi:10.1186/2047-2994-2-31.
World Health Organization. World health report: the world health report 2007—a safer future: global public health security in the 21st century. 2007. http://www.who.int/whr/2007/overview/en/. Accessed 15 July 2016.
Tacconelli E, De Angelis G, Cataldo MA, Mantengoli E, Spanu T, Pan A, et al. Antibiotic usage and risk of colonization and infection with antibiotic-resistant bacteria: a hospital population-based study. Antimicrob Agents Chemother. 2009;53(10):4264–9. doi:10.1128/AAC.00431-09.
Pallecchi L, Bartoloni A, Paradisi F, Rossolini GM. Antibiotic resistance in the absence of antimicrobial use: mechanisms and implications. Expert Rev Anti-Infect Ther. 2008;6(5):725–32. doi:10.1586/14787210.6.5.725.
Davies J, Davies D. Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev. 2010;74(3):417–33. doi:10.1128/MMBR.00016-10.
Hawkey PM. The origins and molecular basis of antibiotic resistance. BMJ. 1998;317(7159):657–60.
Pruden A, Pei RT, Storteboom H, Carlson KH. Antibiotic resistance genes as emerging contaminants: studies in northern Colorado. Environ Sci Technol. 2006;40(23):7445–50. doi:10.1021/es0604131.
Berglund B. Environmental dissemination of antibiotic resistance genes and correlation to anthropogenic contamination with antibiotics. Infect Ecol Epidemiol. 2015;5:28564. doi:10.3402/iee.v5.28564.
Li B, Zhang X, Guo F, Wu W, Zhang T. Characterization of tetracycline resistant bacterial community in saline activated sludge using batch stress incubation with high-throughput sequencing analysis. Water Res. 2013;47(13):4207–16. doi:10.1016/j.watres.2013.04.021.
Mao DQ, Yu S, Rysz M, Luo Y, Yang FX, Li FX, et al. Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water Res. 2015;85:458–66. doi:10.1016/j.watres.2015.09.010.
Li J, Cheng W, Xu L, Strong PJ, Chen H. Antibiotic-resistant genes and antibiotic-resistant bacteria in the effluent of urban residential areas, hospitals, and a municipal wastewater treatment plant system. Environ Sci Pollut Res Int. 2015;22(6):4587–96. doi:10.1007/s11356-014-3665-2.
Chen B, Hao L, Guo X, Wang N, Ye B. Prevalence of antibiotic resistance genes of wastewater and surface water in livestock farms of Jiangsu Province, China. Environ Sci Pollut Res Int. 2015;22(18):13950–9. doi:10.1007/s11356-015-4636-y.
Peng S, Wang Y, Zhou B, Lin X. Long-term application of fresh and composted manure increase tetracycline resistance in the arable soil of eastern China. Sci Total Environ. 2015;506–507:279–86. doi:10.1016/j.scitotenv.2014.11.010.
Fahrenfeld N, Ma YJ, O’Brien M, Pruden A. Reclaimed water as a reservoir of antibiotic resistance genes: distribution system and irrigation implications. Front Microbiol. 2013;4:130. doi:10.3389/fmicb.2013.00130.
Burch TR, Sadowsky MJ, LaPara TM. Fate of antibiotic resistance genes and class 1 integrons in soil microcosms following the application of treated residual municipal wastewater solids. Environ Sci Technol. 2014;48(10):5620–7. doi:10.1021/es501098g.
Huang X, Liu CX, Li K, Liu F, Liao DR, Liu L, et al. Occurrence and distribution of veterinary antibiotics and tetracycline resistance genes in farmland soils around swine feedlots in Fujian Province, China. Environ Sci Pollut Res. 2013;20(12):9066–74. doi:10.1007/s11356-013-1905-5.
He LY, Ying GG, Liu YS, Su HC, Chen J, Liu SS, et al. Discharge of swine wastes risks water quality and food safety: antibiotics and antibiotic resistance genes from swine sources to the receiving environments. Environ Int. 2016;92–93:210–9. doi:10.1016/j.envint.2016.03.023.
Brooks JP, Adeli A, McLaughlin MR. Microbial ecology, bacterial pathogens, and antibiotic resistant genes in swine manure wastewater as influenced by three swine management systems. Water Res. 2014;57:96–103. doi:10.1016/j.watres.2014.03.017.
Cheng WX, Chen H, Su C, Yan SH. Abundance and persistence of antibiotic resistance genes in livestock farms: a comprehensive investigation in eastern China. Environ Int. 2013;61:1–7. doi:10.1016/j.envint.2013.08.023.
Zhang SH, Gu J, Wang C, Wang PF, Jiao SJ, He ZL, et al. Characterization of antibiotics and antibiotic resistance genes on an ecological farm system. J Chem-Ny. 2015. doi:10.1155/2015/526143.
Sui QW, Zhang JY, Chen MX, Tong J, Wang R, Wei YS. Distribution of antibiotic resistance genes (ARGs) in anaerobic digestion and land application of swine wastewater. Environ Pollut. 2016;213:751–9. doi:10.1016/j.envpol.2016.03.038.
Ruuskanen M, Muurinen J, Meierjohan A, Parnanen K, Tamminen M, Lyra C, et al. Fertilizing with animal manure disseminates antibiotic resistance genes to the farm environment. J Environ Qual. 2016;45(2):488–93. doi:10.2134/jeq2015.05.0250.
Wang H, Li HY, Gilbert JA, Li HB, Wu LH, Liu M, et al. Housefly larva vermicomposting efficiently attenuates antibiotic resistance genes in swine manure, with concomitant bacterial population changes. Appl Environ Microbiol. 2015;81(22):7668–79. doi:10.1128/Aem.01367-15.
Rodriguez-Mozaz S, Chamorro S, Marti E, Huerta B, Gros M, Sanchez-Melsio A, et al. Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res. 2015;69:234–42. doi:10.1016/j.watres.2014.11.021.
Li J, Shao B, Shen JZ, Wang SC, Wu YN. Occurrence of chloramphenicol-resistance genes as environmental pollutants from swine feedlots. Environ Sci Technol. 2013;47(6):2892–7. doi:10.1021/es304616c.
Li L, Sun J, Liu BT, Zhao DH, Ma J, Deng H, et al. Quantification of lincomycin resistance genes associated with lincomycin residues in waters and soils adjacent to representative swine farms in China. Front Microbiol. 2013;4:364. doi:10.3389/fmicb.2013.00364.
Alexander J, Knopp G, Dotsch A, Wieland A, Schwartz T. Ozone treatment of conditioned wastewater selects antibiotic resistance genes, opportunistic bacteria, and induce strong population shifts. Sci Total Environ. 2016;559:103–12. doi:10.1016/j.scitotenv.2016.03.154.
Luby EM, Moorman TB, Soupir ML. Fate and transport of tylosin-resistant bacteria and macrolide resistance genes in artificially drained agricultural fields receiving swine manure. Sci Total Environ. 2016;550:1126–33. doi:10.1016/j.scitotenv.2016.01.132.
Yuan QB, Guo MT, Yang J. Monitoring and assessing the impact of wastewater treatment on release of both antibiotic-resistant bacteria and their typical genes in a Chinese municipal wastewater treatment plant. Environ Sci Process Impacts. 2014;16(8):1930–7. doi:10.1039/c4em00208c.
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, et al. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ. 2013;447:345–60. doi:10.1016/j.scitotenv.2013.01.032.
Bondarczuk K, Markowicz A, Piotrowska-Seget Z. The urgent need for risk assessment on the antibiotic resistance spread via sewage sludge land application. Environ Int. 2016;87:49–55. doi:10.1016/j.envint.2015.11.011.
Xu LK, Ouyang WY, Qian YY, Su C, Su JQ, Chen H. High-throughput profiling of antibiotic resistance genes in drinking water treatment plants and distribution systems. Environ Pollut. 2016;213:119–26. doi:10.1016/j.envpol.2016.02.013.
Thiele-Bruhn S. Pharmaceutical antibiotic compounds in soils—a review. J Plant Nutr Soil Sci. 2003;166(2):145–67. doi:10.1002/jpln.200390023.
Tang XJ, Lou CL, Wang SX, Lu YH, Liu M, Hashmi MZ, et al. Effects of long-term manure applications on the occurrence of antibiotics and antibiotic resistance genes (ARGs) in paddy soils: evidence from four field experiments in south of China. Soil Biol Biochem. 2015;90:179–87. doi:10.1016/j.soilbio.2015.07.027.
Zhu YG, Johnson TA, Su JQ, Qiao M, Guo GX, Stedtfeld RD, et al. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci USA. 2013;110(9):3435–40. doi:10.1073/pnas.1222743110.
Chen CQ, Li J, Chen PP, Ding R, Zhang PF, Li XQ. Occurrence of antibiotics and antibiotic resistances in soils from wastewater irrigation areas in Beijing and Tianjin, China. Environ Pollut. 2014;193:94–101. doi:10.1016/j.envpol.2014.06.005.
Dalkmann P, Broszat M, Siebe C, Willaschek E, Sakinc T, Huebner J, et al. Accumulation of pharmaceuticals, Enterococcus, and resistance genes in soils irrigated with wastewater for zero to 100 years in central Mexico. PLoS One. 2012;7(9):e45397. doi:10.1371/journal.pone.0045397.
Negreanu Y, Pasternak Z, Jurkevitch E, Cytryn E. Impact of treated wastewater irrigation on antibiotic resistance in agricultural soils. Environ Sci Technol. 2012;46(9):4800–8. doi:10.1021/es204665b.
Chen Q, An X, Li H, Su J, Ma Y, Zhu YG. Long-term field application of sewage sludge increases the abundance of antibiotic resistance genes in soil. Environ Int. 2016;92–93:1–10. doi:10.1016/j.envint.2016.03.026.
Rahube TO, Marti R, Scott A, Tien YC, Murray R, Sabourin L, et al. Persistence of antibiotic resistance and plasmid-associated genes in soil following application of sewage sludge and abundance on vegetables at harvest. Can J Microbiol. 2016;62(7):600–7. doi:10.1139/cjm-2016-0034.
Svara F, Rankin DJ. The evolution of plasmid-carried antibiotic resistance. BMC Evol Biol. 2011;11:130. doi:10.1186/1471-2148-11-130.
Lorenz MG, Gerjets D, Wackernagel W. Release of transforming plasmid and chromosomal DNA from two cultured soil bacteria. Arch Microbiol. 1991;156(4):319–26.
Cai P, Huang Q, Zhang X, Chen H. Adsorption of DNA on clay minerals and various colloidal particles from an Alfisol. Soil Biol Biochem. 2006;38(3):471–6. doi:10.1016/j.soilbio.2005.05.019.
Cai P, Huang QY, Zhang XW. Microcalorimetric studies of the effects of MgCl2 concentrations and pH on the adsorption of DNA on montmorillonite, kaolinite and goethite. Appl Clay Sci. 2006;32(1–2):147–52. doi:10.1016/j.clay.2005.11.004.
Cai P, Huang QY, Zhu J, Jiang DH, Zhou XY, Rong XM, et al. Effects of low-molecular-weight organic ligands and phosphate on DNA adsorption by soil colloids and minerals. Colloids Surface B. 2007;54(1):53–9. doi:10.1016/j.colsurfb.2006.07.013.
Saeki K, Kunito T, Sakai M. Effects of pH, ionic strength, and solutes on DNA adsorption by andosols. Biol Fertil Soils. 2010;46(5):531–5. doi:10.1007/s00374-010-0447-y.
Heuer H, Focks A, Lamshoeft M, Smalla K, Matthies M, Spiteller M. Fate of sulfadiazine administered to pigs and its quantitative effect on the dynamics of bacterial resistance genes in manure and manured soil. Soil Biol Biochem. 2008;40(7):1892–900. doi:10.1016/j.soilbio.2008.03.014.
Heuer H, Solehati Q, Zimmerling U, Kleineidam K, Schloter M, Muller T, et al. Accumulation of sulfonamide resistance genes in arable soils due to repeated application of manure containing sulfadiazine. Appl Environ Microbiol. 2011;77(7):2527–30. doi:10.1128/AEM.02577-10.
Furuya EY, Lowy FD. Antimicrobial-resistant bacteria in the community setting. Nat Rev Microbiol. 2006;4(1):36–45. doi:10.1038/nrmicro1325.
Davison J. Genetic exchange between bacteria in the environment. Plasmid. 1999;42(2):73–91. doi:10.1006/plas.1999.1421.
Pietramellara G, Ascher J, Borgogni F, Ceccherini MT, Guerri G, Nannipieri P. Extracellular DNA in soil and sediment: fate and ecological relevance. Biol Fertil Soils. 2009;45(3):219–35. doi:10.1007/s00374-008-0345-8.
Khanna M, Stotzky G. Transformation of Bacillus-subtilis by DNA bound on montmorillonite and effect of Dnase on the transforming ability of bound DNA. Appl Environ Microbiol. 1992;58(6):1930–9.
Baquero F, Negri MC, Morosini MI, Blazquez J. Antibiotic-selective environments. Clin Infect Dis. 1998;27 Suppl 1:S5–11.
Sandegren L. Selection of antibiotic resistance at very low antibiotic concentrations. Ups J Med Sci. 2014;119(2):103–7. doi:10.3109/03009734.2014.904457.
Seiler C, Berendonk TU. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol. 2012;3:399. doi:10.3389/fmicb.2012.00399.
Gullberg E, Albrecht LM, Karlsson C, Sandegren L, Andersson DI. Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. MBio. 2014;5(5):e01918–14. doi:10.1128/mBio.01918-14.
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV. Co-selection of antibiotic and metal resistance. Trends Microbiol. 2006;14(4):176–82. doi:10.1016/j.tim.2006.02.006.
Henriques I, Tacao M, Leite L, Fidalgo C, Araujo S, Oliveira C, et al. Co-selection of antibiotic and metal(loid) resistance in Gram-negative epiphytic bacteria from contaminated salt marshes. Mar Pollut Bull. 2016;109(1):427–34. doi:10.1016/j.marpolbul.2016.05.031.
Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ. Co-occurrence of resistance genes to antibiotics, biocides and metals reveals novel insights into their co-selection potential. BMC Genomics. 2015;16:964–77. doi:10.1186/s12864-015-2153-5.
Ji X, Shen Q, Liu F, Ma J, Xu G, Wang Y, et al. Antibiotic resistance gene abundances associated with antibiotics and heavy metals in animal manures and agricultural soils adjacent to feedlots in Shanghai, China. J Hazard Mater. 2012;235–236:178–85. doi:10.1016/j.jhazmat.2012.07.040.
Hu HW, Wang JT, Li J, Li JJ, Ma YB, Chen D, et al. Field-based evidence for copper contamination induced changes of antibiotic resistance in agricultural soils. Environ Microbiol. 2016;18(11):3896–909. doi:10.1111/1462-2920.13370.
Udikovic-Kolic N, Wichmann F, Broderick NA, Handelsman J. Bloom of resident antibiotic-resistant bacteria in soil following manure fertilization. Proc Natl Acad Sci USA. 2014;111(42):15202–7. doi:10.1073/pnas.1409836111.
Elrashidi MA, Mays MD, Fares A, Seybold CA, Harder JL, Peaslee SD, et al. Loss of nitrate-nitrogen by runoff and leaching for agricultural watersheds. Soil Sci. 2005;170(12):969–84. doi:10.1097/01.ss.0000187353.24364.a8.
Turtola E, Jaakkola A. Loss of phosphorus by surface runoff and leaching from a heavy clay soil under Barley and Grass Ley in Finland. Acta Agric Scand B Soil Plant. 1995;45(3):159–65.
Forney DR, Strahan J, Rankin C, Steffin D, Peter CJ, Spittler TD, et al. Monitoring pesticide runoff and leaching from four farming systems on field-scale coastal plain watersheds in Maryland. Abstr Pap Am Chem Soc. 1998;215:U26–7.
Dolliver H, Gupta S. Antibiotic losses in leaching and surface runoff from manure-amended agricultural land. J Environ Qual. 2008;37(3):1227–37. doi:10.2134/jeq2007.0392.
Rysz M, Alvarez PJ. Transport of antibiotic-resistant bacteria and resistance-carrying plasmids through porous media. Water Sci Technol. 2006;54(11–12):363–70.
Chen CQ, Li J, DeVries SL, Zhang PF, Li XQ. Transport of antibiotic resistance plasmids in porous media. Vadose Zone J. 2015;14(3). doi:10.2136/vzj2014.06.0068.
Joy SR, Bartelt-Hunt SL, Snow DD, Gilley JE, Woodbury BL, Parker DB, et al. Fate and transport of antimicrobials and antimicrobial resistance genes in soil and runoff following land application of swine manure slurry. Environ Sci Technol. 2013;47(21):12081–8. doi:10.1021/es4026358.
Fahrenfeld N, Knowlton K, Krometis LA, Hession WC, Xia K, Lipscomb E, et al. Effect of manure application on abundance of antibiotic resistance genes and their attenuation rates in soil: field-scale mass balance approach. Environ Sci Technol. 2014;48(5):2643–50. doi:10.1021/es404988k.
Hong PY, Yannarell AC, Dai QH, Ekizoglu M, Mackie RI. Monitoring the perturbation of soil and groundwater microbial communities due to pig production activities. Appl Environ Microbiol. 2013;79(8):2620–9. doi:10.1128/Aem.03760-12.
Pruden A, Arabi M, Storteboom HN. Correlation between upstream human activities and riverine antibiotic resistance genes. Environ Sci Technol. 2012;46(21):11541–9. doi:10.1021/es302657r.
Andrews JH, Harris RF. The ecology and biogeography of microorganisms on plant surfaces. Annu Rev Phytopathol. 2000;38:145–80. doi:10.1146/annurev.phyto.38.1.145.
Rosenblueth M, Martinez-Romero E. Bacterial endophytes and their interactions with hosts. Mol Plant-Microbe Interact. 2006;19(8):827–37. doi:10.1094/Mpmi-19-0827.
Heuer H, Schmitt H, Smalla K. Antibiotic resistance gene spread due to manure application on agricultural fields. Curr Opin Microbiol. 2011;14(3):236–43. doi:10.1016/j.mib.2011.04.009.
Marti R, Scott A, Tien YC, Murray R, Sabourin L, Zhang Y, et al. Impact of manure fertilization on the abundance of antibiotic-resistant bacteria and frequency of detection of antibiotic resistance genes in soil and on vegetables at harvest. Appl Environ Microbiol. 2013;79(18):5701–9. doi:10.1128/AEM.01682-13.
Wyttenbach A, Tobler L. Effect of surface contamination on results of plant analysis. Commun Soil Sci Plant Anal. 1998;29(7–8):809–23. doi:10.1080/00103629809369987.
Paungfoo-Lonhienne C, Lonhienne TGA, Mudge SR, Schenk PM, Christie M, Carroll BJ, et al. DNA is taken up by root hairs and pollen, and stimulates root and pollen tube growth. Plant Physiol. 2010;153(2):799–805. doi:10.1104/pp.110.154963.
Azanu D, Mortey C, Darko G, Weisser JJ, Styrishave B, Abaidoo RC. Uptake of antibiotics from irrigation water by plants. Chemosphere. 2016;157:107–14. doi:10.1016/j.chemosphere.2016.05.035.
Kumar K, Gupta SC, Baidoo SK, Chander Y, Rosen CJ. Antibiotic uptake by plants from soil fertilized with animal manure. J Environ Qual. 2005;34(6):2082–5. doi:10.2134/jeq2005.0026.
Bassil RJ, Bashour II, Sleiman FT, Abou-Jawdeh YA. Antibiotic uptake by plants from manure-amended soils. J Environ Sci Health B. 2013;48(7):570–4. doi:10.1080/03601234.2013.774898.
Ye M, Sun M, Wan J, Feng Y, Zhao Y, Tian D, et al. Feasibility of lettuce cultivation in sophoroliplid-enhanced washed soil originally polluted with Cd, antibiotics, and antibiotic-resistant genes. Ecotoxicol Environ Saf. 2016;124:344–50. doi:10.1016/j.ecoenv.2015.11.013.
Ye M, Sun M, Feng Y, Li X, Schwab AP, Wan J, et al. Calcined eggshell waste for mitigating soil antibiotic-resistant bacteria/antibiotic resistance gene dissemination and accumulation in bell pepper. J Agric Food Chem. 2016;64(27):5446–53. doi:10.1021/acs.jafc.6b00866.
Sancheza HM, Echeverria C, Thulsiraj V, Zimmer-Faust A, Flores A, Laitz M, et al. Antibiotic resistance in airborne bacteria near conventional and organic beef cattle farms in California, USA. Water Air Soil Pollut. 2016;227(8):280. doi:10.1007/s11270-016-2979-8.
Heim S, Lleo M, Bonato B, Guzman CA, Canepari P. The viable but nonculturable state and starvation are different stress responses of Enterococcus faecalis, as determined by proteome analysis. J Bacteriol. 2002;184(23):6739–45.
Pasquaroli S, Zandri G, Vignaroli C, Vuotto C, Donelli G, Biavasco F. Antibiotic pressure can induce the viable but non-culturable state in Staphylococcus aureus growing in biofilms. J Antimicrob Chemother. 2013;68(8):1812–7. doi:10.1093/jac/dkt086.
Giusti L. A review of waste management practices and their impact on human health. Waste Manage. 2009;29(8):2227–39. doi:10.1016/j.wasman.2009.03.028.
Beneragama N, Iwasaki M, Lateef SA, Yamashiro T, Ihara I, Umetsu K. Survival of multidrug-resistant bacteria in thermophilic and mesophilic anaerobic co-digestion of dairy manure and waste milk. Anim Sci J. 2013;84(5):426–33. doi:10.1111/asj.12017.
Mitchell SM, Ullman JL, Bary A, Cogger CG, Teel AL, Watts RJ. Antibiotic degradation during thermophilic composting. Water Air Soil Pollut. 2015;226(2):1–12. doi:10.1007/s11270-014-2288-z.
Zheng NG, Huang N, Wang WW, Yu M, Chen XY, Yao YL, et al. Effects of thermophilic composting on antibiotic resistance genes (ARGs) of swine manure source. Huan Jing Ke Xue. 2016;37(5):1986–92.
Wang J, Ben W, Zhang Y, Yang M, Qiang Z. Effects of thermophilic composting on oxytetracycline, sulfamethazine, and their corresponding resistance genes in swine manure. Environ Sci Process Impacts. 2015;17(9):1654–60. doi:10.1039/c5em00132c.
Wang H, Sangwan N, Li HY, Su JQ, Oyang WY, Zhang ZJ, et al. The antibiotic resistome of swine manure is significantly altered by association with the Musca domestica larvae gut microbiome. ISME J. 2016. doi:10.1038/ismej.2016.103.
Ramaswamy J, Prasher SO, Patel RM, Hussain SA, Barrington SF. The effect of composting on the degradation of a veterinary pharmaceutical. Bioresource Technol. 2010;101(7):2294–9. doi:10.1016/j.biortech.2009.10.089.
Pruden A, Larsson DG, Amezquita A, Collignon P, Brandt KK, Graham DW, et al. Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environ Health Perspect. 2013;121(8):878–85. doi:10.1289/ehp.1206446.
Guo MT, Yuan QB, Yang J. Ultraviolet reduction of erythromycin and tetracycline resistant heterotrophic bacteria and their resistance genes in municipal wastewater. Chemosphere. 2013;93(11):2864–8. doi:10.1016/j.chemosphere.2013.08.068.
Yuan QB, Guo MT, Yang J. Fate of antibiotic resistant bacteria and genes during wastewater chlorination: implication for antibiotic resistance control. PLoS One. 2015;10(3):e0119403. doi:10.1371/journal.pone.0119403.
Shi P, Jia S, Zhang XX, Zhang T, Cheng S, Li A. Metagenomic insights into chlorination effects on microbial antibiotic resistance in drinking water. Water Res. 2013;47(1):111–20. doi:10.1016/j.watres.2012.09.046.
Munir M, Wong K, Xagoraraki I. Release of antibiotic resistant bacteria and genes in the effluent and biosolids of five wastewater utilities in Michigan. Water Res. 2011;45(2):681–93. doi:10.1016/j.watres.2010.08.033.
Li JN, Cheng WX, Xu LK, Jiao YN, Baig SA, Chen H. Occurrence and removal of antibiotics and the corresponding resistance genes in wastewater treatment plants: effluents’ influence to downstream water environment. Environ Sci Pollut Res. 2016;23(7):6826–35. doi:10.1007/s11356-015-5916-2.
Tao CW, Hsu BM, Ji WT, Hsu TK, Kao PM, Hsu CP, et al. Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci Total Environ. 2014;496:116–21. doi:10.1016/j.scitotenv.2014.07.024.
Zhang T, Yang Y, Pruden A. Effect of temperature on removal of antibiotic resistance genes by anaerobic digestion of activated sludge revealed by metagenomic approach. Appl Microbiol Biotechnol. 2015;99(18):7771–9. doi:10.1007/s00253-015-6688-9.
Ashbolt NJ, Amezquita A, Backhaus T, Borriello P, Brandt KK, Collignon P, et al. Human Health Risk Assessment (HHRA) for environmental development and transfer of antibiotic resistance. Environ Health Perspect. 2013;121(9):993–1001. doi:10.1289/ehp.1206316.
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MO, Dantas G. The shared antibiotic resistome of soil bacteria and human pathogens. Science. 2012;337(6098):1107–11. doi:10.1126/science.1220761.
Elliott LF, Ellis JR. Bacterial and viral pathogens associated with land application of organic wastes. J Environ Qual. 1977;6(3):245–51.
Brooks JP, Maxwell SL, Rensing C, Gerba CP, Pepper IL. Occurrence of antibiotic-resistant bacteria and endotoxin associated with the land application of biosolids. Can J Microbiol. 2007;53(5):616–22. doi:10.1139/W07-021.
Zerzghi H, Gerba CP, Brooks JP, Pepper IL. Long-term effects of land application of class B biosolids on the soil microbial populations, pathogens, and activity. J Environ Qual. 2010;39(1):402–8. doi:10.2134/jeq2009.0307.
Jechalke S, Kopmann C, Rosendahl I, Groeneweg J, Weichelt V, Krogerrecklenfort E, et al. Increased abundance and transferability of resistance genes after field application of manure from sulfadiazine-treated pigs. Appl Environ Microbiol. 2013;79(5):1704–11. doi:10.1128/AEM.03172-12.
Sommer MO. Microbiology: barriers to the spread of resistance. Nature. 2014;509(7502):567–8. doi:10.1038/nature13342.
Kim SE, Moon JS, Choi WS, Lee SH, Kim SU. Monitoring of horizontal gene transfer from agricultural microorganisms to soil bacteria and analysis of microbial community in soils. J Microbiol Biotechnol. 2012;22(4):563–6.
de Vries J, Meier P, Wackernagel W. The natural transformation of the soil bacteria Pseudomonas stutzeri and Acinetobacter sp. by transgenic plant DNA strictly depends on homologous sequences in the recipient cells. FEMS Microbiol Lett. 2001;195(2):211–5.
Nielsen KM. Barriers to horizontal gene transfer by natural transformation in soil bacteria. APMIS Suppl. 1998;84:77–84.
Lenski RE, Levin BR. Constraints on the coevolution of bacteria and virulent phage—a model, some experiments, and predictions for natural communities. Am Nat. 1985;125(4):585–602. doi:10.1086/284364.
Palchevskiy V, Finkel SE. Escherichia coli competence gene homologs are essential for competitive fitness and the use of DNA as a nutrient. J Bacteriol. 2006;188(11):3902–10. doi:10.1128/Jb.01974-05.
Finkel SE, Kolter R. DNA as a nutrient novel role for bacterial competence gene homologs. J Bacteriol. 2001;183(21):6288–93. doi:10.1128/Jb.183.21.6288-6293.2001.
Goodman AE, Hild E, Marshall KC, Hermansson M. Conjugative plasmid transfer between bacteria under simulated marine oligotrophic conditions. Appl Environ Microbiol. 1993;59(4):1035–40.
Hill KE, Top EM. Gene transfer in soil systems using microcosms. FEMS Microbiol Ecol. 1998;25(4):319–29. doi:10.1016/S0168-6496(97)00101-3.
Su JQ, Wei B, Ou-Yang WY, Huang FY, Zhao Y, Xu HJ, et al. Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol. 2015;49(12):7356–63. doi:10.1021/acs.est.5b01012.
Yang Y, Li B, Zou S, Fang HH, Zhang T. Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Res. 2014;62:97–106. doi:10.1016/j.watres.2014.05.019.
Maguire RO, Kleinman PJA, Dell CJ, Beegle DB, Brandt RC, McGrath JM, et al. Manure application technology in reduced tillage and forage systems: a review. J Environ Qual. 2011;40(2):292–301. doi:10.2134/jeq2009.0228.
Kulesza SB, Maguire RO, Xia K, Cushman J, Knowlton K, Ray P. Manure injection affects the fate of pirlimycin in surface runoff and soil. J Environ Qual. 2016;45(2):511–8. doi:10.2134/jeq2015.06.0266.
Soni B, Bartelt-Hunt SL, Snow DD, Gilley JE, Woodbury BL, Marx DB, et al. Narrow grass hedges reduce tylosin and associated antimicrobial resistance genes in agricultural runoff. J Environ Qual. 2015;44(3):895–902. doi:10.2134/jeq2014.09.0389.
Acknowledgments
This study was supported by NIFA Competitive Grant no. 2014–05280 from the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
This article is part of the Topical Collection on Land Pollution
Rights and permissions
About this article
Cite this article
Chen, C., Xia, K. Fate of Land Applied Emerging Organic Contaminants in Waste Materials. Curr Pollution Rep 3, 38–54 (2017). https://doi.org/10.1007/s40726-017-0048-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40726-017-0048-6