Abstract
Arabidopsis gun4 mutant is defected in retrograde signaling and is characterized by reduced levels of chlorophyll. However, the impact of this mutation on plant photosynthetic performance remains unclear. We carried out a physiological characterization of gun4 in order to investigate the role of GUN4 for plant photosynthetic performance under light stress. The gun4 plants showed reduced minimal fluorescence in dark-adapted leaves and quantum yield of unregulated energy dissipation of photosystem II (PSII) under non-light-stress condition. The effective quantum yield of the PSII (ΦPSII) and photochemical quenching (qL) were higher in gun4 plants. Higher values of ΦPSII were also observed in gun4 under different light intensities. However, the rate of net carbon assimilation and stomatal conductance were lower in gun4. No differences were detected between the genotypes in the total absorption of 14CO2 as well as in the percentage of 14C flux to basic amino acids, sugars, starch, and cell wall. After light stress, the potential quantum yield of PSII decreased only in wild type and the non-photochemical quenching was higher in gun4. Taken together, the results suggest that gun4 transfers more efficiently the excess of light energy absorbed despite a reduction in chlorophyll and carotenoids content and has greater capacity to dissipate the excess of energy absorbed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Pathways that communicate chloroplast to nucleus, called retrograde signaling, has been revealed through gun (g enome un coupled) mutants of Arabidopsis thaliana. Retrograde signaling involves plastids signals that induce nuclear gene expression. There are five genes GUN in Arabidopsis (GUN 1-5), which four of these (GUN2-5) encode proteins related to tetrapyrrole biosynthetic pathway (Susek and Chory 1992; Susek et al. 1993; Mochizuki et al. 2001; Larkin et al. 2003). GUN4 encodes a chloroplast protein that activates Mg-chelatase enzyme (MgChl), responsible by protoporphyrinIX (ProtoIX) to Mg-protoporphyrinIX (Mg-ProtoIX) conversion in the chlorophyll biosynthetic pathway (Fig. 1) (Tanaka and Tanaka 2007). Furthermore, GUN4 is required for posttranslational control of tetrapyrrole biosynthesis for promoting the interaction between H-subunit of MgChl enzyme (MgChl-H) and chloroplast membranes (Adhikari et al. 2009; Peter and Grimm 2009). The phenotype of Arabidopsis gun4 mutant was characterized by reduced levels of Mg-ProtoIX, chlorophyll and carotenoids (Larkin et al. 2003). In the other hand, the overexpression of GUN4 in tobacco increased chlorophyll and Mg-ProtoIX levels as well as increased MgChl enzyme activity (Peter and Grimm 2009). Interestingly, GUN4 mutation in cyanobacteria leads to reduced both Fe-chelatase enzyme activity and Heme levels. It suggests that GUN4 is also involved in the biosynthesis of Heme groups, controlling the flow of substrate for chlorophyll or Heme biosynthetic pathway (Wilde et al. 2004; Masuda and Fujita 2008). Although the function of GUN4 and their precursors- and- products for both retrograde signaling and chlorophyll synthesis has been previously demonstrated in Chlamydomonas and Arabidopsis (Koussevitzky et al. 2007; Czarnecki et al. 2012; Formighieri et al. 2012; Zhou et al. 2012), the mechanisms by which GUN4 are involved in retrograde signaling and the impact of GUN4 mutation on photosynthetic process remains poorly understood (Brzezowski et al. 2014).
Schematic representation of a summarized tetrapyrrole and chlorophyll biosynthetic pathway showing the local of GUN4 mutation. Glu l-glutamate, ALA 5-aminolevulinic acid, ProtoIX nprotoporphyrin IX, MgChl magnesium chelatase, FeChl ferrochelatase, Mg-ProtoIX Mg-protoporphyrin IX, Mg-Proto IX ME Mg-protoporphyrin IX monomethyl ester, Divinyl ProtoChlP divinyl Protochlorophyllide, Mono ChlP monovinyl protohlorophylide; Chl a––Chlorophyll a; Heme––heme groups
Pathways that communicate organelles (plastids and mitochondria) to nucleus are important once coordinate the development and function of the organelles with the whole plant cell (Pfannschmidt 2010). The communication between chloroplast and nucleus involves the exportation of signaling factors from chloroplast to cytosol where the signaling pathway transduces the signal to the nucleus (Kleine et al. 2009). It has been reported that changes in the chloroplast redox potential, the accumulation of reactive oxygen species, Mg-ProtoIX as well as 5-aminolevulinic acid (ALA) biosynthesis could be involved in the retrograde signaling from chloroplast to nucleus (Nott et al. 2006; Zhang 2007; Larkin and Ruckle 2008; Czarnecki et al. 2012). The accumulation of Mg-ProtoIX was reported as a possible signal from chloroplast that regulate light harvesting chlorophyll binding protein-II (LHCBII) genes in the nucleus (Strand et al. 2003; Larkin et al. 2003; Koussevitzky et al. 2007; Larkin and Ruckle 2008). However, strongly evidences suggest that this retrograde signaling cannot be related to Mg-ProtoIX accumulation, indicating that other unknown signal is involved in the chloroplast to nucleus signaling (Mochizuki et al. 2008; Moulin et al. 2008). The current idea is that there is no only one signal in retrograde signaling but rather the result of continuously sensed metabolic and genetic activities in the organelles (Czarnecki et al. 2012). Given these controversies concerning retrograde signaling a more carefully characterization of gun mutants are needed to understand the function of these genes in retrograde signaling and photosynthetic process, especially under light stress conditions.
The maintenance of plants growth under excessive amounts of light has been reported mainly as result of increased photosynthetic capacity of the plant and/or through the activation of process that dissipate the excess of energy absorbed in order to reduce the susceptibility of photo-oxidative damage (Apel and Hirt 2004; Walters 2005; Li et al. 2009). In this context, mutations in GUN genes can induce the overexpression of light harvesting chlorophyll binding (LHCB) proteins especially under stress conditions (Mochizuki et al. 2001; Peter and Grimm 2009; Brzezowski et al. 2014). LHCB are important proteins related to the energy transfer from light harvest complex (LHC) of photosystem II (PSII) to the reaction center of PSII (Teramoto et al. 2001). Given the importance of GUN4 for biosynthesis of photosynthetic pigments and the strong connection between GUN genes and LHCB proteins it’s surprisingly that the effect of GUN4 mutation on light energy transfer in LHCII and plant photochemical performance is still unclear. Therefore, we carried out a photosynthetic characterization of gun4 mutant in order to investigate the role of GUN4 gene for the plant photosynthetic performance under light stress conditions.
2 Materials and methods
2.1 Plant material and growth conditions
All Arabidopsis thaliana plants used in this study were of the columbia ecotype (Col-0). The gun4 knockout mutant (Gene ID: AT3G59400) (SALK_011461) were previously described (Mochizuki et al. 2001; Larkin et al. 2003). Seeds of wild type (WT) and gun4 mutant (homozygous) were surface-decontaminated by shaking 70 % ethanol for 5 min, rinsed with sterile distilled H2O. In sequence treated with 100 % ethanol for 1 min, rinsed three times with sterile distilled H2O. The seeds were then allowed to germinate in Petri dishes containing MS medium (25 cm3) (Murashige and Skoog 1962) and cultivated under photoperiod of 14 h illumination and light intensity of 120 µmol m−2 s−1, with day/night temperatures of 20 ± 3 °C. After 7 days the plants were transferred to recipient (0.1 L) containing commercial soil mixture, keeping in the room with the same conditions described above. We carried out a photosynthetic curve under different light intensities in completely expanded leaves of WT Arabidopsis plants in order to choose the level of light stress would be imposed to the plants. After this, five-week-old plants were subjected to light stress for 0, 14 or 28 h under 800 µmol photons m−2 s−1 using white lamps of 1,000 W in ambient with controlled temperature. It was disposed a water layer among the plants and light source in order to avoid heat excess. Unless otherwise described the fully expanded 9th–12th rosette leaves of six-week-old plants were harvested for subsequent analysis.
2.2 Gas exchange analysis and chlorophyll a fluorescence measurements
The gas exchange analysis were carried out in completely expanded leaves of WT and gun4 mutant using infra-red gas analyzer (IRGA, Licor 6400, Lincoln, USA) according Flexas et al. (2007). It was estimated net photosynthetic rate (A, µmol CO2 m−2 s−1), stomatal conductance (g s, mol H2O m−2 s−1), substomatal CO2 concentration (C i µmol CO2 mol−1), and transpiration rate (E, mmol m−2 s−1) at 0 and 14 h of light stress.
The chlorophyll a fluorescence measurements were performed in completely expanded leaf of WT and gun4 mutant by modulated light fluorometer system (MINI-PAM, Walz, Effeltrich, Germany). For established fluorescence parameters, initially measured minimal (Fo) and maximal fluorescence (Fm) in dark adapted leaves. It was estimated maximal quantum yield of PSII (Fv/Fm) according to Kitajima and Butler (1975). After, it was measured the parameters of light adapted leaves which was used to estimate effective quantum yield of PSII (ΦPSII) (Genty et al. 1989), photochemical quenching of lake model (qL) (qL = (Fm′ − F) × Fo′/(Fm′ − Fo′) × F) (Kramer et al. 2004), non-photochemical quenching (qN = (Fm − Fm′)/(Fm − Fo′) (Bilger and Schreiber 1986), quantum yield of non-regulated non-photochemical energy loss in PSII (ΦNO = F/Fm), quantum yield of regulated non-photochemical energy loss in PSII (ΦNPQ = (F/Fm′) − (F/Fm)). The parameters ΦPSII, ΦNO, and ΦNPQ were estimated according Hendrickson et al. (2004) in which ΦPSII + ΦNO + ΦNPQ = 1. The chlorophyll a fluorescence parameters were also determined in light responses curves. The parameters were estimated from different light intensities (0, 80, 160, 260, 370, 560, 800, 1,060 e 1,660 μmol of photons m−2 s−1).
2.3 Determination of photosynthetic pigments
Immediately after stress treatment, the rosettes of Arabidopsis were frozen at liquid N2 and used for determination of photosynthetic pigments. Total chlorophylls and carotenoids were extracted in acetone:water 80 % (v/v) and their concentration determined spectrophotometrically following Lichtenthaler (1987).
2.4 14CO2 feeding experiment
The redistribution of the photosynthetic metabolic flux was determined in leaves exposed to 14CO2. The 14CO2 was imposed to leaves using oxygen electrode (Hansatech, Kings Lynn, UK) under 400 μmol photons m−2 s−1, at 25 °C, for 60 min. The preparation of the radioactive solution containing 14C as well as the exposition to leaves was realized according DaMatta et al. (2008). After the exposition, leaves were frozen in liquid N2 and store at −80 °C. The extractions as well as the flux distribution between starch, cell wall, basic amino acids, organic acids, and sugars were realized according DaMatta et al. (2008). The incorporated radioactivity was determined using a liquid scintillation analyzer (Beckman LS 6500; Beckman Instruments, Fullerton, CA, EUA).
2.5 Co-expression analysis
The analyses of the co-expression network were performed using microarray data from on line platform ATTED-II (http://atted.jp/) according Obayashi et al. (2011). The network of GUN4 (At3g59400) co-expressed genes was obtained using CoexViewer tool of ATTED-II 5.2 according Obayashi et al. (2009). The data are available in http://atted.jp/top_draw.shtml#CoexViewer. A complete list of genes co-expressed with GUN4 gene is available in ATTED-II on line platform (http://atted.jp/). Further information’s about the significance of the mutual rank values for the gene co-expression can be found in Obayashi and Kinoshita (2009).
2.6 Statistical analysis
To analyze the effect of light stress in each genotype, the data were statistically compared by ANOVA and Dunnet test at 5 % of significance (P < 0.05). Between genotypes, in the same stress treatment, the data were statistically compared by Student t test (P < 0.05). All statistical analyses were performed using the Saeg system version 9.1. The number of replicates (n) is indicated in each figure.
3 Results
3.1 Photosynthetic pigments content are reduced in gun4 plants
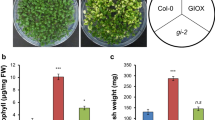
The gun4 mutant showed a phenotype with light green/yellowish leaves (Figure S1) according with lower chlorophyll and carotenoids content in the absence of light stress (Table 1). After 14 h of light stress, no significant differences in pigments content were observed in each genotype. However, the gun4 mutant still showed lower total chlorophyll, chlorophyll b and carotenoids content than WT (Table 1).
Light stress has no effect on carotenoids content in both genotypes. However, the chlorophyll content was significantly reduced in both genotypes after 28 h of light stress, except to chlorophyll b content in gun4 mutant (Table 1).
3.2 GUN4 is co-expressed in a network composed by photosystem II related proteins
We carried out an in silico analysis of GUN4 (At3g59400) gene expression from microarray data available using different web platforms. The Arabidopsis eFP Browser on-line platform (Winter et al. 2007) showed that GUN4 is preferentially expressed in photosynthetic tissues, with high expression in chloroplast of leaf mesophyll cells (data not show). The co-expression network analysis by web platform ATTED-II (Obayashi et al. 2011) showed that there is a high co-expression between GUN4 and genes related to photosynthesis, structure and function of both photosystems I and II, proteins of light harvest complex as well as proteins of chlorophyll biosynthetic pathway (Fig. 2). Among different genes co-expressed with GUN4, highlights to genes of light harvesting chlorophyll binding proteins (LHCB2.3, LHCB3, LHCB4, LHCB4.2, LHCB5, LHCB6, LHCA1, LHCA3), genome uncoupled 5 gene (GUN5), non-photochemical quenching related protein (NPQ4), and subunits of photosystems I and II (PSII protein, PSAD-1, PSAD-2, PSAH1, PSAH2, PSBTN, PSAO, PSAF, PSBO2, PSB28, PSBX, PSBW, PSAL, PASK) (Table S1).
3.3 gun4 plants has greater photochemical efficiency in photosystem II under non-stress conditions
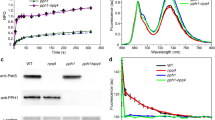
The gun4 mutant showed reduced minimal (Fo) and maximal (Fm) fluorescence of dark-adapted leaves (Fig. 3a, b) as well as reduced quantum yield of unregulated energy dissipation of photosystem II (ΦNO) (data not shown) under non-stress conditions. However, gun4 plants showed increases in non-photochemical quenching (qN) (Fig. 3c), effective quantum yield of PSII (ΦPSII) (Fig. 3e), as well as in photochemical quenching (qL) (Fig. 3f). Higher values of ΦPSII and lower of ΦNO were also found in gun4 under different light intensities (Fig. 4).
Mean values and standard error of minimal (Fo) (a) and maximal (Fm) (b) fluorescence of dark-adapted leaves, non-photochemical quenching (qN) (c), potential quantum yield of the PSII (Fv/Fm) (d), effective quantum yield of the PSII (ΦPSII) (e), and photochemical quenching (qL) (f) in leaves of wild type (WT) and gun4 mutant under different light stress condition. Values followed by the same letter in each genotype and different light stress conditions are not statistically different by Dunnet test (P < 0.05). Asterisks indicate statistic difference between genotypes in the same stress condition by Student t test (P < 0.05) (n = 5)
Mean values and standard error of unregulated energy dissipation of photosystem II (ΦNO) and photochemical quantum yield of PSII (ΦPSII) (b) in leaves of wild type (WT) and gun4 mutant under different light intensities. Asterisks indicate statistic difference between genotypes in the same light condition by Student t test (P < 0.05) (n = 6)
3.4 gun4 has greater non-photochemical dissipation of light excess
It was observed significant differences between WT and gun4 in qN (higher values in gun4) and Fo and Fm (lower values in gun4) independent of the stress level (Fig. 3a–c). Increased qN was observed in both genotypes after 14 h of light stress. However, only WT showed significant increases in Fo and significant reduction in potential quantum yield of the PSII (Fv/Fm) after 14 h of stress (Fig. 3a, d). After 28 h of light stress, all photochemical parameters analyzed were significantly different of the non-stress condition, with exception for qN in WT. However, Fo, Fm, and qN were significantly different between WT and gun4 mutant. After this severe stress, the Fv/Fm values were significantly reduced in both genotypes, but higher decrease was observed in WT (74 %) compared to gun4 mutant (54 %) (Fig. 3 D). The gun4 mutant maintains non-photochemical dissipation of light excess through a significant higher qN under light stress (Fig. 3c). Furthermore, there was no difference in Fo between 14 and 28 h of light stress in gun4 plants (Fig. 3a).
3.5 gun4 plants has intrinsic water use efficiency increased despite strong reduction in photosynthetic rates
The gun4 mutant showed decreased net photosynthetic rate (A) (43 %), stomatal conductance (g s) (61 %), and transpiration rate (E) (46 %) compared to WT plants. These reductions lead to a phenotype with increased intrinsic water use efficiency (WUEi), i.e. higher A/g s relationship (Fig. 5). Due to the mild reductions in A and E in WT after light stress, no differences were observed in these parameters between WT and gun4 plants. However, g s was still lower in gun4 after light stress which leads to a higher A/g s relationship in gun4 (Fig. 5). Given the strong light stress imposed on plants whereas that 28 h of light stress dramatically reduced all photochemical parameters (Fv/Fm, ΦPSII, and qL), no net photosynthetic rate was detected in leaves of both genotypes after 28 h of light stress (data not shown).
Mean values and standard error of gas exchange parameters in leaves of wild type (WT) and gun4 mutant submitted to 0 or 14 h of light stress. A net photosynthetic rate, g s stomatal conductance, E transpiration rate, WUE i intrinsic water use efficiency. Asterisks indicate statistic difference between genotypes in the same stress condition by Student t test (P < 0.05) (n = 5)
3.6 14CO2 incorporation in WT and gun4 leaves
We carried out a feeding experiment in whole leaves submitted to saturate 14CO2 concentration. Under these conditions, total 14C incorporation was not different between WT and gun4 mutant (Table 2). No differences were found in the percentage of 14C flux redistribution to basic amino acids, sugars, starch, and cell wall. However, mild reduction in the percentage of 14C to organic acids was observed in gun4 mutant (Table 2).
4 Discussion
4.1 gun4 has greater light energy transfer efficiency in PSII despite lower both chlorophyll and carotenoid contents
Given the role of GUN4 for chlorophyll biosynthesis, the mutation in this gene leads to reduced level of photosynthetic pigments in Arabidopsis leaves. In turn it may lead to small LHCII and reduced capacity of light energy absorption in gun4 plants, evidenced by the lower Fo values compared to WT. There are evidences in plants and cyanobacteria that the mutation in genes of chlorophyll biosynthetic pathway, including GUN4, result in smaller LHCII due the less components of light harvesting complex and less subunits of PSII (Havaux et al. 2007; Sobotka et al. 2008; Hansson and Jensen 2009). In this context, gun4 phenotype is similar to other mutants of chlorophyll biosynthetic pathway, such as chl27 and sg1. CHL27 codifies the enzyme Mg-ProtoIX monomethyl ester ciclase (EC 1.14.13.81) (Fig. 1). The chl27 mutant has less chlorophyll content and reduced both LHCI (30 % of reduction) and LHCII (50 % of reduction) content. However, the effective quantum yield of both PSI and PSII increase in chl27 (Hansson and Jensen 2009). The photochemical efficiency of chl27 mutant is greater than WT a despite of minor capacity of light absorption, similar to observed here in gun4 mutant. Interestingly, gun4 and chl27 plants showed a flexibility of PSII to increase the capacity of energy transfer despite of fewer molecules for light absorption.
Light stress has been reported as signal to reduce the expression of nuclear genes that codify light-harvesting chlorophyll a/b-binding proteins (LHCB) via a pathway that involve GUN4 and ABI4 transcription factor (abscisic acid-insensitive 4) (Larkin et al. 2003; Larkin and Ruckle 2008). Probably this is a photo-protection response in order to reduce light excess absorption in PSII and minimize oxidative damages caused by excess of light in WT plants. Furthermore, gun mutants exhibited high-level expression of LHCB genes under stress (Susek et al. 1993; Mochizuki et al. 2001; Eberhard et al. 2008; Peter and Grimm 2009; Brzezowski et al. 2014). It suggests that the expression of GUN and LHCB genes are highly coordinated. Indeed, the analysis of GUN4 co-expression network showed that many genes related to photosynthesis and structure of photosystems are co-expressed with GUN4, including many genes of LHCB family (Table S1). Similarly, the single sg1 mutant and the double sg1 gun4 mutant presented low levels of chlorophyll but high levels of genes related to photosystem II (psbA, psbB) and carbon fixation (RbcS, RbcL) (Hu et al. 2014). It suggests that GUN4, like SG1, is important not only for chlorophyll biosynthesis but also for expression of genes related to the photosynthetic process; which was indeed showed in this work. The up-regulation of LHCB genes in gun4 mutant under stress may optimize the light energy transfer from LHCII to reaction center of PSII, contributing to the greater light energy transfer efficiency in LHCII/PSII despite low chlorophyll and carotenoids content. Furthermore, in the absence of light stress, gun4 plants showed lower values of ΦNO than WT. This reduction in constitutive loss of energy can be a flexibility of gun4 LHCII to maximize the energy transfer to reaction center of PSII, contributing for the greater photochemical capacity of gun4 plants.
4.2 GUN4 mutation minimize the effects of light stress
The decrease in Fv/Fm and the increase in Fo in both genotypes after 28 h of light stress suggest the occurrence of photoinhibition on photochemical apparatus (Maxwell and Johnson 2000; Müller et al. 2001). The increases in Fo can be related to damages in the reaction center of PSII and/or in the LHCII (Demmig-Adams and Adams 1992; Bolhàr-Nordenkampf and Öquist 1993). Damages on PSII light stress-induced can also be associated with increases in unregulated energy dissipation of PSII (ΦNO) (Klughammer and Schreiber 2008). Indeed it was observed increases in ΦNO in both genotypes after 14 h of light stress (data not shown). It is important to highlight that although light stress was sufficient to induce damages on PSII of both WT and gun4 mutant, the decreases observed in Fv/Fm and the increases in Fo after severe light stress were minor in gun4 plants. Furthermore, WT plants were unable to keep a high qN value after 28 h of light stress as observed in gun4 plants. Thus, considering that gun4 mutant absorbs less light energy (lower Fo values) and has greater capacity to dissipate energy (higher qN values), it suggests that the oxidative damage on PSII light stress-induced is minor in gun4 plants. However, the mechanism by which gun4 plants dissipate more efficiently the excess of energy absorbed is still unknown.
4.3 Total 14CO2 leaf absorption suggests a diffusive limitation for photosynthesis in gun4 mutant
Although gun4 plants showed lower values of stomatal conductance (g s) and net photosynthetic rate (A) compared to WT in the absence of stress, no differences were detected between the genotypes in the total 14CO2 fixation as well as in starch fraction after 14CO2 fixation, a photosynthetic product accumulated in the light (Zeeman et al. 2007). It suggests that the lower capacity for CO2 fixation observed in gun4 plants in absence of stress can be related to the diffusional limitation of CO2 rather than a biochemical limitation, once the feeding experiment was carried out under saturating levels of CO2 where stomata present no barrier to CO2 assimilation.
5 Conclusions
Taken together, the results of this work showed that GUN4 mutation have high impact on gas exchange through the leaves, photosynthetic pigment contents, and on light energy transfer capacity of the PSII. It suggests that this gene is important not only for the chlorophyll biosynthesis but also for process that regulates the transfer of energy from LHCII to reaction center of PSII. Furthermore, because of the phenotype displayed by gun4 mutant differs from other gun mutants, it indicates that gun4 phenotype cannot be explained solely by the reduction in chlorophyll content. The flexibility of gun4 PSII to maximize light energy transfer reflects an acclimation of LHCII in order to compensate the reduced ability of light absorption. It highlights the importance of gun4 mutant to investigate the mechanisms that control LHCII plasticity, an important aspect to understand plant light stress tolerance.
Abbreviations
- ΦPSII :
-
Effective quantum yield of PSII
- GUN:
-
Genome uncoupled
- gun4 :
-
Genome uncoupled 4 mutant
- LHCB:
-
Light harvesting chlorophyll binding proteins
- LHC:
-
Light harvesting complex
- LHCII:
-
Light harvesting complex of the complex II
- Fm :
-
Maximal fluorescence in dark adapted leaves
- Fo :
-
Minimal fluorescence in dark adapted leaves
- MgChl:
-
Mg-chelatase enzyme
- Mg-ProtoIX:
-
Mg-protoporphyrinIX
- A :
-
Net photosynthetic rate
- qN:
-
Non-photochemical quenching
- ΦP :
-
Photochemical quantum yield of PSII
- qL:
-
Photochemical quenching of lake model
- PSII:
-
Photosystem II
- Fv/Fm :
-
Potential quantum yield of PSII
- ProtoIX:
-
ProtoporphyrinIX
- ΦNO :
-
Quantum yield of non-regulated non-photochemical energy loss in PSII
- g s :
-
Stomatal conductance
- C i :
-
Substomatal CO2 concentration
- E :
-
Transpiration
- WT:
-
Wild type
References
Adhikari ND, Orler R, Chory J, Froehlich JE, Larkin RM (2009) Porphyrins promote the association of GENOMES UNCOUPLED 4 and Mg-chelatase subunit with chloroplast membranes. J Biolog Chem 284(37):24783–24796
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Bilger W, Schreiber U (1986) Energy-dependent quenching of dark level chlorophyll fluorescence in intact leaves. Photosynth Res 10:303–308
Bolhàr-Nordenkampf HR, Öquist GO (1993) Chlorophyll fluorescence as a tool in photosynthesis research. In: Hall DO, Scurlock JMO, Bolhàr-Nordenkampf HR, Leegood RC, Long SP (eds) Photosynthesis and production in a changing environment. Chapman & Hall, London, pp 193–205
Brzezowski P, Schlicke H, Richter A, Dent RM, Niyogi KK, Grimm B (2014) The GUN4 protein plays a regulatory role in tetrapyrrole biosynthesis and chloroplast-tonucleus signalling in Chlamydomonas reinhardtii. Plant J. doi:10.1111/tpj.12560
Czarnecki O, Gläßer C, Chen JG, Mayer KFX, Grimm B (2012) Evidence for a contribution of ALA synthesis to plastid-to-nucleus signaling. Front Plant Sci 3:1–19
DaMatta FM, Cunha RL, Antunes WC, Martins SCV, Araujo WL, Fernie AR, Moraes GABK (2008) In field-grown coffee trees source–sink manipulation alters photosynthetic rates, independently of carbon metabolism, via alterations in stomatal function. New Phytol 178:348–357
Demmig-Adams B, Adams WW (1992) Photoprotection and other responses of plants to high light stress. Annu Rev Plant Physiol Plant Mol Biol 43:599–626
Eberhard S, Finazzi G, Wollman FA (2008) The dynamics of photosynthesis. Annu Rev Genet 42:463–515
Flexas J, Ortuño MF, Ribas-Carbo M, Diaz-Espejo A, Flórez-Sarasa ID, Medrano H (2007) Mesophyll conductance to CO2 in Arabidopsis thaliana. New Phytol 175:501–511
Formighieri C, Ceol M, Bonente G, Rochaix JD, Bassi R (2012) Retrograde signaling and photoprotection in a gun4 mutant of Chlamydomonas reinhardtii. Mol Plant 5(6):1242–1262
Genty B, Briantais JM, Baker NR (1989) The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochim Biophys Acta 990:87–92
Hansson A, Jensen PE (2009) Chlorophyll limitation in plants remodels and balances the photosynthetic apparatus by changing the accumulation of photosystems I and II through two different approaches. Physiol Plant 135:214–228
Havaux M, Dall’Osto L, Bassi R (2007) Zeaxanthin has enhanced antioxidant capacity with respect to all other xanthophylls in Arabidopsis leaves and functions independent of binding to PSII antennae. Plant Physiol 145:1506–1520
Hu Z, Xu F, Guan L, Qian P, Liu Y, Zhang H, Huang Y, Hou S (2014) The tetratricopeptide repeat-containing protein slow green1 is required for chloroplast development in Arabidopsis. J Exp Bot 65(4):1111–1123
Kitajima M, Butler WL (1975) Quenching of chlorophyll fluorescence and primary photochemistry in chloroplasts by dibromothymoquinone. Biochim Biophys Acta 376:105–115
Kleine T, Voigt C, Leister D (2009) Plastid signalling to the nucleus: messengers still lost in the mists? Trends Genet 25(4):185–192
Klughammer C, Schreiber U (2008) Complementary PSII quantum yield calculated from simple fluorescence parameters measured by PAM fluorometry and saturation pulse method. PAM Appl Notes 1:27–35
Koussevitzky S, Nott TC, Mockler FH, Sachetto-Martins G, Surpin MJ, Chory J (2007) Multiple signals from damaged chloroplasts converge on a common pathway to regulate nuclear gene expression. Science 316:715–719
Kramer DM, Johnson G, Kiirats O, Edwards GE (2004) New fluorescence parameters for the determination of QA redox state and excitation energy fluxes. Photosynth Res 79:209–218
Larkin RM, Ruckle ME (2008) Integration of light and plastid signals. Curr Opin Plant Biol 11:1–7
Larkin RM, Alonso JM, Ecker JR, Chory J (2003) GUN4, a regulator of chlorophyll synthesis and intracellular signaling. Science 299:902–906
Li Z, Wakao S, Fisher BB, Niyogi KK (2009) Sensing and responding to excess light. Annu Rev Plant Biol 60:239–260
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzimol 148:350–382
Masuda T, Fujita Y (2008) Regulation and evolution of chlorophyll metabolism. Photochem Photobiol Sci 7:1131–1149
Maxwell K, Johnson GN (2000) Chlorophyll fluorescence-a practical guide. J Exp Bot 51:659–668
Mochizuki N, Brusslan JÁ, Larkin R, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelataseH subunit in plastid-nucleus signal transduction. Proc Natl Acad Sci USA 98:2053–2058
Mochizuki N, Tanaka R, Tanaka A, Masuda T, Nagatani A (2008) The steady state level of Mg-protoporphyrin IX is not a determinant of plastid-to-nucleus signaling in Arabidopsis. Proc Natl Acad Sci USA 105:15184–15189
Moulin M, McCormac AC, Terry MJ, Smith AG (2008) Tetrapyrrole profiling in Arabidopsis seedlings reveals that retrograde plastid nuclear signaling is not due to Mg-protoporphyrin IX accumulation. Proc Natl Acad Sci USA 105:15178–15183
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nott A, Jung HS, Koussevitzky S, Chory J (2006) Plastid-to-nucleus retrograde signalling. Annu Rev Plant Biol 57:739–759
Obayashi T, Kinoshita K (2009) Rank of correlation coefficient as a comparable measure for biological significance of gene coexpression. DNA Res 16(2009):249–260
Obayashi T, Hayashi S, Saeki M, Ohta H, Kinoshita K (2009) ATTED-II provides coexpressed gene networks for Arabidopsis. Nucleic Acids Res 37:D987–D991
Obayashi T, Nishida K, Kasahara K, Kinoshita K (2011) ATTED-II updates: condition-specific gene coexpression to extend coexpression analyses and applications to a broad range of flowering plants. Plant Cell Physiol 52:213–219
Peter E, Grimm B (2009) GUN4 is required for posttranslational control of plant tetrapyrrole biosynthesis. Mol Plant 2(6):1198–1210
Pfannschmidt T (2010) Plastidial retrograde signalling––a true ‘‘plastid factor’’ or just metabolite signatures? Trends Plant Sci 15:427–435
Sobotka R, Dühring U, Komenda J, Peter E, Gardian Z, Tichy M, Grimm B, Wilde A (2008) Assembly of photosynthetic complexesprotein for chlorophyll metabolism and importance of the cyanobacterial Gun4. J Biol Chem 283:25794–25802
Susek RE, Chory J (1992) A tale of two genomes: role of a chloroplast signal in coordinating nuclear and plastid genome expression. Aust J Plant Physiol 19:387–399
Susek RE, Ausubel FM, Chory J (1993) Signal transduction mutants of Arabidopsis uncouple nuclear CAB and RBCS gene expression from chloroplast development. Cell 74:787–799
Tanaka R, Tanaka A (2007) Tetrapyrrole biosynthesis in higher plants. Annu Rev Plant Biol 58:321–346
Teramoto H, Ono T, Minagawa J (2001) Identification of Lhcb gene family encoding the light-harvesting chlorophyll-a/b proteins of photosystem II in Chlamydomonas reinhardtii. Plant Cell Physiol 42:849–856
Walters RG (2005) Towards an understanding of photosynthetic acclimation. J Exp Bot 56(411):435–447
Wilde A, Mikolajczyk S, Alawady A, Lokstein H, Grimm B (2004) The gun4 gene is essential for cyanobacterial porphyrin metabolism. FEBS Lett 571:119–123
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV et al (2007) An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS One 2(8):e718
Zeeman SC, Smith SM, Smith AM (2007) The diurnal metabolism of leaf starch. Biochem J 401:13–28
Zhang DP (2007) Signaling to the nucleus with a loaded GUN. Science 316:700–701
Zhou S, Sawicki A, Willows RD, Luo M (2012) C-terminal residues of Oryza sativa GUN4 are required for the activation of the ChlH subunit of magnesium chelatase in chlorophyll synthesis. FEBS Lett 586:205–210
Acknowledgments
This research was supported by the CNPq (National Council for Technological and Scientific Development, Brazil) and FAPEMIG (Foundation for Research Assistance of the Minas Gerais State, Brazil). Scholarships granted by CNPq, CAPES and FAPEMIG are gratefully acknowledged. DMD was recipient of a scholarship from CAPES.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Daloso, D.M., Antunes, W.C., Santana, T.A. et al. Arabidopsis gun4 mutant have greater light energy transfer efficiency in photosystem II despite low chlorophyll content. Theor. Exp. Plant Physiol. 26, 177–187 (2014). https://doi.org/10.1007/s40626-014-0025-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40626-014-0025-z