Abstract
Purpose of Review
Over recent decades, the number of outbreaks caused by fungi has increased for humans, plants (including important crop species) and animals. Yet this problem is compounded by emerging antifungal drug resistance in pathogenic species. Resistance develops over time when fungi are exposed to drugs either in the patient or in the environment.
Recent Findings
Novel resistant variants of fungal pathogens that were previously susceptible are evolving (such as Aspergillus fumigatus) as well as newly emerging fungal species that are displaying antifungal resistance profiles (e.g. Candida auris and Trichophyton indotineae).
Summary
This review highlights the important topic of emerging antifungal resistance in fungal pathogens and how it evolved, as well as how this relates to a growing public health burden.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungi encapsulate a wide range of diversity, with an estimated 3.8 million species within this kingdom [1] and are currently classified into eight phyla: zoosporic fungi (Cryptomycota, Chytridiomycota and Blastocladiomycota), Microsporidia, Zoopagomycota, Mucoromycota and Dikarya fungi (Ascomycota and Basidiomycota). Reports of emerging pathogens in all phyla are on the increase, due to altered pathogenicity or availability of new biological niches [2]. In addition, there are increased reports of antifungal drug resistance amongst emerged human fungal pathogens and is often found in emerging fungal pathogens. For example, Cryptococcus gattii has experienced a shift in species distribution to the Pacific Northwest (PNW) [3], and more recently, Candida auris has emerged as an important nosocomial infection [4] on all continents except Antarctica [5]. Fungi demonstrate complex life cycles, enabling genomes to undergo recombination in sexual and parasexual life cycles, and clonal propagation during asexual reproduction. This results in novel genetic diversity and expansion of pathogenic lineages capable of overcoming host defences, survival in non-endemic regions and drug resistance.
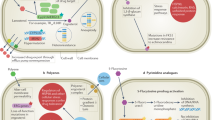
The overall burden of fungal disease equates to more than a billion people worldwide [6], causing equally diverse diseases, from superficial infections and allergic syndromes to life-threatening invasive diseases. Treatment of disease is fought with four antifungal drug classes in our arsenal: azoles, echinocandins, polyenes and pyrimidine analogues. Yet clinicians and mycologists alike may say we are losing the battle, with rising drug resistance and tolerance causing treatment failure [7]. The increasing public health threat posed by resistance and treatment failure has been recognised by the World Health Organisation (WHO) [8••] and the US Centers for Disease Control and Prevention (CDC) [9]. A Global Action Plan [10] to address antimicrobial resistance has been endorsed by the WHO and stresses the importance of a One Health approach to tackle it. This plan has called for coordination between human and veterinary medicine, as well as agricultural, environmental and financial sectors, and has certainly helped raise antifungal drug resistance to the top of the global public health agenda.
In this review, we focus on emerging drug-resistant fungal pathogens as well as previously susceptible fungi that have recently evolved resistance and the potential contributors to this evolution.
Emerging Antifungal Drug Resistance in Previously Susceptible Pathogens
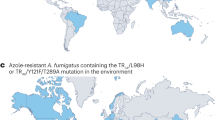
Whilst present in a variety of environments, including soil, compost and decaying organic matter, the filamentous mould Aspergillus fumigatus can also cause acute and chronic forms of invasive aspergillosis in immunocompromised individuals [11]. With over 300,000 life-threatening infections in susceptible patient cohorts each year, azole antifungal drugs are the most effective treatment [12]. However, over the last two decades, clinicians have seen a rise in the prevalence of azole-resistant A. fumigatus in patients, mirroring a rise in azole resistance environmentally [13]. Whilst initially responsive to azole antifungal drug therapy, susceptibility has given way to resistance.
It is thought that at least two-thirds of patients with azole-resistant infections have not received azole drug therapy [14], suggesting an environmental route of acquisition [15, 16]. The most prominent set of polymorphisms conferring resistance to azoles is the 34 base pair (bp) tandem repeat and L98H amino acid substitution in cyp51a, TR34/L98H (Table 1), which has been recovered both within clinical samples and environmentally. Much research has been conducted to establish whether azole-resistant A. fumigatus has an agricultural origin [17, 18] or evolved independently in the clinic [19].
If azole-resistant A. fumigatus isolates originated in the environment and were subsequently acquired by patients, it would be logical to assume clinical isolates would show resistance to agricultural fungicides. Kang et al. showed clinical A. fumigatus isolates carry alleles conferring resistance to agricultural fungicides such as benzimidazoles (MBCs) and quinone outside inhibitors (QoIs) [20••]; this subsequently provides strong evidence for an agricultural origin of isolates acquired by patients. This finding was quickly followed by a study by Rhodes et al. that showed multiple clinical isolates with azole drug-resistant genotypes that matched environmental isolates with very high similarity [21••]. Since no evidence has been presented to convincingly show patients transmit A. fumigatus into the environment, it is further evidence for the acquisition of azole-resistant isolates by at-risk patients from the environment. Other fungal diseases have been shown to be acquired from the environment, such as Coccidioides spp. [22] and Cryptococcus spp. [23], further bolstering support for this argument.
A. fumigatus not only represents a human public health threat, but also a threat to wildlife and conservation efforts. Recently, a single strain of A. fumigatus was the cause of a mass aspergillosis outbreak in an endangered parrot species [24]. Despite not being drug-resistant, the outbreak still resulted in 21 infections and 9 deaths, in a population of only 211. A known opportunistic pathogen of penguins [25], azole drug-resistant A. fumigatus has been isolated from a zoo environment and from Humboldt penguins themselves [26]. However, little research is dedicated to the impact of aspergillosis on wildlife species in terms of clinical outcome and treatment failure, certainly highlighting a gap in our current knowledge.
Given the above evidence, it is likely that resistance has emerged in the previously susceptible A. fumigatus due to the dual use of azoles in the environment and the clinic, driving the evolution of antifungal resistance [2]. The TR34/L98H resistance allele, along with TR46/Y121F/T289A (Table 1), is associated with itraconazole and voriconazole resistance respectively and has been isolated within and outside the clinic [27]. A contributing factor for the generation of novel drug resistance alleles could be attributed to the high recombination rate observed in A. fumigatus [28]; a recent example of a pan-azole resistance phenotype, attributed to TR34/L98H/T289A/I364V/G448S, could be the result of such recombination [29]. Thus, the pandemic potential of drug-resistant A. fumigatus has already been demonstrated, primarily due to human activity by using clinical antifungals in horticultural products [30], and global spread is likely due to ease of dispersal via conidia on air currents [2].
Emerging Drug-Resistant Fungal Pathogens
We have recently witnessed entirely new fungal species emerge and spread globally, already resistant to multiple antifungal drugs. Candida auris has received much public attention and scientific research focus and was recently listed as a “critical priority” fungal pathogen in the inaugural WHO Fungal Priority Pathogen List (FPPL) [8••]. Fungicide exposure and climate change have been hypothesised as factors explaining the rapid worldwide emergence of C. auris, which displays high intrinsic levels of fluconazole resistance and varying levels of resistance to other antifungal drugs [31].
First described in 2009 in Japan [4], C. auris has since been reported in over 40 countries [32]. Initial genomic analyses revealed four geographically restricted clades emerging almost simultaneously, I (South Asia), II (East Asia), III (South Africa) and IV (South America) [33], which have since been found elsewhere in the world. Interestingly, isolates from Clade II do not tend to cause invasive infections and are susceptible to antifungal agents. In recent years, a further two new clades, V from Iran [34, 35] and VI from Singapore [36], have also been described. Yet unlike A. fumgiatus, C. auris has emerged with intrinsic fluconazole antifungal drug resistance [37] and has since evolved drug resistance to other azoles and drug classes [38••].
C. auris possesses different characteristics to set it apart from other Candida species, such as a lack of morphological switching between yeast and hyphal forms [39, 40], which may explain colonisation of distinct niches. For example, C. auris primarily colonises the skin, whereas C. albicans colonises and infects mucosal surfaces [37]. Antifungal drug tolerance is attributed to biofilm formation in Candida species due to drug sequestration by the biofilm mannan-glucan polysaccharide matrix [41] and plays a key role in intrinsic resistance in C. auris [33, 42, 43]. Previous studies have shown that Clade II isolates seem to have evolved host adaptation and are particularly adapted to the ear, resulting in non-invasive clinical phenotypes that are drug-susceptible [44] and incapable of forming biofilms [45].
Whole genome sequencing has helped unravel the basis of the resistance mechanisms existing and evolving within C. auris (Table 1). The main cause of fluconazole resistance and therefore treatment failure is centred on amino acid substitutions within the ERG11 gene, which are also restricted by genetic clade: Y132F and K143R in Clade I, only Y132F in Clades IV and V and F126L in Clade III [33, 35]. Currently, no known mutations in ERG11 have been associated with Clade VI isolates, but given the susceptibility of all three identified isolates in this clade, it remains unlikely any will be found [36]. Similarly, given the drug susceptibility of Clade II isolates, no ERG11 mutations are associated with this clade [33]. Mutations within ERG11 account for the majority of azole drug resistance and were initially attributed as the causation of intrinsic fluconazole resistance. Mutations caused by amino acid substitutions have also been described in TAC1b [46], along with overexpression of CDR1 and MDR1 [47], but the necessary experiments to confirm these contributions have not yet been carried out in all cases.
Since the first genomic analyses were completed on C. auris in 2017 by Lockhart et al. [33], more than 5000 genomes have been made publicly available. Genomic data has allowed the mining of confirmed and probable mutations linked to antifungal drug resistance (Table 1). Of greatest abundance are the clade-specific ERG11 mutations conferring fluconazole resistance. Yet over time, mutations conferring resistance to other antifungal drug classes have been described: the first European outbreak detailed flucytosine resistance due to F211I in FUR1 and echinocandin resistance due to S639Y in FKS1 [48]. This later mutation is situated in the hotspot 1 (HS1) region of FKS1, and other amino acid substitutions (S639P and S639F) have been described at the same loci [49, 50]. Recently, pan-drug class resistance has been described in a single patient undergoing multi-visceral transplantation, with ERG11 K143R, FKS1 S639Y and a novel FUR1[1Δ33] mutation [38••]. This combination conferred resistance to azoles, echinocandins and flucytosine, respectively. In addition, resistance to amphotericin B was also observed, yet no distinct variants were identified, suggesting unknown genetic elements were driving resistance to this drug. The recent emergence of a C. auris isolate resistant to four classes of antifungal drugs is concerning; further research is needed to ascertain patient outcome consequences and the mechanism(s) behind novel drug resistance mutation evolution.
The question of how resistance emergences in C. auris has been explored through a combination of in vitro microevolution and in silico genomic approaches. Serial drug exposures have shown rapid adaptation leading to tolerance of echinocandins (FKS1 codon deletion and mutations in FKS1, ERG3 and CIS3), azoles (chromosome 5 aneuploidy, containing TAC1b, ERG11 contig duplication, mutations in TAC1b), polyenes (MEC3 mutations) and concurrently azoles and polyenes (ERG3/ERG11 nonsense mutations) [51]. Sub-telomeric deletions, aneuploidies and SNP acquisitions were also seen on fluconazole exposure with evidence of a potential hypermutator phenotype resulting in missense or nonsense impact mutations in one strain with an associated MLH1 variant (A216V) [52]. However, evidence for recombination in C. auris has not been convincingly demonstrated since major clades diverged, according to testing for phylogenetic incompatibility and linkage equilibrium testing [53]. The use of bioinformatic approaches in public repository data is limited by the availability of phenotypic data, which might be used in genome-wide association studies. One group used 356 strains to test several machine learning clustering methods to rank the contribution of specific mutations to resistance, including in several genes that have not been explored or described [54].
More recently, Trichophyton indotineae has emerged as a novel dermatophyte, posing a significant health threat in India due to its extensive infection and treatment failures [55, 56]. Fungal infections caused by dermatophytosis that affect the skin and nails and have a global prevalence of 20–25% [57]. Initially described as Trichophyton mentagrophytes genotype VIII, T. indotineae has recently been classified as a distinct species with unique characteristics, including high terbinafine (TERB) resistance and anthropophilic transmission [55].
T. indotineae causes highly inflammatory, often widespread, dermatophytosis affecting much of the lower body, groins, limbs and face [55]. The emergence and isolation of this species took place sequentially, starting in Australia in 2008, then spreading to Oman in 2010, and later to Iran in 2016 [55]. Around 2017–2018, a drug-resistant variant of T. indotineae emerged in India, setting off a global spread from the Indian subcontinent [55, 56]. Reported cases are seen in several countries, including India, Bahrain, United Arab Emirates, Oman, Iran, Germany, France, Italy, Belgium, Switzerland, Greece, Denmark, Finland, China, Australia, Canada, Japan, Vietnam and the United States of America [55, 58,59,60,61,62,63,64,65,66,67,68,69].
The development of resistance in T. indotineae is primarily linked to the excessive and improper use of over-the-counter corticosteroid-antifungal medications [70]. As such, T. indotineae now exhibits a high frequency of resistance to conventional antifungals, including allylamines (which includes TERB) and azoles (itraconazole and fluconazole) [71,72,73,74].
TERB interacts with squalene epoxidase (SQLE), a key enzyme encoded by the ERG1 gene, in which SQLE plays a pivotal role in initiating the biosynthesis of ergosterol [75, 76]. Mutations in the ERG1/SQLE gene at specific amino acid positions, such as Phe397Leu (F397L) (associated with minimum inhibitory concentration (MIC) values ranging ≥10 to ≥15 µg/mL) and Leu393Phe (L393F) (associated with MIC values in access of ≥40 µg/mL), have been identified to result in high MIC values to TERB [71]. Furthermore, when double mutations occur at Phe397Leu and Ala448Thr (F397L/A448T) within the ERG1 gene, they result in a combined resistance to TERB [71]. This leads to MIC values that are higher than those observed in wild-type strains or single mutants with the F397L single mutation (with MIC ranges typically falling between 15 to ≥30 µg/mL) [71].
Azoles exert their antifungal effects by binding to the lanosterol 14-α demethylase enzyme, encoded by the ERG11 gene [71]. These proteins are categorised under the heme-containing monooxygenases within the cytochrome P450 superfamily, CYP51 [71]. Thus, point mutations within the ERG11 gene lead to alterations in amino acid sequences at specific positions, ultimately conferring resistance to particular azole drugs [71, 77, 78]. However, no research has yet detected mutations within ERG11. However, the presence of identified tandem repeats in the promoter region and the overexpression of Cyp51B suggests the reduced sensitivity of T. indotineae strains to azoles [79]. Intriguingly, ERG1/SQLE mutants carrying the A448T mutation remain resistance-sensitive to TERB but, instead, show azole resistance patterns [71, 73, 80]. The underlying reason for this observed correlation remains undisclosed.
To better understand the TERB mechanisms of resistance in this newly emerging species T. indotineae, one study proposes that the diversity of resistance mechanisms may be linked to the TruMDR2 and salA genes, initially identified in other fungal species, thus suggesting promising directions for future research [81]. Our current understanding of T. indotineae’s resistance mechanisms is limited, but preliminary findings suggest that TruMDR2 encodes an ABC transporter known to be part of the multidrug-resistance class of transporters and has been shown to be responsive to various drugs in T. rubrum [82]. On the other hand, the salA gene, identified in UV-induced TERB-resistant mutants of Aspergillus nidulans, encodes a salicylate 1-monooxygenase and contributes to TERB resistance by conferring resistance upon transformation [83]. Hence, the study suggests a similar role in modulating TERB susceptibility in Trichophyton rubrum due to its potential involvement in naphthalene ring degradation mechanisms [83]. Further exploration of these genes in T. indotineae could provide valuable insights into its unique resistance mechanisms.
Contributions to the Development of Drug Resistance
Certain fungal species, such as Candida spp., exhibit intrinsic drug resistance, yet acquired resistance is also increasing in prevalence. Selection for drug resistance in T. indotineae has been attributed to the ease of availability of over-the-counter medications and incomplete treatment courses. The use of low-dosage exposure along with incomplete coverage of topical creams has also been suggested as a contributing factor [84]. Studies have also confirmed the acquisition of fungi from the environment, such as A. fumigatus [20••, 21••], Cryptococcus spp. [23] and Coccidioides [22]; it can therefore be assumed that environmental exposure to antifungal drugs could also contribute to emerging resistance, mirroring the public health threat posed by antibiotic resistance.
Evolutionary pressure demonstrated by the usage of fungicides in the environment will also likely pre-select resistant fungi before they emerge as pathogens of humans. Some azole fungicides used to protect crops from fungal disease have displayed long half-lives in the soil, potentially due to hydrophobic properties: for example, triadimenol has a half-life in the soil varying from 110 to 375 days [85]. Given that chemicals, including azole fungicides, present in topsoil could be washed into waterways [86] and wastewater [87], fungi have the potential to be exposed in many habitats. Fungicide usage is not limited to crop protection, and these chemicals are also used in the horticultural and timber industries [88], providing ample opportunities for fungal communities to be exposed and develop resistance.
Anthropogenic activity will likely contribute to the initial emergence and subsequent dispersal of these pre-selected drug-resistant fungi. Whilst the acquisition of drug resistance is usually associated with reduced capacity for pathogenicity and reduced overall fitness, we are indeed now witnessing highly fit drug-resistant fungal pathogens, e.g. C. auris. It is hypothesised that climate change causing a warming planet will also lead to the selection of fungal species capable of infection in humans [31], and anthropogenic activity will change species distribution ranges.
Conclusion
Since its first description nearly 15 years ago, C. auris has now been reported around the world, with the exception to Antarctica, creating a global public health due to high levels of antifungal drug resistance and mortality. At this time, over 1000 publications have been written on C. auris, of which nearly 15% feature or mention genomics (PubMed accessed 10th October 2023). The medical and research communities have worked together to not only discover and describe this important pathogen but have also applied cutting-edge technologies to address the question of why C. auris has emerged and how it is evolved to further drug resistance. This presents a roadmap for One Health stakeholders to follow for collaboration and intervention to prevent T. indotineae from following a similar trajectory to C. auris.
Other fungal pathogens will likely emerge, either following the initially susceptible or intrinsically resistant trajectories of A. fumigatus and C. auris respectively. With continued use of antifungal drugs in the environment and clinically, regardless of the origins of emergence, resistance will be sure to follow. Recent reports have mentioned Candida africana as an important emerging agent of vulvovaginal candidiasis [89]; whilst drug resistance is still displaying low prevalence [89], isolated cases are reporting oral candidiasis and altered drug susceptibility profiles [90], perhaps alluding to the potential for a more severe public health problem.
Yet humans are not solely the recipient of drug-resistant fungal pathogens; wildlife and plants are also targets, whether opportunistic or otherwise, thus increasing the opportunity of not only the evolution of further drug resistance, but also “spillover” events into other species. Whilst transmission of a fungal pathogen from the environment to an animal and then on to humans is not a regularly occurring event, there is both potential and precedent for this to occur [91].
As guardians of this planet, humans have a collective responsibility to work together to prevent catastrophe caused by antimicrobial resistance, including antifungal drug resistance. An integrated One Health approach is crucial not only to aid our understanding of the roads leading to drug resistance, but also how we can mitigate the spread across host species at risk of infection. Improved antimicrobial stewardship in medical, agricultural and veterinarian treatment planning is an obvious choice, limiting the use of drug classes for clinical or agricultural usage (and preventing dual-use) and thus slowing the selection pressure and drug resistance evolution. But we can be more ambitious, learning from the success of open-access genomic surveillance platforms generated as part of the COVID-19 pandemic [92], to track the emergence and spread of fungal pathogens globally and across habitats. We can also embrace new technologies, such as rapid “real-time” sequencing for resistance diagnostic test development.
Given the increase in novel emerging fungal pathogens as well as antifungal drug resistance in recent years, a co-ordinated global response between multiple industries is desperately needed. We call on funding agencies to support the creation of these networks to tackle this urgent problem.
Data Availability
No data is associated with this paper as it is a review.
References
Papers of particular interest, published recently, have been highlighted as: •• Of major importance
Hawksworth DL, Lücking R. Fungal diversity revisited: 2.2 to 3.8 million species. Microbiol Spectr. 2017;5(4):5.4.10.
Fisher MC, Alastruey-Izquierdo A, Berman J, Bicanic T, Bignell EM, Bowyer P, et al. Tackling the emerging threat of antifungal resistance to human health. Nat Rev Microbiol [Internet]. 2022 Mar 29 [cited 2022 Jul 6]; Available from: https://www.nature.com/articles/s41579-022-00720-1
Fraser JA, Giles SS, Wenink EC, Geunes-Boyer SG, Wright JR, Diezmann S, et al. Same-sex mating and the origin of the Vancouver Island Cryptococcus gattii outbreak. Nature. 2005;437(7063):1360–4.
Satoh K, Makimura K, Hasumi Y, Nishiyama Y, Uchida K, Yamaguchi H. Candida auris sp. nov., a novel ascomycetous yeast isolated from the external ear canal of an inpatient in a Japanese hospital. Microbiol Immunol. 2009;53(1):41–4.
Rhodes J. Rapid worldwide emergence of pathogenic fungi. Cell Host Microbe. 2019;26(1):12–4.
Brown GD, Denning DW, Gow NAR, Levitz SM, Netea MG, White TC. Hidden killers: human fungal infections. Sci Transl Med. 2012;4(165):165rv13.
Berman J, Krysan DJ. Drug resistance and tolerance in fungi. Nat Rev Microbiol. 2020;18(6):319–31.
•• WHO. WHO fungal priority pathogens list to guide research, development and public health action.World Health Organ. 2022. The inaugural fungal pathogen priority list from the World Health Organisation, detailing which fungal pathogens contribute to the greatest public health threats
CDC. Centers for Disease Control and Prevention. 2022 [cited 2022 Aug 17]. The biggest antibiotic-resistant threats in the U.S. Available from: https://www.cdc.gov/drugresistance/biggest-threats.html
WHO, FAO, OIE. Monitoring and evaluation of the global action plan on antimicrobial resistance.
Van De Veerdonk FL, Gresnigt MS, Romani L, Netea MG, Latgé JP. Aspergillus fumigatus morphology and dynamic host interactions. Nat Rev Microbiol. 2017;15(11):661–74.
Van Rhijn N, Bromley M. The consequences of our changing environment on life threatening and debilitating fungal diseases in humans. J Fungi. 2021;7(5):367.
Resendiz Sharpe A, Lagrou K, Meis JF, Chowdhary A, Lockhart SR, Verweij PE, et al. Triazole resistance surveillance in Aspergillus fumigatus. Med Mycol. 2018;56(suppl_1):S83-92.
Snelders E, Huis in’t Veld RAG, Rijs AJMM, Kema GHJ, Melchers WJG, Verweij PE. Possible environmental origin of resistance of Aspergillus fumigatus to medical triazoles. Appl Environ Microbiol. 2009;75(12):4053–7.
Berger S, Chazli YE, Babu AF, Coste AT. Azole resistance in aspergillus fumigatus: a consequence of antifungal use in agriculture? Front Microbiol. 2017;8:7463–6.
Chowdhary A, Kathuria S, Xu J, Meis JF. Emergence of azole-resistant aspergillus fumigatus strains due to agricultural azole use creates an increasing threat to human health. PLoS Pathog. 2013;9(10):e1003633.
Sewell TR, Zhu J, Rhodes J, Hagen F, Meis JF, Fisher MC, et al. Nonrandom distribution of azole resistance across the global population of Aspergillus fumigatus. mBio. 2019;10(3):209–11.
Sewell TR, Zhang Y, Brackin AP, Shelton JMG, Rhodes J, Fisher MC. Elevated prevalence of azole-resistant Aspergillus fumigatus in urban versus rural environments in the United Kingdom. Antimicrob Agents Chemother. 2019;63(9).
Hollomon D. Does agricultural use of azole fungicides contribute to resistance in the human pathogen Aspergillus fumigatus ? Pest Manag Sci. 2017;73(10):1987–93.
•• Kang SE, Sumabat LG, Melie T, Mangum B, Momany M, Brewer MT. Evidence for the agricultural origin of resistance to multiple antimicrobials in Aspergillus fumigatus , a fungal pathogen of humans. G3 GenesGenomesGenetics. 2022;12(2):jkab427. First study to prove the environmental to clinical route for acquisition of A. fumigatus by humans
•• Rhodes J, Abdolrasouli A, Dunne K, Sewell TR, Zhang Y, Ballard E, et al. Population genomics confirms acquisition of drug-resistant Aspergillus fumigatus infection by humans from the environment. Nat Microbiol. 2022;7(5):663–74. Recent study confirming humans can acquire drug resistant A. fumigatus direct from the environment
Fisher MC, Rannala B, Chaturvedi V, Taylor JW. Disease surveillance in recombining pathogens: multilocus genotypes identify sources of human Coccidioides infections. Proc Natl Acad Sci U S A. 2002;99(13):9067–71.
Vanhove M, Beale MA, Rhodes J, Chanda D, Lakhi S, Kwenda G, et al. Genomic epidemiology of Cryptococcus yeasts identifies adaptation to environmental niches underpinning infection across an African HIV/AIDS cohort. Mol Ecol. 2017;26(7):1991–2005.
Winter DJ, Weir BS, Glare T, Rhodes J, Perrott J, Fisher MC, et al. A single fungal strain was the unexpected cause of a mass aspergillosis outbreak in the world’s largest and only flightless parrot. iScience. 2022;25(12):105470.
Alvarez-Perez S, Mateos A, Dominguez L, Martinez-Nevado E, Blanco JL, Garcia ME. Polyclonal Aspergillus fumigatus infection in captive penguins. Vet Microbiol. 2010;144(3–4):444–9.
Debergh H, Becker P, Vercammen F, Lagrou K, Haesendonck R, Saegerman C, et al. Pulmonary aspergillosis in Humboldt penguins—susceptibility patterns and molecular epidemiology of clinical and environmental Aspergillus fumigatus isolates from a Belgian Zoo, 2017–2022. Antibiotics. 2023;12(3):584.
Verweij PE, Lucas JA, Arendrup MC, Bowyer P, Brinkmann AJF, Denning DW, et al. The one health problem of azole resistance in Aspergillus fumigatus: current insights and future research agenda. Fungal Biol Rev. 2020;34(4):202–14.
Auxier B, Debets AJM, Stanford FA, Rhodes J, Becker FM, Reyes Marquez F, et al. The human fungal pathogen Aspergillus fumigatus can produce the highest known number of meiotic crossovers. PLOS Biol. 2023;21(9):e3002278.
Abdolrasouli A, Rhodes J. Phenotypic variants of azole-resistant Aspergillus fumigatus that co-exist in human respiratory samples are genetically highly related. Mycopathologia. 2022;187(5–6):497–508.
Dunne K, Hagen F, Pomeroy N, Meis JF, Rogers TR. Inter-country transfer of triazole-resistant Aspergillus fumigatus on plant bulbs. Clin Infect Dis. 2017.
Casadevall A, Kontoyiannis DP, Robert V. On the emergence of Candida auris: climate change, azoles, swamps, and birds. mBio. 2019;10(4):41–7.
Sanyaolu A, Okorie C, Marinkovic A, Abbasi AF, Prakash S, Mangat J, et al. Candida auris : an overview of the emerging drug-resistant fungal infection. Infect Chemother. 2022;54(2):236.
Lockhart SR, Etienne KA, Vallabhaneni S, Farooqi J, Chowdhary A, Govender NP, et al. Simultaneous emergence of multidrug resistant Candida auris on three continents confirmed by whole genome sequencing and epidemiological analyses. Clin Infect Dis. 2017;64(2):134–40.
Chow NA, Groot T de, Badali H, Abastabar M, Chiller TM, Meis JF. Potential fifth clade of Candida auris, Iran, 2018 - volume 25, number 9—September 2019 - Emerging Infectious Diseases journal - CDC. [cited 2022 Aug 17]; Available from: https://wwwnc.cdc.gov/eid/article/25/9/19-0686_article
Spruijtenburg B, Badali H, Abastabar M, Mirhendi H, Khodavaisy S, Sharifisooraki J, et al. Confirmation of fifth Candida auris clade by whole genome sequencing. Emerg Microbes Infect. 2022;11(1):2405–11.
Suphavilai C, Ko KKK, Lim KM, Tan MG, Boonsimma P, Keat Chu JJ, et al. Discovery of the sixth Candida auris clade in Singapore [Internet]. Infectious Diseases (except HIV/AIDS); 2023 Aug [cited 2023 Sep 28]. Available from: http://medrxiv.org/lookup/doi/10.1101/2023.08.01.23293435
Rhodes J, Fisher MC. Global epidemiology of emerging Candida auris. Curr Opin Microbiol. 2019;1(52):84–9.
•• Jacobs SE, Jacobs JL, Dennis EK, Taimur S, Rana M, Patel D, et al. Candida auris pan-drug-resistant to four classes of antifungal agents. Antimicrob Agents Chemother. 2022;30:e00053-22. Recent study demonstrating C. auris isolates resistant to four major antifungal drug classes, resulting in patient mortality
Chowdhary A, Sharma C, Meis JF. Candida auris: a rapidly emerging cause of hospital-acquired multidrug-resistant fungal infections globally. PLoS Pathog. 2017;13(5):e1006290.
Borman AM, Szekely A, Johnson EM. Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic Candida species. mSphere. 2016;1(4):9.
Mitchell KF, Zarnowski R, Sanchez H, Edward JA, Reinicke EL, Nett JE, et al. Community participation in biofilm matrix assembly and function. Proc Natl Acad Sci. 2015;112(13):4092–7.
Sherry L, Ramage G, Kean R, Borman A, Johnson EM, Richardson MD, et al. Biofilm-forming capability of highly virulent, multidrug-resistant Candida auris. Emerg Infect Dis. 2017;23(2):328–31.
Lockhart SR. Candida auris and multidrug resistance: defining the new normal. Fungal Genet Biol. 2019;131:103243.
Sekizuka T, Iguchi S, Umeyama T, Inamine Y, Makimura K, Kuroda M, et al. Clade II Candida auris possess genomic structural variations related to an ancestral strain. PLoS ONE. 2019;14(10):e0223433-22.
Oh BJ, Shin JH, Kim MN, Sung H, Lee K, Joo MY, et al. Biofilm formation and genotyping of Candida haemulonii, Candida pseudohaemulonii, and a proposed new species (Candida auris) isolates from Korea. Med Mycol. 2011;49(1):98–102.
Li J, Coste AT, Liechti M, Bachmann D, Sanglard D, Lamoth F. Novel ERG11 and TAC1b mutations associated with azole resistance in Candida auris. Antimicrob Agents Chemother. 2021;65(5):e02663-20.
Rybak JM, Doorley LA, Nishimoto AT, Barker KS, Palmer GE, Rogers PD. Abrogation of triazole resistance upon deletion of CDR1 in a clinical isolate of Candida auris. Antimicrob Agents Chemother. 2019;63(4):e00057-19.
Rhodes J, Abdolrasouli A, Farrer RA, Cuomo CA, Aanensen DM, Armstrong-James D, et al. Genomic epidemiology of the UK outbreak of the emerging human fungal pathogen Candida auris. Emerg Microbes Infect. 2018;7(1):43.
Chowdhary A, Prakash A, Sharma C, Kordalewska M, Kumar A, Sarma S, et al. A multicentre study of antifungal susceptibility patterns among 350 Candida auris isolates (2009–17) in India: role of the ERG11 and FKS1 genes in azole and echinocandin resistance. J Antimicrob Chemother. 2018;13:e1006290.-9.
Berkow EL, Lockhart SR. Activity of CD101, a long-acting echinocandin, against clinical isolates of Candida auris. Diagn Microbiol Infect Dis. 2017.
Carolus H, Pierson S, Muñoz JF, Subotić A, Cruz RB, Cuomo CA, et al. Genome-wide analysis of experimentally evolved Candida auris reveals multiple novel mechanisms of multidrug resistance. MBio. 2021;12(2):e03333-20
Burrack LS, Todd RT, Soisangwan N, Wiederhold NP, Selmecki A. Genomic diversity across Candida auris clinical isolates shapes rapid development of antifungal resistance In Vitro and In Vivo. mBio. 2022;13(4):e00842-22.
Wang Y, Xu J. Population genomic analyses reveal evidence for limited recombination in the superbug Candida auris in nature. Comput Struct Biotechnol J. 2022;20:3030–40.
Li D, Wang Y, Hu W, Chen F, Zhao J, Chen X, et al. Application of machine learning classifier to Candida auris drug resistance analysis. Front Cell Infect Microbiol. 2021;15(11):742062.
Uhrlaß S, Verma SB, Gräser Y, Rezaei-Matehkolaei A, Hatami M, Schaller M, et al. Trichophyton indotineae—an emerging pathogen causing recalcitrant dermatophytoses in India and worldwide—a multidimensional perspective. J Fungi. 2022;8(7):757.
Singh A, Masih A, Monroy-Nieto J, Singh PK, Bowers J, Travis J, et al. A unique multidrug-resistant clonal Trichophyton population distinct from Trichophyton mentagrophytes/Trichophyton interdigitale complex causing an ongoing alarming dermatophytosis outbreak in India: genomic insights and resistance profile. Fungal Genet Biol. 2019;133:103266.
Brown GD, Denning DW, Levitz SM. Tackling human fungal infections. Science. 2012;336(6082):647–647.
Jia S, Long X, Hu W, Zhu J, Jiang Y, Ahmed S, et al. The epidemic of the multiresistant dermatophyte Trichophyton indotineae has reached China. Front Immunol. 2023;16(13):1113065.
Astvad KMT, Hare RK, Jørgensen KM, Saunte DML, Thomsen PK, Arendrup MC. Increasing terbinafine resistance in Danish trichophyton isolates 2019–2020. J Fungi. 2022;8(2):150.
Nenoff P, Verma SB, Ebert A, Süß A, Fischer E, Auerswald E, et al. Spread of terbinafine-resistant trichophyton mentagrophytes type VIII (India) in Germany–“the tip of the iceberg?” J Fungi. 2020;6(4):207.
Jabet A, Brun S, Normand AC, Imbert S, Akhoundi M, Dannaoui E, et al. Extensive dermatophytosis caused by terbinafine-resistant Trichophyton indotineae, France. Emerg Infect Dis. 2022;28(1):229–33.
Klinger M, Theiler M, Bosshard PP. Epidemiological and clinical aspects of Trichophyton mentagrophytes/Trichophyton interdigitale infections in the Zurich area: a retrospective study using genotyping. J Eur Acad Dermatol Venereol. 2021;35(4):1017–25.
Posso-De Los Rios CJ, Tadros E, Summerbell RC, Scott JA. Terbinafine resistant Trichophyton Indotineae isolated in patients with superficial dermatophyte infection in Canadian patients. J Cutan Med Surg. 2022;26(4):371–6.
Sacheli R, Hayette MP. Antifungal resistance in dermatophytes: genetic considerations, clinical presentations and alternative therapies. J Fungi. 2021;7(11):983.
Siopi M, Efstathiou I, Theodoropoulos K, Pournaras S, Meletiadis J. Molecular epidemiology and antifungal susceptibility of trichophyton isolates in Greece: emergence of terbinafine-resistant trichophytonmentagrophytes type VIII locally and globally. J Fungi. 2021;7(6):419.
Fattahi A, Shirvani F, Ayatollahi A, Rezaei-Matehkolaei A, Badali H, Lotfali E, et al. Multidrug-resistant Trichophyton mentagrophytes genotype VIII in an Iranian family with generalized dermatophytosis: report of four cases and review of literature. Int J Dermatol. 2021;60(6):686–92.
Ngo TMC, Ton Nu PA, Le CC, Ha TNT, Do TBT, Tran Thi G. First detection of trichophyton indotineae causing tinea corporis in Central Vietnam. Med Mycol Case Rep. 2022;36:37–41.
Caplan AS, Chaturvedi S, Zhu Y, Todd GC, Yin L, Lopez A, et al. First reported U.S. cases of tinea caused by Trichophyton indotineae — New York City, December 2021–March 2023.
Bortoluzzi P, Prigitano A, Sechi A, Boneschi V, Germiniasi F, Esposto MC, et al. Report of terbinafine resistant Trichophyton spp. in Italy: clinical presentations, molecular identification, antifungal susceptibility testing and mutations in the squalene epoxidase gene. Mycoses. 2023;66(8):680–7.
Shaw D, Singh S, Dogra S, Jayaraman J, Bhat R, Panda S, et al. MIC and Upper Limit of wild-type distribution for 13 antifungal agents against a Trichophyton mentagrophytes-Trichophyton interdigitale complex of Indian origin. Antimicrob Agents Chemother. 2020;64(4):e01964-19.
Burmester A, Hipler U, Elsner P, Wiegand C. Point mutations in the squalene epoxidase erg1 and sterol 14-α demethylase erg11 gene of T indotineae isolates indicate that the resistant mutant strains evolved independently. Mycoses. 2022;65(1):97–102.
Singh S, Chandra U, Anchan VN, Verma P, Tilak R. Limited effectiveness of four oral antifungal drugs (fluconazole, griseofulvin, itraconazole and terbinafine) in the current epidemic of altered dermatophytosis in India: results of a randomized pragmatic trial*. Br J Dermatol. 2020;183(5):840–6.
Ebert A, Monod M, Salamin K, Burmester A, Uhrlaß S, Wiegand C, et al. Alarming India-wide phenomenon of antifungal resistance in dermatophytes: a multicentre study. Mycoses. 2020;63(7):717–28.
Rudramurthy SM, Shankarnarayan SA, Dogra S, Shaw D, Mushtaq K, Paul RA, et al. Mutation in the squalene epoxidase gene of Trichophyton interdigitale and Trichophyton rubrum associated with allylamine resistance. Antimicrob Agents Chemother. 2018;62(5):e02522-17.
Gupta AK, Venkataraman M, Hall DC, Cooper EA, Summerbell RC. The emergence of Trichophyton indotineae : implications for clinical practice. Int J Dermatol. 2023;62(7):857–61.
Nenoff P, Verma SB, Vasani R, Burmester A, Hipler U, Wittig F, et al. The current Indian epidemic of superficial dermatophytosis due to Trichophyton mentagrophytes —a molecular study. Mycoses. 2019;62(4):336–56.
Song J, Zhang S, Lu L. Fungal cytochrome P450 protein Cyp51: what we can learn from its evolution, regulons and Cyp51-based azole resistance. Fungal Biol Rev. 2018;32(3):131–42.
Rosam K, Monk BC, Lackner M. Sterol 14α-demethylase ligand-binding pocket-mediated acquired and intrinsic azole resistance in fungal pathogens. J Fungi. 2020;7(1):1.
Yamada T, Yaguchi T, Maeda M, Alshahni MM, Salamin K, Guenova E, et al. Gene Amplification of CYP51B : a new mechanism of resistance to azole compounds in Trichophyton indotineae. Antimicrob Agents Chemother. 2022;66(6):e00059-22.
Burmester A, Hipler U, Uhrlaß S, Nenoff P, Singal A, Verma SB, et al. Indian Trichophyton mentagrophytes squalene epoxidase erg1 double mutants show high proportion of combined fluconazole and terbinafine resistance. Mycoses. 2020;63(11):1175–80.
Mohammadi LZ, Shams-Ghahfarokhi M, Salehi Z, Razzaghi-Abyaneh M. Terbinafine resistance is associated with newly developed point mutations in squalene epoxidase gene in Trichophyton rubrum and T. mentagrophytes/T. indotineae species complex [Internet]. In Review; 2023 Jun [cited 2024 Feb 4]. Available from: https://www.researchsquare.com/article/rs-3003123/v1
Fachin AL, Ferreira-Nozawa MS, Maccheroni W, Martinez-Rossi NM. Role of the ABC transporter TruMDR2 in terbinafine, 4-nitroquinoline N-oxide and ethidium bromide susceptibility in Trichophyton rubrum. J Med Microbiol. 2006;55(8):1093–9.
Graminha MAS, Rocha EMF, Prade RA, Martinez-Rossi NM. Terbinafine resistance mediated by salicylate 1-monooxygenase in Aspergillus nidulans. Antimicrob Agents Chemother. 2004;48(9):3530–5.
Fisher MC, Gurr SJ, Cuomo CA, Blehert DS, Jin H, Stukenbrock EH, et al. Threats posed by the fungal kingdom to humans, wildlife, and agriculture. mBio. 2020;11(3):e00449-20 Chowdhary A, editor.
Sen P, Vijay M, Singh S, Hameed S, Vijayaraghvan P. Understanding the environmental drivers of clinical azole resistance in Aspergillus species. Drug Target Insights. 2022;16(1):25–35.
Hof H. Critical Annotations to the use of azole antifungals for plant protection. Antimicrob Agents Chemother. 2001;45(11):2987–90.
Chen ZF, Ying GG. Occurrence, fate and ecological risk of five typical azole fungicides as therapeutic and personal care products in the environment: a review. Environ Int. 2015;84:142–53.
Jørgensen LN, Heick TM. Azole use in agriculture, horticulture, and wood preservation – is it indispensable? Front Cell Infect Microbiol. 2021;7(11):730297.
Farahyar S, Izadi S, Razmjou E, Falahati M, Roudbary M, Ashrafi-Khozani M, et al. Low prevalence of antifungal resistant Candida africana, in the C. albicans complex causing vulvovaginal candidiasis. Heliyon. 2020;6(3):e03619.
Lotfali E, Mardani M, Abolghasemi S, Darvishnia D, Rabiei MM, Ghasemi R, et al. Isolation of Candida africana in oral candidiasis: first report among cancer patients in Iran. Curr Med Mycol [Internet]. 2020 Apr 18 [cited 2023 Sep 28]; Available from: https://publish.kne-publishing.com/index.php/CMM/article/view/2695
Snider R, Landers S, Levy ML. The ringworm riddle: an outbreak of Microsporum canis in the nursery. Pediatr Infect Dis J. 1993;12(2):145–8.
An integrated national scale SARS-CoV-2 genomic surveillance network. Lancet Microbe. 2020;1(3):e99–100.
Funding
JR is supported by a Wellcome Trust fellowship (219551/Z/19/Z).
Author information
Authors and Affiliations
Contributions
All authors prepared and reviewed the manuscript.
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
Not applicable.
Conflict of Interest
JR has previously received honoraria from Gilead Sciences.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Medical Mycology
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hui, S.T., Gifford, H. & Rhodes, J. Emerging Antifungal Resistance in Fungal Pathogens. Curr Clin Micro Rpt 11, 43–50 (2024). https://doi.org/10.1007/s40588-024-00219-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-024-00219-8