Abstract
Background
Psoriasis and vitiligo are both chronic, skin-specific diseases classified as autoimmune diseases due to the involvement of several biochemical pathways in their pathogenesis, similar to those altered in other autoimmune diseases. The role of miRNAs in regulating skin autoimmune function has yet to be fully characterized.
Aim
The aim of this study was to assess the expression profile of a panel of 11 circulating immune-related miRNAs in patients with autoimmune skin diseases, specifically psoriasis and vitiligo, and correlate their expression signature with the clinicopathological features of the diseases.
Subjects and Methods
Relative gene expression quantification for 11 immune-related circulating miRNAs in plasma was done for 300 subjects—100 patients with psoriasis, 100 patients with vitiligo and 100 normal healthy volunteers—followed by different modalities of bioinformatics analysis for the results.
Results
The expression levels of all the studied immune-related miRNAs were elevated in both autoimmune skin disorders, with much higher levels of expression in psoriasis than in vitiligo patients. There was a significant correlation between most of the studied miRNAs, suggesting shared target genes and/or pathways. Moreover, all the studied miRNAs showed significant results as biomarkers for autoimmune skin disease, with miRNA-145 being the best candidate. Regarding the clinicopathological data, miRNA-7, miRNA-9, miRNA-145, miRNA-148a, and miRNA-148b were positively correlated with age. All the miRNAs were inversely correlated with obesity and disease duration.
Conclusion
This study highlights the critical role of miRNAs in skin-specific autoimmune diseases that proved to be potential biomarkers for autoimmune skin disorders, warranting their exploration as therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The panel of 11 miRNAs under study comprising miRNA-7, miRNA-9, miRNA-23b, miRNA-124, miRNA-145, miR148a, miRNA-148b, miRNA-155, miRNA-181a, miRNA-203a, and miRNA-320a could serve as potential biomarkers in autoimmune skin disorders, warranting their exploration as therapeutic targets. |
The expression levels of all the studied immune-related miRNAs were elevated in both autoimmune skin disorders, with much higher levels of expression in psoriasis than in vitiligo patients. |
There was a significant correlation between most of the studied miRNAs, suggesting shared target genes and/or pathways. |
MiRNAs under study were correlated with various clinicopathological features in both psoriasis and vitiligo. |
1 Introduction
Autoimmune diseases are a group of chronic conditions that destroy the body’s own healthy tissues through immune mechanisms [1]. Although the primary etiology of most autoimmune diseases remains unspecified, they are known to be associated with a combination of environmental, genetic, and epigenetic factors [2,3,4]. Skin is affected by several autoimmune diseases, including psoriasis and vitiligo.
Psoriasis and vitiligo are both chronic, skin-specific diseases with evidence of genetic predisposition, which have been classified as autoimmune diseases due to the involvement of several biochemical pathways in their pathogenesis, similar to those altered in other autoimmune diseases [5]. Psoriasis involves epidermal thickening due to keratinocyte hyperproliferation [6], while vitiligo is characterized by skin depigmentation due to the selective destruction of melanocytes resulting in depigmented skin patches [7].
Psoriasis and vitiligo share several features. They have many immunological signaling pathways in common, including the Th1 and Th17 pathways and regulatory T cell (Treg) alterations [8,9,10,11,12]. At the genetic level, both diseases are associated with NALP1 gene polymorphisms [13, 14]. Moreover, there are clinicopathological similarities such as skin patches, neuropeptide involvement, absence of organ-specific autoantibodies, positive family history of cardiovascular disease, and the Koebner phenomenon [15,16,17,18].
MicroRNAs (miRNAs) are small (18–25 nucleotides [nt]), highly conserved, non-coding RNA (ncRNA) sequences [19]. Recent advances in molecular medicine have led to the discovery of thousands of miRNAs that are relevant to translational research [20]. The primary function of miRNAs is to block messenger RNA (mRNA) translation into protein via binding to complementary sequences in mRNA. A single miRNA may target multiple genes, and multiple miRNAs may target a single gene [21]. MiRNAs are thus potent modulators of gene expression and regulate up to 60% of protein-coding genes [22].
The role of miRNAs in regulating skin functions has been uncovered but has yet to be fully characterized [23]. Recent research efforts have implicated miRNAs in several skin physiological processes, such as keratinocyte proliferation, melanogenesis, wound healing, and skin ageing [24]. Altered miRNA expression could thus result in skin disease. Since miRNAs are expressed in the skin and biological fluids (e.g., plasma, serum, urine), they could be considered potential non-invasive biomarkers with implications for diagnosis and prognosis and the prediction of therapeutic responses [23]. Recent research efforts have implicated miRNAs in several skin physiological processes, such as keratinocyte proliferation, melanogenesis, wound healing, and skin ageing [24]. Altered miRNA expression could thus result in skin disease. Since miRNAs are expressed in the skin and biological fluids (e.g., plasma, serum, urine), they could be considered potential non-invasive biomarkers with implications for diagnosis and prognosis and the prediction of therapeutic responses.
In this regard, the present study aimed to assess the expression profile of a panel of 11 circulating immune-related miRNAs in patients with the autoimmune skin diseases psoriasis and vitiligo. We also correlated the expression signature with the clinical features of the diseases to shed light on their role in disease pathogenesis and to evaluate their role as disease biomarkers.
2 Subjects and Methods
2.1 Study Population
The current study included 300 subjects. The study subjects were divided into three groups:
-
(i)
Psoriasis patients (Group 1) comprising 100 patients diagnosed with chronic plaque psoriasis of both sexes, aged above 16 years. Immunosuppressed patients or those with serious chronic illnesses or autoimmune diseases were excluded. Psoriasis severity was assessed using Psoriasis Activity Score Index (PASI) score. Psoriasis was rated as mild if the PASI score was < 10, moderate if the PASI score was 10–20 and severe if PASI was > 20.
-
(ii)
Vitiligo patients (Group 2) comprised 100 patients, both males and females, diagnosed with non-segmental vitiligo and assessed by a dermatologist according to the criteria of the Vitiligo Area Severity Index (VASI) and the Vitiligo European Task Force (VETF). Patients with other hypopigmentation disorders were excluded. Patients in both groups were recruited to the Dermatology Clinic. Clinicopathological data were obtained for both groups, including personal history, disease onset age, disease duration, severity, and medication history. The size, site, pattern, and distribution of individual lesions for both groups were examined via detailed dermatological examination.
-
(iii)
Control group (Group 3) comprised 100 healthy volunteers with no history of any autoimmune or chronic diseases.
2.2 The Selection of the Circulating Immune-Related miRNAs Under Study
Selection of the 11 circulating immune-related miRNAs was based on results of the online bioinformatics tools, miR2Disease (http://www.mir2disease.org/) [25] and HMDD (http://www.cuilab.cn/) [26], and available literature [20, 24]. Circulating immune-related miRNAs under study are listed in Supplementary file 1 (see electronic supplementary material [ESM]).
2.3 Blood Sample Collection, miRNA Isolation, and miRNA Quality Assessment
Ethylene diamine tetra-acetic acid (EDTA) anticoagulant vacutainers were used to collect 3 mL of venous blood from each subject. Samples were centrifuged and 100 μL of separated plasma was added to a 500-μL qiazole reagent. Total RNA, including miRNA, was extracted from the plasma-qiazole mixture following the manufacturer’s protocol using Qiagen miRNeasy mini kit (Catalog no. 217004, Qiagen, Hilden, Germany). NanoDrop 2000 1C (NanoDrop Tech., Inc. Wilmington, USA) was used to determine miRNA purity and concentration.
2.4 Relative Gene Expression Quantification of the Circulating Immune-Related miRNAs
The isolated miRNA was subjected to two-step relative gene expression quantification. The first step was reverse transcription (RT), where miScript II RT Kit (Catalog no. 218161, Qiagen, Hilden, Germany) was used to generate complementary DNA (cDNA) from isolated miRNA. Veriti™ 96-Well Thermal Cycler (Applied Biosystems, Thermo Fischer, Waltham, USA) was used to carry the RT reaction for 1 hour at 37 °C, followed by brief incubation at 95 °C for inactivating the reaction.
The second step was SYBR Green-based real-time PCR using miScript SYBR Green PCR Kit with a universal reverse primer (Catalog no. 218076, Qiagen, Hilden, Germany) for the circulating immune-related miRNAs under study. Primer sequences for the 11 circulating immune-related miRNAs (miRNA-7, miRNA-9, miRNA-23b, miR124, miRNA-145, miRNA-148a, miRNA-148b, miRNA-155, miRNA-181a, miRNA-203a, and miRNA-320a) are described in Supplementary file 1 with the experiment set up and conditions (see ESM).
2.5 Gene Expression Data Analysis
The 11 circulating immune-related miRNA expression fold changes were calculated for patients’ samples relative to the comparable controls and estimated using the Livak method [27].
2.6 Circulating Immune-Related miRNAs Predictive Significance Testing
The discriminative significance of the circulating immune-related miRNAs under study was analyzed using the receiver operating characteristic (ROC) curve to assess the diagnostic accuracy of the transcripts under study as biomarkers in both psoriasis and vitiligo.
2.7 Scoring of Disease Prioritization for Circulating Immune-Related miRNAs in Relation to Autoimmune Skin Disease
Utilizing data from the MalaCards database, the human disease database, relationship analysis for vitiligo and psoriasis was carried out using the GeneAnalytics software gene set option [28]. First, the number of miRNAs under study that matched psoriasis and vitiligo was normalized by the number of miRNAs associated primarily with psoriasis and vitiligo. Then the quality and type of the miRNA–psoriasis or miRNA–vitiligo relations, with each messenger RNA (mRNA) in both diseases as represented in MalaCards, was used to determine the disease matching score parameters. VarElect software [29] was used to prioritize the genes of the circulating immune-related miRNAs under study based on their relevance to the autoimmune skin diseases under study, vitiligo and psoriasis. VarElect score is based in part on the weight each gene currently has in relation to vitiligo and psoriasis based on prior research. The weight associated with autoimmune skin diseases was calculated using the frequency (term frequency) at which the miRNA gene is associated with psoriasis and vitiligo in comparison with all other miRNAs’ genes under investigation (inverse document frequency).
2.8 Function and Pathway Enrichment Analysis of the Circulating Immune-Related miRNAs Under Study
The functional enrichment analysis of the 11 circulating immune-related miRNAs under research was performed using the Gene analytics software (https://geneanalytics.genecards.org) [28]. The immune-related circulating miRNAs under study and the gene ontology (GO) concepts that matched them were listed in the order of the matching scores. Higher scores indicated better matches, and the reported score for each circulating immune-related miRNA is a transformation (log2) of the derived p-value. GO, consisting of cellular components, biological processes, and molecular functions terms, was searched for via pathway analysis on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database to determine the affected pathways. The pathway enrichment analysis was carried out using the software Database for Annotation Visualization and Integrated Discovery (DAVID) (https://david.ncifcrf.gov/) [30] and DIANA Tool mirPath v.3. (http://snf-515788.vm.okeanos.grnet.gr/) [31].
2.9 MiRNA Regulatory Network Construction
Using miRTargetLink 2.0 (Version 2.0, available at https://ccb-compute.cs.uni-saarland.de/), the targets of the circulating immune-related miRNAs with statistically significant differential expression were predicted [32]. MiRNA targets that were statistically significant and homogeneous were retained, and negative correlations between miRNA and mRNA pairings were also taken into account.
2.10 Statistical Analysis
The software Statistical Package for the Social Sciences (SPSS) for Windows, version 26.0, was used (IBM Corp., New York, USA). G*Power 3.1.9.2 was used to calculate the sample size and research power. At a total sample size of 300, the power specified for the gene expression study design choice was 90%. While categorical variables were shown as frequencies and percentages, continuous variables were shown as means and standard deviations. The data profile was checked for uniformity and outliers. Mann-Whitney and Student t tests were employed as necessary to compare cases and controls. The correlation coefficient was calculated using the Spearman correlation test. When analyzing two-sided p-values for correlation, Spearman's rank test was used. Statistical significance was defined as a two-tailed p-value = 0.05.
3 Results
3.1 Description of the Study Population
The baseline variables of the 300 study participants are depicted in Table 1. The age of the study participants ranged from 16 to 60 years. The mean age for vitiligo patients was 35.41 ± 15.05 years; for psoriasis patients was 41.8 ± 14.08 years; and for controls was 39.17 ± 11.65 years. When patients were stratified according to age, 42% of vitiligo patients were in the 18–30 years group, while psoriasis patients were more evenly distributed in the different age groups (p = 0.002). Regarding special habits, 55%, 57%, and 61% were smokers among controls, vitiligo patients, and psoriasis patients, respectively, with significant differences among the study groups. Obesity was more represented in vitiligo patients than in other study groups (p < 0.001). Finally, gender and family history did not show statistically significant differences between psoriasis and vitiligo patients.
3.2 Clinical Features of Autoimmune Skin Disease Patients
Table 2 shows disease features of psoriasis and vitiligo patients. The duration of disease was longer in vitiligo patients compared with psoriasis patients (p < 0.001). VASI, Vitiligo Disease Activity Score (VIDA), psoriasis severity, and PASI score showed statistically high significant values (p < 0.001) in relation to the diseases.
3.3 Circulating Immune-Related miRNAs Expression Signature in Autoimmune Skin Disease
Figures 1 and 2 display the expression levels of 11 immune-related miRNAs studied in psoriasis and vitiligo patients. The level of expression of all studied miRNAs was significantly higher in psoriasis patients than in vitiligo patients. MiRNA-7 showed the highest expression level in psoriasis (Log2FC 10.7), followed by miRNA-23b and miRNA-145 (Log2FC 10.2), miRNA-148b (Log2FC 9.8), miRNA-181a (Log2FC 9.0), miRNA-148a (Log2FC 8.6), miRNA-124 (Log2FC 8.3), miRNA-203a (Log2FC 8.1), miRNA-9 (Log2FC 8.0), miRNA-320a (Log2FC 6.6), and miRNA-155 (Log2FC 5.5) in descending order. In vitiligo patients, miRNA-155 was the highest expressed miRNA (Log2FC 4.1) followed by miRNA-23b and miRNA-124 (Log2FC 3.4), miRNA-9 (Log2FC 3.1), miRNA-203a (Log2FC 2.9), miRNA-181a (Log2FC 2.4), miRNA-320a (Log2FC 1.7), miRNA-148a (Log2FC 1.6), miRNA-7 (Log2FC 0.8), miRNA-148b (Log2FC 0.7), and miRNA-145 (Log2FC 0.2) in descending order.
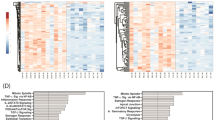
Expression signature of the circulating immune-related miRNAs under study in autoimmune skin disease. The heatmap illustrates the expression levels (Log2fold change) of the 11 immune-related miRNAs (miRNA-7, miRNA-9, miRNA-23b, miRNA-124, miRNA-145, miRNA-148a, miRNA-148b, miRNA-155, miRNA-181a, miRNA-203a, and miRNA-320a) in autoimmune skin disease patients: vitiligo (n = 100) and psoriasis (n = 100). Color grades are shown, with highest expression corresponding to deep red and lowest to deep blue. “V” stands for values in vitiligo patients while “P” stands for values in psoriasis patients
Relative expression levels of the circulating immune-related miRNAs under study in autoimmune skin disease: vitiligo and psoriasis. Eleven miRNAs were analyzed: miRNA-7, miRNA-9, miRNA-23b, miRNA-124, miRNA-145, miRNA-148a, miRNA-148b, miRNA-155, miRNA-181a, miRNA-203a, and miRNA-320a. SNORD68 and RNU6B were used as endogenous controls. The values are represented as median (Q1 and Q3) using Whiskers and bars. “Mann–Whitney U test was used for comparison”. *p < 0.05 was considered statistically significant
3.4 Predictive Significance of the Circulating Immune-Related miRNAs Under Study in Autoimmune Skin Disease
The 11 studied circulating immune-related miRNAs were examined for being potential biomarkers in autoimmune skin disease using ROC curve analysis (Fig. 3). The area under the curve (AUC) of the miRNAs ranged from 0.620 to 0.985, all being significantly associated with autoimmune skin disease (p < 0.001) with miRNA-145 representing the largest (AUC 0.985), followed by miRNA-148b (AUC 0.982), miRNA-7 (AUC 0.979), miRNA-148a (AUC 0.931), miRNA-23b (AUC 0.913), miRNA-9 (AUC 0.881), miRNA-203a (AUC 0.864), miRNA-320a (AUC 0.849), miRNA-181a (AUC 0.848), miRNA-124 (AUC 0.847), and miRNA-155 (AUC 0.620) in descending order.
3.5 Correlation Between the Expression of the Circulating Immune-Related miRNAs and the Clinical Features of Autoimmune Skin Disease
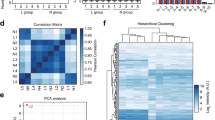
Patients with autoimmune skin diseases showed diverse distributions in the circulating immune-related miRNAs’ expression levels. Table 3 summarizes the analysis of the immune-related Spearman's rank correlation of the 11 immune-related miRNAs in autoimmune skin disease. Nearly all of the examined miRNAs showed a significant connection with autoimmune skin disease, with a Spearman correlation coefficient of 0.59 and a two-tailed significance of p < 0.001.
When the expression levels of the studied miRNAs were correlated with the clinical features of autoimmune skin disease, miRNA-7, miRNA-9, miRNA-145, miRNA-148a, and miRNA-148b were positively correlated with the age of the patient. In contrast, miRNA-7 was correlated with different age groups. All the studied miRNAs showed an inverse correlation between obesity and disease duration. MiRNA-7 and miRNA-320a showed a significant correlation with age at disease onset. Finally, miRNA-320a was correlated with gender (Table 4).
Several correlations were also found among the clinical features of our study participants, as shown in Table 5, where the patient's age was positively correlated with the age of onset of disease, VASI, and VIDA. Age groups showed significant correlations with VASI. Gender showed a significant correlation with the age of onset of disease. Obesity and family history showed a significant correlation with disease duration. VASI was significantly correlated with the patient's age, age group, obesity, and disease duration, while VIDA was only correlated with the patient's age. Finally, PASI and psoriasis severity showed a significant direct correlation.
3.6 Prioritization Score of Circulating Immune-Related miRNA Genes in Relation to Autoimmune Skin Disease
Figure 4 displays the prioritization scores of the circulating immune-related miRNA genes under study classified into two categories, directly and indirectly related to disease, according to detected links between immune-related miRNA genes and both psoriasis and vitiligo. This list is based on the association between these genes and the shared pathways, interaction networks, paralog relations, domain sharing, and the mutual publications of the autoimmune skin diseases under study. Table 6 lists the genes implicated with the indirectly related category of miRNA genes.
Prioritization score of eleven circulating immune-related miRNA genes in autoimmune skin disease. A Prioritization score for miRNA genes in vitiligo. B Prioritization score for miRNA genes in psoriasis. All scores were generated with the VarElect tool (https://ve.genecards.org)
3.7 Pathway and Function Enrichment Analysis for the Circulating Immune-Related miRNAs
A pathway enrichment analysis was performed based on annotated gene targets to detect the pathways targeted by the differentially expressed immune-related miRNAs in autoimmune skin disease. MirPath v.3 database was used to perform functional pathway analysis. KEGG enrichment analysis (Fig. 5) revealed the following pathways: extracellular matrix (ECM)-receptor interaction, proteoglycans in cancer, fatty acid biosynthesis, glioma, hippo signaling pathway, adherens junction, pathways in cancer, fatty acid elongation, TGF-β signaling pathway, and transcriptional misregulation in cancer.
The GO biological processes enriched in our analysis were distinctly associated with gene silencing by miRNA, miRNA-mediated inhibition of translation, negative regulation of gene expression, inflammatory response, interleukin (IL)-8 and IL-11 production, microglial cell activation, low-density lipoprotein particle clearance, receptor signaling pathway via JAK-STAT, and positive regulation of macrophage activation as shown (Fig. 6A).
“Gene ontology matching scores results for eleven immune-related miRNAs in autoimmune skin disease. A Biological processes in matching score. B Cellular component according to matching score. C Molecular function according to matching score. All scores were generated using gene analytics tool. (https://geneanalytics.genecards.org)”. MRNA messenger RNA, MiRNA microRNA, RISC RNA-induced silencing complex, UTR untranslated region
The extracellular space, extracellular vesicles, RNA-induced silencing complex (RISC), extracellular exosomes, apical section of the cell, and perinuclear region of the cytoplasm were all associated with the circulating immune-related miRNAs under study (Fig. 6B). The molecular activities of these miRNAs included mRNA binding involved in posttranscriptional gene silencing, mRNA 3'-UTR binding, high-density lipoprotein particle binding, RNA polymerase II complex binding, and single-stranded RNA binding (Fig. 6C).
3.8 Circulating Immune-Related miRNA‐mRNA Regulatory Network in Autoimmune Skin Disease
Our network analysis revealed the connections between the target genes of the circulating immune-related miRNAs under investigation. Initially, 10,962 target genes were identified in the miRNA-target gene network made up of the 11 immune-related miRNAs (Supplementary file 2, see ESM). Then, these were selected to only keep targets that had at least two shared strongly validated targets. Using miRTargetLink 2.0 (https://ccb-web.cs.uni-saarland.de/mirtargetlink/network.php), a final list of 30 target genes was obtained (Fig. 7).
“MiRNAs-target gene network analysis for eleven immune-related miRNAs in autoimmune skin disease. The targets used were the strongly validated miRNAs in the literature with an additional filter of minimum 2 shared targets using miRTargetLink 2.0 (https://ccb-compute.cs.uni-saarland.de/mirtargetlink2)”
According to our miRNA‐mRNA regulatory network analysis results, STAT3, IRS1, TGFB2, IRS2, EGFR, MSH3, NDUFA4, SMAD2, RGS5, IGF1R, STAT1, KLF4, BCL2, CDKN1A, CDKN1B, MCL1, XIAP, NRAS, CDK6, SAMHD1, PPP3CA, PTEN, ETS1, MYC, ABCC1, SMAD3, BAX, TGFBR2, CCND2, and FOS are the critical genes involved in autoimmune skin disease in relation to the studied immune-related miRNAs.
4 Discussion
Psoriasis and vitiligo are skin-restricted autoimmune diseases with several clinicopathological, immunological, and genetic features [33]. MiRNAs are short gene-regulatory RNA molecules implicated in most physiological and pathological processes, including differentiation, proliferation, migration, and survival [34]. MiRNAs are abundantly expressed in the skin [35] and are involved in skin development, maintenance, and homeostasis through regulating cell proliferation, differentiation, and immune regulation [36]. In addition, they are becoming essential targets for disease diagnosis and treatment [37]. They have been shown to regulate keratinocyte differentiation and proliferation, as well as melanogenesis and development and function of immune cells [38], processes relevant to the pathogenesis of psoriasis and vitiligo, respectively. Dysregulated miRNA expression has been demonstrated in several skin disorders, including the autoimmune skin diseases psoriasis and vitiligo [36, 37].
In this study, we aimed to explore the circulating miRNA repertoire of the autoimmune skin disorders psoriasis and vitiligo and to identify potential biomarkers for the diseases. Eleven immune-related miRNAs were selected to be examined in psoriasis (n = 100) and vitiligo (n = 100) patients as compared with healthy controls (n = 100). The studied miRNAs were miRNA-7, miRNA-9, miRNA-23b, miRNA-124, miRNA-145, miR148a, miRNA-148b, miRNA-155, miRNA-181a, miRNA-203a, and miRNA-320a. The expression levels of all the studied immune-related miRNAs were elevated in both autoimmune skin disorders, with much higher levels of expression in psoriasis than in vitiligo patients (Figs 1 and 2).
4.1 Differentially Expressed miRNAs in Vitiligo
MiRNA-155 upregulation has been found in the serum [39] and lesional skin [40] of vitiligo patients. Šahmatova et al. demonstrated that miRNA-155 was stimulated by vitiligo-associated cytokines (TNF-α, IFN-α, IFN-γ and IL-1β) in human primary melanocytes and keratinocytes and the upregulated miRNA-155 inhibited the expression of melanogenesis-associated genes (TYRP1, YWHAE, SDCBP and SOX10 in melanocytes, and YWHAE in keratinocytes) and altered interferon-regulated genes (SOCS1, IRF1 and IFITM1) in melanocytes and keratinocytes [40]. Moreover, miRNA-155 has been shown to increase the differentiation of Treg cells by stimulating forkhead box P3 (Foxp3) transcription [41]. Tregs are suppressive T cells that inhibit autoimmune CD8+ T cells and can damage melanocytes [42]. MiRNA-155 has also been shown to decrease CD8+ T-cell growth and promote melanocyte proliferation via increasing Treg cells [43]. In addition, IL-1β stimulated upregulation of miRNA-155 resulted in reduced expression in melanoma cells of the microphthalmia-associated transcription factor (MIFT) [44], a melanocyte-specific transcription factor involved in melanocyte survival, proliferation, and differentiation [45]. Therefore, miRNA-155 is a main miRNA in melanocyte and immune cell function, both involved in vitiligo. Treatment with narrowband ultraviolet B (UVB) has been shown to reduce miRNA-155 expression in peripheral mononuclear cells (PMNCs) from active non-segmental vitiligo (NSV) patients [46], making it a potential target for vitiligo treatment. Consistently, our data shows circulating miRNA-155 as being the highest expressed miRNA in vitiligo patients (Log2FC 5.5) and of moderate discriminative value (AUC 0.620) in autoimmune skin disease.
The miRNA-9 expression has been shown to be increased in the lesional skin [47,48,49] and serum [49] of vitiligo patients. Raia et al. also demonstrated a positive correlation between circulating miRNA-9 levels with disease extent and VASI score in Egyptian vitiligo patients [49]. MiRNA-9 could target the expression of sirtuin 1 (SIRT1), which is known to protect against ageing and other stress-related diseases, implicating a role in destroying melanocytes [47]. Su et al. have also shown that the miRNA-9 upregulation in lesional vitiligo skin was associated with decreased expression of adhesion molecules (E-cadherin and β1 integrin) and that exposing the normal human keratinocyte cell line HaCaT cells to UVB decreased the expression of miRNA-9 and increased the expression of IL-10, E-cadherin and β1 integrin, thereby antagonizing the miRNA-9 migration-inhibitory effect on pigment cells during UVB-induced repigmentation [48], making miRNA-9 an essential therapeutic target in vitiligo. This is consistent with our findings, where the circulating miRNA-9 was the third highest expressed miRNA in our vitiligo patients (Log2FC 3.1), is positively correlated with patient age and negatively correlated with obesity and disease duration, and is of excellent discriminative value using ROC curve analysis (AUC 0.881) in autoimmune skin disease.
MiRNA-124 is a tumor suppressor in several tumors, including melanoma [50]. To our knowledge, miRNA-124 has not been examined for vitiligo. However, melanoma and vitiligo have long been considered related diseases [51]. Downregulated miRNA-124 expression has been demonstrated in melanoma cells associated with upregulated RACK1. Through RACK1 degradation, overexpression of miRNA-124 resulted in impaired melanoma cell proliferation, migration, invasion, and increased apoptosis [50]. This may be relevant to vitiligo immunopathogenesis. Our findings have demonstrated that increased circulating miRNA-124 expression levels, being the second highest expressed (Log2FC 3.4) in vitiligo patients, is positively correlated with age and negatively correlated with obesity and disease duration and is of excellent discriminative value in autoimmune skin disease (AUC 0.847).
The expression level of miRNA-181a has been shown to be increased in PBMCs from active NSGV patients [44]. This is consistent with our current and previous findings in vitiligo patients [52], where it is the fifth highest expressed miRNA in vitiligo (Log2FC 2.4), is inversely correlated with obesity and disease duration, and is of excellent discrimination value in autoimmune skin disease (AUC 0.848).
Collectively, these data on the role of miRNA-155 and miRNA-9 in vitiligo pathogenesis support our data that these miRNAs are directly related to vitiligo pathogenesis (Fig. 4). Other miRNAs were indirectly related to vitiligo with varying degrees (Table 6). Implicated genes related to melanogenesis (MITF, POMC), oxidative stress defense (PRDX5), apoptosis (FAS, p53, MYC), DNA repair (NBN, p53, and ATM), autoimmunity-related genes (TNF, IL2, IL6, IL1B, IFNG, TGFBR2, HLA-C, and STAT3), and cell adhesion (ITGA5) are closely related to the pathogenesis of vitiligo [53].
4.2 Differentially Expressed miRNAs in Psoriasis
MiRNA-203a is a keratinocyte-derived miRNA involved in the balance of keratinocyte proliferation and differentiation and is considered a marker of keratinocyte differentiation [54]. MiRNA-203a has reportedly been upregulated in psoriasis skin lesions [55,56,57] and plasma [58]. MiRNA-203a in keratinocytes is believed to inhibit skin immune responses by downregulating pro-inflammatory cytokine genes TNF-α and IL-24 [59]. Moreover, miRNA-203a inhibits the expression of suppressor of cytokine signaling 3 (SOCS-3) with subsequent upregulation of signal transducer and activator of transcription-3 (STAT-3), a transcription factor in keratinocytes that regulates keratinocyte proliferation and differentiation [60]. In addition, miRNA-203 inhibits the expression of liver X receptor-α (LXR-α) and peroxisome proliferator-activated receptor-γ (PPAR-γ) in psoriatic lesions. In contrast, the overexpression of LXR-α and PPAR-γ inhibits keratinocyte proliferation, suggesting a role for the miRNA-203a-LXR-α /PPARγ axis in keratinocyte hyperproliferation in psoriasis [61]. In our study, circulating miRNA-203a was upregulated in psoriasis (Log2FC 8.1). It was significantly higher in psoriasis when compared with vitiligo (Log2FC 2.9) (p < 0.001), reflecting its crucial role in keratinocyte differentiation and proliferation and making it a target for psoriasis therapy. MiRNA-203a was inversely correlated with obesity and disease duration and was of excellent discriminative value (AUC 0.864) in autoimmune skin disease.
MiRNA-155 has also been shown to be upregulated in skin lesions [62,63,64] and PMNCs [66] of psoriasis. The role of miRNA-155 in psoriasis could be explained by promoting keratinocyte proliferation and inhibiting apoptosis through downregulating PTEN [62, 66]. Moreover, the knockdown of miR155 in the human keratinocyte line HaCaT cells significantly increased cells in the G0/G1 phase and decreased those in the G2/M phase [62], promoting cell differentiation rather than proliferation. In our study, circulating miRNA-155 was upregulated in psoriasis (Log2FC 5.5) and was significantly higher in psoriasis than in vitiligo (Log2FC 4.1) (p = 0.004). According to our findings, miRNA-155 has been identified as one of the top four priority miRNAs in both vitiligo and psoriasis. Taken together, miRNA-155 plays multiple roles in keratinocyte proliferation, apoptosis, melanogenesis, and immune cell regulation, making it a potential therapeutic choice in autoimmune skin disease.
The miRNA-145 circulating level has been shown to be downregulated in psoriasis and could be protective against psoriasis [67, 68]. MiRNA-145 inhibits keratinocyte proliferation by downregulating the MLK3 gene via regulating NF-kB and STAT-3 [68]. Moreover, miRNA-145 inhibits keratinocyte proliferation and stimulates apoptosis through inhibiting Wnt/β-catenin signaling, which is involved in psoriasis pathogenesis [69], implicating an inhibitory role for miRNA-145 in psoriasis [68]. This is consistent with our finding that miRNA-145 is directly involved in psoriasis pathogenesis and is of the highest predictive value among the 11 immune-related miRNAs under study (AUC 0.985, p < 0.001) in autoimmune disease. However, it contrasts with our data on circulating miRNA-145 level, which is significantly upregulated in psoriasis (Log2FC 10.2) and is significantly higher in psoriasis compared with vitiligo (Log2FC 0.2) (p < 0.001).
MiRNA-9 expression levels were shown to be upregulated in the serum of psoriasis patients and were positively correlated with disease extent and PASI [70] but downregulated in the skin of psoriasis patients [70, 71]. Interestingly, miRNA-9 has been shown to influence Th17 differentiation by inhibiting the expression of negative regulators of Th17 differentiation [72]. The serum data is consistent with our findings, where the circulatory miRNA-9 expression is upregulated in psoriasis patients (Log2FC 8.0) and is significantly higher in psoriasis compared with vitiligo (Log2FC 3.1, p < 0.001). When we prioritized the 11 circulating immune-related miRNAs under study, miRNA-9 was in the top four in both psoriasis and vitiligo, directly involved in vitiligo and indirectly in psoriasis (Fig. 4). Moreover, miRNA-203a is one of the genes implicated with miRNA-9 in relation to psoriasis (Table 6); we have already explained its essential role in our previous work [73].
Collectively, this supports our data that miRNA-203 and miRNA-155 are directly involved in psoriasis pathogenesis (Fig. 4). On the other hand, the genes implicated with the miRNAs indirectly involved in psoriasis (Table 6) are related to chemokines (CXCL8), cytokines and/or autoimmunity genes (NOD2, TNF, IL6, TNFRSF1A, IL4, IL1B, IFNG, IFIH1, and DDX58), apoptosis (Tp53), and miRNAs associated with psoriasis such as miRNA-203a, miRNA-200a, miRNA-141, miRNA-30 and miRNA-LET-7e [74, 75].
In silico analysis of the 11 immune-related miRNAs in the studied autoimmune skin diseases (Figs 5, 6 and 7) revealed that the KEGG targeted pathways and GO biological processes, as well as the cellular components and target genes, have been consistently shown to be involved in psoriasis and vitiligo pathogenesis [53, 75,76,77,78]. Nevertheless, further research is necessary to identify the network of intricate interactions between each of these miRNAs and the molecules/genes/pathways involved in autoimmune disease in general and vitiligo and psoriasis in particular.
Finally, we emphasize that miRNAs are involved in the regulation of various cellular and physiological processes including adipose tissue development and metabolism, insulin secretion and function, and energy balance. Dysregulated miRNA expression, thus, has been incriminated in the pathogenesis of obesity and obesity-related diseases [79,80,81].
Our findings show that the studied miRNAs were inversely correlated with obesity and disease duration. Interestingly, several miRNAs evaluated in this study have been shown to be involved in obesity-related processes. miRNA-203 is a key regulator of the development of brown adipocytes [82] and is dysregulated in relation to diet [83]. miRNA-155 inhibits adipogenic differentiation and the development of the thermogenic program in brown adipocytes [83] and participates in the amplification of inflammatory status in adipocytes [81]. miRNA-9 regulates the release of insulin in pancreatic β-cells in rat and mouse models [79]. miRNA-124 is involved in pancreatic development, as well as in insulin expression and release [79]. miRNA-320 regulates insulin resistance in insulin-resistant adipocytes [79]. miRNA-148a, normally induced during adipogenesis, is downregulated in cells isolated from mice models of obesity [79]. The expression of miRNA-145 correlates with key metabolic parameters, including fasting plasma glucose, and circulating leptin and adiponectin levels [79], and is involved in adipocyte inflammatory responses via stimulating the expression of TNF-α in adipocytes [81]. Our findings may support the recent suggested connections between obesity and autoimmune disease [84]. Nevertheless, further research is necessary to establish the common signaling and genetic pathways involved.
4.3 Study Limitations
This study is mono-center, so selection bias cannot be disregarded. Moreover, our results are limited by the ethnicity of the study participants. Finally, further in vivo and in vitro research is necessary to examine the molecular relationships uncovered by in silico analysis. Therefore, larger multi-center clinical and functional studies are recommended.
5 Conclusion
This study highlights the critical role of miRNAs in skin-specific autoimmune diseases, namely psoriasis and vitiligo, through examining the expression repertoire of 11 circulatory immune-related miRNAs, which were found to be upregulated in both diseases, more so in psoriasis, and were correlated with various clinicopathological features of both, and proved to be potential biomarkers for autoimmune skin disorders, warranting their exploration as therapeutic targets.
References
Goris A, Liston A. The immunogenetic architecture of autoimmune disease. Cold Spring Harb Perspect Biol. 2012;4(3):a007260. https://doi.org/10.1101/cshperspect.a007260.
McLafferty E, Hendry C, Alistair F. The integumentary system: anatomy, physiology and function of skin. Nurs Stand. 2012;27(3):35–42. https://doi.org/10.7748/ns2012.10.27.7.35.c9358.
Kolarsick PA, Kolarsick MA, Goodwin C. Anatomy and physiology of the skin. JDNA. 2011;3(4):203–13.
Bolon B. Cellular and molecular mechanisms of autoimmune disease. Toxicol Pathol. 2012;40(2):216–29. https://doi.org/10.1177/0192623311428481.
Jiang S, Hinchliffe TE, Wu T. Biomarkers of an autoimmune skin disease-psoriasis. Genom Proteom Bioinform. 2015;13(4):224–33. https://doi.org/10.1016/j.gpb.2015.04.002.
Albanesi C. Immunology of psoriasis. In: Immunology (Fifth Edition), Principles and Practice, 2019, pp 871-878.e1. https://www.sciencedirect.com/science/article/pii/B9780702068966000648
Seneschal J, Harris JE, Le Poole IC, Passeron T, Speeckaert R, Boniface K. Editorial: immunology of vitiligo. Front Immunol. 2021;12:711080. https://doi.org/10.3389/fimmu.2021.711080.
Speeckaert R, Lambert J, Grine L, Van Gele M, De Schepper S, van Geel N. The many faces of interleukin-17 in inflammatory skin diseases. Br J Dermatol. 2016;175(5):892–901. https://doi.org/10.1111/bjd.14703.
Yen H, Chi CC. Association between psoriasis and vitiligo: a systematic review and meta-analysis. Am J Clin Dermatol. 2019;20(1):31–40. https://doi.org/10.1007/s40257-018-0394-1.
Ono S, Tanizaki H, Otsuka A, et al. Coexistent skin lesions of vitiligo and psoriasis vulgaris. Immunohistochemical analyses for IL-17A-producing cells and regulatory T cells. Acta Derm Venereol. 2014;94(3):329–30. https://doi.org/10.2340/00015555-1713.
Das D, Akhtar S, Kurra S, Gupta S, Sharma A. Emerging role of immune cell network in autoimmune skin disorders: an update on pemphigus, vitiligo and psoriasis. Cytokine Growth Factor Rev. 2019;45:35–44. https://doi.org/10.1016/j.cytogfr.2019.01.001.
Mukhatayev Z, Ostapchuk YO, Fang D, Le Poole IC. Engineered antigen-specific regulatory T cells for autoimmune skin conditions. Autoimmun Rev. 2021;20(3):102761. https://doi.org/10.1016/j.autrev.2021.102761.
Sharon C, Jiquan S. Association of NLRP1 and NLRP3 gene polymorphism with psoriasis. Our Dermatol Online. 2020;11(3):275–83. https://doi.org/10.7241/ourd.20204.13.
Saleh AA, Shehata WA, Abd-Elhafiz HI, Soliman SE. Potential impact of TNFAIP3 rs6920220 and DEFB1 rs1800972 gene polymorphisms on vitiligo in Egyptian patients. Meta Gene. 2022;1(31): 101002.
Sandhu K, Kaur I, Kumar B. Psoriasis and vitiligo. J Am Acad Dermatol. 2004;51(1):149–50. https://doi.org/10.1016/j.jaad.2003.12.014.
Sharquie KE, Sharquie IK, Al Hamza AN. Psoriasis, pityriasis alba, and vitiligo (PPV) are a triad of one disease: new observation. Our Dermatol Online. 2021;12(3):314–23. https://doi.org/10.7241/ourd.20213.21.
Sagi L, Trau H. The Koebner phenomenon. Clin Dermatol. 2011;29(2):231–6. https://doi.org/10.1016/j.clindermatol.2010.09.014.
Arunachalam M, Dragoni F, Colucci R, et al. Non-segmental vitiligo and psoriasis comorbidity - a case-control study in Italian patients. J Eur Acad Dermatol Venereol. 2014;28(4):433–7. https://doi.org/10.1111/jdv.12117.
Hüttenhofer A, Schattner P, Polacek N. Non-coding RNAs: hope or hype? Trends Genet. 2005;21(5):289–97. https://doi.org/10.1016/j.tig.2005.03.007.
Guarnieri DJ, DiLeone RJ. MicroRNAs: a new class of gene regulators. Ann Med. 2008;40(3):197–208. https://doi.org/10.1080/07853890701771823.
Peter ME. Targeting of mRNAs by multiple miRNAs: the next step. Oncogene. 2010;29(15):2161–4. https://doi.org/10.1038/onc.2010.59.
Catalanotto C, Cogoni C, Zardo G. MicroRNA in control of gene expression: an overview of nuclear functions. Int J Mol Sci. 2016;17(10):1712. https://doi.org/10.3390/ijms17101712.
Deng X, Su Y, Wu H, et al. The role of microRNAs in autoimmune diseases with skin involvement. Scand J Immunol. 2015;81(3):153–65. https://doi.org/10.1111/sji.12261.
Khan AQ, Ahmad F, Raza SS, et al. Role of non-coding RNAs in the progression and resistance of cutaneous malignancies and autoimmune diseases. Semin Cancer Biol. 2022;83:208–26. https://doi.org/10.1016/j.semcancer.2020.07.003.
Jiang Q, Wang Y, Hao Y, et al. miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res. 2009;37(Database issue):D98–104. https://doi.org/10.1093/nar/gkn714.
Huang Z, Shi J, Gao Y, et al. HMDD v3.0: a database for experimentally supported human microRNA-disease associations. Nucleic Acids Res. 2019;47(D1):D1013–7. https://doi.org/10.1093/nar/gky1010.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.1262.
Ben-Ari Fuchs S, Lieder I, Stelzer G, et al. GeneAnalytics: an integrative gene set analysis tool for next generation sequencing. RNAseq and Microarray Data OMICS. 2016;20(3):139–51. https://doi.org/10.1089/omi.2015.0168.
Stelzer G, Plaschkes I, Oz-Levi D, Alkelai A, Olender T, Zimmerman S, Twik M, Belinky F, Fishilevich S, Nudel R, Guan-Golan Y. VarElect: the phenotype-based variation prioritizer of the GeneCards Suite. BMC Genomics. 2016;17(2):195–206. https://doi.org/10.1186/s12864-016-2722-2.
Dennis G, Sherman BT, Hosack DA, et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4:R60. https://doi.org/10.1186/gb-2003-4-9-r60.
Vlachos IS, Zagganas K, Paraskevopoulou MD, et al. DIANA-miRPath v3.0: deciphering microRNA function with experimental support. Nucleic Acids Res. 2015;43(W1):W460-W466. https://doi.org/10.1093/nar/gkv403
Kern F, Aparicio-Puerta E, Li Y, et al. miRTargetLink 2.0-interactive miRNA target gene and target pathway networks. Nucleic Acids Res. 2021;49(W1):W409–16. https://doi.org/10.1093/nar/gkab297.
Canu D, Shourick J, Andreu N, et al. Demographic and clinical characteristics of patients with both psoriasis and vitiligo in a cohort of vitiligo patients: a cross-sectional study. J Eur Acad Dermatol Venereol. 2021;35(10):e676–9. https://doi.org/10.1111/jdv.17383.
O’Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol (Lausanne). 2018;9:402. https://doi.org/10.3389/fendo.2018.00402.PMID:30123182;PMCID:PMC6085463.
Yi R, Fuchs E. MicroRNA-mediated control in the skin. Cell Death Differ. 2010;17(2):229–35. https://doi.org/10.1038/cdd.2009.92.
Singhvi G, Manchanda P, Krishna Rapalli V, Kumar Dubey S, Gupta G, Dua K. MicroRNAs as biological regulators in skin disorders Biomed. Pharmacother. 2018;108:996–1004. https://doi.org/10.1016/j.biopha.2018.09.090.
Li X, Ponandai-Srinivasan S, Nandakumar KS, et al. Targeting microRNA for improved skin health. Health Sci Rep. 2021;4:e374. https://doi.org/10.1002/hsr2.374.
Lee A-Y. The role of microRNAs in epidermal barrier. Int J Mol Sci. 2020;21(16):5781. https://doi.org/10.3390/ijms21165781.
Issa YW, Salih SM. Impact of miRNA-155, miRNA-145 and miRNA-328 on pigmentary process in Iraqi patients with vitiligo. Gene Rep. 2020;21:100955. https://doi.org/10.1016/j.genrep.2020.100955.
Šahmatova L, Tankov S, Prans E, et al. MicroRNA-155 is dysregulated in the skin of patients with vitiligo and inhibits melanogenesis-associated genes in melanocytes and keratinocytes. Acta Derm Venereol. 2016;96(6):742–7. https://doi.org/10.2340/00015555-2394.
Kempinska-Podhorodecka A, Milkiewicz M, Wasik U, Ligocka J, Zawadzki M, Krawczyk M, Milkiewicz P. Decreased expression of vitamin D receptor affects an immune response in primary biliary cholangitis via the VDR-miRNA155-SOCS1 pathway. Int J Mol Sci. 2017;18:E289. https://doi.org/10.3390/ijms18020289.
Lan Q, Zhou X, Fan H, Chen M, Wang J, Ryffel B, Brand D, Ramalingam R, Kiela PR, Horwitz DA, et al. Polyclonal CD4+Foxp3+ Treg cells induce TGFβ-dependent tolerogenic dendritic cells that suppress the murine lupus-like syndrome. J Mol Cell Biol. 2012;4:409–19. https://doi.org/10.1093/jmcb/mjs040.
Lv M, Li Z, Liu J, et al. MicroRNA 155 inhibits the proliferation of CD8+ T cells via upregulating regulatory T cells in vitiligo. Mol Med Rep. 2019;20(4):3617–24. https://doi.org/10.3892/mmr.2019.10607.
Arts N, Cané S, Hennequart M, et al. microRNA-155, induced by interleukin-1ß, represses the expression of microphthalmia-associated transcription factor (MITF-M) in melanoma cells. PLoS ONE. 2015;10(4):e0122517. https://doi.org/10.1371/journal.pone.0122517.
Vance KW, Goding CR. The transcription network regulating melanocyte development and melanoma. Pigment Cell Res. 2004;17(4):318–25. https://doi.org/10.1111/j.1600-0749.2004.00164.x.
Parihar AS, Tembhre MK, Sharma VK, Gupta S, Chattopadhyay P, Deepak KK. Effect of narrowband ultraviolet B treatment on microRNA expression in active nonsegmental generalized vitiligo. Br J Dermatol. 2020;183(1):167–9. https://doi.org/10.1111/bjd.18890. (Epub 2020 Mar 11 PMID: 31975367).
Mansuri MS, Singh M, Dwivedi M, Laddha NC, Marfatia YS, Begum R. MicroRNA profiling reveals differentially expressed microRNA signatures from the skin of patients with nonsegmental vitiligo. Br J Dermatol. 2014;171(5):1263–7. https://doi.org/10.1111/bjd.13109.
Su M, Yi H, He X, Luo L, Jiang S, Shi Y. miRNA-9 regulates melanocytes adhesion and migration during vitiligo repigmentation induced by UVB treatment. Exp Cell Res. 2019;384(1):111615. https://doi.org/10.1016/j.yexcr.2019.111615.
Raia NMA, Shaker OG, Hassan ZM, Abd Elrahim TA. Is there a relation between long non-coding RNA MALAT-1 and miRNA-9 in Egyptian patients with Vitiligo? Exp Dermatol. 2022;31(3):381–3. https://doi.org/10.1111/exd.14487.
Shen C, Hua H, Gu L, et al. miRNA-124 functions as a melanoma tumor suppressor by targeting RACK1. Onco Targets Ther. 2019;12:9975–86. https://doi.org/10.2147/OTT.S225120.
Failla CM, Carbone ML, Fortes C, Pagnanelli G, D’Atri S. Melanoma and vitiligo: in good company. Int J Mol Sci. 2019;20(22):5731. https://doi.org/10.3390/ijms20225731.
Abdallah HY, Abdelhamid NR, Mohammed EA, et al. Investigating melanogenesis-related microRNAs as disease biomarkers in vitiligo. Sci Rep. 2022;12(1):13526. https://doi.org/10.1038/s41598-022-17770-3.
Marchioro HZ, Silva de Castro CC, Fava VM, Sakiyama PH, Dellatorre G, Miot HA. Update on the pathogenesis of vitiligo. An Bras Dermatol. 2022;97(4):478–90. https://doi.org/10.1016/j.abd.2021.09.008.
Hildebrand J, Rütze M, Walz N, et al. A comprehensive analysis of microRNA expression during human keratinocyte differentiation in vitro and in vivo. J Invest Dermatol. 2011;131(1):20–9. https://doi.org/10.1038/jid.2010.268.
Zibert JR, Løvendorf MB, Litman T, Olsen J, Kaczkowski B, Skov L. MicroRNAs and potential target interactions in psoriasis. J Dermatol Sci. 2010;58(3):177–85. https://doi.org/10.1016/j.jdermsci.2010.03.004.
Sonkoly E, Wei T, Janson PC, et al. MicroRNAs: novel regulators involved in the pathogenesis of psoriasis? PLoS ONE. 2007;2(7):e610. https://doi.org/10.1371/journal.pone.0000610.
Joyce CE, Zhou X, Xia J, et al. Deep sequencing of small RNAs from human skin reveals major alterations in the psoriasis miRNAome. Hum Mol Genet. 2011;20(20):4025–40. https://doi.org/10.1093/hmg/ddr331.
Mostafa SA, Mohammad MHS, Negm WA, et al. Circulating microRNA203 and its target genes’ role in psoriasis pathogenesis. Front Med (Lausanne). 2022;9:988962. https://doi.org/10.3389/fmed.2022.988962.
Primo MN, Bak RO, Schibler B, Mikkelsen JG. Regulation of pro-inflammatory cytokines TNFα and IL24 by microRNA-203 in primary keratinocytes. Cytokine. 2012;60(3):741–8. https://doi.org/10.1016/j.cyto.2012.07.031.
Sonkoly E, Wei T, Pavez Loriè E, et al. Protein kinase C-dependent upregulation of miRNA-203 induces the differentiation of human keratinocytes. J Invest Dermatol. 2010;130(1):124–34. https://doi.org/10.1038/jid.2009.294.
Xiao Y, Wang H, Wang C, Zeng B, Tang X, Zhang Y, et al. miRNA-203 promotes HaCaT cell overproliferation through targeting LXR-α and PPAR-γ. Cell Cycle. 2020;19:1928–40. https://doi.org/10.1080/15384101.2020.1783934.
Deng JN, Li YQ, Liu Y, et al. Exosomes derived from plasma of septic patients inhibit apoptosis of T lymphocytes by down-regulating bad via hsa-miRNA-7-5p. Biochem Biophys Res Commun. 2019;513(4):958–66. https://doi.org/10.1016/j.bbrc.2019.04.051.
Xu L, Leng H, Shi X, Ji J, Fu J, Leng H. MiRNA-155 promotes cell proliferation and inhibits apoptosis by PTEN signaling pathway in the psoriasis. Biomed Pharmacother. 2017;90:524–30. https://doi.org/10.1016/j.biopha.2017.03.105.
Wang H, Zhang Y, Luomei J, Huang P, Zhou R, Peng Y. The miRNA-155/GATA3/IL37 axis modulates the production of proinflammatory cytokines upon TNF-α stimulation to affect psoriasis development. Exp Dermatol. 2020;29:647–58. https://doi.org/10.1111/exd.14117.
El-Komy M, Amin I, El-Hawary MS, Saadi D, Shaker O. Upregulation of the miRNA-155, miRNA-210, and miRNA-20b in psoriasis patients and their relation to IL-17. Int J Immunopathol Pharmacol. 2020;34:2058738420933742. https://doi.org/10.1177/2058738420933742.
García-Rodríguez S, Arias-Santiago S, Blasco-Morente G, Orgaz-Molina J, Rosal-Vela A, Navarro P, et al. Increased expression of microRNA-155 in peripheral blood mononuclear cells from psoriasis patients is related to disease activity. J Eur Acad Dermatol Venereol. 2017;31:312–22. https://doi.org/10.1111/jdv.13861.
Li Y, Man X, You L, Xiang Q, Li H, Xu B, et al. Downregulation of PTEN expression in psoriatic lesions. Int J Dermatol. 2014;53:855–60. https://doi.org/10.1111/ijd.12061.
Yan JJ, Qiao M, Li RH, Zhao XT, Wang XY, Sun Q. Downregulation of miRNA-145-5p contributes to hyperproliferation of keratinocytes and skin inflammation in psoriasis. Br J Dermatol. 2019;180(2):365–72. https://doi.org/10.1111/bjd.17256.
Wang Y, Cao Y. miRNA-145-5p inhibits psoriasis progression by regulating the Wnt/β-catenin pathway. Am J Transl Res. 2021;13(9):10439–48 (PMID: 34650713; PMCID: PMC8507052).
Yu X, Yan N, Li Z, Hua Y, Chen W. FGF19 sustains the high proliferative ability of keratinocytes in psoriasis through the regulation of Wnt/GSK-3β/β-catenin signalling via FGFR4. Clin Exp Pharmacol Physiol. 2019;46(8):761–9. https://doi.org/10.1111/1440-1681.13103.
Elamir AM, Shaker OG, El-Komy MH, Mahmoud Sharabi M, Aboraia NM. The role of LncRNA MALAT-1 and MiRNA-9 in psoriasis. Biochem Biophys Rep. 2021;26:101030. https://doi.org/10.1016/j.bbrep.2021.101030.
Chicharro P, Rodríguez-Jiménez P, Llamas-Velasco M, et al. Expression of miRNA-135b in psoriatic skin and its association with disease improvement. Cells. 2020;9(7):1603. https://doi.org/10.3390/cells9071603.
Shirani F, Baghi M, Rostamian Delavar M, Shoaraye Nejati A, Eshaghiyan A, Nasr-Esfahani MH, Peymani M, Ghaedi K. Upregulation of miRNA-9 and miRNA-193b over human Th17 cell differentiation. Mol Genet Genomic Med. 2022;8:e1538. https://doi.org/10.1002/mgg3.1538.
Abdallah HY, Tawfik NZ, Soliman NH, Eldeen LAT. The lncRNA PRINS-miRNA-mRNA axis gene expression profile as a circulating biomarker panel in psoriasis. Mol Diagn Ther. 2022;26(4):451–65. https://doi.org/10.1007/s40291-022-00598-y.
Negroni A, Pierdomenico M, Cucchiara S, Stronati L. NOD2 and inflammation: current insights. J Inflamm Res. 2018;11:49–60. https://doi.org/10.2147/JIR.S137606.
Bhattacharjee O, Ayyangar U, Kurbet AS, Ashok D, Raghavan S. Unraveling the ECM-immune cell crosstalk in skin diseases. Front Cell Dev Biol. 2019;7(7):68. https://doi.org/10.3389/fcell.2019.00068.
Custurone P, Di Bartolomeo L, Irrera N, Borgia F, Altavilla D, Bitto A, Pallio G, Squadrito F, Vaccaro M. Role of cytokines in vitiligo: pathogenesis and possible targets for old and new treatments. Int J Mol Sci. 2021;22(21):11429. https://doi.org/10.3390/ijms222111429.
Zhou X, Chen Y, Cui L, Shi Y, Guo C. Advances in the pathogenesis of psoriasis: from keratinocyte perspective. Cell Death Dis. 2022;13(1):81. https://doi.org/10.1038/s41419-022-04523-3.
Williams MD, Mitchell GM. MicroRNAs in insulin resistance and obesity. Exp Diabetes Res. 2012. https://doi.org/10.1155/2012/484696.
Iacomino G, Siani A. Role of microRNAs in obesity and obesity-related diseases. Genes Nutr. 2017;12:23. https://doi.org/10.1186/s12263-017-0577-z.
Landrier JF, Derghal A, Mounien L. MicroRNAs in obesity and related metabolic disorders. Cells. 2019;8(8):859. https://doi.org/10.3390/cells8080859.
Heyn GS, Corrêa LH, Magalhães KG. The impact of adipose tissue–derived miRNAs in metabolic syndrome, obesity, and cancer. Front Endocrinol. 2020;11:563816. https://doi.org/10.3389/fendo.2020.563816.
almer JD, Soule BP, Simone BA, Zaorsky NG, Jin L, Simone NL. MicroRNA expression altered by diet: Can food be medicinal? Ageing Res Rev. 2014;17:16–24. https://doi.org/10.1016/j.arr.2014.04.005.
Tsigalou C, Vallianou N, Dalamaga M. Autoantibody Production in Obesity: Is There Evidence for a Link Between Obesity and Autoimmunity? Curr Obes Rep. 2020;9(3):245–54. https://doi.org/10.1007/s13679-020-00397-8. (PMID: 32632847).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Committee of Ethics approval no 5091 was granted from the Faculty of Medicine, Suez Canal University, Ismailia, Egypt, and the study was conducted according to the Declaration of Helsinki’s guidelines.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Authors' contributions
AHY, EA and TNZ designed the study, TNZ collected the clinical data, SNH collected the patients’ samples, AHY, ElA, KRM, FS, and SNH carried out the experiments, AHY, EA, KiRM, FS and SNH analyzed and interpreted the patient data. All authors discussed the results, contributed to the final manuscript and approved it.
Conflicts of interest statement
Abdallah HY, Faisal S, Tawfik NZ, Soliman NH, Kishk RM, and Ellawindy A declare that they have no conflicts of interest that might be relevant to the contents of this manuscript.
Code availability
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Abdallah, H.Y., Faisal, S., Tawfik, N.Z. et al. Expression Signature of Immune-Related MicroRNAs in Autoimmune Skin Disease: Psoriasis and Vitiligo Insights. Mol Diagn Ther 27, 405–423 (2023). https://doi.org/10.1007/s40291-023-00646-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-023-00646-1