Abstract
Staphylococcus pseudintermedius (SP) is the major pathogen incriminated in the skin infections of dog. Identification of SP requires molecular methods. The incidence of methicillin resistant SP (MRSP) is increasing worldwide and it is a growing concern in treating pet animals. The prevalence of SP and MRSP from skin infections of dog in India has not been studied previously. Hence, the present study was aimed to isolate SP from common skin infections of dog in Chennai, India and to characterize these isolates. A total of 53 SP organisms were isolated from 91 samples of skin infection accounting for 59 % of isolation rate. Labrador was a major breed from which isolation was made. Panton–Valentine leucotoxin (Luk-I) and S. intermedius exfoliative toxin (siet) genes were detected in all SP isolates but staphylococcal protein A homologue (spsQ) gene was detected only in 36 % of the SP isolates. Out of 53 isolates, 17 % were found to be strong and 19 % to be moderate producers of biofilm and 28 % were classified as MRSP due to possession of the mecA gene. Most isolates were sensitive to tetracycline and ciprofloxacin and least sensitive to erythromycin and trimethoprim/sulphamethaxazole. The authors first time reported the isolation of MRSP, characterization of SP isolates by detecting virulence genes, biofilm forming ability and susceptibility to antimicrobials in Chennai, India.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial pyoderma and otitis are the most common dermatological problems encountered in small animal clinical practice. Staphylococcus sp. is predominantly incriminated in canine skin disease. Among them, coagulase positive Staphylococcus pseudintermedius (SP) is the most important opportunistic commensal pathogen in dogs. SP is a normal inhabitant of skin and mucosal surface of dogs and is commonly obtained from the nares, mouth, groin, axilla and perianal areas [1]. SP is a leading cause of skin and ear infections and it also causes infections of other body tissues and cavities, post-operative wound infections, urinary tract infections and necrotizing faciitis in dogs and cats [2]. SP was first described in 2005 by Devriese [3]. It has been categorized under Staphylococcus intermedius group (SIG) which includes S.intermedius, S.pseudintermedius and S.delphini as they could not be differentiated phenotypically. SP can be differentiated from other members biochemically [4] but there is no phenotypic marker. Hence, molecular methods such as PCR–RFLP [5, 6], MALDI-TOF MS [7] and multiplex PCR [8] are required for species differentiation. PFGE [9], MLST [9], Spa typing [10] and SCCmec [11] are used for strain typing. Incidence of methicillin resistant SP (MRSP) is increasing worldwide but prevalence differs in various geographical locations. Colonization by SP isolates on the skin of dog owners, veterinarians and attending staff has been reported [12]. Moreover, cases of human infections caused by SP have been increasing due to the available molecular methods to identify SP in diagnostic/academic laboratories [13]. As there is no previous study on the isolation and characterization of SP isolates from India, the present study was attempted to determine the prevalence of SP from skin infections of dogs in Chennai, India, to identify MRSP and to detect their virulence genes and antimicrobial susceptibility.
Material and Methods
Bacteria Isolation and Identification
Samples were collected between February 2013 and February 2014 from dogs with skin infections, brought to the Dermatology Unit of Teaching Veterinary Clinical Complex, Madras Veterinary College, Chennai. Isolation and identification of S. pseudintermedius isolates was done as described previously [14].
Detection of Virulence Genes
Virulence factor genes such as S. intermedius exfoliative toxin (siet), Panton Valentine-like toxins (LukS-I and LukF-I) and staphylococcal protein A homologue (SpsQ) were detected by PCR using primers and conditions as given in the Table 1. PCR was performed in a reaction volume of 10 µl containing approximately 100–150 ng of genomic DNA, 5 pmol of each primer and 2 × master mix (Ampliqon, Denmark). Cycling conditions were 94 °C for 3 min, followed by 30 cycles of denaturation at 94 °C for 30 s, annealing at 55/60 °C for 30 s, extension at 72 °C for 30 s and a final extension cycle of 5 min at 72 °C. PCR products were loaded on a 2 % agarose gel for electrophoresis, visualized with ethidium bromide and documented.
Antimicrobial Susceptibility Test
Antimicrobial susceptibility testing was performed by disk diffusion method on Mueller–Hinton agar and isolates were classified as sensitive, intermediate and resistant based on recommendations of CLSI document M100-S21 (M2) [15]. Ampicillin (10 µg), cephotaxime (30 µg), ciprofloxacin (5 µg), erythromycin (15 µg), oxacillin (1 µg), tetracycline (30 µg), trimethoprime (1.25 µg)/sulphamethaxazole (23.75 µg) antimicrobial discs used in the present study were procured from HiMedia Pvt Ltd, India.
Biofilm Formation
Biofilm formation was carried out as per the method described by Hassan et al. [16] using a quantitative spectrophotometric microliter plate assay. The interpretation of biofilm production was done based on the OD value as given below:
OD of isolate ≤ OD of control (ODc) = No biofilm formation; ODc < OD of isolate = Weak biofilm formation; 2× ODc ≤ OD of isolate = Moderate biofilm formation; 4× ODc ≤ OD of isolate = High biofilm formation.
Results and Discussion
Prevalence of S. pseudintermedius Isolates
A total of 91 samples were collected between February 2013 and February 2014 from various skin infections of dogs of different breeds, age and sex. SP was isolated from 53 (59 %) animals. The major breed from which SP was isolated was Labrador (30 %) followed by mixed breed dogs (21 %), Pug (11 %), Spitz (11 %), German Shepherd (9 %) and 17 % of other breeds. A higher rate of isolation of SP have been reported in pyoderma cases from Japan (76 % in 2009) [17], Germany (76 %) [18] and south Korea (61 %) [19]. An almost similar rate of isolation of 52 % from both diseased and healthy dogs was noticed in Poland [21] and 55 % from healthy dogs in Tunisia [22]. However, a lower isolation rate was observed with 40 % in Guide dog school in Finland [23], 16 % from healthy and diseased dogs in south China [24] and 26.5 % from pyoderma cases in north China [25].
Generally, pyoderma caused by SP is either simple infection or complex infection that is associated with underlying disease such as allergies (flea allergy, atopic dermatitis, food allergy), internal disease (hypothyroidism or hyperadrenocortism), seborrhoeic conditions (folliculitis) or parasitic disease (demodicosis). In the present study, 25 isolates were from dogs with simple pyoderma and others with complex pyoderma. Ten dogs were infected with demodicosis, 5 dogs each with otitis and allergy and 4 dogs each with other parasitic infection and other causes.
Age did not significantly influence skin infection caused by SP. Prevalence of SP infection was 40 % in dogs less than 1 year old, 32 % in dogs 2–4 years old and 28 % in dogs more than 4 years old. SP isolates obtained from females (66 %) were twice than that of males (34 %). A similar higher incidence of nasal colonization of S.aureus in females than male dogs was observed in Hong Kong [26]. In another study in USA, no difference in the susceptibility to infection caused by SP among male and female dogs was noticed [27]. The rate of colonization of S. aureus in humans was reported more in males than females in Bangalore city [28] and Andhra Pradesh [29] in India; whereas, the prevalence of MRSA was found to be higher in females than that of males in Agra [30] and Himachal Pradesh [31] in India. Hence, these results of the previous studies on SP or S. aureus colonization/infection in humans and dogs have been variable in regards to age or sex predilection and there is no conclusive evidence.
S. intermedius Exfoliative Toxin (SIET)
All the S. pseudintermedius isolates possessed the S. intermedius exfoliative toxin (siet) gene and this result corroborates with the studies of canine SP isolates of Korea [32], Poland [21] and Tunisia [22]. Dogs injected with purified SIET develop clinical signs such as erythema, exfoliation and crusting, which are signs of canine pyoderma [33]. S. aureus exfoliative toxins are extremely specific serine proteases and function as ‘molecular scissors’ during skin infection and cleave desmosomal cadherins only in the superficial layers of the skin, which is directly responsible for the clinical manifestation of staphylococcal scalded skin syndrome in human. Recent reports demonstrated that 3–4 % of methicillin sensitive S. aureus (MSSA) strains carry the eta or etb gene [34, 35], whereas around 10 % of methicillin resistant S. aureus (MRSA) are eta positive [35]. However, the significance of all SP strains isolated from both healthy and diseased cases possessing siet gene is not known.
Panton and Valentine Leucocidin (PVL) Like Toxin (Luk-I)
Panton and valentine leucocidin (PVL) found in certain strains of S. aureus is a bicomponent—LukS-PV and LukF-PV, pore forming leukotoxin that causes leukocyte destruction and tissue necrosis. A similar bicomponent leukotoxin Luk-I, encoded by two genes, lukS/F, was also detected in SP and toxins were found to be cytotoxic to various polymorphonuclear cells, monocytes and macrophages [20]. All the SP isolates characterized in the present study possessed Luk-I genes. Other studies involving SP isolates from both healthy and diseased dogs were also positive for Luk S/F genes [21, 22, 36, 37]. The clinical sequelae of PVL-positive S. aureus infections tend to be more severe than PVL-negative S. aureus [38]. However, such clarity could not be established in SP isolates as it is present in both healthy and diseased dogs. Studies to detect S. aureus PVL gene in SP isolates from diseased dogs in Switzerland [39] and in Belgium [40] revealed that none of the SP isolates were PVL- positive.
Staphylococcal Protein A Homologue (spsQ)
Staphylococcal protein A homologue gene (spsQ) was detected in 19 (36 %) of the isolates and among them, 8 isolates were MRSP. A higher prevalence of protein A homologue and Clumping Factor of 54.5 % canine S. intermedius strains was observed on latex agglutination test in Japan [41]. However, a lower prevalence of 14.2 % SP isolates from infected dogs and 1.4 % from healthy dogs of Poland was reported for protein A by dot blot assay [21]. These SP isolates from dogs of Poland were also characterized for siet, Luk-I, thermonuclease and agr virulence genes by PCR. Protein A was the only phenotypic pathogenicity factor that distinguished infected and non-infected dogs in their study and it has unambiguously been confirmed that the strains from infected dogs synthesize protein A markedly and more frequently than those from healthy dogs. In the present study, SP from healthy dogs was not studied. Staphylococcal protein A is a cell wall anchored surface protein with four or five domains that each can bind to the Fc region of IgG. The interaction between protein A and IgG coats the surface of the cell with IgG molecules that cannot be recognized by the neutrophil Fc receptor and activates the complement by the classical pathway as a result of incorrect orientation. The study explains the anti-phagocytic effect of protein A in vitro and for that reason it is considered a virulence factor in several models of animal infection.
In S. aureus, staphylococcal protein A is encoded by spa gene where as in SP, two orthologues of spsA–spsP and spsQ were detected [42]. In that study, the prevalence of spsP and spsQ were tested in 20 SP isolates and it was found that spsQ was present in 12/20 strains whereas spsP was present only in 8/20 isolates which also possessed spsQ orthologues. In the present study, primers originally designed to target spsQ were used and further studies are required to understand the prevalence of spsP orthologue and typing of SP isolates of Indian origin. The significance of possessing two orthologues of protein A in SP and its relevance in pathophysiology of SP colonization and infection in dogs need to be investigated. The Xr repeat region spa is widely used to type S.aureus strains for epidemiological analysis and the same technique was adapted to type SP isolates by targeting the Xr repeat region of spsQ [10].
Biofilm Formation
The majority of S. pseudintermedius isolates evaluated in the present study were either weak (30 %, 16/53) or had no ability (34 %, 18/53) to produce biofilm, with only 17 % (9/53) being classified as strong and 19 % (10/53) as moderate biofilm producers. There was no difference between MRSP and MSSP isolates in biofilm formation. However, the number of MRSP isolates evaluated in the study was relatively less. In a Norwegian study [43], all 23 MRSP isolates analyzed produced biofilm and belonged to sequence type (ST) 71, producing significantly more biofilm as compared to other STs. Similarly, all 20 MRSP isolates evaluated by Dicicco et al. [44] formed biofilm and also reported that clarithromycin was ineffective in eradicating MRSP biofilm at therapeutic doses. Out of 140 SP isolates from dogs in Canada and United States, 96 % were able to produce biofilm and the biofilm production was not significantly different amongst isolates from clinical infections as compared to isolates obtained from colonized dogs [45]. Biofilm production by S. pseudintermedius plays an important role in the pathophysiology of disease and in potential colonization. It could be a contributing factor in the rapid and worldwide emergence of MRSP [11].
Antimicrobial Susceptibility
Among the 53 SP isolates, 15 isolates (28 %) were confirmed to be methicillin resistant S. pseudintermedius (MRSP) with the detection of mecA gene. MRSP was first reported in 2005 and since then more isolates of MRSP have been isolated from various countries with differing prevalence rate. The prevalence of MRSP identified in the present study is comparatively lesser than those reported from south China (48 %) [24], north China (44 %) [25] and USA (63.3 %) [46]. However, the prevalence of MRSP varies significantly between regions, as it was observed to be 21 % in Italy [47], 18 % in Korea [19], 11.4 % in Japan [17], 4.6 and 20 % in dogs with various clinical conditions and bitches with different reproductive disorders in Lithuania [48], 7.5 % in Croatia [36], 7.4 % in Germany [18], and no methicillin resistance was detected in SP isolates from canine pyoderma cases in West Indies [49].
In the present study, one mecA negative SP isolate was found to be resistant to oxacillin (Table 2). A similar observation of mecA negative oxacillin resistant SP isolates was reported by Feng et al. [24] and Kang et al. [50]. Hawraa et al. [51] and Elhassan et al. [52] also reported mecA-negative oxacillin resistant staphylococci. The first MRSA strain encoding a divergent mecA gene (now designated as mecC) was discovered from bovine mastitis milk sample and it has also been found in animal and human isolates [53]. The presence of mecC gene was not evaluated in the isolated SP organisms used in the present study. However, these findings provided clear evidence that there are mechanisms other than the presence of mecA gene responsible for methicillin resistance. Hence, it is highly recommended to use both phenotypical traditional Kirby-Bauer method and genotypic mecA and mecC detection by PCR to identify methicillin resistant staphylococci.
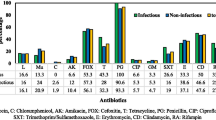
Among other tested antimicrobials, SP isolates were most susceptible to tetracycline (60 %), ciprofloxacin (51 %) and cefotaxime (39 %) and least sensitive to erythromycin (21 %) and trimethoprim/sulphamethoxazole (26 %). Feng et al. [24] reported that more than 70 % of the SP isolates from south China were resistant to erythromycin, trimethoprim and penicillin and >50 % isolates were resistant to ciprofloxacin and enrofloxacin. Out of 74 SP isolates from Korea, the highest antibiotic resistance was observed towards penicillin, tetracycline, trimethoprim and erythromycin [18]. More than 60 % of the SP isolates from Japan in 2009 were found to be resistant to ampicillin and >40 % of the isolates were resistant to kanamycin, tetracycline, enrofloxacin and ofloxacin and only 27 % of the isolates were resistant to erythromycin [16]. Antimicrobial susceptibility study of 106 SP isolates from Croatia revealed that the resistance was more to ampicillin (78 %) followed by kanamycin (42 %), tetracycline (37.7 %), erythromycin (37.7 %), clindamycin (32 %) and chloramphenicol (26.4 %) [20]. Differences in geographical locality, investigation period, method of antimicrobial susceptibility test carried out, variation in the treatment of choice with available antimicrobial drugs in the pertinent country might be the possible reasons for the variations in the reported resistance rate to different antimicrobials.
The present study is the first comprehensive investigation on the isolation of SP and detection of various virulence genes, biofilm forming ability and antimicrobial susceptibility in Indian SP isolates. Further research is required to understand the prevalent type of SP in this geographical region and genetic background of resistance profile for antimicrobials.
Conclusion
The present study revealed the moderate prevalence of MRSP in dogs in India. Among the virulence factors evaluated, SP isolates differed in the presence of staphylococcal protein A homologue gene and biofilm forming ability. But, there was no difference in these factors between SP and MRSP isolates of India. Characterization of other potential virulence factors and the zoonotic capability of this organism to the populations should be ascertained and the public should be educated in how to handle pet dogs and cats that may have staphylococcal infections.
References
Rubin JE, Chirino-Trejo M (2011) Prevalence, sites of colonization, and antimicrobial resistance among Staphylococcus pseudintermedius isolated from healthy dogs in Saskatoon, Canada. J Vet Diagn Investig 23:351–354
Weese JS, van Duijkeren E (2010) Methicillin-resistant Staphylococcus aureus and Staphylococcus pseudintermedius in veterinary medicine. Vet Microbiol 140:418–429
Devriese LA, Vancanneyt M, Baele M, Vaneechoutte M, De Graef E, Snauwaert C, Cleenwerck I, Dawyndt P, Swings J, Decostere A, Haesebrouck F (2005) Staphylococcus pseudintermedius sp. nov., a coagulase-positive species from animals. Int J Syst Evol Microbiol 55(Pt 4):1569–1573
Sasaki T, Kikuchi K, Tanaka Y, Takahashi N, Kamata S, Hiramatsu K (2007) Reclassification of phenotypically identified Staphylococcus intermedius strains. J Clin Microbiol 45:2770–2778
Bannoehr J, Franco A, Iurescia M, Battisti A, Fitzgerald JR (2009) Molecular diagnostic identification of Staphylococcus pseudintermedius. J Clin Microbiol 47:469–471
Blaiotta G, Fusco V, Ercolini D et al (2010) Diversity of Staphylococcus species strains based on partial kat (catalase) gene sequences and design of a PCR-restriction fragment length polymorphism assay for identification and differentiation of coagulase-positive species (S. aureus, S. delphini, S. hyicus, S. intermedius, S. pseudintermedius, and S. schleiferi subsp. coagulans). J Clin Microbiol 48:192–201
Decristophoris P, Fasola A, Benagli C et al (2011) Identification of Staphylococcus intermedius group by MALDI-TOF MS. Syst Appl Microbiol 34:45–51
Sasaki T, Tsubakishita S, Tanaka Y et al (2010) Multiplex-PCR method for species identification of coagulase-positive staphylococci. J Clin Microbiol 48:765–769
Black CC, Solyman SM, Eberlein LC et al (2009) Identification of a predominant multilocus sequence type, pulsed-field gel electrophoresis cluster, and novel staphylococcal chromosomal cassette in clinical isolates of mecA-containing, methicillin-resistant Staphylococcus pseudintermedius. Vet Microbiol 139:333–338
Moodley A, Stegger M, Ben Zakour NL, Fitzgerald JR, Guardabassi L (2009) Tandem repeat sequence analysis of staphylococcal protein A (spa) gene in methicillin-resistant Staphylococcus pseudintermedius. Vet Microbiol 135:320–326
Perreten V, Kadlec K, Schwarz S, Andersson UG, Finn M, Greko C et al (2010) Clonal spread of methicillin-resistant Staphylococcus pseudintermedius in Europe and North America: an international multicentre study. J Antimicrob Chemother 65:1145–1154
vanDuijkeren E, Kamphuis M, van der Mije IC et al (2011) Transmission of methicillin-resistant Staphylococcus pseudintermedius between infected dogs and cats and contact pets, humans and the environment in households and veterinary clinics. Vet Microbiol 150:338–343
Stegmann R, Burnens A, Maranta CA, Perreten V (2010) Human infection associated with methicillin-resistant Staphylococcus pseudintermedius ST71. J Antimicrob Chemother 65:2047–2048
AnandaChitra M, Jayanthy C, Nagarajan B (2015) Detection and sequence analysis of accessory gene regulator genes of Staphylococcus pseudintermedius isolates. Veterinary World 8(7):902–907
Clinical and Laboratory Standards Institute (2011) Performance standards for antimicrobial susceptibility testing; Twenty-first informational supplement. In: CLSI document M100-S21 (M2), vol 30, nos 1, 15. Wayne, Pennsylvania
Hassan A, Usman J, Kaleem F, Omair M, Khalid A, Iqbal M (2011) Evaluation of different detection methods of biofilm formation in the clinical isolates. Braz J Infect Dis. doi:10.1590/S1413-86702011000400002
Onuma K, Tanabe T, Sato H (2011) Antimicrobial resistance of Staphylococcus pseudintermedius isolates from healthy dogs and dogs affected with pyoderma in Japan. Vet Dermatol 23:17–e5
Ruscher C, Lübke-Becker A, Wleklinski CG, Soba A, Wieler LH, Walther B (2009) Prevalence of methicillin-resistant Staphylococcus pseudintermedius isolated from clinical samples of companion animals and equidaes. Vet Microbiol 136:197–201
Yoon JW, Lee KJ, Lee SY, Chae MJ, Park JK, Yoo JH et al (2010) Antibiotic resistance profiles of Staphylococcus pseudintermedius isolates from canine patients in Korea. J Microbiol Biotechnol 20:1764–1768
Futagawa-Saito K, Sugiyama T, Karube S, Sakurai N, Ba-Thein W, Fukuyasu T (2004) Prevalence and characterization of leukotoxin-producing Staphylococcus intermedius in isolates from dogs and pigeons. J Clin Microbiol 42:5324–5326
Garbacz K, Zarnowska S, Piechowicz L, Haras K (2013) Pathogenicity potential of Staphylococcus pseudintermedius strains isolated from canine carriers and from dogs with infection signs. Virulence 4(3):255–259
Gharsa H, Ben Slama K, Lozano C, Gómez-Sanz E, Klibi N, Ben Sallem R, Gómez P, Zarazaga M, Boudabous A, Torres C (2013) Antimicrobial resistance, virulence genes, and genetic lineages of Staphylococcus pseudintermedius in healthy dogs in Tunisia. Microb Ecol 66:363–368
Grönthal T, Ollilainen M, Eklund M, Piiparinen H, Gindonis V, Junnila J, Koulumies LS, Liimatainen R, Rantala M (2015) Epidemiology of methicillin resistant Staphylococcus pseudintermedius in guide dogs in Finland. Acta Vet Scand 57:37. doi:10.1186/s13028-015-0129-8
Feng Y, Tian W, Lin D, Luo Q, Zhou Y, Yang T, Deng Y, Liu YH, Liu JH (2012) Prevalence and characterization of methicillin-resistant Staphylococcus pseudintermedius in pets from South China. Vet Microbiol 160:517–524
Wang Y, Yang J, Logue CM, Liu K, Cao X, Zhang W, Shen J, Wu C (2012) Methicillin-resistant Staphylococcus pseudintermedius isolated from canine pyoderma in North China. J Appl Microbiol 112:623–630
Boost MV, O’Donoghue MM, James A (2008) Prevalence of Staphylococcus aureus carriage among dogs and their owners. Epidemiol Infect 136:953–964
Bryan J, Frank LA, Rohrbach BW, Burgette LJ, Cain CL, Bemis DA (2012) Treatment outcome of dogs with meticillin-resistant and meticillin-susceptible Staphylococcus pseudintermedius pyoderma. Vet Dermatol 23:361–e65
Bouchiat C, El-Zeenni N, Chakrakodi B, Nagaraj S, Arakere G, Etienne J (2015) Epidemiology of Staphylococcus aureus in Bangalore, India: emergence of the ST217 clone and high rate of resistance to erythromycin and ciprofloxacin in the community. New Microbe New Infect 7:15–20
Rao BN, Srinivas B (2012) A prospective study of methicillin resistant Staphylococcus aureus (MRSA) in a teaching hospital of rural setup. J Pharm Sci Innov 1(2):37–40
Goyal A, Diwakar MK, Bhooshan S, Goyal S, Agrawal A (2013) Prevalence and antimicrobial susceptibility pattern of methicillin-resistant Staphylococcus aureus [MRSA] isolates at a Tertiary Care Hospital in Agra, North India—a systemic annual review. IOSR J Dent Med Sci 11(6):80–84
Sharma S, Mall A (2011) The prevalence, antibiogram and characterization methicillin resistant Staphylococcus aureus among the patients from the Doon Valley hospitals. Afr J Microbiol Res 5(21):3446–3451
Yoon JW, Lee GJ, Lee SY, Park C, Yoo JH, Park HM (2010) Prevalence of genes for enterotoxins, toxic shock syndrome toxin 1 and exfoliative toxin among clinical isolates of Staphylococcus pseudintermedius from canine origin. Vet Dermatol 5:484–489
Terauchi R, Sato H, Hasegawa T, Yamaguchi T, AizawaC Maehara N (2003) Isolation of exfoliative toxin from Staphylococcus intermedius and its local toxicity in dogs. Vet Microbiol 94:19–29. doi:10.1016/S0378-1135(03)00048-8
Megevand C, Gervaix A, Heininger U, Berger C, Aebi C et al (2010) Molecular epidemiology of the nasal colonization by methicillin-susceptible Staphylococcus aureus in Swiss children. Clin Microbiol Infect 16:1414–1420
Sila J, Sauer P, Kolar M (2009) Comparison of the prevalence of genes coding for enterotoxins, exfoliatins, panton–valentine leukocidin and tsst-1 between methicillin-resistant and methicillin susceptible isolates of Staphylococcus aureus at the university hospital in Olomouc. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub 153:215–218
Matanović K, eMekić S, Šeol B (2012) Antimicrobial susceptibility of Staphylococcus pseudintermedius isolated from dogs. Veterinarski Arhiv 82(5):505–517
Ruscher C, Lübke-Becker A, Semmler T, Wleklinski CG, Paasch A, Soba A, Stamm I, Kopp P, Wieler LH, Walther B (2010) Widespread rapid emergence of a distinct methicillin- and multidrug-resistant Staphylococcus pseudintermedius (MRSP) genetic lineage in Europe. Vet Microbiol 26:340–346
Gillet Y, Issartel B, Vanhems P, Godail-Gamot et al (2002) Association between Staphylococcus aureus strains carrying gene for panton–valentine leukocidin and highly lethal necrotising pneumonia in young immunocompetent patients. Lancet 359:753–759
Descloux S, Rossano A, Perreten V (2008) Characterization of new staphylococcal cassette chromosome mec (SCCmec) and topoisomerase genes in fluoroquinolone- and methicillin-resistant Staphylococcus pseudintermedius. J Clin Microbiol 46(5):1818–1823
Bardiau M, Yamazaki K, Ote I, Misawa N, Mainil JG (2013) Characterization of methicillin-resistant Staphylococcus pseudintermedius isolated from dogs and cats. Microbiol Immuno 57(7):496–501. doi:10.1111/1348-0421.12059
Futagawa-Saito K, Ba-Thein W, Sakurai N, Fukuyasu T (2006) Prevalence of virulence factors in Staphylococcus intermedius isolates from dogs and pigeons. BMC Vet Res 2:4. doi:10.1186/1746-6148-2-4
Bannoehr J, Ben Zakour NL, Reglinski M, Inglis NF, Prabhakaran S, Fossum E et al (2011) Genomic and surface proteomic analysis of the canine pathogen Staphylococcus pseudintermedius reveals proteins that mediate adherence to the extracellular matrix. Infect Immun 79(8):3074–3086
Osland AM, Vestby LK, Fanuelsen H et al (2012) Clonal diversity and biofilm forming ability of methicillin-resistant Staphylococcus pseudintermedius. J Antimicrob Chemother 67:841–848
Dicicco M, Neethirajan S, Singh A, Weese JS (2012) Efficacy of clarithromycin on biofilm formation of methicillin-resistant Staphylococcus pseudintermedius. BMC Vet Res 8:225
Singh A, Walkar M, Rousaaeau J’, Weese JS (2013) Characterization of the biofilm forming ability of Staphylococcus pseudintermedius from dogs. BMC Vet Res 9:93
Black CC (2010) Methicillin resistance in Staphylococcus pseudintermedius. Ph D Dissertation, University of Tennessee. http://trace.tennessee.edu/utk_graddiss/775
De Lucia M, Moodley A, Latronico F, Giordano A, Caldin M, Fondati A, Guardabassi L (2011) Prevalence of canine methicillin resistant Staphylococcus pseudintermedius in a veterinary diagnostic laboratory in Italy. Res Vet Sci 91:346–348
Ruzauskas M, Couto N, Kerziene S, Siugzdiniene R, Klimiene I, Virgailis M, Pomba C (2015) Prevalence, species distribution and antimicrobial resistance patterns of methicillin-resistant staphylococci in Lithuanian pet animals. Acta Vet Scand 57:27. doi:10.1186/s13028-015-0117-z
Hariharan H, Gibson K, Peterson R, Frankie M, Matthew V, Daniels J, Martin NA, Andrews L, Paterson T, Sharma RN (2014) Staphylococcus pseudintermedius and Staphylococcus schleiferi subspecies coagulans from canine pyoderma cases in Grenada, West Indies, and their susceptibility to beta-lactam drugs. Vet Med Int. doi:10.1155/2014/850126
Kang MH, Chae MJ, Yoon JW, Kim SG, Lee SY, Yoo JH, Park HM (2014) Antibiotic resistance and molecular characterization of ophthalmic Staphylococcus pseudintermedius isolates from dogs. J Vet Sci 15(3):409–415. doi:10.4142/jvs.2014.15.3.409
Hawraa WA, Al-Dulaimi T, Al-Marzoqi AH (2014) Phenotypic detection of resistance in Staphylococcus aureus isolates: detection of (mec A and fem A) gene in methicillin resistant Staphylococcus aureus (MRSA) by polymerase chain reaction. J Nat Sci Res 4:112–118
Elhassan MM, Ozbak HA, Hemeg HA, Elmekk MA, Ahmed LM (2015) Absence of the mecA Gene in methicillin resistant Staphylococcus aureus isolated from different clinical specimens in Shendi City. Bio Med Res Int, Sudan. doi:10.1155/2015/895860
García ÁL, Holden MTM, Lindsay H, Webb CR, Brown DFJ et al (2011) Meticillin resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: a descriptive study. Lancet Infect Dis 11:595–603
Acknowledgments
The authors thankfully acknowledge the Vice-Chancellor, TANUVAS for providing the infrastructure and facilities to carry out this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
AnandaChitra, M., Jayanthy, C. & Nagarajan, B. Virulence Genes Detection and Antimicrobial Susceptibility of Staphylococcus pseudintermedius Isolates from Canine Skin Infection in Chennai, India. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 88, 355–361 (2018). https://doi.org/10.1007/s40011-016-0760-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-016-0760-9