Abstract
Textile industries produce a large volume of dye-contaminated wastewater through the dyeing process, which poses a high risk to the environment. The reactive dyes are commonly used as colorant organic substances as these are chemically stable and give bright colors to fabrics. This study aims to optimize the decolorization of Reactive Red 21 dye from water by Pseudomonas aeruginosa strain 23N1. The two-level factorial design has been applied to select the levels of operational parameters, which has been further optimized by rotatable central composite design-based response surface methodology. The concentration of yeast extract has shown a significant positive effect (highest coefficient estimate of 25.55) on decolorization by the strain, whereas added glucose and peptone as nutrients for bacteria have shown negative effect (coefficient estimate of − 1.71 and − 1.62, respectively) because the bacteria preferably have utilized glucose and peptone as prime carbon/nitrogen source instead of dye molecules. Quadratic model generated through central composite design using experimental data to predict decolorization percentage has shown a good correlation coefficient (R2 = 0.999) and adjusted correlation coefficient (Adj. R2 = 0.992). The decolorization percentages from model validation experiments are achieved as 93.5 ± 0.4% and 91.5 ± 0.5% for initial dye concentrations of 50 mg/L and 150 mg/L, respectively, and have demonstrated satisfactory correlation with model predicted data. Based on the result obtained, it can be suggested that the strain 23N1 could be efficiently used to decolourize RR21 dye-contaminated wastewater.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rapid industrialization has introduced varieties of wastewaters, contaminated with different toxic organic and inorganic constituents resulting in deterioration of the environment and imbalanced ecosystem (Colak et al. 2011; Gupta and Ali 2012; Mishra and Maiti 2018; Basheer 2018). Among all industries, textile industrial wastewater has considered as a great concern due to its large volume and complexity of the toxicants present in it (Wang et al. 2017; Korenak et al. 2018; Ali et al. 2018). An account over 7 × 105 metric tons of synthetic dyes is produced and used worldwide yearly in dyeing process (Colak et al. 2009). Out of which, about > 15% (i.e., 1–5% loss in production and 1–10% loss in handling), i.e., 280 kilotons of dyes, remained unfixed and carried out through wastewater (Yenikaya et al. 2010; Liu et al. 2017a). The commercial dyes, especially reactive azo dyes, have extensive use in dyeing because of their superior characteristics like capable of making brilliant color shades, high wet fastness profiles, less energy requirement and easy to apply (Naseer et al. 2016). A small concentration of dye (10–15 mg/L) in wastewater is even significantly visible and reduces the transparency, gas solubility and the esthetic value of water bodies (Maqbool et al. 2016). However, the conventional physiochemical treatment techniques such as adsorption (Khan et al. 2011; Dehghani et al. 2016; Ali et al. 2016a, b, c), coagulation and flocculation (Slokar and Marechal 1997; Ali et al. 2012), photochemical oxidation (Yang and Wyatt 1998; Burakova et al. 2018), electrochemical oxidation (Pelegrini et al. 1999), and ozonation and membrane filtration (Xu and Lebrun 1999) have been investigated to treat textile wastewater. But those processes have certain demerits like requirement of high energy and cost, inefficient to remove the reactive dyes and produce hazardous by-products (Dafale et al. 2008). Biological treatment is considered a better method, as this is environmentally friendly and efficiently removes or transforms the dye molecules to non-toxic chemicals. In this regard, numerous numbers of potential microorganisms (bacteria, fungi and algae) have been investigated by the researchers to treat dye-contaminated wastewater (Lim et al. 2013a, b; Yang et al. 2016; Das and Kumar 2018). The efficiency of microbial dye decolorization depends on the selected microorganism and their adaptability into the process conditions of culture (Deive et al. 2010; Ma et al. 2013). However, the microorganism like fungi and algae exhibits certain limitations, including extended hydraulic retention period to complete decolorization process and involve adsorption phenomenon to remove dye molecules rather than environmental friendly biotransformation into non-toxic molecules (Kalyani et al. 2009). In contrast, the bacteria could efficiently remove dyes through complete or partial mineralization mechanism under preferable operating conditions along with rapid growth of bacterial population (Bedekar et al. 2014). The growing microorganisms have specific nutritional needs, and when the growth-promoting nutrients are sufficiently available, microorganism even responds well to the toxicity stress and exhibits higher percentage of decolorization (Garg et al. 2012). Nevertheless, it is not easy to predict the optimal condition of process parameters (nutritional ingredients and its amount in the culture at particular pH and temperature) of decolorization using bacteria (Banat et al. 1996). Therefore, studies are needed to elucidate the impact of such operational factors on the growth and performance of microorganism for dye decolorization. In the recent past decades, the effect of experimental parameters on microbial dye decolorization was used to investigate with traditional single-factor optimization method (Zhao et al. 2010). This conventional approach could not significantly analyze the effect of interaction between or among experimental factors. Thus, the statistical methods like two-level factorial experimental design and response surface methodology (RSM) could solve the complexities involved in experimental design and explore the relationship among response and various experimental factors (Du et al. 2010; Ekka et al. 2016). RSM is a statistical technique, in which central composite design (CCD) is extensively used as an analytical optimization tool to achieve the desired response (Kuppusamy et al. 2017). It also reduces the number of experimental trials required to examine the effects of various parameters in dye decolorization process.
Based on the available literature, it has been revealed that dye alone could not be used as the sole carbon and nitrogen source for most of the bacterial species since the organism requires additional carbon and nitrogen sources to co-metabolize the dye (Senthilkumar et al. 2013). A very few bacterial species have the capability to cleave azo bond and could utilize the dye molecules as the sole carbon and energy source (Telke et al. 2010). In the present study, the decolorization of Reactive Red 21 (RR21) dye has been investigated using Pseudomonas aeruginosa strain 23N1 that exhibits the capability to grow using azo dye as the sole carbon and energy source. The synthetic RR21 dye is used widely in textile and leather industries, which adds biotic risk due to environmental contamination through wastewater. Thus, the decolourization of reactive RR21 (azo dye) has been investigated using P. aeruginosa strain 23N1 in the presence and absence of other carbon and nitrogen sources. The effects of operational parameters have been analyzed using two-level factorial design, and the optimization of culture condition has been performed using CCD-based response surface methodology. Later, the kinetics study of decolorization has been also carried out to elucidate the activity of bacteria in decolorization as a function of time. This study aims to optimize a bacterial-based dye decolorization process. It is also to be noted that this study first reports the decolorization of RR21 dye-contaminated water and that by P. aeruginosa strain 23N1. The experiments in this study were performed in Indian Institute of Technology Roorkee, at Saharanpur Campus, Saharanpur, India, in 2017.
Materials and methods
Dyes and chemicals
The azo dye namely Reactive Red 21 (Remazol Red BB or RR21) of CAS number 11099-79-9 (Molecular formula C26H19N4Na3O15S4), used in this study was supplied by Sisco Research Laboratories Pvt. Ltd., Delhi, India, and chemical structure is shown in Fig. S1 of supplementary information (SI). All other chemicals were procured from Himedia Laboratories, Mumbai, India.
Microbial strain and culture conditions
The bacteria P. aeruginosa strain 23N1 of accession number MCC 2622 was procured from National Centre for Cell Sciences (NCCS), Pune, India. The pure culture was maintained in HK34b nutrient agar medium [peptone (5 g/L), NaCl (5 g/L), meat extract (1.5 g/L), yeast extract (1.5 g/L) and agar (15 g/L)]. For inoculums preparation, the strain 23N1 was routinely grown aerobically under shaking condition of 150 rpm, at 30 °C in 100-mL Erlenmeyer flask containing 50 mL broth (0.35% (w/v), yeast extract and 2% (w/v) peptone) for 24 h. This culture was used as inoculums (volume in % (v/v)) in subsequent experiments unless specified (Pathak et al. 2014). The stock solution (1000 mg/L) of Reactive Red 21 dye was prepared by dissolving 1 g/L of dye in double distilled water, and dye solution of desired concentration (50 mg/L, 100 mg/L, 150 mg/L and 200 mg/L) for respective experimental trial was prepared by diluting the stock solution. The initial pH of the solutions was adjusted using 1 N NaOH and 1 N HCl. All the statistical schematics of decolorization experiments were performed in triplicates, in 100-mL Erlenmeyer flask containing 50 mL dye solution supplemented with broth nutrient of particular concentration and inoculums, under static condition. The duration of culture incubation was 48 h, which was kept constant for all experimental runs. The operational variables like pH, incubation temperature, inoculation volume, the concentration of dye, carbon source, nitrogen source, salt concentration and yeast extract concentration have been chosen to investigate the relationship and interactive effects among its on dye decolorization using the regular two-level factorial and RSM design methods.
Analytical techniques
The sample collected from the experimental flask to measure residual dye concentration was first centrifuged at 10,000 rpm for 20 min. The resultant supernatant was analyzed using UV spectrophotometer at 537 nm (λmax of RR21) to detect the residual dye concentration in mg/L. A control of particular experimental run performed under the same condition. The following equation was used to evaluate the dye decolorization percentage:
where Concin is initial dye concentration and Concf is final dye concentration of the supernatant solution of respective sample. The mean value of the responses (% decolorization) from triplicates of respective experimental trial was considered for statistical analysis.
Statistical optimization of decolorization conditions
The decolorization study of RR21 dye-containing water was carried out as per the experimental trials produced through statistical designs using Design-Expert software 7.0 (version 7.1.6, Stat-Ease Inc., Minneapolis, USA). Firstly, the conventional two-level factorial design was used to select the minimum and maximum level of operational parameters and examined their effect on decolorization by strain 23N1. Secondly, after identifying the significant levels of influential factors from two-level factorial design, the response surface methodology (RSM) was carried out to evaluate the required decolorization process condition to achieve an optimal response. In RSM, central composite design (CCD) was chosen for optimization study as it includes three groups of design points as two-level factorial design points, centre points and axial points (Maqbool et al. 2016). Moreover, this technique is cost-effective as it reduces the number of experimental trials required to examine the effect of operational factors and their interaction on a response (Du et al. 2010). The elaborate description of these statistical techniques has written in S1.1 and S.1.2 sections of SI.
Kinetics study
The kinetic study was carried out under an analyzed optimal condition for 50 mg/L and 150 mg/L of initial dye concentrations. The samples of the kinetic study were collected at different predetermined times up to 60 h. A 3 mL sample was withdrawn from culture flask and centrifuged to get the supernatant for analyzing the residual dye concentration. The UV–visible spectra of RR21 dye solutions were recorded in the range of wavelength 200 to 700 nm (λmax of dye: 537 nm) after collecting experimental supernatant solutions during kinetic study to investigate the spectral changes.
Results and discussion
In the preliminary study, the decolorization of RR21 dye by P. aeruginosa strain 23N1 is carried out under both static and shaking (150 rpm) conditions with initial dye solution of 50 mg/L for 48 h of the incubation period. The decolorization percentage of RR21 dye is obtained about 90% under static condition and 55% under shaking condition, when other parameters kept constant (pH: 9, temperature: 40 °C, salt: 3 g/L, and yeast extract: 8 g/L). The lower decolorization might be achieved due to lower azoreductase enzyme activity under shaking aerobic conditions (Oturkar et al. 2011). This observation indicates that strain 23N1 may require a certain amount of oxygen. Similar observation was reported by Sheth and Dave (2009), who investigated the decolorization of Reactive Red BS by P. aeruginosa NGKCTS. Therefore, the static aerobic condition is selected in the subsequent experiments of RR21 dye decolorization using strain 23N1.
Regular two-level factorial design for screening the operational factors
In the screening study, six factors (A-pH, B-glucose, C-peptone, D-salt, E-yeast extract and F-inoculation volume) have been considered for conventional two-level factorial design. All experiments have been carried out using 50 mg/L of initial dye solution incubated at 30 °C for 48 h. The decolorization of RR21 by the strain 23N1 in different culture supplements has shown wide variation in decolorization percentages from 1.46 to 89.97% (Table 1), indicating the high significance of the selected factors within the lower (− 1) and upper (+1) levels taken in this optimization study. The ANOVA results of operational factors, which include F-value, p value of the selected factors and coefficient of the first-order model, are shown in Table S1 of SI. The result indicates that the model with F-value 35.9 is found to be statistically significant (p value < 0.05). The model reliability is verified by computing the correlation coefficient (R2 = 0.987) and adjusted coefficient (Adj R2 = 0.959), which suggests the 95.9% variability in the response variables, that can be elucidated through its relation with the input variables exhibited in the model (Maqbool et al. 2016). The adequacy precision value obtained as 17.98, which indicates an adequate signal for the model to be used to navigate in design space. Moreover, the significance of model is verified by p values < 0.05, and in this case, D, E, F, AE, BE and CE are found to be significant model terms. The estimated coefficient of the factors, standard error, confidence limits and variance inflation factor (VIF) is enlisted in Table S2 of SI. The significance of regression coefficient of each factor is determined by constructing their confidence limit. Among the selected factors, when glucose and peptone are added to culture medium, both have exhibited negative influence on decolorization and their coefficient values are estimated as − 1.71 and − 1.62, respectively. In the presence of glucose and peptone, bacteria have preferably used these as carbon and nitrogen sources instead of dye molecules. Thus, glucose and peptone are not required in culture medium to achieve better decolorization of RR21 dye using strain 23N1. Although the coefficient estimates (1.76) for pH are found lower than the other significant model terms, it has shown positive influence on decolorization. Among the significant model terms, yeast extract observed as the highest decisive influential factor (coefficient estimate = 25.55) on decolorization. The VIF value of 1 obtained for all factors, which indicates the magnitude of multicollinearity in the ordinary least squares regression analysis. Based on the result obtained, the four factors such as pH, salt concentration, the concentration of yeast extract and inoculation volume are revealed out to be major influential factors. These four most influential factors along with dye concentration and temperature have been considered for optimization process through RSM.
Optimization of experimental conditions by central composite design
CCD is a well-known second-order experimental design used for sequential experimentation to get reasonable information for testing the goodness of fit using a reduced number of design points. The complete design matrix and analysis of variance (ANOVA) are shown in Tables 2 and 3, respectively. The actual decolorization percentage is varied widely ~ 0% to 91.5% under defined test conditions. Using the rotatable method, a total number of 33 batch trials (triplet of each trial) of experiments are carried out. The significance of highest order polynomial model is identified through evaluation of “partial sum of squares-Type III” for response data. The analyses provide the information that additional terms in data are significant and the model is not aliased. Besides a moderately lower value of the coefficient of variation (CV% = 5.88), the model has exhibited the high reliability of the experiments (Amini et al. 2008). The lack of fit F-value as 0.027 implies that lack of fit is not significant relative to the pure error. There is an 87.68% chance that a “lack-of-fit F-value” of this large could occur due to the noise. The lack-of-fit test suggests the better use of quadratic model against a two-factor interaction model. Moreover, the F-value of 156.55 along with p value < 0.0001 obtained from quadratic model indicates that applied quadratic model here is highly significant (Dasgupta et al. 2015). The significance of model is reinforced by a higher coefficient of correlation (R2 = 0.999), predicted correlation coefficient (Pred. R2 = 0.987), adjusted correlation coefficient (Adj. R2 = 0.992) and adequacy precision (Adeq. precision = 43.67), which indicate an adequate signal (Hafshejani et al. 2014). ANOVA result of this quadratic model suggests that the model could be worked out to navigate the design space. In this case, C, D, E, AD, AE, BC, BE, BF, CE, CF, EF, A2, B2, C2, D2 and E2 are significant model terms and those have p value < 0.05 (Li et al. 2014). By applying multiple regressions, an appropriate second-order polynomial equation of quadratic model is obtained in terms of the coded factors as:
The fitted model equation includes the contribution of all six factors in the form of six linear terms such as A, B, C, D, E and F. The six quadratic terms are obtained as A2, B2, C2, D2, E2 and F2. The fifteen interaction terms are obtained as AB, BC, DE, CF and others. In the quadratic model equation, all the linear terms have a positive contribution to the response. Although the interaction terms BC, CD, CF and EF have negative contribution along with coefficients of quadratic terms B2 and C2, corresponding linear terms have shown a positive significant. Furthermore, the significance of correlation coefficients of model terms is obtained by constructing the confidence limits, shown in Table S3 of SI. All the regression coefficients are achieved within the confidence limits. Among the selected factors, pH and temperature have displayed the lowest coefficient value as 0.68 and 0.57, respectively, and those are found as nonsignificant factors in the studied range. However, it is known that the bacteria will grow well at temperature range 25–40 °C and in slightly alkaline medium (Prasad and Aikat 2014). Thus, these factors are needed to be optimized during the experiment to provide a suitable environment for bacterial growth. Among the significant model terms, amount of yeast extract has shown the highest positive influence (coefficient estimate = 20.87) on decolorization than any other factors. Based on the impacts of these variables and contribution of factors in the above equation, it can be suggested that this quadratic model could elucidate the interrelationship among all investigated factors and well predict the corresponding response as percentage of decolorization.
Usually, it should be validated to ensure the suitability of the generated model for providing an adequate estimate of required process condition to achieve a target response for real system (Lim et al. 2013a, b). As mentioned above, the diagnostic plots are constructed for a better understanding of ANOVA and to judge the adequacy of a model, shown in Fig. S2 of SI. The normal plot of residual is constructed between internally studentized residuals and normal percentage probability values, shown in Fig. S2a of SI. The graph shows that the residuals follow a normal distribution along the straight line, which reveals that a transformation of the response may provide a better analysis (Jadhav et al. 2015). The graph of actual values versus predicted values indicates a good fit, shown in Fig. S2b of SI. The Box–Cox plot is an indication of selecting the correct power law transformation based on the best lambda, shown in Fig. S2c of SI. In this study, the 95% confidence interval around this lambda includes value 1. Thus, it is not required to transform further the observed response for model fitting (Dasgupta et al. 2015; Ayed et al. 2017). The plot of Cook’s distances (shown in Fig. S2d of SI) provides a measure of changes in regression if any case is deleted, which helps in identifying an outlier. However, no outlier is found in this study, as the Cook’s distances are within < 1 value. Moreover, all leverage numerical values obtained within the standard range (shown in Fig. S2e of SI) indicate the potential for a design point to influence the model fit.
The perturbation plot exhibiting the comparative effect of all the factors at a particular point in the design space is illustrated in Fig. 1. This plot helps to identify the factors that most affect the response. A significant steep slope or curvature of factors A, D, B, C and E shows that the decolorization response is sensitive to those factors. The higher gradient of factor E (concentration of yeast extract) indicates its high sensitivity to the response. The yeast extract acts as a nitrogen source and vitamins and contributes to the other growth factors which are essential for bacterial growth (Maqbool et al. 2016). Thus, factor E is found to be one of the most influential on the decolorization percentage of RR21 dye.
Effect of operational parameters on decolorization of dye
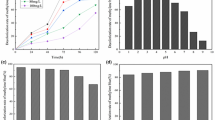
It is well known that the glucose and peptone have been considered an important component of dye decolorization medium (Khan et al. 2013; Sheth and Dave 2009). In this study, the two-level factorial analysis revealed that glucose and peptone have the negative effect on decolorization of RR21. Thus, these were not added in culture medium in the subsequent studies. In contrast, the amount of salt and yeast extract has shown positive influence on decolorization. Imran et al. 2016 reported the enhanced decolorization of azo dye in medium incorporated with yeast extract, which simulates the activity of azoreductase enzyme responsible for the degradation of dye. In the recent past, two opinions have been argued to classify the roles of dyes in cellular metabolic processes: One supports that dyes are not carbon sources, while another considers it contrarily (Deive et al. 2010). The present result reveals that the bacteria have used dye as a carbon source from culture medium when containing no other external carbon sources, resulting in high decolorization compared to solutions containing carbon sources like glucose and peptone because in glucose- and peptone-supplemented medium, bacteria have used up those nutrients as a prime carbon source instead of dye molecules. Moreover, the effect of interaction among operational parameters (temperature, pH, salt concentration, dye concentration, the concentration of yeast extracts and inoculation volume) on RR21 decolorization by the strain is studied through CCD. The 3D response surface plots are constructed during CCD analysis at certain limited operational condition (pH: 9, salt: 6 g/L, dye concentration: 100 mg/L, temperature: 40 °C, yeast extract concentration: 9 g/L and inoculation volume: 5.5%) and are shown in Fig. 2. The 3D response surface plots used to understand the interactive effects of inoculation volume, dye concentration and temperature on the response are shown in Fig. 3. In the subsequent study, the operating conditions such as pH 9, 4 g/L of salt, 100 mg/L of initial dye concentration, temperature 40 °C and 9 g/L of yeast extract are considered. The 2D contour plots of individual operational factors obtained during CCD analysis are shown in Fig. S3 of SI. The plots are constructed based on the regression model, keeping one factor constant at its zero level, while varying the other two factors to see the effects of those factors on percentage decolorization of RR21 dye by strain 23N1. The interactive effect of salt concentration and pH on the decolorization RR21 is shown in Fig. 2a. The elliptical or saddle shape of the plot indicates the significant interaction between corresponding factors (Wang and Liu 2008). An increase in decolorization percentage observed in alkaline pH > 7 medium supplemented with a salt concentration around 6 mg/L, which indicates that strain could effectively function in high salt medium (Liu et al. 2017b). Figure 2b demonstrates the less significant interaction between pH and concentration of yeast extract, whereas the amount of yeast extract shows significantly effect on percent decolorization of dye. In this consequence, increasing the concentration of yeast extract and alkalinity of medium up to pH 9 could significantly increase the decolorization performance (Chen et al. 2011; Krishnan et al. 2016). A similar influence on the response is also observed in case of interaction between dye and salt concentrations as shown in Fig. 2c, which exhibit decreases in decolorization percentage with a decrease in salt concentration (Wang et al. 2013). Besides, a weak interaction has been observed between dye and salt concentrations. The temperature and dye concentration have shown strong influence individually on decolorization performance, although their interactive influence is weak (Fig. 2d). The high decolorization percentage is observed at a temperature near 40 °C, but it has been inhibited on either increasing the temperature beyond 40 °C or lowering below 20 °C. At higher temperatures above 40 °C and below 20 °C, the metabolic activity of bacteria reduces, which ultimately decreases the decolorization performance (Sharma et al. 2009a, b; Wei et al. 2011). Moreover, it is well known that bacteria could grow significantly in the temperature range of 25–40 °C (Pearce et al. 2003; Prasad and Aikat 2014). Furthermore, the decolorization process has shown significant changes due to the slight change in either temperature or concentration of yeast extract, as shown in Fig. 2e. This indicates that the higher decolorization only could be achieved at optimum temperature when sufficient concentration of yeast extract has been added into the medium (Pandey et al. 2007). However, the concentration of yeast extract should increase with an increase in dye concentration to achieve higher decolorization, as shown in Fig. 2f. The effect of interaction between inoculation volume and temperature shown in Fig. 3a is found to be very significant. It has been observed that a small change in percentage of inoculation volume could exhibit a higher change in decolorization at a higher temperature (40 °C). Zhao et al. (2010) reported a similar trend, who investigated the bio-decolorization of Acid Red GR by Dyella ginsengisoli LA-4. At a lower temperature, it is required to increase the percentage of inoculation volume by sufficiently higher value to achieve the same extent of decolorization using same initial dye concentration, as shown in Fig. 3b. The response clearly indicates the strong impact of temperature on the performance of bacteria during decolorization process.
3D response plots for operational factors exhibiting interactive effects on decolorization of RR21 using Pseudomonas aeruginosa 23N1: a pH and salt; b amount of yeast extract and pH; c amount of salt and dye concentration; d dye concentration and temperature; e amount of yeast extract and temperature; f amount of yeast extract and dye concentration
Validation of optimized quadratic model
To validate the experimental design model, some confirmatory experiments were performed under optimized static condition. In the first confirmatory test, the optimal operating conditions are taken as follows: pH 9, temperature 40 °C, 50 mg/L of dye concentration, 6 g/L of salt, 8.4 g/L of yeast extract and 7.6% of inoculation volume, while, for the second experiment, the operational conditions were fixed as follows: pH 8.8, temperature 40 °C, 150 mg/L of dye concentration, 3.7 g/L of salt concentration, 9 g/L of yeast extract loading and 3% of inoculation volume. Both the experiments are conducted in triplet to validate the proposed quadratic model. The predicted responses of decolorization of RR21 from the quadratic model for first and second confirmatory experimental runs are obtained as 93.9% and 91.4%, respectively. In this study, the mean values of actual decolorization percentage for first and second confirmatory experimental runs are found to be as 93.5 ± 0.4% and 91.5 ± 0.5%, respectively. Thus, it is confirmed that the proposed RSM quadratic model could be used to assess the required operational factors to achieve a desire decolorization of RR21 by P. aeruginosa strain 23N1 or vice versa.
Kinetics study of decolorization of dye
The kinetic study is performed to investigate the rate of decolorization of RR21 dye by the strain 23N1. The optimal operational factors considered for decolorization kinetic study experiment are same as the confirmatory experiment described in earlier section. The time intervals considered for the collection and analysis of samples are as 1 h, 2 h, 3 h, 4 h, 6 h, 8 h, 12 h, 16 h, 20 h, 24 h, 28 h, 32 h, 36 h, 40 h, 48 h and 60 h. During this study, it is observed that rate of decolorization of dye increases after 8 h of incubation and gradually achieves higher rate during the period 12–40 h. The curves of decolorization percentage as a function of ln(t) (t is the incubation time an hour) in a kinetic study are presented in Fig. 4a. It is clear from Fig. 4a that kinetic data for both initial dye concentrations (50 and 150 mg/L) have shown same kinetic behavior. For both cases, three distinct kinetic steps are visible as: first, slow decolorization from 0 to 12 h; second, most rapid decolorization step between 12 and 48 h; and third, very slow step beyond 48 h. The slopes of the three regions are obtained through linear regression which reveals the rate of decolorization of the corresponding step. The first step has shown slower kinetics compared to the second step due to lag phase for generation of sufficient bacteria to consume dyes molecules as food. The second step of the kinetic is fastest due to more availability of high amount of dye molecules and availability of sufficient amount of matured microorganism. The kinetic rate in the last step, beyond 48 h, is slowest due to unavailability of the dye molecules in the culture media. The slopes for the three steps with an initial dye concentration of 50 mg/L are obtained as 5.3, 54.16 and 0.089, respectively. It is shown that the rate of log/exponential phase is highest among other two steps of the decolorization process. The slowest rate is observed in the last steps, where dye concentration diminished almost. Furthermore, the decolorization percentage data obtained during the kinetic study for initial dye solution of 50 mg/L and 150 mg/L are also fitted well to the empirical kinetic model, elaborately discussed in S.2. section of SI. The empirical kinetic model graph plotted between decolorization percentages versus time (h) for initial dye solution of 50 mg/L and 150 mg/L, and it is shown in Fig. S4 of SI.
Based on the available literature reported on the topic of microbial decolorization of dyes, it has been found that decolorization by the living cell occurs due to either the biodegradation, bioaccumulation or biotransformation of the dyes (Khataee et al. 2010). In such cases, spectral changes arise during the UV–visible analysis of experimental samples, which might lead to either disappearance of absorption peak or appearance of a new absorption peak, while if bioaccumulation causes the decolorization, then the absorption peak will decrease in the equal proportion of percent decolorization during the experimental period (Parshetti et al. 2010; Ong et al. 2012). The samples collected during the kinetic study are used to analyze the spectral changes in absorption peak of dye using UV–visible analysis (200–700 nm). The maximum absorption peak of the RR21 dye solution obtained at 537 nm, which remained unaltered for 8 h of the incubation period. However, a change in absorption peak is significantly observed at 12 h and in later time intervals of the incubation period. The spectral change data obtained during analysis are presented in Fig. 4b. The absorption peak almost diminished after 40 h of the incubation period. Thus, it could be concluded that the decolorization of RR21 dye could be due to the bioaccumulation/degradation by the strain 23N1. However, the mechanism of decolorization needs to be confirmed by the analysis of intermediate metabolites through advance analytical techniques such as HPLC and GC–MS to elucidate the metabolic system involved in this process.
Conclusion
The two-level factorial design and response surface methodology (RSM) are successfully applied to screen out and optimize the influential operational factors to achieve efficient decolorization of Reactive Red 21 (RR21) dye by P. aeruginosa strain 23N1 from water. During the response analysis in two-level factorial design, glucose and peptone are found to exhibit a negative effect on decolorization of RR21 dye. The result obtained from rotatable central composite design (CCD)-based RSM clearly indicates the high positive significance of yeast extract (coefficient estimate of 20.87) supplement for decolorization of RR21 dye. The quadratic model equation obtained from CCD could be applied to elucidate the decolorization process of RR21 by the studied bacterial strain. Moreover, the maximum decolorization potential of strain is found to be 93.5 ± 0.4% and 91.5 ± 0.5% for initial dye solution of 50 and 150 mg/L, respectively, in the presence of yeast extract and salt-supplemented alkaline medium under a static condition at 40 °C for 48 h of the incubation period. Furthermore, UV–visible analysis of RR21 dye decolorization with respect to time indicates the spectral change particularly in the maximum absorption peak of dye, which reveals that decolorization might be due to the biodegradation/accumulation of dye by the strain 23N1. However, the mechanism of decolorization needs to be elucidated by identification of involved microbial machinery, metabolic pathway and resultant products, through biological and advanced analytical techniques. Based on the results obtained in this study, it is suggested that P. aeruginosa strain 23N1 could use for effective treatment of RR21 dye-contaminated wastewater.
References
Ali I, Gupta VK, Khan TA, Asim M (2012) Removal of arsenate from aqueous solution by electro-coagulation method using Al–Fe electrodes. Int J Electrochem Sci 7:1898–1907
Ali I, AL-Othman ZA, Alwarthan A (2016a) Molecular uptake of congo red dye from water on iron composite nano particles. J Mol Liq 224:171–176
Ali I, AL-Othman ZA, Alwarthan A (2016b) Green synthesis of functionalized iron nano particles and molecular liquid phase adsorption of Ametryn from water. J Mol Liq 221:1168–1174
Ali I, AL-Othman ZA, Alwarthan A (2016c) Removal of secbumeton herbicide from water on composite nanoadsorbent. Desal Water Treat 57:10409–10421
Ali I, Alharbi OML, Alothman ZA, Badjah AY, Alwarthan A, Basheer AA (2018) Artificial neural network modelling of amido black dye sorption on iron composite nano material: kinetics and thermodynamics studies. J Mol Liq 250:1–8
Amini M, Younesi H, Bahramifar N, Lorestani AAZ, Ghorbani F, Daneshi A, Sharifzadeh M (2008) Application of response surface methodology for optimization of lead biosorption in an aqueous solution by Aspergillus niger. J Hazard Mat 154:694–702
Ayed L, Bekir K, Achour S, Cheref A, Bakhrouf A (2017) Exploring bioaugmentation strategies for azo dye CI Reactive Violet 5 decolorization using bacterial mixture: dye response surface methodology. Water Environ J 31:80–89
Banat IM, Nigam P, Singh D, Marchant R (1996) Microbial decolorization of textile dye-containing effluents: a review. Bioresour Technol 58:217–227
Basheer AA (2018) Chemical chiral pollution: impact on the society and science and need of the regulations in the 21st century. Chirality 30:402–406. https://doi.org/10.1002/chir.22808
Bedekar PA, Saratale RG, Saratale GD, Govindwar SP (2014) Oxidative stress response in dye degrading bacterium Lysinibacillus sp. RGS exposed to Reactive Orange 16, degradation of RO16 and evaluation of toxicity. Environ Sci Pollut Res 21:11075–11085
Burakova EA, Ali I, Dyachkova TP, Rukhov AV, Tugolukov EN, Galunin EV, Tkachev AG, Basheer AA (2018) Novel and economic method of carbon nanotubes synthesis on a nickel magnesium oxide catalyst using microwave radiation. J Mol Liq 253:340–346
Chen G, Huang MH, Chen L, Chen DH (2011) A batch decolorization and kinetic study of reactive black 5 by a bacterial strain Enterobacter sp. GY-1. Int Biodeterior Biodegrad 65:790–796
Colak F, Atar N, Olgun A (2009) Biosorption of acidic dyes from aqueous solution by Paenibacillus macerans: kinetic, thermodynamic and equilibrium studies. Chem Eng J 150:122–130
Colak F, Atar N, Yazıcıoglu D, Olgun A (2011) Biosorption of lead from aqueous solutions by Bacillus strains possessing heavy-metal resistance. Chem Eng J 173:422–428
Dafale N, Wate S, Meshram S, Nandy T (2008) Kinetic study approach of remazol black-B use for the development of two-stage anoxic–oxic reactor for decolorization/biodegradation of azo dyes by activated bacterial consortium. J Hazard Mat 159:319–328
Das AJ, Kumar R (2018) Bioslurry phase remediation of petroleum-contaminated soil using potato peels powder through biosurfactant producing Bacillus licheniformis J1. Int J Environ Sci Technol 15:525–532
Dasgupta J, Singh M, Sikder J, Padarthi V, Chakraborty S, Curcio S (2015) Response surface-optimized removal of Reactive Red 120 dye from its aqueous solutions using polyethyleneimine enhanced ultrafiltration. Ecotoxicol Environ Saf 121:271–278
Dehghani MH, Sanaei D, Ali I, Bhatnagar A (2016) Removal of chromium(VI) from aqueous solution using treated waste newspaper as a low-cost adsorbent: kinetic modeling and isotherm studies. J Mol Liq 215:671–679
Deive FJ, Dominguez A, Barrio T, Moscoso F, Moran P, Longo MA, Sanroman MA (2010) Decolorization of dye reactive black 5 by newly isolated thermophilic microorganisms from geothermal sites in Galicia (Spain). J Hazard Mater 182:735–742
Du LN, Yang YY, Li G, Wang S, Jia XM, Zhao YH (2010) Optimization of heavy metal-containing dye Acid Black 172 decolorization by Pseudomonas sp. DY1 using statistical designs. Int Biodeterior Biodegrad 64:566–573
Ekka B, Sahu MK, Patel RK, Dash P (2016) Titania coated silica nanocomposite prepared via encapsulation method for the degradation of Safranin-O dye from aqueous solution: optimization using statistical design. Water Resour Ind. https://doi.org/10.1016/j.wri.2016.08.001
Garg SK, Tripathi M, Singh SK, Tiwari JK (2012) Biodecolorization of textile dye effluent by Pseudomonas putida SKG-1 (MTCC 10510) under the conditions optimized for monoazo dye orange II color removal in simulated minimal salt medium. Int Biodeterior Biodegrad 74:24–35
Gupta VK, Ali I (2012) Environmental water: advances in treatment, remediation and recycling. Elsevier, Amsterdam. ISBN 9780444594037
Hafshejani MK, Ogugbue CJ, Morad N (2014) Application of response surface methodology for optimization of decolorization and mineralization of triazo dye Direct Blue 71 by Pseudomonas aeruginosa. 3 Biotech 4:605–619
Imran M, Arshad M, Negm F, Khalid A, Shaharoona B, Hussain S, Nadeem SM, Crowley DE (2016) Yeast extract promotes decolorization of azo dyes by stimulating azoreductase activity in Shewanella sp. Strain IFN4. Ecotoxicol Environ Saf 124:42–49
Jadhav SB, Chougule AS, Shah DP, Pereira CS, Jadhav JP (2015) Application of response surface methodology for the optimization of textile effluent biodecolorization and its toxicity perspectives using plant toxicity, plasmid nicking assays. Clean Technol Environ Policy 17:709–720
Kalyani DC, Telke AA, Dhanve RS, Jadhav JP (2009) Ecofriendly biodegradation and detoxification of Reactive Red 2 textile dye by newly isolated Pseudomonas sp. SUK1. J Hazard Mater 163:735–742
Khan TA, Sharma S, Ali I (2011) Adsorption of Rhodamine B dye from aqueous solution onto acid activated mango (Mangifera indica) leaf powder: equilibrium, kinetic and thermodynamic studies. J Toxicol Environ Health Sci 3:286–297
Khan R, Bhawana P, Fulekar MH (2013) Microbial decolorization and degradation of synthetic dyes: a review. Rev Environ Sci Biotechnol 12:75–97
Khataee AR, Dehghan G, Ebadi A, Zarei M, Pourhassan M (2010) Biological treatment of a dye solution by Macroalgae Chara sp.: effect of operational parameters, intermediates identification and artificial neural network modeling. Bioresour Technol 101:2252–2258
Korenak J, Ploder J, Trcek J, Helix-Nielsen C, Petrinic I (2018) Decolourisations and biodegradations of model azo dye solutions using a sequence batch reactor, followed by ultrafiltration. Int J Environ Sci Technol 15:483–492
Krishnan J, Kishore AA, Suresh A, Madhumeetha B, Prakash DG (2016) Effect of pH, inoculum dose and initial dye concentration on the removal of azo dye mixture under aerobic conditions. Int Biodeterior Biodegrad. https://doi.org/10.1016/j.ibiod.2016.11.024
Kuppusamy S, Sethurajan M, Kadarkarai M, Aruliah R (2017) Biodecolorization of textile dyes by novel, indigenous Pseudomonas stutzeri MN1 and Acinetobacter baumannii MN3. J Environ Chem Eng 5:716–724
Li H, Zhang R, Tang L, Zhang J, Mao Z (2014) Evaluation of Bacillus sp. MZS10 for decolorizing Azure B dye and its decolorization mechanism. J Environ Sci 26:1125–1134
Lim CK, Bay HH, Aris A, Majid ZA, Ibrahim Z (2013a) Biosorption and biodegradation of Acid Orange 7 by Enterococcus faecalis strain ZL: optimization by response surface methodological approach. Environ Sci Pollut Res 20:5056–5066
Lim CK, Bay HH, Aris A, Majid ZA, Ibrahim Z (2013b) Biosorption and biodegradation of Acid Orange 7 by Enterococcus faecalis strain ZL: optimization by response surface methodological approach. Environ Sci Pollut Res 20:5056–5066
Liu C, You Y, Zhao R, Sun D, Zhang P, Jiang J, Zhu A, Liu W (2017a) Biosurfactant production from Pseudomonas taiwanensis L1011 and its application in accelerating the chemical and biological decolorization of azo dyes. Ecotoxicol Environ Saf 145:8–15
Liu W, Liu C, Liu L, You Y, Jiang J, Zhou Z, Dong Z (2017b) Simultaneous decolorization of sulfonated azo dyes and reduction of hexavalent chromium under high salt condition by a newly isolated salt tolerant strain Bacillus circulans BWL1061. Ecotoxicol Environ Saf 141:9–16
Ma C, Zhou S, Lu Q, Yang G, Wang D, Zhuang L, Li F, Lei F (2013) Decolorization of Orange I under alkaline and anaerobic conditions by a newly isolated humus-reducing bacterium, Planococcus sp. MC01. Int Biodeterior Biodegrad 83:17–24
Maqbool Z, Hussain S, Ahmad T, Nadeem H, Imran M, Khalid A, Abid M, Martin-Laurent F (2016) Use of RSM modeling for optimizing decolorization of simulated textile wastewater by Pseudomonas aeruginosa strain ZM130 capable of simultaneous removal of reactive dyes and hexavalent chromium. Environ Sci Pollut Res 23:11224–11239
Mishra S, Maiti A (2018) The efficacy of bacterial species to decolourise reactive azo, anthraquinone and triphenylmethane dyes from wastewater: a review. Environ Sci Poll Res. 25:8286–8314
Naseer A, Nosheen S, Kiran S, Kamal S, Javaid MA, Mustafa M, Tahir A (2016) Degradation and detoxification of Navy Blue CBF dye by native bacterial communities: an environmental bioremedial approach. Desalination Water Treat 57:24070–24082
Ong SA, Toorisaka E, Hirata M, Hano T (2012) Decolorization of Orange II using an anaerobic sequencing batch reactor with and without co-substrates. J Environ Sci 24:291–296
Oturkar CC, Nemade HN, Mulik PM, Patole MS, Hawaldar RR, Gawai KR (2011) Mechanistic investigation of decolorization and degradation of reactive red 120 by Bacillus lentus BI377. Bioresour Technol 102:758–764
Pandey A, Singh P, Iyengar L (2007) Bacterial decolorization and degradation of azo dyes. Int Biodeterior Biodegrad 59:73–84
Parshetti GK, Telke AA, Kalyani DC, Govindwar SP (2010) Decolorization and detoxification of sulfonated azo dye methyl orange by Kocuria rosea MTCC 1532. J Hazard Mater 176:503–509
Pathak H, Soni D, Chauhan K (2014) Evaluation of in vitro efficacy for decolorization and degradation of commercial azo dye RB-B by Morganella sp. HK-1 isolated from dye contaminated industrial landfill. Chemosphere 105:126–132
Pearce CI, Lloyd JR, Guthrie JT (2003) The removal of colour from textile wastewater using whole bacterial cells: a review. Dyes Pigments 58:179–196
Pelegrini R, Peralto-Zamora P, De Andrade AR, Reyers J, Duran N (1999) Electrochemically assisted photocatalytic degradation of reactive dyes. Appl Catal B: Environ 22:83–90
Prasad SS, Aikat K (2014) Study of bio-degradation and bio-decolorization of azo dye by Enterobacter sp. SXCR. Environ Technol 35:956–965
Senthilkumar S, Prabhu HJ, Perumalsamy M (2013) Response surface optimization for biodegradation of textile azo dyes using isolated bacterial strain Pseudomonas sp. Arab J Sci Eng 38:2279–2291
Sharma P, Singh L, Dilbaghi N (2009a) Optimization of process variables for decolorization of Disperse Yellow 211 by Bacillus subtilis using Box–Behnken design. J Hazard Mater 164:1024–1029
Sharma S, Malik A, Satya S (2009b) Application of response surface methodology (RSM) for optimization of nutrient supplementation for Cr(VI) removal by Aspergillus lentulus AML05. J Hazard Mater 164:1198–1204
Sheth NT, Dave SR (2009) Optimization for enhanced decolorization and degradation of reactive red BS C.I. 111 by Pseudomonas aeruginosa NGKCTS. Biodegradation 20:827–836
Slokar YM, Marechal AM (1997) Methods of decolorization of textile wastewaters. Dyes Pigments 37:335–356
Telke A, Joshi S, Jadhav S, Tamboli D, Govindwar S (2010) Decolorization and detoxification of Congo red and textile industry effluent by an isolated bacterium Pseudomonas sp. SU-EBT. Biodegradation 21:283–296
Wang ZW, Liu XL (2008) Medium optimization for antifungal active substances production from a newly isolated Paenibacillus sp. using response surface methodology. Bioresour Technol 99:8245–8251
Wang ZW, Liang JS, Liang Y (2013) Decolorization of reactive black 5 by a newly isolated bacterium Bacillus sp. YZU1. Int Biodeterior Biodegrad 76:41–48
Wang N, Chu Y, Wu F, Zhao Z, Xu X (2017) Decolorization and degradation of Congo red by a newly isolated white rot fungus, Ceriporia lacerata, from decayed mulberry branches. Int Biodeterior Biodegrad 117:236–244
Wei Y, Mu J, Zhu X, Gao Q, Zhang Y (2011) Decolorization of dye solutions with Ruditapes philippinarum conglutination mud and the isolated bacteria. J Environ Sci 23(Supplement):S142–S145
Xu Y, Lebrun RE (1999) Treatment of textile dye plant effluent by nanofiltration membrane. Sep Sci Technol 34:2501–2519
Yang Y, Wyatt DT (1998) Decolorization of textile dyestuffs using UV/H2O2 photochemical oxidation technology. Text Chem Color 30:27–35
Yang X, Zheng J, Lu Y, Jia R (2016) Degradation and detoxification of the triphenylmethane dye malachite green catalyzed by crude manganese peroxidase from Irpex lacteus F17. Environ Sci Pollut Res 23:9585–9597
Yenikaya C, Atar E, Olgun A, Atar N, İlhan S, Colak F (2010) Biosorption study of anionic dyes from aqueous solutions using Bacillus amyloliquefaciens. Eng Life Sci 10:233–241
Zhao L, Zhou J, Jia Y, Chen J (2010) Biodecolorization of Acid Red GR by a newly isolated Dyella ginsengisoli LA-4 using response surface methodology. J Hazard Mater 181:602–608
Acknowledgement
The authors acknowledge the financial support from Ministry of Human Resource Development (MHRD), Government of India, for student scholarship and funding support through a project (IMPRINT scheme, Project No. 6125) along with Ministry of Urban Development, New Delhi, India.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest is reported by the authors.
Additional information
Editorial responsibility: Necip Atar.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mishra, S., Maiti, A. Optimization of process parameters to enhance the bio-decolorization of Reactive Red 21 by Pseudomonas aeruginosa 23N1. Int. J. Environ. Sci. Technol. 16, 6685–6698 (2019). https://doi.org/10.1007/s13762-018-2023-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13762-018-2023-1