Abstract
Plants have been gifted with intricate regulatory networks to carry on with their sessile life form. Often such networks involve delicate association between various proteins. The WD40 proteins, which are present abundantly in several eukaryotes, act as scaffolding molecules assisting proper activity of other proteins. They comprise several stretches of 44–60 amino acid residues and often terminate with a WD dipeptide. They function in several cellular, metabolic and molecular pathways, biologically playing important roles in plant development and also during stress signaling. Moreover, some WD40 (named DWD) proteins also function as substrate receptors in Cullin4 RING dependent E3 ubiquitin ligase mediated proteosomal degradation and DNA damage repair mechanism. In this review, we have discussed the various aspects of these proteins that affect their highly diversified functions in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
WD40 protein—definition and architecture
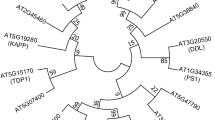
Formation of large heteromeric protein complexes involves regulatory interactions, which are mainly controlled by scaffolding proteins in living system. The WD40 proteins, which are also named as WD-repeats, are such group of proteins which are a prominent feature of diverse protein-protein interactions (Gibson 2009). These proteins are very abundant in eukaryotic organisms but are rarely present in prokaryotes (e.g. Thermomonospora curvata and Cyanobacterium synechocystis; Stirnimann et al. 2010). The SMART database predicted 349 WD40 domain containing proteins in humans (Letunic et al. 2009). In plants like Arabidopsis thaliana, Oryza sativa and Setaria italica more than 200 putative WD40 domain containing proteins have been predicted (Van Nocker and Ludwig 2003; Ouyang et al. 2012), which can help out to understand their cellular networking system and roles in biological molecular machinery. A phylogenetic tree of some representative WD40 genes from these plants (including some of the functionally characterized genes from Arabidopsis) brings out three major WD40 groups (Fig. 1).
Phylogenetic tree of WD40 repeats proteins. The deduced full-length amino acid sequences of 141 WD40 genes (having 81 Arabidopsis thaliana, 59 Oryza sativa and 1 Setaria italica) were aligned by ClustalW and the phylogenetic tree was constructed using MEGA 5.05 by the Neighbor-Joining (NJ) method with 1,000 bootstrap replicates. The evolutionary distances were computed using the Poisson correction method and the optimal tree with the sum of branch length = 77.88547303. Genes are mentioned by their accession numbers and names of the functionally characterized genes are given in parenthesis
Structurally, the WD40 domain is characterized by presence of several copies of WD40 repeats with each repeat containing 44–60 residue units (Fig. 2a). Each unit contains a glycine-histidine (GH) dipeptide about 11–24 residues from its N-terminus and terminates with Trp-Asp (WD) doublet residues at the C-terminus (Neer et al. 1994; Smith et al. 1999). Each of the repeat folds into four-stranded anti-parallel β-sheet and is thought to arise from intragenic duplication and recombination events and diversify during evolution (Andrade et al. 2001). It was therefore thought that the sequences and structures of the repeats in the same protein are more similar to each other than those from different proteins.
Structure of WD40 protein. a WD proteins contain several repeats of 44–60 amino acid residues and each repeat comprises four antiparallel β sheets with conserved GH residues at N-terminal and WD at C-terminal. The residues shown by stars are hydrophobic amino acid frequently present in top surface and participate in interaction with other peptide (b) Seven-bladed β-propeller structure (most stable form) made by using PyMOL (http://www.pymol.org) and (c) Four stranded antiparallel β sheet structure
Generally, the WD40 domains have been shown to contain seven or multiples of seven repeats forming a highly stable β-propeller structure (Fig. 2b). Some researchers have also predicted proteins enclosing as low as two to as many as sixteen repeats (Van Nocker and Ludwig 2003; Saeki et al. 2006; Mishra et al. 2012). On the basis of the geometry modeling, it was predicted that the 7-fold β-propeller is the most stable β-sheet geometry and this form always dominates in the resolved WD40 structures and identified WD40 proteins (Murzin 1992). Proteins with less than 7 repeats form an incomplete β-propeller structure and require additional WD-repeats from its neighbours to stabilize themselves. For instance, two complex structures, Seh1-Nup85 and Sec13-Nup145 where Seh1 (component of nuclear pore complex) and Sec13 (component of the Coat protein II) display a six-bladed open propeller structure, whereas the two Nup (component of nucleoporin pair) proteins possess a three stranded β-blade as well as helical trunk and crow modules. Domain invasion motif (DIM) of the Nup proteins inserts into the opening between blades of corresponding protein and hence completes the seven-bladed WD40 propeller structure (Hsia et al. 2007; Brohawn et al. 2008).
The role of WD40 protein is to provide a rigid platform for interactions of proteins with other cellular components. These interactions aid in controlling several vital functions of the cell, such as signaling cascades, cellular transport and apoptosis (Xu and Min 2011). The sequences to the exterior of the repeats help to establish protein specificity. They also coordinate downstream events, such as ubiquitination and histone methylation and have also been known to function in several plant developmental processes.
The WD-repeat is also called β-transducin repeat, since the first crystal structure having WD40 repeat protein was bovine β-transducin (one subunit of the trimeric G protein transducin complex; Lambright et al. 1996; Sondek et al. 1996). This Gβ subunit is involved in signal transduction and forms a seven-blade β-propeller structure containing seven WD40 repeats. In plants some other WD40 proteins include the Tightly Associated Factor (TAFII), CULLIN4-E3 ubiquitin ligase, Transparent Testa Glabra 1 (TTG1), Constitutively Photomorphogenic 1 (COP1) and Suppressor of PhyA-105 (SPA1), Anthocyanin11 (AN11), and Pale Aleurone Color1 (PAC1). These proteins have reported to be involved in light signaling/photomorphogenesis and flowering (von Arnim et al. 1997; Hoecker et al. 1999; Holm et al. 2002), respectively. Some others like, GTP binding protein β1 is involved in auxin response and cell division (Ullah et al. 2003), Fasciata2 (FAS2) functions in meristem maintenance (Kaya et al. 2001), Leunig (LUG) functions in floral development (Franks et al. 2002), while Fertilization-Independent Endosperm (FIE) regulates seed development and flowering (Sorensen et al. 2001). Another WD40 protein Cyclophilin71 which interacts with histone H3 has a role in chromatin-based gene silencing and organogenesis (Li et al. 2007).
Abundance of WD40 domains in eukaryotes and their action as common interactor
Among the different eukaryotic proteomes, WD40 domains form one of the most copious types along with RNA recognition motif, Zinc fingers, RING-finger and immunoglobulin (Stirnimann et al. 2010). The WD40 proteins occupy 1% of the total proteome in Homo sapiens, 1.4% in Saccharomyces cereviasiae and >0.8% in Arabidopsis (Stirnimann et al. 2010). Some extra functional or catalytic domains are often found in most WD40 proteins such as those involved in protein degradation (F-BOX, Really Interesting New Gene-finger; RING, and suppressors of cytokine signaling domain; SOCS), microtubule dynamics (Lissencephaly type 1-like Homology; LisH), phospholipid binding (Fab1, YOTB/ZK632.12, Vac1 and EEA1 a class of zinc finger domains; FYVE) and vesicle coating (CLathrin Heavy chain repeat homology; CLH) (Stirnimann et al. 2010). The functionality of WD40 protein lies mainly in top surface residues which are created by domain part of protein. WD40 propellers are large domains of about 300 amino acids having three distinct surfaces available for interactions: the top region of the propeller, which is defined as the part of the structure, where the loops connecting D and A strands of the WD-repeats lie, the bottom region, and the circumference (Stirnimann et al. 2010) (Fig. 2c). The most common interacting site of WD40 protein is located on top small surface (formed by N-terminal of repeat) of propeller structure. Three residues of the repeat close to the propeller axis are normally concerned in this interaction. They are positioned at the beginning of strand A (mostly two residues) and at the end of strand B (mostly one residue; Xu and Min 2011). Deletion mutation leads to the conclusion that there is no apparent folding order for each repeat and that the order in which repeats fold might vary among different WD40 proteins, or even within the same protein (Garcia-Higuera et al. 1998). Single amino acid substitutions in one repeat that affect local folding thus might not have drastic consequences for the overall folding process. Structural clustering of interaction interfaces (Aloy et al. 2003) within structural domains indicates that there are 30 interfaces (distinct families with diverse function) for 86 distinct WD40 domains or 0.34 interfaces per domain. The other common families have much lower values (0.07 or 86/1216 for kinases; 0.06 or 15/242 for PDZ domains; 0.10 or 31/310 for SH3 domains), signifying greater interface diversity for the WD40 domain family. Structure and their interacting peptide are mentioned in Table 1.
Protein–protein and protein–peptide interactions involve the entry site to the central channel of the β-propeller, corresponding to the ‘supersite’ for the β-propeller superfold, where the majority of interaction partners (including small molecules) bind (Russell et al. 1998). N- or C-terminal extensions of the β-propeller also pack against the entry sites of the central channel, while the core of the channel is not accessible for interactions. WD40 domains can thus act as large interaction platforms for multiple proteins, making them preferably suited to be hubs in cellular interaction networks. Thus, in comparison to other domains, the proteins containing WD motif are components of several interaction pairs. Datasets of yeast two-hybrid [i.e. binary interaction (Yu et al. 2008)] and of mass spectroscopy/tandem-affinity purification (i.e. multi-protein complex) experiments (Collins et al. 2007) provide evidence for highly interaction mediated by these proteins. Remarkably, the number of WD40 domain-containing complexes found in the MS/TAP datasets easily surpasses the corresponding number of binary interactions in the yeast two-hybrid datasets, suggesting that WD40 proteins act as scaffolds for assemblies of larger complexes.
DWD (DDB1-binding WD40) protein
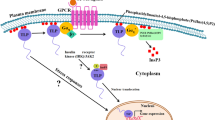
A subset of WD40 proteins have been named as DWD [Damaged DNA binding (DDB) WD40] based on their interaction with DDB1 and CULLIN4 (CUL4). These proteins contain 16 conserved amino acids referred as “DWD box” (Angers et al. 2006; Higa et al. 2006). They have some key characteristics, such as: difference in the number of motifs, importance of the Arg residue following the WD dipeptide (Scrima et al. 2008) and competence of binding with DDB1 provided for some additional residues besides the Arg. DDB complex was first identified as a heterodimer involved in DNA repair, and consists of the 127 kDa subunit DDB1 and a 48 kDa subunit DDB2. DDB1 is recognized as the linker protein in cullin4-RING ubiquitin ligase complexes (CRL4) and thus play additional roles beyond DNA repair. The term DCAF (DDB1-CUL4 Associated Factor) is used for all DDB1 interactors except CUL4. DWD proteins have been reported as substrate receptors for CRL4, a family of multi-subunit E3 ligases. This CUL4 E3 ligase is involved in ubiquitination of target proteins for degradation by the 26S proteasome and also UV damage repair mechanism. In our own molecular docking analysis we found that the circumference of β–propeller structure of SiWD40 (Setaria italica WD40) was involved in an interaction with a SiCullin4 protein (Mishra et al. 2012). The CRLs are a huge subfamily of CUL4-based E3 ubiquitin ligases. It consists of three core subunits: CULLIN4 (CUL4), a RING finger protein Regulator of Cullins1 (ROC1)/RING-BOX1 (RBX1), and UV-Damaged DNA Binding Protein1 (DDB1) (Lee and Zhou 2007) (Fig. 3). RING fingers help in transfer of ubiquitin directly from the E2 to the substrate by allosterically activating the E2. Two different isoforms of CUL4 are expressed from a single gene in Arabidopsis, one (CUL4-L) has 50 amino acids extra at the N-terminus as compared to the other (CUL4-S) (Jackson and Xiong 2009).
Assembly of Cullin 4 RING E3 ligase (CLR4 ligase). Cullin 4 E3 is multisubunit ubiquitin ligase function in proteosomal degradation pathway and UV DNA damage repair mechanism. In the case of cullin4-E3 complex, DNA Damaged Binding1 (DDB1) and DDB1 WD40 (DWD) proteins are utilized as adaptors and substrate receptors, respectively. Here RING fingers (RBX1) help the transfer of ubiquitin (Ub) directly from the E2 to the substrate by allosterically activating the E2 and DWD transfer substrate from E2 to DDB1
In CRL4 ligase, DDB1 serves as the adaptor protein connecting the CUL4/RBX1 to the substrate receptors. In cullin E3 ligase, various types of DWD (also known as DCAF) protein serve as substrate receptors crucial for the recognition of variety of substrate (Lee and Zhou 2007; He et al. 2006). Most DWD proteins contain WD40 repeats and within the repeats, a conserved 16 amino acid block (called the DWD domain) generates the docking site for DDB1. In the 16 amino acid block, there are four highly conserved residues: Asp (or Glu) 7, Trp (or Tyr) 13, Asp (or Glu) 14, and Arg (or Lys) 16 (Fig. 4). There are hydrophobic residues at positions 1, 2, 10, 12, and 15 and small residues at positions 3, 4, and 5 (He et al. 2006). Database searches using the DWD box predicted that there are as many as 90 DWD proteins in human genome (He et al. 2006). Likewise 33 DWD proteins have been predicted in fission yeast, 36 in C. elegans, 75 in Drosophila, 78 in rice, 85 in Arabidopsis and 82 in foxtail millet (He et al. 2006; Lee et al. 2008). Further, in Arabidopsis, 101 DWD domains were identified, of which 69 had one DWD domain and 16 had two. This correlates with the fact that DWD proteins usually possess one and sometimes two but rarely three DWD motifs (He et al. 2006). Additionally, in rice, 96 DWD motifs were present of which 14 had two while two of the proteins had three motifs. Moreover, some other conserved domains were found in 15 Arabidopsis and 9 rice DWD proteins which may regulate interaction with substrate proteins.
AtDDB2 and AtCSA-1 (an ortholog of the mammalian Cockayne Syndrome type-A), encode two related WD40 proteins. Both along with RBX1 form CRL4 complex, [CSA1-DDB2-CUL4A/B-RBX1 (CRL4DDB2/CSA) ligases], were reported as crucial components in UV-B DNA damage repair mechanism in Arabidopsis thaliana (Biedermann and Hellmann 2010; Zhang et al. 2010; Fischer et al. 2011). Here CUL4-CSA1 ubiquitin ligase targets CDT1 (chromatin licensing and DNA replication factor 1) for ubiquitination in response to UV irradiation. It also ubiquitinates histone H2A, histone H3 and histone H4 at sites of UV-induced DNA damage (Wang et al. 2006). Histone ubiquitination assists their exclusion from the nucleosome assembly and consequently helps in DNA repair. Hence ubiquitination of histone does not lead to degradation, instead assist in repair mechanism.
Role of WD40 protein in development and cellular diversity
A protein complex comprising MYB and bHLH transcription factors linked with a WD40 repeat protein regulates the anthocyanin biosynthetic pathway in Antirrhinum majus, Petunia hybrida and Arabidopsis thaliana (de Vetten et al. 1997; Walker et al. 1999; Quattrocchio et al. 1999; Morita et al. 2006; Schwinn et al. 2006). In Arabidopsis, the WD40 repeat protein TTG1 is vital to several aspects of epidermal cell fate, positively regulating trichome formation, anthocyanin production, seed-coat pigmentation and seed-coat mucilage production, and negatively regulating hypocotyl stomatal-cell identity and root-hair formation (perhaps by initiating the differentiation of alternative cell fates). Ramsay and Glover (2005) suggested TTG1 to function as a scaffold for protein–protein interactions between the bHLH and MYB proteins. A seven WD-repeats nucleolar protein named yaozhe (YAO) is essential for the correct positioning of the first zygotic division plane and plays a significant role in gametogenesis in Arabidopsis. Its counterparts in yeast and human are components of the U3 small nucleolar ribonucleoproteins (snoRNP) complex, hence YAO is probably involved in rRNA processing in plants as well (Li et al. 2010). Another five WD40-repeat protein OsLIS-L1 contains a LisH domain at C terminus along with a domain of four WD40 repeats homologous to the β subunit of trimeric G-proteins (Gβ). OsLIS-L1 plays an important role in male gametophyte formation and the first internode elongation in rice (Gao et al. 2011). Another WD40 repeat protein (NEDD1) from Arabidopsis was found to support cell division by interacting with the γ-tubulin complex (Zeng et al. 2009). This protein arranges spindle microtubules (MTs) preferentially towards spindle poles and phragmoplast MTs toward their minus ends and helps in organization of spindles during mitosis (Zeng et al. 2009). The An11 family of WD40 proteins is highly conserved between plant and animals. In plants, the An11 protein is known to be involved in the regulation of anthocyan biosynthesis (de Vetten et al. 1997).
Role in post-translational modification
Embryonic Ectoderm Development (EED) is a WD40 protein responsible for the methyltransferase activity of Polycomb Repressor Complex 2 (PRC2; a histone methyltransferase) complex and directs it’s binding to histone H3 tail (Margueron et al. 2009; Xu et al. 2010). The interaction of EED with H3K27me3 marks enhanced histone methyltransferase activity of PRC2, which in turn maintains transcriptional silencing whereas interaction with H1K26me3 inhibits the PRC2 methyltransferase activity (Xu et al. 2010). Chromatin-based gene silencing provides a vital mechanism for the regulation of gene expression. Li et al. (2007) identified a WD40 repeat protein named CYCLOPHILIN71 (CYP71), and showed its utility in gene repression and organogenesis in Arabidopsis. Methylation of H3K27 of chromatin histone H3 is directly mediated by CYP71 in some regulatory genes, including KNAT1 (knotted1-like gene) and STM (SHOOT MERISTEMLESS) that regulate meristem activity and organogenesis in Arabidopsis. Because CYP71 is highly conserved among eukaryotes ranging from fission yeast to mammals, its homologs may function as universal histone remodeling factors required for epigenetic gene silencing (Zhu et al. 2008). MSI1 a WD40 protein in Arabidopsis has also been proposed to exhibit pleiotropic phenotypes by epigenetic regulation (Hennig et al. 2003; Alexandre et al. 2009). WD40 β-propeller is a class of ubiquitin-binding domains especially found in member of the F box family of SCF ubiquitin E3 ligase adaptors (Pashkova et al. 2010).
Role in abiotic stress tolerance
Zhu et al. (2008) reported that an Arabidopsis WD40-repeat protein (HOS15), functioned through chromatin histone deacetylation for repressing abiotic stress related genes. ABA and abiotic stress treatment increases the expression level of HOS15. Salt and ABA treatment leads to upregulation of many stress-responsive genes in hos15 mutant which is higher than wild type. Hence chromatin remodeling by means of HOS15 (caused by deacetylation of histone H4) signifies the role of this gene in abiotic stress tolerances in plants (Zhu et al. 2008). A salt-responsive WD40 protein in rice (LOC_Os08g38880) having five WD40 repeats is regulated by salt stress and contains several stress responsive cis-acting elements in upstream region of gene (Huang et al. 2008). BnSWD1 (Brassica napus salt responsive WD40 1) contains eight WD40 repeats and is found to be conserved in all eukaryotes. This gene expresses at high levels under salt-stress conditions and is also upregulated after various hormonal treatments such as abscisic acid, salicylic acid, and methyl jasmonate (Lee et al. 2010).
The myoinositol signaling pathway has been correlated to stress and developmental processes in plants. WD40 repeat region of a myoinositol polyphosphate 5-phosphatase (At1g05630) was found to interact with the sucrose nonfermenting-1-related kinase, which has a central role in sugar, metabolic, stress, and developmental signals in Arabidopsis (Ananieva et al. 2008). Further, we have recently shown that a WD40 repeat containing gene (homologous with salt responsive WD40 protein of rice) is upregulated during different abiotic stresses in foxtail millet. It also shows the DNA binding activity with an AP2 domain containing protein, SiDREB gene (Mishra et al. 2012).
Conclusions
The WD40 domain is one of the most abundant and highly conserved domains across eukaryotes. Given their rich interaction surfaces, these proteins function as an adaptor in many different protein complexes or protein-DNA complexes in very diverse cellular processes. They mediate molecular recognition events mainly through the smaller top surface of domain which comprises three residues and form the transient complexes with other peptide. DWD protein function as substrate receptor for CUL4 mediated E3 ligase dependant in proteasomal degradation of protein. Some WD40 proteins control the post-translation event through methylation and uniquitination of histone protein. Besides, these proteins also play role in development specific event and abiotic stress tolerance in plant. Due to larger subunits assemblies and lack of measurable intrinsic activity (such as catalysis), these proteins are difficult to handle. However, structural details and functional mechanism of most WD40 protein in plants is still unclear which is critical for understanding their role in cellular process. We thus definitely need to explore more about the WD40 domain protein complexes in the future.
Abbreviations
- DIM:
-
Domain invasion motif
- TAF:
-
Tightly associated factors
- DDB1:
-
DNA Damaged Binding1
- EED:
-
Embryonic Ectoderm Development
References
Alexandre C, Moller-Steinbach Y, Schonrock N, Gruissem W, Hennig L (2009) Arabidopsis MSI1 is required for negative regulation of the response to drought stress. Mol Plant 2:675–687
Aloy P, Ceulemans H, Stark A, Russell RB (2003) The relationship between sequence and interaction divergence in proteins. J Mol Biol 332:989–998
Ananieva EA, Gillaspy GE, Ely A, Burnette RN, Erickson FL (2008) Interaction of the WD40 domain of a myoinositol polyphosphate 5-phosphatase with SnRK1 links inositol, sugar, and stress signaling. Plant Physiol 148:1868–1882
Andrade MA, Perez-Iratxeta C, Ponting CP (2001) Protein repeats: structures, functions, and evolution. J Struct Biol 134:117–131
Angers S, Li T, Yi X, MacCoss MJ, Moon RT, Zheng N (2006) Molecular architecture and assembly of the DDB1-CUL4A ubiquitin ligase machinery. Nature 443:590–593
Biedermann S, Hellmann H (2010) The DDB1a interacting proteins ATCSA-1 and DDB2 are critical factors for UV-B tolerance and genomic integrity in Arabidopsis thaliana. Plant J 62:404–415
Brohawn SG, Leksa NC, Spear ED, Rajashankar KR, Schwartz TU (2008) Structural evidence for common ancestry of the nuclear pore complex and vesicle coats. Science 322:1369–1373
Collins SR, Kemmeren P, Zhaog XC, Greenblatth JF, Spencerg F, Holstegee FC, Weissman JS, Krogan NJ (2007) Toward a comprehensive atlas of the physical interactome of Saccharomyces cerevisiae. Mol Cell Proteomics 6:439–450
Couture JF, Collazo E, Trievel RC (2006) Molecular recognition of histone H3 by the WD40 protein WDR5. Nat Struct Mol Biol 13:698–703
Davis TL, Bonacci TM, Sprang SR, Smrcka AV (2005) Structural and molecular characterization of a preferred protein interaction surface on G protein βγ subunits. Biochemistry 44:10593–10604
de Vetten N, Quattrocchio F, Mol J, Koes R (1997) The an11 locus controlling flower pigmentation in petunia encodes a novel WD-repeat protein conserved in yeast, plants, and animals. Genes Dev 11:1422–1434
Eytan E, Moshe Y, Braunstein I, Hershko A (2006) Roles of the anaphase-promoting complex/cyclosome and of its activator Cdc20 in functional substrate binding. Proc Natl Acad Sci USA 103:2081–2086
Fischer ES, Scrima A, Böhm K, Matsumoto S, Lingaraju GM, Faty M, Yasuda T, Cavadini S, Wakasugi M, Hanaoka F, Iwai S, Gut H, Sugasawa K, Thomä NH (2011) The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147:1024–1039
Franks RG, Wang C, Levin JZ, Liu Z (2002) SEUSS, a member of a novel family of plant regulatory proteins, represses floral homeotic gene expression with LEUNIG. Development 129:253–263
Gao X, Chen Z, Zhang J, Li X, Chen G, Li X, Wu C (2011) OsLIS-L1 encoding a lissencephaly type-1-like protein with WD40 repeats is required for plant height and male gametophyte formation in rice. Planta 235:713–727
Garcia-Higuera I, Gaitatzes C, Smith TF, Neer EJ (1998) Folding a WD repeat propeller. Role of highly conserved aspartic acid residues in the G protein beta subunit and Sec13. J Biol Chem 273:9041–9049
Gibson TJ (2009) Cell regulation: determined to signal discrete cooperation. Trends Biochem Sci 34:471–482
He YJ, McCall CM, Hu J, Xiong Y (2006) DDB1 functions as a linker to recruit receptor WD40 proteins to CUL4–ROC1 ubiquitin ligases. Genes Dev 20:2949–2954
Hennig L, Taranto P, Walser M, Schonrock N, Gruissem W (2003) Arabidopsis MSI1 is required for epigenetic maintenance of reproductive development. Development 130:2555–2565
Higa LA, Wu M, Ye T, Kobayashi R, Sun H, Zhang H (2006) CUL4-DDB1 ubiquitin ligase interacts with multiple WD40-repeat proteins and regulates histone methylation. Nat Cell Biol 8:1277–1283
Hoecker U, Tepperman JM, Quail PH (1999) SPA1, a WD-repeat protein specific to phytochrome A signal transduction. Science 284:496–499
Holm M, Hardtke CS, Gaudet R, Deng XW (2001) Identification of a structural motif that confers specific interaction with the WD40 repeat domain of Arabidopsis COP1. EMBO J 20:118–127
Holm M, Ma LG, Qu LJ, Deng XW (2002) Two interacting bZIP proteins are direct targets of COP1-mediated control of light dependent gene expression in Arabidopsis. Genes Dev 16:1247–1259
Hsia KC, Stavropoulos P, Blobel G, Hoelz A (2007) Architecture of a coat for the nuclear pore membrane. Cell 131:1313–1326
Huang J, Wang MM, Bao YM, Sun SJ, Pan LJ, Zhang HS (2008) SRWD: a novel WD40 protein subfamily regulated by salt stress in rice (Oryza sativa L.). Gene 424:71–79
Jackson S, Xiong Y (2009) CRL4s: the CUL4-RING E3 ubiquitin ligases. Trends Biochem Sci 34:562–570
Jennings BH, Pickles LM, Wainwright SM, Roe SM, Pearl LH, Ish-Horowicz D (2006) Molecular recognition of transcriptional repressor motifs by the WD domain of the Groucho/TLE corepressor. Mol Cell 22:645–655
Kaya H, Shibahara KI, Taoka KI, Iwabuchi M, Stillman B, Araki T (2001) FASCIATA genes for chromatin assembly factor-1 in Arabidopsis maintain the cellular organization of apical meristems. Cell 104:131–142
Lambright DG, Sondek J, Bohm A, Skiba NP, Hamm HE, Sigler PB (1996) The 2.0A° crystal structure of a heterotrimeric G protein. Nature 379:311–319
Lee J, Zhou P (2007) DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol Cell 26:775–780
Lee JH, Terzaghia W, Gusmarolia G, Charrona JBF, Yoon HJ, Chen H, He YJ, Xiong Y, Deng XW (2008) Characterization of Arabidopsis and rice DWD proteins and their roles as substrate receptors for CUL4-RING E3 ubiquitin ligases. Plant Cell 20:152–167
Lee S, Lee J, Paek KH, Kwon SY, Cho HS, Kim SJ, Park JM (2010) A novel WD40 protein, BnSWD1, is involved in salt stress in Brassica napus. Plant Biotechnol Rep 4:165–172
Letunic I, Doerks T, Bork P (2009) SMART 6: recent updates and new developments. Nucleic Acids Res 37:D229–D232
Li H, He Z, Lu G, Lee SC, Alonso J, Ecker JR, Luan S (2007) A WD40 domain cyclophilin interacts with histone H3 and functions in gene repression and organogenesis in Arabidopsis. Plant Cell 19:2403–2416
Li HJ, Liu NY, Shi DQ, Liu J, Yang WC (2010) YAO is a nucleolar WD40-repeat protein critical for embryogenesis and gametogenesis in Arabidopsis. BMC Plant Biol 10:169
Margueron R, Justin N, Ohno K, Sharpe ML, Son J, Drury WJ, Voigt P, Martin SR, Taylor WR, De Marco V, Pirrotta V, Reinberg D, Gamblin SJ (2009) Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461:762–767
Mishra AK, Puranik S, Bahadur RP, Prasad M (2012) The DNA-binding activity of an AP2 protein is involved in transcriptional regulation of a stress-responsive gene, SiWD40, in foxtail millet. Genomics. doi:10.1016/j.ygeno.2012.06.012
Morita Y, Saitoh M, Hoshino A, Nitasaka E, Iida S (2006) Isolation of cDNAs for R2R3- MYB, bHLH and WDR transcriptional regulators and identification of c and ca mutations conferring white flowers in the Japanese morning glory. Plant Cell Physiol 47:457–470
Mukai S, Ghaedi K, Fujiki Y (2002) Intracellular localization, function, and dysfunction of the peroxisome-targeting signal type 2 receptor, Pex7p, in mammalian cells. J Biol Chem 277:9548–9561
Murzin AG (1992) Structural principles for the propeller assembly of beta-sheets: the preference for seven-fold symmetry. Proteins 14:191–201
Murzina NV, Pei XY, Zhang W, Sparkes M, Vicente-Garcia J, Pratap JV, McLaughlin SH, Ben-Shahar TR, Verreault A, Luisi BF, Laue ED (2008) Structural basis for the recognition of histone H4 by the histone-chaperone RbAp46. Structure 16:1077–1085
Neer EJ, Schmidt CJ, Nambudripad R, Smith TF (1994) The ancient regulatory-protein family of WD-repeat proteins. Nature 371:297–300
Oliver AW, Swift S, Lord CJ, Ashworth A, Pearl LH (2009) Structural basis for recruitment of BRCA2 by PALB2. EMBO Rep 10:990–996
Orlicky S, Tang X, Willems A, Tyers M, Sicheri F (2003) Structural basis for phosphodependent substrate selection and orientation by the SCFCdc4 ubiquitin ligase. Cell 112:243–256
Ouyang Y, Huang X, Lu Z, Yao J (2012) Genomic survey, expression profile and co-expression network analysis of OsWD40 family in rice. BMC Genomics 13:100
Pashkova N, Gakhar L, Winistorfer SC, Yu L, Ramaswamy S, Piper RC (2010) WD40 repeat propellers define a ubiquitin-binding domain that regulates turnover of F box proteins. Mol Cell 40:433–443
Quattrocchio F, Wing J, van der Woude K, Souer E, de Vetten N, Mol J, Koes R (1999) Molecular analysis of the anthocyanin2 gene of petunia and its role in the evolution of flower colour. Plant Cell 11:1433–1444
Ramsay NA, Glover BJ (2005) MYB–bHLH–WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10:63–70
Russell RB, Sasieni PD, Sternberg MJE (1998) Supersites within superfolds. Binding site similarity in the absence of homology. J Mol Biol 282:903–918
Saeki M, Irie Y, Ni L, Yoshida M, Itsuki M, Kamisaki M (2006) Monad, a WD40 repeat protein, promotes apoptosis induced by TNF-α. Biochem Biophys Res Commun 342:568–572
Schwinn K, Venail J, Shang Y, Mackay S, Alm V, Butelli E, Oyama R, Bailey P, Davies K, Martin C (2006) A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 18:831–851
Scrima A, Konícková R, Czyzewski BK, Kawasaki Y, Jeffrey PD, Groisman R, Nakatani Y, Iwai S, Pavletich NP, Thomä NH (2008) Structural basis of UV DNA-damage recognition by the DDB1-DDB2 complex. Cell 135:1213–1223
Smith TF, Gaitatzes C, Saxena K, Neer EJ (1999) The WD repeat: a common architecture for diverse functions. Trends Biochem Sci 24:181–185
Sondek J, Bohm A, Lambright DG, Hamm HE, Sigler PB (1996) Crystal structure of a G-protein beta gamma dimer at 2.1A0 resolution. Nature 379:369–374
Sorensen MB, Chaudhury AM, Robert H, Bancharel E, Berger F (2001) Polycomb group genes control pattern formation in plant seed. Curr Biol 11:277–281
Stankewich MC, Stabach PR, Morrow JS (2006) Human Sec31B: a family of new mammalian orthologues of yeast Sec31p that associate with the COPII coat. J Cell Sci 119:958–969
Stirnimann CU, Petsalaki E, Russell RB, Müller CW (2010) WD40 proteins propel cellular networks. Trends Biochem Sci 35:565–574
Ullah H, Chen JG, Temple B, Boyes DC, Alonso JM, Davis KR, Ecker JR, Jones AM (2003) The beta-subunit of the Arabidopsis G protein negatively regulates auxin-induced cell division and affects multiple developmental processes. Plant Cell 15:393–409
Van Nocker S, Ludwig P (2003) The WD-repeat protein superfamily in Arabidopsis: conservation and divergence in structure and function. BMC Genomics 4:50
von Arnim AG, Osterlund MT, Kwok SF, Deng XW (1997) Genetic and developmental control of nuclear accumulation of COP1, a repressor of photomorphogenesis in Arabidopsis. Plant Physiol 114:779–788
Walker AR, Davison PA, Bolognesi-Winfield AC, James CM, Srinivasan N, Blundell TL, Esch JJ, Marks MD, Gray JC (1999) The TRANSPARENT TESTA GLABRA1 locus, which regulates trichome differentiation and anthocyanin biosynthesis in Arabidopsis, encodes a WD40 repeat protein. Plant Cell 11:1337–1350
Wang H, Zhai L, Xu J, Joo HY, Jackson S, Erdjument-Bromage H, Tempst P, Xiong Y, Zhang Y (2006) Histone H3 and H4 ubiquitylation by the CUL4-DDB-ROC1 ubiquitin ligase facilitates cellular response to DNA damage. Mol Cell 22:383–394
Wu G, Xu G, Schulman BA, Jeffrey PD, Harper JW, Pavletich NP (2003) Structure of a beta-TrCP1-Skp1-beta-catenin complex: destruction motif binding and lysine specificity of the SCF (beta-TrCP1) ubiquitin ligase. Mol Cell 11:1445–1456
Xu C, Min J (2011) Structure and function of WD40 domain proteins. Protein Cell 2:202–214
Xu C, Bian C, Yang W, Galka M, Ouyang H, Chen C, Qiu W, Liu H, Jones AE, MacKenzie F, Pan P, Li SS, Wang H, Min J (2010) Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc Natl Acad Sci USA 107:19266–19271
Yonezawa K, Tokunaga C, Oshiro N, Yoshino K (2004) Raptor, a binding partner of target of rapamycin. Biochem Biophys Res Commun 313:437–441
Yu H, Braun P, Yıldırım MA, Lemmens I, Venkatesan K, Sahalie J, Hirozane-Kishikawa T, Gebreab F, Li N, Simonis N, Hao T, Rual JF, Dricot A, Vazquez A, Murray RR, Simon C, Tardivo L, Tam S, Svrzikapa N, Fan C, de Smet AS, Motyl A, Hudson ME, Park J, Xin X, Cusick ME, Moore T, Boone C, Snyder M, Roth FP, Barabási AL, Tavernier J, Hill DE, Vidal M (2008) High-quality binary protein interaction map of the yeast interactome network. Science 322:104–110
Zeng CJ, Lee YR, Liu B (2009) The WD40 repeat protein NEDD1 functions in microtubule organization during cell division in Arabidopsis thaliana. Plant Cell 21:1129–1140
Zhang X, Rashid R, Wang K, Shan S (2010) Sequential checkpoints govern substrate selection during cotranslational protein targeting. Science 328:757–760
Zhu J, Jeong JC, Zhu Y, Sokolchik I, Miyazaki S, Zhu JK, Hasegawa PM, Bohnert HJ, Shi H, Yun DJ, Bressan RA (2008) Involvement of Arabidopsis HOS15 in histone deacetylation and cold tolerance. Proc Natl Acad Sci USA 105:4945–4950
Acknowledgments
Grateful thanks are due to the Director, National Institute of Plant Genome Research (NIPGR), New Delhi, India for providing facilities. The authors work in this area was supported by NIPGR core grant and Department of Biotechnology (DBT), Govt. of India. Mr Awdhesh Kumar Mishra and Ms Swati Puranik acknowledge the fellowships form the Council of Scientific and Industrial Research, and NIPGR, New Delhi, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mishra, A.K., Puranik, S. & Prasad, M. Structure and regulatory networks of WD40 protein in plants. J. Plant Biochem. Biotechnol. 21 (Suppl 1), 32–39 (2012). https://doi.org/10.1007/s13562-012-0134-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-012-0134-1