Abstract
1,10-Phenanthroline, phen, is a molecule with a wide variety of chemical and biological activities, being interesting from a pharmacological standpoint. Its metal complexes have been extensively studied during the last century. Considering the vast applications of phen and its metallic complexes for the development of potential metallodrugs, it becomes of utmost importance to study the molecule’s interaction with different molecular targets. So far, in-depth analyses of phen and metal–phen complexes’ interaction with DNA are abundant. Nevertheless, little has been reported on the interaction of this molecule with the first molecular “target” that it finds in the cell, the lipid membrane. Here, we present the characterization of the interaction of the phen molecule with model membranes of 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) or 1,2-dipalmitoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (sodium salt) (DPPG) using a combination of differential scanning calorimetry and molecular dynamics simulations. Our results indicate that phen diffuses into the membranes, being stabilized around the middle of the hydrocarbon chains in DPPC, while in DPPG, it is placed near the center of the bilayer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The molecule 1,10-phenanthroline, phen (Fig. 1), presents a rich variety of biological activity. For instance, it is cytotoxic against tumor and leukemia cells. It induces the death of human ovarian carcinoma A2780 cell line (IC50 2.7 μM) [1] and murine leukemia cells (IC50 2.5 μM) [2]. It has recently been reported that phen can kill osteosarcoma cancer stem cells, responsible for osteosarcoma cancer relapse (IC50 2.9 μM) [3]. It also presents antifungal activity, and its reported mechanism of action includes mitochondrial damage. For instance, its minimum inhibitory concentration against the opportunistic pathogen C. albicans is 1.25–2.5 μg/mL, making phen an interesting new antifungal compound [4, 5]. Phen also presents antibacterial activity [6]. It is not mutagenic according to the AMES test [7]. The phen molecule is relatively tolerated in mammals, with doses up to 150 mg/kg weight tolerated by mice [8]. All the characteristics mentioned above make phen an interesting molecule from a pharmacological standpoint.
Phenanthroline forms highly stable metallic complexes with most transition metals in various oxidation states [9]. Akin to phen, its metal complexes also present a wide range of biological activity [10,11,12], usually more potent than the phen itself. The biological activity of this effective chelating agent boosted the use of metal–phen complexes for pharmacological ends, being widely employed in medicinal inorganic chemistry. Hitherto, various complexes, including phen or its derivatives, present biological activity, especially antitumor and antibacterial [6, 13, 14]. Levina et al. [15] proposed that the copper–phen complex may act as an ionophore, delivering copper into the cell.
The interaction of biologically active compounds with cellular membranes influences their bioavailability, even when membranes are not regarded as the biological target of these compounds. Considering the vast amount of information about phen and metallic complexes containing phen with biological activity, it has become clear the need to study the complexes’ interactions with the first “target” they find in the cell: the membrane. Moreover, the study of the free phen can also contribute to further studies of the interaction of phen-containing complexes and membranes, as recommended for coordination compound biological studies [15]. To the best of our knowledge, there are no such studies of isolated phen and membranes in the field.
As a part of our studies on new copper-containing compounds with potential applications for cancer treatment [16,17,18,19,20,21,22,23,24,25,26,27,28], we previously studied the cytotoxicity of copper complexes containing different ligands, including phen. However, as pointed out above, the initial step of analyzing the interaction between phen and membranes has been missing. Here, we investigate such interaction by experimental and theoretical methods, trying to understand the changes in the membrane induced by the phen at the molecular level.
The membrane models used were 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and 1,2-dipalmitoyl-sn-glycero-3-phospho-(1 − ′-rac-glycerol) (sodium salt) (DPPG). DPPC represents one of the most abundant lipid components of cell plasma membranes. The use of DPPG allows the comparison with the effects on negatively charged membranes, also common in cellular membranes [29]. The effects of phen on the phase transition of lipid bilayer models were studied by differential scanning calorimetry (DSC) and molecular dynamics simulations. In order to aid in the interpretation of the experimental results, molecular dynamics simulations were carried out. The information obtained is potentially useful for understanding the mechanism of interaction of phen with cellular membranes.
2 Experimental
2.1 Materials
Lipids 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and 1,2-dipalmitoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (sodium salt) (DPPG) were obtained from Avanti Polar Lipids, Inc. HEPES buffer (4-(2-hydroxyethyl)-1-piperizineethanesulfonic) and phenanthroline (phen) were purchased from Sigma-Aldrich. All reagents were used without further purification.
2.2 Sample Preparation
Lipid films were formed by drying chloroform/methanol (2/1, v/v) stock solutions of phospholipids under nitrogen flow and centrifuged under vacuum for at least 2 h to ensure complete removal of the organic solvent. The films were hydrated by adding 20 mM HEPES buffer pH = 7.4, vortexed for 15 min, and sonicated to a final concentration of 5 mM. Phen was dissolved in the same buffer and added to the mixture to a final concentration of 0.5 mM. Before measurements, it was left at least for 15 min at 60 °C.
2.3 Calorimetric Measurements
Differential scanning calorimetry (DSC) experiments were carried out in a VP-DSC MicroCal MicroCalorimeter. A 5 mM lipid suspension in 20 mM HEPES buffer pH = 7.4 with 0.5 mM sample concentration in a 10:1 lipid:compound relationship was used. The mixture was stabilized at 25 °C for 20 min before each measurement. Scans were recorded in the heating cycle only and with temperatures varying from 25 to 60 °C using a 20 °C/h heating velocity. Two scans were performed at each experiment to ensure reproducibility. Baseline correction and endotherm peak integration were performed on MicroCal Origin software.
2.4 Molecular Biomodeling
Molecular biomodeling and simulations were carried out using a workstation with an Intel (R) CoreTM i7-6700 CPU @ 3.40 GHz processor, with a Graphic Processor Unity (GPU) NVIDIA-GEFORCE GTX 1080 ti. The MD calculations were performed using NAMD [30] (Not Another Molecular Dynamics simulator) 2018 CUDA version 9010. System topology (CHARMM36 force field) [31, 32] was built using VMD [33].
The initial structure of the DPPC phospholipid bilayer was generated using a replacement method with the CHARMM-GUI web-based membrane builder (http://www.charmm-gui.org) [34, 35]. The lipid bilayer, containing 136 DPPC molecules, was oriented with its plane (4284 Å2) parallel to the xy plane and with the DPPC polar groups pointing towards the water phase. The latter consisted of 8898 tip3p water molecules, which were considered rigid, and the SETTLE constriction algorithm [36] was used for their simulation. The dimensions of the periodic cell were 65.00 × 65.00 × 110.00 Å. Its center was set to the center of mass.
The input structure for 1,10-phenanthroline was optimized at b3lyp/6–31 + G* level with Gaussian 09 [37]. To build the initial structure of the 1,10-phenanthroline-loaded DPPC membrane, the final MD conformation of the lipid bilayer was employed, and 4 solute molecules were randomly placed inside the water phase, two on each side of the bilayer.
2.5 Molecular Dynamics Simulations
The simulations involved three steps. Firstly, 10,000-step energy minimization was performed until an RMS gradient below 1 kcal mol‒1 Å‒2 was reached to avoid the system collapse at the beginning of the MD. Secondly, an equilibration step of 10 ns at 310 K and 1 atm was run, letting the system adjust its density by varying only the z-dimension. Finally, a 100 ns (membrane) and 300 ns (phen-loaded membrane) MD simulation was performed under the same conditions, keeping the ratio of the unit cell in the xy plane constant while allowing fluctuations along all axes.
Trajectories were run up with a time step of 2 fs and a time sampling of 2 ps. Non-bonded interactions were calculated with a switch distance of 8 Å and a cutoff of 10 Å (pair list distance = 12 Å). Long-range distance electrostatic interactions were taken into account using the particle mesh Ewald (PME) technique [38], with a grid size of 1 Å and a cutoff of 10 Å. Langevin dynamics was used to simulate the canonical ensemble (NPT), with a Langevin piston coupled to the heat bath (pressure = 1 atm; period = 100 fs; oscillation decay time = 50 fs).
To estimate the free energy change during the release of phen from the membrane into the cytosol, we carried out steered molecular dynamics simulations using the Colvars module (https://www.tandfonline.com/doi/full/10.1080/00268976.2013.813594). For that, a moving restrain was applied to a phen molecule, to progressively steer it along the z axis from the center of the membrane (0 Å) to the outside (40 Å), applying a harmonic potential of force constant 20 kcal/mol Å2 over 12,000,000 simulation steps.
2.6 Analyses and Visualization
All analyses took into account the last 30 ns of the trajectories using MEMBPLUGIN [39] within VMD [33]. The results were rendered with Discovery Studio Visualizer [40].
3 Results and Discussion
3.1 Calorimetric Profile
The phase transition temperature and thermodynamics of lipid phase transitions are sensitive to the presence of exogenously added molecules, making calorimetric methods an adequate strategy to determine whether the added molecule disturbs the packing of the membrane. Therefore, the thermograms corresponding to DSC measurements of DPPC and DPPG membranes in the absence and presence of phen are presented in Fig. 2. Table 1 summarizes the thermodynamical parameters obtained from the DSC studies.
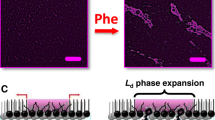
As the temperature of DPPC bilayers increased, two thermal transitions were registered in measurements in the presence or absence of phen. The first weak, broad peak corresponds to a pre-transition (P) between the gel phase Lβ′ and the ripple phase Pβ′ (around 35 °C). At higher temperatures, a sharp peak corresponding to the main transition (M) between the Pβ′ and the liquid–crystalline phase Lα (at approximately 41 °C) is observed. In the DPPC bilayer, the addition of phen induced a shift towards lower temperatures and a broadening of the pre-transition. A broader peak and, therefore, a decrease of the cooperative unit are related to the phen being placed in the region of the cooperative unit, that is, around carbons C1–C8 [41]. The temperature shift can be related to the polar region of the lipids being disturbed [41, 42]. The DPPC main transition temperature was not modified, while ΔHCAL M and ΔT1/2 M increased. This behavior suggests that the phen can intercalate between the fatty acid chains, reducing the transition’s cooperativity (reflected in the increase of the ΔT1/2), therefore being placed in the C1-C8 region and inducing more substantial van der Waals interactions.
DPPG bilayers presented only a sharp peak corresponding to the gel to liquid–crystalline phase transition around 39 °C. The presence of phen in the membrane was evidenced by an increase of the enthalpy variation associated with the main transition (ΔHCAL M) and also the appearance of a broad low-energy transition seen as a shoulder on the lower temperature side of the main peak (Fig. 2B). They can be related to a degree of phen intercalation into the acyl chains leading to augmented van der Waals interactions. There is no evidence of phen being placed in the cooperative unit region in this lipid.
3.2 Computational Results
To computationally assess the interaction of phen with a DPPC bilayer, we modeled the system by MD simulations. We used a step-by-step approach, starting by generating an equilibrated and representative model of the lipid membrane. As it is depicted in Fig. S1a–d, the bilayer system is equilibrated after a 70 ns MD simulation, giving rise to a stable membrane model (Fig. S2). The parameters averaged over the production phase (last 30 ns of the MD trajectory) were also calculated to validate the computational model against previous experimental evidence. The results were satisfactory: the averaged calculated area per phospholipid and membrane thickness was 57.63 ± 0.02 Å2 and 41.47 ± 0.01 Å, in complete agreement with the reported values of 47.9 – 64 Å2 and 38.3 – 44.2 Å, respectively, registered between 20 and 50 °C [43]. Besides, the lateral diffusion coefficient (D) calculated from the MD simulation was 7.6 × 10‒8 cm2 s‒1, compared to the experimental results for fluorescent lipid analogs in DPPC membranes and vesicles (between 10‒8 and 10‒7 cm2 s‒1) [44]. Moreover, the calculated values for the order parameter ‒SCD (Fig. S3) varied around 0.21, in line with the empirical value found for phospholipid bilayers in the fluid phase (‒SCD ≈ 0.2) [45].
Starting from this equilibrated model of the DPPC bilayer, MD simulations in the presence of 1,10-phenanthroline and the results are depicted in Figs. 3 and 4. During the 295 ns MD simulation at 310 K, two of the four phen molecules placed in the water phase diffused into the lipid membrane (Fig. 3). According to the distance between the z coordinates of the phen and membrane centers of mass (Δz) (Fig. 4a), the two phen molecules entered the lipid phase around 20 ns and 45–85 ns. Both were stabilized inside the membrane at an average distance of 11.1–11.2 Å from the center of the bilayer, interacting with the middle of the hydrophobic hydrocarbon tails through dispersion forces and C–H/π interactions. This observation agrees with the calorimetric results, which evidenced a reduction of the cooperative unit in the presence of phen, indicating its localization in the cooperative unit region (C1–C8) and increased dispersion forces. Molecular dynamics results also agree with previous experimental evidence, which shows that other polycyclic aromatic hydrocarbons like anthracene, benz[a]pyrene, and dibenz[a,h]anthracene prefer the lipid phase [46,47,48].
a Distance between the z coordinates of the phen and membrane centers of mass (Δz), b membrane area per DPPC molecule, and c bilayer thickness of the phen-DPPC membrane system monitored along with the MD simulation. In a, the curves for the four phen molecules are plotted with different colors (black, red, green, and blue), and the membrane–water boundary is depicted as a dashed line. The time range during which the solute molecules entered the lipid bilayer is highlighted. DPPC hydrogen atoms are omitted for clarity. Atom color code: C (gray), N (blue), O (red), P (orange)
The phen molecules placed inside the lipid bilayer also interact, through H-bonds, with the water molecules linked to the DPPC polar headgroups, pulling them towards the center of the membrane (compare Figs. 3 and S2). This phenomenon causes the formation of undulations over the surface of the lipid membrane. This effect can be responsible for modifying the calorimetric profile related to the polar heads observed for phen in DPPC.
The presence of phen molecules inside the lipid phase significantly influences the bilayer structure. According to Fig. 4b–c, the area per phospholipid decreased, while the bilayer thickness increased. Indeed, the averaged calculated area per phospholipid and membrane thickness was 51.90 ± 0.01 Å2 and 44.023 ± 0.006 Å, in comparison with the analogous parameters calculated for the pure DPPC membrane: 57.63 ± 0.02 Å2 and 41.47 ± 0.01 Å, respectively. This is in line with previous computational evidence for similar ligands [46]. Additionally, the computational results showed that the phen molecules inside the membrane increased the order of the phospholipid chains (lower SCD values) (see Fig. S3). Consequently, the phospholipid bilayer becomes less fluid, confirmed by the decrease in the lateral diffusion coefficient from 7.6 × 10‒8 cm2 s‒1 (DPPC bilayer) to 2.1 × 10‒8 cm2 s‒1 (DPPC bilayer + phen). This effect has been previously observed for analogous systems [46], potentially being the structural basis behind the sharp increase of the interdigitation width upon insertion of the phen ligand (Fig. S4). The release of phen from the membrane into the cytosol involved a free energy change, estimated using steered molecular dynamics, is 13.4 kcal/mol. This suggests that the release of phen from the membrane would likely require the participation of other molecules or processes (for instance, the transition to a more fluid membrane state).
4 Conclusions
1,10-Phenanthroline can diffuse into the DPPC and DPPG model membranes. It induces modifications in the membrane, which can be part of the diverse biological effects of the phen. In DPPC membranes, it appears to be stabilized near the middle of the hydrocarbon chains interfering with the cooperative unit, while in DPPG membranes, it appears to be stabilized in the deeper, hydrophobic region of the membrane. Molecular dynamics were able to explain the effects observed by DSC at the molecular level. In addition, it shows that the insertion of the ligands triggers an increase in the interdigitation, with a concomitant reduction of membrane fluidity.
Despite the extensive use of inorganic compounds in medicinal chemistry, the essential aspects governing their cytotoxicity and mechanism of action have remained poorly understood [49]. The general rules that dictate the behavior of inorganic systems in medicinal chemistry have not yet emerged. That is likely the result of the significant reactivity and plasticity shown by metal binding in those compounds. To that end, understanding the free ligands’ behavior compared with the complexes can give crucial information on the more complicated situations found in the cell.
In particular, phen inhibits cellular DNA synthesis, which was assumed to be its only mechanism of cytotoxic action [50, 51]. This idea is being revisited with the increasing use of phen and related ligands on metallodrugs. The impact of cellular uptake of phen and congeners on cytotoxicity is underexplored. Diffusion is the most accepted model of transport [49]. As there are plenty of factors that may influence the activity, in our contribution, we started from a basic aspect trying to understand phen interaction with model membranes as a step to understanding its biological activity. Whether or not the observed membrane modifications may contribute to the mechanism of action cannot be yet fully answered. Notwithstanding, they are a very likely part of the phen uptake process.
References
L. Messori et al., Gold (III) complexes as potential antitumor agents: solution chemistry and cytotoxic properties of some selected gold (III) compounds. J. Med. Chem. 43(19), 3541–3548 (2000)
M. Pitié et al., Cytostatic activity of 1, 10-phenanthroline derivatives generated by the clip-phen strategy. ChemBioChem 6(4), 686–691 (2005)
P. Robin, K. Singh, K. Suntharalingam, Gallium (III)-polypyridyl complexes as anti-osteosarcoma stem cell agents. Chem. Commun. (2020)
B. Coyle et al., Mode of anti-fungal activity of 1, 10-phenanthroline and its Cu (II), Mn (II) and Ag (I) complexes. Biometals 16(2), 321–329 (2003)
M. McCann et al., Insights into the mode of action of the anti-Candida activity of 1, 10-phenanthroline and its metal chelates. Met.-Based Drugs 7(4), 185–193 (2000)
L. Viganor et al., The antibacterial activity of metal complexes containing 1, 10-phenanthroline: potential as alternative therapeutics in the era of antibiotic resistance. Curr. Top. Med. Chem. 17(11), 1280–1302 (2017)
C. Deegan et al., In vitro cancer chemotherapeutic activity of 1, 10-phenanthroline (phen),[Ag2 (phen) 3 (mal)]· 2H2O,[Cu (phen) 2 (mal)]· 2H2O and [Mn (phen) 2 (mal)]· 2H2O (malH2= malonic acid) using human cancer cells. Cancer Lett. 247(2), 224–233 (2007)
M. McCann et al., In vitro and in vivo studies into the biological activities of 1, 10-phenanthroline, 1, 10-phenanthroline-5, 6-dione and its copper (II) and silver (I) complexes. Toxicol. Res. 1(1), 47–54 (2012)
W.W. Brandt, F.P. Dwyer, E.D. Gyarfas, Chelate complexes of 1, 10-phenanthroline and related compounds. Chem. Rev. 54(6), 959–1017 (1954)
A.A. Adeniyi, P.A. Ajibade, The anticancer activities of some nitrogen donor ligands containing bis-pyrazole, bipyridine, and phenanthroline moiety using docking methods. Bioinorg. Chem. Appl. (2018)
R.M. Gandra et al., Antifungal Potential of copper (II), manganese (II) and silver (I) 1, 10-phenanthroline chelates against multidrug-resistant fungal species forming the Candida haemulonii complex: impact on the planktonic and biofilm lifestyles. Front. Microbiol. 8, 1257 (2017)
C. Santini et al., Advances in copper complexes as anticancer agents. Chem. Rev. 114(1), 815–862 (2014)
Z. Albobaledi et al., Mixed ligand Cu (II) complexes of an unsymmetrical Schiff base ligand and N-donor heterocyclic co-ligands: investigation of the effect of co-ligand on the antibacterial properties. Inorganica Chim. Acta. 499, 119185 (2020)
A. Casini, A. Vessières, S.M. Meier-Menches, Metal-based anticancer agents. Royal Soc. Chem. 14, (2019)
A. Levina, D.C. Crans, P.A. Lay, Speciation of metal drugs, supplements and toxins in media and bodily fluids controls in vitro activities. Coord. Chem. Rev. 352, 473–498 (2017)
G. Facchin et al., Interaction of Cu-dipeptide complexes with calf thymus DNA and antiproliferative activity of [Cu (ala-phe)] in osteosarcoma-derived cells. Polyhedron 28(12), 2329–2334 (2009)
S. Iglesias et al., Towards the development of new copper compounds for the treatment of cancer: study of the cytotoxic activity of Cu(L-dipeptide)(1,10-phenanthroline) complexes. Proc. 12th Int. Symp. Metal Ions Biol. Med. 1, (2013)
N. Alvarez et al., Synthesis, structural characterization and DNA interaction of new copper-terpyridine complexes. Polyhedron 68, 295–302 (2014)
S. Iglesias et al., Structural characterization and cytotoxic activity of heteroleptic copper (II) complexes with L-dipeptides and 5-NO2-phenanthroline. Crystal Structure of [Cu (Phe-Ala)(5-NO2-Phen)]. 4H2O. Struct. Chem. Crystallogr. Commun. (2015)
G. Facchin et al., Experimental and theoretical studies of copper complexes with isomeric dipeptides as novel candidates against breast cancer. J. Inorg. Biochem. 162, 52–61 (2016)
N. Alvarez et al., Synthesis, structural characterization and cytotoxic activity against tumor cells of heteroleptic copper (I) complexes with aromatic diimines and phosphines. Inorg. Chim. Acta 466, 559–564 (2017)
N. Alvarez et al., Development of copper (II)-diimine-iminodiacetate mixed ligand complexes as potential antitumor agents. Inorg. Chim. Acta. (2018)
G. Facchin et al., Structural and spectroscopic characterization of two new Cu (II)-dipeptide complexes. Z. Naturforsch. B 55(12), 1157–1162 (2000)
G. Facchin et al., Synthesis and characterization of three new Cu (II)-dipeptide complexes. J. Inorg. Biochem. 89(3), 174–180 (2002)
G. Facchin et al., Cu (II) complexation with His–Gly and His–Ala. X-ray structure of [Cu (his–gly)2(H2O)2]· 6H2O. Inorg. Chim. Acta 355, 408–413 (2003)
E.D. Vieira et al., Weak exchange interaction supported by a biologically relevant long chemical bridge in a Cu-peptide model compound. Inorg. Chem. 45(7), 2942–2947 (2006)
G. Facchin et al., Structural characterization of a series of new Cu-dipeptide complexes in solid state and in solution. Polyhedron 25(13), 2597–2604 (2006)
S. Iglesias et al., Synthesis, structural characterization and cytotoxic activity of ternary copper(II)–dipeptide–phenanthroline complexes. A step towards the development of new copper compounds for the treatment of cancer. J. Inorg. Biochem. 139, 117–123 (2014)
A.C. Alves et al., Biophysics in cancer: the relevance of drug-membrane interaction studies. Biochim. Biophys. Acta Biomembr. 1858(9), 2231–2244 (2016)
J.C. Phillips et al., Scalable molecular dynamics with NAMD. J. Comput. Chem. 26(16), 1781–1802 (2005)
K. Vanommeslaeghe et al., CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31(4), 671–690 (2010)
J.B. Klauda et al., Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114(23), 7830–7843 (2010)
W. Humphrey, A. Dalke, K. Schulten, VMD: visual molecular dynamics. J. Mol. Graph. 14(1), 33–38 (1996)
E.L. Wu et al., CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 35(27), 1997–2004 (2014)
S. Jo et al., CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29(11), 1859–1865 (2008)
S. Miyamoto, P.A. Kollman, Settle: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13(8), 952–962 (1992)
M.J. Frisch et al., Gaussian 09 (Gaussian Inc, Wallingford CT USA, 2009)
T. Darden, D. York, L. Pedersen, Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98(12), 10089–10092 (1993)
R. Guixà-González et al., MEMBPLUGIN: studying membrane complexity in VMD. Bioinformatics 30(10), 1478–1480 (2014)
BIOVIA, Dassault Systèmes, Discovery Studio Visualizer, v16.1.015350, San Diego: Dassault Systèmes (2015)
M.K. Jain, N.M. Wu, Effect of small molecules on the dipalmitoyl lecithin liposomal bilayer: III. Phase transition in lipid bilayer. J. Membr. Biol. 34(1), 157–201 (1977)
B.Z. Chowdhry et al., Effect of lanthanum ions on the phase transitions of lecithin bilayers. Biophys. J . 45(3), 633–635 (1984)
J.F. Nagle, S. Tristram-Nagle, Structure of lipid bilayers. Biochim. Biophys. Acta Rev. Biomembr. 1469(3), 159–195 (2000)
P.F. Fahey, W.W. Webb, Lateral diffusion in phospholipid bilayer membranes and multilamellar liquid crystals. Biochemistry 17(15), 3046–3053 (1978)
L.S. Vermeer et al., Acyl chain order parameter profiles in phospholipid bilayers: computation from molecular dynamics simulations and comparison with 2H NMR experiments. Eur. Biophys. J. 36(8), 919–931 (2007)
H.I. Padilla-Chavarría, T.R.C. Guizado, A.S. Pimentel, Molecular dynamics of dibenz[a, h]anthracene and its metabolite interacting with lung surfactant phospholipid bilayers. Phys. Chem. Chem. Phys. 17(32), 20912–20922 (2015)
T.R. Sosnowski et al., Alteration of surface properties of dipalmitoyl phosphatidylcholine by benzo[a]pyrene: a model of pulmonary effects of diesel exhaust inhalation. J. Biomed. Nanotechnol. 8(5), 818–825 (2012)
J.-R. Ren et al., Comparative transmembrane transports of four typical lipophilic organic chemicals. Biores. Technol. 101(22), 8632–8638 (2010)
A.C. Hachey, D. Havrylyuk, E.C. Glazer, Biological activities of polypyridyl-type ligands: implications for bioinorganic chemistry and light-activated metal complexes. Curr. Opin. Chem. Biol. 61, 191–202 (2021)
N. Berger, E.S. Johnson, S.A.M. Skinner, Ortho-phenanthroline inhibition of DNA synthesis in mammalian cells. Exp. Cell Res. 96(1), 145–155 (1975)
C. Krishnamurti, L.A. Saryan, D.H. Petering, Effects of ethylenediaminetetraacetic acid and 1, 10-phenanthroline on cell proliferation and DNA synthesis of Ehrlich ascites cells. Can. Res. 40(11), 4092–4099 (1980)
Funding
The authors are grateful to Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) (Grant no. 2015/50366-7 and 2020/15542-7), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) (Grant no. 306682/2018-4), CAPES/UdelaR (grant no. 050/2013), and Programa de Desarrollo de las Ciencias Básicas (PEDECIBA, Uruguay) for the financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Sergio Mascarenhas - A Polymath in Physics
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Alvarez, N., Freddi, P., Castellani, S. et al. New Insights into the Biophysical Behavior of an Old Molecule: Experimental and Theoretical Studies of the Interaction Between 1,10-Phenanthroline and Model Phospholipid Membranes. Braz J Phys 52, 111 (2022). https://doi.org/10.1007/s13538-022-01114-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13538-022-01114-7