Abstract
Purpose
Ovarian metastasis of gastric cancer (GC), commonly referred to as Krukenberg tumors, leads to a poor prognosis. However, the cause of metastasis remains unknown. Here, we present an integrated single-cell RNA sequencing (scRNA-Seq) analysis of the immunological microenvironment of two paired clinical specimens with ovarian metastasis of GC.
Methods
scRNA-Seq was performed to determine the immunological microenvironment in ovarian metastasis of gastric cancer. CellChat was employed to analyze cell-cell communications across different cell types. Functional enrichment analysis was done by enrichKEGG in clusterProfiler. GEPIA2 was used to assess the influence of certain genes and gene signatures on prognosis.
Results
The ovarian metastasis tissues exhibit a heterogenous immunological microenvironment compared to the primary tumors. Exhaustion of T and B cells is observed in the ovarian metastasis tissues. Compared to the paired adjacent non-tumoral and primary tumors, the ratio of endothelial cells and fibroblasts is high in the ovarian metastasis tissues. Compared to primary ovarian cancers, we identify a specific group of tumor-associated fibroblasts with MFAP4 and CAPNS1 expression in the ovarian metastatic tissues of GC. We further define metastasis-related-endothelial and metastasis-related-fibroblast signatures and indicate that patients with these high signature scores have a poor prognosis. In addition, the ovarian metastasis tissue has a lower level of intercellular communications compared to the primary tumor.
Conclusion
Our findings reveal the immunological microenvironment of ovarian metastasis of gastric cancer and will promote the discovery of new therapeutic strategies for ovarian metastasis in gastric cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Gastric cancer (GC) is one of the most commonly diagnosed cancers and a leading cause of cancer-related death [1,2,3]. Ovarian metastasis of GC, also referred to Krukenberg tumors, is one unique type of peritoneal metastasis in female patients, with an incidence ranging from 0.3% to 6.7% [4,5,6]. Autopsy results showed that the incidence rate of ovarian metastasis ranges from 33% to 41% [7, 8]. Although multiple therapies including chemoradiotherapy, targeted therapy, and immune therapy have been developed, GC with ovary metastasis has a poor prognosis with a median overall survival of less than one year [4,5,6]. However, the underlying mechanism of organ-specific metastatic is still unclear.
Dynamic and reciprocal interactions of the components of tumor microenvironment (TME) in GC play a critical role in cancer cell metastasis and therapy resistance [9, 10]. Immunosuppressive TME promotes gastric cancer angiogenesis, metabolic reprogramming, extracellular matrix remodeling, and epithelial-mesenchymal transition [11,12,13]. Thus, a detailed and precise elaboration of the TME is of great significance for developing new treatments of ovarian metastasis in GC.
Single-cell RNA sequencing (scRNA-Seq) technology has been widely utilized to visualize the heterogeneity of tumor cells and TME in various cancers, including GC [14,15,16]. However, considering clinical limitation, the TME of ovarian metastases of GC remain poorly characterized. Herein, we performed scRNA-Seq and in-depth bioinformatic analyses on two primary GC tumors with the matched adjacent non-tumor and ovarian metastasis tissues. The comprehensive profiles of the TME obtained allowed us to identify a complex yet unique tumor microenvironment of GC with ovarian metastasis.
2 Materials and methods
2.1 Data collection
The 10x genomics scRNA-Seq data of ovarian tumors were from GEO (https://www.ncbi.nlm.nih.gov/geo/) database (GSE184880), including 7 treatment-naïve patients with high-grade serous ovarian cancer at early or late stages and 5 age-matched nonmalignant ovarian samples. Samples have more than 50 fibroblasts were used.
The original data of BD Rhapsody scRNA-Seq in this study was from one primary gastric tumor with the paired adjacent non-tumor and ovarian metastasis tissues derived from a 52-year-old woman affected by ovarian metastasis of gastric cancer at RenJi Hospital, Shanghai Jiao Tong University School of Medicine. The consent forms and the proposed studies were approved by Institutional Review Board of Renji Hospital (LY2022-065-B). Before all sample collection, the patient was treated with 6-cycle chemotherapy in combination with immunotherapy. Fresh specimens of primary tumor, adjacent tissue and ovarian metastases were obtained from the surgery. Matched adjacent gastric tissues from sites displaced at least several centimeters from the tumor were used.
We further downloaded a 10x genomics scRNA-Seq data (GEO: GSE234129) from another primary gastric tumor (MDA_Pt5-Ca) with the paired adjacent non-tumor (MDA_Pt5-Ad) and ovarian metastasis tissues (MDA_Pt5-Ov) from treatment-naïve patient (MDA_Pt5) [17].
2.2 Preparation of single-cell suspensions
Tissues were cut into small pieces and washed with phosphate-buffered saline (PBS), and then dissociated using a Human Tumor Dissociation kit (Miltenyi Biotech, Germany) according to the manufacturer’s protocol. Dissociated cells were passed through a MACS smart strainer (70 μm) and incubated with RBC lysis buffer (Solarbio, China) for 5 min. The dissociated cells were sorted into PBS + 0.04% bovine serum albumin (BSA). All samples showed a viability of around 80% on average as confirmed by trypan blue exclusion. The single-cell suspensions were counted using the Countess II Automated Cell Counter (Thermo Fisher Scientific, USA) and then used for single-cell experiments.
2.3 Library preparation and single-cell RNA sequencing
Single-cell RNA-Seq was performed according to the manufacturer’s instructions of BD Rhapsody. scRNA-Seq libraries were generated using the BD Rhapsody™ Targeted mRNA and AbSeq Amplification Kit (Cat. # 633774). Single-cell capture was achieved by random distribution of a single-cell suspension across > 200,000 microwells through a limited dilution approach. Beads with oligonucleotide barcodes were added to saturation so that a bead was paired with a cell in a microwell. Cell lysis buffer was added so that poly-adenylated RNA molecules hybridized to the beads. Beads were collected into a single tube for reverse transcription. Upon cDNA synthesis, each cDNA molecule was tagged on the 5′ end (that is, the 3′ end of an mRNA transcript) with a molecular index and cell label indicating its cell of origin. Whole transcriptome libraries were prepared using the BD Resolve single-cell whole-transcriptome amplification workflow. In brief, second strand cDNA was synthesized, followed by ligation of the adaptor for universal amplification. 20 cycles of PCR were used to amplify the adaptor-ligated cDNA products. Sequencing libraries were prepared using random priming PCR of the whole-transcriptome amplification products to enrich the 3′ end of the transcripts linked with the cell label and molecular indices. The libraries were sequenced on NovaSeq 6000.
2.4 Single-cell RNA expression quantification, quality control, and clustering analysis
Raw sequencing data were aligned and quantified using the BD WTA Rhapsody Analysis Pipeline. The human genome hg38 was used as the reference genome. The outputs for each sample were converted to a Seurat object using the R Seurat package for downstream analysis [18]. To remove low-quality cells, we used the following criteria: less than 200 unique genes expressed, more than 5000 unique genes expressed or more than 20% of reads mapping to mitochondria. Then we combined all samples by the function of IntegrateData and the integrated data was scaled through the function of ScaleData. Then, we performed a principal component analysis using RunPCA, selected the first 20 principal components and used them to perform UMAP dimensionality reduction through RunUMAP. Cell clustering was performed using the FindClusters function with resolution set to 0.2 and the clusters were annotated by the expression of canonical marker genes. FindAllMarkers with default parameters except for min.pct set to 0.25 (expression in above 25% of the corresponding cluster) was used to define marker genes for each cluster. Functional enrichment analysis was done by enrichKEGG in clusterProfiler [19]. To further identify sub-clusters and annotate them as cell subtypes in each major cell type, the above-mentioned step (integration, normalization, dimensionality reduction, and clustering) were repeated for each major cell type.
2.5 Survival analysis
Survival analysis was performed using GEPIA2 [20] (http://gepia2.cancer-pku.cn). Briefly, the TCGA-STAD tumor samples were divided into two groups along with low (< median) and high (> median) target gene expression for all patients, Kaplan-Meier plots and log-rank tests were used for overall survival analysis.
2.6 Cell-cell communications analysis
CellChat was used to analyze cell-cell communications across different cell types in the tumor ecosystems with default parameters [21]. To compare different cellular communications among the primary tumor, matched normal tissue, and ovarian metastasis, cell groups in different samples were lifted to the same, then performed comparison.
2.7 Statistical analysis
GraphPad Prism 8.4 software (GraphPad Software Inc., San Diego, CA, USA) was used for statistical analyses. A minimum of 3 independent biological replicates was analyzed using Student’s two-tailed t-test or one-way ANOVA (Newman-Keuls post hoc test). Kaplan-Meier survival analysis was performed using the log-rank test. P < 0.05 was considered significant.
3 Results
3.1 Single-cell transcriptomic analyses of the primary tumors and the paired adjacent non-tumoral and ovarian metastasis tissues from two patients with gastric cancer
To better understand the underlying mechanisms that drive the progression and ovarian metastasis of gastric cancer, we performed single-cell RNA-Seq analysis of paired clinical specimens from one GC patient with ovarian metastasis after chemotherapy and immunotherapy (Pt1). We also downloaded another scRNA-Seq dataset of paired clinical specimens from one treatment-naïve GC patient (Pt5, GSE234129) [17]. In total, we got 6 paired samples from two patients: two primary tumors (i.e., P), two adjacent non-tumoral samples (i.e., N), and two paired ovarian metastasis tissues (i.e., M) (Fig. 1A). After quality filtering, a total of 10,363 cells was identified with a median of 1140 genes detected per cell, among which 2771, 2303, and 5289 cells were collected from N, P, and M, respectively. After data integration, dimensionality reduction, and unsupervised cell clustering, we identified 8 cell clusters and annotated as a specific tumor microenvironment (TME) cell type based on the expression of well-known marker genes: fibroblasts (3537; DCN, COL1A1, COL1A2), B cells (2116; CD19, CD79A), T cells (1348; CD3D, CD3E), endothelial cells (886; VWF, PECAM1), myeloid cells (876; CD14, CD68), epithelial cells (920; EPCAM, KRT8), pericyte cells (547; RGS5, TAGLN), and proliferative cells (133; MKI67) (Fig. 1B and C). The cell number and cell proportion of each cell type were shown in Fig. 1D and E. In general, samples received treatments had a higher proportion of stromal cells like fibroblasts and endothelial cells, while immune cells like B cells and T cells make up most of the treatment-naïve samples. Meanwhile, irrelevant to whether received treatments, ovarian metastasis tissues had a higher proportion of stromal cells than adjacent non-tumoral tissues and primary tumors (Fig. 1E). This indicates that the ovarian metastasis tissue has a different TME heterogeneity from the gastric cancer primary tumor and treatments impair the TME.
Single-cell transcriptomic analyses of the primary tumor and the paired adjacent non-tumoral and ovarian metastasis tissues from two patients with gastric cancer. (A) Schematic of the study design and workflow. Six fresh specimens (two primary GC tumors: P, two adjacent non-tumor: N, and two ovarian metastasis tissues: M) were collected from two patients. (B) Uniform manifold approximation and projection (UMAP) of cells from all six samples colored by sample origin (left panel) and cell type (right panel), respectively. (C) Dotplot of cell type-specific marker genes across all cell types. (D) The number of cells in each cell type across all samples. (E) The proportion of each cell type in each sample
3.2 Exhaustion of T and B cells is related to gastric cancer progression and ovarian metastasis
First, we compared the tumor immune microenvironment among the adjacent non-tumoral tissue, primary tumor, and ovarian metastasis. In total 1348 T cells (269 from N, 244 from P, and 835 from M) were subjected to integration and unsupervised clustering to reveal subtypes. Four subclusters were identified, C0, C1, C2, and C3 (Fig. 2A). CD8B, CD8A, and GZMK were highly expressed in C0 cluster. TYROBP, FCGR3A, and KLRF1 were highly expressed in C1 cluster. CTLA4, LTB, and CD4 were enriched in C2 cluster. KIR2DL4, KRT86, and CD7 were specifically expressed in C4 cluster (Fig. 2B, D). Consistent with the marker genes, the ovarian metastasis tissues showed a low proportion of C0 cluster with CD8+ T cells compared to the adjacent non-tumoral tissue and the primary tumor (Fig. 2C). In addition, the ovarian metastasis tissues had a higher proportion of C1 cluster with TYROBP+ and FCGR3A+ T cells (Fig. 2C). TYROBP was a poor prognostic factor in GC and low-grade glioma [22, 23]. FCGR3A has been reported as an immune-oncogenic molecule that correlated with tumor immune infiltration levels and affected drug sensitivity in most cancers [24].
Exhaustion of T cells is related to gastric cancer progression and ovarian metastasis. (A) UMAP of T cells in different specimens by their distinct origination. (B) Heatmap shows expression of marker genes in each subcluster. (C) Proportion of each T cell subcluster in each sample. (D) Expression of CD8A, FCGR3A, CTLA4, and KIR2DL4 mRNAs across T cells
Next, we performed B cell analysis. Five subclusters were identified from 2116 B cells (1050 from N, 896 from P, and 170 from M) (Fig. 3A). DUSP5 and EGR1 were highly expressed in C0 cluster. IGKC and MZB1 were enriched in C1 cluster. ACTG1 and TMSB4X were expressed in C2 cluster. MS4A1 and HLA-DRA were expressed in C3 cluster. CPA3 and TPSAB1 were enriched in C4 cluster (Fig. 3B). Ovarian metastasis tissues showed the lowest proportion in C1 cluster with IGKC+ and MZB1+ B cells (Fig. 3C). In C2 cluster with ACTG1+ B cells, the ovarian metastasis tissues had a higher proportion compared to primary tumors and non-tumoral tissues (Fig. 3C). IGKC+ plasma/B cells predicts a prolonged survival in both esophageal and gastric cancer [25], and MZB1 acted as a suppressor of GC and low MZB1 expression in the primary GC tissues was predictive of recurrence after curative resection [26]. While upregulation of ACTG1 promoted epithelial-mesenchymal transition in GC [27]. These results indicate that the exhaustion of T and B cells is related to gastric cancer progression and ovarian metastasis.
3.3 M0 macrophage infiltration is high in the ovarian metastasis tissues of GC
We identified 4 subclusters from 876 myeloid cells (196 from N, 229 from P, and 451 from M): M0 macrophages (CD68, CD14), M2 macrophages (MRC1, CD163), Monocytes (S100A8, S100A9), and DC (dendritic cell, CD1C, CLEC10A) (Fig. 4A, B). Differentially expressed genes (DEGs) in M0 macrophages were enriched in the KEGG pathways including ribosome and oxidative phosphorylation. DEGs in M2 macrophages were enriched in the lysosome, phagosome, and antigen processing and presentation. DEGs in DC were enriched in ribosome, phagosome, and antigen processing and presentation. However, DEGs in Monocytes were enriched in osteoclast differentiation, IL-17 signaling pathway, and TNF signaling pathway (Fig. 4C). The proportion of M0 macrophages was higher in the ovarian metastasis tissues compared to the primary tumors and adjacent non-tumoral tissues (Fig. 4D). Meanwhile, the proportion of M2 macrophages decreased from non-tumoral tissues to primary tumors and ovarian metastasis tissues had the lowest (Fig. 4D). This data shows that macrophage infiltration may be important for ovarian metastasis in GC.
M0 macrophage infiltration is high in the ovarian metastasis tissue in GC. (A) UMAP of myeloid cells in different specimens by their distinct origination. (B) Heatmap shows expression of marker genes in each subcluster. (C) Dotplot shows enriched KEGG pathways in each subcluster. (D) Proportion of each subcluster in each sample
3.4 Endothelial cells are rich in the ovarian metastasis tissues
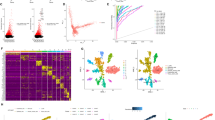
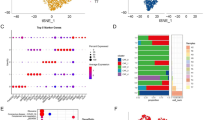
A total of 886 endothelial cells was clustered into 4 subclusters (52 from N, 110 from P, and 724 from M) (Fig. 5A). These subclusters showed different gene expression paradigms. RGCC, PLPP1, and CXCL12 were highly expressed in C0 cluster. ACKR1, HLA-DRA, and SLCO2B1 were highly expressed in C1 cluster. C7, COL3A1, and DCN were highly expressed in C2 cluster, while CCL21, PDPN, and NTS were highly expressed in C3 cluster (Fig. 5B). Besides, DEGs in each subcluster were enriched in different KEGG pathways. C0 cluster enriched in Leukocyte transendothelial migration, focal adhesion, and regulation of actin cytoskeleton. C1 cluster enriched in ribosome, antigen processing and presentation, allograft rejection. C2 cluster enriched in focal adhesion and ribosome. C3 cluster enriched in focal adhesion, regulation of actin cytoskeleton and endocytosis (Fig. 5C). These subclusters were distributed differently across the adjacent non-tumoral tissue, primary, and metastasis tumors. C0 cluster made up most of the adjacent non-tumoral tissue and primary tumor and nearly 40% of the metastasis tissue. C2 and C3 clusters constituted little of both three samples, and C1 cluster occupied nearly half of the metastatic tumor and a very small proportion in the adjacent non-tumoral tissue and primary tumor (Fig. 5D). Although all three samples contained a large proportion of C0 cluster, these cells still showed different expression patterns among three specimens, Gene Ontology analysis showed that genes highly expressed in the metastasis tumor enriched in ECM-receptor interaction, focal adhesion, PI3K-Akt signaling pathway, and estrogen signaling pathway, while in the adjacent non-tumoral tissue enriched in MAPK signaling pathway, IL-17 signaling pathway and Th17 cell differentiation, for the primary tumor enriched in cell adhesion molecules, antigen processing and presentation, and phagosome (Fig. 5E). We also found that VEGFA, VEGFB, VEGFC mRNAs were highly expressed in the metastasis tissue (Supplementary Fig. S1). We further defined the genes specifically highly expressed in endothelial cells from the metastasis sample as metastasis-related-endothelial signature, the Cancer Genome Atlas Stomach Adenocarcinoma (TCGA-STAD) samples with high signature values showed poor prognosis, while it was not statistically significant (Fig. 5F). High expression of CD300LG, IGFBP7, or CXCR4 was related to a poor prognosis in the TCGA-STAD dataset (Fig. 5F). This data indicates that endothelial cells are rich in the ovarian metastasis tissue in gastric cancer.
Endothelial cells are rich in the ovarian metastasis tissue. (A) UMAP of endothelial cells in different specimens by their distinct origination. (B) Dotplot shows expression of marker genes in each subcluster. (C) Dotplot shows enriched KEGG pathways in each subcluster. (D) Proportion of each subcluster in each sample. (E) Marker genes and enriched KEGG pathways of N, P, and M in C0. (F) A high level of the endothelial signature, CD300LG, IGFBP7, or CXCR4 mRNA expression is associated with a poor prognosis in TCGA-STAD. P < 0.001, by a log-rank test
3.5 Fibroblast infiltration is associated with ovarian metastasis of GC
As above mentioned (Fig. 1E), fibroblast cells composed the majority of TME in the metastasis tissue. Thus, we further investigated the role of fibroblast cells on ovarian metastasis in GC. In total 3385 fibroblast cells with expression of VIM, DCN, and COL1A1 genes [cancer-associated fibroblasts (CAFs), Supplementary Fig. S2] were integrated and clustered into 6 subclusters (560 from N, 294 from P, and 2531 from M) (Fig. 6A). The marker genes of each cluster were shown in (Fig. 6B). DEGs in these subclusters showed different pathway enrichment (Fig. 6C). C0 cluster was the major part of the ovarian metastasis tissue and enriched Ribosome biogenesis (Fig. 6C, D). The cell fraction of C0 cluster was also higher in the metastasis tissue than that in the adjacent non-tumoral tissue and the primary tumor (Fig. 6D). In contrast, the cell fraction of C1 cluster was low in the ovarian metastasis tissue compared to the adjacent non-tumoral tissue and the primary tumor (Fig. 6D). C1 cluster enriched Focal adhesion, TGF-beta signaling pathway, and PI3K-Akt signaling pathway (Fig. 6C). Like endothelial cells, fibroblast cells in C1 cluster also showed different gene expression between the ovarian metastasis tissue and the primary tumor (Fig. 6E). Thus, we defined genes specifically highly expressed in fibroblast cells from metastasis samples as metastasis-related-fibroblast signature. Although the statistical analysis was not significant, TCGA-STAD samples with high values of the metastasis-related-fibroblast signature showed poor prognosis (Fig. 6F). In addition, TCGA-STAD samples highly expressed PLCXD3, GPC3, and MCC have worse survival outcomes (Fig. 6F). These data demonstrate that fibroblast infiltration is associated with ovarian metastasis of GC.
Fibroblast infiltration is associated with ovarian metastasis of GC. (A) UMAP of fibroblast cells in different specimens by their distinct origination. (B) Dotplot shows expression of marker genes in each subcluster. (C) The dot plot shows enriched KEGG pathways in each subcluster. (D) Proportion of each subcluster in each sample. (E) Marker genes and enriched KEGG pathways of N, P, and M in C1 cluster. (F) A high level of the fibroblast signature, PLCXD3, GPC3, or MCC mRNA expression is associated with a poor prognosis in TCGA-STAD
To explore the differences of the CAFs of ovarian metastasis of GC from primary ovarian tumors, we downloaded an ovarian tumor dataset (GSE184880), which included 7 treatment-naïve patients with high-grade serous ovarian cancer (HGSOC) at early or late stages and 5 age-matched nonmalignant ovarian samples [28]. After quality control and data integration, we got 19,356 fibroblast cells and grouped into 7 subclusters (Fig. 7A). Intriguingly, we found a subcluster C4, which composed nearly 40% of the CAFs in the ovarian metastasis tissue of GC while absent in normal ovarian tissues and less than 20% of primary ovary tumors (Fig. 7B). We further analyzed genes highly expressed in C4 cluster and found that Microfibrillar-associated protein 4 (MFAP4) [29] and Calpain Small Subunit 1 (CAPNS1) [30] were specifically expressed in this cluster CAFs (Fig. 7C). KEGG pathway analysis revealed that thermogenesis, oxidative phosphorylation, regulation of actin cytoskeleton, and glycolysis/gluconeogenesis were enriched in C4 cluster (Fig. 7D). These data demonstrate that MFAP4- and CAPNS1-positive CAFs may be important for ovarian metastasis of GC.
Difference of fibroblasts among ovarian metastasis of GC, primary ovarian tumors, and normal ovarian tissues. (A) UMAP of fibroblast cells in two ovary metastasis tissues of GC, 5 primary ovarian tumors, and 5 normal ovarian tissues. (B) Proportion of each subcluster in each condition. (C) Dotplots show expression of marker genes in each subcluster. (D) The dot plot shows enriched KEGG pathways in C4 cluster
3.6 Intercellular communications are reduced in ovarian metastasis tissues compared to the primary tumors
To explore the mechanism of the accumulation of M0 macrophage, endothelial, and fibroblasts in ovarian metastasis, we analyzed the differential intercellular communications in the adjacent non-tumoral tissue, primary tumor, and ovarian metastasis tissue. Compared with the normal tissue, the number and intensity of proteins in fibroblasts and endothelial interacted with T and B cells increased in the primary tumor (Fig. 8A, B). In contrast, the number and intensity of proteins in fibroblasts and endothelial interacted with T and B cells decreased in the ovarian metastasis tissue compared to the primary tumor (Fig. 8C, D). For example, the interaction between COL1A1, COL1A2, and COL4A1 in fibroblasts and CD44, SDC1 in B cells decreased in ovarian metastasis tissues, CSF1 in fibroblasts and CSF1R in macrophage decreased, HLA-A, HLA-C, HLA-E in fibroblasts and CD8A in T cells decreased (Fig. 8E). These data indicates that the intercellular communications are reduced in the ovarian metastasis tissue compared to the primary tumor.
Intercellular communications are reduced in ovarian metastasis tissue compared to the primary tumor. (A) Differential number of interactions of primary tumor compared to adjacent non-tumor sample. (B) Interaction strength of N and P respectively. (C) Differential number of interactions of ovarian metastasis compared to primary tumor. (D) Interaction strength of P and M respectively. (E) Decreased signaling ligand-receptor pairs between fibroblasts and other cell types in ovarian metastasis compared to primary tumor
4 Discussion
A comprehensive description of tumor microenvironment will promote to find new strategies to improve the prognosis of ovarian metastasis of GC. Although the heterogeneity and microenvironment of primary GC and lymph node metastasis has been analyzed in several studies [31,32,33], the TME of ovarian metastasis of GC remains elusive. In this study, we reveal the heterogeneity of cellular compositions, gene expression patterns, and TME among the primary tumor, the matched adjacent non-tumoral and ovarian metastasis tissues from two patients with gastric cancer using scRNA-Seq.
In our study, the cellular compositions vary greatly between the ovarian metastasis tissues and primary tumors, while the primary tumors showed a similar pattern with the adjacent non-tumor tissues. Samples received treatments had a higher proportion of stromal cells like fibroblasts and endothelial cells, while immune cells like B cells and T cells make up most of the treatment-naïve samples. Meanwhile, irrelevant to whether received treatments, ovarian metastasis tissues had a higher proportion of stromal cells than adjacent non-tumoral tissues and primary tumors. In line with the metastatic phenotype, the ratio of B cells and cytotoxic T cells was lower in the ovarian metastasis tissues.
The polarization of macrophages between M1 and M2 phenotypes played an important role in tumor metastasis [34, 35]. However, our results indicate that the proportion of unpolarized macrophages (M0) is high in the ovarian metastasis tissue compared to the primary tumor and the adjacent non-tumor tissue, while M2 macrophages show the opposite pattern. More clinical specimen analyses are warranted for this observation.
Endothelial cells play a vital role in tumor metastasis [36]. In our study, we defined genes highly expressed in endothelial cells in the ovarian metastasis as a metastasis-related-endothelial signature, and patients with a high signature value in the TCGA-STAD dataset show a significantly worse survival than those with low signature value. Specifically, patients with high expression of CD300LG, IGFBP7, or CXCR4 had a poor prognosis. Since IGFBP7 promoted tumor progression in colon cancer [37] and CXCR4 played as essential role in metastasis in breast cancer and colon cancer [38,39,40], it is suggested that IGFBP7 and CXCR4 may be new prognostic factors for ovarian metastasis of gastric cancer.
The diverse functions of cancer-associated fibroblasts (CAFs) have been studied in various tumors [41]. In this study, we indicate that the ratio of the CAFs is high in the ovarian metastasis tissue compared to the primary tumor and the adjacent non-tumor tissue, which may be related to the ovarian tissue specificity. We also identify the genes highly expressed in the CAFs in the ovarian metastasis tissue as a metastasis-related-fibroblast signature and found that patients with high signature values in TCGA-STAD dataset show significantly poorer survival. We further characterized that a subgroup of CAFs with MFAP4 and CAPNS1 expression may be related to ovarian metastasis in GC. MFAP4 encodes a microfibrillar-associated protein 4 and is specifically expressed in early embryonic macrophages in zebrafish [42], which also regulates elastic fiber homeostasis, integration signaling, and cancer [43]. CAPNS1 is important for autophagy [30], IL-6 expression [44], inflammation and angiogenesis [45].
In summary, to our knowledge, we are the first to present a comprehensive description of the TME of two paired ovarian metastasis of gastric cancer, which will promote to uncover new therapeutic strategies for ovarian metastasis of gastric cancer.
5 Conclusions
Our findings provide a comprehensive microenvironment landscape of ovarian metastasis of gastric cancer and will promote to uncover new therapeutic strategies for this specific metastasis cancer.
Data availability
Processed raw count matrixes of scRNA-seq were placed on figshare (https://figshare.com/s/19dcb72884c9b49f366c).
References
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, CA Cancer J. Clin. 71, 209–249 (2021). https://doi.org/10.3322/caac.21660
E.C. Smyth, M. Nilsson, H.I. Grabsch, N.C. van Grieken, F. Lordick, Lancet 396, 635–648 (2020). https://doi.org/10.1016/S0140-6736(20)31288-5
M. Alsina, V. Arrazubi, M. Diez, J. Tabernero, Nat. Rev. Gastroenterol. Hepatol. 20, 155–170 (2023). https://doi.org/10.1038/s41575-022-00703-w
M. Zulfiqar, J. Koen, S. Nougaret, C. Bolan, W. VanBuren, M. McGettigan, C. Menias, AJR Am. J. Roentgenol. 215, 1020–1029 (2020). https://doi.org/10.2214/AJR.19.22184
H. Xie, B.J. Erickson, S.P. Sheedy, J. Yin, J.M. Hubbard, J. Gastrointest. Oncol. 12, 226–236 (2021). https://doi.org/10.21037/jgo-20-364
R. Lionetti, M. De Luca, A. Travaglino, A. Raffone, L. Insabato, G. Saccone, M. Mascolo, M. D’Armiento, F. Zullo, F. Corcione, Arch. Gynecol. Obstet. 300, 15–23 (2019). https://doi.org/10.1007/s00404-019-05167-z
J. Wang, Y.K. Shi, L.Y. Wu, J.W. Wang, S. Yang, J.L. Yang, H.Z. Zhang, S.M. Liu, Int. J. Gynecol. Cancer. 18, 825–832 (2008). https://doi.org/10.1111/j.1525-1438.2007.01078.x
H.K. Kim, D.S. Heo, Y.J. Bang, N.K. Kim, Gynecol. Oncol. 82, 105–109 (2001). https://doi.org/10.1006/gyno.2001.6210
V. Kumar, K. Ramnarayanan, R. Sundar, N. Padmanabhan, S. Srivastava, M. Koiwa, T. Yasuda, V. Koh, K.K. Huang, S.T. Tay, S.W.T. Ho, A.L.K. Tan, T. Ishimoto, G. Kim, A. Shabbir, Q. Chen, B. Zhang, S. Xu, K.P. Lam, H.Y.J. Lum, M. Teh, W.P. Yong, J.B.Y. So, P. Tan, Cancer Discov. 12, 670–691 (2022). https://doi.org/10.1158/2159-8290.CD-21-0683
Y. Li, X. Hu, R. Lin, G. Zhou, L. Zhao, D. Zhao, Y. Zhang, W. Li, Y. Zhang, P. Ma, H. Ren, X. Liao, P. Niu, T. Wang, X. Zhang, W. Wang, R. Gao, Q. Li, G. Church, J. He, Y. Chen, Theranostics. 12, 3818–3833 (2022). https://doi.org/10.7150/thno.71833
Y. Sun, S. Chen, Y. Lu, Z. Xu, W. Fu, W. Yan, Commun. Biol. 7, 63 (2024). https://doi.org/10.1038/s42003-024-05765-x
X. Wang, X. Shi, H. Lu, C. Zhang, X. Li, T. Zhang, J. Shen, J. Wen, Adv. Sci. (Weinh) 9, e2200546 (2022) https://doi.org/10.1002/advs.202200546
A. Sathe, S.M. Grimes, B.T. Lau, J. Chen, C. Suarez, R.J. Huang, G. Poultsides, H.P. Ji, Clin. Cancer Res. 26, 2640–2653 (2020). https://doi.org/10.1158/1078-0432.CCR-19-3231
P. Zhang, M. Yang, Y. Zhang, S. Xiao, X. Lai, A. Tan, S. Du, S. Li, Cell. Rep. 27, 1934–1947 (2019). https://doi.org/10.1016/j.celrep.2019.04.052
K. Sun, R. Xu, F. Ma, N. Yang, Y. Li, X. Sun, P. Jin, W. Kang, L. Jia, J. Xiong, H. Hu, Y. Tian, X. Lan, Nat. Commun. 13, 4943 (2022). https://doi.org/10.1038/s41467-022-32627-z
B. Kang, J. Camps, B. Fan, H. Jiang, M.M. Ibrahim, X. Hu, S. Qin, D. Kirchhoff, D.Y. Chiang, S. Wang, Y. Ye, Z. Shen, Z. Bu, Z. Zhang, H.G. Roider, Genome Biol. 23, 265 (2022). https://doi.org/10.1186/s13059-022-02828-2
R. Wang, S. Song, J. Qin, K. Yoshimura, F. Peng, Y. Chu, Y. Li, Y. Fan, J. Jin, M. Dang, E. Dai, G. Pei, G. Han, D. Hao, Y. Li, D. Chatterjee, K. Harada, M.P. Pizzi, A.W. Scott, G. Tatlonghari, X. Yan, Z. Xu, C. Hu, S. Mo, N. Shanbhag, Y. Lu, M. Sewastjanow-Silva, A.A. Fouad Abdelhakeem, G. Peng, S.M. Hanash, G.A. Calin, C. Yee, P. Mazur, A.N. Marsden, A. Futreal, Z. Wang, X. Cheng, J.A. Ajani, L. Wang, Cancer Cell. 41, 1407–1426 (2023). https://doi.org/10.1016/j.ccell.2023.06.005
Y. Hao, S. Hao, E. Andersen-Nissen, W.M. Mauck 3rd, S. Zheng, A. Butler, M.J. Lee, A.J. Wilk, C. Darby, M. Zager, P. Hoffman, M. Stoeckius, E. Papalexi, E.P. Mimitou, J. Jain, A. Srivastava, T. Stuart, L.M. Fleming, B. Yeung, A.J. Rogers, J.M. McElrath, C.A. Blish, R. Gottardo, P. Smibert, R. Satija, Cell 184, 3573–3587 (2021). https://doi.org/10.1016/j.cell.2021.04.048
T. Wu, E. Hu, S. Xu, M. Chen, P. Guo, Z. Dai, T. Feng, L. Zhou, W. Tang, L. Zhan, X. Fu, S. Liu, X. Bo, G. Yu, Innovation (Camb). 2, 100141 (2021). https://doi.org/10.1016/j.xinn.2021.100141
Z. Tang, B. Kang, C. Li, T. Chen, Z. Zhang, Nucleic Acids Res. 47, W556-W560 (2019). https://doi.org/10.1093/nar/gkz430
S. Jin, C.F. Guerrero-Juarez, L. Zhang, I. Chang, R. Ramos, C.H. Kuan, P. Myung, M.V. Plikus, Q. Nie, Nat. Commun. 12, 1088 (2021). https://doi.org/10.1038/s41467-021-21246-9
J. Jiang, Y. Ding, M. Wu, X. Lyu, H. Wang, Y. Chen, H. Wang, L. Teng, Front. Oncol. 10, 1765 (2020). https://doi.org/10.3389/fonc.2020.01765
J. Lu, Y. Peng, R. Huang, Z. Feng, Y. Fan, H. Wang, Z. Zeng, Y. Ji, Y. Wang, Z. Wang, BMC Cancer. 21, 723 (2021). https://doi.org/10.1186/s12885-021-08456-6
L. Li, Z. Huang, K. Du, X. Liu, C. Li, D. Wang, Y. Zhang, C. Wang, J. Li, Front. Pharmacol. 13, 900699 (2022). https://doi.org/10.3389/fphar.2022.900699
R. Fristedt, D. Borg, C. Hedner, J. Berntsson, B. Nodin, J. Eberhard, P. Micke, K. Jirström, J. Gastrointest. Oncol. 7, 848–859 (2016). https://doi.org/10.21037/jgo.2016.11.07
M. Kanda, C. Tanaka, D. Kobayashi, H. Tanaka, D. Shimizu, M. Shibata, H. Takami, M. Hayashi, N. Iwata, Y. Niwa, S. Yamada, T. Fujii, G. Nakayama, M. Fujiwara, Y. Kodera, Int. J. Cancer. 139, 2290–2298 (2016). https://doi.org/10.1002/ijc.30286
H. Wang, J. Min, Y. Ding, Z. Yu, Y. Zhou, S. Wang, A. Gong, M. Xu, Biol. Proced. Online. 26, 1 (2024). https://doi.org/10.1186/s12575-023-00228-9
J. Xu, Y. Fang, K. Chen, S. Li, S. Tang, Y. Ren, Y. Cen, W. Fei, B. Zhang, Y. Shen, W. Lu, Clin. Cancer Res. 28, 3590–3602 (2022). https://doi.org/10.1158/1078-0432.CCR-22-0296
R. Kanaan, M. Medlej-Hashim, R. Jounblat, B. Pilecki, G.L. Sorensen, Matrix Biol. 111, 1–25 (2022). https://doi.org/10.1016/j.matbio.2022.05.008
F. Demarchi, C. Bertoli, T. Copetti, E.L. Eskelinen, C. Schneider, Autophagy. 3, 235–237 (2007). https://doi.org/10.4161/auto.3661
R. Wang, M. Dang, K. Harada, G. Han, F. Wang, M. Pool Pizzi, M. Zhao, G. Tatlonghari, S. Zhang, D. Hao, Y. Lu, S. Zhao, B.D. Badgwell, M. Blum Murphy, N. Shanbhag, J.S. Estrella, S. Roy-Chowdhuri, A.A.F. Abdelhakeem, Y. Wang, G. Peng, S. Hanash, G.A. Calin, X. Song, Y. Chu, J. Zhang, M. Li, K. Chen, A.J. Lazar, A. Futreal, S. Song, J.A. Ajani, L. Wang, Nat. Med. 27, 141–151 (2021). https://doi.org/10.1038/s41591-020-1125-8
Y. Qian, E. Zhai, S. Chen, Y. Liu, Y. Ma, J. Chen, J. Liu, C. Qin, Q. Cao, J. Chen, S. Cai, Int. J. Cancer. 151, 1367–1381 (2022). https://doi.org/10.1002/ijc.34172
H. Jiang, D. Yu, P. Yang, R. Guo, M. Kong, Y. Gao, X. Yu, X. Lu, X. Fan, Clin. Transl Med. 12, e730 (2022). https://doi.org/10.1002/ctm2.730
B. Kutryb-Zajac, G. Harasim, A. Jedrzejewska, O. Krol, A. Braczko, P. Jablonska, P. Mierzejewska, J. Zielinski, E.M. Slominska, R.T. Smolenski, Int. J. Mol. Sci. 22 (2021). https://doi.org/10.3390/ijms22073764
W. Li, X. Zhang, F. Wu, Y. Zhou, Z. Bao, H. Li, P. Zheng, S. Zhao, Cell. Death Dis. 10, 918 (2019). https://doi.org/10.1038/s41419-019-2131-y
K.E. de Visser, J.A. Joyce, Cancer Cell. 41, 374–403 (2023). https://doi.org/10.1016/j.ccell.2023.02.016
C. Rupp, M. Scherzer, A. Rudisch, C. Unger, C. Haslinger, N. Schweifer, M. Artaker, H. Nivarthi, R. Moriggl, M. Hengstschlager, D. Kerjaschki, W. Sommergruber, H. Dolznig, P. Garin-Chesa, Oncogene 34, 815–825 (2015). https://doi.org/10.1038/onc.2014.18
Y.M. Li, Y. Pan, Y. Wei, X. Cheng, B.P. Zhou, M. Tan, X. Zhou, W. Xia, G.N. Hortobagyi, D. Yu, M.C. Hung, Cancer Cell. 6, 459–469 (2004). https://doi.org/10.1016/j.ccr.2004.09.027
D.D. Cave, X. Hernando-Momblona, M. Sevillano, G. Minchiotti, E. Lonardo, Theranostics. 11, 5686–5699 (2021). https://doi.org/10.7150/thno.54027
J.A. Burger, T.J. Kipps, Blood 107, 1761–1767 (2006). https://doi.org/10.1182/blood-2005-08-3182
A. Costa, Y. Kieffer, A. Scholer-Dahirel, F. Pelon, B. Bourachot, M. Cardon, P. Sirven, I. Magagna, L. Fuhrmann, C. Bernard, C. Bonneau, M. Kondratova, I. Kuperstein, A. Zinovyev, A.M. Givel, M.C. Parrini, V. Soumelis, A. Vincent-Salomon, F. Mechta-Grigoriou, Cancer Cell. 33, 463–479 (2018). https://doi.org/10.1016/j.ccell.2018.01.011
A. Zakrzewska, C. Cui, O.W. Stockhammer, E.L. Benard, H.P. Spaink, A.H. Meijer, Blood 116, e1–11 (2010). https://doi.org/10.1182/blood-2010-01-262873
A. Mohammadi, G.L. Sorensen, B. Pilecki, Cells 11, (2022) https://doi.org/10.3390/cells11132115
B.P. Fairfax, F.O. Vannberg, J. Radhakrishnan, H. Hakonarson, B.J. Keating, A.V. Hill, J.C. Knight, Hum. Mol. Genet. 19, 720–730 (2010). https://doi.org/10.1093/hmg/ddp530
C. Yi, W. Wu, D. Zheng, G. Peng, H. Huang, Z. Shen, X. Teng, Cell. Death Dis. 11, 533 (2020). https://doi.org/10.1038/s41419-020-02737-x
Funding
This work was supported in part by National Key Research and Development Program of China (2022YFA1104001), National Natural Science Foundation of China (32371004 and 82072896), the Program of Shanghai Academic/Technology Research Leader (21XD1403100) to H.F.; Program of Shanghai Cancer Institute Master and Ph.D. (SB22-14) to G.C; and Shanghai Municipal Committee of Science and Technology (21Y11913200) to X.X.
Author information
Authors and Affiliations
Contributions
H.F. and X.X. designed and supervised the project. G.C., M.Z., X.L., C.X., Q.S., and B.S. performed experiments. G.C. and H.F. interpreted and/or reviewed the data and wrote the manuscript. M.Z., C.X., Q.S., B.S., and X.X. edited the manuscript. All authors contributed to the article and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval
The study was conducted according to the Ethical Principles of Measures for Ethical Review of Biomedical Research Involving Human Beings and the Declaration of Helsinki.
Consent to participate
All patients provided written informed consent prior to participating in any study-specific procedures.
Consent for publication
All authors read and approved the final manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, G., Zhang, M., Lin, X. et al. Single-cell RNA transcriptomic analyses of tumor microenvironment of ovarian metastasis in gastric cancer. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00974-2
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00974-2