Abstract
Theta glass electrospray emitters can rapidly mix solutions to investigate fast reactions that occur as quickly as 1 μs, but emitters with submicron tips have the unusual properties of desalting protein ions and affecting the observed abundances of some proteins as a result of protein–surface interactions. The role of protein physical properties on ion signal was investigated using 1.7 ± 0.1 μm and 269 ± 7 nm emitters and 100 mM aqueous ammonium acetate or ammonium bicarbonate solutions. Protein ion desalting occurs for both positive and negative ions. The signal of a mixture of proteins with the 269 nm tips is time-dependent and the order in which ions of each protein is observed is related to the expected strengths of the protein–surface interactions. These results indicate that it is not just the high surface-to-volume ratio that plays a role in protein adsorption and reduction or absence of initial ion signal, but the small diffusion distance and extremely low flow rates of the smaller emitters can lead to complete adsorption of some proteins and loss of signal until the adsorption sites are filled and the zeta potential is significantly reduced. After about 30 min, signals for a protein mixture from the two different size capillaries are similar. These results show the advantages of submicron emitters but also indicate that surface effects must be taken into account in experiments using such small tips or that coating the emitter surface to prevent adsorption should be considered.

Graphical Abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nanoelectrospray ionization (nanoESI) enables low sample flow rates resulting in little sample consumption and is widely used in applications where low sample consumption is essential [1, 2]. The electric field strength at the tip of small emitters is high, and droplets can be efficiently produced from solutions with high surface tension, such as water, even without other methods to facilitate droplet formation, such as sheath gas flow or addition of low surface tension solvents [3, 4]. The ability to form droplets directly from aqueous solutions containing high concentrations of buffer makes nanoESI especially important in native mass spectrometry [5, 6]. NanoESI can be implemented in a number of different ways, including borosilicate or fused silica emitters with or without metal coatings and microfabricated devices. Microfabricated devices can incorporate buffer exchange and nanoLC for proteomics applications [7, 8]. Theta glass borosilicate nanoESI emitters have two separate barrels and can be used to rapidly mix two solutions during the electrospray ionization process [9,10,11,12,13,14]. The reaction time can be controlled by changing droplet size, which depends on flow rate [10, 12, 13] or by changing the distance of the emitter tip to the ESI interface [14]. Flow rates can be controlled by varying the backing pressure on the emitter and/or by changing the emitter tip size. Reaction times of 100s of ms have been achieved with emitter tip sizes ~4 μm [14] and reaction times as low as 1 μs has been achieved with ~300 nm tips [12, 14]. These small tips have made possible the fastest mixing times of two different solutions to date.
In addition to enabling fast mixing, the submicron tips have some unusual properties with nanoESI, one of which appears to be highly advantageous. Fewer salt adducts are formed with smaller tips, and this effect has been reported from both water/methanol/acetic acid solutions in which proteins are denatured [15, 16] as well as from buffered aqueous solutions in which proteins have native conformations and activities and nonvolatile salts are present at high (≥150 mM) concentration [17, 18]. The phenomenon of ion desalting in small tips was investigated using traditional single barrel borosilicate emitters from buffered aqueous solutions containing high levels of nonvolatile salts and buffers. Charge-state distributions of proteins and protein complexes could be readily observed in nanoESI spectra of solutions containing 150 mM KCl and 25 mM Tris-HCl buffer at pH 7 using 0.5 μm tips but not with 1.7 μm tips [17]. Similar results were obtained with aqueous solutions with six commonly used nonvolatile buffers containing ≥150 mM of Na+ [18]. These results suggest a mechanism for salt adduct reduction in which initial ESI droplets are sufficiently small that they contain on average fewer than one protein molecule per droplet. The majority of droplets contain salt but no protein, and the droplets that do contain a protein molecule have a much lower salt-to-protein ratio than that in the initial solution. The lowering of the salt-to-protein ratio in smaller droplets may result in less extensive adduction of nonvolatile salts to protein ions. Nonvolatile salts are often removed from solution prior to ESI owing to their adverse effects on mass spectrometry performance, but submicron tips can be used to form protein and protein complex ions directly from solutions containing high concentrations of nonvolatile salts and buffers that are used to mimic either the intracellular or the extracellular environment [17, 18].
The relative abundances of protein ions can depend on tip size. For example, a mixture of myoglobin and cytochrome c in 100 mM aqueous ammonium bicarbonate resulted in the formation of ions of both proteins with 1.5 μm theta emitter tips but the formation of only myoglobin ions with 310 nm emitter tips [19]. It was proposed that the absence of cytochrome c signal in the smaller tips was from absorption due to the high surface-to-volume ratio, which enhances interactions between the positively charged proteins in solution and the negatively charged glass surface near neutral pH [19].
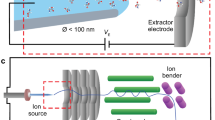
Here, we investigate the unusual phenomenon of selective protein ion signal observed when using submicron theta glass emitter tips and show that the ion desalting is also effective for negative ions. Emitters with smaller tips have lower volume-to-surface ratios at or near the emitter tip. Owing to the central divider, theta-glass emitters have even greater surface areas than single barrel emitters of the same size. The use of theta glass emitters in this study has the advantage of previously measured droplet lifetimes, and this information can be useful for distinguishing effects induced by the interfacial region between droplet formation and the mass spectrometer [12]. In this study, we compare two theta emitter sizes, 269 nm and 1.7 μm in diameter (Supplementary Figure S-1a, b) to explore the protein–glass surface interactions of proteins that have different conformations and physical properties. We also show that the effects of these surface–interactions and the resulting protein ion signal is time-dependent. These effects should be taken into consideration or eliminated by surface modifications in future studies that use submicron emitters for fast mixing or for ion desalting of biological samples.
Experimental
Nanoelectrospray ionization emitters are made from borosilicate theta capillaries (1.2 mm o.d./0.9 mm i.d., 0.15 mm septum thickness, Sutter Instruments, Novato, CA, USA) by pulling the tips to o.d. of 269 ± 7 nm or 1.7 ± 0.1 μm in the long dimension (the short dimension is 82% and 78% of the long dimension, respectively) with a Flaming/Brown micropipette puller (model P-87, Sutter Instruments, Novato, CA, USA). Tip diameters are measured with a scanning electron microscope (Hitachi TM-1000 SEM, Schaumburg, IL, USA). Both channels are filled with the same aqueous ammonium acetate (AA) or ammonium bicarbonate (ABC) solution containing proteins. Two 0.127 mm diameter platinum wires are inserted into the two barrels of a theta emitter and are in contact with the sample solution. Ion formation is initiated by placing the emitter tips 1 to 2 mm distant from the instrument orifice and applying the lowest possible (±0.6–1 kV) voltage to the platinum wires in order to produce stable ion formation. A nitrogen gas backing pressure of 10 psi is applied to the end of the theta emitters. Mass spectra are acquired with an LTQ mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) with a capillary temperature of 265 °C.
Reduced/alkylated RNase A solution is prepared by dissolving lyophilized RNase A powder in a 100 mM ABC (pH 8.7) solution containing 6 M guanidine hydrochloride, 15 mM dithiothreitol, and 100 mM iodoacetamide. The sample is reacted in the dark for approximately 2 h and buffer exchanged once into 100 mM ABC using a biospin column (Bio-Rad, Hercules, CA, USA). The final concentration of the stock solution of reduced/alkylated RNase A is 200 μM. Reduced/alkylated lysozyme is prepared by first reducing the lysozyme in 6 M guanidine hydrochloride, 15 mM dithiothreitol, and 100 mM ABC solution (pH 8.8) for 1/2 h at 37 °C, then reacted in the dark with 150 mM iodoacetic acid at room temperature for about 2 h. The 200 μM stock solution of reduced/alkylated lysozyme is buffer exchanged once into 100 mM ABC using a biospin column (Bio-Rad, Hercules, CA, USA). The change in the isoelectric point for the reduced proteins compared with the intact protein is computed using ExPASy (SIB Swiss Institute of BioInformatics) [20] by replacing cysteine residues with acidic or basic residues in the protein sequence and computing the shift in isoelectric point for the two forms of the protein.
Lyophilized protein powders of equine cytochrome c, bovine pancreatic ribonuclease A (RNase A), egg white lysozyme, bovine ubiquitin, ammonium acetate, ammonium bicarbonate, guanidine hydrochloride, dithiothreitol, iodoacetamide, and iodoacetic acid are from Sigma (St. Louis, MO, USA). Protein solutions were prepared at different concentrations either in 100 mM ABC or 100 mM AA.
Results and Discussion
Protein Conformation and Signal with Small Tips
Many factors affect protein signal in ESI, including solvents, sequence, conformation, instrument parameters, etc. [21,22,23,24]. Recent results indicate that interactions between positively charged proteins and the negatively charged emitter surface at small tip size can also be a contributing factor [19]. The role of emitter tip size was investigated with theta glass capillaries with tip sizes of 1.7 ± 0.1 μm and 269 ± 7 nm. The effects of protein conformation on the nanoESI signal with these two different size theta emitters were investigated by reducing internal disulfide bonds of bovine pancreatic ribonuclease A (RNase A) and capping the S-H groups by alkylation to prevent reformation of these bonds. RNase A is a 13.7 kDa protein with a high isoelectric point (pI 9.6) [25, 26] and it has a net positive charge in ammonium acetate and ammonium bicarbonate solutions. Results from circular dichroism [27, 28], hydrogen-deuterium exchange [29, 30], and X-ray scattering [31] show that reduced RNase A is significantly unfolded whereas intact RNase A has a compact, folded structure. Ubiquitin (pI 6.8) [32] has a net negative charge in ABC and has no net charge in AA. It is used as an internal standard because it should not interact significantly with the negatively charged surfaces of the borosilicate emitters. Mass spectra of the mixture of 10 μM RNase A and 1 μM ubiquitin from 100 mM ABC using 1.7 μm and 269 nm theta emitters are shown in Figure 1a and d, respectively. The relative abundances of these two proteins differ slightly with the two different tips. The charge-state distributions of both proteins are shifted to slightly lower charge with the 269 nm emitters. Higher charging with small tips has been reported previously, and this has been attributed to electrothermal supercharging or higher electric field with small tips [16, 19, 33]. Here, this effect may be due to the different droplet sizes and thus lifetimes of these droplets. The 269 nm theta emitters form smaller droplets that have lifetimes of around 1 μs whereas droplets from the 1.7 μm emitters are larger and have lifetimes of about 26 μs [12]. The emitter tips are approximately 1 to 2 mm away from the instrument orifice. In order for the small droplets formed from 269 nm theta emitters to enter the instrument orifice, they must have a velocity that is at least 1000 m/s. Thus, ions formed from these small droplets are likely desolvated or nearly desolvated before entering the heated capillary of the instrument. In contrast, the larger droplets formed from the 1.7 μm emitters have sufficient lifetimes that they likely enter the heated ion transfer tube. Heating of the droplet inside the heated transfer tube can destabilize the folded form of proteins, which leads to higher protein charge states in the mass spectra using the 1.7 μm theta emitters. There is no effect of tip size on protein charge state in AA, which is consistent with electrothermal supercharging results that suggest that thermal protein unfolding occurs less readily in AA [34, 35].
Electrospray ionization mass spectra of ubiquitin with RNase A (a, d), reduced RNase A (b, e), and both lysozyme and reduced lysozyme (c, f) in 100 mM ABC with 1.7 μm theta emitters (a-c) and 269 nm theta emitters (d-f). The charge states of each protein are color coded and labeled. * indicates polydimethylsiloxane clusters that are present as an impurity
Mass spectra of a mixture of 10 μM reduced carbamidomethylated RNase A (“amide form”) and 5 μM ubiquitin in 100 mM ABC are shown in Figure 1b and e for the two different tip sizes. The charge-state distribution of reduced RNase A is bimodal with the higher charge states (10+ to 18+) most abundant, indicating that reduced RNase A is predominantly unfolded in solution. Both ubiquitin and reduced RNase A ions are formed with the 1.7 μm theta emitters. In contrast, the abundance of reduced RNase A ions is dramatically lower with the 269 nm theta emitters. The abundance of reduced RNase A ions relative to the total protein ion abundance (reduced RNaseA and ubiquitin) is 82% ± 1% and 5.1% ± 0.3% with the 1.7 μm and 269 nm theta emitters, respectively. The significantly lower abundance of reduced RNase A compared with folded RNase A with the 269 nm theta emitters indicates that the unfolded conformation of the protein is likely the origin of the very low signals with the 269 nm emitters.
Effects of conformation on protein ion signal and tip size were further investigated by measuring ESI mass spectra of 10 μM egg white lysozyme, 5 μM reduced carboxymethylated lysozyme (“acid form”), and 2.5 μM ubiquitin mixture in 100 mM ABC (Figure 1c and f). Lysozyme is a 14.7 kDa basic protein (pI 11.4) with four disulfide bonds and has a net positive charge in these solutions [36, 37]. The conformation of reduced lysozyme is similar to that of lysozyme denatured in 4 M guanidine hydrochloride [38]. Charge-state distributions for all three proteins are produced with the 1.7 μm theta emitters. In contrast, there is no signal for reduced lysozyme with the 269 nm theta tips. The absence of signal for reduced lysozyme but not intact lysozyme in the 269 nm tips is consistent with the results for reduced/intact RNase A, providing additional evidence that the ion signal for largely denatured proteins with isoelectric points higher than 7 that have net positive charges in these solutions is significantly reduced with submicron emitters.
To determine the effect of protein charge, mass spectra of reduced α-lactalbumin (four disulfide bonds) with carboxymethylated S-H groups (“acid form”) were obtained with both 1.7 μm and 269 nm tips from ABC. The ion signal for carboxymethylated α-lactalbumin is comparable to but slightly lower with the 269 nm tips (Supplementary Figure S-2). The pI of carboxymethylated α-lactalbumin is lower than 4.2, so this unfolded protein has a net negative charge in these solutions [39]. The observation that significantly lower signal is observed for unfolded proteins with a net positive charge in solution but only slightly lower signal is observed for unfolded proteins with a net negative charge in solution is consistent with positively charged proteins interacting strongly with the negatively charged surface of the smaller emitters where the surface-to-volume ratio is greatest. Unfolded proteins have potentially more sites of interaction with surfaces and may be expected to interact more strongly when the number of positive charges exceeds the number of negative charges on the protein.
Another feature of the 269 nm theta emitters is that the protein ions have fewer salt adducts (Figure 1d–f) compared with those formed by the 1.7 μm theta emitters (Figure 1a–c). This effect of small tips on desalting protein ions has been reported with both buffered aqueous solutions with ≥150 mM Na+ or K+ [17, 18] and water/methanol/acetic acid solutions [15, 16]. These results are consistent with the hypothesis that the smaller initial droplets formed with the 269 nm emitters have fewer than one protein per droplet so that the initial droplets have a much lower salt-to-protein ratio compared with that in the original solution or to the larger droplets formed from 1.7 μm tips, which have on average more than one protein molecule. The lower salt-to-protein ratio in the initial droplet leads to fewer salt adducts on the desolvated protein ions [17].
Time Evolution of Protein Signals
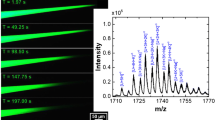
To further investigate the origin of the extraordinarily low signal for unfolded proteins in the 269 nm emitter tips, experiments focusing on time as a variable were conducted. NanoESI mass spectra of a mixture of 2.5 μM ubiquitin, 10 μM lysozyme, 5 μM reduced lysozyme, and 5 μM cytochrome c in 100 mM ABC and 100 mM AA were continuously acquired for 30 min with both 1.7 μm and 269 nm emitters. A time sequence of mass spectra obtained from 100 mM ABC, each signal averaged for 6 min over the course of 30 min, is shown in Figure 2. With the 1.7 μm theta emitters, all four proteins are present in the first 6-min signal averaged mass spectrum (Figure 2a). The charge-state distributions are similar to reported values for ubiquitin (5+ to 7+), lysozyme (6+ to 8+), reduced lysozyme (bimodal distribution 8+ to 17+), and cytochrome c (7+ to 8+) from ABC [34, 40, 41]. With the 269 nm theta emitters, charge-state distributions of only ubiquitin and lysozyme are observed for the first 12 min (Figure 2b, c), consistent with the results shown in Figure 1f. The peaks for both proteins are narrower attributable to fewer salt adducts. After 12 min, ions of reduced lysozyme appear (Figure 2d, e), and cytochrome c ions appear after 24 min (Figure 2f). The mass spectrum acquired after 24 min with a 269 nm theta emitter (Figure 2f) is comparable to the mass spectrum initially obtained with the 1.7 μm theta emitters, albeit with fewer salt adducts to the protein ions (Figure 2a). The same results are obtained for these proteins when 100 mM AA is used instead indicating that this effect is not unique to ABC (Supplementary Figure S-3). The protein signal of reduced lysozyme and cytochrome c takes many minutes to appear in mass spectra obtained with the 269 nm theta emitters, but there is signal for all four proteins with the 1.7 μm emitters at the start (Supplementary Figure S-3). Mortensen and Williams found that cytochrome c signal was not obtained from 100 mM ABC but was from 100 mM AA with a ~300 nm theta emitter [19]. Time was not a parameter considered in the previous studies, which may account for this earlier observation.
NanoESI mass spectra of ubiquitin (black), lysozyme (red), reduced lysozyme (blue) and cytochrome c (green) in 100 mM ABC solution obtained 1.7 μm theta emitters (a) and 269 nm theta emitters (b-f). Each mass spectrum is signal averaged for six minutes. * indicates polydimethylsiloxane clusters that are present as an impurity
Acidification of solutions can occur when nanoESI occurs for a long time [42, 43]. In these experiments, voltage is applied to the two platinum wires that are in contact with the protein solution. The platinum wire is 3–5 mm away from the emitter tip where droplet formation occurs. Thus, it takes time for excess protons to diffuse to the end of the emitter tip. The effect of acidification on protein signal and charge state was investigated by spraying the same mixture of the four proteins in the ABC solution from 1.7 μm theta emitters for 30 min. No significant change of charge-state distributions occurs after 30 min (Supplementary Figure S-4). Similarly, the protein charge states that are formed with the 269 nm tips at the end of 30 min (Figure 2f) are similar to those formed with the larger tips at the start and in the end (Figure 2a and Supplementary Figure S-4). These results indicate that acidification does not play an important role in the time evolution of protein signal observed with the 269 nm tips.
The time evolution of the protein ion signal in 1-min intervals for lysozyme, reduced lysozyme, cytochrome c, and ubiquitin is shown in Figure 3. Both the signal of each protein relative to that of ubiquitin and the fraction of signal of each protein relative to the total ion abundance are shown in Figure 3a and b, respectively. The relative ion abundance of each protein relative to the absolute ion abundance of ubiquitin shown on the y-axis is defined as
where I i is the total abundance of a protein ion of interest and I j is the total abundance of ubiquitin ions. These data show that there is significant variability of signal with these small tips but there are clear trends in the ion abundances with time. Data for two additional emitters and three emitters for AA show the same trends of ion abundance with time (Supplementary Figure S-5a, b). Initially, only ubiquitin ions are formed in both ABC and AA solutions. Lysozyme ions appear 1 to 5 min after nanoESI is initiated and ubiquitin ions are observed (Supplementary Figure S-4). The time required for reduced lysozyme signal to appear depends on the individual theta emitter, and ranges from several minutes to 15 min. The signal for both forms of lysozyme increases with time. Cytochrome c signal appears after 25 min to 30 min with either AA or ABC solutions. The order of appearance of the four protein ions is consistently ubiquitin, lysozyme, reduced lysozyme, and cytochrome c for both ABC and AA buffer using the 269 nm theta emitters.
These results show that reduced lysozyme and cytochrome c ions can be formed from the 269 nm theta emitters with both ABC and AA solutions, but it takes significantly longer for these ions to appear than when these ions are formed with the 1.7 μm theta emitters. The significant delay of ion formation for reduced lysozyme and cytochrome c indicates that these two proteins do not enter the droplets when nanoESI is initiated with a 269 nm theta emitter. We hypothesize that these proteins present in the solution near the emitter tip interact with the inner surface of the glass emitter, thereby making them unavailable to enter the initial droplets that are formed when nanoESI is initiated. The hydroxyl groups at the glass surface are deprotonated when the solution pH is greater than 3 [44, 45]. The pH of the ammonium bicarbonate and ammonium acetate solutions are 7.8 and 6.8, respectively. Thus, the inner surface of the emitter is deprotonated and has a net negative charge with these solutions. Ubiquitin has an isoelectric point of 6.8, so it has no net charge in the AA solution and has a net negative charge in ABC solution. Cytochrome c has a slightly lower isoelectric point (pI 10.04) [46] than lysozyme (pI 11.4) but does not have disulfide bonds. Both proteins are positively charged in AA or ABC solution. Calculations indicate that the additional acidic groups on the eight cysteine residues of lysozyme lower the isoelectric point from 11.4 to 8.6. Thus, reduced carboxymethylated lysozyme has lower pI than cytochrome c, but a net positive charge in the aqueous buffer solutions. The favorable electrostatic interactions between positively charged proteins and the negatively charged emitter inner surface can lead to a decrease in concentration of free protein in solution and a loss of signal for these proteins in the initial electrospray droplets.
It is interesting to compare lysozyme and cytochrome c because both proteins are positively charged in solution, are similar in size, and have similar radii of gyration (Rg) (13.1 Å for folded cytochrome c [47] and 13.3 Å for folded lysozyme [48, 49]). However, lysozyme ions appear a short time after nanoESI is initiated, but cytochrome c appears after ~25 min with the 269 nm emitter tips. There are more basic residues on one side of cytochrome c and more acidic residues on the other side resulting in a high dipole moment for this protein [50]. In contrast, lysozyme has more uniformly dispersed basic and acidic residues resulting in a lower dipole moment (Figure 4) [50]. The lower dipole moment and lower density of basic residues on one side of lysozyme may lead to a weaker interaction with the negatively charged borosilicate glass surface than cytochrome c.
A schematic illustration of protein conformation and the distribution of charges of reduced lysozyme, lysozyme, cytochrome c, and ubiquitin. Ubiquitin has a net negative charge in 100 mM ABC and is neutral in 100 mM AA, thus it should have minimal interactions with the surface. Cytochrome c has net positive charge, and has more positive charges on one side of the molecule. Lysozyme also has a net positive charge but the positive charges are more uniformly distributed over the molecule. Reduced lysozyme has a net positive charge and has a flexible conformation that allows more interactions with the glass surface
The order of appearance of the protein ion signal with the 269 nm emitter tips is different from that in cation exchange chromatography, where lysozyme binds more strongly to a stationary phase consisting of agarose or cellulose beads coated with anionic polymers [51]. In contrast, Moerz et al. showed that at the pH of the ABC and AA buffers, more than twice as much cytochrome c absorbs onto SBA-15 silica particles than lysozyme [52]. The latter result is consistent with our observations for a nearly identical material although differences in surface roughness may also affect these results.
The binding affinities between the emitter surface and the different proteins depend not only on protein net charge and size, but it also depends on the accessibility of the charged amino acid residues. The conformation of reduced lysozyme is considerably more flexible than that of intact lysozyme, which has limited structural flexibility owing to the four internal disulfide bonds. More conformational flexibility should lead to a greater number of possible interactions between protonated basic residues and the negatively charged surface and, hence, a greater electrostatic interaction for reduced lysozyme and the glass surface (Figure 4). Reduced lysozyme appears sooner in mass spectra than cytochrome c possibly because reduced lysozyme has lower pI than cytochrome c. This may also be due to higher repulsive interactions between the negatively charged amino acid groups in reduced lysozyme, which must also be stronger owing to their closer proximity to the glass surface (Figure 4). Thus, the order in which these proteins are observed in time appears to be related to their relative binding strengths of these proteins to the anionic surface of the emitters.
In order to estimate the surface coverage corresponding to adsorption of all protein onto the emitter surface, the surface area and volume of solution in each channel up to 1.0 mm from the tip was estimated. For a 600 nm tip, the open area of each channel is about 150 nm in the long dimension (Supplementary Figure S-1c). Thus, the open area of the 269 nm theta emitter cross-section was estimated as a split circle with 135 nm diameter. The surface area was estimated by using a volume element of a half cylinder with a radius of 68 nm and length of 1.0 mm. With 20 μM total concentration of proteins with a net positive charge, only about 0.3% of the surface of the capillary would be covered with protein if every protein in solution in this volume adhered to the surface (modeled with cytochrome c with a diameter of 3.4 nm). Thus, there is more than sufficient capillary surface area in these 269 nm emitters to completely remove protein from the initial solutions at the capillary tips. Even with the 1.7 μm theta emitters, less than 2% of the surface would be covered if all of the protein in solution adhered to the capillary surface. The low surface coverage for both the 269 nm and the 1.7 μm theta emitters indicates that diffusion plays an important role. The diffusion distance is six times longer in the 1.7 μm theta emitters than it is in the 269 nm emitters. Moreover, the solution flow rate in the 1.7 μm theta emitters (~1000 pL/s) is one order of magnitude higher than the 269 nm theta emitters (~100 pL/s) [12]. As a result, a much smaller fraction of protein molecules can interact with the capillary surface inside the 1.7 μm theta emitters owing to both the longer diffusion length and significantly faster flow rate. Thus, the surface effects observed with the smaller capillaries are significant not just because of the higher surface-to-volume ratio but also because of the smaller diffusion length and significantly lower flow rate that enable a greater fraction of the proteins to interact with the surface. As solution flows through the emitter tip, more protein in solution can bind to the surface until the surface is sufficiently coated with positively charged protein ions. This will significantly lower the zeta potential and result in significantly lower protein–surface interactions. Competitive binding of proteins should result in the displacement of weaker binding proteins consistent with the order of appearance of protein ions in these experiments with time.
Competing Protein–Surface Interaction
In order to investigate potential competitive binding of proteins to the emitter surfaces, solutions of just cytochrome c (10 μM) and ubiquitin (2.5 μM) without other proteins present were prepared in AA and ABC. Results with both positive and negative ions were obtained and are consistent (Figure 5a–f and Supplementary Figure S-6a–f). Results for negative ions are shown to illustrate that the ion desalting effect with the small tips also occurs for protein anions. Both ubiquitin and cytochrome c anions are initially formed with the 1.7 μm theta emitters (Figure 5a, b). With the 269 nm emitters, only ubiquitin ions are formed initially. No cytochrome c ions are observed for the first 8 min, but ions for both proteins are observed from 9 to 13 min (Figure 5e and f). The slightly lower charge states formed from ABC may reflect differences in protein conformation or stabilities in these different buffers. Cytochrome c signal is delayed with the 269 nm theta emitters for both buffers when either positive or negative ions are formed, consistent with our proposed mechanism that proteins that are positively charged in solution can interact with the negatively charged glass surface prior to the droplet formations.
Cytochrome c ions appear at a shorter time in this solution (10 μM) than the one that contains 5 μM reduced lysozyme and 5 μM cytochrome c, and appear at a time comparable to where reduced lysozyme is first observed in the latter solution. These results are consistent with cytochrome c displacing reduced lysozyme on the surface when both these proteins are present at lower concentration.
Conclusions
The effects of protein structure and charge on ion signal were investigated using 1.7 μm and 269 nm theta glass emitters. Significantly lower salt adduction to both positive and negative ions occurs with the submicron emitters. This effect is attributed to a lower salt-to-protein ratio in nanoESI droplets that are sufficiently small that most droplets contain salt but no protein, which lowers the salt-to-protein ratio in small nanodrops that do contain a protein molecule. The signal for a mixture of proteins does not depend on time over the course of 30 min with the 1.7 μm emitters but a significant time dependence is observed with the 269 nm tips. The signal for unfolded forms of a protein that has a net positive charge in solution is initially reduced signficantly or is absent compared with the folded forms of the same protein with the 269 nm tips, but signals for these proteins appear after times as long as 25 min. The same effect is not observed for an unfolded form of a protein that is negatively charged in solution, indicating that the charge of the protein in solution plays a key role. These results are all consistent with positively charged proteins in solution interacting with and adhering to the negatively charged glass surface of the emitters, thereby reducing the concentration of these proteins in the initial electrospray droplets. A key finding of this work is that it is not just the high surface to volume ratio of the small emitters that is important because both the small and large emitters have sufficient surface areas to absorb all the protein in the first 1.0 mm distance from the tip. However, the diffusion length of the smaller emitters is significantly shorter and the flow rate is significantly slower. These factors enable the protein in the small emitters to interact with and adhere to the emitter surface more efficiently. The use of small emitters appears to be highly advantageous for desalting protein ions and for mixing two solutions at a fast rate that is not achievable with conventional mixing apparatus. However, analyte–surface interactions can occur with these small emitters and these effects should be considered in experiments using such small tips. These interactions could potentially be significantly reduced or even eliminated by derivatizing the emitter surface to produce little or no net surface charge. The charge of the glass surface depends on pH and is deprotonated above pH 3. Thus, the surface effects observed with the submicron emitters are significant when using buffered aqueous solutions near neutral pH that are commonly used in native mass spectrometry. These effects should be considerably less significant for solutions consisting of water/methanol/acetic acid solutions when the pH is below 3 and the hydroxyl groups at the glass surface are predominately protonated with no net charge. These effects should also be slightly less significant for single channel emitters compared with theta glass emitters that have the same overall diameter owing to the lower surface area and longer diffusion distance.
References
Valaskovic, G.A., Kelleher, N.L., Little, D.P., Aaserud, D.J., McLafferty, F.W.: Attomole-sensitivity electrospray source for large-molecule mass spectrometry. Anal. Chem. 67, 3802–3805 (1995)
Gong, X., Zhao, Y., Cai, S., Fu, S., Yang, C., Zhang, S., Zhang, X.: Single cell analysis with probe ESI-mass spectrometry: detection of metabolites at cellular and subcellular levels. Anal. Chem. 86, 3809–3816 (2014)
Wilm, M., Mann, M.: Analytical properties of the nanoelectrospray ion source. Anal. Chem. 68, 1–8 (1996)
Juraschek, R., Dülcks, T., Karas, M.: Nanoelectrospray – more than just a minimized-flow electrospray ionization source. J. Am. Soc. Mass Spectrom. 10, 300–308 (1999)
Hernández, H., Robinson, C.V.: Determining the stoichiometry and interactions of macromolecular assemblies from mass spectrometry. Nat. Protoc. 2, 715–726 (2007)
Heck, A.J.R.: Native mass spectrometry: a bridge between interactomics and structural biology. Nat. Methods. 5, 927–933 (2008)
Zhang, S., Van Pelt, C.K.: Chip-based nanoelectrospray mass spectrometry for protein characterization. Expert Rev. Proteom. 1, 449–468 (2004)
Nissila, T., Sainiemi, L., Franssila, S., Ketola, R.A.: Fully polymeric integrated microreactor/electrospray ionization chip for on-chip digestion and mass spectrometric analysis. Sensors Actuators B Chem. 143, 414–420 (2009)
Mark, L.P., Gill, M.C., Mahut, M., Derrick, P.J.: Dual nano-electrospray for probing solution interactions and fast reactions of complex biomolecules. Eur. J. Mass Spectrom. 18, 439–446 (2012)
Mortensen, D.N., Williams, E.R.: Theta-glass capillaries in electrospray ionization: Rapid mixing and short droplet lifetimes. Anal. Chem. 86, 9315–9321 (2014)
Fisher, C.M., Hilger, R.T., Zhao, F., Mcluckey, S.A.: Electroosmotically driven solution mixing in borosilicate theta glass nESI emitters. J. Mass Spectrom. 50, 1063–1070 (2015)
Mortensen, D.N., Williams, E.R.: Ultrafast (1 μs) mixing and fast protein folding in nanodrops monitored by mass spectrometry. J. Am. Chem. Soc. 138, 3453–3460 (2016)
Mortensen, D.N., Williams, E.R.: Microsecond and nanosecond polyproline II helix formation in aqueous nanodrops measured by mass spectrometry. Chem. Commun. 52, 12218–12221 (2016)
Jansson, E.T., Lai, Y.-H., Santiago, J.G., Zare, R.N.: Rapid hydrogen–deuterium exchange in liquid droplets. J. Am. Chem. Soc. 139, 6851–6854 (2017)
Schmidt, A., Karas, M., Dülcks, T.: Effect of different solution flow rates on analyte ion signals in nano-ESI MS, or: When does ESI turn into nano-ESI? J. Am. Soc. Mass Spectrom. 14, 492–500 (2003)
Hu, J., Guan, Q.-Y., Wang, J., Jiang, X.-X., Wu, Z.-Q., Xia, X.-H., Xu, J.-J., Chen, H.-Y.: Effect of nanoemitters on suppressing the formation of metal adduct ions in electrospray ionization mass spectrometry. Anal. Chem. 89, 1838–1845 (2017)
Susa, A.C., Xia, Z., Williams, E.R.: Small emitter tips for native mass spectrometry of proteins and protein complexes from nonvolatile buffers that mimic the intracellular environment. Anal. Chem. 89, 3116–3122 (2017)
Susa, A.C., Xia, Z., Williams, E.R.: Native mass spectrometry from common buffers with salts that mimic the extracellular environment. Angew. Chem. Int. Ed. 56, 7912–7915 (2017)
Mortensen, D.N., Williams, E.R.: Electrothermal supercharging of proteins in native MS: effects of protein isoelectric point, buffer, and nanoESI-emitter tip size. Analyst. 141, 5598–5606 (2016)
Gasteiger, E., Hoogland, C., Gattiker, A., Duvaud, S., Wilkins, M.R., Appel, R.D., Bairoch, A.: Protein identification and analysis tools on the ExPASy Server. In: Walker, J.M. (ed.) The Proteomics Protocols Handbook, pp. 571–607. Humana Press, Totowa (2005)
Liang, Z., Yang, Q., Zhang, W., Zhang, L., Zhang, Y.: Effects of experimental parameters on the signal intensity of capillary electrophoresis electrospray ionization mass spectrometry in protein analysis. Chromatographia. 57, 617–621 (2003)
Cech, N.B., Enke, C.G.: Practical implications of some recent studies in electrospray ionization fundamentals. Mass Spectrom. Rev. 20, 362–387 (2001)
Iavarone, A.T., Jurchen, J.C., Williams, E.R.: Effects of solvent on the maximum charge state and charge state distribution of protein ions produced by electrospray ionization. J. Am. Soc. Mass Spectrom. 11, 976–985 (2000)
Kuprowski, M.C., Konermann, L.: Signal response of coexisting protein conformers in electrospray mass spectrometry. Anal. Chem. 79, 2499–2506 (2007)
Smyth, D.G., Stein, W.H., Moore, S.: The sequence of amino acid residues in bovine pancreatic ribonuclease: revisions and confirmations. J. Biol. Chem. 238, 227–234 (1963)
Tanford, C., Hauenstein, J.D.: Hydrogen ion equilibria of ribonuclease. J. Am. Chem. Soc. 78, 5287–5291 (1956)
Takahashi, S., Kotani, T., Yoneda, M., Ooi, T.: A circular dichroic spectral study on disulfide-reduced pancreatic ribonuclease A and its renaturation to the active enzyme. J. Biochem. 82, 1127–1133 (1977)
Qi, P.X., Sosnick, T.R., Englander, S.W.: The burst phase in ribonuclease A folding and solvent dependence of the unfolded state. Nat. Struct. Biol. 5, 882–884 (1998)
Woodward, C.K., Rosenberg, A.: Oxidized RNase as a protein model having no contribution to the hydrogen exchange rate from conformational restrictions. Proc. Natl. Acad. Sci. USA. 66, 1067–1074 (1970)
Bai, Y., Milne, J.S., Mayne, L., Englander, S.W.: Primary structure effects on peptide group hydrogen exchange. Prot. Struct. Funct. Genet. 17, 75–86 (1993)
Jacob, J., Dothager, R.S., Thiyagarajan, P., Sosnick, T.R.: Fully reduced ribonuclease A does not expand at high denaturant concentration or temperature. J. Mol. Biol. 367, 609–615 (2007)
Rechesteiner, M.C.: Ubiquitin. Plenum Press, New York (1988)
Yuill, E.M., Sa, N., Ray, S.J., Hieftje, G.M., Baker, L.A.: Electrospray ionization from nanopipette emitters with tip diameters of less than 100 nm. Anal. Chem. 85, 8498–8502 (2013)
Sterling, H.J., Cassou, C.A., Susa, A.C., Williams, E.R.: Electrothermal supercharging of proteins in native electrospray ionization. Anal. Chem. 84, 3795–3801 (2012)
Cassou, C.A., Williams, E.R.: Anions in electrothermal supercharging of proteins with electrospray ionization follow a reverse Hofmeister series. Anal. Chem. 86, 1640–1647 (2014)
Canfield, R.E.: The amino acid secpence of egg white lysozyme. J. Biol. Chem. 228, (1963)
Wetter, L.R., Deutsch, H.F.: Immunological studies on egg white proteins. IV. Immunochemical and physical studies of lysozyme. J. Biol. Chem. 192, 237–242 (1951)
Millett, I.S., Doniach, S., Plaxco, K.W.: Toward a taxonomy of the denatured state: Small angle scattering studies of unfolded proteins. Adv. Prot. Chem. 62, 241–262 (2002)
Bramaud, C., Aimar, P., Daufin, G.: Whey protein fractionation: Isoelectric precipitation of α-lactalbumin under gentle heat treatment. Biotechnol. Bioeng. 56, 391–397 (1997)
Going, C.C., Xia, Z., Williams, E.R.: New supercharging reagents produce highly charged protein ions in native mass spectrometry. Analyst. 140, 7184–7194 (2015)
Robinson, E.W., Sellon, R.E., Williams, E.R.: Peak deconvolution in high-field asymmetric waveform ion mobility spectrometry (FAIMS) to characterize macromolecular conformations. Int. J. Mass Spectrom. 259, 87–95 (2007)
Gatlin, C.L., Turecek, F.: Acidity determination in droplets formed by electrospraying methanol-water solutions. Anal. Chem. 66, 712–718 (1994)
Van Berkel, G.J., Zhou, F., Aronson, J.T.: Changes in bulk solution pH caused by the inherent controlled-current electrolytic process of an electrospray ion source. Int. J. Mass Spectrom. Ion Processes. 162, 55–67 (1997)
Brinker, C.J.: Hydrolysis and condensation of silicates: effects on structure. J. Non. Cryst. Solids. 100, 31–50 (1988)
Behrens, S.H., Grier, D.G.: The charge of glass and silica surfaces. J. Chem. Phys. 115, 6716–6721 (2001)
Barlow, G.H., Margoliash, E.: Electrophoretic behavior of mammalian-type cytochromes c. J. Biol. Chem. 241, 1473–1477 (1966)
Shiu, Y.-J., Jeng, U.-S., Huang, Y.-S., Lai, Y.-H., Lu, H.-F., Liang, C.-T., Hsu, I.-J., Su, C.-H., Su, C., Chao, I., Su, A.-C., Lin, S.-H.: Global and local structural changes of cytochrome c and lysozyme characterized by a multigroup unfolding process. Biophys. J. 94, 4828–4836 (2008)
Hamill, A.C., Wang, S.C., Lee, C.T.: Probing lysozyme conformation with light reveals a new folding intermediate. Biochemistry. 44, 15139–15149 (2005)
Ghosh, A., Brinda, K.V., Vishveshwara, S.: Dynamics of lysozyme structure network: probing the process of unfolding. Biophys. J. 92, 2523–2535 (2007)
Bharti, B., Findenegg, G.H.: Protein-specific effects of binding to silica nanoparticles. Chem. Lett. 41, 1122–1124 (2012)
DePhillips, P., Lenhoff, A.M.: Determinants of protein retention characteristics on cation-exchange adsorbents. J. Chromatogr. A. 933, 57–72 (2001)
Moerz, S.T., Huber, P.: PH-dependent selective protein adsorption into mesoporous silica. J. Phys. Chem. C. 119, 27072–27079 (2015)
Acknowledgments
This material is based upon work supported by the National Science Foundation Division of Chemistry under grant number CHE-1609866. The authors also thank Dr. Anna Susa and Dr. Daniel Mortensen for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(PDF 1172 kb)
Rights and permissions
About this article
Cite this article
Xia, Z., Williams, E.R. Protein-Glass Surface Interactions and Ion Desalting in Electrospray Ionization with Submicron Emitters. J. Am. Soc. Mass Spectrom. 29, 194–202 (2018). https://doi.org/10.1007/s13361-017-1825-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-017-1825-6