Abstract
Dissociations of z 4 ions from pentapeptides AAXAR where X = H, Y, F, W, and V produce dominant z 2 ions that account for >50 % of the fragment ion intensity. The dissociation has been studied in detail by experiment and theory and found to involve several isomerization and bond-breaking steps. Isomerizations in z 4 ions proceed by amide trans→cis rotations followed by radical-induced transfer of a β-hydrogen atom from the side chain, forming stable Cβ radical intermediates. These undergo rate-determining cleavage of the Cα–CO bond at the X residue followed by loss of the neutral AX fragment, forming x 2 intermediates. The latter were detected by energy-resolved resonant excitation collision-induced dissociation (CID) and infrared multiphoton dissociation (IRMPD) experiments. The x 2 intermediates undergo facile loss of HNCO to form z 2 fragment ions, as also confirmed by energy-resolved CID and IRMPD MS4 experiments. The loss of HNCO from the x 2 ion from AAHWR is kinetically hampered by the Trp residue that traps the OCNH radical group in a cyclic intermediate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Dissociations of fragment ions from electron transfer dissociation (ETD) of peptide cations continue to attract considerable attention because of their potential for providing peptide primary sequence information [1]. In the preceding report [2] we analyzed resonant excitation collision-induced dissociation mass spectra of z-type ions produced by ETD of a set of over 40 penta-, nona-, and decapeptides of the tryptic type that were C-terminated with arginine and contained different amino acid residues in the peptide chain. Upon collisional activation, the z ions underwent radical-induced dissociations that depended on the nature of the side chains in the amino acid residues. In particular, z n ions with amino acid residues that contained benzylic (His, Phe, Tyr, Trp) or tertiary (Val) hydrogens in the Cβ positions showed enhanced backbone dissociations in positions shifted two or three amino acid residues toward the C-terminus, forming abundant z n-2 or z n-3 ions in the MS3 mass spectra [2]. This type of dissociation was thought to be analogous to cascade dissociations of peptide ions upon electron capture [3] that can be driven by the substantial energy associated with electron-ion recombination. In keeping with previous studies [4–6], we adopted the proposed mechanism of backbone dissociations, involving radical-induced hydrogen migration from the side chain forming intermediate radical Ι (Scheme 1). According to Scheme 1, a transfer of a Cβ hydrogen atom activates the adjacent Cα–CO bond in I for dissociation forming an x intermediate which eliminates HNCO forming the truncated z n-2 or z n-3 fragment ion.
Although this multistep mechanism is chemically plausible, it raises several questions not addressed by previous survey-like studies. First, the Scheme 1 mechanism involves a hydrogen atom migration forming intermediate I, raising the question of the pertinent transition state (TS) energies and relative stabilities of the reactants and the I intermediate. A second question concerns the relative stability of the x ions and the TS for the HNCO loss. Finally, to assess the overall kinetics of these cascade dissociations, it is necessary to compare the reaction flux through the transition states and obtain the pertinent rate constants that allow one to pinpoint the rate determining step.
In this Part 2 report, we address these questions by experiment and theory. z-Type ions from AAHWR were studied using energy-resolved collision-induced and infrared multiphoton dissociation to assess the energetics and kinetics of the consecutive dissociations. Electronic structure theory calculations using two different density functionals and a perturbational Møller-Plesset theory were used to locate transition states and obtain activation energies for the isomerizations and dissociations of z 4 ions from AAHAR, AAHWR, and AAYAR. The energy data were used for Rice-Ramsperger-Kassel-Marcus (RRKM) calculations of unimolecular rate constants to assess the branching ratios for the dissociations.
2 Experimental
2.1 Materials and Methods
The peptides were custom-synthesized by NEOPeptide Laboratories (Cambridge, MA, USA) or CHI Scientific (Maynard, MA, USA) and used as received. ETD mass spectra were measured on a dual-cell quadrupole linear ion trap (QLT) outfitted with a chemical ionization source for the generation of fluoranthene anion radicals for use as an ETD reagent (LTQ XL, Thermo Fisher Scientific, San Jose, CA, USA) [7, 8]. Selected measurements were carried out using a hybrid mass spectrometer comprising both ion trap (QLT) and Orbitrap mass analyzers and likewise equipped with a chemical ionization source for the generation of fluoranthene radicals for use as an ETD reagent (LTQ Orbitrap Velos; Thermo Fisher Scientific). Peptide solutions (10 μM) in 50/50/1 water/methanol/acetic acid were electrosprayed to form precursor cations for ETD. Electrospray ionization of the pentapeptides produced mainly doubly protonated (M + 2H)2+ ions and minor singly protonated (M + H)+ ions, typically in a ≥ 10:1 ratio.
IRMPD and energy-resolved CID experiments were carried out on a modified dual-cell QLT outfitted with a CI source for the generation of fluoranthene anion radicals, as described previously [8]. The mass spectrometer was modified to allow for the introduction of IR photons to the ion trapping region of the QLT. We inserted a ZnSe window on the rear flange of the instrument to transmit IR photons through a stainless steel aperture (diameter = 0.065 in.) concentric with the trapping volume of the QLT. To allow the passage of IR photons to the dual cell QLT, the ion volume of the CI source was also modified with the addition of a hole (diameter = 0.070 in.) in the section of the ion volume farthest from the ion trap and the enlargement of the hole in the section of the ion volume closest to the QLT, both also concentric with the QLT trapping volume. IR photons were generated using a Firestar T-100 Synrad 120-W CO2 continuous wave laser (Mukilteo, WA, USA). Doubly protonated peptide precursor ions were produced by electrospray and subjected to ETD (~70 ms reaction with 5 × 105 fluoranthene radical anions) to form z 4 ions. The z 4 ions were re-isolated and activated by using resonant excitation (CID) within the high-pressure QLT (bath gas He). For the energy-resolved experiments, CID normalized collision energies (NCE) of 0 to 50 correspond roughly to supplemental AC excitation voltages of 0 to ~2.5 V. Infrared multiphoton dissociation was carried out by operating the laser in a mode whereby it was externally triggered via a TTL pulse from pin14 of the J1 connector. Laser power was set as indicated in the text.
2.2 Calculations
All calculations were performed using the Gaussian 09 suite of programs [9]. Initial guesses for the z ion structures were taken from the optimized peptide cation-radicals under the assumption that the fragments preserve the local hydrogen bonding patterns in the charge-reduced ions, as suggested by a recent study of z ion structures [10]. Geometries were optimized with density functional theory calculations using the hybrid B3LYP functional [11–13] in the spin-unrestricted format and the 6-31 + G(d,p) basis set. The optimized structures were confirmed as local energy minima or first-order saddle points by harmonic frequency calculations. Two sets of single-point energies were calculated using the larger 6-311++G(2d,p) basis set and B3LYP and M06-2X functionals [14]. The latter functional has been claimed to give improved transition state energies for kinetic calculations. In addition, Møller-Plesset theory [15], UMP2(frozen core), with the 6-311++G(2d,p) basis set was used to obtain single-point energies, which were corrected for contribution of higher spin states by the standard spin annihilation procedure [16, 17]. The UB3LYP and PMP2 single-point energies were averaged (B3-PMP2) to provide improved relative energies for cation-radicals, as reported previously [18–20]. Unimolecular rate constants were calculated for reactions occurring on the calculated B3-PMP2 and M06-2X potential energy surfaces, including zero-point vibrational energy corrections, within the formalism of the RRKM theory [21]. These RRKM calculations used the program of Zhu and Hase [22] that had been modified to deal with larger molecular systems with up to 1000 atoms, and recompiled to run under Windows XP [23] and Windows 7. The RRKM rate constants were obtained by direct count of quantum states at internal energies that were increased in 2 kJ mol–1 steps from the transition state up to 400 kJ mol–1 above the reactant. Rotations were treated adiabatically, and the calculated microscopic rate constants k(E,J,K) were then Boltzmann-averaged over the thermal distribution of rotational states at 298 K.
3 Results
3.1 Consecutive Dissociations of z 4 Ions
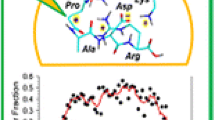
We first present and discuss the features of z ion dissociations as observed in the MS3 mass spectra. The z 4 and z 5 fragment ions produced by ETD of (AAXAR + 2H)2+ (X = F, Y, H, W, and V) (Figure 1a) undergo collision-induced dissociations producing dominant z 2 ions that amounted to 15 %–65 % of the total fragment intensity in the MS3 mass spectra (Figure 1b) [2]. A peculiar feature of these dissociations, when starting from z 4 ions, was the presence of minor x 2 peaks (at 8 %–10 % relative to the z 2 ion intensity) that were broadened, and their m/z values were reduced by ≥0.2 m/z units from the expected m/z 273.1 nominal value. This phenomenon was observed for MS3 dissociations of all these z 4 ions, as illustrated for AAHAR (Figure 1b), AAYAR, AAFAR, and AAVAR (Figure S1, Supplementary Data) and reported previously for AAWAR [2]. We speculated that the x 2 ion was metastable and dissociated during ejection from the linear ion trap, so that its detection was slightly offset during the scan, resulting in a lower apparent m/z value [24, 25]. This was consistent with the CID mass spectra obtained at longer detection times on an Orbitrap instrument, which showed the x 2 peak at the correct m/z 273.1427 but at a substantially reduced relative intensity (0.9 % relative to the z 2 ion at m/z 230.1370, Figure S2, Supplementary Data). The difference in the exact masses, Δm = 43.0057 Da, fits the theoretical mass of HNCO (43.0058 Da).
In contrast to CID of z 4 ions from ETD of the F, Y, H, W, and V-containing AAXAR peptides, the z 4 ions from AAHWR (m/z 553) showed an MS3 spectrum that contained both the x 2 and z 2 ions at correct m/z values (388.1844 and 345.1786, respectively) and with substantial relative intensities (Figure S3, Supplementary Data). This enabled the re-isolation of the x 2 ion at m/z 388 and investigation of its dissociations in MS4 experiments and also to study the consecutive dissociations of the z 4 ions as a function of the excitation energy.
The energy-resolved MS3 mass spectra of the •AHWR z 4 ion were measured at NCE ranging from 0 to 50. The energy-dependent relative intensities of the z 4 , x 2 , and z 2 ions are shown in Figure 2. The graph shows different onsets for the formation of the x 2 ion at NCE = 11 and for the z 2 ion at NCE ≈ 14. At NCE = 15, the dissociation mainly produces the x 2 ions at an intensity ratio of [x 2 ]/[z 2 ] = 8.8. This ratio drops to 1.7 at NCE = 30 and further decreases at higher collision energies. These results are consistent with the consecutive nature of the z 2 ion formation, whereby the internal energy of the x 2 intermediate increases at higher excitations of the z 4 parent ion to drive the loss of HNCO.
The results were even more dramatic for IRMPD which nonselectively deposits internal energy to all absorbing ions in the ion trap rather than just the mass-selected z 4 precursor ion. The relative intensities of the z 4 , x 2 , and z 2 ions were monitored at three laser power levels (36, 48, and 60 W) and at variable irradiation times ranging from 0 to 50 ms. Figure 3a–c illustrates the time-dependent relative intensities at the 36, 48, and 60 W power settings. The x 2 ion shows an early onset which is followed by that of the z 2 ion. The ion intensities reached maxima at different irradiation times, which were ca. 15 ms for the x 2 ion and ca. 25 ms for the z 2 ion at 48 W (Figure 3b). Control experiments were performed to assess the photo-dissociative depletion of the x 2 and z 2 ions. In this measurement, the ions were independently generated by MS3-CID and irradiated with the CO2 laser at 60 W. Figure S4 (Supplemental Data) shows very similar depletion curves, which indicate that the measured relative intensities are not strongly affected by secondary dissociations. This result is consistent with the calculated IR spectra of the x 2 and z 2 ions, which show only weak absorption bands in the region of CO2 laser wavelengths (900–1100 cm–1, Figure S5).
The CID and IRMPD MS4 mass spectra of mass-selected x 2 ions at m/z 388 showed a dominant dissociation by loss of HNCO forming the z 2 ion at m/z 345 (Figure S6 and S7). The other fragments in the CID and IRMPD spectra in Figure S6 and S7, appearing at m/z 301, 300, 257, 230, 200, and 158, were mainly produced by consecutive dissociations of the z 2 ion. This was corroborated by the IRMPD spectrum of mass-selected m/z 345 ions, which gave the same fragments as those in the MS3 spectra of the x 2 ions.
3.2 Mechanistic Considerations for the Consecutive z 4 → z 2 Dissociations
The experimental data showed different dissociations of z 4 ions from the AAXAR (X = H, F, Y, W, V) and AAHWR peptides in that the former proceeded rapidly to z 2 fragments, whereas the latter formed an abundant x 2 intermediate, which needed further activation to eliminate HNCO. To rationalize these observations, we performed detailed calculations exploring the potential energy surfaces along several reaction paths in z 4 ions from AAHAR and AAYAR and the x 2 ions from AAHWR. The z 4 ion from AAHAR is used here as a case in point. The relative and dissociation energies were obtained at several levels of theory including two density functionals and a perturbational Møller-Plesset (MP2, frozen core) treatment. The energies calculated at all levels of theory are shown in Table 1. The data used in text refer to the averaged B3LYP and spin-projected MP2 (B3-PMP2) values, which provided relative and transition state energies at the chemical accuracy limit (±4 kJ mol–1) for a number of radical systems [19, 20]. The B3-PMP2 and M06-2X relative energies are also plotted in the comprehensive potential energy diagram, collating several pathways we investigated (Figure 4). The energy data and ion structures for the AAYAR system are given in Table S1 and Scheme S1 of the Supplemental Data.
Potential energy surface (kJ mol–1) for isomerizations and dissociations of z 4 ions from AAHAR. The energies are from B3-PMP2/6-311++G(2d,p) single-point calculations and include zero-point corrections. The energies in parentheses are from single-point M06-2X/6-311++G(2d,p) calculations including zero-point corrections. The color-coding is as follows: black lines: cis pathway; blue lines: trans pathway; pink lines: enol pathway
3.3 Pathways to Cα–CO Bond Dissociation in AAHAR
The trans-Pathway
To form an x 2 intermediate, the z 4 ion from AAHAR must break the Cα–CO bond at the His residue. Activation of this bond can involve the formation of a Cβ radical at the His residue by β-hydrogen atom transfer to the original Cα radical site in the z 4 reactant as postulated previously [4, 5] and analyzed in detail for a z 4 ion from AHDAL [6]. Inspection of the z 4 ion structure shows that the His β-hydrogen atom transfer is sterically impossible in the amide all-trans conformer (1), which must first undergo a trans → cis rotation of the terminal Ala amide group (Scheme 2) [6]. The relevant transition state for the rotation (TS1) was calculated to be 36 kJ mol–1 above 1 and led to cis-amide isomer 2. Note that the TS1 energy is substantially lower than typical activation energies for amide group rotations in peptides (80–100 kJ mol–1) [26–28]. This lowering of the amide rotational barrier in z ions is caused by the conjugation of the Cα radical with the adjacent amide group that results in the weakening of the CO–NH double-bond character, as discussed previously [6]. The cis isomer 2 is comparably stable as 1 (Table 1). The His Hβ migration in 2 requires 69 kJ mol–1 in TS2, forming a His Cβ radical 3. Radical 3 is more stable than 1 and 2 at all levels of theory. It can gain further stabilization by cis → trans rotation of the terminal Ala amide group, forming a trans isomer (4) at –38 kJ mol–1 relative to 1. The amide rotation in 3 forming 4 proceeds through TS3 which is 91 kJ mol–1 above 4. Note that the CO–NH bond of the rotating amide group in 3 is not weakened by the nonconjugated Cβ radical, and therefore the TS3 energy is close to values typical for peptides [26–28].
In contrast, the Cβ radical in 4 weakens the adjacent Cα–CO bond, which can dissociate through TS4 which is 124 kJ mol–1 above 4 and 86 kJ mol–1 above 1 (Scheme 3). Dissociation of the Cα–CO bond forms a complex (5) of the incipient x 2 ion (6) and a truncated neutral Ala-His fragment (7). Complex 5 was slightly less stable than 1. However, since we did not study more conformers of 5, we cannot exclude the existence of other, lower energy structures. The important feature is that the dissociation of 5 to 6 and 7, which requires the incipient fragment separation but no covalent bond cleavage, is substantially endothermic. This makes the thermochemical threshold for these fragments the highest point on the potential energy surface of the above-described trans-pathway.
B3LYP/6-31 + G(d,p) optimized structures of reaction intermediates from AAHAR. The atoms are color-coded as in Scheme 2
The cis-Pathway
We considered several alternatives to the trans-pathway. One obvious alternative is for the cis-β-radical 3 to undergo a Cα–CO bond cleavage to form an ion–molecule complex (8) of the incipient cis-Ala-His neutral fragment and the x 2 ion (6) (Scheme 3). The relevant TS5 (108 kJ mol–1 relative to 3) requires a substantially higher energy than does TS3 for the competing amide rotation in 3 (63 kJ mol–1 relative to 3), see Table 1. Likewise, the cis-complex 8 is 11 kJ mol–1 above the trans-complex 5. However, the dissociation threshold for the formation of the cis-Ala-His neutral fragment (9) and 6 is somewhat lower (114 kJ mol–1 relative to 1) than for the trans fragment 7 (128 kJ mol–1 relative to 1). Thus, if the fragment separation in complexes 5 and 8 is the rate determining step for the formation of the x 2 ion 6, the cis-pathway is kinetically preferred.
The cis-pathway, which was also studied for the isomerizations and dissociations of the z 4 ion from AAYAR (y1, Scheme S1), showed some noteworthy similarities and differences from the z 4 ion from AAHAR. The migration of the tyrosine benzylic hydrogen ion in the cis-z 4 ion (y2) to form the benzylic radical y3 required 82 kJ mol–1 in TS1y (Table S1), which was only slightly higher than the TS2 energy (Table 1). However, the relative energy of the benzylic radical y3 was higher than that of the histidine-Cβ radical 3. The Cα–CO bond cleavage in y3 can proceed through TS2y, which is at 111 kJ mol–1 relative to y1 and, thus, higher than the analogous TS5 from AAHAR. The largest difference between the two systems was in the relative energies of the complexes, y4 and 8, where the former was substantially less stable than y1. This can be attributed to the different type of noncovalent bonding in these complexes. Complexes 5 and 8 gain stabilization by hydrogen bonding between the histidine imidazole ring and the amide hydrogen of the dangling carbamoyl group (Scheme 3). No such interaction occurs between the tyrosine phenol group and the incipient x 2 fragment in complex y4 (Scheme S1).
The Enol Pathway
Another alternative pathway we addressed differed from the previous two in that the first step, the His β-H transfer, proceeded to the oxygen of the terminal amide group rather than to the Cα position (Scheme S2). The amide group in 1 is conjugated with the Cα radical and displays non-negligible spin density at the oxygen atom (11 %), which presumably can make it an H-atom receptor in a radical driven migration. Note that the His β-H transfer to the neighboring amide oxygen can proceed in 1 without a trans → cis amide rotation. However, the TS6 energy (127 kJ mol–1) is higher than that of TS2, and the enol intermediate 10 is substantially less stable than 1. This indicates that the enol pathway would have a reversible first step favoring a reverse 10 → 1 hydrogen migration, which would adversely affect the kinetics of the overall dissociation. Perhaps the most serious factor disfavoring the enol pathway is the high threshold energy for the formation of an enol form of the Ala-His neutral fragment (11) at 217 kJ mol-1 relative to 1 (Table 1).
A common feature of the above-mentioned pathways is the formation of the x 2 intermediate (6), which is experimentally observed as a low-intensity metastable m/z 273 fragment ion. Its further dissociation by loss of O = C = NH is calculated to be slightly exothermic (Table 1), but requires additional 70 kJ mol–1 in TS7 and involves a stable ion-molecule complex (12, Scheme 3).
The Cyclization Cascade Pathway
The previous study of cascade dissociations in ECD by O’Connor and coworkers [3] proposed radical-driven cyclization reactions to create new radical centers in the peptide fragments that would activate adjacent bonds for further dissociations. We analyzed similar mechanisms for the dissociation of the z 4 ion from AAHAR, as shown in Scheme S3. A cyclization reaction between the Ala Cα radical and the His amide group must involve amide trans → cis rotation through TS1, which is a low energy process (Scheme 2). The cis–amide 2 can attack the His amide oxygen atom in TS8 forming a cyclic aminoketyl radical (13),which is substantially less stable (+97 kJ mol–1) than 2 (Table 1). The cyclic radical 13 can undergo competitive ring opening back to 2 or proceed by N–Cα bond cleavage through TS9, yielding directly the z 2 fragment ion and the cyclic neutral fragment 14. We also investigated the possibility of a Cα radical attack at the amide nitrogen atom followed by N–Cα bond cleavage (Scheme S3). However, the pertinent TS10 for the radical attack at the amide nitrogen required a very high energy (Table 1).
Dissociation Kinetics
The efficiency of the cascade dissociation pathways depends on the forward and reverse reaction flux through the pertinent transition states as quantified by the respective rate constants. The rate constants were calculated using RRKM for the B3-PMP2 TS energies and used to express the molar fractions of z 2 fragment ions produced on the time scale of the experiment which was chosen as 300 ms. The calculated rate constants pertinent to the formation of the x 2 intermediate in the cis pathway (log k, s–1) are shown in Figure 5a. As expected, the amide cis → trans rotations are fast (k 1, k -1), leading to rapid equilibration of 1 and 2. The rate constants for the reversible Hβ migration through TS2 (k 2, k -2) favor the formation of the Cβ radical 3. The rate determining steps are the Cα–CO bond cleavage through TS5 (k 3) and the subsequent separation of the incipient products in complex 8, forming the x 2 fragment ion. The consecutive loss of HNCO from the x 2 ion is fast and shows a rapid rise of the pertinent rate constant (k 4) with the x 2 internal energy.
RRKM kinetics of z 4 and x 2 ion isomerizations and dissociations for the AAHAR peptide. (a) RRKM rate constants (log k, s–1) for reactions: k 1: 1 → TS1; k -1: 2 → TS1; k 2: 2 → TS2; k -2: 3 → TS2; k 3: 3 → TS5; k 4: 6 → TS7. (b) RRKM rate constants for reactions: k 5: 2 → TS8; k -5: 13 → TS8; k 6: 13 → TS9. (c) Energy-dependent molar fractions of x 2 intermediate (blue curve) and z 2 fragment ions (red curve) at 300 ms reaction time. The purple curve shows the molar fraction of z 2 ions formed at 300 ms by the cyclization cascade pathway
The cyclization pathway shows different kinetics (Figure 5b). The rate determining step (k 5) is the cyclization through TS8 forming intermediate 13. The latter can undergo competitive Cα–CO bond cleavage through TS9 to directly produce the z 2 fragment (k 6) or undergo reverse ring opening (k -5). The k -5 and k 6 rate constants indicate that the reaction branching in 13 should favor the ring opening to cis-z 4 , not fragmentation to z 2 .
The overall reaction flux on the 300 ms time scale is illustrated by the calculated molar fractions of 6, and the z 2 fragments produced by the Scheme 2 (cis) and Scheme S3 (cyclization) pathways (Figure 5c). Note that while the formation of 8 was treated rigorously by solving the complex kinetics of reversible, competitive, and consecutive reactions, the rate constant for the subsequent fragment separation, 8 → 6 + 9, was not calculated because this dissociation is likely to be continuously endothermic without a well-defined saddle point on the potential energy surface. We estimated the pertinent rate constant as being equal to k 3 (Figure 5a) because both the 3 → 8 and 8 → 6 + 9 reactions are simple dissociations requiring similar energies (Table 1). A reverse fragment recombination in the complex, 8 → 3, was considered slow because of entropy loss in the transition state. Under these assumptions, the overall formation of the x 2 intermediate showed a substantial kinetic shift (Figure 5c, blue curve), reaching 50 % conversion to 6 at 223 kJ mol–1 internal energy, which was 109 kJ mol–1 above the thermochemical threshold. The dissociation was nearly complete (>99 %) at >242 kJ mol–1 internal energy. We presumed that fragments 6 and 9 formed from 8 equipartitioned the available non-fixed internal energy according to the number of their vibrational degrees of freedom (108/177 or 61 % for 6). For example, 1 having 223 kJ mol–1 of internal energy and dissociating at 50 % efficiency is estimated to form the intermediate 6 with (223 – 114) × 0.61 = 66 kJ mol–1 internal energy and likewise for dissociations of 1 having higher internal energies. Vibrational excitation in 6 exceeding 70 kJ mol–1, which was required to reach TS7, was calculated to lead to a rapid dissociation by loss of HNCO which showed a steep increase of the rate constant with internal energy (k 4, Figure 5a). Hence, intermediate 6 formed by dissociation of 1 having >260 kJ mol–1 internal energy was calculated to undergo nearly 100 % loss of HNCO on the 300 ms time scale forming the z 2 fragment ion (Figure 5c, red curve). The narrow interval of kinetic metastability for 6 (ca. 220–261 kJ mol–1) was consistent with the low relative intensity of the x 2 (m/z 273) ions in the MS3 mass spectrum. In other words, most z 4 ions that acquired excessive internal energy to dissociate on the time-scale of the experiment produced x 2 intermediates with internal energies that were sufficient to drive their fast subsequent dissociation by loss of HNCO.
The formation of the z 2 fragment ion through the cyclization pathway displays a very large kinetic shift, reaching 50 % conversion at internal energies ≥400 kJ mol–1 (Figure 5c purple curve). This indicates that the cyclization pathway may not be kinetically competitive. We note that although the calculated rate constants strongly depend on the quality of the potential energy surface, the conclusion regarding the preference of the cis over the cyclization pathway also makes chemical sense. Schemes 1 and 2 show that the dissociation is facilitated by the Cβ–H transfer, which explains the activating role of the His, Phe, Tyr, Trp, and Val residues. In contrast, the role of the His side-chain group in the Scheme S3 mechanism appears to be one of a spectator with no obvious involvement in the rate-determining ring cyclization.
The Tryptophan Effect on Dissociations of z 4 Ions from AAHWR
Dissociation of this z 4 ion produced an unusually stable x 2 intermediate at m/z 388 (Figure S3). We reasoned that this effect could be due to the radical-trapping properties of the indole ring [29–31] and investigated the potential energy surface in the pertinent x 2 cation-radical (w1). The optimized structures are shown in Scheme 4, the relative energies are given in Table 2. The dissociation by loss of HNCO from w1 was found to have a TSw1 energy (75 kJ mol–1, Table 2) that was nearly identical to that for the HNCO loss from the x 2 ion from AAHAR (TS7, Table 1). Thus, the effect of the Trp residue on TSw1 was weak. However, the optimized structure of w1 showed that the O = C-NH radical group can reach to the C-2', C-3' and C-10' positions of the indole ring for radical-driven cyclizations. The TS energy for cyclization to C-2' was calculated as 54 kJ mol–1 (TSw2) to give the most stable radical w2 which was –24 kJ mol–1 relative to w1 (Table 2, Scheme 4). Cyclizations at C-3' and C-10' analogously gave radicals w3 and w4, respectively, which however, were comparably stable as w1 (Table 2). This order of tryptophan radical stabilities is quite analogous to those reported for H-atom adducts which also preferred a C-2' adduct as the lowest energy isomer [31].
The existence of the low-energy isomer w2 and TSw2 for cyclization has a pronounced effect on the dissociation kinetics of the x 2 ion. RRKM calculations on the B3-PMP2 and M06-2X potential energy surfaces (Figure 6a and c, respectively) show that the cyclization in w1 (k 7) forming w2 was faster than both the loss of HNCO (k 8) and the ring opening in w2 (k -7) to re-form w1. Solving the kinetics for the competitive and consecutive reactions gave the breakdown diagrams for m/z 388 ions which are plotted for combined molar fractions of w1 and w2 as a function of the internal energy of w1 at reaction times ranging from 50 μs to 300 ms. (Figure 6b and d). The breakdown diagrams show substantial kinetic shifts for the depletion of w1 and w2 by loss of HNCO even at the 300 ms reaction time. The B3-PMP2 data (Figure 6b) indicate a kinetic shift of ΔE kin = 85 kJ mol–1 for 50 % depletion of the m/z 388 ion (w1 + w2) after 300 ms. The M06-2X data (Figure 6d), which gave higher TS energies for the HNCO loss and ring opening in w2 (Table 2), indicate a 150 kJ mol–1 kinetic shift under the same conditions. This is to be compared with the HNCO loss from 6 which shows 50 % dissociation over 300 ms at 86 kJ mol–1 internal energy, giving a much smaller kinetic shift of 16 kJ mol–1.
RRKM kinetics of x 2 ion isomerizations and dissociations for the AAHWR peptide. (a) and (c) Rate constants for reactions: k 7: w1 → TSw2; k -7: w2 → TSw2; k 8: w1 → TSw1 on the (a) B3-PMP2/6-311++G(2d,p) and (c) M06-2X/6-311++G(2d,p) potential energy surfaces. (b) (B3-PMP2) and (d) (M06-2X): calculated breakdown curves for m/z 388 (w1 + w2) ions at 50 μs, 1, 10, 100, and 300 ms reaction times
In summary, the reason for the lack of dissociation of w1 is the competing radical cyclization to isomer w2. Ions w1 and w2 appear at the same m/z 388 and, therefore, are not distinguished in the mass spectrum. The calculated energy-dependent ratio of w1/w2 molar fractions shows that the majority of the non-dissociating x 2 ions should be the w2 isomer (Figure S8, Supplementary Data).
4 Conclusions
Experimental and computational analysis of cascade dissociations of z 4 ions from pentapeptides AAHAR, AAYAR, and AAHWR indicate that these proceed with involvement of β-hydrogen atoms in the benzylic positions of the side chains. Cyclization mechanisms, analogous to those postulated previously, are not compatible with the dissociation kinetics for z 4 ions. Intermediates of the x 2 type were detected by IRMPD and energy-resolved CID experiments in complete agreement with the kinetic analysis. The rate determining step in the dissociation is the Cα–CO bond dissociation at the activating residue (H, Y, F, V, W) in the z 4 ion. Loss of HNCO from the x 2 intermediate is fast, leaving a narrow energy window of metastability for the x 2 ions. The loss of HNCO is hampered by the Trp residue that functions as a kinetic trap for the OCNH radical group.
References
Li, W., Song, C., Bailey, D.J., Tseng, G.C., Coon, J.J., Wysocki, V.H.: Statistical analysis of electron transfer dissociation pairwise fragmentation patterns. Anal. Chem. 83, 9540–9545 (2011)
Chung, T.W., Renjie Hui, R., Ledvina, A.R., Coon, J.J., Turecek, F.: Cascade dissociations of peptide cation radicals. Part 1. Scope and effects of amino acid residues in penta-, nona-, and decapeptides. J. Am. Soc. Mass Spectrom., preceding article in this issue (2012)
Leymarie, N., Costello, C.E., O’Connor, P.B.: Electron capture dissociation initiates a free radical reaction cascade. J. Am. Chem. Soc. 125, 8949–8958 (2003)
Liu, J., Liang, X., McLuckey, S.A.: On the value of knowing a z• ion for what it is. J. Proteome Res. 7, 130–137 (2008)
Sun, Q., Nelson, H., Stolz, B.M., Julian, R.R.: Side chain chemistry mediates backbone fragmentation in hydrogen deficient peptide radicals. J. Proteome Res. 8, 958–966 (2009)
Chung, T.W., Tureček, F.: Backbone and side chain-specific dissociations of z ions from nontryptic peptides. J. Am. Soc. Mass Spectrom. 21, 1279–1295 (2010)
Pekar, S.T., Blethrow, J.D., Schwartz, J.C., Merrihew, G.E., MacCoss, M.J., Swaney, D.L., Russell, J.D., Coon, J.J., Zabrouskov, V.: Dual-pressure linear ion trap mass spectrometer improving the analysis of complex protein mixtures. Anal. Chem. 81, 7757–7765 (2009)
Ledvina, A.R., Beauchene, N.A., McAlister, G.C., Syka, J.E.P., Schwartz, J.C., Griep-Raming, J., Westphall, M.S., Coon, J.J.: Activated ion electron transfer dissociation improves the ability of electron transfer dissociation to identify peptides in a complex mixture. Anal. Chem. 82, 10068–10074 (2010)
Kitao, O., Nakai, H., Vreven, T., Montgomery Jr., J.A., Peralta, J.E., Ogliaro, F., Bearpark, M., Heyd, J.J., Brothers, E., Kudin, K.N., Staroverov, V.N., Kobayashi, R., Normand, J., Raghavachari, K., Rendell, A., Burant, J.C., Iyengar, S.S., Tomasi, J., Cossi, M., Rega, N., Millam, J.M., Klene, M., Knox, J.E., Cross, J.B., Bakken, V., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R.E., Yazyev, O., Austin, A.J., Cammi, R., Pomelli, C., Ochterski, J.W., Martin, R.L., Morokuma, K., Zakrzewski, V.G., Voth, G.A., Salvador, P., Dannenberg, J.J., Dapprich, S., Daniels, A.D., Farkas, O., Foresman, J.B., Ortiz, J.V., Cioslowski, J., Fox, D.J.: Gaussian 09, Revision A.02. Gaussian, Inc., Wallingford (2009)
Moss, C.L., Chamot-Rooke, J., Brown, J., Campuzano, I., Richardson, K., Williams, J., Bush, M., Bythell, B., Paizs, B., Tureček, F.: Assigning structures to gas-phase peptide cations and cation-radicals. An infrared multiphoton dissociation, ion mobility, electron transfer and computational study of a histidine peptide ion. J. Phys. Chem. B 106, 3445–3456 (2012)
Becke, A.D.: New mixing of Hartree-Fock and local density-functional theories. J. Chem. Phys. 98, 1372–1377 (1993)
Becke, A.D.: Density functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993)
Stephens, P.J., Devlin, F.J., Chabalowski, C.F., Frisch, M.J.: Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 98, 11623–11627 (1994)
Zhao, Y., Truhlar, D.G.: The M06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four M06-class functionals and 12 other functionals. Theor. Chem. Acc. 120, 215–241 (2008)
Møller, C., Plesset, M.S.: A note on an approximation treatment for many-electron systems. Phys. Rev. 46, 618–622 (1934)
Schlegel, H.B.: Potential energy curves using unrestricted Moller-Plesset perturbation theory with spin annihilation. J. Chem. Phys. 84, 4530 (1986)
Mayer, I.: Spin-projected UHF method. IV. Comparison of potential curves given by different one-electron methods. Adv. Quantum Chem. 12, 189 (1980)
Tureček, F.: Proton affinity of dimethyl sulfoxide and relative stabilities of C2H6OS molecules and C2H7OS+ Ions. A comparative G2(MP2) ab initio and density functional theory study. J. Phys. Chem. A 102, 4703 (1998)
Polášek, M., Tureček, F.: Hydrogen atom adducts to nitrobenzene. Formation of the phenylnitronic radical in the gas phase and energetics of Wheland intermediates. J. Am. Chem. Soc. 122, 9511–9524 (2000)
Wolken, J.K., Yao, C., Tureček, F., Polce, M.J., Wesdemiotis, C.: Cytosine neutral molecules and cation-radicals in the gas-phase. Structures, energetics, ion chemistry, and neutralization-reionization mass spectrometry. Int. J. Mass Spectrom 267, 30–42 (2007)
Gilbert, R.G., Smith, S.C.: Theory of Unimolecular and Recombination Reactions, pp. 52–132. Blackwell Scientific Publications, Oxford (1990)
Zhu, L., Hase, W.L.: Quantum Chemistry Program Exchange. Indiana University: Bloomington; Program No. QCPE 644 (1994)
Frank, A.J., Sadílek, M., Ferrier, J.G., Tureček, F.: Sulfur oxoacids and radicals in the gas phase. A variable-time neutralization-photoexcitation-reionization mass spectrometric and ab initio/RRKM study. J. Am. Chem. Soc. 119, 12343 (1997)
McClellan, J.E., Murphy, J.P.I.I.I., Mulholland, J.J., Yost, R.A.: Effects of fragile ions on mass resolution and on isolation for tandem mass spectrometry in the quadrupole ion trap mass spectrometer. Anal. Chem. 74, 402–412 (2002)
Swaney, D.L., McAlister, G.C., Wirtala, M., Schwartz, J.C., Syka, J.E.P., Coon, J.J.: Supplemental activation method for high-efficiency electron-transfer dissociation of doubly protonated peptide precursors. Anal. Chem. 79, 477–485 (2007)
Jhon, J.S., Kang, Y.K.: Imide cis-trans isomerization of N-acetyl-N'-methylprolineamide and solvent effects. J. Phys. Chem. A 103, 5436–5439 (1999)
Shi, T., Spain, S.M., Rabenstein, D.L.: Unexpectedly fast cis-trans isomerization of Xaa-Pro peptide bonds in disulfide-constrained cyclic peptides. J. Am. Chem. Soc. 126, 790–796 (2004)
Fischer, G.: Chemical aspects of peptide bond isomerization. Chem. Soc. Rev. 29, 119–127 (2000)
Byers, J.H., Campbell, J.E., Knapp, F.H., Thissell, J.G.: Radical aromatic substitution via atom-transfer addition. Tetrahedron Lett. 40, 2677–2680 (1999)
Walden, S.E., Wheeler, R.A.: Distinguishing features of indolyl radical and radical cation: implications for tryptophan radical studies in proteins. J. Phys. Chem. 100, 1530–1535 (1996)
Gregersen, J.A., Tureček, F.: Mass-spectrometric and computational study of tryptophan radicals (Trp + H)• produced by collisional electron transfer to protonated tryptophan in the gas phase. Phys. Chem. Chem. Phys. 12, 13434–13447 (2010)
Acknowledgments
F.T. thanks the National Science Foundation for support (grant CHE-1055132). Instrumentation support by the University of Washington Proteomics Resource (Dr. Priska von Haller) for the ETD mass spectra measurements is gratefully appreciated, along with access to the ETD-equipped Thermo-LTQ Orbitrap Velos by the Institute for Systems Biology (Drs. Richard Johnson and Rob Moritz). The work at the University of Wisconsin was supported by National Institutes of Health (NIH) grant RO1 GM080148.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ledvina, A.R., Chung, T.W., Hui, R. et al. Cascade Dissociations of Peptide Cation-Radicals. Part 2. Infrared Multiphoton Dissociation and Mechanistic Studies of z-Ions from Pentapeptides. J. Am. Soc. Mass Spectrom. 23, 1351–1363 (2012). https://doi.org/10.1007/s13361-012-0409-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-012-0409-8