Abstract
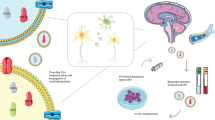
Extracellular vesicles (EVs), once considered a pathway for cells to remove waste, have now emerged as an important mechanism for intercellular communication. EVs are particularly appealing in understanding the central nervous system (CNS) communication, given that there are very diverse cell types in the CNS and constant communications among various cells to respond to the frequently changing environment. While they are heterogeneous and new vesicles are continuously to be discovered, EVs are primarily classified as plasma membrane-derived microvesicles (MVs) and endosome-derived exosomes. Secretion of EVs has been shown from all CNS cell types in vitro and intercellular EV signaling has been implicated in neural development, axon integrity, neuron to glia communication, and propagation of protein aggregates formed by disease pathogenic proteins. However, significant hurdles remain to be tackled in understanding their physiological and pathological roles as well as how they can be developed as biomarkers or new therapeutics. Here we provide our summary on the known cell biology of EVs and discuss opportunities and challenges in understanding EV biology in the CNS and particularly their involvement in ALS pathogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Extracellular vesicles (EV) are small membrane bound vesicles, typically within the size range of 50–1000 nm, released by different cell types under physiological and pathological conditions. Surrounded by a phospholipid bilayer membrane, EVs protect molecules from degradation and enable delivery to both adjacent and distant cells. The existence of EVs in tissue fluids was first discovered in the early 1940s [1] and EV secretion was initially thought to be a way for cells to remove intracellular molecular wastes [2, 3]. However, the discovery in 1996 that EVs were capable of inducing antigen-specific MHC II T Cell responses began to implicate an intercellular communication role for EVs [4]. This notion was further strongly supported in 2006 [5] and 2007 [6] when two studies showed that EVs (especially exosomes) contain mRNAs and microRNAs (miRs) which are able to change recipient cell functions after being transferred into recipient cells. Since then, the role of EVs has been studied extensively in many cell types especially in cancer where it is now thought to play an important role in influencing the tumor microenvironment and regulating metastasis [7, 8]. Intercellular communication in the CNS is highly complex, partially due to the extensive heterogeneity of CNS cell types and its constantly changing responses to the environment. EVs secreted from different CNS cell types have been examined and their involvement in mediating such complex intercellular communications in the CNS has started to be revealed [9]. In addition, EVs have been implicated in neurodegenerative diseases especially in Alzheimer’s disease (AD) and amyotrophic lateral sclerosis (ALS), as a potential propagation mechanism of abnormal protein aggregates [10]. Here, we will discuss current understanding of the roles of EVs in the CNS, especially their involvement in ALS pathogenesis and diagnostics, as well as hurdles that need to be cleared for potential EV-mediated therapeutic opportunities for treating ALS and other neural diseases.
Biogenesis and Secretion of Major EV Types
Although all known EVs share the common phospholipid bilayer membrane structure and the enclosed vesicle morphology, EVs are highly heterogenous in size, origin, cargo composition, and surface markers [11]. These differences result from drastically distinct biogenesis, sorting, and secretion mechanisms that can be very cell-type dependent. As a result, EV heterogeneity affects the delivery of various bioactive molecules (proteins, lipids, and nucleic acids) to recipient cells and their subsequent functions in recipient cells [12]. Depending on their origins, EVs are generally classified as plasma membrane-derived microvesicles (MVs) and endosome-originated exosomes [13] which is primarily discussed here as they are currently better understood. Although other more specialized secreted vesicles, such as exomeres [14], arrestin domain containing protein 1 [ARRDC1]-mediated microvesicles (ARMMs) [15], and supermeres [16]. have recently been identified, their origins and secretion regulations remain to be further characterized and therefore will not be discussed here. It is noted that the use of “EVs” and “exosomes” is often mixed and interchangeable in the literatures, here we keep the terminology consistent with its use in original literatures. In other occasions when it is not so clear or it can be both, we used “EVs/exosomes.”
MVs, also known as ectosomes, microparticles, oncosomes, or shedding vesicles, are typically within the range of 100 to 1000 nm in diameter and were originally described as a secreted product from platelets [17]. While further characterization of the detailed mechanisms for the biogenesis of MVs is needed, it is well-accepted that MVs are formed by the outward budding of the parent cell’s plasma membrane [18]. Specifically, certain lipids and membrane proteins (either associated or inserted) become clustered in particular microdomains of the plasma membrane, which further recruits cytosolic proteins and RNAs to be packaged in MVs for release [18]. Lipid movement and membrane dynamics play critical roles in MV biogenesis [19]. However, it is possible that other protein complexes are involved, such as the endosomal sorting complex required for transport (ESCRT). How exactly cytosolic proteins and RNAs are sorted into MVs remains very little understood, but Rab GTPase (typically Rab11 or Rab 35)-mediated intracellular trafficking of recycling endosomes has been implicated in this sorting process [20]. It has been shown that the small GTPase ADP-ribosylation factor 6 (ARF6) regulates selective recruitment of β1 integrin receptors, MHC class I molecules, and membrane type 1‑matrix metalloproteinase 1 (MT1‑MMP), into tumor cell-derived MVs [21]. The tetraspanin family protein CD9 also plays a role in regulating protein targeting in tumor cell-derived MVs [22, 23]. Because of their plasma membrane origin, MVs mostly contain plasma membrane-derived receptors and lipids [24]. MV membranes contain large amounts of cholesterol, diacylglycerol, and phosphatidylserine. Various membrane proteins such as CD40, integrins, and selectins are also frequently found on the surface of MVs [25].
MVs are released to the extracellular environment through a fission process in which the interaction between actin and myosin plays a key role [26]. As a result, mechanisms regulating the contraction of the actin-myosin (actomyosin) complex are involved in MV release [27]. For example, the activation of small GTP binding proteins leads to the phosphorylation of myosin light chain (MLC) and actomyosin contraction, facilitating the release of MVs from the membrane of cancer cells [28]. Similarly, Rho GTPase activity and Rho-associated protein kinase (ROCK) also promote MV biogenesis in tumor cells by regulating actin dynamics [29]. In addition, intracellular Ca2+ levels regulate MV biogenesis and secretion [18]. Several Ca2+-dependent enzymes including aminophospholipid translocases (flippases and floppases) and scramblases can rearrange the membrane phospholipids, in particular, to expose phosphatidylserine (PS) from the inner leaflet to the outside of the membrane [30]. At the steady state, lipid translocases maintain an asymmetric distribution of phospholipids at the plasma membrane. An increase in intracellular Ca2+ levels disrupts the balance and promotes external exposure of PS, which causes physical bending of the membrane [30] and changes the underlying actin cytoskeleton to facilitate the budding and release of MVs [27]. Although apoptotic bodies have also been considered MVs in certain literature [31], it is important to note that MVs discussed here only refer to plasma membrane-derived vesicles from non-apoptotic cells.
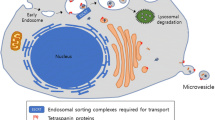
Exosomes are typically the smaller subgroup of EVs, ranging from 50–200 nm in diameter. Exosomes are originated as intraluminal vesicles (ILVs) that are inwardly budded from endosomal multivesicular bodies (MVBs) during the endosome maturation process [11]. Based on the genetic analysis, especially RNA interference (RNAi)-based screening [32], ESCRT-dependent mechanisms underlying exosome biogenesis have been fairly well-established [11]. In particular, ESCRT-III is required for the scission of ILVs into the MVB lumen [33]. Depletion of the ESCRT-0 components HRS or signal transducing adapter molecule 1 (STAM1) in HeLa cells also decreases the secretion of exosomes, as measured by the expression of their surface marker tetraspanin protein CD63 [34]. By interacting with their cellular adaptor proteins Syntenin-1 and ESCRT accessory protein ALG‑2 interacting protein X (ALIX), plasma membrane proteoglycans such as syndecans also regulates the biogenesis of exosomes [35]. However, given the large number of ESCRT components and accessory proteins, it remains to be investigated how these components and accessory proteins are specifically involved in exosome biogenesis of different cell types. While it is clear that ESCRT complexes regulate exosome biogenesis, depletion of ESCRT components only decrease but not completely diminish the secretion of exosomes, indicating that there are ESCRT-independent mechanisms in regulating exosome biogenesis [11]. Several lipid pathways have been identified to regulate exosome biogenesis [36]. Among these, the best-studied and well-demonstrated lipid pathway is neutral type II sphingomyelinase (nSMase)-mediated hydrolysis of sphingomyelin to ceramide [37]. Selective pharmacological and genetic inhibition of this pathway is able to significantly suppress exosome biogenesis in many cell types in vitro and in vivo [37, 38]. Other lipids, including cholesterol, diacylglycerol, and phosphatidic acid etc., have also been implicated in regulating exosome (or MV) formation [36].
As exosomes originate from endosomal MVB structures, how intracellular proteins and RNAs are sorted into ILVs for release becomes an important cellular process for exosome-mediated intercellular communication. Similar to MV biogenesis, membrane microdomains that contain membrane protein cargoes are formed with the coordination of ESCRT-0 and -I and also recruit cytosolic proteins through the binding of adaptors such as syntenin [18, 39]. Cytosolic proteins can also be sorted into ILVs by binding to chaperones heat shock 70 kDa protein (HSP70) and heat shock cognate 71 kDa protein (HSC70) [40, 41], which are often found in exosomes. By interacting with membrane lipid rafts, glycosylphosphatidylinositol (GPI)-anchored proteins are also found in ILVs and secreted exosomes [42]. In addition, ubiquitylation or farnesylation have been proposed to modify cytosolic proteins for sorting into ILVs [43, 44] but remains to be further investigated. As tetraspanin family proteins CD63 (and CD81) are highly enriched on the surface of ILVs, they also help the formation of membrane microdomains and facilitate protein targeting into ILVs [18]. In parallel to protein sorting into ILVs, another important class of cargo, nucleic acids especially miRs are also selectively packaged into ILVs/exosomes [6]. While different mechanisms have been proposed, for example, ESCRT‑II may act as an RNA-binding complex [45], membrane microdomains may sequester RNA-binding proteins such as the RNA-induced silencing complex (RISC) and argonaute 2 (AGO2) [46], future studies are needed to elucidate the specificity of such mechanisms. On the other hand, specific motifs on miRs have been identified to bind with RNA-binding proteins such as hnRNPA2B1 [47] or YBX1 [48]. A recent new study further defined additional sequence motifs that determine whether miRs are secreted in exosomes or retained inside cells [49].
Unlike the direct outward budding of MVs from the plasma membrane, exosome release from MVBs is a sophisticatedly regulated process that involves the secretory/degradatory balance of MVBs, intracellular MVB transport, and SNARE-dependent fusion with the plasma membrane [13]. As the formation of ILVs occurs inside MVBs, the intermediate endosomal structure destined to fuse with lysosomes for degradation, factors that regulate MVB/lysosome fusion significantly affect exosome secretion. Although largely unclear, a higher pH level inside MVBs, resulting from abolished V1-ATPase proton transporter activity [50], appears to promote MVBs towards the secretory path. Similarly, MVB fusion with the autophagosome has also been thought to promote its degradation, which prevents exosome secretion [51]. To secrete exosomes, MVBs need to be transported close to the plasma membrane, which is primarily regulated by different members of the Rab GTPase family, such as Rab27a/b, Rab11, and Rab35, depending on cell types [52,53,54]. MVBs subsequently dock on the plasma membrane and fuse with the plasma membrane to release exosomes in a SNARE-dependent manner [55]. Overall, both MV and exosome biogenesis are closely associated with ESCRT complexes, which are also involved in multiple intracellular vesicle trafficking pathways, such as recycling endosomes formation/transport, ER-Golgi secretory vesicles [18]. As a result, the experimental manipulation of these pathways may not selectively affect MV/exosome biogenesis, and studies for searching selective MV or exosome biogenesis/secretion regulators are needed.
Isolation and Characterization of EVs
Detection and characterization of EVs and analysis of their cargoes are essential in better understanding the role of EV-mediated intercellular communication from experimental samples such as culture medium or tissue homogenates. EVs isolated from clinical samples, including blood plasma (or serum), saliva, urine, and cerebrospinal fluid (CSF), especially under pathological conditions, may also serve as a new class of biomarkers for diagnosis, screening, and prognosis purposes [56]. As described above, EV composition can be highly heterogeneous based on size, density, and surface markers [13, 34]. Especially, EVs from biofluids have a mixed cellular origin [56] and are often associated with cell debris, lipoproteins, and other secreted smaller vesicles such as exomeres, making the selective, consistent, and high-yield isolation of EVs a challenging task. Currently, there are no standardized methods for EV isolation. Here, we will focus only on common EV isolation approaches that can be easily set up in the lab (as summarized in Fig. 1), based on physical properties of EVs and the experimental purpose. The pros and cons for each method are also discussed. Additional isolation approaches such as Exochip, microfluidic filtering, and others are being tested [57] and thus not included here.
Ultracentrifugation (Including Density Gradient Centrifugation)
Serial centrifugation steps from 2,000 g to 100,000 g have been commonly used in many early EV studies and are still considered the most commonly used isolation method at present, based on a survey investigating current practices for isolation, purification, and characterization of EVs [58]. This method is based on the typical 100–1000 nm size and the 1.08–1.22 g/mL density range of EVs [13]. The serial steps with increased centrifugation speed allow removal of cell debris and other larger organelles or protein complexes. The centrifugation method is often simple and relatively fast. However, increasing evidence has begun to show secreted extracellular matrix and lipoproteins can be co-pelleted together with EVs during the ultracentrifugation [59, 60]. To better separate potential extracellular protein contaminates from EVs, density gradient ultracentrifugation has been employed to specifically isolate EVs from a particular density layer, which is particularly useful for samples with a complex composition, such as tissue homogenates. However, the force of high-speed centrifugation force and prolonged time, can similarly damage the membrane structure of EVs as observed in ultracentrifugation [61], which potentially leads to lower EV yield. In addition, high-speed centrifugation force causes EV aggregation [61] that interferes with subsequent characterization or functional applications.
Polymeric Precipitation
To overcome the low yield in ultracentrifuge-based EV isolation, polymeric precipitation of EVs was developed to maximize EV yield (especially from small volume biofluids) [62]. In this method, aqueous polyethylene glycol is mixed with biofluids to precipitate EVs. Although this method is convenient and effective in obtaining a high EV yield, this precipitation often leads to a crude preparation of EVs with many non-vesicular contaminants co-precipitated with EVs. In addition, there are concerns that EVs precipitated with polymeric mixtures may have altered EV properties, as observed from the characterization of precipitated EVs [63].
Size Exclusion Chromatography (SEC)
Size exclusion chromatography is emerging as a promising new method for isolating EVs based on the drastic size difference between EVs (greater) and extracellular proteins (smaller), which therefore elute in separate fractions during elution (Fig. 1). As a result, SEC is able to substantially increase the purity of isolated EVs with no or minimal protein contamination. It also helps maintain the vesicle integrity for downstream applications. Because of the typical smaller size of exosomes (50-200um), SEC combined with filtration (0.22 \(\mu\)m membrane) further allows preferential isolation of exosomes over MVs.
Immunoaffinity Purification
Various tetraspanin proteins CD63, CD81, and CD9 have been well validated as conserved surface markers on EVs secreted from different cells of origin [11]. Proteomic analysis has begun to further identify cell-type specific surface markers for EVs, especially exosomes [34, 39]. These surface markers make it possible to develop immunoaffinity based isolation of EVs. The surface marker-based isolation can completely remove potential contamination of extracellular proteins. As CD9 is preferentially expressed on the surface of MVs but not exosomes [64], CD9-based immunoaffinity beads allow selective MV isolation. This approach is particularly advantageous for the isolation of cell-type specific EVs by using cell-type specific surface markers. On the other hand, EV yields from this approach tend to be lower and it requires highly specific and efficient antibodies. It is also difficult to remove the antibody without disrupting EVs, preventing certain downstream applications.
Depending on experimental purposes, EVs can be analyzed using a range of different approaches. Biochemically, EVs are detected based on their specific cargo or surface proteins by immunoblotting. EV morphology and size are typically characterized by transmission electron microscopy. EV quantity and size can be characterized by nanoparticle-tracking analysis (NTA) [65] and tunableresistive pulse sensing (TRPS) [66], as well as specific surface marker-based flow cytometry.
Intercellular EV Signaling in CNS Development and Physiology
The CNS is the most complex and sophisticated biological system, controlling physiology and behavior in phylogenetically diverse animal species. As a consequence of this complexity, communications among multiple classes of electrically excitable neurons and functionally heterogeneous glial cells are essential to control CNS development, physiology, and behavior. In the past decade, EVs especially exosomes secreted from various CNS cell types have emerged as a novel and important intercellular communication pathway in the CNS (Fig. 2) [9]. Depending on the source and recipient cell types, here we highlighted EV signaling among different CNS cell types.
Currently reported EV-mediated intercellular communication in CNS cell-types. Representative specific signals identified in CNS cell-derived EVs/exosomes were shown. miRs: microRNA; The size of the arrows indicates relative number of studies reported so far about the indicated intercellular communications. N: neurons; A: astroglia; O: Oligodendrocytes; M: microglia
Neuron to Neuron (N–N) EV Signaling
In 2006, a seminal study first revealed that cultured neurons release exosomes that can be promoted by neuronal depolarization [67]. Proteomic analysis of exosomes isolated from human iPSC-derived neural cultures found 20 proteins that are involved in neuronal proliferation and development, such as Cadherin 2 (CDH2), cell division cycle 42 (CDC42), and nerve growth factor receptor (NGFR), etc. Interestingly, these proteins were missing in exosomes isolated from neural cultures harboring loss-of-function mutations of methyl CpG binding protein 2 (MECP2) [68], a major cause for the rare neurodevelopmental disorder Rett syndrome. Interestingly, exosomes from control neural cultures are able to rescue neurodevelopmental deficits in vitro and promote hippocampal neurogenesis in vivo [68]. Recently, in vivo transmission electron microscopy, tracing, and proteomic experiments also demonstrated the involvement of exosomes in mediating transfer of transneuronally transported proteins from retinogeniculate inputs to excitatory lateral geniculate nucleus (LGN) neurons and further to neurons in visual cortex [69]. By analyzing sequence homology with retrotransposon Gag protein, it was found that post-synaptic Arc protein forms EV to facilitate transfer of mRNA between hippocampal neurons and participate in activity-dependent translation [70]. Additional Gag homology proteins such as PEG10 was also recently found to be able to package mRNA for intercellular transfer in neurons [71]. EV-mediated intercellular signaling has also been found in invertebrate model organisms, as evidenced by the transfer of Wingless (Wg) from synaptic boutons to a specialized muscle region, the sub synaptic reticulum (SSR), in the neuromuscular junction of fly larva [72].
Neuron to Astroglia (N-G) EV Signaling
In addition to early studies that demonstrated neuron to neuron EV communications, EVs especially exosomes, have been increasingly shown to participate in neuron to glia communication. We previously showed that cortical neurons secrete miR-124-containing exosomes that are internalized into astroglia and up-regulate astroglial GLT1 expression in culture [73] and in vivo [74]. By employing a cell-type specific fluorescent labeling of ILVs and exosomes [74], we showed that ILVs are predominantly localized in soma and dendrites of neurons in vitro and in vivo [74]. The post-synaptic somatodendritically secreted miR-containing neuronal exosomes to astroglia is in stark contrast to the typical pre-synaptically released neurotransmitter-mediated activation of receptors on astroglial surface [75]. In this mode of communication, the signals are miRs especially the neuron-specific miR-124 but not neurotransmitters. This exosomal mode of communication also alters genetic regulation in astroglia to regulate astroglial (glutamate uptake) functions, which can be slower but long-lasting. This is distinct from the typical gliotransmission response following neurotransmitter-activated astroglial surface receptors [76], which is faster but typically short-lived. Thus, exosome and neurotransmitter-mediated neuron to astroglia signaling can be complementary in responding to changes in neuronal activity in different CNS physiological context. How neuronal exosomes signal to other glial cells is currently less understood. Exosomes derived from a neuroblastoma cell line PC12 appear to promote phagocytic activity of a murine microglial cell line by increasing complement component 3 expression (77).
Glia to Neuron (G-N) EV Signaling
Despite the well-established modulatory roles of astroglial factors on neuronal synaptogenesis and synaptic transmission during development and in maintaining CNS physiology [78], whether astroglial exosome signals play a role in regulating neuronal functions is at the beginning. astroglia-derived EVs are found to influence the dendritic complexity of hippocampal neurons [79]. A recent study, by ultracentrifugation method, identified extracellular matrix protein fibulin-2 as astrocyte EV cargo to promote synapse formation in a TGF\(\beta\)-dependent manner [80]. In contrast, multiple studies have shown that oligodendrocytes (OLs) releases EVs especially following neuronal glutamatergic signaling [81]. OL-derived EVs are enriched in a number of myelin proteins, such as myelin proteolipid protein, cyclic nucleotide 3’- phosphodiesterase, myelin-associated glycoprotein, and myelin oligodendroglial glycoprotein [82] which are internalized into neurons to support neuronal resistance to oxidative stress and to promote axonal transport and long-term axonal maintenance [81, 83]. OL-derived EVs also facilitate the clearance of myelin debris by being taken up into microglia [84]. Microglial EVs were also found to promote excitatory neurotransmission by enhancing sphingolipid metabolism [85]. Consistently, a separate study found that EVs secreted from microglia carry endocannabinoid N-arachidonoylethanolamine (AEA) on their surface, which stimulates type-1 cannabinoid receptors (CB1) on GABAergic neurons to inhibit their presynaptic transmission [86]. As our current understanding of intercellular MV/exosome signaling is largely gained from cancer/immune cells, and cell-type specific regulators (such as different Rab GTPases in mediating intracellular vesicle trafficking) can be involved in MV/exosome biogenesis and secretion, it is important to specifically investigate how MVs/exosomes are generated and secreted from different CNS cell types. Indeed, neuronal activity has been shown to promote exosome secretion in neurons [67] and extracellular ATP has been identified as a strong stimulant for microglial EV biogenesis [87, 88]. Additional CNS specific signals that stimulate release of EVs from different CNS cell types needs to be better characterized.
Involvement of EV Signaling in ALS Pathogenesis
Involvement of EVs in different neurodegenerative diseases and neural injury has been widely observed [89], suggesting that EV signaling is likely a shared pathogenic mechanism for CNS pathology. As the pathogenic roles of EVs in other neurodegenerative diseases and stroke have been previously summarized elsewhere [89, 90], here we will focus on the pathogenic role of EVs in ALS.
ALS is a typical progressive neurodegenerative disease in which upper and lower motor neurons (UMNs and LMNs) are degenerated. Current understanding of pathogenic mechanisms of ALS has mostly been gained from studies of familial ALS, which is caused by mutations of individual genes, including Sod1, Tardbp, and C9orf72, etc. [91]. Previous studies from human postmortem tissues and animal models, especially transgenic SOD1 mutant models, have suggested that altered glia to neuron signaling pathways, such as induction of a pro-inflammatory cytokine environment [92], reduced glutamate uptake and metabolic support [93, 94], play important roles in the pathogenesis of ALS. In particular, astroglia conditioned medium (ACM) from mouse astroglia expressing human SOD1 (hSOD1) mutant or from human ALS patient brain astroglia is able to substantially modulate health and survival of either primary or embryonic stem cell-derived motor neurons, despite the elusive identity of such ACM toxic factors [95,96,97]. Interestingly, an early study found that exosomes secreted from mutant SOD1 overexpressing astroglia in vitro carry mutant SOD1 protein which is transferred to spinal neurons through exosomes [98]. These astroglial exosomes are also toxic to motor neurons [98], implicating that astroglial exosomes could be potential astroglia-secreted toxic factors.
In ALS and other typical neurodegenerative diseases, abnormal protein aggregation has been recognized a prominent pathological feature in both human post-mortem tissues and animal models [99]. These aggregates are typically formed due to conformational changes, such as misfolding of disease pathogenic proteins, which are caused by either genetic mutations or disease-associated post-translational modifications. Similar to prion protein propagation, abnormal aggregates are able to transmit across cells and induce protein conformational changes in recipient cells to continuously propagate aggregates which eventually spread to many CNS regions [100]. How abnormal protein aggregates transmit across cells is less understood. Various mechanisms such as trans-synaptic transmission, astroglial gap junctions, and microglial phagocytosis have all been proposed [101, 102]. As EVs are constantly released from various CNS cells, including neurons where the majority of protein aggregates are formed, it is appealing to test whether misfolded proteins are contained in EVs and transmitted through EVs. Spreading of disease-relevant misfolded proteins or protein aggregates has been observed or proposed in the CNS in vitro and in vivo (Fig. 3). By overexpression of either WT or mutant SOD1 in NSC-34 and HEK cells, it was found that misfolded SOD1 proteins (either from WT or mutant SOD1) were associated with secreted EVs, and such misfolded SOD1 proteins were transferred across cells in an EV-dependent manner [103]. Additional in vivo studies further showed that astroglia and neurons are main sources of misfolded SOD1-containing EVs isolated from SOD1G93A mouse and human SOD1 familial ALS patient spinal cords [104].
Potential involvement of CNS cell-type specific EVs in the propagation of various protein aggregates in neurodegenerative diseases. Stressed neuron-derived aggregate-containing EVs can either be taken up directly into healthy neurons or be phagocytosed into microglia or astroglia and then secreted in EVs to be taken up by healthy neurons. Representative disease-relevant mutant protein aggregates were shown. EVs secreted from neurons, microglia, or astroglia are in grey, blue, and purple, respectively. EVs in red indicate protein aggregate-containing EVs from different cell types
In addition to the misfolded SOD1 protein, TDP-43 protein (encoded by the Tardbp gene) also forms pathological intracellular aggregates that can be found in CNS tissues of many ALS and frontal temporal dementia (FTD) patients [105]. TDP-43 protein contains a glycine-rich and prion-like domain in its C terminus that can form oligomers to be released and taken up in neurons to induce redistribution of nuclear TDP-43 to the cytoplasm [106, 107], a characteristic feature of TDP43 pathology. In a TDP-43 overexpressing HEK cell model, TDP43 oligomers were found in exosomes which were preferentially taken up by naïve HEK-293 cells and exerted higher toxicity than free TDP-43 [108]. By employing microfluidic neuronal cultures, additional anterograde and retrograde trans-synaptic spreading of TDP-43 was also observed [108], suggesting that both exosome-dependent and independent mechanisms are involved in TDP-43 intercellular transmission. Examination of exosomes isolated from control and ALS brains found substantially increased full-length TDP-43 and especially its C-terminal fragment in ALS brain exosomes [109]. Subsequent exposure of a neuronal cell line Neuro2a to ALS (but not control) brain exosomes further caused cytoplasmic redistribution of TDP-43, supporting the role of exosomes in propagating TDP-43 proteinopathy [109]. Despite these in vitro observations, in vivo administration of the exosome biogenesis inhibitor GW4869 exacerbated the disease phenotypes of human TDP-43A315T transgenic mice [109]. As GW4869 likely inhibits exosome biogenesis broadly in a non-cell type dependent manner, it is possible that beneficial exosome signaling is also blocked, leading to exacerbated disease phenotypes. Additional cell-type specific inhibition of exosome signaling will help dissect out the contributions of exosomes from different cell types. Recently, aggregates formed by dipeptide repeat proteins (DPRs) translated in a non-ATG-dependent fashion (RAN-translation) from hexanucleotide repeat expanded C9orf72 mRNAs have been found in post-mortem ALS and FTD brain tissues [110, 111]. Depending on the specific genetic codon frame and orientation, 5 DPRs—poly(GA), poly(GP), poly(GR), poly(PA), and poly(PR)—can be generated [112]. Among these diverse DPRs, poly(GA), poly(GP), poly(GR), and poly(PA) have been shown to partially transmit cell-to-cell in an exosome-dependent manner [113].
While there are a considerable number of reports to support a pathological role of EVs/exosomes in the propagation of abnormal protein aggregates and even in mediating toxicity to neurons in ALS, it is important to point out that the majority of these studies were carried out in cultured cell lines or primary cells and rely on the overexpression system. In addition, all of these studies employed ultracentrifugation-based approaches to prepare EVs/exosomes – a method that is likely to co-isolate directly secreted proteins (including misfolded proteins) with EVs, which complicates the interpretation of results. Indeed, direct secretion of protein signals from CNS cell types has been widely observed. Misfolded SOD1 has been shown to be directly secreted by Golgi-derived secretory vesicles [114]. Moreover, how cell-type specific exosomes are involved in the propagation of misfolded proteins in ALS remains unclear. For example, are misfolded proteins phagocytosed into glia first and then secreted in their exosomes? Or directly taken up by neurons in vivo? Potential involvement of CNS cell-type specific EVs in the propagation of protein aggregates is shown in Fig. 3. Further studies using isolation approaches that better separate EVs from secreted proteins such as SEC or immunoaffinity-based isolation should be performed to validate these observations.
In addition to the emerging roles of EVs in the CNS pathogenesis of ALS, as peripheral axon degeneration especially denervation from muscle is widely observed in ALS, whether EVs derived from peripheral Schwann cells and muscle myotube cells have a role in ALS has begun to be examined. Schwann cells are the major myelinating cells in the peripheral nervous system (PNS), how Schwann cells are involved in ALS remains unclear. Selective expression of mutant SOD1G93A in Schwann cells induces no obvious changes to locomotion or axonal degeneration [115]. On the other hand, selective deletion of mutant SOD1G37R in Schwann cells leads to unexpected acceleration of disease pathology [116]. Despite the unclear roles of Schwann cells in ALS, Schwann cell-derived exosomes are able to stimulate neurite growth of diabetic dorsal root ganglia (DRG) neurons after intravenously administered via tail vein [117]. These exosomes also promote axonal regeneration after sciatic nerve injury in vivo [118], indicating a therapeutic effect for axon injury conditions including ALS. On the other hand, smaller size (mean 123 nm), exosome-like vesicles isolated form the myotube cell cultures of ALS (C9orf72 mutation or sporadic) patients but not healthy controls reduced the survival of motor neurons by 31%, decreased neurite length and branching [119]. Specific mechanisms or exosome cargos that mediate these effects remain to be defined in future studies.
Potential of EVs in ALS Diagnostics
The search for and validation of biomarkers for the diagnosis and monitoring of disease progression of ALS has been one of the central goals in ALS research. Based on proposed pathogenic mechanisms of ALS, individual biomarkers that are associated with excitotoxicity, oxidative stress, inflammation, neurodegeneration, and metabolic dysfunction have been identified [120, 121]. However, the clinical application of these biomarkers is currently limited due to insufficient patient cohorts or the lack of specificity for ALS. As a result, there is a need to search for and develop new biomarkers for diagnostics and monitoring disease progression of ALS. Ever since EVs were found to mediate intercellular communication, it has drawn substantial interest toward examining EV cargoes as potential biomarkers for various disease conditions [122].
Although EVs have been detected in various biofluids including blood plasma (or serum), CSF, saliva, and urine [123], CSF and blood have been the primary sample sources for testing EV isolation and cargo changes in neurodegenerative diseases including ALS [89]. Misfolded SOD1 has been consistently detected in CSF of ALS patients [124]; however, whether it is contained in EVs in CSF has not yet been determined. On the other hand, both the full-length and C-terminal TDP-43 fragments were detected in EVs prepared from a cohort of ALS/FTD patient CSF samples [125]. A comprehensive proteomic analysis of exosome-enriched fractions isolated from CSF of control and ALS patients found increase of nucleolar complex protein 2 homolog (NOC2L), programmed cell death 6-interacting protein (PDCD6IP), and versican core protein (VCAN) and decrease of alpha-1-antichymotrypsin (SERPINA3), receptor-type tyrosine-protein phosphatase zeta (PTPRZ1), complement C1q subcomponent subunit C (C1QC), coiled-coil domain-containing protein 19, mitochondrial (CCDC19), myosin light chain 6B (MYL6B), Macrophage receptor MARCO, IgG Fc-binding protein (FCGBP), folate receptor alpha (FOLR1), reelin, complement factor B (CFB), and charged multivesicular body protein 4a (CHMP4A) in the ALS patient samples [126]. As low amounts of pathological proteins are typically detected in EVs from patient CSF samples, changes in exosomal RNAs, especially miRs from ALS CSF were also investigated. A RNA-Seq study revealed that differentially expressed exosomal mRNAs between control and ALS CSF samples are primarily involved in the ubiquitin–proteasome pathway, oxidative stress response, and the unfolded protein response pathways [127]. We recently found elevated exosomal miR-124 secretion from SOD1G93A spinal neurons and a positive correlation between increased exosomal miR-124 levels and disease severity especially in male ALS patients [128].
Although the CNS is well insulated from the periphery largely through the blood–brain barrier (BBB), the integrity of BBB decreases due to aging as well as the elevated neuroinflammation that is commonly observed in neurodegenerative diseases including ALS [129, 130]. In addition, recent exciting progress in understanding brain lymphatics, i.e., meningeal lymphatics and the glymphatic system [131, 132], have unveiled that meningeal lymphatic vessels are the preferred route for CSF uptake and drainage to cervical lymph nodes (cLNs) in model animals (rodent, primates) and humans, potentially opening a new way for communication between the periphery and the CNS. This also allows the potential exit of CNS-derived exosomes in disease conditions. This is particularly appealing for CNS diseases and injury due to the difficult accessibility of the CNS in patients. Thus, CNS-derived exosomes in peripheral circulation potentially provide a promising “window” in detecting disease-associated early changes in the CNS. By analyzing either (presumably) neuron-derived exosomes using a surface marker, L1 CAM [133], or all blood plasma (or serum) exosomes, attempts have been carried out to test changes in blood exosomal proteins or RNAs in ALS [134]. For example, a longitudinal analysis of ALS patient (n = 18) plasma found increased exosomal TDP-43 levels as disease progresses [135]. Several other studies have also found altered miRs in blood of sporadic or familial ALS patients [136, 137]. Currently, dynamic changes of CNS-derived exosomes into peripheral circulation have not been well examined in animal models of neurodegenerative disease including ALS. Blood exosomes are also highly mixed and derived from very heterogeneous cell origins. The marker L1CAM, that has been widely used to selectively isolate neuronal exosomes from blood samples, may not be specifically associated with neuronal exosomes, as shown in a recent study [138]. Moreover, many detected disease pathogenic proteins or miRs are often not selectively expressed in the CNS, casting doubt on whether changes in blood exosome cargoes are closely correlated with specific brain disease changes. Nevertheless, these attempts provided new knowledge about specific changes in blood exosome cargoes in ALS and potential new insights into the pathogenesis of ALS. Future studies are required to specifically trace and analyze CNS-derived exosomes in the peripheral circulation in ALS.
Conclusion and Future Perspectives
Studies in the past decade have demonstrated that secreted EVs, especially exosomes, represent an exciting new mechanism in mediating intercellular communication that play important roles in a range of (patho)physiological conditions. In comparison to typical Golgi budded secretory vesicle-mediated release of proteins or synaptic vesicle released neurotransmitters, EV communication is advantageous in that it better protects signals (especially nucleic acids) from degradation by encapsulating them into the vesicles. EVs also facilitate long-distance signaling and allow selective targeting to certain recipient cells. This is particularly relevant to the CNS, in which certain proteins and mRNAs/miRs (and possibly lipids) are selectively expressed in highly heterogeneous (or subsets of) cell types in the CNS. EVs provide a way for these selectively expressed biomolecules to be delivered into distinctive CNS cell types, as shown in neuronal exosomal miR-124 to astroglia communication [74], to exert ectopic functions. This is also highly relevant to ALS, as altered neuron to glial communication is well-established in ALS pathogenesis. In addition, while still at the very early stage, EV mediated therapeutics already exhibit great potential. Exosomes isolated from various cell sources especially mesenchymal stem cells (MSC) are able to promote axon regeneration following injury [139]. EVs and their cargoes are also considered to mediate the regenerative potential of stem cells [140]. EVs also hold the potential to become a new platform for RNA delivery, especially across the BBB for CNS delivery with surface modifications [141].
With the increasingly recognized functions of EVs and their great potential in therapeutics, it is equally important to realize that many hurdles need to be tackled to advance our understanding of EV cell biology and its involvement in ALS pathogenesis, and to develop EVs as potential therapeutics. Currently, most EV/exosome knowledge has been gained based on culture models or human CSF samples. Exosome signaling in situ in the CNS remains essentially unexplored. Several newly developed Cre-dependent cell-type specific exosome reporter mice will help examine the in vivo distribution of EVs from different CNS cells (74, 142, 143). In addition, how MV/exosome pathways are regulated in different CNS cell types and how selectively it targets to recipient CNS cell types, as well as its alterations in pathological conditions remains largely unknown, making it difficult to dissect out the role of this pathway in disease pathogenesis. Given the complexity of efficient and selective isolation of EVs, especially in a cell-type specific manner from in vivo sources, and the characterization of their heterogeneity, interdisciplinary collaboration with complementary expertise will be needed to further develop new approaches and rigorously examine the specific roles of EVs in CNS physiology and pathology.
References
Chargaff E, West R. The Biological Significance of the Thromboplastic Protein of Blood. J Biol Chem. 1946;166(1):189–97.
Trams EG, Lauter CJ, Salem N Jr, Heine U. Exfoliation of membrane ecto-enzymes in the form of micro-vesicles. Biochim Biophys Acta. 1981;645(1):63–70.
Harding C, Heuser J, Stahl P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. J Cell Biol. 1983;97(2):329–39.
Kleijmeer MJ, Raposo G, Geuze HJ. Characterization of MHC Class II Compartments by Immunoelectron Microscopy. Methods. 1996;10(2):191–207.
Ratajczak J, Miekus K, Kucia M, Zhang J, Reca R, Dvorak P, et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia. 2006;20(5):847–56.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9(6):654–9.
Wendler F, Favicchio R, Simon T, Alifrangis C, Stebbing J, Giamas G. Extracellular vesicles swarm the cancer microenvironment: from tumor-stroma communication to drug intervention. Oncogene. 2017;36(7):877–84.
Han L, Lam EW, Sun Y. Extracellular vesicles in the tumor microenvironment: old stories, but new tales. Mol Cancer. 2019;18(1):59.
Budnik V, Ruiz-Canada C, Wendler F. Extracellular vesicles round off communication in the nervous system. Nat Rev Neurosci. 2016;17(3):160–72.
Lim YJ, Lee SJ. Are exosomes the vehicle for protein aggregate propagation in neurodegenerative diseases? Acta Neuropathol Commun. 2017;5(1):64.
Colombo M, Raposo G, Thery C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu Rev Cell Dev Biol. 2014;30:255–89.
Willms E, Cabanas C, Mager I, Wood MJA, Vader P. Extracellular Vesicle Heterogeneity: Subpopulations, Isolation Techniques, and Diverse Functions in Cancer Progression. Front Immunol. 2018;9:738.
Mathieu M, Martin-Jaular L, Lavieu G, Thery C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat Cell Biol. 2019;21(1):9–17.
Zhang H, Freitas D, Kim HS, Fabijanic K, Li Z, Chen H, et al. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat Cell Biol. 2018;20(3):332–43.
Wang Q, Yu J, Kadungure T, Beyene J, Zhang H, Lu Q. ARMMs as a versatile platform for intracellular delivery of macromolecules. Nat Commun. 2018;9(1):960.
Zhang Q, Jeppesen DK, Higginbotham JN, Graves-Deal R, Trinh VQ, Ramirez MA, et al. Supermeres are functional extracellular nanoparticles replete with disease biomarkers and therapeutic targets. Nat Cell Biol. 2021;23(12):1240–54.
Wolf P. The nature and significance of platelet products in human plasma. Br J Haematol. 1967;13(3):269–88.
van Niel G, D’Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol. 2018;19(4):213–28.
Hugel B, Martinez MC, Kunzelmann C, Freyssinet JM. Membrane microparticles: two sides of the coin. Physiology (Bethesda). 2005;20:22–7.
Blanc L, Vidal M. New insights into the function of Rab GTPases in the context of exosomal secretion. Small GTPases. 2018;9(1–2):95–106.
Muralidharan-Chari V, Clancy J, Plou C, Romao M, Chavrier P, Raposo G, et al. ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr Biol. 2009;19(22):1875–85.
Buschow SI, Nolte-’t Hoen EN, van Niel G, Pols MS, ten Broeke T, Lauwen M, et al. MHC II in dendritic cells is targeted to lysosomes or T cell-induced exosomes via distinct multivesicular body pathways. Traffic. 2009;10(10):1528–42.
Chairoungdua A, Smith DL, Pochard P, Hull M, Caplan MJ. Exosome release of beta-catenin: a novel mechanism that antagonizes Wnt signaling. J Cell Biol. 2010;190(6):1079–91.
Doyle LM, Wang MZ. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells. 2019;8(7).
Lv Y, Tan J, Miao Y, Zhang Q. The role of microvesicles and its active molecules in regulating cellular biology. J Cell Mol Med. 2019;23(12):7894–904.
Tricarico C, Clancy J, D’Souza-Schorey C. Biology and biogenesis of shed microvesicles. Small GTPases. 2017;8(4):220–32.
Bonifacino JS, Glick BS. The mechanisms of vesicle budding and fusion. Cell. 2004;116(2):153–66.
Schlienger S, Campbell S, Claing A. ARF1 regulates the Rho/MLC pathway to control EGF-dependent breast cancer cell invasion. Mol Biol Cell. 2014;25(1):17–29.
Li B, Antonyak MA, Zhang J, Cerione RA. RhoA triggers a specific signaling pathway that generates transforming microvesicles in cancer cells. Oncogene. 2012;31(45):4740–9.
Al-Nedawi K, Meehan B, Micallef J, Lhotak V, May L, Guha A, et al. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat Cell Biol. 2008;10(5):619–24.
Hristov M, Erl W, Linder S, Weber PC. Apoptotic bodies from endothelial cells enhance the number and initiate the differentiation of human endothelial progenitor cells in vitro. Blood. 2004;104(9):2761–6.
Colombo M, Moita C, van Niel G, Kowal J, Vigneron J, Benaroch P, et al. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J Cell Sci. 2013;126(Pt 24):5553–65.
Hurley JH. ESCRT complexes and the biogenesis of multivesicular bodies. Curr Opin Cell Biol. 2008;20(1):4–11.
Kowal J, Arras G, Colombo M, Jouve M, Morath JP, Primdal-Bengtson B, et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc Natl Acad Sci U S A. 2016;113(8):E968–77.
Baietti MF, Zhang Z, Mortier E, Melchior A, Degeest G, Geeraerts A, et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat Cell Biol. 2012;14(7):677–85.
Donoso-Quezada J, Ayala-Mar S, Gonzalez-Valdez J. The role of lipids in exosome biology and intercellular communication: Function, analytics and applications. Traffic. 2021;22(7):204–20.
Trajkovic K, Hsu C, Chiantia S, Rajendran L, Wenzel D, Wieland F, et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science. 2008;319(5867):1244–7.
Dinkins MB, Dasgupta S, Wang G, Zhu G, Bieberich E. Exosome reduction in vivo is associated with lower amyloid plaque load in the 5XFAD mouse model of Alzheimer’s disease. Neurobiol Aging. 2014;35(8):1792–800.
Kugeratski FG, Hodge K, Lilla S, McAndrews KM, Zhou X, Hwang RF, et al. Quantitative proteomics identifies the core proteome of exosomes with syntenin-1 as the highest abundant protein and a putative universal biomarker. Nat Cell Biol. 2021;23(6):631–41.
Thery C, Boussac M, Veron P, Ricciardi-Castagnoli P, Raposo G, Garin J, et al. Proteomic analysis of dendritic cell-derived exosomes: a secreted subcellular compartment distinct from apoptotic vesicles. J Immunol. 2001;166(12):7309–18.
Geminard C, De Gassart A, Blanc L, Vidal M. Degradation of AP2 during reticulocyte maturation enhances binding of hsc70 and Alix to a common site on TFR for sorting into exosomes. Traffic. 2004;5(3):181–93.
de Gassart A, Geminard C, Fevrier B, Raposo G, Vidal M. Lipid raft-associated protein sorting in exosomes. Blood. 2003;102(13):4336–44.
Buschow SI, Liefhebber JM, Wubbolts R, Stoorvogel W. Exosomes contain ubiquitinated proteins. Blood Cells Mol Dis. 2005;35(3):398–403.
Luhtala N, Aslanian A, Yates JR 3rd, Hunter T. Secreted Glioblastoma Nanovesicles Contain Intracellular Signaling Proteins and Active Ras Incorporated in a Farnesylation-dependent Manner. J Biol Chem. 2017;292(2):611–28.
Irion U, St JD. bicoid RNA localization requires specific binding of an endosomal sorting complex. Nature. 2007;445(7127):554–8.
Gibbings DJ, Ciaudo C, Erhardt M, Voinnet O. Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat Cell Biol. 2009;11(9):1143–9.
Villarroya-Beltri C, Gutierrez-Vazquez C, Sanchez-Cabo F, Perez-Hernandez D, Vazquez J, Martin-Cofreces N, et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat Commun. 2013;4:2980.
Shurtleff MJ, Temoche-Diaz MM, Karfilis KV, Ri S, Schekman R. Y-box protein 1 is required to sort microRNAs into exosomes in cells and in a cell-free reaction. Elife. 2016;5.
Garcia-Martin R, Wang G, Brandao BB, Zanotto TM, Shah S, Kumar Patel S, et al. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature. 2021.
Janvier K, Pelchen-Matthews A, Renaud JB, Caillet M, Marsh M, Berlioz-Torrent C. The ESCRT-0 component HRS is required for HIV-1 Vpu-mediated BST-2/tetherin down-regulation. PLoS Pathog. 2011;7(2):e1001265.
Dias MV, Teixeira BL, Rodrigues BR, Sinigaglia-Coimbra R, Porto-Carreiro I, Roffe M, et al. PRNP/prion protein regulates the secretion of exosomes modulating CAV1/caveolin-1-suppressed autophagy. Autophagy. 2016;12(11):2113–28.
Ostrowski M, Carmo NB, Krumeich S, Fanget I, Raposo G, Savina A, et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat Cell Biol. 2010;12(1):19–30; sup pp 1–13.
Hsu C, Morohashi Y, Yoshimura S, Manrique-Hoyos N, Jung S, Lauterbach MA, et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J Cell Biol. 2010;189(2):223–32.
Savina A, Fader CM, Damiani MT, Colombo MI. Rab11 promotes docking and fusion of multivesicular bodies in a calcium-dependent manner. Traffic. 2005;6(2):131–43.
Gross JC, Chaudhary V, Bartscherer K, Boutros M. Active Wnt proteins are secreted on exosomes. Nat Cell Biol. 2012;14(10):1036–45.
Ayers L, Pink R, Carter DRF, Nieuwland R. Clinical requirements for extracellular vesicle assays. J Extracell Vesicles. 2019;8(1):1593755.
Shao H, Im H, Castro CM, Breakefield X, Weissleder R, Lee H. New Technologies for Analysis of Extracellular Vesicles. Chem Rev. 2018;118(4):1917–50.
Gardiner C, Di Vizio D, Sahoo S, Thery C, Witwer KW, Wauben M, et al. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J Extracell Vesicles. 2016;5:32945.
Sluijter JPG, Davidson SM, Boulanger CM, Buzas EI, de Kleijn DPV, Engel FB, et al. Extracellular vesicles in diagnostics and therapy of the ischaemic heart: Position Paper from the Working Group on Cellular Biology of the Heart of the European Society of Cardiology. Cardiovasc Res. 2018;114(1):19–34.
Li P, Kaslan M, Lee SH, Yao J, Gao Z. Progress in Exosome Isolation Techniques. Theranostics. 2017;7(3):789–804.
Sodar BW, Kovacs A, Visnovitz T, Pallinger E, Vekey K, Pocsfalvi G, et al. Best practice of identification and proteomic analysis of extracellular vesicles in human health and disease. Expert Rev Proteomics. 2017;14(12):1073–90.
Shin H, Han C, Labuz JM, Kim J, Kim J, Cho S, et al. High-yield isolation of extracellular vesicles using aqueous two-phase system. Sci Rep. 2015;5:13103.
Gamez-Valero A, Monguio-Tortajada M, Carreras-Planella L, Franquesa M, Beyer K, Borras FE. Size-Exclusion Chromatography-based isolation minimally alters Extracellular Vesicles’ characteristics compared to precipitating agents. Sci Rep. 2016;6:33641.
Andreu Z, Yanez-Mo M. Tetraspanins in extracellular vesicle formation and function. Front Immunol. 2014;5:442.
Sokolova V, Ludwig AK, Hornung S, Rotan O, Horn PA, Epple M, et al. Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B Biointerfaces. 2011;87(1):146–50.
Coumans FA, van der Pol E, Boing AN, Hajji N, Sturk G, van Leeuwen TG, et al. Reproducible extracellular vesicle size and concentration determination with tunable resistive pulse sensing. J Extracell Vesicles. 2014;3:25922.
Faure J, Lachenal G, Court M, Hirrlinger J, Chatellard-Causse C, Blot B, et al. Exosomes are released by cultured cortical neurones. Mol Cell Neurosci. 2006;31(4):642–8.
Sharma P, Mesci P, Carromeu C, McClatchy DR, Schiapparelli L, Yates JR 3rd, et al. Exosomes regulate neurogenesis and circuit assembly. Proc Natl Acad Sci U S A. 2019;116(32):16086–94.
Schiapparelli LM, Sharma P, He HY, Li J, Shah SH, McClatchy DB, et al. Proteomic screen reveals diverse protein transport between connected neurons in the visual system. Cell Rep. 2022;38(4):110287.
Pastuzyn ED, Day CE, Kearns RB, Kyrke-Smith M, Taibi AV, McCormick J, et al. The Neuronal Gene Arc Encodes a Repurposed Retrotransposon Gag Protein that Mediates Intercellular RNA Transfer. Cell. 2018;173(1):275.
Segel M, Lash B, Song J, Ladha A, Liu CC, Jin X, et al. Mammalian retrovirus-like protein PEG10 packages its own mRNA and can be pseudotyped for mRNA delivery. Science. 2021;373(6557):882–9.
Korkut C, Ataman B, Ramachandran P, Ashley J, Barria R, Gherbesi N, et al. Trans-synaptic transmission of vesicular Wnt signals through Evi/Wntless. Cell. 2009;139(2):393–404.
Morel L, Regan M, Higashimori H, Ng SK, Esau C, Vidensky S, et al. Neuronal exosomal miRNA-dependent translational regulation of astroglial glutamate transporter GLT1. J Biol Chem. 2013;288(10):7105–16.
Men Y, Yelick J, Jin S, Tian Y, Chiang MSR, Higashimori H, et al. Exosome reporter mice reveal the involvement of exosomes in mediating neuron to astroglia communication in the CNS. Nat Commun. 2019;10(1):4136.
Araque A, Carmignoto G, Haydon PG, Oliet SH, Robitaille R, Volterra A. Gliotransmitters travel in time and space. Neuron. 2014;81(4):728–39.
Halassa MM, Fellin T, Haydon PG. The tripartite synapse: roles for gliotransmission in health and disease. Trends Mol Med. 2007;13(2):54–63.
Bahrini I, Song JH, Diez D, Hanayama R. Neuronal exosomes facilitate synaptic pruning by up-regulating complement factors in microglia. Sci Rep. 2015;5:7989.
Chung WS, Allen NJ, Eroglu C. Astrocytes Control Synapse Formation, Function, and Elimination. Cold Spring Harb Perspect Biol. 2015;7(9):a020370.
Chaudhuri AD, Dastgheyb RM, Yoo SW, Trout A, Talbot CC Jr, Hao H, et al. TNFalpha and IL-1beta modify the miRNA cargo of astrocyte shed extracellular vesicles to regulate neurotrophic signaling in neurons. Cell Death Dis. 2018;9(3):363.
Patel MR, Weaver AM. Astrocyte-derived small extracellular vesicles promote synapse formation via fibulin-2-mediated TGF-beta signaling. Cell Rep. 2021;34(10):108829.
Fruhbeis C, Frohlich D, Kuo WP, Amphornrat J, Thilemann S, Saab AS, et al. Neurotransmitter-triggered transfer of exosomes mediates oligodendrocyte-neuron communication. PLoS Biol. 2013;11(7):e1001604.
Domingues HS, Falcao AM, Mendes-Pinto I, Salgado AJ, Teixeira FG. Exosome Circuitry During (De)(Re)Myelination of the Central Nervous System. Front Cell Dev Biol. 2020;8:483.
Fruhbeis C, Kuo-Elsner WP, Muller C, Barth K, Peris L, Tenzer S, et al. Oligodendrocytes support axonal transport and maintenance via exosome secretion. PLoS Biol. 2020;18(12):e3000621.
Fitzner D, Schnaars M, van Rossum D, Krishnamoorthy G, Dibaj P, Bakhti M, et al. Selective transfer of exosomes from oligodendrocytes to microglia by macropinocytosis. J Cell Sci. 2011;124(Pt 3):447–58.
Antonucci F, Turola E, Riganti L, Caleo M, Gabrielli M, Perrotta C, et al. Microvesicles released from microglia stimulate synaptic activity via enhanced sphingolipid metabolism. EMBO J. 2012;31(5):1231–40.
Gabrielli M, Battista N, Riganti L, Prada I, Antonucci F, Cantone L, et al. Active endocannabinoids are secreted on extracellular membrane vesicles. EMBO Rep. 2015;16(2):213–20.
Drago F, Lombardi M, Prada I, Gabrielli M, Joshi P, Cojoc D, et al. ATP Modifies the Proteome of Extracellular Vesicles Released by Microglia and Influences Their Action on Astrocytes. Front Pharmacol. 2017;8:910.
Asai H, Ikezu S, Tsunoda S, Medalla M, Luebke J, Haydar T, et al. Depletion of microglia and inhibition of exosome synthesis halt tau propagation. Nat Neurosci. 2015;18(11):1584–93.
Hill AF. Extracellular Vesicles and Neurodegenerative Diseases. J Neurosci. 2019;39(47):9269–73.
Howitt J, Hill AF. Exosomes in the Pathology of Neurodegenerative Diseases. J Biol Chem. 2016;291(52):26589–97.
Brown RH Jr, Al-Chalabi A. Amyotrophic Lateral Sclerosis. N Engl J Med. 2017;377(16):1602.
Philips T, Robberecht W. Neuroinflammation in amyotrophic lateral sclerosis: role of glial activation in motor neuron disease. Lancet Neurol. 2011;10(3):253–63.
Morrison BM, Lee Y, Rothstein JD. Oligodendroglia: metabolic supporters of axons. Trends Cell Biol. 2013;23(12):644–51.
Rothstein JD, Martin LJ, Kuncl RW. Decreased glutamate transport by the brain and spinal cord in amyotrophic lateral sclerosis. N Engl J Med. 1992;326(22):1464–8.
Di Giorgio FP, Carrasco MA, Siao MC, Maniatis T, Eggan K. Non-cell autonomous effect of glia on motor neurons in an embryonic stem cell-based ALS model. Nat Neurosci. 2007;10(5):608–14.
Nagai M, Re DB, Nagata T, Chalazonitis A, Jessell TM, Wichterle H, et al. Astrocytes expressing ALS-linked mutated SOD1 release factors selectively toxic to motor neurons. Nat Neurosci. 2007;10(5):615–22.
Haidet-Phillips AM, Hester ME, Miranda CJ, Meyer K, Braun L, Frakes A, et al. Astrocytes from familial and sporadic ALS patients are toxic to motor neurons. Nat Biotechnol. 2011;29(9):824–8.
Basso M, Pozzi S, Tortarolo M, Fiordaliso F, Bisighini C, Pasetto L, et al. Mutant copper-zinc superoxide dismutase (SOD1) induces protein secretion pathway alterations and exosome release in astrocytes: implications for disease spreading and motor neuron pathology in amyotrophic lateral sclerosis. J Biol Chem. 2013;288(22):15699–711.
Peng C, Trojanowski JQ, Lee VM. Protein transmission in neurodegenerative disease. Nat Rev Neurol. 2020;16(4):199–212.
Guo JL, Lee VM. Cell-to-cell transmission of pathogenic proteins in neurodegenerative diseases. Nat Med. 2014;20(2):130–8.
Walker LC, Diamond MI, Duff KE, Hyman BT. Mechanisms of protein seeding in neurodegenerative diseases. JAMA Neurol. 2013;70(3):304–10.
Maragakis NJ, Rothstein JD. Mechanisms of Disease: astrocytes in neurodegenerative disease. Nat Clin Pract Neurol. 2006;2(12):679–89.
Grad LI, Yerbury JJ, Turner BJ, Guest WC, Pokrishevsky E, O’Neill MA, et al. Intercellular propagated misfolding of wild-type Cu/Zn superoxide dismutase occurs via exosome-dependent and -independent mechanisms. Proc Natl Acad Sci U S A. 2014;111(9):3620–5.
Silverman JM, Christy D, Shyu CC, Moon KM, Fernando S, Gidden Z, et al. CNS-derived extracellular vesicles from superoxide dismutase 1 (SOD1)(G93A) ALS mice originate from astrocytes and neurons and carry misfolded SOD1. J Biol Chem. 2019;294(10):3744–59.
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science. 2006;314(5796):130–3.
Budini M, Buratti E, Stuani C, Guarnaccia C, Romano V, De Conti L, et al. Cellular model of TAR DNA-binding protein 43 (TDP-43) aggregation based on its C-terminal Gln/Asn-rich region. J Biol Chem. 2012;287(10):7512–25.
Zhang YJ, Xu YF, Cook C, Gendron TF, Roettges P, Link CD, et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc Natl Acad Sci U S A. 2009;106(18):7607–12.
Feiler MS, Strobel B, Freischmidt A, Helferich AM, Kappel J, Brewer BM, et al. TDP-43 is intercellularly transmitted across axon terminals. J Cell Biol. 2015;211(4):897–911.
Iguchi Y, Eid L, Parent M, Soucy G, Bareil C, Riku Y, et al. Exosome secretion is a key pathway for clearance of pathological TDP-43. Brain. 2016;139(Pt 12):3187–201.
Ash PE, Bieniek KF, Gendron TF, Caulfield T, Lin WL, Dejesus-Hernandez M, et al. Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron. 2013;77(4):639–46.
Zu T, Liu Y, Banez-Coronel M, Reid T, Pletnikova O, Lewis J, et al. RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc Natl Acad Sci U S A. 2013;110(51):E4968–77.
Nguyen L, Cleary JD, Ranum LPW. Repeat-Associated Non-ATG Translation: Molecular Mechanisms and Contribution to Neurological Disease. Annu Rev Neurosci. 2019;42:227–47.
Westergard T, Jensen BK, Wen X, Cai J, Kropf E, Iacovitti L, et al. Cell-to-Cell Transmission of Dipeptide Repeat Proteins Linked to C9orf72-ALS/FTD. Cell Rep. 2016;17(3):645–52.
Urushitani M, Sik A, Sakurai T, Nukina N, Takahashi R, Julien JP. Chromogranin-mediated secretion of mutant superoxide dismutase proteins linked to amyotrophic lateral sclerosis. Nat Neurosci. 2006;9(1):108–18.
Turner BJ, Ackerley S, Davies KE, Talbot K. Dismutase-competent SOD1 mutant accumulation in myelinating Schwann cells is not detrimental to normal or transgenic ALS model mice. Hum Mol Genet. 2010;19(5):815–24.
Lobsiger CS, Boillee S, McAlonis-Downes M, Khan AM, Feltri ML, Yamanaka K, et al. Schwann cells expressing dismutase active mutant SOD1 unexpectedly slow disease progression in ALS mice. Proc Natl Acad Sci U S A. 2009;106(11):4465–70.
Wang L, Chopp M, Szalad A, Lu X, Zhang Y, Wang X, et al. Exosomes Derived From Schwann Cells Ameliorate Peripheral Neuropathy in Type 2 Diabetic Mice. Diabetes. 2020;69(4):749–59.
Lopez-Verrilli MA, Picou F, Court FA. Schwann cell-derived exosomes enhance axonal regeneration in the peripheral nervous system. Glia. 2013;61(11):1795–806.
Anakor E, Milla V, Connolly O, Martinat C, Pradat PF, Dumonceaux J, et al. The Neurotoxicity of Vesicles Secreted by ALS Patient Myotubes Is Specific to Exosome-Like and Not Larger Subtypes. Cells. 2022;11(5).
Turner MR, Kiernan MC, Leigh PN, Talbot K. Biomarkers in amyotrophic lateral sclerosis. Lancet Neurol. 2009;8(1):94–109.
Chen X, Shang HF. New developments and future opportunities in biomarkers for amyotrophic lateral sclerosis. Transl Neurodegener. 2015;4:17.
Xu K, Liu Q, Wu K, Liu L, Zhao M, Yang H, et al. Extracellular vesicles as potential biomarkers and therapeutic approaches in autoimmune diseases. J Transl Med. 2020;18(1):432.
Liang Y, Lehrich BM, Zheng S, Lu M. Emerging methods in biomarker identification for extracellular vesicle-based liquid biopsy. J Extracell Vesicles. 2021;10(7):e12090.
Tokuda E, Takei YI, Ohara S, Fujiwara N, Hozumi I, Furukawa Y. Wild-type Cu/Zn-superoxide dismutase is misfolded in cerebrospinal fluid of sporadic amyotrophic lateral sclerosis. Mol Neurodegener. 2019;14(1):42.
Ding X, Ma M, Teng J, Teng RK, Zhou S, Yin J, et al. Exposure to ALS-FTD-CSF generates TDP-43 aggregates in glioblastoma cells through exosomes and TNTs-like structure. Oncotarget. 2015;6(27):24178–91.
Hayashi N, Doi H, Kurata Y, Kagawa H, Atobe Y, Funakoshi K, et al. Proteomic analysis of exosome-enriched fractions derived from cerebrospinal fluid of amyotrophic lateral sclerosis patients. Neurosci Res. 2020;160:43–9.
Otake K, Kamiguchi H, Hirozane Y. Identification of biomarkers for amyotrophic lateral sclerosis by comprehensive analysis of exosomal mRNAs in human cerebrospinal fluid. BMC Med Genomics. 2019;12(1):7.
Yelick J, Men Y, Jin S, Seo S, Espejo-Porras F, Yang Y. Elevated exosomal secretion of miR-124–3p from spinal neurons positively associates with disease severity in ALS. Exp Neurol. 2020;333:113414.
Kinney JW, Bemiller SM, Murtishaw AS, Leisgang AM, Salazar AM, Lamb BT. Inflammation as a central mechanism in Alzheimer’s disease. Alzheimers Dement (N Y). 2018;4:575–90.
Winkler EA, Sengillo JD, Sagare AP, Zhao Z, Ma Q, Zuniga E, et al. Blood-spinal cord barrier disruption contributes to early motor-neuron degeneration in ALS-model mice. Proc Natl Acad Sci U S A. 2014;111(11):E1035–42.
Da Mesquita S, Fu Z, Kipnis J. The Meningeal Lymphatic System: A New Player in Neurophysiology. Neuron. 2018;100(2):375–88.
Jessen NA, Munk AS, Lundgaard I, Nedergaard M. The Glymphatic System: A Beginner’s Guide. Neurochem Res. 2015;40(12):2583–99.
Mustapic M, Eitan E, Werner JK Jr, Berkowitz ST, Lazaropoulos MP, Tran J, et al. Plasma Extracellular Vesicles Enriched for Neuronal Origin: A Potential Window into Brain Pathologic Processes. Front Neurosci. 2017;11:278.
Ferrara D, Pasetto L, Bonetto V, Basso M. Role of Extracellular Vesicles in Amyotrophic Lateral Sclerosis. Front Neurosci. 2018;12:574.
Chen PC, Wu D, Hu CJ, Chen HY, Hsieh YC, Huang CC. Exosomal TAR DNA-binding protein-43 and neurofilaments in plasma of amyotrophic lateral sclerosis patients: A longitudinal follow-up study. J Neurol Sci. 2020;418:117070.
Freischmidt A, Muller K, Zondler L, Weydt P, Mayer B, von Arnim CA, et al. Serum microRNAs in sporadic amyotrophic lateral sclerosis. Neurobiol Aging. 2015;36(9):2660 e15–20.
Liguori M, Nuzziello N, Introna A, Consiglio A, Licciulli F, D’Errico E, et al. Dysregulation of MicroRNAs and Target Genes Networks in Peripheral Blood of Patients With Sporadic Amyotrophic Lateral Sclerosis. Front Mol Neurosci. 2018;11:288.
Norman M, Ter-Ovanesyan D, Trieu W, Lazarovits R, Kowal EJK, Lee JH, et al. L1CAM is not associated with extracellular vesicles in human cerebrospinal fluid or plasma. Nat Methods. 2021;18(6):631–4.
Zhang Y, Chopp M, Liu XS, Katakowski M, Wang X, Tian X, et al. Exosomes Derived from Mesenchymal Stromal Cells Promote Axonal Growth of Cortical Neurons. Mol Neurobiol. 2017;54(4):2659–73.
Lamichhane TN, Sokic S, Schardt JS, Raiker RS, Lin JW, Jay SM. Emerging roles for extracellular vesicles in tissue engineering and regenerative medicine. Tissue Eng Part B Rev. 2015;21(1):45–54.
Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJ. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol. 2011;29(4):341–5.
Neckles VN, Morton MC, Holmberg JC, Sokolov AM, Nottoli T, Liu D, et al. A transgenic inducible GFP extracellular-vesicle reporter (TIGER) mouse illuminates neonatal cortical astrocytes as a source of immunomodulatory extracellular vesicles. Sci Rep. 2019;9(1):3094.
Sheller-Miller S, Choi K, Choi C, Menon R. Cyclic-recombinase-reporter mouse model to determine exosome communication and function during pregnancy. Am J Obstet Gynecol. 2019;221(5):502 e1- e12.
Acknowledgements
We thank Rachel Jarvis’s proofreading for the manuscript. This work was supported by NIH grants R01NS125490, R01NS118747, RF1AG059610, and RF1AG057882 (YY).
Required Author Forms
Disclosure forms provided by the authors are available with the online version of this article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kim, G., Chen, X. & Yang, Y. Pathogenic Extracellular Vesicle (EV) Signaling in Amyotrophic Lateral Sclerosis (ALS). Neurotherapeutics 19, 1119–1132 (2022). https://doi.org/10.1007/s13311-022-01232-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13311-022-01232-9