Abstract
It has been reported that miR-615-5p was upregulated in hepatocellular carcinoma (HCC) preventing both growth and migration. However, the underlying mechanism by which miR-615-5p played a role in HCC remains unknown. Here, in our present study, to investigate the mechanism of miR-615-5p, bioinformatic prediction and luciferase reporter assay were employed to ascertain the downstream target of miR-615-5p finding that the serine hydromethyltransferase 2 (SHMT2) was the direct downstream target. Knockdown or overexpression of miR-615-5p can lead to increasing or decreasing expression of SHMT2 in HCC cells. Besides, knockdown or overexpression of SHMT2 can suppress or promote both proliferation and migration of HCC cells, indicating that miR-615-5p can directly and negatively regulate the SHMT2 in HCC cells. In addition, to understand the clinicopathological significance of SHMT2 expression in HCC, immunohistochemistry was performed. It was found that SHMT2 expression was significantly associated with poor prognosis and TNM stage. Together, our results for the first time showed that miR-615-5p prevents proliferation and migration through negatively regulating SHMT2 in HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is a highly lethal cancer, which has been ranked as the fifth most common malignancy and the third leading cause of cancer-related mortality worldwide [1]. Despite the tremendous progress in diagnosis and multimodality treatment in the past decades, the prognosis of HCC patients remains gloomy, mainly because of its high recurrent [2] and metastatic rate [3]. Therefore, there is an urgent need to unravel the molecular mechanism by which how HCC cells metastasize and grow.
To date, numerous studies have found that microRNAs (miRs) have been involved and played an important role in the carcinogenesis of HCC [4, 5], with some miRs being upregulated and others downregulated in HCC. Despite some miRs overexpressed in cancer tissues may be usually as oncogenes to promote tumorigenesis and others downregulated in cancer tissues may have a tumor suppressive role, it is not necessarily the case. Take miR-615-5p for example. miR-615-5p has been reported to be overexpressed in HCC tissues and cell lines compared with normal controls; however, whose overexpression alleviated both the proliferation and migration of HCC cells in vitro [6], indicating the tumor suppressor role in HCC [6–8]. Given the rather limited data regarding miR-615-5p in the setting of HCC, the molecular mechanism by which miR-615-5p plays a role in HCC remains unknown.
In our study, our bioinformatics analysis found that serine hydromethyltransferase 2 (SHMT2) was a novel downstream target of miR-615-5p. To explore the relationship between miR-615-5p and SHMT2, we performed relevant experiments in the following present study.
Materials and methods
Clinical tissues
The present study was approved by the Medical Ethics Committee of Jiangsu Cancer Hospital, the Affiliated Cancer Hospital of Nanjing Medical University. One hundred cases of HCC and paired normal control tissues were collected and retrieved from the Department of General Surgery in Jiangsu Cancer Hospital. Signed informed consents were obtained from all subjects involved. None of the recruited patients received treatment before surgery and for all patients whose clinical-pathological information was available. Representative hematoxylin and eosin (H&E)-stained slides from each patient were retrospectively reviewed blindly and separately by two pathologists. All fresh tissues were separately excised by experienced pathologists and were frozen in liquid nitrogen within 30 min after surgery and stored at −80 °C until analysis.
Cell culture
The human HCC cell lines HepG2, Hep3B, Huh1, Huh7, and HLE as well as normal human liver epithelial cell line THLE3 were all obtained from American Type Culture Collection (ATCC; Manassas, VA, USA). These cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Invitrogen, Carlsbad, CA, USA) supplemented with 10 % fetal bovine serum (FBS) and penicillin/streptomycin in a humidified incubator at 37 °C with an atmosphere of 5 % CO2, unless otherwise stated. Transfection was performed with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s guidance.
Small interfering RNA and shRNA
Both miR-615-5p mimics and inhibitor sequences were designed and synthesized by Genepharm company (Genepharm, Shanghai, China), so did SHMT2. All the small interference RNA (siRNA) sequences against SHMT2 and scramble control as well as miR-615-5p mimics and inhibitor sequences are listed in Supplementary Table 1.
Real-time quantitative reverse transcription PCR
Total RNA of the cultured cells was prepared using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). The quality and quantity of RNA was determined using a NanoDrop® ND-1000 spectrophotometer (Thermal Fisher, Wilmington, DE, USA). Reverse transcription (RT) was performed using random primers of the SuperScript III First-Strand Synthesis SuperMix kit (Invitrogen, Carlsbad, CA, USA), following the manufacture’s protocols. All the samples were tested with SYBR Green PCR Master Mix (Takara, Dalian, China). U6 expression was assayed for normalization, and miR-615-5p relative gene expression determinations were determined using the comparative delta-delta CT method (2−ΔΔCT) using IQ5 software. The expression of miR-615-5p was normalized with U6. Both the primers involved were designed and synthesized by Shangong company (Shangong, Shanghai, China). All the primers are listed in Supplementary Table 1.
Luciferase reporter assay
HLE and HepG2 cells were maintained in a 24-well plate and co-transfected with miR-615-5p mimics or mutation scramble and pMirTarget-SHMT2-3′-UTR-WT or pMirTarget-SHMT2-3′-UTR-MT (OriGene, USA). Forty-eight hours later, cell lysates were prepared and the relative dual-luciferase activity was examined with the Dual-Luciferase Reporter Assay System (Promega, WI, USA).
Methylthiazolyl blue tetrazolium
Cell viability was examined using the methylthiazolyl blue tetrazolium (MTT) assay (Shangong, Shanghai, China), according to the standard protocol after transfection for 24, 48, 72, and 96 h. Huh7 and HepG2 cells were plated in 96-well plates at a density of 5 × 103 cells/well. After transfection, cell proliferation was assessed. Cells were incubated for 4 h in 20 μL MTT at 37 °C. The color was developed by incubating the cells in 150 μL dimethyl sulfoxide (DMSO); the absorbance was detected at 490 nm wavelength. The data were obtained from three independent experiments.
Wound healing assay
Cell migration ability was calculated using the wound healing assay. HCC cells were plated in a 6-well plate at a concentration of 4 × 105 cells/well and allowed to form a confluent monolayer for 24 h. After the transfection, the monolayer was scratched with a sterile pipette tip (10 μL), washed with serum-free medium to remove floated and detached cells, and photographed (time 0 and 48 h) using an inversion fluorescence microscope (Olympus, Japan).
Western blot
Cell lysates were prepared in RIPA lysis buffer (BioTeke, Beijing, China). Equal amount of total cell lysates was separated by 10 % SDS-PAGE and transferred to PVDF membranes incubated with primary antibodies against SHMT2 (no. 12762; Cell Signaling Technology, USA) and β-actin (sc-47778; Santa Cruz, CA, USA) overnight. Membranes were subsequently probed with AP-conjugated secondary antibodies, the blots were visualized with chemiluminescence with SuperSignal West Femto Chemiluminescent Substrate (Thermo Scientific, USA), and images were captured with a Bio-Rad camera system (Bio-Rad, USA).
Immunohistochemistry
H&E-stained slides and unstained slides for immunohistochemical analysis were prepared from formalin-fixed, paraffin-embedded blocks of HCC tissues. Immunohistochemical stains were performed using heat-induced epitope retrieval, an avidin-biotin complex method. The rabbit anti-SHMT2 antibody (TA327002; OriGene, USA) was diluted with 1:200 following the recommendation. The sections were evaluated by light microscopic examination, and cellular localization of the protein and immunostaining level in each section were assessed by two pathologists. The staining patterns were scored as follows: negative, represented as –; weak (less than 30 % of cells with positive staining), represented as +; moderately positive (more than 30 % but less than 60 % of cells with positive staining), represented as ++; and strongly positive (more than 60 % of cells with positive staining), represented as +++, according to the signal intensity. Both negative and weak immunostaining were categorized into low expression, whereas moderate and strong staining were categorized into high expression.
Statistics
All the results were expressed as mean ± standard deviation (SD) based on at least three independent experiments. Statistical analysis was performed using SPSS 17.0 (SPSS, Chicago, IL, USA). For statistical comparisons, independent t test was employed. p value less than 0.05 was considered to be statistically significant, whereas it is taken to be extremely significant when p value was less than 0.01.
Results
miR-615-5p was markedly upregulated in HCC tissues and prevents both growth and migration of HCC cells
Having been reported to be overexpressed in HCC, the expression level of miR-615-5p was confirmed first of all in our study. Quantitative reverse transcription PCR (qRT-PCR) result reproduced that miR-615-5p was significantly upregulated in HCC tissues compared with paired normal control tissues (Fig. 1a). Moreover, we have also analyzed the clinicopathological significance of miR-615-5p expression. It was found that miR-615-5p was significantly and negatively associated with metastasis and TNM stage (Supplementary Table 2). To investigate the biological role of miR-615-5p in HCC cells, a panel of HCC cell lines was employed with miR-615-5p being different basal levels (Fig. 1b). To confirm the role in the proliferation and migration of HCC cells that has been scarcely reported, both MTT and wound healing assay were carried out. On the basis of successful knockdown or overexpression of miR-615-5p by transfection with miR-615-5p inhibitor or its mimics (Supplementary Fig. 1), MTT results showed that knockdown of miR-615-5p can remarkably promote the proliferation in Huh7 (Fig. 1c) and HepG2 (Fig. 1d) cells where an endogenous level of miR-615-5p was relatively higher (Fig. 1b), whereas re-expression of miR-615-5p can significantly suppress the proliferation in HLE cells where a basal level of miR-615-5p was comparatively higher (Fig. 1e), suggesting that miR-615-5p can prevent the growth of HCC cells. Meanwhile, wound healing assay found that knockdown or overexpression of miR-615-5p can significantly promote or prevent the migration of HCC cells (Fig. 1f, g), confirming that miR-615-5p can prevent the migration of HCC cells.
miR-615-5p was markedly upregulated in HCC tissues and prevents both growth and migration of HCC cells. a miR-615-5p was significantly upregulated in HCC tissues (90 cases) as compared with paired normal control tissues (90 cases), as exemplified by qRT-PCR. b The basal expression status of miR-615-5p in HCC cell lines (HepG2, Hep3B, Huh1, Huh7, and HLE) as well as normal human liver epithelial cell line (THLE3), detected by qRT-PCR. c In Huh7 cells where the basal level of miR-615-5p was comparatively higher, the proliferative variation of Huh7 was evaluated after transfection with miR-615-5p as well as siRNA scramble sequence. d Similarly, as did in Huh7, proliferative variation of HepG2 where the basal level of miR-615-5p was also comparatively higher was assayed after transfection with miR-615-5p as well as siRNA scramble sequence. e In parallel, the proliferation of HLE was detected using the MTT assay after being artificially upregulated. f The qualitative assay of migratory variations of Huh7, HepG2 cells transfected with miR-615-5p scramble and inhibitor, and HLE cells transfected with miR-615-5p scramble and mimics. g The quantitative assay of wound healing of the HCC cell line Huh7, HepG2 cells transfected with miR-615-5p inhibitor, and HLE cells transfected with miR-615-5p mimics. All the experiments were done independently in triplicate, similar results were obtained, and representative figures are shown here. Independent sample t test was employed to analyze the statistical significance between the two groups, and one-way ANOVA was used to compare the statistical significance among the three groups. *p < 0.05; **p < 0.01; ***p < 0.001, in comparison with the control group
SHMT2 was directly and negatively regulated by miR-615-5p
To understand how miR-615-5p promoted both growth and migration in HCC cells, we found using two different online bioinformatics prediction tools (www.microrna.org [9] and www.targetscan.com [10]) that SHMT2 can be a downstream target of miR-615-5p (Fig. 2a), followed by verification using the luciferase reporter assay (Fig. 2b). To make clear the endogenous expression level of SHMT2 in the panel of HCC cell lines involved, western blot was employed (Fig. 2c). To confirm the regulatory relationship between miR-615-5p and SHMT2, the expression variation of SHMT2 was detected after ectopic expression of miR-615-5p in HepG2, Huh7, and HLE whose basal level of miR-615-5p was starkly in contrast (Fig. 2d–f). It can be seen that miR-615-5p was able to directly and negatively regulate SHMT2 expression in HCC cells, indicating that SHMT2 was a direct and negative downstream target of miR-615-5p.
SHMT2 was directly and negatively regulated by miR-615-5p. a Bioinformatics prediction that SHMT2 was a putative downstream target of miR-615-5p, using online software (www.microrna.org and www.targetscan) independently and respectively. b Luciferase reporter assay was used to confirm that SHMT2 was the downstream target of miR-615-5p. c The basal expression status of SHMT2 in HCC cell lines HepG2, Hep3B, Huh1, Huh7, and HLE as well as normal human liver epithelial cell line THLE3, detected by western blotting. The molecular weight of SHMT2 was indicated at 55 kDa according to the instruction of primary antibody against SHMT2. β-Actin was 46 kDa. Fifty micrograms of the total protein was loaded per lane, and the exposure time was 60 s. d Expression variation of SHMT2 in HepG2 cells after downregulation of miR-615-5p by transfection with miR-615-5p inhibitor as compared with control. e Similarly, as did in HepG2 cells, the expression variation of SHMT2 in Huh7 was detected after being transfected with miR-615-5p inhibitor. f In parallel, the expression variation of SHMT2 in HLE was detected after being overexpressed of miR-615-5p. All the experiments were done independently in triplicate, similar results were obtained, and representative figures are shown here. Independent sample t test was employed to analyze the statistical significance between the two groups, and one-way ANOVA was used to compare the statistical significance among the three groups. *p < 0.05; **p < 0.01; ***p < 0.001, in comparison with the control group
SHMT2 can promote proliferation and migration in HCC cells
To investigate the biological role of SHMT2 in HCC cells, both loss-of-function and gain-of-function strategies were employed. First of all, on the basis of successful knockdown of SHMT2 on mRNA (Supplementary Fig. 3) and protein level (Fig. 3a, b), both MTT and wound healing assay were carried out to explore the biological role of SHMT2 in the proliferation and migration of HCC cells. MTT assay showed that knockdown of SHMT2 was markedly capable of preventing (Fig. 3c, d) the proliferation. Wound healing assay showed that knockdown of SHMT2 can significantly inhibit (Fig. 3e–h) the migratory ability, suggesting that SHMT2 was able to promote both proliferation and migration in HCC cells. To further validate whether or not miR-615-5p and SHMT2 were in the same axis, we performed the rescue experiment after knockdown of SHMT2 with miR-615-5p inhibitor in HLE and Huh1 cells. It can be seen that transfection of miR-615-5p inhibitor can entirely rescue the phenotype after knockdown of SHMT2, in terms of both expression and proliferation (Supplementary Fig. 4).
SHMT2 can promote proliferation and migration in HCC cells. a SHMT2 was knocked down by transfection with siRNAs against SHMT2 and siRNA scramble, as negative control, in HLE cells. b Similarly, SHMT2 was knocked down using siRNAs against SHMT2 and siRNA scramble in Huh1 cells. c On the basis of successful knockdown of SHMT2, proliferative ability was evaluated using the MTT assay in HLE cells. d As did in HLE cells, proliferation of Huh1 was assayed using MTT methods. e On the ground of successful knockdown of SHMT2 in HLE, migratory variations of HLE were detected using the wound healing assay. f As did in HLE, the migration variation of Huh1 was assayed with the wound healing assay in the absence of serum at 0 and 48 h, respectively. g Quantitative assay of wound healing experiment of HLE cells. h Quantitative assay of wound healing experiment of Huh1 cells. All the experiments were done independently in triplicate, similar results were obtained, and representative figures are shown here. Independent sample t test was employed to analyze the statistical significance between the two groups, and one-way ANOVA was used to compare the statistical significance among the three groups. *p < 0.05; **p < 0.01; ***p < 0.001, in comparison with the control group
SHMT2 was significantly overexpressed in HCC tissues
To understand the expression status of SHMT2 in HCC tissues, IHC was conducted with HCC tissue microarray. It was discovered that SHMT2 was significantly upregulated in HCC tissues (Fig. 4) in comparison with paired normal control tissues (Supplementary Fig. 3). Next, to make clear the clinicopathological significance of SHMT2 expression in HCC, we have analyzed the statistical association between SHMT2 expression and clinicopathological parameters. It was found that overexpression of SHMT2 was significantly associated with metastasis (Table 1).
SHMT2 was significantly upregulated in HCC tissues. Immunohistochemistry of SHMT2 was carried out with primary antibody (TA327002; OriGene, USA) against SHMT2 on HCC tissue microarray. The antibody was diluted at 1:200 following the recommendation of the manufacturer. The positive immunostaining of SHMT2 was mainly sublocalized both cytoplasmically and membraneously. The upper panel represents the macroscopic view of HCC tissue microarray whose magnification was 100-fold; the lower panel was the magnified at 400-fold where the red boxed in the upper panel
SHMT2 was significantly associated with poor prognosis
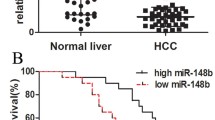
To analyze the association between SHMT2 expression and overall prognosis, Kaplan-Meier survival analysis was performed. It was found that overexpression of SHMT2 was remarkably associated with overall poor prognosis (Fig. 5a), suggesting that SHMT2 could be used as a potential prognostic biomarker for patients diagnosed with HCC. The Kaplan-Meier survival curves for the progression-free survival (PFS) showed that the SHMT2-positive patients tended to have a poorer PFS than the SHMT2-negative patients, although the difference between the two groups was not significant. No difference in PFS was observed between the two groups (Fig. 5b).
SHMT2 was significantly associated with overall poor prognosis of patients diagnosed with HCC. a Kaplan-Meier survival curve was conducted to statistically analyze the difference of overall prognosis between patients with high expression of SHMT2 and low expression of SHMT2. b Kaplan-Meier survival curve was performed to statistically analyze the difference of progression-free survival between patients with high expression of SHMT2 and low expression of SHMT2. Log rank test was employed. p < 0.05 was taken as statistically significant for the difference
Discussion
In our present study, we for the first time found that it is through targeting SHMT2 that miR-615-5p prevents both proliferation and migration in HCC cells and that overexpression of SHMT2 was not only able to promote both growth and migration in HCC cells but also was significantly associated with metastasis and poor prognosis in HCC tissues, suggesting that SHMT2 could be used as a potential prognostic biomarker in HCC.
The original report regarding miR-615 in the setting of malignant tumor came from prostate cancer [11], in which miR-615 was mentioned and found to be epigenetically activated in prostate cancer cells. Later, El Tayebi HM and colleagues for the first time detected and found that despite miR-615-5p was overexpressed both in HCC cell lines and tissues compared with corresponding normal controls where miR-615-5p was hardly detectable, actually, it can prevent both the growth and migration of HCC cells in vitro [6], indicating its tumor suppressor roles in HCC. What is consistent with the role of miR-615-5p in HCC is that miR-615-5p was discovered to be epigenetically inactivated and plays a role as a tumor suppressor in pancreatic ductal adenocarcinoma through negative regulation of insulin-like growth factor 2 (IGF2) [7], whereas Jiang Y et al. found that miR-615-5p can be negatively regulated by the CDX2 gene in pancreatic ductal adenocarcinoma [12]. Meanwhile, in another similar study in pancreatic ductal adenocarcinoma, miR-615-5p was confirmed as a tumor suppressor in the proliferation, migration, and invasion through negative regulation of the AKT2 gene [13]. Besides, the miR-615-5p/IGF2 axis was found to be negatively regulated by PU.1 in HCC [8]. In the present study, first of all, we have confirmed the expression status of miR-615-5p using a qRT-PCR technique and found that miR-615-5p was significantly upregulated in HCC tissues and cell lines, which was entirely supported by and in agreement with a previous report by El Tayebi HM et al., in HCC [6]. In terms of the expression status of miR-615-5p in HCC tissues compared with normal control, both our study and that of El Tayebi HM et al. [6] were entirely in contrast with that in pancreatic ductal adenocarcinoma [7, 13] where miR-615-5p was detected as significantly overexpressed in normal tissues in comparison with cancer tissues. Furthermore, in in vitro HCC cell lines, we for the first time found that SHMT2 was a novel downstream target that was able to be directly and negatively regulated by miR-615-5p, which was distinctively different from previous reports. Given the inconsistency between the expression level of miR-615-5p and its reported role as a tumor suppressor in HCC, there may be two possibilities from both two aspects. First of all, the inconsistency may be associated with the way the miR-615-5p was evaluated by qRT-PCR on the messenger RNA level. The total RNA was extracted using TRIzol reagent on the ground of total block of clinical tissue, which is to say that the whole block of clinical tissue to be detected histologically includes various types of mixture of cells, which could seriously contaminate the tumor cells where miR-615-5p was. Considering that the spatial-temporal specificity of microRNA (miRNA) expression and function, no wonder that miRNA that was found to play a tumor-suppressing role was unexpectedly observed to be upregulated in tumor tissues, whereas miRNA that was supposed to be upregulated in tumor tissues owing to its observed tumor-promoting role, actually, was found to be downregulated in tumor tissues compared with paired normal control tissues; therefore, given the disadvantage of qRT-PCR in the evaluation of miRNA, in situ hybridization approach may be more appropriate and complementary to qRT-PCR that is regardless of specific tissue architecture; secondly, in addition to the technical aspect, the gene miR-615-5p as such deserves consideration. It is well established that any tumor-suppressing gene whose expression level could be remarkably elevated in tumor tissues after mutation takes place. As a classical example, the elevated tumor suppressor gene p53, also aliased for TP53, in cancerous tissues was usually taking the form of mutational TP53 rather than wild-type TP53. The possibility might also apply to miR-615-5p. We cannot rule out the possibility that mutation takes place in miR-615-5p we have detected, and moreover, we are not sure whether the qRT-PCR amplified in our study is wild type or mutational, which left to be confirmed by DNA sequencing. Given above, both the two aspects could potentially lead to the phenomenon that miR-615-5p, despite a suppressing tumor, was found to be upregulated in HCC tissues.
SHMT2, abbreviated for serine hydromethyltransferase 2, was discovered to be in charge of catalyzing the conversion of serine to glycine with the transfer of β-carbon from serine to tetrahydrofolate (THF) to form 5,10-methylene-THF [14]. Suppression of SHMT2 was shown to be able to block cancer cell proliferation [15]. In the setting of malignancies, experimental evidence regarding SHMT2 has been limited and scarcely reported in cancers, let alone in the HCC. SHMT2 was first mentioned and reported to be a potential downstream target of miR-193 in breast cancer [16]. In another study performed in breast cancer, SHMT2 was mentioned as a potential prognostic indicator for patients diagnosed with breast cancer [17], which was wholly consistent with and supported by the finding of Lee GY et al. [18] that elevated expression of SHMT2 was shown to be associated with poor prognosis in human cancer, supporting the important role of SHMT2 in the control of cell proliferation in several cancer types [19, 20] and as a hot target for anticancer therapy. Until now, there has been no relevant report whatsoever concerning SHMT2 in HCC. In our present study, we for the first time identified that SHMT2 was a novel downstream target that was able to be directly and negatively regulated by miR-615-5p. Besides, we have also firstly found that SHMT2 was significantly upregulated in HCC tissues and, clinicopathologically, whose overexpression was remarkably associated with metastasis and poor prognosis, suggesting that SHMT2 could be used as an unfavorable prognostic biomarker in HCC. In addition, SHMT2 was found to be able to promote both the proliferation and migration in HCC cells in our study, which was fully in line with earlier relevant published reports [18–20].
Despite our study for the first time identified that miR-615-5p directly and negatively regulated SHMT2 in HCC, there are still several limitations that have to be acknowledged. First of all, both the clinical sample sizes of HCC tissues and cell lines were limited and may therefore lead to potentially biased and insufficient conclusions [21, 22]; secondly, given that the specificity of primary antibody that should have been evaluated and tested prior to being used [23, 24] otherwise may contribute to biased even wrong final results, however, we failed to test and evaluate before the experimentation; thirdly, given that prevalent cross-contamination or variation existed in in vitro cancer cell lines that could lead to irreproducibility [25, 26], all the cancer cell lines had better should have been sequenced to make genetic purity; however, we also failed to have all the HCC cell lines we have used sequenced using a short tandem repeat (STR) method [27].
In conclusion, in our study, we for the first time found that it is direct and negative targeting SHMT2 that miR-615-5p prevents both proliferation and migration in HCC and that SHMT2 expression was significantly associated with unfavorable prognosis and tumor metastasis in HCC tissue level and promotes both proliferation and migration of HCC cell in vitro.
References
Galun D, Basaric D, Zuvela M, Bulajic P, Bogdanovic A, Bidzic N, et al. Hepatocellular carcinoma: from clinical practice to evidence-based treatment protocols. World J Hepatol. 2015;7(20):2274–91.
Mazzola A, Costantino A, Petta S, Bartolotta TV, Raineri M, Sacco R, Brancatelli G, Camma C, Cabibbo G. Recurrence of hepatocellular carcinoma after liver transplantation: an update. Future Oncol. 2015.
Palmer DH, Johnson PJ. Evaluating the role of treatment-related toxicities in the challenges facing targeted therapies for advanced hepatocellular carcinoma. Cancer Metastasis Rev. 2015;34(3):497–509.
Mao B, Wang G. MicroRNAs involved with hepatocellular carcinoma (review). Oncol Rep. 2015.
Giordano S, Columbano A. MicroRNAs: new tools for diagnosis, prognosis, and therapy in hepatocellular carcinoma? Hepatology. 2013;57(2):840–7.
El Tayebi HM, Hosny KA, Esmat G, Breuhahn K, Abdelaziz AI. miR-615-5p is restrictedly expressed in cirrhotic and cancerous liver tissues and its overexpression alleviates the tumorigenic effects in hepatocellular carcinoma. FEBS Lett. 2012;586(19):3309–16.
Gao W, Gu Y, Li Z, Cai H, Peng Q, Tu M, et al. miR-615-5p is epigenetically inactivated and functions as a tumor suppressor in pancreatic ductal adenocarcinoma. Oncogene. 2015;34(13):1629–40.
Song LJ, Zhang WJ, Chang ZW, Pan YF, Zong H, Fan QX, et al. PU.1 is identified as a novel metastasis suppressor in hepatocellular carcinoma regulating the miR-615-5p/IGF2 axis. Asian Pac J Cancer Prev: APJCP. 2015;16(9):3667–71.
Betel D, Wilson M, Gabow A, Marks DS, Sander C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 2008;36(Database issue):D149–153.
Welten SM, Bastiaansen AJ, de Jong RC, de Vries MR, Peters EA, Boonstra MC, et al. Inhibition of 14q32 microRNAs miR-329, miR-487b, miR-494, and miR-495 increases neovascularization and blood flow recovery after ischemia. Circ Res. 2014;115(8):696–708.
Hulf T, Sibbritt T, Wiklund ED, Bert S, Strbenac D, Statham AL, et al. Discovery pipeline for epigenetically deregulated miRNAs in cancer: integration of primary miRNA transcription. BMC Genomics. 2011;12:54.
Jiang Y, Zhang Y, Li F, Du X, Zhang J. CDX2 inhibits pancreatic adenocarcinoma cell proliferation via promoting tumor suppressor miR-615-5p. Tumour Biol: J Int Soc Oncodev Biol Med. 2015.
Sun Y, Zhang T, Wang C, Jin X, Jia C, Yu S, et al. MiRNA-615-5p functions as a tumor suppressor in pancreatic ductal adenocarcinoma by targeting AKT2. PLoS One. 2015;10(4):e0119783.
Hebbring SJ, Chai Y, Ji Y, Abo RP, Jenkins GD, Fridley B, et al. Serine hydroxymethyltransferase 1 and 2: gene sequence variation and functional genomic characterization. J Neurochem. 2012;120(6):881–90.
di Salvo ML, Contestabile R, Paiardini A, Maras B. Glycine consumption and mitochondrial serine hydroxymethyltransferase in cancer cells: the heme connection. Med Hypotheses. 2013;80(5):633–6.
Leivonen SK, Rokka A, Ostling P, Kohonen P, Corthals GL, Kallioniemi O, et al. Identification of miR-193b targets in breast cancer cells and systems biological analysis of their functional impact. Mol Cellular Proteomics. 2011;10(7):M110 005322.
Antonov A, Agostini M, Morello M, Minieri M, Melino G, Amelio I. Bioinformatics analysis of the serine and glycine pathway in cancer cells. Oncotarget. 2014;5(22):11004–13.
Lee GY, Haverty PM, Li L, Kljavin NM, Bourgon R, Lee J, et al. Comparative oncogenomics identifies PSMB4 and SHMT2 as potential cancer driver genes. Cancer Res. 2014;74(11):3114–26.
Kim D, Fiske BP, Birsoy K, Freinkman E, Kami K, Possemato RL, et al. SHMT2 drives glioma cell survival in ischaemia but imposes a dependence on glycine clearance. Nature. 2015;520(7547):363–7.
Ye J, Fan J, Venneti S, Wan YW, Pawel BR, Zhang J, et al. Serine catabolism regulates mitochondrial redox control during hypoxia. Cancer Dis. 2014;4(12):1406–17.
Kaplan RM, Chambers DA, Glasgow RE. Big data and large sample size: a cautionary note on the potential for bias. Clin Trans Sci. 2014;7(4):342–6.
Bridges AJ, Holler KA. How many is enough? Determining optimal sample sizes for normative studies in pediatric neuropsychology. Child Neuropsychol. 2007;13(6):528–38.
Baker M. Reproducibility crisis: blame it on the antibodies. Nature. 2015;521(7552):274–6.
Holmseth S, Zhou Y, Follin-Arbelet VV, Lehre KP, Bergles DE, Danbolt NC. Specificity controls for immunocytochemistry: the antigen preadsorption test can lead to inaccurate assessment of antibody specificity. J Histochem Cytochem. 2012;60(3):174–87.
Jiang T, Wang H. Identification of cross-contamination in SH-SY5Y cell line. Hum Cell. 2014;27(4):176–8.
International Cell Line Authentication C. Cell line cross-contamination: WSU-CLL is a known derivative of REH and is unsuitable as a model for chronic lymphocytic leukaemia. Leuk Res. 2014;38(8):999–1001.
Dirks WG, MacLeod RA, Nakamura Y, Kohara A, Reid Y, Milch H, et al. Cell line cross-contamination initiative: an interactive reference database of STR profiles covering common cancer cell lines. Int J Cancer. 2010;126(1):303–4.
Acknowledgments
The present work was supported by the National Science Foundation of China (NFSC no. 81373990 and 81402523).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Xiaoyu Wu and Liang Deng contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 2469 kb)
Rights and permissions
About this article
Cite this article
Wu, X., Deng, L., Tang, D. et al. miR-615-5p prevents proliferation and migration through negatively regulating serine hydromethyltransferase 2 (SHMT2) in hepatocellular carcinoma. Tumor Biol. 37, 6813–6821 (2016). https://doi.org/10.1007/s13277-015-4506-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4506-8