Abstract
The cullin proteins are a family of scaffolding proteins that associate with RING proteins and ubiquitin E3 ligases and mediate substrate–receptor bindings. Thus, cullin proteins regulate the specificity of ubiquitin targeting in the regulation of proteins involved in various cellular processes, including proliferation, differentiation, and apoptosis. There are seven cullin proteins that have been identified in eukaryotes: CUL1, CUL2, CUL3, CUL4A, CUL4B, CUL5, and CUL7/p53-associated parkin-like cytoplasmic protein. All of these proteins contain a conserved cullin homology domain that binds to RING box proteins. Cullin–RING ubiquitin ligase complexes are activated upon post-translational modification by neural precursor cell-expressed, developmentally downregulated protein 8. The aberrant expression of several cullin proteins has been implicated in many cancers though the significance in gastric cancer has been less well investigated. This review provides the first systematic discussion of the associations between all members of the cullin protein family and gastric cancer. Functional and regulatory mechanisms of cullin proteins in gastric carcinoma progression are also summarized along with a discussion concerning future research areas. Accumulating evidence suggests a critical role of cullin proteins in tumorigenesis, and a better understanding of the function of these individual cullin proteins and their targets will help identify potential biomarkers and therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer (GC) is the fourth most common cancer and the third leading cause of cancer-related death worldwide [1]. The highest reported incidences of GC are in Eastern Asia, with China alone accounting for 42 % of all newly diagnosed cases [2]. A majority of patients with GC are diagnosed at advanced clinical stages with a high rate of lymph node metastasis [3]. Even with radical resection, the 5-year survival rate remains low, largely due to postoperative metastasis [4, 5].
Although the mechanisms underlying the development and progression of GC are not completely understood, numerous abnormal cellular processes have been implicated. One implicated mechanism involves the ubiquitin (Ub)–proteasome system, which regulates the function of numerous proteins and almost all cellular processes [6]. The Ub–proteasome system comprises a pathway of post-translational modifications that can potentially regulate the proliferation, invasion, and growth of cancer cells. At the same time, the pathway drives the degradation of anti-oncogenic proteins that also contribute to tumorigenesis.
The cullin protein family includes scaffolding proteins that mediate substrate–receptor bindings and determine the specificity of Ub targeting for the regulation of various cellular processes [7], including proliferation, differentiation, and apoptosis. Many cullin proteins have been implicated in various cancers, including breast, skin, and colorectal cancer and have therefore emerged as a hot spot in cancer research (Table 1). Nevertheless, the specific function of cullin proteins in GC has not been clarified. To the end, recent research monitoring the dynamics and specificity of cullin proteins will be reviewed in this article. Furthermore, potential mechanisms involving these cullin proteins in gastric tumorigenesis will be discussed. A better understanding of the functional and regulatory mechanisms of these proteins in gastric carcinoma progression may ultimately help identify more potent novel biomarkers for the diagnosis and treatment of GC.
Ub–proteasome system
The three-step Ub–proteasome pathway begins with the activation of Ub by the ATP-dependent E1 Ub-activating enzyme, which is then conjugated to an E2 Ub enzyme and attached to a target protein by an E3 Ub ligase in preparation for its degradation by a 26S proteasome (Fig. 1). Currently, two E1 enzymes have been identified in the human genome, as well as 30–40 E2 enzymes and more than 600 E3 ligases [8, 9]. Ubiquitination is a diverse, dynamic, reversible, and specific modification of targeted proteins [10]. Ub-binding domains, present in ~200 proteins, are responsible for recognizing these Ub signals [11, 12]. Monoubiquitination, or attachment of a single Ub to a target, is typically nonproteolytic and controls various cellular processes, such as receptor transport, endocytosis, gene expression, and DNA repair [13]. On the other hand, polyubiquitination has various effects depending on the type of linkage; Ub linkage via lysine (K)63 results in modification of the target protein function [14], whereas linkage of K11 or K48 residues initiates 26S proteasome degradation [9, 15, 16] (Fig. 1). The length and structure of the Ub chain also affect the interaction between the proteasome and substrate [17]. The highly specific degradation of target proteins is mainly determined by the largest family of Ub E3 ligases, the cullin–RING E3 ligases (CRLs), which regulate the proteolysis of 20 % of ubiquitinated proteins [18–20].
Ubiquitin–proteasome system. Ubiquitin (Ub) is activated by an ATP-dependent ubiquitin-activating enzyme (E1), then conjugated to Ub-conjugating enzyme (E2), and ligated to a substrate by the Ub-ligating enzyme (E3). A single Ub (monoubiquitination) or multiple types of Ub chains in different combinations (polyubiquitination) can be attached to the targeted substrate protein. Monoubiquitination typically has nonproteolytic functions, whereas polyubiquitination via lysine (K) 63-linked Ub chains can modify protein function and linkage of K11 or K48 residues can trigger degradation by the 26S proteasome
Cullin protein family
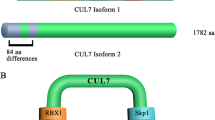
Cullins are core scaffold proteins for CRLs that utilize distinct adaptors to recruit substrates for degradation via substrate receptors at the N-terminus(NTD). At the C-terminus(CTD), the cullin proteins interact with the RING domain of RING box proteins (RBX or ROC), which recruit E2-conjugated Ub to the complex (Fig. 2) [21]. There are seven cullin proteins that have been identified in eukaryotes: CUL1, CUL2, CUL3, CUL4A, CUL4B, CUL5, and CUL7/p53-associated parkin-like cytoplasmic protein (PARC) [22, 23]. These proteins all contain a conserved cullin homology domain, which binds with RBX1/ROC1 [24]. CRLs are activated by post-translational neddylation by the Ub-like neuronal precursor cell-expressed, developmentally downregulated protein 8 [25].
Cullin–RING ubiquitin ligases. Cullin recruits substrates for degradation via adaptors and substrate receptors (SR) at its N-terminal domain (NTD), and interacts with the RING domain of RING box protein (RBX/ROC), which recruits E2-conjugated ubiquitin (Ub) at the C-terminal domain (CTD). The distance between the E2-conjugated ubiquitin and substrate is roughly 50 A ̊when the conformation is inactivated, and it is difficult for ubiquitylation. All cullin-based ubiquitin ligases are activated by neuronal precursor cell-expressed, developmentally downregulated protein 8 (NEDD8); post-translational neddylation reduces the distance between the E2-conjugated Ub and substrate to promote ubiquitination. MLN4924 is a small molecule inhibitor of NEDD8 that blocks neddylation and subsequently inactivates the cullin–RING ubiquitin ligase. Positive sign: promotes the neddylation; negative sign: inhibits the neddylation
Cul1
CUL1 was the first and most extensively characterized member of the cullin family. It primarily associates with RBX1, which binds with E2-conjugated Ub at the C-terminus, and S-phase kinase-associated protein (SKP) 1, which recruits substrates with F-box-containing substrate receptors [26, 27]. First implicated in skin cancer [28], CUL1 was recently shown to regulate the proliferation of colorectal cancer cells while promoting their migration and invasion [29]. By now, aberrant expression of CUL1 has been found in many types of cancer including GC which is closely associated with poor patient prognosis. Indeed, CUL1 expression has been implicated in GC via associations with F-box proteins. While the majority of the 69 F-box proteins identified in the human genome remain undefined [30], they are characterized by a 40-amino-acid F-box domain, which provides the linkage to SKP1 and CUL1 [31]. F-boxes are divided into three types: FBXW contains WD40 repeats, FBXL contains leucine-rich repeats, and FBXO contains various undefined repeats [32]. The main representation is Skp2, FBXW7, and FBX8. FBX8 is the novel component of the F-box family. Some of these function as an oncoprotein, whereas others function as a tumor suppressor.
Cul1 → FBXW7 → GC:
FBXW7 is transcriptionally controlled by p53, which plays a key role in the initiation of cell cycle arrest following the degradation of cyclin E, and regulates cell cycle metabolism via c-Myc degradation [33]. Notably, 6 % of the GC cases screened show allelic loss and mutation of FBXW7 [34]. Yokobori et al. [33] showed that mutation of p53 and low FBXW7 expression contributed to malignant potential in primary GC. In addition, the loss of cell division control protein 4/FBXW7 appears in both early-onset and conventional GCs [35], which may be mediated by the subsequent upregulation of c-Myc. For example, Calcagno et al. [36] found that mRNA and protein levels of c-Myc were increased, and FBXW7 was reduced in GC cell lines. These data suggest that abnormal regulation of c-Myc and FBXW7 influences the invasive ability of GC cells.
Cul1 → FBX8 → GC:
FBX8 is a novel component of the F-box family [37], which is downregulated in GC tissues and associated with invasion, metastasis, and poor survival time in GC patients [38]. Cellular invasion may be promoted by interaction with c-Myc [37] or regulation of ADP-ribosylation factor 6 [39].
Cul1 → SKP2 → GC:
SKP2 is necessary for the regulation of cyclins (cyclins D1 and E) and cyclin-dependent kinase (CDK) inhibitors (p27 and p21), which drive cells from the G1 to the S phase of the cell division cycle [40]. Inactivation of SKP2 has been shown to weaken or eliminate the oncogenic effect from the loss of p53, retinoblastoma, and phosphatase and tensin homolog tumor suppressor proteins [41–43]. A previous study showed that Skp2 overexpression modulates the malignant phenotype of gastric carcinoma probably through p27 proteolysis [44]. Furthermore, SKP2 depletion markedly suppresses the growth and metastasis of gastric carcinomas by inducing cell cycle arrest via upregulation of p27, p21, and p57 and downregulation of cyclin E and CDK2 [44].
Bai et al. [45] reported that knockdown CUL1 in GC cells markedly reduced cell proliferation and arrested the cell cycle in the G1 phase by inhibiting the expression of cyclins, possibly the result of p27 accumulation. In contrast, the overexpression of CUL1 promoted cell invasion and metastasis via activation of the Src/focal adhesion kinase pathway, a known target for GC progression and prognosis [45–47].
Cul1 → FBX6 → GC:
FBX6 is a crucial component of the Ub metabolic system and is closely related to the cell cyclins [48, 49]. Although the mechanism is unknown, high levels of FBX6 influence the cell cycle of GC cells and promote their development and proliferation, while negatively controlling apoptosis and invasion [50].
Cul1 → FBXL5 → GC:
FBXL5 is known to target various substrates for proteasomal degradation and regulate several cellular processes [51–53]. A study by Cen et al. [54] demonstrated that FBXL5 is involved in GC cell migration via degradation of cortactin. Cortactin is a substrate of Src kinase, which can also regulate cancer cell migration and invasion [55]. Indeed, a phosphomimetic (S405A/S418A) nondegradable mutant form of cortactin was protected from FBXL5-induced degradation, resulting in enhanced GC cell migration [54].
Cul1 → RBX1 → GC:
RBX1 is involved in cancer development via the regulation of transcription factors, signal transducers, cell cycle regulators, and oncogenes/tumor suppressors [56–58]. Migita et al. [56] showed that RBX1 is overexpressed in GC tissues and correlates with a higher risk of recurrence and postoperative overall survival. They also showed that there is a significant correlation between the level of CUL1 and GC differentiation. In addition, a target of RBX1, microRNA-194, was recently shown to be downregulated in GC, and the expressions of both were associated with GC patient survival time [59]. These data indicate that RBX1 may act as an oncogene by modulating the proliferation and migration of GC cells via miR-194.
CDK-associated CUL1 (CAC1) → p53 → GC
CAC1 is a novel member of the cullin family that promotes GC cell proliferation and attenuates anti-cancer effects [60]. In their study, Kong et al. [60] showed that GC cell lines have higher expression of CAC1 compared to control gastric epithelial mucosa cells. The same study revealed that reducing the expression of CAC1 attenuates CDK2 activity and interrupts the G1–S cell cycle phase transition. Another study showed that inactivation of CAC1 results in excessive accumulation of p53 and BAX and downregulates BCL2 levels, thus enhancing cisplatin-induced apoptosis [61].
Cul2/5
CUL2 interacts with RBX1 as a substrate receptor for the von Hippel–Lindau tumor suppressor and targets hypoxia-inducible factor 1α in common human cancers [62]. CUL5 interacts with RBX2 to bind to suppressor of cytokine signaling (SOCS) box proteins, as well as with a virion infectivity factor [63, 64]. Together, CUL2 and CUL5 assemble into a complex using elongins B and C as adaptor proteins [23, 65].
Cul2 → miR-574-3p → GC:
CUL2 is required for normal vasculogenesis and involved in cell cycle regulation. A microRNA, miR-574-3p, is shown to be decreased in the early stages or differentiation of GC [66]. CUL2 expression is downregulated in GC cells transfected with miR-574-3p that show decreased proliferation, invasion, and migration. The levels of CUL2 in GC may, in part, be mediated by miR-574-3p. Furthermore, it suggests that the dynamic changes in the expression levels of CUL2 may reflect different requirement for CUL2 across GC at different development stages or differentiation [67].
Cul5 → SOCS → GC
CUL5 may be involved in GC due to its ability to bind SOCS proteins that regulate the Ub-mediated degradation of proteins [68]. SOCS6 negatively regulates cell growth and promotes apoptosis, and a study by Lai et al. [69] showed that it is downregulated in GC due to loss of heterozygosity and promoter hypermethylation. A recent study found that SOCS1 and SOCS3 are also downregulated in GC and contribute to a poor prognosis [70]. It was postulated that the downregulation of these proteins might impact proinflammatory cytokine signaling. Further studies are needed to identify the precise role of CUL5 in regulating these effects.
Cul3
Cul3 → SPOP → Hedgehog/Gli2 → GC
Unlike the other family members, CUL3 can directly bind to BTB-domain-containing proteins, such as speckle-type POZ protein (SPOP), and integrate the functions of both adaptors and substrate receptors [71–73]. The Hedgehog (Hh) signaling pathway is crucial for the growth and patterning of various tissues during embryonic development. This pathway is a highly coordinated network of Hh proteins and transcription factor Gli2. It has been found that significant level of sonic Hh is expressed in gastric mucosa. SPOP promotes apoptosis of GC cells via inhibition of the Hh/Gli2 pathway, which may relate to the HIB/CUL3 complex [74–76]. In contrast, the proliferation and migration of human GC cells are enhanced by downregulation of SPOP [77].
Cul4
CUL4 has two highly homologous members, CUL4A and CUL4B, which target proteins regulating many processes, such as cellular progression, DNA damage response, DNA replication control, and chromatin remodeling, for degradation [78–80]. CUL4A is highly expressed in many cancers and is associated with their development and patient prognosis via regulation of p21, p27, p53, P-glycoprotein, damage-specific DNA-binding protein (DDB)2, and zinc finger E-box binding homeobox 1 [81–86], and CUL4B induces polyubiquitination of p53 [87]. CUL4A initiates DNA repair by interacting with DDB1 to recruit substrates for ubiquitination and stimulate multiple regulators of the cell cycle and DNA replication pathways [88]. However, the role of CUL4 in GC has not been studied extensively and remains unknown.
Cul4A → MDM2/COP1 → p53 → GC:
CUL4A with DDB1 is involved in regulating the stability of p53 and murine double minute 2 (MDM2) [89], which is in turn considered an oncoprotein because it mediates the transcriptional transactivation and polyubiquitination of p53 [90, 91]. Furthermore, it was found that the T309G MDM2 polymorphism is associated with an increased susceptibility to GC [92]. Similar to MDM2, constitutive photomorphogenic 1 (COP1) is an oncoprotein-targeting E3 Ub ligases containing RING finger, coiled-coil, and WD40-repeat domains [90, 93]. Elevated levels of COP1 are closely related to the overall survival and prognosis of GC patients, possibly as a result of p53 degradation [93]. However, Sawada et al. [94] found low COP1 expression in the tumor tissues of patients with a poor prognosis, and in vitro experiments indicated that this may be due to altered c-Jun-mediated regulation of matrix metalloproteinases.
Cul7/PARC
Like CUL1, CUL7 complexes include SKP1, FBWL8, and RBX1. PARC is highly homologous to CUL7 and negatively regulates p53 by cytoplasmic sequestration [95, 96]. Inactivation of the CUL7/FBXW8 complex leads to increased levels of insulin receptor substrate 1 and subsequent suppression of cell growth [97]. CUL7 has been implicated in 3M syndrome [95] though its role in GC needs further research.
Perspectives
Cullin proteins function like oncogenes or suppressors in gastric carcinogenesis mediated by their adaptors and substrates, so they may serve as attractive targets for the treatment of GC. Indeed, studies have investigated the effects of interfering with the activation of cullin proteins as a possible therapeutic approach. For example, MLN4924 is a type of proteasome inhibitor that inhibits NEDD8-activating enzyme, the enzyme responsible for CRL neddylation. This inhibitor induced apoptosis of cancer cells and demonstrated anti-proliferative, anti-migratory, and anti-invasive effects in in vitro and in vivo xenograft models [98]. Furthermore, it has been used in phase I clinical trials for the treatment of several human malignancies [98, 99]. Attempts to find additional agents are hindered by the wide variety of substrates and receptors that comprise CRL complexes. It is important to identify first which of these are related to GC in order to target them or use them as potential biomarkers for early detection and prognosis prediction.
There are still many important questions that need to be addressed. First, though in vitro studies are important for revealing potential molecular mechanisms, they are restricted to evaluating limited substrates involved in GC. Second, very few of the clinical trials include the early stages of GC, making it difficult to identify targets for prevention and diagnosis of GC. Another important issue involves the diversity of the cullin proteins. For example, although CUL1 and CUL4 have similar structures, the mechanisms involved in the degradation of some of their common substrates, such as CDT1/2, differ [100–102]. Thus, more extensive research may be needed when evaluating the causal role of other cullin proteins in gastric carcinogenesis, such as CUL4A and CUL4B. Nevertheless, further studies are required to define the mechanisms by which cullin proteins are involved in the tumorigenesis and oncogenesis of GC.
Conclusion
The review presents the first systematic description of the associations between all members of the cullin protein family and GC. This family of scaffolding proteins associates with Ub E3 ligases and marks highly specific target proteins for degradation. Many of the target proteins include cell cycle regulators, tumor suppressors, and oncogenes, resulting in a diversity of processes that can promote tumorigenesis. Although many studies have implicated various members of the cullin family of proteins in GC, the majority of studies have focused on CUL1, which mediates GC development and tumorigenesis with FBOX, FBXW7, and FBX8 functioning as tumor suppressors, and FBX6, FBXL5, and SKP2 identified as oncoproteins (Table 2). CUL2/5 interacts with SOCS proteins to act as suppressors, and CUL3 interacts with BTB domains to act as an oncoprotein. However, there are few related studies focusing on CUL4B and CUL7. Further studies are needed to clarify the roles of each of the cullin proteins to provide a better framework for the development of biomarkers and therapeutic targets. As Cullin proteins play different roles in gastric carcinogenesis mediated by their adaptors and substrates, more studies should be focused on the functions of these individual cullin proteins and their targets. Given the fact that cancer occurs in the gastric surface and the critical role of cullin proteins in tumorigenesis, better understanding of the functional and regulatory mechanisms of cullin proteins in gastric carcinoma progression will help identify potential biomarkers for early detection and prognosis prediction. Meanwhile, it also can help to facilitate the development of novel therapeutic targets and proteasome inhibitors to target cullin proteins in GC.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108. doi:10.3322/caac.21262.
Lin Y, Ueda J, Kikuchi S, Totsuka Y, Wei WQ, Qiao YL, et al. Comparative epidemiology of gastric cancer between Japan and China. World J Gastroenterol. 2011;17:4421–8. doi:10.3748/wjg.v17.i39.4421.
Fang WL, Huang KH, Lan YT, Chen MH, Chao Y, Lo SS, et al. The risk factors of lymph node metastasis in early gastric cancer. Pathol Onocl Res. 2015. doi:10.1007/s12253-015-9920-0.
Coburn NG, Lourenco LG, Rossi SE, Gunraj N, Mahar AL, Helyer LK, et al. Management of gastric cancer in Ontario. J Surg Oncol. 2010;102:54–63. doi:10.1002/jso.21561.
Chen Y, Awan N, Haveman JW, Apostolou C, Chang DK, Merrett ND. Gastric cancer: Australian outcomes of multi-modality treatment with curative intent. ANZ J Surg. 2014. doi:10.1111/ans.12693.
Voutsadakis IA. The ubiquitin-proteasome system and signal transduction pathways regulating epithelial mesenchymal transition of cancer. J Biomed Sci. 2012;19:67. doi:10.1186/1423-0127-19-67.
Liu J, Nussinov R. The mechanism of ubiquitination in the cullin-RING E3 ligase machinery: conformational control of substrate orientation. PLoS Comput Biol. 2009;5, e1000527. doi:10.1371/journal.pcbi.1000527.
Qi J, Kim H, Scortegagna M, Ronai ZA. Regulators and effectors of Siah ubiquitin ligases. Cell Biochem Biophys. 2013;67:15–24. doi:10.1007/s12013-013-9636-2.
Chen ZJ, Sun LJ. Nonproteolytic functions of ubiquitin in cell signaling. Mol Cell Biol. 2009;33:275–86. doi:10.1016/j.molcel.2009.01.014.
Ikeda F, Dikic I. Atypical ubiquitin chains: new molecular signals. ‘protein modifications: beyond the usual suspects’ review series. EMBO Rep. 2008;9:536–42. doi:10.1038/embor.2008.93.
Dikic I, Wakatsuki S, Walters KJ. Ubiquitin-binding domains—from structures to functions. Nat Rev Mol Cell Biol. 2009;10:659–71. doi:10.1038/nrm2767.
Ikeda F, Crosetto N, Dikic I. What determines the specificity and outcomes of ubiquitin signaling? Cell. 2010;143:677–81. doi:10.1016/j.cell.2010.10.026.
Sadowski M, Suryadinata R, Tan AR, Roesley SN, Sarcevic B. Protein monoubiquitination and polyubiquitination generate structural diversity to control distinct biological processes. IUBMB Life. 2012;64:136–42. doi:10.1002/iub.589.
Sekiyama N, Jee J, Isogai S, Akagi K, Huang TH, Ariyoshi M, et al. NMR analysis of Lys63-linked polyubiquitin recognition by the tandem ubiquitin-interacting motifs of Rap80. J Biomol NMR. 2012;52:339–50. doi:10.1007/s10858-012-9614-9.
Hofmann K. Ubiquitin-binding domains and their role in the DNA damage response. DNA Repair (Amst). 2009;8:544–56. doi:10.1016/j.dnarep.2009.01.003.
Jin L, Williamson A, Banerjee S, Philipp I, Rape M. Mechanism of ubiquitin-chain formation by the human anaphase-promoting complex. Cell. 2008;133:653–65. doi:10.1016/j.cell.2008.04.012.
Kravtsova-Ivantsiv Y, Sommer T, Ciechanover A. The lysine48-based polyubiquitin chain proteasomal signal: not a single child anymore. Angew Chem Int Ed Engl. 2013;52:192–8. doi:10.1002/anie.201205656.
Lee EK, Diehl JA. SCFs in the new millennium. Oncogene. 2014;33:2011–8. doi:10.1038/onc.2013.144].
Komander D. The emerging complexity of protein ubiquitination. Biochem Soc Trans. 2009;37(Pt 5):937–53. doi:10.1042/BST0370937].
Pan Y, Xu H, Liu R, Jia L. Induction of cell senescence by targeting to Cullin-RING Ligases (CRLs) for effective cancer therapy. Int J Biochem Mol Biol. 2012;3:273–81.
Lu A, Pfeffer SR. A CULLINary ride across the secretory pathway: more than just secretion. Trends Cell Biol. 2014;24(7):389–99. doi:10.1016/j.tcb2014.02.011.
Zimmerman ES, Schulman BA, Zheng N. Structural assembly of cullin-RING ubiquitin ligase complexes. Curr Opin Struct Biol. 2010;20:714–21. doi:10.1016/j.sbi.2010.08.010.
Lee J, Zhou P. Cullins and cancer. Genes Cancer. 2010;1:690–9. doi:10.1177/1947601910382899.
Sarikas A, Hartmann T, Pan ZQ. The cullin protein family. Genome Biol. 2011;12:220. doi:10.1186/gb-2011-12-4-220.
Watson IR, Irwin MS, Ohh M. NEDD8 pathways in cancer, Sine Quibus Non. Cancer Cell. 2011;19:168–76. doi:10.1016/j.ccr.2011.01.002.
Petroski MD, Deshaies RJ. Function and regulation of cullin-RING ubiquitin ligases. Nat Rev Mol Cell Biol. 2005;6:9–20. doi:10.1038/nrm1547.
Zheng N, Schulman BA, Song L, Miller JJ, Jeffrey PD, Wang P, et al. Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature. 2002;416:703–9. doi:10.1038/416703a.
Xie CM, Wei W, Sun Y. Role of SKP1-CUL1-F-box-protein (SCF) E3 ubiquitin ligases in skin cancer. J Genet Genomics. 2013;40:97–106. doi:10.1016/j.jgg.2013.02.001.
Wang W, Chen Y, Deng J, Zhou J, Gu X, Tang Y, et al. Cullin1 is a novel prognostic marker and regulates the cell proliferation and metastasis in colorectal cancer. J Cancer Res Clin Oncol. 2015. doi:10.1007/s00432-015-1931-4.
Skaar JR, D’Angiolella V, Pagan JK, Pagano M. SnapShot: F box proteins II. Cell. 2009;137:1358. doi:10.1016/j.cell.2009.05.040. 1358 e1351.
Skaar JR, Pagan JK, Pagano M. Mechanisms and function of substrate recruitment by F-box proteins. Nat Rev Mol Cell Biol. 2013;14:369–81. doi:10.1038/nrm3582.
Wang Z, Liu P, Inuzuka H, Wei W. Roles of F-box proteins in cancer. Nat Rev Cancer. 2014;14:233–47. doi:10.1038/nrc3700].
Yokobori T, Mimori K, Iwatsuki M, Ishii H, Onoyama I, Fukagawa T, et al. p53-Altered FBXW7 expression determines poor prognosis in gastric cancer cases. Cancer Res. 2009;69:3788–94. doi:10.1158/0008-5472.CAN-08-2846].
Akhoondi S, Sun D, von der Lehr N, Apostolidou S, Klotz K, Maljukova A, et al. FBXW7/hCDC4 is a general tumor suppressor in human cancer. Cancer Res. 2007;67:9006–12. doi:10.1158/0008-5472.CAN-07-1320].
Milne AN, Leguit R, Corver WE, Morsink FH, Polak M, de Leng WW, et al. Loss of CDC4/FBXW7 in gastric carcinoma. Cell Oncol. 2010;32:347–59. doi:10.3233/CLO-2010-523].
Calcagno DQ, Freitas VM, Leal MF, de Souza CR, Demachki S, Montenegro R, et al. MYC, FBXW7 and TP53 copy number variation and expression in gastric cancer. BMC Gastroenterol. 2013;13:141. doi:10.1186/1471-230X-13-141.
Cho HJ, Oh YJ, Kwon J, Kwon JY, Kim KS, Kim H. c-Myc stimulates cell invasion by inhibiting FBX8 function. Mol Cells. 2010;30:355–62. doi:10.1007/s10059-010-0134-8.
Wu P, Wang F, Wang Y, Men H, Zhu X, He G, et al. Significance of FBX8 in progression of gastric cancer. Exp Mol Pathol. 2015;98:360–6. doi:10.1016/j.yexmp.2015.03.015.
Yano H, Kobayashi I, Onodera Y, Luton F, Franco M, Mazaki Y, et al. Fbx8 makes Arf6 refractory to function via ubiquitination. Mol Biol Cell. 2008;19:822–32. doi:10.1091/mbc.E07-08-0763.
Nakayama KI, Nakayama K. Ubiquitin ligases: cell-cycle control and cancer. Nat Rev Cancer. 2006;6:369–81. doi:10.1038/nrc1881.
Lin HK, Chen Z, Wang G, Nardella C, Lee SW, Chan CH, et al. Skp2 targeting suppresses tumorigenesis by Arf-p53-independent cellular senescence. Nature. 2010;464:374–9. doi:10.1038/nature08815.
Wang Z, Gao D, Fukushima H, Inuzuka H, Liu P, Wan L, et al. Skp2: a novel potential therapeutic target for prostate cancer. Biochim Biophys Acta. 1825;2012:11–7. doi:10.1016/j.bbcan.2011.09.002.
Wang H, Bauzon F, Ji P, Xu X, Sun D, Locker J, et al. Skp2 is required for survival of aberrantly proliferating Rb1-deficient cells and for tumorigenesis in Rb1+/− mice. Nat Genet. 2010;42:83–8. doi:10.1038/ng.498.
Wei Z, Jiang X, Liu F, Qiao H, Zhou B, Zhai B, et al. Downregulation of Skp2 inhibits the growth and metastasis of gastric cancer cells in vitro and in vivo. Tumour Biol. 2013;34:181–92. doi:10.1007/s13277-012-0527-8.
Bai J, Zhou Y, Chen G, Zeng J, Ding J, Tan Y, et al. Overexpression of Cullin1 is associated with poor prognosis of patients with gastric cancer. Hum Pathol. 2011;42:375–83. doi:10.1016/j.humpath.2010.09.003.
Humar B, Fukuzawa R, Blair V, Dunbier A, More H, Charlton A, et al. Destabilized adhesion in the gastric proliferative zone and c-Src kinase activation mark the development of early diffuse gastric cancer. Cancer Res. 2007;67:2480–9. doi:10.1158/0008-5472.CAN-06-3021.
Giaginis CT, Vgenopoulou S, Tsourouflis GS, Politi EN, Kouraklis GP, Theocharis SE. Expression and clinical significance of focal adhesion kinase in the two distinct histological types, intestinal and diffuse, of human gastric adenocarcinoma. Pathol Oncol Res. 2009;15:173–81. doi:10.1007/s12253-008-9120-2.
Ilyin GP, Rialland M, Pigeon C, Guguen-Guillouzo C. cDNA cloning and expression analysis of new members of the mammalian F-box protein family. Genomics. 2000;67:40–7. doi:10.1006/geno.2000.6211.
Zhang YW, Brognard J, Coughlin C, You Z, Dolled-Filhart M, Aslanian A, et al. The F box protein Fbx6 regulates Chk1 stability and cellular sensitivity to replication stress. Mol Cell. 2009;35:442–53. doi:10.1016/j.molcel.2009.06.030.
Zhang L, Hou Y, Wang M, Wu B, Li N. A study on the functions of ubiquitin metabolic system related gene FBG2 in gastric cancer cell line. J Exp Clin Cancer Res. 2009;28:78. doi:10.1186/1756-9966-28-78.
Moroishi T, Nishiyama M, Takeda Y, Iwai K, Nakayama KI. The FBXL5-IRP2 axis is integral to control of iron metabolism in vivo. Cell Metab. 2011;14:339–51. doi:10.1016/j.cmet.2011.07.011.
Vashisht AA, Zumbrennen KB, Huang X, Powers DN, Durazo A, Sun D, et al. Control of iron homeostasis by an iron-regulated ubiquitin ligase. Science. 2009;326:718–21. doi:10.1126/science.1176333.
Salahudeen AA, Thompson JW, Ruiz JC, Ma HW, Kinch LN, Li Q, et al. An E3 ligase possessing an iron-responsive hemerythrin domain is a regulator of iron homeostasis. Science. 2009;326:722–6. doi:10.1126/science.1176326.
Cen G, Ding HH, Liu B, Wu WD. FBXL5 targets cortactin for ubiquitination-mediated destruction to regulate gastric cancer cell migration. Tumour Biol. 2014;35:8633–8. doi:10.1007/s13277-014-2104-9.
MacGrath SM, Koleske AJ. Cortactin in cell migration and cancer at a glance. J Cell Sci. 2012;125:1621–6. doi:10.1242/Jcs.093781.
Migita K, Takayama T, Matsumoto S, Wakatsuki K, Tanaka T, Ito M, et al. Prognostic impact of RING box protein-1 (RBX1) expression in gastric cancer. Gastric Cancer. 2014;17:601–9. doi:10.1007/s10120-013-0318-y.
Ohta T, Michel JJ, Schottelius AJ, Xiong Y. ROC1, a homolog of APC11, represents a family of cullin partners with an associated ubiquitin ligase activity. Mol Cell. 1999;3:535–41.
Skowyra D, Koepp DM, Kamura T, Conrad MN, Conaway RC, Conaway JW, et al. Reconstitution of G1 cyclin ubiquitination with complexes containing SCFGrr1 and Rbx1. Science. 1999;284:662–5.
Chen X, Wang Y, Zang W, Du Y, Li M, Zhao G. miR-194 targets RBX1 gene to modulate proliferation and migration of gastric cancer cells. Tumour Biol. 2014;36:2393–401. doi:10.1007/s13277-014-2849-1.
Kong Y, Kejun N, Yin Y. Identification and characterization of CAC1 as a novel CDK2-associated cullin. Cell Cycle. 2014;8:3552–61. doi:10.4161/cc.8.21.9955.
Zheng Q, Zhao LY, Kong Y, Nan KJ, Yao Y, Liao ZJ. CDK-associated Cullin 1 can promote cell proliferation and inhibit cisplatin-induced apoptosis in the AGS gastric cancer cell line. World J Surg Oncol. 2013;11:5. doi:10.1186/1477-7819-11-5.
Park SW, Chung NG, Hur SY, Kim HS, Yoo NJ, Lee SH. Mutational analysis of hypoxia-related genes HIF1alpha and CUL2 in common human cancers. APMIS. 2009;117:880–5. doi:10.1111/j.1600-0463.2009.02550.x.
De Maio FA, Rocco CA, Aulicino PC, Bologna R, Mangano A, Sen L. APOBEC3-mediated editing in HIV type 1 from pediatric patients and its association with APOBEC3G/CUL5 polymorphisms and Vif variability. AIDS Res Hum Retroviruses. 2012;28:619–27. doi:10.1089/AID.2011.0291.
Bulatov MEM, Chatterjee S, Knebel A, Shimamura S, Konijnenberg A, Johnson C, et al. Biophysical studies on interactions and assembly of full-size E3 ubiquitin ligase: suppressor of cytokine signaling 2 (SOCS2)-elongin BC-cullin 5-ring box protein 2 (RBX2). J Biol Chem. 2015;290:4178–91. doi:10.1074/jbc.M114.616664.
Mahrour N, Redwine WB, Florens L, Swanson SK, Martin-Brown S, Bradford WD, et al. Characterization of Cullin-box sequences that direct recruitment of Cul2-Rbx1 and Cul5-Rbx2 modules to Elongin BC-based ubiquitin ligases. J Biol Chem. 2008;283:8005–13. doi:10.1074/jbc.M706987200.
Guo J, Miao Y, Xiao B, Huan R, Jiang Z, Meng D, et al. Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. J Gastroenterol Hepatol. 2009;24:652–7. doi:10.1111/j.1440-1746.2008.05666.x.
Su Y, Ni Z, Wang G, Cui J, Wei C, Wang J, et al. Aberrant expression of microRNAs in gastric cancer and biological significance of miR-574-3p. Int Immunopharmacol. 2012;13:468–75. doi:10.1016/j.intimp.2012.05.016.
Kazi JU, Ronnstrand L. Suppressor of cytokine signaling 2 (SOCS2) associates with FLT3 and negatively regulates downstream signaling. Mol Oncol. 2013;7:693–703. doi:10.1016/j.molonc.2013.02.020.
Lai R-H, Hsiao Y-W, Wang M-J, Lin H-Y, Wu C-W, Chi C-W, et al. SOCS6, down-regulated in gastric cancer, inhibits cell proliferation and colony formation. Cancer Lett. 2010;288:75–85. doi:10.1016/j.canlet.2009.06.025.
Li G, Xu J, Wang Z, Yuan Y, Li Y, Cai S, et al. Low expression of SOCS-1 and SOCS-3 is a poor prognostic indicator for gastric cancer patients. J Cancer Res Clin Oncol. 2015;141:443–52. doi:10.1007/s00432-014-1838-5.
Furukawa M, He YJ, Borchers C, Xiong Y. Targeting of protein ubiquitination by BTB-Cullin 3-Roc1 ubiquitin ligases. Nat Cell Biol. 2003;5:1001–7. doi:10.1038/ncb1056.
Stogios PJ, Downs GS, Jauhal JJ, Nandra SK, Prive GG. Sequence and structural analysis of BTB domain proteins. Genome Biol. 2005;6:R82. doi:10.1186/gb-2005-6-10-r82.
Zhuang M, Calabrese MF, Liu J, Waddell MB, Nourse A, Hammel M, et al. Structures of SPOP-substrate complexes: insights into molecular architectures of BTB-Cul3 ubiquitin ligases. Mol Cell. 2009;36:39–50. doi:10.1016/j.molcel.2009.09.022.
Merchant JL. Hedgehog signalling in gut development, physiology and cancer. J Physiol. 2012;590(Pt 3):421–32. doi:10.1113/jphysiol.2011.220681.
Zeng C, Wang Y, Lu Q, Chen J, Zhang J, Liu T, et al. SPOP suppresses tumorigenesis by regulating Hedgehog/Gli2 signaling pathway in gastric cancer. J Exp Clin Cancer Res. 2014;33:75. doi:10.1186/s13046-014-0075-8.
Zeng Q, Zhang L, Wang B, Ou CY, Chien CT, Jiang J. A Hedgehog-induced BTB protein modulates Hedgehog signaling by degrading Ci/Gli transcription factor. Dev Cell. 2006;10:710–29. doi:10.1016/j.devcel.2006.05.004.
Kim MS, Je EM, Oh JE, Yoo NJ, Lee SH. Mutational and expressional analyses of SPOP, a candidate tumor suppressor gene, in prostate, gastric and colorectal cancers. APMIS. 2013;121:626–33. doi:10.1111/apm.12030.
Lee J, Zhou P. Pathogenic role of the CRL4 ubiquitin ligase in human disease. Front Oncol. 2012;2:21. doi:10.3389/fonc.2012.00021.
Kerzendorfer C, Whibley A, Carpenter G, Outwin E, Chiang SC, Turner G, et al. Mutations in Cullin 4B result in a human syndrome associated with increased camptothecin-induced topoisomerase I-dependent DNA breaks. Hum Mol Genet. 2010;19:1324–34. doi:10.1093/hmg/ddq008.
Kerzendorfer C, Hart L, Colnaghi R, Carpenter G, Alcantara D, Outwin E, et al. CUL4B-deficiency in humans: understanding the clinical consequences of impaired Cullin 4-RING E3 ubiquitin ligase function. Mech Ageing Dev. 2011;132:366–73. doi:10.1016/j.mad.2011.02.003.
Hannah J, Zhou PB. The CUL4A ubiquitin ligase is a potential therapeutic target in skin cancer and other malignancies. Chin J Cancer. 2013;32:478–82. doi:10.5732/cjc.012.10279.
Hung MS, Mao JH, Xu Z, Yang CT, Yu JS, Harvard C, et al. Cul4A is an oncogene in malignant pleural mesothelioma. J Cell Mol Med. 2011;15:350–8. doi:10.1111/j.1582-4934.2009.00971.x.
Xu Y, Wang Y, Ma G, Wang Q, Wei G. CUL4A is overexpressed in human pituitary adenomas and regulates pituitary tumor cell proliferation. J Neurooncol. 2014;116:625–32. doi:10.1007/s11060-013-1349-2.
Wang Y, Wen M, Kwon Y, Xu Y, Liu Y, Zhang P, et al. CUL4A induces epithelial-mesenchymal transition and promotes cancer metastasis by regulating ZEB1 expression. Cancer Res. 2014;74:520–31. doi:10.1158/0008-5472.CAN-13-2182.
Wang Y, Ma G, Wang Q, Wen M, Xu Y, He X, et al. Involvement of CUL4A in regulation of multidrug resistance to P-gp substrate drugs in breast cancer cells. Molecules. 2013;19:159–76. doi:10.3390/molecules19010159.
Ren S, Xu C, Cui Z, Yu Y, Xu W, Wang F, et al. Oncogenic CUL4A determines the response to thalidomide treatment in prostate cancer. J Mol Med (Berl). 2012;90:1121–32. doi:10.1007/s00109-012-0885-0.
Thirunavukarasou A, Singh P, Govindarajalu G, Bandi V, Baluchamy S. E3 ubiquitin ligase Cullin4B mediated polyubiquitination of p53 for its degradation. Mol Cell Biochem. 2014;390:93–100. doi:10.1007/s11010-014-1960-3.
Lee J, Zhou P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol Cell. 2007;26:775–80. doi:10.1016/j.molcel.2007.06.001.
Banks D, Wu M, Higa LA, Gavrilova N, Quan J, Ye T, et al. L2DTL/CDT2 and PCNA interact with p53 and regulate p53 polyubiquitination and protein stability through MDM2 and CUL4A/DDB1 complexes. Cell Cycle. 2014;5:1719–29. doi:10.4161/cc.5.15.3150.
Mendoza M, Mandani G, Momand J. The MDM2 gene family. Biomol Concepts. 2014;5:9–19. doi:10.1515/bmc-2013-0027.
Nag A, Bagchi S, Raychaudhuri P. Cul4A physically associates with MDM2 and participates in the proteolysis of p53. Cancer Res. 2004;64:8152–5. doi:10.1158/0008-5472.CAN-04-2598.
Shen W, Hu P, Cao JQ, Liu XX, Shao JH. MDM2 oncogene, E3 ubiquitin protein ligase T309G polymorphism and risk of oesophageal or gastric cancer: meta-analysis of 15 studies. J Int Med Res. 2014;42:1065–76. doi:10.1177/0300060514527910.
Li YF, Wang DD, Zhao BW, Wang W, Huang CY, Chen YM, et al. High level of COP1 expression is associated with poor prognosis in primary gastric cancer. Int J Biol Sci. 2012;8:1168–77. doi:10.7150/ijbs.4778.
Sawada G, Ueo H, Matsumura T, Uchi R, Ishibashi M, Mima K, et al. Loss of COP1 expression determines poor prognosis in patients with gastric cancer. Oncol Rep. 2013;30:1971–5. doi:10.3892/or.2013.2664.
Jackson PK. Regulating microtubules and genome stability via the CUL7/3M syndrome complex and CUL9. Mol Cell. 2014;54:713–5. doi:10.1016/j.molcel.2014.05.024.
Guo H, Wu F, Wang Y, Yan C, Su W. Overexpressed ubiquitin ligase Cullin7 in breast cancer promotes cell proliferation and invasion via down-regulating p53. Biochem Biophys Res Commun. 2014;450:1370–6. doi:10.1016/j.bbrc.2014.06.134.
Xu X, Sarikas A, Dias-Santagata DC, Dolios G, Lafontant PJ, Tsai SC, et al. The CUL7 E3 ubiquitin ligase targets insulin receptor substrate 1 for ubiquitin-dependent degradation. Mol Cell. 2008;30:403–14. doi:10.1016/j.molcel.2008.03.009.
Kuo KL, Ho IL, Shih TH, Wu JT, Lin WC, Tsai YC, et al. MLN4924, a novel protein neddylation inhibitor, suppresses proliferation and migration of human urothelial carcinoma: In vitro and in vivo studies. Cancer Lett. 2015;363:127–36. doi:10.1016/j.canlet.2015.01.015.
Soucy TA, Smith PG, Milhollen MA, Berger AJ, Gavin JM, Adhikari S, et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature. 2009;458:732–6. doi:10.1038/nature07884.
Arias EE, Walter JC. Replication-dependent destruction of Cdt1 limits DNA replication to a single round per cell cycle in Xenopus egg extracts. Genes Dev. 2005;19:114–26. doi:10.1101/gad.1255805.
Li X, Zhao Q, Liao R, Sun P, Wu X. The SCF(Skp2) ubiquitin ligase complex interacts with the human replication licensing factor Cdt1 and regulates Cdt1 degradation. J Biol Chem. 2003;278:30854–8. doi:10.1074/jbc.C300251200.
Abbas T, Mueller AC, Shibata E, Keaton M, Rossi M, Dutta A. CRL1-FBXO11 promotes Cdt2 ubiquitination and degradation and regulates Pr-Set7/Set8-mediated cellular migration. Mol Cell. 2013;49:1147–58. doi:10.1016/j.molcel.2013.02.003.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Rights and permissions
About this article
Cite this article
Chen, P., Yao, GD. The role of cullin proteins in gastric cancer. Tumor Biol. 37, 29–37 (2016). https://doi.org/10.1007/s13277-015-4154-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4154-z