Abstract
Papillary thyroid cancer (PTC) is a predominant type of thyroid cancer. Ionizing radiation is the only well-established risk factor and may result in double-strand breaks. The x-ray repair cross-complementing group 3 (XRCC3) gene plays a vital role in DNA repair through homologous recombination. We aimed at investigating the association between XRCC3 genetic polymorphisms and PTC susceptibility. Eighty-three PTC patients and 367 controls in a Chinese population were enrolled in the study. Tag single-nucleotide polymorphisms (SNPs) were identified by HaploView 4.2 software. Genomic DNAs were isolated from peripheral blood samples by using TaqMan Blood DNA kits. The genotyping of XRCC3 SNPs was performed by TaqMan SNPs genotyping assay. Odds ratios (ORs) and corresponding 95 % confidence intervals (CIs) were calculated to evaluate the association between XRCC3 SNPs and PTC susceptibility. The statistical analyses were conducted by using SPSS 13.0 software. Four tag-SNPs were initially identified by HaploView 4.2 software. Only one SNP (rs861539) was shown to be significantly associated with increased risk of PTC. There was a significant difference in smoking and drinking status between PTC cases and controls. And the stratified analysis suggested that the polymorphisms of rs861539 in XRCC3 were correlated with PTC risk in the four subgroups of smokers (ex-smokers included), non-smokers, drinkers (ex-drinkers included), and non-drinkers. The meta-analysis showed that only two studies reported a significant association between XRCC3 polymorphisms and PTC risk. In this study, we find a significant association between rs861539 polymorphisms and PTC susceptibility. However, there were inconsistent results in previous published studies. Therefore, further studies in a large population are required to gain insights into the PTC risk conferred by XRCC3 SNPs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thyroid cancer is the most common endocrine malignancy worldwide, which accounts for 90 % of endocrine cancer cases [1]. The incidence rate has been continuously increasing in the past decades and it is the fifth most prevalent cancer in women at present [2]. Thyroid cancer has three main subtypes, namely papillary thyroid cancer (PTC), follicular thyroid carcinoma (FTC), and medullar thyroid carcinoma (MTC) [3]. PTC is the predominant type of thyroid cancer and accounts for almost 90 % of all cases [4]. PTC emerges as an irregular form of solid or cystic mass or nodule in a normal thyroid parenchyma. To date, ionizing radiation is the only well-established risk factor for thyroid cancer [5]. However, exposure to ionizing radiation does not necessarily lead to the development of thyroid cancer since not only environmental factors but also genetic predisposition could contribute to the occurrence of thyroid cancer [6]. In other words, individuals with diverse genetic backgrounds showed different sensitivity to ionizing radiation [7].
DNA double-strand breaks (DSBs) are severe DNA lesions in eukaryotic cells, leading to the genome instability [8]. DSBs may occur in exposure to exogenous agents, such as ionizing radiation, alkylating agents, etc. [9]. DNA damage repair is a fundamental mechanism to maintain the normal physiological function, especially when the cells are exposed to stress or exogenous damage. Hence, the dysfunction of DNA repair may result in the malignant tumor development. There were two major mechanisms to repair DSBs, homologous recombination repair (HRR) and the non-homologous end-joining (NHEJ) pathways. The precise mechanisms of regulation in the two DSBs repair pathways are not well elucidated yet, but a number of genes are known to regulating these pathways. X-ray repair cross-complementing group 3 (XRCC3) is a key member of DNA repair gene responsible for maintaining the genome stability and repairing the DNA double-strand breaks caused by ionizing radiation damage [10]. XRCC3 repairs the DSBs through HRR pathway [11]. During HRR, the XRCC3 is essential for Rad51 recruitment and enables the Rad51 protein to assemble as multimers to the sites of DSBs [12]. The XRCC3 also play an important role in the enzymatic resolution of the Holliday junction [13]. Reduced HRR and hypersensitivity to ionizing radiation, UV radiation, genotoxic alkylating agents, and cross-linking agents were observed in XRCC3 deficient cell lines [14]. Defects in the HRR-related genes have been revealed to be correlated with carcinogenesis [15]. Thus, single-nucleotide polymorphisms (SNP) in the XRCC3 may contribute to the altered activity of encoding proteins or even loss of protein functions and ultimately influence the individual genetic predisposition to cancer. A number of studies have investigated the association between XRCC3 polymorphisms and risk of diverse cancers, such as colorectal cancer [16, 17], bladder cancer [18], ovarian cancer [19], etc. The most intensively studied polymorphisms in the XRCC3 gene is C241T, namely C18067T or rs861539, which causes a substitution from Thr to Met at codon 241. Several studies have been conducted to examine the correlation between rs861539 polymorphisms in XRCC3 and PTC cancer risks [20–23]. Nonetheless, no conclusive finding was reported due to the inconsistent results between studies. The inconsistency may probably be due to limited sample sizes and ethnicities with different genetic backgrounds.
The aim of our study is to investigate the association between XRCC3 polymorphisms and PTC risk in the Chinese population. Additionally, we also summarized the previous studies in the meta-analysis to further assess the PTC risk conferred by XRCC3 SNPs.
Methods and materials
Study subjects
The study population consisted of 550 subjects, including 183 PTC cases and 367 healthy controls. PTC patients were recruited from Shandong Provincial Qianfoshan Hospital and the control group was healthy volunteers for annual physical examination. All the participants were Chinese resided in the surrounding regions of Sichuan Province. The medical records of PTC patients were under review, in order to ensure that the PTC patients had no previous cancer diagnosis or treatment. Moreover, the diagnosis of PTC patients as well as PTC subtype determination was confirmed by histopathological examination. The medical records of healthy controls were also reviewed to ensure that they have no previous or current diagnosis of cancer or thyroid-related diseases. The protocol of our study was approved by the ethic committee of Shandong Provincial Qianfoshan Hospital and written informed consents were obtained from all participants prior to execute any experimental procedure.
Tag-SNPs selection
Tag-SNPs of XRCC3 in Han Chinese population were initially retrieved through the HapMap database (Hapmap Data Rel 24/Phase II Nov08, on NCBI B36, dbSNP b126) and further analyzed with HaploView 4.2 (Broad Institute, Cambridge, MA, USA). Furthermore, tag-SNPs were identified based on the selection criteria, which the minor allele frequency must be greater than 0.05 and linkage disequilibrium r 2 must be greater than 0.8.
DNA extraction and SNP genotyping
Five milliliters of peripheral blood was collected from each subject in our study. QIAamp DNA Blood Kit (QIAGEN, Hilden, Germany) were used to extract the genomic DNAs from the blood samples. The SNP genotyping was performed using Sequenom MassARRAY iPLEX platform (Sequenom Inc., San Diego, CA, USA) with matrix-assisted laser desorption ionization time of flight (MALDI-TOF) mass spectrometry. PCR primers were designed by MassARRAY Assay Design Version 3.1 (Sequenom Inc., San Diego, CA, USA). The 5-μl PCR reactions were performed in 384-well plates, in which the reaction mixture includes 10 ng genomic DNA, 100 nM each designed primers, 5 U Taq DNA polymerase, 100 nM dNTP mix, 10× PCR buffer, and 25 mM MgCl2. The conditions of PCR thermal cycles were as follows: (1) initial denaturation 94 °C for 15 min; (2) 45 thermal cycles of denaturation (94 °C, 20 s), annealing (56 °C, 20 s), and extension (72 °C for 60 s); (3) final extension at 72 °C for 3 mins. After removal of unincorporated dNTPs by 1.7 U shrimp alkaline phosphatase (SAP), the plate were placed at 37 °C for 20 mins and then 85 °C for 5 mins. The iPLEX reaction mix includes the PCR amplication products, 10 × iPLEX Buffer Plus, iPLEX Termination Mix, Primer mix, and iPLEX enzyme. The iPLEX reaction was carried out under the conditions as follows: (1) initial denaturation at 94 °C for 30 s; (2) 40 cycles at 94 °C for 5 s, [52 °C for 5 s and 80 °C for 5 s (repeated five times per cycle)]; and (3) final incubation at 72 °C for 3 min. The iPLEX reaction products were desalted by resin treatment and water. The iPLEX product was subsequently spotted on a SpectroCHIP (Sequenom, Inc., San Diego, CA), and the data were analyzed by SpectroTYPER 4.0 software (Sequenom, Inc., San Diego, CA).
Statistical analyses
Difference of clinical characteristics (age, sex, smoking and alcohol status) between PTC cases and controls were tested by chi-square test. The Hardy-Weinberg equilibrium of selected tag-SNPs was assessed by goodness-of-fit chi-square test. Genetic models such as homozygous comparison (MM vs. WW), dominant model (WM + MM vs. WW), recessive model (MM vs. WW + WM) and allele model (M vs. W) were constructed to compare the allelic and genotypic frequencies between PTC cases and controls. Odds ratio (OR) and the 95 % confidence interval (95 % CI) were used to estimate the PTC risk conferred by XRCC3 polymorphisms. In meta-analysis, heterogeneity between studies was assessed by I 2 and Q test. When I 2 < 0.05 and P > 0.05 for Q test, fixed effects model should be used; otherwise, the random effects model should be employed. Funnel plot were used to evaluate the publication bias. A two-tailed P value less than 0.05 was considered as statistically significant. All statistical analyses were performed using SPSS 20.0 software (SPSS, Inc., Chicago, IL, USA). Meta-analysis was conducted with STATA version 11.0 (Stata, College Station, TX, USA).
Results
Clinical features of study population
In this cohort study, 550 participants include 183 PTC patients and 367 healthy controls. Comparing the clinical features between PTC cases and controls, we found a similar age and sex ratio in two groups (Table 1). However, there was significant difference in the consumption of tobacco and alcohol between PTC patients and healthy controls. Additionally, the proportion of current smoker and drinker was markedly higher in PTC patients compared with healthy controls. In regards to the cancer-related characteristics of PTC patients, the PTC patients were mainly diagnosed at the stage of T2 or T3. Sixty-eight PTC patients have no clinical manifestation of regional lymph node metastasis, nonetheless, 75 PTC patients have tumor cells found in one lymph node and the rest of the 40 PTC patients have tumor cells invaded in two or three nearby lymph nodes. Most of the PTC patients have primary tumors located in either right lobe or left lobe; 72.1 % of PTC cases have no extra-thyroidal extension.
Tags SNP selection
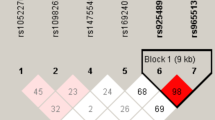
Sixteen tag-SNPs were identified from HapMap databases, with MAF > 0.05 and r 2 > 0.08 as selection criteria. The linkage disequilibrium of 16 retrieved tag-SNPs was illustrated in Fig. 1. As it is shown, four SNPs (rs1799794, rs56377012, rs1799796, and rs861539) which can represent all genetic variants of XRCC3 were selected by HaploView 4.0 (Broad Institute, Cambridge, MA). The relative locations of four SNPs were shown in Fig. 2. The polymorphism of rs1799794 in XRCC3 was located in the 5′ untranslated region (5′UTR). The rs56377012 mutation was localized in exon1, and the polymorphisms of rs1799796 and rs861539 were situated within exon 5.
Association between the XRCC3 polymorphisms and risk of PTC
The XRCC3 allelic and genotypic frequencies of PTC cases and healthy controls were shown in Table 2. No significant deviation from Hardy-Weinberg Equilibrium was observed in the allelic and genotypic frequencies of PTC cases and healthy controls (Table 3). Analysis of the four XRCC3 SNPs showed a significant correlation between rs861539 polymorphisms and increased PTC risk. In the allele model (T vs. C), T mutant allele was significant associated with increased risk of PTC (OR = 1.65, 95 % CI = 1.24–2.19, P = 0.001). In the homozygous model (TT vs. CC), the individuals with TT mutants were more likely to develop PTC than CC wild types carriers (OR = 2.93, 95 % CI = 1.55–5.56, P = 0.001). In the dominant model (CT + TT vs. CC), the combined genotypes CT + TT were associated with higher risk of PTC than CC wild type (OR = 1.59, 95 % CI = 1.11–2.28, P = 0.011). In the recessive model (TT vs. CT + CC), TT mutants were shown to associate with increased risk of PTC (OR = 2.62, 95 % CI = 1.40–4.88, P = 0.002). However, the polymorphisms of rs1799796, rs56377012, and rs1799794 were reported to have no apparent association with PTC susceptibility.
Stratified analysis of XRCC3 in rs861539 and PTC risk
As shown in Table 3, it was noted that PTC cases have significantly higher proportion of current tobacco or alcohol users than healthy controls. Hence, we further investigated the impact of rs861539 polymorphism on the PTC occurrence stratified by smoking and drinking status. Stratified analysis (Table 4) showed that there was a significantly higher frequency of T allele and CT or TT genotypes in the subgroup of smokers or ex-smokers. The polymorphisms of rs861539 in XRCC3 were significantly associated with PTC risk in the subgroup of non-smokers under the allelic, homozygous, and recessive models. In regards to the drinking status, we observed a significant association between XRCC3 variants of rs861539 and PTC susceptibility in the subgroup of current or former drinkers under the allelic, homozygous, and recessive models. As for the subgroup of non-drinkers, T allele was shown to be a risk factor compared with C allele under the allele model. Meanwhile, the TT and TC mutants were significantly associated with PTC risk compared with CC wild types under the dominant model.
Meta-analysis of the association between XRCC3 polymorphisms in rs861539 and PTC risk
Four eligible studies were identified from the comprehensive literature retrieval, which investigated the association between XRCC3 polymorphisms in rs861539 and PTC risk [20–23]. The results of the meta-analysis were summarized in Table 5. Sturgis et al. reported the significant correlation between XRCC3 polymorphisms in rs861539 and PTC risk under allelic (OR = 1.16; 95 % CI = 1.15–2.27; P = 0.006) and dominant models (OR = 2.10; 95 % CI = 1.31–3.38; P = 0.002). Bastos et al. also revealed that the rs861539 polymorphisms in XRCC3 were associated with PTC susceptibility under the recessive model (OR = 2.35; 95 % CI = 1.24–4.44; P = 0.007). However, Siraj et al. and Akulevich et al. reported no significant association between XRCC3 polymorphisms in rs861539 and PTC risk.
Discussion
In the present study, we conducted this hospital-based cohort study to investigate the association between XRCC3 SNPs and PTC susceptibility. Our study involved 183 PTC patients and 367 healthy controls. Our results showed that only rs861539 polymorphisms were shown to significantly associate with higher risk of PTC. Comparing the clinicopathological parameters of cases and controls, we found a significant difference in smoking and drinking status between PTC patients and healthy controls. Therefore, we further performed a stratified analysis based on the smoking and drinking status in PTC patients and healthy controls. Our data indicated that XRCC3 polymorphisms in rs861539 were significantly associated with increased risk of PTC in the subgroup of current, former, and non-smokers. Similar results were observed in the subgroup of current, former and non-smokers as well. In the meta-analysis of previous studies, the rs861539 polymorphisms of XRCC3 were reported to correlate with PTC susceptibility in the studies of Sturgis et al. and Bastos et al. The other two eligible studies suggested no association between rs861539 polymorphisms of XRCC3 and PTC risk.
XRCC3, located in chromosome 14q32.3, is a member of Rad51 family that enhances chromosome stability and repair DNA damage through homologous recombination. Genomic instability is a hallmark of most cancers, caused by abnormal DNA damage repair as well as the defects in apoptosis induction and cell cycle checkpoints [24]. In the process of DNA repair, Ku70/Ku80 heterodimer (KU) and MRE11-RAD50-NBS1 complex (MRN) function as sensor proteins to detect the DSB (Fig. 3) [25]. After binding to the DSB sites, DSB repair was activated through signal transduction by KU or MRN. XRCC3, along with Rad51 and BRCA2, initiate the homologous recombination [26]. Effective DNA repair can restore the normal cell function and cell proliferation. However, ineffective DNA repair can result in cell cycle arrest, apoptosis, or even cancer [27]. Hence, the XRCC3 mutation might induce the genomic instability due to ineffective DNA repair, ultimately leading to the development of cancer.
Accumulating evidences have suggested that XRCC3 is implicated in the DSBs repair. The C241T mutation in XRCC3 is likely to be marginally but not significantly associated with impaired DNA repair capacity [28]. Hence, a number of studies have investigated the association between XRCC3 polymorphisms and cancer. The C241T polymorphism in the exon5 of XRCC3 gene was the most intensively studied genetic variant, which was reported to associate with colorectal cancer [16, 17], ovarian cancer [19], acute myeloid leukemia [29, 30], glioma [31], bladder cancer [18, 32], and melanoma skin cancer [33]. However, other studies reported no conclusive evidence of the correlation between XRCC3 polymorphisms and risk of lung cancer [34, 35], gastric cancer [36, 37], breast cancer [38–40], as well as prostate cancer [41]. As for thyroid cancer, several studies have been conducted to examine the possible association between XRCC3 SNPs and thyroid cancer risk. Sturgis et al. and Bastos et al. reported a positive association between XRCC3 variants and thyroid cancer susceptibility [21, 23]. In contrast, Siraj et al. and Akulevich N et al. found no evident correlation between XRCC3 polymorphism and thyroid cancer risk [20, 22].
In view of the mutation site in XRCC3, the rs56377012, rs1799796 and rs861539 are located in the coding regions (exon1 and exon5) while the rs1799794 is situated in the 5′UTR. Even though the four polymorphisms were located in the regulatory and coding regions, only rs861539 were shown to associate with increased PTC risk. The SNP of rs861539 contributes to Thr241Met amino acid substitution, which may possibly alter the function of XRCC3 or its interaction with other proteins implicated in the process of DNA damage repair.
Some inherited limitation in our study needed to be addressed. First, since our study was a hospital-based cohort research, the possibility of selection bias in the recruitment process cannot be ruled out. Besides, our study was conducted only based on Asian ethnicity. Due to the existence of different genetic backgrounds between ethnic groups, the genetic predisposition to PTC induced by XRCC3 mutation may vary among different ethnicities. Moreover, our study was conducted on a relatively small population, as well as the eligible studies included in our meta-analysis. Since there were few studies published on the association between rs861539 polymorphisms in XRCC3 and PTC risk, only four qualified studies were included in our study and thus the results of meta-analysis can only serve as a supporting evidence for our study. Further large-scale studies are needed to further validate our results and shed light on the etiological mechanism of PTC induced by XRCC3 polymorphisms.
In conclusion, our study suggested that the polymorphism of rs861539 in XRCC3 was associated with increased risk of PTC. Due to the significant difference in smoking and drinking status between PTC cases and controls, we performed a stratified analysis based on smoking and drinking status, of which the results indicated that the polymorphisms of rs861539 in XRCC3 was correlated with PTC susceptibility in the four subgroups (current/former smokers, non-smokers, current/former drinkers, and non-drinkers). In addition, the meta-analysis showed that two out of four eligible studies have reported a significant association between rs861539 polymorphisms in XRCC3 and PTC susceptibility.
References
Adam MA, Pura J, Goffredo P, Dinan MA, Hyslop T, Reed SD, et al. Impact of extent of surgery on survival for papillary thyroid cancer patients younger than 45 years. J. Clin Endocrinol Metab. 2014;100:115–121.
Vigneri R, Malandrino P, Vigneri P. The changing epidemiology of thyroid cancer: why is incidence increasing? Curr Opin Oncol. 2015;27:1–7.
Nikiforov YE, Nikiforova MN. Molecular genetics and diagnosis of thyroid cancer. Nat Rev Endocrinol. 2011;7:569–80.
Siegel R, DeSantis C, Virgo K, Stein K, Mariotto A, Smith T, et al. Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin. 2012;62:220–41.
Schlumberger M, Cailleux AF, Suarez HG, de Vathaire F. Irradiation and second cancers. The thyroid as a case in point. C R Acad Sci III. 1999;322:205–13.
Czene K, Lichtenstein P, Hemminki K. Environmental and heritable causes of cancer among 9.6 million individuals in the Swedish family‐cancer database. Int J Cancer. 2002;99:260–6.
Frigerio N, Stowe R. Carcinogenic and genetic hazard from background radiation. In: Biological and environmental effects of low-level radiation. 1976.
Kaina B. DNA damage-triggered apoptosis: critical role of DNA repair, double-strand breaks, cell proliferation and signaling. Biochem Pharmacol. 2003;66:1547–54.
Wolff S, Afzal V, Wiencke JK, Olivieri G, Michaeli A. Human lymphocytes exposed to low doses of ionizing radiations become refractory to high doses of radiation as well as to chemical mutagens that induce double-strand breaks in DNA. Int J Radiat Biol. 1988;53:39–48.
Thacker J, Zdzienicka MZ. The < i > XRCC</i > genes: expanding roles in DNA double-strand break repair. DNA Repair. 2004;3:1081–90.
Brenneman MA, Weiss AE, Nickoloff JA, Chen DJ. < i > XRCC3</i > is required for efficient repair of chromosome breaks by homologous recombination. Mutation Research/DNA Repair. 2000;459:89–97.
Bishop DK, Ear U, Bhattacharyya A, Calderone C, Beckett M, Weichselbaum RR, et al. Xrcc3 is required for assembly of Rad51 complexes in vivo. J Biol Chem. 1998;273:21482–8.
Liu Y, Masson J-Y, Shah R, O’Regan P, West SC. RAD51C is required for Holliday junction processing in mammalian cells. Science. 2004;303:243–6.
Jones NJ, Cox R, Thacker J. Isolation and cross-sensitivity of x-ray-sensitive mutants of V79-4 hamster cells. Mutation Research/DNA Repair Reports. 1987;183:279–86.
Goode EL, Ulrich CM, Potter JD. Polymorphisms in DNA repair genes and associations with cancer risk. Cancer Epidemiol Biomark Prev. 2002;11:1513–30.
Jin M-J, Chen K, Song L, Fan C-H, Chen Q, Zhu Y-M, et al. The association of the DNA repair gene < i > XRCC3</i > Thr241Met polymorphism with susceptibility to colorectal cancer in a Chinese population. Cancer Genet Cytogenet. 2005;163:38–43.
Krupa R, Blasiak J. An association of polymorphism of DNA repair genes XRCC1 and XRCC3 with colorectal cancer. J Exp Clin Cancer Res. 2004;23:285–94.
Mittal RD, Gangwar R, Mandal RK, Srivastava P, Ahirwar DK. Gene variants of XRCC4 and XRCC3 and their association with risk for urothelial bladder cancer. Mol Biol Rep. 2012;39:1667–75.
Auranen A, Song H, Waterfall C, DiCioccio RA, Kuschel B, Kjaer SK, et al. Polymorphisms in DNA repair genes and epithelial ovarian cancer risk. Int J Cancer. 2005;117:611–8.
Akulevich NM, Saenko VA, Rogounovitch TI, Drozd VM, Lushnikov EF, Ivanov VK, et al. Polymorphisms of DNA damage response genes in radiation-related and sporadic papillary thyroid carcinoma. Endocr Relat Cancer. 2009;16:491–503.
Bastos HN, Antao MR, Silva SN, Azevedo AP, Manita I, Teixeira V, et al. Association of polymorphisms in genes of the homologous recombination DNA repair pathway and thyroid cancer risk. Thyroid. 2009;19:1067–75.
Siraj AK, Al-Rasheed M, Ibrahim M, Siddiqui K, Al-Dayel F, Al-Sanea O, et al. RAD52 polymorphisms contribute to the development of papillary thyroid cancer susceptibility in Middle Eastern population. J Endocrinol Invest. 2008;31:893–9.
Sturgis EM, Zhao C, Zheng R, Wei Q. Radiation response genotype and risk of differentiated thyroid cancer: a case-control analysis. Laryngoscope. 2005;115:938–45.
Lengauer C, Kinzler KW, Vogelstein B. Genetic instabilities in human cancers. Nature. 1998;396:643–9.
Polo SE, Jackson SP. Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications. Genes Dev. 2011;25:409–33.
Zhou B-BS, Elledge SJ. The DNA damage response: putting checkpoints in perspective. Nature. 2000;408:433–9.
Hartwell LH, Kastan MB. Cell cycle control and cancer. Science. 1994;266:1821–8.
Araujo FD, Pierce AJ, Stark JM, Jasin M. Variant XRCC3 implicated in cancer is functional in homology-directed repair of double-strand breaks. Oncogene. 2002;21:4176–80.
Hamdy MS, El-Haddad AM, El-Din NMB, Makhlouf MM, Abdel-Hamid SM. RAD51 and XRCC3 gene polymorphisms and the risk of developing acute myeloid leukemia. J Investig Med. 2011;59:1124–30.
Seedhouse C, Faulkner R, Ashraf N, Das-Gupta E, Russell N. Polymorphisms in genes involved in homologous recombination repair interact to increase the risk of developing acute myeloid leukemia. Clin Cancer Res. 2004;10:2675–80.
Zhou K, Liu Y, Zhang H, Liu H, Fan W, Zhong Y, et al. XRCC3 haplotypes and risk of gliomas in a Chinese population: a hospital‐based case‐control study. Int J Cancer. 2009;124:2948–53.
Matullo G, Guarrera S, Carturan S, Peluso M, Malaveille C, Davico L, et al. DNA repair gene polymorphisms, bulky DNA adducts in white blood cells and bladder cancer in a case‐control study. Int J Cancer. 2001;92:562–7.
Winsey SL, Haldar NA, Marsh HP, Bunce M, Marshall SE, Harris AL, et al. A variant within the DNA repair gene XRCC3 is associated with the development of melanoma skin cancer. Cancer Res. 2000;60:5612–6.
David-Beabes GL, Lunn RM, London SJ. No association between the XPD (Lys751G1n) polymorphism or the XRCC3 (Thr241Met) polymorphism and lung cancer risk. Cancer Epidemiol Biomark Prev. 2001;10:911–2.
Misra RR, Ratnasinghe D, Tangrea JA, Virtamo J, Andersen MR, Barrett M, et al. Polymorphisms in the DNA repair genes < i > XPD, XRCC1, XRCC3</i > and < i > APE/ref-1</i > and the risk of lung cancer amongmale smokers in Finland. Cancer Lett. 2003;191:171–8.
Duarte MC, Colombo J, Rossit ARB, Caetano A, Borim AA, Wornrath D, et al. Polymorphisms of DNA repair genes XRCC1 and XRCC3, interaction with environmental exposure and risk of chronic gastritis and gastric cancer. World J Gastroenterol. 2005;11:6593.
Shen H, Wang X, Hu Z, Zhang Z, Xu Y, Hu X, et al. Polymorphisms of DNA repair gene < i > XRCC3</i > Thr241Met and risk of gastric cancer in a Chinese population. Cancer Lett. 2004;206:51–8.
Brooks J, Shore RE, Zeleniuch-Jacquotte A, Currie D, Afanasyeva Y, Koenig KL, et al. Polymorphisms in RAD51, XRCC2, and XRCC3 are not related to breast cancer risk. Cancer Epidemiol Biomark Prev. 2008;17:1016–9.
Jacobsen NR, Nexø BA, Olsen A, Overvad K, Wallin H, Tjønneland A, et al. No association between the DNA repair gene XRCC3 T241M polymorphism and risk of skin cancer and breast cancer. Cancer Epidemiol Biomark Prev. 2003;12:584–5.
Thyagarajan B, Anderson KE, Folsom AR, Jacobs Jr DR, Lynch CF, Bargaje A, et al. No association between XRCC1 and XRCC3 gene polymorphisms and breast cancer risk: Iowa Women’s Health Study. Cancer Detect Prev. 2006;30:313–21.
Ritchey JD, Huang W-Y, Chokkalingam AP, Gao Y-T, Deng J, Levine P, et al. Genetic variants of DNA repair genes and prostate cancer: a population-based study. Cancer Epidemiol Biomark Prev. 2005;14:1703–9.
Conflict of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Kai Yuan and Meiling Huo are first co-authors of this manuscript.
Kai Yuan and Meiling Huo contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yuan, K., Huo, M., Sun, Y. et al. Association between x-ray repair cross-complementing group 3 (XRCC3) genetic polymorphisms and papillary thyroid cancer susceptibility in a Chinese Han population. Tumor Biol. 37, 979–987 (2016). https://doi.org/10.1007/s13277-015-3882-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3882-4