Abstract
Gastric cancer is still the second leading cause of cancer-related death worldwide, even though its incidence and mortality have declined over the recent few decades. Epigenetic control using histone deacetylase inhibitors, such as trichostatin A (TSA), is a promising cancer therapy. This study aimed to assess the messenger RNA (mRNA) levels of three histone deacetylases (HDAC1, HDAC2, and HDAC3), two histone acetyltransferases (GCN5 and PCAF), and two possible targets of these histone modifiers (MYC and CDKN1A) in 50 matched pairs of gastric tumors and corresponding adjacent nontumors samples from patients with gastric adenocarcinoma, as well as their correlations and their possible associations with clinicopathological features. Additionally, we evaluated whether these genes are sensitive to TSA in gastric cancer cell lines. Our results demonstrated downregulation of HDAC1, PCAF, and CDKN1A in gastric tumors compared with adjacent nontumors (P < 0.05). On the other hand, upregulation of HDAC2, GCN5, and MYC was observed in gastric tumors compared with adjacent nontumors (P < 0.05). The mRNA level of MYC was correlated to HDAC3 and GCN5 (P < 0.05), whereas CDKN1A was correlated to HDAC1 and GCN5 (P < 0.05 and P < 0.01, respectively). In addition, the reduced expression of PCAF was associated with intestinal-type gastric cancer (P = 0.03) and TNM stages I/II (P = 0.01). The increased expression of GCN5 was associated with advanced stage gastric cancer (P = 0.02) and tumor invasion (P = 0.03). The gastric cell lines treated with TSA showed different patterns of histone deacetylase and acetyltransferase mRNA expression, downregulation of MYC, and upregulation of CDKN1A. Our findings suggest that alteration of histone modifier genes play an important role in gastric carcinogenesis, contributing to MYC and CDKN1A deregulation. In addition, all genes studied here are modulated by TSA, although this modulation appears to be dependent of the genetic background of the cell line.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite the fact that overall rates of gastric cancer (GC) continue to decline worldwide, the majority of patients are still diagnosed with advanced disease in Western countries [1-3]. In these cases, surgical resection of the primary tumor offers limited value for a cure and has high morbidity rates [3]. New strategies for early diagnosis and new therapeutic methods in GC continue to be explored.
Epigenetic mechanisms have increasingly drawn attention in cancer studies with therapeutic purposes [4]. Acetylation of the amino-terminal tails of nucleosomal histones is the best characterized posttranslational modification of the chromatin structure [5]. Reversible (de)acetylation of chromatin plays a crucial role in the regulation of gene transcription by the activity of two specific enzymes, histone acetyltransferases (HATs), and histone deacetylases (HDACs). The presence of acetylated lysine in histone tails is associated with a more relaxed chromatin state and gene transcription activation, while the deacetylation of lysine residues is associated with a more condensed chromatin state and transcriptional gene silencing [6]. In addition, HATs and HDACs participate in protein–protein interactions, as coactivators/adaptors or corepressors, respectively, in regulation of transcription [5, 7, 8]. Changes in the expression pattern of HDACs and HATs have the potential to affect the structure and integrity of the genome, which have been described in several human tumors [9-13]. Although regulation of gene expression is an important form of regulating enzyme activity, little is known regarding the role of HAT and HDAC messenger RNA (mRNA) expression in GC. To the best our knowledge, only two studies have reported the expression pattern of HDAC1 and HDAC2 in GC [14, 15]. Moreover, only one study reported the expression of HAT TIP60 in GC [16].
HDAC inhibitors are a class of anticancer drugs that have been used in clinical treatment of solid tumors, although increasing evidence has shown their worth in hematological malignancies [4, 17]. In cancer, global hypoacetylation induces repression of tumor suppressor genes, and HDAC inhibitors are considered highly effective in upregulating tumor suppressor gene expression [18]. Trichostatin A (TSA) is one of the most potent HDAC inhibitors, acting specifically in HDAC classes I (HDAC1-3 and 8) and II (HDAC4-7, 9, and 10). In GC cell lines, as in other tumor cells, TSA may inhibit cell growth and induce apoptosis through modulation of genes involved in these processes [19-21]. Moreover, TSA appears to increase the chemosensitivity of anticancer drugs in GC cell lines [22]. Despite the knowledge regarding the biological effects of TSA in cancer, the exact mechanism of its action is not fully understood. Furthermore, there is no information on HDAC and HAT expression in response to TSA in GC cells. Determination of differences in HDAC and HAT mRNA expression upon treatment with TSA may help understand the mechanism of regulation of these genes and of the drug action in GC cells.
The purpose of this study was to assess the mRNA levels of three HDACs (HDAC1, HDAC2, and HDAC3), two HATs (GCN5, also known as KAT2A, and PCAF, also known as KAT2B), and two possible targets of these histone modifiers (MYC and CDKN1A, also known as p21) in matched pairs of gastric tumors and corresponding adjacent nontumor tissues, as well as their correlations and their possible associations with clinicopathological features. In addition, we evaluated whether these genes are sensitive to TSA treatment in two GC cell lines.
Methods
Tissue specimens
Fifty matched pairs of GC and corresponding adjacent nontumor tissues were obtained from patients with gastric adenocarcinoma who underwent gastric resection and were used in the analysis of the expression of histone modifier genes HDAC1, HDAC2, HDAC3, GCN5, and PCAF. Of the 50 pairs of gastric tissues, 43 and 46 pairs were evaluated for the expression of possible targets of these histone modifiers, MYC and CDKN1A, respectively. The gastric samples were obtained from João de Barros Barreto University Hospital (HUJBB) and São Paulo Hospital (HSP), Brazil, from 2009 to 2013. None of the patients had a history of exposure to either chemotherapy or radiotherapy before surgery, and there was no other co-occurrence of diagnosed cancers. Written informed consent with approval of the ethics committees of HUJBB and HSP was obtained from all patients prior to sample collection.
Gastric tumors were classified according to Lauren and were staged using criteria by tumor–node–metastasis (TNM) staging [23]. The presence of Helicobacter pylori, a class I carcinogen, in tumor and nontumor samples was detected by polymerase chain reaction (PCR). A 150-bp fragment corresponding to 16S-rRNA from H. pylori was amplified as previously described [24] using DNA purified simultaneously with RNA. In each PCR experiment, positive and negative controls were included. Table 1 shows the clinicopathological characteristics of the patients diagnosed with GC.
Cell lines and TSA treatment
The ACP02 and ACP03 cell lines used in this study were previously established by our research group from primary gastric adenocarcinomas classified as diffuse and intestinal types, respectively [25]. The cell lines were cultured at 37 °C in RPMI media 1640 (GIBCO®, Grand Island, NY, USA) supplemented with 10 % fetal bovine serum (GIBCO) and 0.02 mg/mL kanamycin (GIBCO).
Before cell line treatment, we first evaluated the best dosage of TSA using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. Briefly, cells were seeded in duplicate on 96-well plates, and after they reached 80 % confluence, they were incubated with 250, 350, or 500 nM of TSA (Sigma-Aldrich, St. Louis, MO, USA) for 24 h. Untreated cells were used as controls. The cells were then incubated with 100 μL of 0.5 mg/mL MTT (Invitrogen, Eugene, OR, USA) for 3 h at 37 °C. After removal of MTT, 100 μL dimethylsulfoxide (Mallinckrodt, Phillipsburg, NJ, USA) was added and homogenized for 10 min. Optical density was detected in the NanoDrop ND-1000 (Thermo Scientific, Wilmington, DE, USA) using the UV/VIS absorbance module with 595 nm in λ1/Abs1 and 600 nm in λ2/Abs2. Average optical density of the untreated cells was considered 100 % survival. The cell viability in treated cells was expressed as a percentage relative to the untreated cells.
All of the three tested doses resulted in reduction of the cell populations compared with the respective controls (Fig. 1). Both cell lines showed a decrease in cell viability of approximately 70 % when treated with 250 nM of TSA. Therefore, this dose was selected for the evaluation of effects of TSA in the ACP02 and ACP03 cell lines.
For both cell lines, TSA treatment was performed in duplicate. In each replicate, cells in the same passage were cultivated in 75-cm2 cell culture flasks until they reached 80 % confluence. The growth medium was then replaced with a medium containing 250 nM of TSA (Sigma-Aldrich). Untreated cells were used as controls. After 24 h, the cells were trypsinized and washed twice with phosphate-buffered saline (GIBCO).
RNA extraction and cDNA synthesis
Total RNA was extracted from the tissue samples and cell lines using the AllPrep DNA/RNA/Protein Kit (Qiagen, Hilden, Germany) and the RNeasy Plus Mini Kit (Qiagen), respectively, according to the manufacturer’s instructions. The RNA concentration and quality were measured using the Nanodrop ND-1000 (Thermo Scientific). The integrity of RNA was assessed by gel electrophoresis. The complementary DNA (cDNA) was synthesized using High-Capacity® cDNA Reverse Transcription (Life Technologies, Foster City, CA, USA) by following the manufacturer’s protocol.
Reverse transcription quantitative PCR
The reaction to detect the expression levels of HDAC1 (Hs02621185_s1), HDAC2 (Hs00231032_m1), HDAC3 (Hs00187320_m1), GCN5 (Hs00221499_m1), PCAF (Hs00187332_m1), MYC (Hs00153408_m1), and CDKN1A (HS00355782_m1) was performed in triplicate using TaqMan® hydrolysis probes purchased as Assays-on-demand (Life Technologies, Foster City, CA, USA) and the Applied Biosystems 7500 fast real-time PCR system (Foster City, CA, USA). The reference genes GAPDH + B2M were used for analysis of tissue samples, and ACTB + B2M were used for analysis of cell lines, as previously determined [26]. The relative quantification (RQ) of mRNA expression in tissue samples and cell lines was determined according to Pffafl [27] and Livak e Schmittgen [28], respectively. A sample of a patient without cancer and the respective nontreated cell lines were designated as calibrators.
Statistical analysis
We first evaluated the distribution of data using the Shapiro–Wilk test to determine subsequent appropriate tests for statistical comparison. No data were normally distributed. Therefore, the Wilcoxon and Mann–Whitney U tests were used to compare the expression data between the studied groups and clinicopathological features, respectively. The correlations between genes expression in tumor samples were analyzed by the Spearman test, in which a value below 0.40 was determined as a weak correlation, 0.40–0.59 as a moderate correlation, 0.6–0.79 as a strong correlation, and ≥0.80 as a very strong correlation. Differences were considered significant at P < 0.05. In gastric cancer cell lines treated with TSA, increased or decreased mRNA expression was determined when RQ was > or < 1.0, respectively.
Results and discussion
Gene expression in tissue samples
Upregulation of class I HDAC has been described in different tumors [29-31]. High levels of HDAC genes and aberrant enzyme recruitment can repress the transcription of tumor suppressor genes, resulting in tumor onset and progression [6]. In our study, we observed that gastric tumors had increased HDAC2 expression compared with adjacent nontumor tissues (1.31 ± 0.98 vs 1.03 ± 1.08, Z = −2.23, P = 0.02; Fig. 2a). To the best of our knowledge, we have shown, for the first time, that HDAC2 mRNA levels were increased in primary GC samples, which is consistent with some previous HDAC2 protein findings [32-34].
Relative quantification of histone modifier genes in gastric samples. a HDAC2 expression, b HDAC1 expression, c HDAC3 expression, d GCN5 expression, e PCAF expression, and f comparison of PCAF expression between tumor samples classified as intestinal and diffuse histological types and their respective adjacent nontumoral samples. Data are expressed as median ± interquartile range. RQ, relative quantification; T, tumor gastric sample; N, nontumor gastric sample. *P < 0.05 (differentially expressed between groups)
In addition to repression of tumor suppressor genes by upregulation of HDACs, the loss of class I HDAC activity also contributes to cancer development by inducing oncogene expression [6]. In our study, we observed that gastric tumors had decreased HDAC1 expression compared with adjacent nontumor tissues (0.76 ± 0.61 vs 1.27 ± 1.23, Z = −2.45, P = 0.01; Fig. 2b). Consistent with the notion that HDACs can also function as a tumor suppressor, downregulation of HDAC1 has been described in colorectal cancer and leukemia [35, 36]. Moreover, we found a direct correlation between HDAC1 and CDKN1A (ρ = 0.32, P = 0.04; Table 2). A decreased expression of CDKN1A was also indentified in gastric tumors compared with adjacent nontumor tissues (1.23 ± 0.98 vs 1.45 ± 1.41; Z = −2.54; P = 0.01). CDKN1A is a cyclin-dependent kinase inhibitor implicated in cell cycle regulation [37] and was previously reported as a tumor suppressor in GC [38]. We hypothesize that downregulation of HDAC1 may contribute to the overexpression of a CDKN1A transcriptional inhibitor, resulting in CDKN1A downregulation in GC. Therefore, selective therapeutics targeting HDAC1 may be useful for induction of CDKN1A expression in this neoplasia.
In contrast to our results that showed a significant reduction in HDAC1 expression in GC, a previous study reported upregulation of HDAC1 in this neoplasia by semiquantitative RT-PCR assay, as well as by immunoblot analysis [14]. However, mRNA levels can vary among gastric samples. In our study, we observed downregulation of HDAC1 in most GC samples, but 36 % (18 out of 50) of tumors showed upregulation of this gene in relation to their matched nontumor samples. The expression of HDAC1, as well as for the other studied genes, is probably related to the cellular context and molecular background of gastric cells.
On the other hand, HDAC3 expression did not differ in gastric tumors compared with adjacent nontumor tissues (0.71 ± 0.68 vs 0.87 ± 0.74, Z = −1.90, P = 0.05; Fig. 2c). However, a significant correlation between HDAC3 and MYC was found (ρ = 0.34, P = 0.03; Table 2). An increased expression of MYC was also indentified in gastric tumors compared with adjacent nontumor tissues (1.24 ± 1.46 vs 1.0 ± 0.89, Z = −2.66, P = 0.008). Our research group has previously reported that MYC gene and protein overexpression is a common finding in GC samples from Northern Brazilian population, as well as in some preneoplastic gastric lesions [25, 39-47]. In addition to transcription activation, MYC represses gene activation by recruitment of HDAC3 [48]. Among the several factors influencing the ability of MYC to regulate expression of genes required for cell growth and proliferation, the mRNA level of HDAC3 may participate of this process in GC.
In addition to HDAC deregulation in tumors, HATs have been described to play crucial and complex roles in cancer development. GCN5 and PCAF function as cofactors for several proto-oncoproteins. We observed increased GCN5 expression in gastric tumors compared with adjacent nontumor tissues (1.38 ± 1.81 vs 1.08 ± 1.21, Z = −2.03, P = 0.04; Fig. 2d). GCN5 may promote cell growth and cancerous transformation through its association with transcription factors, such as E2F1 and MYC [13, 49-51]. Our results revealed a significant correlation between GCN5 and MYC mRNA levels (ρ = 0.31, P = 0.046; Table 2). However, considering only intestinal-type tumors, a moderate correlation was identified between the expression level of these genes (ρ = 0.46, P = 0.01), confirming the previous findings reported by our research group that MYC protein expression is more frequently observed in intestinal than diffuse-type gastric tumors [47, 52]. The oncogenic transformation carried out by MYC is not a result of the protein per se, but rather depends on its physical and functional interaction with GCN5 complexes [50]. In addition, the acetylation of MYC by GCN5 leads to an increase in protein stability [53]. Interestingly, MYC appears to affect the global chromatin structure through a direct pathway, recruiting HAT complexes to its numerous binding sites, and an indirect pathway, activating GCN5 transcription [49]. Therefore, elevated expression of GCN5 seems to be associated with MYC overexpression, a frequent finding in GC in our population [52], and it may also contribute to its oncogenic role. In the current study, we also observed that high expression of GCN5 was associated with advanced stages GC (U = 56.5, P = 0.02; Table 1) and tumor invasion (U = 153, P = 0.03; Table 1). Upregulation of GCN5 has also been reported in lung cancer, and it is associated with tumor size and a poor outcome [13]. Therefore, upregulation of GCN5 may have prognostic significance in GC.
Another HAT studied, PCAF, showed lower expression in gastric tumors compared with adjacent nontumor samples (0.76 ± 0.78 vs 1.55 ± 1.61, Z = −4.46, P < 0.001; Fig. 2e). In addition, a moderated correlation between PCAF and CDKN1A mRNA levels was detected (ρ = 0.53, P < 0.001; Table 2). PCAF plays an important role in cell cycle arrest and differentiation by activating CDKN1A expression through acetylation of the transcription factors p53 and MyoD [54-56]. Therefore, PCAF is considered a potential tumor suppressor and was found to be downregulated in a subset of intestinal-type gastric carcinoma [57, 58]. In the present study, intestinal and diffuse histological types showed downregulation of PCAF in gastric tumors compared with adjacent nontumor tissues (P < 0.001 and P = 0.01, respectively; Fig. 2f). However, the intestinal type had a significantly lower level of PCAF expression than the diffuse-type (U = 165, P = 0.03; Table 1), supporting the hypothesis that these two histological GC subtypes follow different genetic pathways and are two distinct entities [59]. In addition, reduced PCAF expression was observed in TNM stages I/II compared with that in stages III/IV (U = 180, P = 0.01; Table 1). However, PCAF expression did not differ between TNM stages III/IV and nontumor samples, suggesting that downregulation of PCAF may be a necessary step for tumor initiation.
In the present study, we detected a significant positive correlation among mRNA levels of the HDACs and HATs in tumor samples (P < 0.05; Table 2), except for between HDAC2 and PCAF expression levels. However, only HDAC1 and GCN5 mRNA levels were strongly correlated (ρ = 0.69). There is a fine balance between acetylation and deacetylation for regulating cellular function of several important acetylated factors, such as IRF, IFN, STAT, HSF, SMAD, MYC, and the E2F family of proteins. Therefore, the smallest change in this balance can alter the life span of the cell [50]. We hypothesize that a dynamic regulation of histone modifier gene expression may occur to allow the survival of GC cells and proliferation in response to different intrinsic and extrinsic factors.
Gene expression modulation by TSA treatment in GC cell lines
Some HDAC inhibitors have been used in preclinical and clinical trials in cancer therapies [4]. In our study, deregulation of HDAC1, HDAC2, GCN5, and PCAF appeared to play an important role in gastric carcinogenesis. Therefore, modulation of the expression of these HDACs and HATs may be an interesting anticancer strategy.
The ACP02 and ACP03 cell lines represent interesting models of gastric carcinogenesis in our population. Our research group previously demonstrated that these cell lines retain, in vitro, genetic alterations present in their parental primary tumors. In addition, these cell lines have genetic alterations commonly detected in GC samples [25, 60].
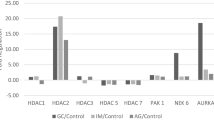
Studies have reported upregulation of HDAC genes induced by TSA treatment [7, 61]. However, our study showed a heterogeneous pattern of expression of HDAC and HAT upon treatment with TSA (Fig. 3). GCN5 was the only gene that showed increased expression in the ACP02 cell line treated with TSA compared with nontreated cells (1.21-fold; Fig. 3). While there are no studies demonstrating regulation of GCN5 by histone acetylation, we suggest a direct or indirect effect of HDAC inhibition in promoting increased transcription of this HAT gene. However, in the ACP03 cell line, downregulation of GCN5 was observed upon treatment with TSA (0.71-fold; Fig. 3), suggesting that there are different genetic and epigenetic factors controlling this gene in our GC cell lines. The other genes investigated (HDAC1, HDAC2, HDAC3, and PCAF) were downregulated in both cell lines treated with TSA compared with their respective nontreated cells, except HDAC2 in ACP02 cell line (Fig. 3). Downregulation of histone modifier genes could result from a direct effect of the absence of HDACs in histones that are insufficiently acetylated, resulting in reduction of transcription. Alternatively, hyperacetylation could result in the transcription of a regulatory factors that negatively regulates the histone modifier genes [62]. In addition, the absence of altered HDAC2 expression in ACP02 cell line treated with TSA compared with nontreated cells (1.04-fold; Fig. 3) suggests differential cell sensitivity of our cell lines to TSA.
Expression of histone deacetylase and acetyltranferase genes and the possible targets of histone modifiers, MYC and CDKN1A, in two TSA-treated gastric cancer cell lines. RQ, relative quantification of gene expression in TSA-treated cell lines calibrated by respective nontreated controls. Data are expressed as mean ± standard deviation
Considering the possible targets of histone modifiers studied here, MYC showed reduced expression in ACP02 and ACP03 cell lines upon treatment with TSA (0.85 and 0.73-fold, respectively, Fig. 3). This finding is interesting once MYC overexpression is a common finding in GC samples from Northern Brazilian population. Corroborating the observation of a moderate correlation between GCN5 and MYC in intestinal-type tumors, the marked decreased of GCN5 expression seems to collaborate with MYC downregulation in ACP03 cell line treated with TSA. On the other hand, CDKN1A demonstrated increased expression in ACP02 and ACP03 cell lines after TSA treatment (1.18 and 1.23-fold, respectively). CDKN1A is one of the major targets in the studies involving cell cycle regulation through class I HDACs inhibitors [63]. Downregulation of HDAC1 expression may play a role in the reduction of cell proliferation in ACP02 and ACP03 cell lines upon treatment with TSA (Figs. 1 and 3). The loss of HDAC1 alone or HDAC1 and HDAC2 may induce cell cycle arrest and apoptosis via hyperacetylation of the CDKN1A promoter and increased p21 levels, as demonstrated in osteosarcoma cell lines [64]. Moreover, a previous study showed that PCAF appears to participate in HDAC1 transcriptional activation [65]. Considering the ACP03 cell line, the lower level of PCAF may have contributed to the marked reduction in HDAC1 expression and increased CDKN1A expression compared with its expression in the ACP02 cell line, in which minor decrease in PCAF expression was observed.
An important limitation concerning the use of TSA as an anticancer drug is the lack of specificity [4]. Most HDAC inhibitors target a panel of HDAC superfamily members, and individual HDAC members regulate diverse cellular pathways [66]. In addition, proteins other than histones are targets of these drugs [4]. The understanding of all genetic and epigenetic factors involved in these processes and the use of this knowledge in cancer therapy are still a challenge. In the current study, we showed that TSA modulates the genes that encode HAT and HDAC enzymes, as well as MYC oncogene and the CDKN1A tumor suppressor, in two GC cell lines. These results suggest that HDACs and HATs control their own and each other’s gene expression.
Conclusions
Our results show that alteration of the histone modifier genes HDAC1, HDAC2, GCN5, and PCAF play an important role in gastric carcinogenesis, contributing to MYC and CDKN1A deregulation. PCAF downregulation is associated with intestinal-type GC, and its deregulation appears to be important in the initial stages of this disease. However, upregulation of GCN5 is associated with a poor prognosis in GC. Our findings also show that histone modifier genes HDAC1, HDAC2, HDAC3, GCN5, and PCAF, as well as MYC and CDKN1A, are modulated by TSA in GC cell lines. However, this modulation appears to be dependent of the genetic background of the cell line.
Abbreviations
- GC:
-
Gastric cancer
- HATs:
-
Histone acetyltransferases
- HDACs:
-
Histone deacetylases
- MTT:
-
(3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide, RQ, Relative quantification
- RT-qPCR:
-
Reverse transcription quantitative polymerase chain reaction
- TNM:
-
Tumor–node–metastasis
- TSA:
-
Trichostatin A
References
Greenlee RT, Hill-Harmon MB, Murray T, Thun M. Cancer statistics, 2001. CA Cancer J Clin. 2001;51(1):15–36.
Jemal A, Tiwari RC, Murray T, Ghafoor A, Samuels A, Ward E, et al. Cancer statistics, 2004. CA Cancer J Clin. 2004;54(1):8–29.
Russell MC, Mansfield PF. Surgical approaches to gastric cancer. J Surg Oncol. 2013;107(3):250–8.
Azad N, Zahnow CA, Rudin CM, Baylin SB. The future of epigenetic therapy in solid tumours-lessons from the past. Nat Rev Clin Oncol. 2013;10(5):256–66.
Legube G, Trouche D. Regulating histone acetyltransferases and deacetylases. EMBO Rep. 2003;4(10):944–7.
Ropero S, Esteller M. The role of histone deacetylases (HDACs) in human cancer. Mol Oncol. 2007;1(1):19–25.
Dangond F, Gullans SR. Differential expression of human histone deacetylase mRNAs in response to immune cell apoptosis induction by trichostatin A and butyrate. Biochem Biophys Res Commun. 1998;247(3):833–7.
Marmorstein R. Structure and function of histone acetyltransferases. Cell Mol Life Sci. 2001;58(5–6):693–703.
Liby P, Kostrouchova M, Pohludka M, Yilma P, Hrabal P, Sikora J, et al. Elevated and deregulated expression of HDAC3 in human astrocytic glial tumours. Folia Biol (Praha). 2006;52(1–2):21–33.
Armas-Pineda C, Arenas-Huertero F, Perezpenia-Diazconti M. Chico-Ponce de Leon F, Sosa-Sainz G, Lezama P, Recillas-Targa F. Expression of PCAF, p300 and Gcn5 and more highly acetylated histone H4 in pediatric tumors. J Exp Clin Cancer Res. 2007;26(2):269–76.
Xie HJ, Noh JH, Kim JK, Jung KH, Eun JW, Bae HJ, et al. HDAC1 inactivation induces mitotic defect and caspase-independent autophagic cell death in liver cancer. PLoS ONE. 2012;7(4):e34265.
Jung KH, Noh JH, Kim JK, Eun JW, Bae HJ, Xie HJ, et al. HDAC2 overexpression confers oncogenic potential to human lung cancer cells by deregulating expression of apoptosis and cell cycle proteins. J Cell Biochem. 2012;113(6):2167–77.
Chen L, Wei T, Si X, Wang Q, Li Y, Leng Y, Deng A, Chen J, Wang G, Zhu S, Kang J. Lysine acetyltransferase GCN5 potentiates the growth of non-small cell lung cancer via promotion of E2F1, Cyclin D1 and Cyclin E1 expression. J Biol Chem. 2013.
Choi JH, Kwon HJ, Yoon BI, Kim JH, Han SU, Joo HJ, et al. Expression profile of histone deacetylase 1 in gastric cancer tissues. Jpn J Cancer Res. 2001;92(12):1300–4.
Kim JK, Noh JH, Eun JW, Jung KH, Bae HJ, Shen Q, et al. Targeted inactivation of HDAC2 restores p16INK4a activity and exerts antitumor effects on human gastric cancer. Mol Cancer Res. 2013;11(1):62–73.
Sakuraba K, Yokomizo K, Shirahata A, Goto T, Saito M, Ishibashi K, et al. TIP60 as a potential marker for the malignancy of gastric cancer. Anticancer Res. 2011;31(1):77–9.
Jiang GM, Wang HS, Zhang F, Zhang KS, Liu ZC, Fang R, et al. Histone deacetylase inhibitor induction of epithelial-mesenchymal transitions via up-regulation of Snail facilitates cancer progression. Biochim Biophys Acta. 2013;1833(3):663–71.
Liu T, Kuljaca S, Tee A, Marshall GM. Histone deacetylase inhibitors: multifunctional anticancer agents. Cancer Treat Rev. 2006;32(3):157–65.
Suzuki T, Yokozaki H, Kuniyasu H, Hayashi K, Naka K, Ono S, et al. Effect of trichostatin A on cell growth and expression of cell cycle- and apoptosis-related molecules in human gastric and oral carcinoma cell lines. Int J Cancer. 2000;88(6):992–7.
Wu ZQ, Zhang R, Chao C, Zhang JF, Zhang YQ. Histone deacetylase inhibitor trichostatin A induced caspase-independent apoptosis in human gastric cancer cell. Chin Med J (Engl). 2007;120(23):2112–8.
Zou XM, Li YL, Wang H, Cui W, Li XL, Fu SB, et al. Gastric cancer cell lines induced by trichostatin A. World J Gastroenterol. 2008;14(30):4810–5.
Zhang X, Yashiro M, Ren J, Hirakawa K. Histone deacetylase inhibitor, trichostatin A, increases the chemosensitivity of anticancer drugs in gastric cancer cell lines. Oncol Rep. 2006;16(3):563–8.
Washington K. 7th edition of the AJCC cancer staging manual: stomach. Ann Surg Oncol. 2010;17(12):3077–9.
Scholte GH, van Doorn LJ, Quint WG, Lindeman J. Polymerase chain reaction for the detection of Helicobacter pylori in formaldehyde-sublimate fixed, paraffin-embedded gastric biopsies. Diagn Mol Pathol. 1997;6(4):238–43.
Leal MF, Nascimento JL M d, da Silva CE, Vita Lamarao MF, Calcagno DQ, Khayat AS, et al. Establishment and conventional cytogenetic characterization of three gastric cancer cell lines. Cancer Genet Cytogenet. 2009;195(1):85–91.
Wisnieski F, Calcagno DQ, Leal MF, Dos Santos LC, Gigek Cde O, Chen ES, Pontes TB, Assumpcao PP, de Assumpcao MB, Demachki S, Burbano RR, Smith Mde A. Reference genes for quantitative RT-PCR data in gastric tissues and cell lines. World J Gastroenterol.19(41):7121-8.
Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29(9):e45.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods (San Diego). 2001;25(4):402–8.
Jin KL, Pak JH, Park JY, Choi WH, Lee JY, Kim JH, et al. Expression profile of histone deacetylases 1, 2 and 3 in ovarian cancer tissues. J Gynecol Oncol. 2008;19(3):185–90.
Xu X, Jin H, Liu Y, Liu L, Wu Q, Guo Y, et al. The expression patterns and correlations of claudin-6, methy-CpG binding protein 2, DNA methyltransferase 1, histone deacetylase 1, acetyl-histone H3 and acetyl-histone H4 and their clinicopathological significance in breast invasive ductal carcinomas. Diagn Pathol. 2012;7:33.
Niegisch G, Knievel J, Koch A, Hader C, Fischer U, Albers P, Schulz WA. Changes in histone deacetylase (HDAC) expression patterns and activity of HDAC inhibitors in urothelial cancers. Urol Oncol. 2012.
Song J, Noh JH, Lee JH, Eun JW, Ahn YM, Kim SY, et al. Increased expression of histone deacetylase 2 is found in human gastric cancer. APMIS. 2005;113(4):264–8.
Nakagawa M, Oda Y, Eguchi T, Aishima S, Yao T, Hosoi F, et al. Expression profile of class I histone deacetylases in human cancer tissues. Oncol Rep. 2007;18(4):769–74.
Mutze K, Langer R, Becker K, Ott K, Novotny A, Luber B, et al. Histone deacetylase (HDAC) 1 and 2 expression and chemotherapy in gastric cancer. Ann Surg Oncol. 2010;17(12):3336–43.
Ozdag H, Teschendorff AE, Ahmed AA, Hyland SJ, Blenkiron C, Bobrow L, et al. Differential expression of selected histone modifier genes in human solid cancers. BMC Genomics. 2006;7:90.
Santoro F, Botrugno OA, Dal Zuffo R, Pallavicini I, Matthews GM, Cluse L, et al. A dual role for Hdac1: oncosuppressor in tumorigenesis, oncogene in tumor maintenance. Blood. 2013;121(17):3459–68.
Stivala LA, Cazzalini O, Prosperi E. The cyclin-dependent kinase inhibitor p21CDKN1A as a target of anti-cancer drugs. Current cancer drug targets.12(2):85-96.
Do Nascimento Borges B, Burbano RM, Harada ML. Absence of CIP1/KIP1 hypermethylation in gastric cancer patients from Northern Brazil. In vivo (Athens, Greece).24(4):579-82.
Calcagno DQ, Leal MF, Taken SS, Assumpcao PP, Demachki S, Smith Mde A, et al. Aneuploidy of chromosome 8 and C-MYC amplification in individuals from northern Brazil with gastric adenocarcinoma. Anticancer Res. 2005;25(6B):4069–74.
Burbano RR, Assumpcao PP, Leal MF, Calcagno DQ, Guimaraes AC, Khayat AS, et al. C-MYC locus amplification as metastasis predictor in intestinal-type gastric adenocarcinomas: CGH study in Brazil. Anticancer Res. 2006;26(4B):2909–14.
Assumpcao PP, Ishak G, Chen ES, Takeno SS, Leal MF, Guimaraes AC, et al. Numerical aberrations of chromosome 8 detected by conventional cytogenetics and fluorescence in situ hybridization in individuals from northern Brazil with gastric adenocarcinoma. Cancer Genet Cytogenet. 2006;169(1):45–9.
Calcagno DQ, Leal MF, Seabra AD, Khayat AS, Chen ES, Demachki S, et al. de Arruda Cardoso Smith M, Burbano RR. Interrelationship between chromosome 8 aneuploidy, C-MYC amplification and increased expression in individuals from northern Brazil with gastric adenocarcinoma. World J Gastroenterol. 2006;12(38):6207–11.
Calcagno DQ, Leal MF, Assumpcao PP, Smith MA, Burbano RR. MYC and gastric adenocarcinoma carcinogenesis. World J Gastroenterol. 2008;14(39):5962–8.
Costa Raiol LC, Figueira Silva EC, Mendes da Fonseca D, Leal MF, Guimaraes AC, Calcagno DQ, et al. Interrelationship between MYC gene numerical aberrations and protein expression in individuals from northern Brazil with early gastric adenocarcinoma. Cancer Genet Cytogenet. 2008;181(1):31–5.
Calcagno DQ, Guimaraes AC, Leal MF, Seabra AD, Khayat AS, Pontes TB, et al. MYC insertions in diffuse-type gastric adenocarcinoma. Anticancer Res. 2009;29(7):2479–83.
Calcagno DQ, Leal MF, Demachki S, Araujo MT, Freitas FW, Oliveira e Souza D, et al. MYC in gastric carcinoma and intestinal metaplasia of young adults. Cancer Genet Cytogenet. 2010;202(1):63–6.
Calcagno DQ, Freitas VM, Leal MF, de Souza CR, Demachki S, Montenegro R, Assumpcao PP, Khayat AS, Smith Mde A, dos Santos AK, Burbano RR. MYC, FBXW7 and TP53 copy number variation and expression in gastric cancer. BMC gastroenterology.13:141.
Kurland JF, Tansey WP. Myc-mediated transcriptional repression by recruitment of histone deacetylase. Cancer Res. 2008;68(10):3624–9.
Knoepfler PS, Zhang XY, Cheng PF, Gafken PR, McMahon SB, Eisenman RN. Myc influences global chromatin structure. EMBO J. 2006;25(12):2723–34.
Nagy Z, Tora L. Distinct GCN5/PCAF-containing complexes function as co-activators and are involved in transcription factor and global histone acetylation. Oncogene. 2007;26(37):5341–57.
Kenneth NS, Ramsbottom BA, Gomez-Roman N, Marshall L, Cole PA, White RJ. TRRAP and GCN5 are used by c-Myc to activate RNA polymerase III transcription. Proc Natl Acad Sci U S A. 2007;104(38):14917–22.
de Souza CR, Leal MF, Calcagno DQ, Costa Sozinho EK, Borges Bdo N, Montenegro RC, et al. de Arruda Cardoso Smith M, Burbano RR. MYC deregulation in gastric cancer and its clinicopathological implications. PLoS One. 2013;8(5):e64420.
Patel JH, Du Y, Ard PG, Phillips C, Carella B, Chen CJ, et al. The c-MYC oncoprotein is a substrate of the acetyltransferases hGCN5/PCAF and TIP60. Mol Cell Biol. 2004;24(24):10826–34.
Sakaguchi K, Herrera JE, Saito S, Miki T, Bustin M, Vassilev A, et al. DNA damage activates p53 through a phosphorylation-acetylation cascade. Genes Dev. 1998;12(18):2831–41.
Liu L, Scolnick DM, Trievel RC, Zhang HB, Marmorstein R, Halazonetis TD, et al. p53 sites acetylated in vitro by PCAF and p300 are acetylated in vivo in response to DNA damage. Mol Cell Biol. 1999;19(2):1202–9.
Sartorelli V, Puri PL, Hamamori Y, Ogryzko V, Chung G, Nakatani Y, et al. Acetylation of MyoD directed by PCAF is necessary for the execution of the muscle program. Mol Cell. 1999;4(5):725–34.
Schiltz RL, Nakatani Y. The PCAF acetylase complex as a potential tumor suppressor. Biochim Biophys Acta. 2000;1470(2):M37–53.
Ying MZ, Wang JJ, Li DW, Yu GZ, Wang X, Pan J, Chen Y, He MX. The p300/CBP associated factor: is frequently downregulated in intestinal-type gastric carcinoma and constitutes a biomarker for clinical outcome. Cancer Biol Ther. 2010;9(4).
Tahara E. Genetic pathways of two types of gastric cancer. IARC Sci Publ. 2004;157:327–49.
Leal MF, Calcagno DQ. Borges da Costa Jde F, Silva TC, Khayat AS, Chen ES, Assumpcao PP, de Arruda Cardoso Smith M, Burbano RR. MYC, TP53, and chromosome 17 copy-number alterations in multiple gastric cancer cell lines and in their parental primary tumors. J Biomed Biotechnol. 2011;2011:631268.
Gray SG, Ekstrom TJ. Effects of cell density and trichostatin A on the expression of HDAC1 and p57Kip2 in Hep 3B cells. Biochem Biophys Res Commun. 1998;245(2):423–7.
Glaser KB, Staver MJ, Waring JF, Stender J, Ulrich RG, Davidsen SK. Gene expression profiling of multiple histone deacetylase (HDAC) inhibitors: defining a common gene set produced by HDAC inhibition in T24 and MDA carcinoma cell lines. Mol Cancer Ther. 2003;2(2):151–63.
Spiegel S, Milstien S, Grant S. Endogenous modulators and pharmacological inhibitors of histone deacetylases in cancer therapy. Oncogene.31(5):537-51.
Senese S, Zaragoza K, Minardi S, Muradore I, Ronzoni S, Passafaro A, et al. Role for histone deacetylase 1 in human tumor cell proliferation. Mol Cell Biol. 2007;27(13):4784–95.
Schuettengruber B, Simboeck E, Khier H, Seiser C. Autoregulation of mouse histone deacetylase 1 expression. Mol Cell Biol. 2003;23(19):6993–7004.
Gray SG, Ekstrom TJ. The human histone deacetylase family. Exp Cell Res. 2001;262(2):75–83.
Acknowledgments
This paper was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP; to MACS, FW, DQC, MFL, and TBP), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq; to MCS and RRB), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES; to COG).
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wisnieski, F., Calcagno, D.Q., Leal, M.F. et al. Differential expression of histone deacetylase and acetyltransferase genes in gastric cancer and their modulation by trichostatin A. Tumor Biol. 35, 6373–6381 (2014). https://doi.org/10.1007/s13277-014-1841-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-1841-0