Abstract
Cerebral palsy (CP) is a non-progressive neurological disease, of which susceptibility is linked to genetic and environmental risk factors. More and more studies have shown that CP might be caused by multiple genetic factors, similar to other neurodevelopmental disorders. Due to the high genetic heterogeneity of CP, we focused on investigating related molecular pathways. Ten children with CP were collected for whole-exome sequencing by next-generation sequencing (NGS) technology. Customized processes were used to identify potential pathogenic pathways and variants. Three pathways (axon guidance, transmission across chemical synapses, protein–protein interactions at synapses) with twenty-three genes were identified to be highly correlated with CP. This study showed that the three pathways associated with CP might be the molecular mechanism of pathogenesis. These findings could provide useful clues for developing pathway-based pharmacotherapies. Further studies are required to confirm potential roles for these pathways in the pathogenesis of CP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cerebral palsy is a neurological disorder with multiple clinical types, such as abnormal movement, stiff muscles, weak muscles, and tremors. Causes and risk factors for CP include: (1) Before pregnancy: premature delivery, intrauterine distress, multiple pregnancy, poor pregnancy, intrauterine infection, intrauterine infection, maternity abnormalities, pregnancy and high disease, abnormal glucose tolerance, maternal age, pregnancy infection, pregnancy trauma, placenta previa, placental abruption, premature rupture of membranes, umbilical cord around the neck, oligoamnios, family history, test-tube baby, maternal chronic diseases. (2) During pregnancy: timing asphyxia, meconium aspiration, amniotic fluid turbidity, prolonged labor, midwifery, inhalation pneumonia, swallowing syndromes. (3) After delivery: postpartum asphyxia, jaundice, assisted ventilation, intracranial hemorrhage, neonatal infection, encephalitis, neonatal convulsion, hydrocephalus, craniocerebral trauma. The main causes of CP are premature birth, choking, jaundice, premature rupture of membranes, miscarriage history, poor pregnancy, intracranial hemorrhage, multiple pregnancies, intrauterine distress. Among these risk factors, premature birth is the top one, followed by choking, jaundice, premature rupture of membranes and poor pregnancy (including risk factors before pregnancy include intrauterine distress, premature birth, low weight, and poor pregnancy, premature rupture of membranes, miscarriage, intrauterine infection, etc.). More and more CP prevention and treatment studies are focused on these key risk factors. However, although many risk factors and well-characterized causes have been clearly put forward, the cause for many children with CP has still remained obscure. Recent studies suggest many cases of CP are associated with genetic alterations (Fahey et al. 2017; MacLennan et al. 2015; McMichael et al. 2015).
In a recent study, several genes have been identified as susceptibility or disease-causing gene using next-generation sequencing. Approximately 14% of 200 unrelated CP cases were reported to have a pathogenic genetic variation (McMichael et al. 2015). Analogous to the autistic spect rum disorders (Catalina Betancur 2011), the genomic architecture of CP may be highly complex, with many genetic and genomic disorders involved. Up to date, studies about genetic variations in CP, either previous CNV (Segel et al. 2015) or WES (McMichael et al. 2015), suggested considerable genetic heterogeneity of CP. It is considered to have varying effects of genetic variations (Fahey et al. 2017). A severely deleterious variation may have a major impact and become sufficient to cause CP in some individuals. A less damaging variation that does not disrupt protein function profoundly may have a minor impact, leading to clinical CP with cumulative effect. Considering the considerable genetic heterogeneity and the varying effects of genetic variations, we thought it should focus our attention on molecular pathways relevant to CP.
In this study, customized filtering processes at the variant level and gene level were applied in turn to obtain the potential genes. We then discovered statistically significantly enriched pathways using the most promising genes. Finally, we identified three molecular pathways and the subset of candidate genes that may contribute to CP.
Materials and methods
Sample collection
In total ten children (eight unrelated individuals and a pair of twins) with CP were selected from our children’s rehabilitation center. The diagnosis of CP patients is determined by a pediatrician physician through a review of clinical examinations or related medical records. Neurodevelopmental retardation (MR) is determined primarily based on a score of < 70 for the evaluation of psychometric indicators. The definition, diagnostic criteria and typing criteria of CP were established according to the 2nd China National Conference of Pediatric Rehabilitation, 9th China National Conference on Rehabilitation of Children with Cerebral Palsy and International Exchange Conference.
The diagnostic criteria for CP are as follows:
-
1.
Dyskinesia is caused by brain damage or developmental defects.
-
2.
Encephalopathy is non-progressive.
-
3.
Symptoms appear in infancy.
-
4.
Phenotypes can be combined with mental retardation, epilepsy, sensory disturbance, communication disorders, behavioral abnormalities and other abnormalities. Patients may have secondary bone and muscle system damage.
-
5.
Progressive disease caused by central dyskinesia and normal pediatric relativistic exercise retardation should be checked carefully and excluded.
Exome sequencing and analysis
For all individuals, whole-blood DNA was enriched for whole-exome sequencing using Agilent SureSelect 50 Mb Human All Exon Kit. The captured fragments were amplified and sequenced using an Illumina HiSeq X Ten with 150 bp paired-end mode. The sequencing reads from WES were aligned to the human genome (NCBI Build37/hg19) with Burrow-Wheeler Aligner (BWA) (Li and Durbin 2010). In order to reduce the duplication which may result from PCR amplification, we marked and removed the duplication using Picard. As a result, high quality data were obtained with 100× average sequencing depth, and 92% target regions had at least 20× depth of coverage. Applications in Genome Analysis Toolkit (GATK) (McKenna et al. 2010) were used to local realign based on indels from 1000 Genomes (Genomes Project et al. 2012). To improve the veracity and reliability, BaseRecalibrator from GATK was used to adjust the base quality score based on dbSNP database (Sherry et al. 2001). And then, we called variants using HaplotypeCaller from GATK with default parameters.
Variant annotation and filtering
A widely annotation software ANNOVAR (Wang et al. 2010) was used to annotate variants with allelic frequencies, inheritance model, pathogenicity, functionality. Many databases, including population databases (dbSNP, Exome Aggregation Consortium 1000 Genomes Project, Exome Variant Server, and Complete Genomics) (Genomes Project et al. 2012; Lek et al. 2016; Sherry et al. 2001), prediction database (dbNSFP, dbscSNV, REVEL, and M-CAP) (Ioannidis et al. 2016; Jagadeesh et al. 2016; Jian et al. 2014; Liu et al. 2011) and disease databases (ClinVar and OMIM) (Hamosh et al. 2005; Landrum et al. 2014), were applied in this annotation. Pathogenicity of variants were predicted based on the ClinVar database and ACMG Standards and Guidelines (Richards et al. 2015). A filtering process was customized for removing likely non-pathogenic variants.
-
1.
Variants with minor allele observations (AO) < 5;
-
2.
Variants with minor allele frequency (MAF) greater than 0.05 in East Asians based on either Exome Aggregation Consortium or 1000 Genomes Project;
-
3.
Variants with pathogenicity predicted as benign or likely benign;
-
4.
Variants with function is synonymous;
-
5.
Variants in intronic or intergenic regions;
-
6.
Indels in short tandem repeats (STR) regions from UCSC (Fujita et al. 2011).
Gene mining
In order to obtain the pathogenic variations associated with the CP phenotype as accurately as possible, we screened the genes from three aspects: gene specific expression, phenotypic correlation, and the tolerance of a gene to amino acid-changing. Brain, as the central nervous system, is the core of perception, motion and cognition. Transcriptomic analysis showed that 74% (n = 14,518) of all human genes (n = 19,613) were expressed in brain, among which 1460 genes exhibited an elevated expression in brain compared to other tissue types (Uhlen et al. 2010). What we focused on was variants in these 1460 genes with elevated expression in the brain. Next, we adopted a web tool Phenolyzer (http://phenolyzer.wglab.org) (Yang et al. 2015) to obtain genes correlated with CP, using “cerebral palsy” as keyword. Phenolyzer, integrating multiple phenotypic-related databases (OMIM, HPO, CTD, etc.), could generate scores for predicting gene-phenotype correlations based on pathway and protein interaction analysis. Recent study suggested genes correlated with mendelian disorders had less tolerance to functional genetic variations compared to those not correlated with any known diseases (Petrovski et al. 2013). Residual variation intolerance score (RVIS) was used to eliminate the high tolerance genes. According to the genic intolerance RVIS percentiles in ExAC, genes which score less than 50 percent were retained for further analysis.
Pathway annotation
Remained genes across eight unrelated individuals were annotated to Reactome (http://www.reactome.org) (Joshi-Tope et al. 2005) pathway for finding susceptibility pathways correlation with CP. p values < 0.05 were considered statistically significant.
Results
Patients
Screening tests for common inherited metabolic diseases such as phenylketonuria (PKU) and amino acid metabolism abnormalities have been carried out by tandem mass spectrometry in these ten children with CP. The possibility of the chromosomal abnormality in children has been ruled out by chromosome analysis (karyotyping) or gene chip technology, and the condition is relatively static. Relatives of ten children have denied “injury” history, denied “virus bacterial infection medication” history, denied “poison, radioactive material, pet” contact history, denied “diabetes” and other medical histories. The general information and clinical manifestations of ten children (nine and ten are identical twins) with CP are shown in Table 1, and brain magnetic resonance images (MRIs) are shown in Fig.S1.
Discovery cohort
After running the quality filters application of GATK, 156484 SNVs and 26547 indels resulted across eight samples with WES (85401–87998 variants per sample). As described by standards and guidelines for validating next-generation sequencing bioinformatics pipelines (Roy et al. 2018), variants representing false positive amplification of the NGS method should be filtered, and clinically insignificant variants and neutral substitutions should not be considered at first. In order to remove variants with a high-frequency in normal populations and reducing the impact of artificial error, the filter was used based on allele frequency and align depth (Variant annotation and filtering in the materials and methods). We next removed benign variants and likely benign variants based on pathogenicity from ClinVar database and ACMG. Variants within an exon and outside the STR domain from UCSC were retained. Through a series of filters, approximately 2% of the raw variants and 2840 genes remained among eight unrelated individuals.
Similar to recent years’ studies (Eilbeck et al. 2017; Farlow et al. 2016), we searched for genes from aspects of tissue specific gene expression, correlation between gene and phenotype, tolerance of a gene to amino acid-changing or loss‑of‑function (LOF) variation, and pathway enrichment analysis. We considered the tissue specificity of CP and obtained 181 genes that are elevated in the brain compared to other tissue types. 1437 of human proteins that show an elevated expression in the brain are from the human protein atlas (Uhlen et al. 2015) (http://www.proteinatlas.org). Phenolyzer, a tool focusing on discovering genes based on specific disease/phenotype terms, was used to acquire genes associated with CP, and then we got 95 genes. Genic intolerance RVIS percentiles were computed to reflect the possible effects of variations at the gene level and filtered to discard those more tolerant genes. After the above filtration steps, 63 genes were retained (Fig. 1). Among these genes, twins identified 5 genes (BAIAP3, DNM1, CDH9, EPHA5, GRIK3).
Customized filter process to obtain variant genes associated with CP. The filtering process included two data processing strategies, variant filtering and gene filtering. The variant filtering was targeted at false positive artifacts and clinically insignificant variants, based on a rich set of metadata, such as minor allele frequencies in human populations, known or predicted pathogenicity, predicted amino acid sequence change. The gene filtering included tissue-specific expression, gene-phenotype correlations, and gene tolerance to amino acid-changing
Pathway annotation
Taking into account the high genetic heterogeneity of CP, we looked for the same pathways that might be caused by different variations. We used a pathway enrichment analysis for the biological interpretation of 63 genes across eight unrelated individuals in the system level and identified two top-level significantly enriched pathways (developmental biology with p = 0.0012 and neuronal system with p = 8.11E−04). Variant genes from the twins were also confirmed to affect these two biological pathways (Table 2).
Among the sub-pathways of developmental biology, axon guidance showed a significance level (p = 2.32E−7). The role of variant genes in Axon guidance is shown in Fig. 2. Axon guidance is the process by which neurons send out axons to reach the correct targets. Many highly conserved families of guidance molecules and their receptors that guide axons were identified to be affected, such as L1CAM interactions (p = 0.0014), netrin-1 signaling (p = 0.0047). Among the sub-pathways of neuronal system, transmission across chemical synapses showed a significance level (p = 0.0107). Chemical synapses are specialized junctions that are used for communication between neurons, neurons and muscle or gland cells. Among these communications, GABA A receptor activation was significantly enriched. Due to the high heterogeneity and complexity of CP, we believe that multiple genes may lead to the corresponding phenotype. Table 2 shows the pathways affected by the variant genes in 10 children with CP. Except case 3, others all had multiple variants in either axon guidance or transmission across chemical synapses pathway. Although case 3 had only one gene DSCAML1 affected axon guidance pathway, he also had another gene, DLGAP3, which affected the sub-pathway of neuronal system, protein–protein interactions at synapses. Finally, three secondary pathways (axon guidance, transmission across chemical synapses, protein–protein interactions at synapses) with 23 candidate genes Table 3 identified may contribute to CP risk based on 8 unrelated individuals and a pair of twins. All variants remained were nonsynonymous SNV and the genotype was heterozygous. While Sanger sequencing was also used to validate the variations of the 23 candidate genes. Single nucleotide variants deleteriousness assessment tool—CADD (Combined Annotation-Dependent Depletion) scores (Kircher et al. 2014), and genes intolerant of loss-of-function scores—pLI (probability that a gene is intolarent to a loss of function mutation), assisted in evaluating the deleteriousness for these variants. Variants which CADD priority score > 20 are considered functional, deleterious, and disease causal variants and genes which RVIS < 25 or pLI ≥ 0.9 are likely intolerant of loss-of-function variation Fig. 3.
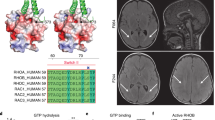
Reactome diagram of axon guidance(Joshi-Tope et al. 2005). The gradient yellow box on the left indicates the degree of gene enrichment. The more yellow the color, the higher the degree of enrichment is. Yellow boxes with square corners are genes we focused on. The role of genes in pathways are as follows: (1) NCAM signaling for neurite out-growth. NCAM participates in intracellular signal transduction cascades, and contribute to neural differentiation and synaptic plasticity. Fyn associates with the 140 kD isoform of NCAM1 in the plasma membrane. SPTBN4, an NCAM1 binding cytoskeletal protein, associates with the activator of Fyn kinase (RPTP-alpha) and activates Fyn. Fyn activation leads to the recruitment and activation of the non-receptor tyrosine kinase FAK. FAK undergo autophosphorylation and multiple tyrosine residues phosphorylation in a Src-dependent manner. Phosphorylated tyrosine 925 recruits the GRB2/SOS complex to activate Ras and initiate Ras-MAPK signaling and acts on neurite outgrown finally. (2) Netrin-1 signaling. Netrins play a key role in neuronal migration and axon guidance. The transmembrane DCC receptor is a Netrin-1 receptor, directly binds to Netrin-1, which induces DCC clustering. FADK1 interacts with DCC and undergoes tyrosine phosphorylation. Activated (phosphorylated) FADK1 recruits src tyrosine kinases Src and Fyn to DCC and leads to the phosphorylation of DCC. DCC interacts with NCK1, which can recruit Rac, Cdc42, Pak and N-WASP to the activated receptor. Thereby mediates Netrin-1 signaling in axon outgrowth and guidance. DSCAM is a cell surface receptor, likely mediates axon pathfinding. AGAP2 interacts with UNC5B to prevents its proapoptotic activity and improves neuronal survival. ROBO interants with its ligand SLIT3 and regulates the midline crossing of axons. (3) Signaling by ROBO receptors. SLIT-ROBO signaling not only involves in axon repulsion, but also involves in cortical dendrite branching. One of the regulatory mechanism is that inhibits the activity of CDC42 and RAC1 by binding to SRGAP, thereby regulates cytoskeletal dynamics of axon guidance receptors. (4) L1CAM interactions. The L1CAM family involves crucial processes in nervous system development. Ankyrins (include ANK1) bind to L1 CAMs and link L1 CAMs/ion channel proteins to the spectrin cytoskeleton (include SPTBN4). This immobilization conduces to adjacent neuron adhesions. The actin spectrin network communicates by voltage gated sodium channels, voltage gated potassium ion channel subunits (KCNQ2 and KCNQ3) and voltage gated sodium ion channel subunits (include SCN3A). L1 binds to NCAM1 and enhances the effect of cell aggregation and adhesion. Neurofascin functions as cell adhesion molecule contributed to axon subcellular targeting and synapse formation during neural development. CNTNAP1 combines with neurofascin form the core structure of paranodal junctions. (5) EPH-Ephrin signaling. During the neural development, correct cell migration direction is important. Hepatocellular carcinoma (include EPHA5) receptors and their ligands (EFNs) take part in the precise control to guide a cell to its destination. EPHA activation by EFNA enhances the catalytic activity of NGEF and increases the RHOA, ROCK activation in cortical neurons resulting in growth cone collapse. (6) RET signaling. RET is essential in axonal growth and axon guidance of developing enteric neurons, motor neurons. REF activates the RAS-RAF-ERK signaling pathway by binds to GFRAs, GDNFs, DOKs, SHC1, GRB2, SOS1 successively and phosphorylation. Also, it’s complex can bind to Rap1GAP to suppresses GDNF-induced activation of ERK and neurite outgrowth
Variants where both CADD, RVIS and pLI were available (n = 24). Different variants identified in DSCAML1 and RAP1GAP respectively are labeled DSCAML1-1 (NM_020693:c.5919G > C), DSCAML1-1 (NM_020693:c.4574G > A), DSCAML1-1 (NM_020693:c.1243G > T), RAP1GAP-1 (NM_001145658:c.1892G > A), RAP1GAP-2 (NM_001145658:c.788A > G)
Discussion
Using WES in eight individuals and a pair of twins, we detected three reactome pathways (axon guidance, transmission across chemical synapses, protein–protein interactions at synapses) that may have a role in CP susceptibility. 23 candidate CP genes participated in these three pathways may contribute to CP risk in eight unrelated individuals and a pair of twins in our study.
Most clinical studies expected to obtain key genes, either susceptible or symptomatic ones, contributing to the pathogenesis of CP. Six genes with variations that included GAD1, KANK1, AP4M1, AP4E1, AP4B1 and AP4S1 are known to cause mendelian forms of CP (Moreno-De-Luca et al. 2012). Research with large samples shows that both maternally inherited micro-duplication of 7q21.13 contained ZNF804B, MGC26647, STEAPl and micro-duplication of 14q23.1 involving DAAM1 gene with uncertain mode of inheritance may cause girl’s spastic diplegia. Studies find that AP4B1 is most closely associated with CP in Chinese CP patients (McMichael et al. 2014; Wang et al. 2013). Variation of GAD1 gene was an important risk factor for mixed CP (Lin et al. 2013). A homozygous p.G367D variation in ADD3, encoding gamma adducing, cause hereditary CP with leading to both abnormal proliferation and migration in cultured patient fibroblasts. Similar to adducin, actin KANK1 also play an integral role in hereditary CP (Kruer et al. 2013). Both ataxia-telangiectasia with novel splicing variations in the ATM gene, thiamine transporter deficiency with the variation in SLC19A3 and biotinidase deficiency caused by BTD genetic abnormality may cause ataxia and movement disorder (Jeong et al. 2014; Kasapkara et al. 2015; Vernau et al. 2015).
These susceptible genes can explain the etiology of some patients, but the vast majority of the genetic causes of CP patients cannot be found. In particular, most patients with CP present a genetic model of the complex disease. It is not comprehensive to determine the common genetic factors of CP patients through susceptibility gene. Due to the huge genetic heterogeneity of CP, these studies lacked strong repeatable verification results, and GWAS research based on the common disease-common variant theory may not be applicable to CP genetic research.
Prior to WES, all children with CP were detected for CNV, based on high-throughput whole genome sequencing technology with low depth. Using whole genome data of multiple healthy people as a reference, the coverage depth was calculated by the sliding window for CNV detection. Since no certain CNVs related to CP were found, we did not show the results. With SNVs and indels analysis, two twins and eight CP detection results showed that two biological pathways involving 23 genes had a common effect. This suggests that CP is not limited to partial genetic variations, and that biological pathways play a role in the pathogenesis of CP. We tend to examine the effects of biological pathways on CP, and other studies of susceptibility genes may be part of the biological pathway, such as glutamate receptor (m Glu Rs), gamma-aminobutyric acid receptor (gamma-GABA) of protein coupled receptors (GPCRs).
As a result, three significantly enriched subpathways of axon guidance and one significantly enriched subpathways of transmission across chemical synapses were identified associated with the disorder. As the participant of axon guidance, L1CAM interactions and netrin-1 signaling were significantly enriched with p-value of 0.0014, 0.0047 respectively. The L1CAM family has been implicated in processes integral to nervous system development, including neurite outgrowth, neurite fasciculation and inter neuronal adhesion. Recent genetic studies support synaptic CAMs were associated with developmental delay and ASD as causal genes or potential susceptibility genes (Betancur et al. 2009). L1CAM, PAK3 and TUBA1A involved in axonal guidance signaling have been reported in patients with CP (McMichael et al. 2015). During the development of the nervous system, netrins play crucial roles in neuronal migration and in axon guidance.
As a participant of transmission across chemical synapses, GABA A receptor activation(GABRG2, GABRB1, GABRA2) is the pathway that most genes annotated with p = 0.0017. GABA A receptor mediated fast synaptic inhibition in the human brain. Many neurological and psychiatric diseases include epilepsy and schizophrenia are considered to be related to GABA A receptor dysfunction. GABA is an excitatory neurotransmitter that acts on GABA locus on GABAA receptors and increases the permeability of chloride ions in the membranes of neurons. The concentration of the extracellular chloride ions is higher than the concentration of chloride ions in the cell. Chloride ions enter the cell by the concentration gradient, resulting in the increase of the cell intima potential and the hyperpolarization, and the excitability also decreases accordingly. GABA inhibits amino acid-induced excitotoxicity, similar to glutamate excitotoxicity, which plays a role in the neuroprotection of ischemic brain injury (Nunez and McCarthy 2004). For clinical treatment, it may be possible to consider targeted use of GABAA receptor antagonists for the treatment and prevention of perinatal brain injury. Other biological pathways to avoid CP can also be considered from the increased expression of EphA receptor inhibits axonal regeneration and induces apoptosis in central nervous system (Nolt et al. 2011). It provides a new idea for targeted prevention and salvage treatment of children born.
Previous studies (McMichael et al. 2015) identified 61 de novo variants, 19 X-chromosome maternally inherited variants, and 4 compound heterozygous variants in 98 case-parent trios. About these 80 genes, 8 genes (TUBA1A, ABLIM2, SCN8A, MAST1, UPF3B, L1CAM, EPHA1, PAK3) take part in axon guidance pathway, 3 genes (SIPA1L1, SLITRK2, IL1RAPL1) participate in protein–protein interactions at synapses and 2 genes (MAOB, ADCY3) are involved in transmission across chemical synapses. In their predicted causative CP genes (n = 14), three genes (L1CAM, PAK3, TUBA1A) were identified (p = 0.006) as being involved in axon guidance. Compared with this study, we additionally identified two pathways related to CP (transmission across chemical synapses, protein–protein interactions at synapses) with 23 related genes. One major limitation of the present study was the small sample size, with only 10 cases, and lacking of function verification. Thus the clinical studies including larger samples on CP and function verification will be a further research direction. Through the above studies, we believe it is a feasible solution that for parents of children with CP, it is necessary to carry out the whole-exome gene sequencing for children to assess the risk of reproduction and to take targeted prevention throughout pregnancy.
Conclusions
We used the customized filter process for variants and genes in turn to identify pathways and genes that may lead to CP. Three molecular pathways (axon guidance, transmission across chemical synapses, protein–protein interactions at synapses) and 23 genes with potential larger effects on CP were found. The potential impact of pathways and genes nominated in this study need to be further explored and studied.
References
Betancur C (2011) Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res 1380:42–77. https://doi.org/10.1016/j.brainres.2010.11.078
Betancur C, Sakurai T, Buxbaum JD (2009) The emerging role of synaptic cell-adhesion pathways in the pathogenesis of autism spectrum disorders. Trends Neurosci 32(7):402–412. https://doi.org/10.1016/j.tins.2009.04.003
Eilbeck K, Quinlan A, Yandell M (2017) Settling the score: variant prioritization and Mendelian disease. Nat Rev Genet 18(10):599–612. https://doi.org/10.1038/nrg.2017.52
Fahey MC, Maclennan AH, Kretzschmar D, Gecz J, Kruer MC (2017) The genetic basis of cerebral palsy. Dev Med Child Neurol 59(5):462–469. https://doi.org/10.1111/dmcn.13363
Farlow JL, Robak LA, Hetrick K, Bowling K, Boerwinkle E, Coban-Akdemir ZH et al (2016) Whole-exome sequencing in familial Parkinson disease. JAMA Neurol 73(1):68–75. https://doi.org/10.1001/jamaneurol.2015.3266
Fujita PA, Rhead B, Zweig AS, Hinrichs AS, Karolchik D, Cline MS et al (2011) The UCSC Genome Browser database: update 2011. Nucleic Acids Res 39(Database issue):D876–D882. https://doi.org/10.1093/nar/gkq963
Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM et al (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491(7422):56–65. https://doi.org/10.1038/nature11632
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA (2005) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33(Database issue):D514–D517. https://doi.org/10.1093/nar/gki033
Ioannidis NM, Rothstein JH, Pejaver V, Middha S, McDonnell SK, Baheti S et al (2016) REVEL: an ensemble method for predicting the pathogenicity of rare missense variants. Am J Hum Genet 99(4):877–885. https://doi.org/10.1016/j.ajhg.2016.08.016
Jagadeesh KA, Wenger AM, Berger MJ, Guturu H, Stenson PD, Cooper DN et al (2016) M-CAP eliminates a majority of variants of uncertain significance in clinical exomes at high sensitivity. Nat Genet 48(12):1581–1586. https://doi.org/10.1038/ng.3703
Jeong H, Huh HJ, Youn J, Kim JS, Cho JW, Ki CS (2014) Ataxia-telangiectasia with novel splicing mutations in the ATM gene. Ann Lab Med 34(1):80–84. https://doi.org/10.3343/alm.2014.34.1.80
Jian X, Boerwinkle E, Liu X (2014) In silico prediction of splice-altering single nucleotide variants in the human genome. Nucleic Acids Res 42(22):13534–13544. https://doi.org/10.1093/nar/gku1206
Joshi-Tope G, Gillespie M, Vastrik I, D’Eustachio P, Schmidt E, de Bono B et al (2005) Reactome: a knowledgebase of biological pathways. Nucleic Acids Res 33(Database issue):D428–D432. https://doi.org/10.1093/nar/gki072
Kasapkara CS, Akar M, Ozbek MN, Tuzun H, Aldudak B, Baran RT, Tanyalcin T (2015) Mutations in BTD gene causing biotinidase deficiency: a regional report. J Pediatr Endocrinol Metab 28(3–4):421–424. https://doi.org/10.1515/jpem-2014-0056
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J (2014) A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 46(3):310–315. https://doi.org/10.1038/ng.2892
Kruer MC, Jepperson T, Dutta S, Steiner RD, Cottenie E, Sanford L et al (2013) Mutations in gamma adducin are associated with inherited cerebral palsy. Ann Neurol 74(6):805–814. https://doi.org/10.1002/ana.23971
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, Maglott DR (2014) ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42(Database issue):D980–D985. https://doi.org/10.1093/nar/gkt1113
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T et al (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536(7616):285–291. https://doi.org/10.1038/nature19057
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5):589–595. https://doi.org/10.1093/bioinformatics/btp698
Lin S, Li T, Zhu D, Ma C, Wang Y, He L et al (2013) The association between GAD1 gene polymorphisms and cerebral palsy in Chinese infants. Cytol Genet 47(5):276–281. https://doi.org/10.3103/s0095452713050071
Liu X, Jian X, Boerwinkle E (2011) dbNSFP: a lightweight database of human nonsynonymous SNPs and their functional predictions. Hum Mutat 32(8):894–899. https://doi.org/10.1002/humu.21517
MacLennan AH, Thompson SC, Gecz J (2015) Cerebral palsy: causes, pathways, and the role of genetic variants. Am J Obstet Gynecol 213(6):779–788. https://doi.org/10.1016/j.ajog.2015.05.034
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
McMichael G, Girirajan S, Moreno-De-Luca A, Gecz J, Shard C, Nguyen LS et al (2014) Rare copy number variation in cerebral palsy. Eur J Hum Genet 22(1):40–45. https://doi.org/10.1038/ejhg.2013.93
McMichael G, Bainbridge MN, Haan E, Corbett M, Gardner A, Thompson S et al (2015) Whole-exome sequencing points to considerable genetic heterogeneity of cerebral palsy. Mol Psychiatry 20(2):176–182. https://doi.org/10.1038/mp.2014.189
Moreno-De-Luca A, Ledbetter DH, Martin CL (2012) Genetic insights into the causes and classification of the cerebral palsies. Lancet Neurol 11(3):283–292. https://doi.org/10.1016/s1474-4422(11)70287-3
Nolt MJ, Lin Y, Hruska M, Murphy J, Sheffler-Colins SI, Kayser MS et al (2011) EphB controls NMDA receptor function and synaptic targeting in a subunit-specific manner. J Neurosci 31(14):5353–5364. https://doi.org/10.1523/JNEUROSCI.0282-11.2011
Nunez JL, McCarthy MM (2004) Cell death in the rat hippocampus in a model of prenatal brain injury: time course and expression of death-related proteins. Neuroscience 129(2):393–402. https://doi.org/10.1016/j.neuroscience.2004.08.006
Petrovski S, Wang Q, Heinzen EL, Allen AS, Goldstein DB (2013) Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Genet 9(8):e1003709. https://doi.org/10.1371/journal.pgen.1003709
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 17(5):405–424. https://doi.org/10.1038/gim.2015.30
Roy S, Coldren C, Karunamurthy A, Kip NS, Klee EW, Lincoln SE et al (2018) Standards and guidelines for validating next-generation sequencing bioinformatics pipelines: a joint recommendation of the Association for Molecular Pathology and the College of American Pathologists. J Mol Diagn 20(1):4–27. https://doi.org/10.1016/j.jmoldx.2017.11.003
Segel R, Ben-Pazi H, Zeligson S, Fatal-Valevski A, Aran A, Gross-Tsur V et al (2015) Copy number variations in cryptogenic cerebral palsy. Neurology 84(16):1660–1668. https://doi.org/10.1212/wnl.0000000000001494
Sherry ST, Ward MH, Kholodov M, Baker J, Phan L, Smigielski EM, Sirotkin K (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29(1):308–311. https://doi.org/10.1093/nar/29.1.308
Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, Forsberg M et al (2010) Towards a knowledge-based human protein atlas. Nat Biotechnol 28(12):1248–1250. https://doi.org/10.1038/nbt1210-1248
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A et al (2015) Proteomics. Tissue-based map of the human proteome. Science 347(6220):1260419. https://doi.org/10.1126/science.1260419
Vernau K, Napoli E, Wong S, Ross-Inta C, Cameron J, Bannasch D et al (2015) Thiamine deficiency-mediated brain mitochondrial pathology in Alaskan Huskies with mutation in SLC19A3.1. Brain Pathol 25(4):441–453. https://doi.org/10.1111/bpa.12188
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164. https://doi.org/10.1093/nar/gkq603
Wang H, Xu Y, Chen M, Shang Q, Sun Y, Zhu D et al (2013) Genetic association study of adaptor protein complex 4 with cerebral palsy in a Han Chinese population. Mol Biol Rep 40(11):6459–6467. https://doi.org/10.1007/s11033-013-2761-6
Yang H, Robinson PN, Wang K (2015) Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat Methods 12(9):841–843. https://doi.org/10.1038/nmeth.3484
Acknowledgements
This project was funded by the Jiangsu Province Association of Maternal and Child Health (JSPAMACH) [Grant No. f201560].
Author information
Authors and Affiliations
Contributions
QZ designed the scheme, performed the experiments, analyzed the data and wrote the manuscript. YN, JW, HY, QZ, LZ, WB, BL, LK, LX and NL participated in the experiments and data analysis.
Corresponding author
Ethics declarations
Conflict of interest
Qingwen Zhu, Yufei Ni, Jing Wang, Honggang Yin, Qin Zhang, Lingli Zhang, Wenjun Bian, Bo Liang, Lingyin Kong, Liming Xuan and Naru Lu declare that they have no conflict of interest.
Ethical approval
The ethical committee of Nantong Municipal Maternal and Child Health Hospital approved this study.
Informed consent
All study participants gave written informed-consent.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhu, Q., Ni, Y., Wang, J. et al. Identification of pathways and genes associated with cerebral palsy. Genes Genom 40, 1339–1349 (2018). https://doi.org/10.1007/s13258-018-0729-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-018-0729-6