Abstract
Plants, being sessile in nature have evolved to combat pathogenic invasion by judicious utilization of cellular events and re-orchestration of existing metabolic pathways. “Autophagy” is a self-elimination procedure for maintaining cellular equipoise as well as recycling of different cytosolic components. The lysosome is a cell organelle filled with the lytic enzyme that has the capability to destroy self and non-self-biological macromolecules. Cells perspicaciously utilize these suicidal enzymes for the perpetual cycling of materials in the cellular milieu. Autophagy not only degenerates inoperative macromolecules but it also can protect the cells from the deleterious effect of different misfolded proteins. Autophagy may be selective or non-selective. Different organelles e.g., mitochondria, peroxisome, chloroplast, etc. can be selectively atrophied by this process. In plants, autophagy has prodigious functions in cellular fitness to senescence. Recently, it has been demonstrated that biotic stress can also be outlasted by autophagy in plants. So, it has become a propitious mechanism in plants for biotic stress tolerance physiology. The present review intends to discuss the mechanism of plant autophagy with special reference to biotic stress regulation in plants.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autophagy is a physiological process by which eukaryotic cells maintain their normal vital cycle by degrading toxic non-self-elements as well as unused, tampered self-components too, into recyclable biomolecules or biounits. In some cases, it is associated with vacuolar trafficking to through away degraded products, which are no longer in use, outside the cell. The concept of cellular autophagy was started with the discovery of the lysosome by Christian de Duve in 1955, He coined the term “autophagy” for this specific cellular event later in 1963 [18, 55].The major breakthrough in autophagic science came through the work of Yoshinori Ohsumi, in yeast. They discovered the basic mechanism behind autophagy by the screening of 15 autophagy mutants (apg mutants) in yeast [126]. Prof. Ohsumi’s group was awarded the Nobel Prize in “Physiology and Medicine” for their groundbreaking work in this field in 2016 [93]. Later on, many genes e.g. autophagy (AUT), cytoplasm-to-vacuole targeting (CVTs), peroxisome degradation by autophagy (PAGs), glucose-induced selective autophagy (GSAs), peroxisome degradation deficient (PDD), pexophagy zeocin resistant (PAZ), etc. were discovered in line and led this cellular phenomenon an emerging field of investigation in biological sciences [36, 94, 112, 124, 125, 147].
Although it was first discovered in yeast, it was unveiled later on that it has also tremendous involvement in animal developmental events and as well as diseases such as neuro-degeneration [19, 96]. Autophagy in plants was deciphered first in Arabidopsis and most of the works were concentrated mainly on morphologic events within plant cells due to autophagy through microscopic analyses [145]. In the past several years, many works have been documented in plants describing mechanistic regulation of plant autophagy. Although, the initial genetic analyses in plant autophagy were focused on finding out principal relatedness with yeast and other animal models, later a huge development in plant autophagy research was held to portray it as an individual branch [28, 31, 81, 92]. Detailed genetic analyses revealed at least 30 autophagy-related genes (ATG) in Arabidopsis. The function of ATP-dependent activating enzyme APG7, which acts on APG8/12 gene families along with many other orthologs was also discovered for the first time in the plant [22, 35]. Autophagy generally functions as an important housekeeping regulator to maintain cellular homoeostasis [130]. Along with the maintenance of normal fitness of the plant, it also contributes in immune response, as autophagy is dramatically induced by different stress responses in plants [7]. Autophagy is largely classified into three types depending on their mechanistic behavior (i) micro autophagy, (ii) macro autophagy, (iii)chaperone-mediated autophagy. The details of each of these procedures are discussed later in this review. This article is further focused on recent advancements in the molecular mechanism of autophagy as well as its relation to combating biotic stress in plants. This would provide a comprehensive idea of contemporary development in autophagy in plant biotic stress biology that will have a remarkable impact on shaping future plant stress biology.
Morphology of autophagic plant cells
Any changes within the cell are best documented with their profound changes in morphological behavior. Autophagy by its degradative nature has some characteristic morphological features as well. In plants, these morphological changes were initially screened in different cells to characterize the associated cellular event. Most of the works were intensive characterization through electron microscopic analyses. Electron microscopic images of autophagic cells were characterized by small intra-vacuolar structures that originated through the invagination of cytoplasmic membranes and were very much similar to auto lysosomal bodies [85, 129]. Later lysosomal fusion with autophagic vesicles was confirmed in plant cells too [83, 84]. In Euphorbia cells, it was shown that these small vacuoles were in tubular structures with many digitate protrusions which in turn capture some portions of cytoplasm within and become autophagosome-like structures, called provacuoles [83, 84]. The organelle-specific autophagy was also documented in plants where degraded chloroplasts were found within autophagosome-like structures [101, 134]. Later on, the degradation of many other cellular organelles through autophagy was also confirmed. In Arabidopsis, degraded endoplasmic reticulum (ER) with ribosomes was observed within autophagosome-like structures due to ER stress. Inositol Requiring Enzyme 1b (IRE1b) was found to be the key regulator of ER stress-induced autophagy in plants [74]. In another study in alga Chlamydomonas reinhardtii plastid autophagy was noticed due to the depletion of chloroplast ClpP protease [107]. In Arabidopsis, it was evidenced that Charged Multivesicular Body Protein (CHMP), CHMP1a, and CHMP1b regulate the autophagic turnover of plastids [119]. Selective autophagy of mitochondria (mitophagy) was evidenced in Saccharomyces ceriviae by transmission electron microscopy and the presence of cytochrome b [15]. Another study demonstrated the function of Autophagy Related 11 in senescence-induced mitophagy in Arabidopsis [62].
Molecular mechanism of autophagy
Macroautophagy is a basic cellular clean-up process where cellular contents are readily degraded by lysosomes, sometimes stored in vacuoles and recycled when required. In plants, macroautophagy is elaborately demonstrated and takes part in major physiological responses. Autophagy is primarily instigated in plants by senescence and starvation [12, 102, 118]. Recently it has been reported that different stress has also major impact on inducing autophagy in plants. The mechanism of autophagy is complex and primarily demonstrated in yeast and mammals. Many homologous autophagy-related genes have also been reported from plants that work in similar fashion as reported in yeast and mammals. Among 40 core ATG, more than 30 genes are also identified in Arabidopsis and many others are investigated as plant-specific [21, 145].The process of autophagy can be subdivided into four major steps, (i) phagophore formation or nucleation, (ii) extension and multimerization of phagophore, (iii)capture of random/selective targets for degradation, and iv] degradation of captured targets.
Phagophore formation or nucleation
The initiation of phagophore formation is usually carried out by a cytosolic membranous structure called PAS (pre-auto-phagosomal structure). This is suspected to originate from the endoplasmic reticulum (ER) in yeast but there is no report of PAS formation in mammals [52].Yeast PAS formation depends on ATG9 and also COPII vesicle was found to be localized onto the ER membrane [121]. Recently, PAS/ autophagosome marker ATG14 and autophagosome formation marker ATG5 were identified in the contact site of ER-mitochondria right after starvation in mammalian cells [32]. This enhances the probability of ER-dependent origin of PAS in mammals. Although, very little is known about the initiation of plant autophagosomes, but it has been reported that atg9 mutants showed minimum accumulation of autophagosomes in plants also [157]. Another protein, SH3 DOMAIN CONTAINING PROTEIN 2 was also found to be co-localized with ER marker in plants that also binds to phosphatidyl inositol tri-phosphate (PI3P)[158]. Autophagosome initiation is further by activation of three complexes, Atg1/ ULK1 (Unc-51-Like Autophagy Activating Kinase 1); ATG9 complex, and PI3K (phosphatidyl inositol 3 kinase)/ Vps34 (VACUOLAR PROTEIN SORTING 34) [144].Atg1 kinase is complexed with Atg13 and Atg17 to form a trimeric complex that in turn regulates the further recruitment of trans-membrane protein Atg9 [106]. This complex regulates the formation and growth of the newly formed phagophore. Vps34 is a class III PI-3 kinase that utilizes phosphatidyl inositol (PI) as only substrate to generate inositol tri-phosphate (PI3P), essential for phagophore initiation and recruitment of other ATG proteins to growing phagophore [137].Beclin I activates the function of Vps34 when binds with it and induces the production of more PI3P[29]. In mammals, UV RAG, BIF1, Atg 14L, Ambra, etc. act as positive regulators of phagophore formation whereas, rubicon, and Bcl2 act as negative regulators [67, 80, 86, 100].
Extension and multimerization of phagophore
Extension of phagophore is achieved through two ubiquitination-like systems e.g., the Atg12-Atg5 conjugation system and the Atg8-Atg3 lipidation system [145].
Atg12–Atg5 conjugation system
The phagophore should extend its length by the addition of lipid tails into it. Atg12 is first activated by the E1 ubiquitination activation enzyme Atg7 in an ATP-dependent manner. Then this Atg12 is carried to Atg10, which acts as an E2 ubiquitination carrier enzyme [53]. The conjugation reaction takes place between the glycine residue of Atg12 and the cystine residue of Atg10. Finally, carboxyterminal glycine is transferred to the 130th lysine residue of Atg5, to form the Atg12–Atg5 complex (Fig. 1). Simultaneously, Atg16L dimer is recruited onto the Atg12-Atg5 complex, and multimeric Atg12–Atg5–Atg16L is formed in the tip of growing phagophore [2, 29].
Atg8-Atg3 lipidation system/ processing of LC3B II
The processing of LC3B II is prevalent in mammalian systems and detailed mechanism in plants is still not completely unveiled. In plants, it is sometimes called the Atg8-PE system. The most crucial step in this process is the activation of the Atg8-Atg3 lipidation system which is also called microtubule-associated protein light chain 3 (LC3B) processing in mammals [29]. In this step, nascent Atg8 (LC3B is the mammalian homolog of Atg8) is cleaved by a cystine protease Atg4 to produce truncated Atg8 with an activated carboxy-terminal glycine residue (LC3B I in mammals). This c-terminal Gly residue is conjugated with Cys residue of Atg7 by a thioester linkage, where Atg7 acts as an E1 enzyme. Atg8 is then carried through the E2 enzyme Atg3 and finally transferred to phosphatidyl inositol (PE) to form PE-Atg8 conjugate (LC3B II in mammals) [29, 42, 145]. This PE-Atg8 is further recruited onto the growing tip of the phagophore with the previously described Atg12-Atg5-Atg16L multimer [29]. This complex recruitment of Atg proteins not only extends the phagophore but also induces curvature to the extending phagophore that helps to engulf cytosolic materials [144]. In mammals, GABA RAP (γ amino butyric type A receptor associated protein) is also modulated in concurrence with LC3B II and produces GABA RAP II that co-localizes with LC3B II [114] (Fig. 1).
Capture of random/ selective targets for degradation
As described above during the formation of autophagosome it captures some amount of cytosolic portions and many cellular parts e.g. protein aggregates, different cell organelles, etc. are also captured with this cytoplasm within the autophagosome non-selectively. In plants, despite starvation and senescence-induced autophagy some stress-induced autophagy is also reported and in all these cases autophagy is rather selective in nature. Selective autophagy is the most interesting part of the whole procedure and the selectivity of cargos in the autophagic body is presently under through scientific scanner. Although many works to solve this problem are going on but still, no such conclusive remarks have come into the picture. It is evident from many studies that ATG8 plays a key role in selective autophagy in yeast, mammals as well as in plants. It has been shown that p62/SQSTM1 helps in selective interaction with LC3B II in mammals during the autophagic degradation of protein aggregates (proteophagy) [105]. Similarly, NBR1 also plays a critical role in proteophagy whereas in yeast Uth1p and Atg32 also play vital roles in selective degradation of mitochondria (mitophagy) [29, 51]. It has been reported that plant NBR1 is also necessary for selective autophagy. In Arabidopsis nbr1 knockout mutants were shown to be defective in selective autophagy of protein aggregates as mutant plants accumulated ubiquitinated insoluble proteins under stress [153]. In another study, it was shown that p62/SQSTM1 and NBR1, although targeted by autophagy but they can also help in cargo receptors for selective autophagy of other substrates. Interestingly, Arabidopsis NBR1 (AtNBR1) has hybrid properties with both p62 and mammalian NBR1 in sequence similarities and domain architecture [122]. A chaperone-associated E3 ubiquitin ligase (CHIP) is a protein known to take part in the degradation of non-native proteins through the 26S proteasome complex in plants, showing its interaction with NBR1 in selective autophagy. In Arabidopsis, knockout chip mutants were highly compromised in autophagy which is further deteriorated in chip nbr1 double mutants [155]. Structural studies have revealed that Atg8 interacts with all the cargo receptors through a WXXL-like motif, known as the Atg8-family interacting motif (AIM). AIM is known to interact with many autophagic receptors e.g. Atg19, p62, Atg32, BRCA1 (NBR1), and Nix [98]. In Nicotiana Joka2, an ortholog of p62 reported taking part in viral-induced selective autophagy of plant proteins [159]. Atg11 is an important scaffold protein that facilitates selective autophagy by interacting with other Atgs e.g. Atg19, Atg32, Atg30, or Atg36. These interactions lead to selective mitochondrial degradation (mitophagy) in Saccharomyces cerevisiae and peroxisomal degradation (pexophagy) in Pichia pastoris [27, 38, 70, 104, 142]. Atg11 is also known to interact with membrane fission GTPase, dynamin-related protein 1 (Dnm1 or Drp1), vacuolar sorting protein 1 (Vps1), and mitochondrial fission 1 (Fis1) [78, 79].The detailed structural and functional overview of Atg11 was wonderfully reviewed by Katarzyna Zientara-Rytter and Suresh Subramani [161] (Fig. 2 and Fig. 3).
Degradation of captured components
This is the final step of autophagosome maturation where an autophagosome fuses itself with a lysosome or vacuole to degrade the contents packed inside it, called autolysosome or autophagic body respectively. In animal cells generally, the autophagosome fuses with the lysosome to produce auto lysosome but in plants or yeast, the autophagosome fuses with vacuole to form the autophagic body. In both cases, the degradation and recycling of the materials take place in a similar way [120]. Despite the above fact, in the tobacco BY-2 cell line, it was demonstrated that starvation also induced distinct autolysosome formation [123]. So, the distinct line of the fate of autophagosomes between plants and animals becomes blurred with the advancement of research. It has long been known that lysosomal-associated membrane protein (LAMP) plays a crucial role in autolysosome formation. It was observed that the depletion of LAMP 2 leads to impaired autolysosome formation in rat pancreatic cells [25]. Phosphatidyl inositol 3 phosphate (PI3P) also plays a critical role in autolysosome formation [136]. In Nicotiana benthamiana NbVPS15 and NbVPS34 exhibited their function in autolysosome formation by regulating phosphatidylinositol 3 phosphates (Ptdlns (3)p)[77]. In another study in the tobacco GFP tracking system using GFP-AtVam3p, sporamin-GFP, and gamma-VM23-GFP fusion protein revealed that endocytosis has a significant effect on autophagosome fusion [141]. Lysosome homeostasis is also important in autolysosome formation. Autophagic lysosome formation (ALR) was found to be significantly controlled by the kinesin motor protein KIF5B [23]. WHAMM is a Wiskott-Aldrich syndrome protein (WASP) which acts as actin nucleation promoting factor (NPF) is also important for autolysosome tubule formation. Knockout of whamm leads to reduced actin scaffolding and autophagosome fusion with the lysosome [14]. Unlike animals, the cytoskeletal system especially actin plays an important role in autophagy in plants. Although, the effect of actin polymerization inhibitors cytochalasin D, latrunculin B, or over-expression of Prolifin 3 did not affect autophagy in Nicotiana benthamiana. Similarly, anti-microfilament drug treatment in Arabidopsis also did not affect basal autophagy but the silencing of Actin7 leads to disruption of autophagy in Atg2,3,5,6 and 7 dependent manner [152]. A small GTPase Rab7 was also found essential for autophagosome-lysosome fusion through the accumulation of phosphatidylinositol 4, 5 bisphosphate (PI (4, 5) P2)[1]. Homologous yeast proteins Ykt6, Vam3, Ypt7, and complex HOPS are involved in the fusion of the autophagosome with vacuole in plants [63, 160]. It was also evident that charged multi-vesicular body protein 1 (CHMP1), FYVE domain protein required for endosomal sorting 1 (FREE1), vacuolar protein sorting 2.1 (Vps2.1), cell death-related endosomal FYVE/SYLF protein 1 (CFS1) and plant exocyst complex component EXO70B1 were also reported to be involved in trafficking as well as the fusion of the autophagosome with vacuole [82].
Role of SNARE complex in autophagosome sequestration
N-ethylmaleimide sensitive factor activating protein receptors (SNAREs) is the large protein family with more than 60 members of yeast and mammals [24, 56] those typically involved in universal membrane fusion and endocytic pathway. In plant studies with SNARE proteins are rare but some have been reported recently for their potential role in membrane fusion. SNAREs with two of their primary types, vesicle-associated v-SNARE (synaptobrevin) and target cell-associated t-SNARE (syntaxin and SNAP 25) found to be involved in autophagosome fusion. In plants, syntaxins e.g., Vam3, Vam7, and VTI1, VTI12 are located on the tonoplast membrane which can interact with syntaxin protein (Ykt6) on the outer membrane of autophagosome [20, 54, 58, 120]. This interaction is facilitated by HOPS and RABG3B also found to be involved [120, 127, 150]. Interestingly, PI3P on the membrane of both tonoplast and autophagosome identified by Ypt7 brings the whole complex together for sequestration in an Atg14-dependent manner[3, 120].
Sugars and autophagy in plants
Sugar is considered to be the indicator of cellular homeostasis and a biological marker of stress. Any physiological imbalance e.g. starvation, senescence, or biotic stress has a direct effect on the sugar contents or carbon balance of a cell [30, 146]. As autophagy in plants largely depends on these cellular conditions, for that reason it is obvious to have an indirect relation with sugars. Interestingly, despite this indirect relation scientists are also establishing a dramatic direct relationship of autophagy with sugar balance [46]. Internal energy sensors, e.g., sucrose non-fermenting -1-related protein kinase (SnRK1)/ AMPK positively control autophagy whereas; the target of rapamycin (TOR) kinase has a negative impact on it [111]. Besides, silent information regulator2 (Sir2) also has sugar-dependent autophagy in plants [133]. As discussed earlier, sugar phosphates imply direct interactions with Atgs, and in plants, they activate SnRK1 and deactivate TOR kinase which in turn activates basic autophagy. Glucose-activated G-proteins also have a tremendous impact on this phenomenon [47]. A study in Arabidopsis showed that AtRGS1 negatively correlated with autophagy as ATG4 and ATG12a promote degradation of AtRGS1 during autophagy [140]. The relationship of sugar, starvation, and autophagy was also beautifully experimented in rice suspension culture where a 46 kDa α-amylase was profusely accumulated in amyloplast upon sucrose starvation that subsequently documented with aggressive autophagic degradation of cytosol [9]. It also evidenced that nocturnal sugar deprivation may lead to targeted autophagy of chloroplasts in Arabidopsis which have developmental roles in plants [43]. In Tripogon loliiformis, an increase in sucrose and trehalose measures concurred with autophagosome formation [132].
Reactive oxygen species (ROS) and autophagy in plants
ROS is the collective assemblage of active chemical compounds with a reactive unpaired electron e.g., superoxide (O2.−), hydroxyl (OH.), oxides of nitrogen (NO., NO2.), peroxyl (RO2.), alkoxyl (RO.), etc. originated as a byproduct of different metabolic activity in living organisms. Mitochondria and peroxisomes are the major reservoirs of ROS in biological organisms [5]. In plants, the electron transport system in chloroplast contributes much to redox homeostasis [6]. Besides, membrane-bound respiratory burst oxidases also control the cellular ROS level [5].ROS also regulates cell death, hence, its connection with autophagy is prominent. ROS generation is inevitable in senescence, starvation, or any biotic or abiotic stress. Active oxygen species are also known to oxidize cytosolic proteins and these deformed proteins are degraded by macroautophagy. Treatment with hydrogen peroxide or methyl viologen (ROS inducer) was found to induce autophagy in Arabidopsis. Macro autophagy defective plants (RNAi-AtATG18a) showed enhanced accumulation of deformed proteins as compared to wild-type plants [138]. In a separate study, it was demonstrated that inhibitors of NADPH oxidase blocked autophagy induced by starvation and salinity stress but not by osmotic stress. It clearly entails that autophagy follows two distinct pathways, NADPH oxidase-dependent and NADPH oxidase-independent [76]. In Arabidopsis, root hydrogen sulfide negatively regulates autophagy under nitrogen-deprived conditions but is unable to mitigate ROS. This study suggested the role of hydrogen sulfide in autophagy is independent of ROS [60]. Mitochondria have an important role in oxidative stress-induced autophagy in plants [91].Glyceraldehyde 3 phosphate dehydrogenase (GAPDH) is a key enzyme in the energy metabolic pathway in plants. In Arabidopsis,GAPDH knockout lines [KO lines] exhibited enhanced cytosolic basal ROS accumulation and autophagy upon Pseudomonas syringae infection [39]. ROS-mediated autophagy was also found to control several physiological and developmental events in plants e.g., stomatal opening and transpiration in plants [88]. Deficiency of essential micronutrients e.g. zinc (Zn+2) may cause physiological imbalance as well as accumulation of cellular ROS. In Arabidopsis thaliana it was observed that autophagy increases the bioavailability of Zn+2which helps to mitigate ROS-mediated cell damage under light stress [115].
The execution of autophagy in biotic stress
Plants are constantly attacked by various pathogens, where these pathogens are facing a co-evolutionary feud with their host plants. Autophagy has an important role in controlling the plant’s innate immunity. It can function either as a guardian or destructor in response to the upcoming pathogen (i.e. biotrophic or necrotrophic) [128]. Autophagy is crucial in the regulation of hypersensitive response (HR) cell death, a type of programmed cell death (PCD) during innate immunity. The production of ROS and nitric oxide (NO) are generally observed at the initial stage of defense response and are necessary for the commencement of HR [16]. Additionally, lipid peroxidation, ion fluxes, cell wall strengthening, and reprogramming of defense gene networks are also recounted [87]. Plant autophagy has been accredited ‘pro-death’ and ‘pro-survival’ function in managing PCD linked to effector‐triggered immunity (ETI). Autophagy arbitrates programmed reprocessing of damaged or degraded organelles like chloroplasts and mitochondria [90]. The unrestrained signalling of death from these flawed organelles, via the release of ROS and cytochrome c, can induce accelerated cell death. The complete photo-damaged chloroplasts are moved to the central vacuole for deterioration, whereas immobile forms were found to concentrate in autophagy-deficient mutants [44]. Recognition of pathogen effectors (also called avirulence (AVR proteins)) by resistance (R) genes encoding R proteins triggers HR PCD [48] which depends upon the type of resistance receptors. Coiled‐coil‐nucleotide binding site (NBS)‐leucine‐rich repeat (LRR)(CC-NB LRR) receptors provoke partial cell death dependent (resistance gene, RPM1 mediated) or independent (resistance gene, RPS2 mediated) autophagy, whereas, Toll/Interleukin‐1 receptor‐NB LRR(TIR-NB LRR) receptors (RPS4 and RPP1) are stringently autophagy-dependent[41].
Plants can take advantage of selective autophagy to escape the pathogenic attack, although some of them surmount the plant’s immune response by encountering host-specific autophagy pathways [13, 31, 37]. Autophagy can restrain HR, lowering the harm caused to healthy tissue [40]. This pro-survival role of autophagy can be associated with its homeostatic function in eradicating toxic by-products released as systemic responses upon pathogen challenge [11, 40, 95, 143]. The pro-survival effect of autophagy has also been reported against necrotrophs, where cell death is induced purposely to derive nutrients from the host [61, 66].
Autophagy is accomplished to perform specialized cellular functions under varied circumstances. The glycolysis enzyme, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), was reported to associate with ATG3 and negatively regulate autophagy [33]. Contrary to this, Bax inhibitor-1 (BI-1), cell death, and endoplasmic reticulum stress manager were observed to interact with ATG6 and positively regulate autophagy [139]. Interestingly, the deletion of GAPDH or deletion of BI-1 both induced TMV-provoked HR on plants carrying the TIR-NLR-receptors[33, 139]. Also, GAPDH silencing produced no HR PCD phenotype upon Pst DC3000 infection. These contradictory results in induction HR might be due to non-autophagy-linked functions of the genes.
The PCD phenotype of the autophagy mutants was reported in the leaves of older plants in comparison to the leaves of younger plants [143]. The increased HR in older leaves was linked to an increase in the phytohormone salicylic acid (SA) level. The study suggested negative regulation of cell death by autophagy in an NPR1-dependent SA signaling during innate immunity [143]. Apart from greater SA concentration, these autophagy-defective plants were also found to promote the growth of incompatible microbes [41, 75]. It was suggested that an increase in bacterial count may release more effectors that may surpass the HR PCD causing a slight delay in cell death at the initial stage of pathogen challenge in atg mutants [41].
Autophagy to combat plant pathogens
Earlier studies provide evidence for autophagy as an endurance strategy in the resistance gene N-mediated HR PCD in Nicotiana benthamiana plants. The N gene provides resistance to the tobacco mosaic virus (TMV) and it may be stimulated by the p50 helicase protein of TMV replicase to generate HR PCD. Autophagy in the resistant cultivar of tobacco plants comes into action upon initial TMV attack, which restricts TMV-induced cell death to only challenge sites in wild-type plants, whereas, the RNAi of Beclin1/ ATG6 or ATG7(autophagy-deficient) dispose of TMV-induced autophagy and causes an ETI-associated PCD by the progression of cell death to uninfected areas [75]. Similar disease expression is reported in Arabidopsis plants where ATG6 silencing hinders autophagy and leads to unrestrained plant cell death when HR PCD is provoked by an avirulent bacterial pathogen (Pseudomonas syringae pv. tomato strain DC3000 (Pst)/AvrRPM1, Pto AvrRPM1) [99]. It was been reported that A. thaliana beclin1/atg6 knock-down lines infected with virulent PtoDC3000 increased disease-associated necrosis, as compared to wild-type plants at late infection stages [99].These studies indicate the ‘anti-death’ action of autophagy in constraining disease‐associated necrosis. The Arabidopsis knockout mutants atg7 and atg9 show slowed PCD by the avirulent strains of bacteria Pto AvrRPS4 or oomycetes Hyaloperonospora arabidopsidis isolate Noco2 [41].
ATG1 protein is produced in Botrytis cinerea during plant colonization and atg1 mutant plants were flawed in appressorium formation [110], indicating the formation of appressorium by the fungal pathogen is dependent on autophagy. Knockout mutants in Magnaporthe oryzae for a small Rab GTPase present in lysosomal membranes were observed to be defective in autophagy and appressorium development [72], speculating an association in autophagy and vesicle transport in phytopathogenic fungi. Recently, it was reported that infection with Phytophthora sojae induced 26 autophagy-related genes significantly, and silencing of the PsATG6a gene in the oomycete curtails its capability to colonize the host plant indicating that autophagy might have an important role in the development and infection mechanism of the pathogen [8]. Similar provocation of autophagy was observed in the haustoria of leaf rust pathogens which was crucial for infection in the host [71].
A large number of growing reports assign a positive role of autophagy activation in resistance towards necrotrophic phytopathogens [50, 59, 61]. In Arabidopsis loss-of-function mutants atg5, atg10, and atg18a, necrosis spread was observed upon infection with the necrotroph, Alternaria brassicicola, along with the generation of ROS and by increased fungal growth [61]. The group suggested autophagy as a ‘pro-survival’ mechanism to restrict the plant’s tissue destruction upon pathogen infection. Contradictory to this, autophagy mutants mostly present enhanced resistance to biotrophic pathogens, due to the closure of the autophagy system causing an increase in SA accumulation and hindered cellular processes under pathogenic stress [34]. Arabidopsis atg mutant plants infected with virulent biotroph, Pseudomonas syringae pv. tomato, exhibit resistance instead of spreading necrosis. SA levels in bacteria unchallenged and challenged atg plants were modestly greater in comparison to wild-type plants, along with the up-regulation of SA-dependent gene expression and camalexin synthesis [61].
Even though autophagy plays a role in plants' antiviral defense system, the insights into molecular mechanisms are not completely known [116]. Recently some interesting reports point out the role of autophagy in providing antiviral immunity in plants. It can induce HR PCD upon an incompatible viral challenge or intervene in the breakdown or removal of viral particles and/or individual proteins from compatible viruses [57, 97]. The RNA-dependent RNA polymerase (RdRp) of the Turnip mosaic virus (TuMV) was reported as a target for selective autophagic degradation [64]. Autophagy can be induced in response to various DNA and RNA viruses reducing virus concentration, indicating the role of autophagy in basal antiviral defences [31, 37, 64]. In a previous report, the beta satellite (DNA-β) derived protein βC1 from Cotton leaf curl Multan virus (CLCuMuV) was found to be deteriorated by host autophagy by employing autophagosomes and association with ATG8 proteins[37]. Therefore, autophagy can be looked up as taking up direct functions in plant immunity by, aiming degradation of viral proteins and particles. The employment of autophagy in plants is well documented in case of abiotic stress tolerance but significant involvement of the same in biotic interaction is also emerging (Table 1).
Reconstruction of host autophagy by pathogens to escape the immune response
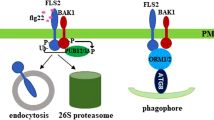
Many phytopathogens have emerged with strategies to modulate host autophagy machinery as an asset to infection, while some eukaryotic microbes utilize their own autophagy system for favorable pathogenesis. TOR kinase is an important control point for autophagy, and TOR knockout and knockdown mutant phenotype suggest TOR plays an essential role in stress confutation in plants [73, 89]. The TOR inhibition is reported by bacterial wilt pathogen Ralstonia solanacearum AWR5 effector thereby inducing autophagy in yeast [103]. PexRD54, an effector of the hemibiotrophic pathogen Phytophthora infestans, binds host plant autophagy protein ATG8CL depleting the autophagy cargo receptor Joka2 out of it. It thus interrupts with NBR1/Joka2 positive regulation of host defenses [13]. The immune response assisted by autophagy is highly complex and varies depending on the host-microbe system under study. Cosmopolitan, necrotrophic fungi, Sclerotinia sclerotiorum, secrete oxalic acid to overcome autophagy-linked host defense, causing unregulated host cell death [45, 49]. Recently, it was seen Pst effector HopM1 induces autophagy and increases the autophagic flux of proteins in A. thaliana to take advantage of infection. The NBR1-mediated selective autophagy pathway induced upon Pst infection restricts disease spread. It was hypothesized that different selective autophagy pathways with pro- and anti-bacterial play a role in defining the outcome of bacterial infection [128].
Some of the viruses are also reported to reprogram host immunity as seen in the case of plant–Polerovirus interactions. An RNA silencing suppressor from Polerovirus, P0, has been linked to causing autophagic degradation of ARGONAUTE 1 (AGO1), an important part of RNA-induced silencing complex (RISC)[4, 17].Similarly, calmodulin-like protein NbCaM in plants induced by geminivirus protein βC1 causes susceptibility by autophagic removal of constituents of host plant RNA silencing machinery [65]. NbCAM in tobacco induces the autophagic degradation of the host Gene Silencing 3 (NbSGS3) protein, which helps in dsRNA synthesis [26].CaMV encodes a suppressor P6 protein that associates TOR kinase [113] and induces TOR activation thereby suppressing oxidative burst and SA-dependent autophagy [162]. Therefore, with respect to pathogenic viruses, autophagy supports the degradation of host RNA silencing machinery, as an infection strategy.
Future perspectives
Plant autophagy is essential for sustaining cellular homeostasis under natural circumstances and is upregulated during all kinds of stress to perpetuate cell life. Though a steady growth is witnessed in the number of studies on autophagy, most of our knowledge is based on studies done in atg knockout mutants. In the past few years, elucidation of the role of autophagy in biotic stress tolerance has provided information regarding the benefit of autophagy on plants. It is also very important to gain a deeper understanding of the activation of the autophagy mechanism in response to upcoming pathogenic stress, and how it functions with other regulatory pathways to induce immunity. The reciprocity between sugar levels and autophagy in plants is composite and depends largely on the species and tissue under observation. Insights into autophagy require targeting specific host cargo receptors and/or pathogenic effectors to elaborate molecular mechanisms involved in regulating autophagy in plant-pathogen interactions. It is better to say that studies should be focussed not only on molecular mechanisms used by host plants for gaining innate immunity but also on pathogens deployed means for using autophagy for cheating plants. Priority should be given to extending the research in autophagy using different plant-pathogen systems to elucidate variation in defense response and pathways modulated.
Data availability
Not applicable.
Code availability
Not applicable.
References
Baba T, Toth DJ, Sengupta N, Kim YJ, Balla T. Phosphatidylinositol 4,5‐bisphosphate controls Rab7 and <scp>PLEKHM</scp> 1 membrane cycling during autophagosome–lysosome fusion. EMBO J. 2019. https://doi.org/10.15252/embj.2019102837.
Barth S, Glick D, Macleod KF. Autophagy: assays and artifacts. J Pathol. 2010;221:117–24. https://doi.org/10.1002/path.2694.
Bas L, Papinski D, Licheva M, Torggler R, Rohringer S, Schuschnig M, et al. Reconstitution reveals Ykt6 as the autophagosomal SNARE in autophagosome–vacuole fusion. J Cell Biol. 2018;217:3656–69. https://doi.org/10.1083/jcb.201804028.
Baumberger N, Tsai CH, Lie M, Havecker E, Baulcombe DC. The polerovirus silencing suppressor P0 Targets ARGONAUTE proteins for degradation. Curr Biol. 2007;17:1609–14. https://doi.org/10.1016/j.cub.2007.08.039.
Bhar A, Gupta S, Chatterjee M, Das S. Redox regulatory networks in response to biotic stress in plants: a new insight through chickpea‐fusarium interplay. In: Pandey GK, editor. Mechanism of Plant Hormone Signaling under Stress. Wiley; 2017. p. 23–43. https://doi.org/10.1002/9781118889022.ch20.
Bhar A, Gupta S, Chatterjee M, Sen S, Das S. Differential expressions of photosynthetic genes provide clues to the resistance mechanism during Fusarium oxysporum f.sp. ciceri race 1 (Foc1) infection in chickpea (Cicer arietinum L.). Eur J Plant Pathol. 2016;148:533–49. https://doi.org/10.1007/s10658-016-1109-1.
Bozhkov PV. Plant autophagy: mechanisms and functions. J Exp Bot. 2018;69:1281–5. https://doi.org/10.1093/jxb/ery070.
Chen L, Zhang X, Wang W, Geng X, Shi Y, Na R, et al. Network and role analysis of autophagy in Phytophthora sojae. Sci Rep. 2017. https://doi.org/10.1038/s41598-017-01988-7.
Chen M, Liu L, Chen Y, Wu H, Yu S. Expression of α-amylases, carbohydrate metabolism, and autophagy in cultured rice cells is coordinately regulated by sugar nutrient. Plant J. 1994;6:625–36. https://doi.org/10.1046/j.1365-313X.1994.6050625.x.
Chen Q, Wu Y, Yu F, Xie Q. Coordinative regulation of ERAD and selective autophagy in plants. Theodoulou F, Orosa B, Trujilo M, Rubio V, editors. Essays Biochem. 2022;66:179–88. https://doi.org/10.1042/EBC20210099.
Coll NS, Smidler A, Puigvert M, Popa C, Valls M, Dangl JL. The plant metacaspase AtMC1 in pathogen-triggered programmed cell death and aging: functional linkage with autophagy. Cell Death Amp Differ. 2014;21:1399–408. https://doi.org/10.1038/cdd.2014.50.
Contento AL, Kim SJ, Bassham DC. Transcriptome profiling of the response of arabidopsis suspension culture cells to suc starvation. Plant Physiol. 2004;135:2330–47. https://doi.org/10.1104/pp.104.044362.
Dagdas YF, Belhaj K, Maqbool A, Chaparro-Garcia A, Pandey P, Petre B, et al. An effector of the Irish potato famine pathogen antagonizes a host autophagy cargo receptor. Elife. 2016. https://doi.org/10.7554/eLife.10856.
Dai A, Yu L, Wang HW. WHAMM initiates autolysosome tubulation by promoting actin polymerization on autolysosomes. Nat Commun. 2019. https://doi.org/10.1038/s41467-019-11694-9.
Deffieu M, Bhatia-Kiššová I, Salin B, Klionsky DJ, Pinson B, Manon S, et al. Increased levels of reduced cytochrome b and mitophagy components are required to trigger nonspecific autophagy following induced mitochondrial dysfunction. J Cell Sci. 2013;126:415–26. https://doi.org/10.1242/jcs.103713.
Delledonne M, Zeier J, Marocco A, Lamb C. Signal interactions between nitric oxide and reactive oxygen intermediates in the plant hypersensitive disease resistance response. Proc Natl Acad Sci. 2001;98:13454–9. https://doi.org/10.1073/pnas.231178298.
Derrien B, Baumberger N, Schepetilnikov M, Viotti C, De Cillia J, Ziegler-Graff V, et al. Degradation of the antiviral component ARGONAUTE1 by the autophagy pathway. Proc Natl Acad Sci. 2012;109:15942–6. https://doi.org/10.1073/pnas.1209487109.
Deter RL, de Duve C. Influence of glucagon, an inducer of cellular autophagy, on some physical properties of rat liver lysosomes. J Cell Biol. 1967;33:437–49. https://doi.org/10.1083/jcb.33.2.437.
Di Bartolomeo S, Nazio F, Cecconi F. The role of autophagy during development in higher eukaryotes. Traffic. 2010;11:1280–9. https://doi.org/10.1111/j.1600-0854.2010.01103.x.
Dilcher M, Köhler B, von Mollard GF. Genetic interactions with the yeast Q-SNARE vti1reveal novel functions for the R-SNARE YKT6. J Biol Chem. 2001;276:34537–44. https://doi.org/10.1074/jbc.M101551200.
Ding X, Zhang X, Otegui MS. Plant autophagy: new flavors on the menu. Curr Opin Plant Biol. 2018;46:113–21. https://doi.org/10.1016/j.pbi.2018.09.004.
Doelling JH, Walker JM, Friedman EM, Thompson AR, Vierstra RD. The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence in arabidopsis thaliana. J Biol Chem. 2002;277:33105–14. https://doi.org/10.1074/jbc.M204630200.
Du W, Su QP, Chen Y, Zhu Y, Jiang D, Rong Y, et al. Kinesin 1 drives autolysosome tubulation. Dev Cell. 2016;37:326–36. https://doi.org/10.1016/j.devcel.2016.04.014.
Fasshauer D, Sutton RB, Brunger AT, Jahn R. Conserved structural features of the synaptic fusion complex: SNARE proteins reclassified as Q- and R-SNAREs. Proc Natl Acad Sci. 1998;95:15781–6. https://doi.org/10.1073/pnas.95.26.15781.
Fortunato F, Bürgers H, Bergmann F, Rieger P, Büchler MW, Kroemer G, et al. Impaired autolysosome formation correlates with lamp-2 depletion: role of apoptosis, autophagy, and necrosis in pancreatitis. Gastroenterology. 2009;137:350-360.e5. https://doi.org/10.1053/j.gastro.2009.04.003.
Fukunaga R, Doudna JA. dsRNA with 5’ overhangs contributes to endogenous and antiviral RNA silencing pathways in plants. EMBO J. 2009;28:545–55. https://doi.org/10.1038/emboj.2009.2.
Gammoh N, Florey O, Overholtzer M, Jiang X. Interaction between FIP200 and ATG16L1 distinguishes ULK1 complex–dependent and –independent autophagy. Nat Struct Amp Mol Biol. 2012;20:144–9. https://doi.org/10.1038/nsmb.2475.
Gao C, Zhuang X, Cui Y, Fu X, He Y, Zhao Q, et al. Dual roles of an Arabidopsis ESCRT component FREE1 in regulating vacuolar protein transport and autophagic degradation. Proc Natl Acad Sci. 2015;112:1886–91. https://doi.org/10.1073/pnas.1421271112.
Glick D, Barth S, Macleod KF. Autophagy: cellular and molecular mechanisms. J Pathol. 2010;221:3–12. https://doi.org/10.1002/path.2697.
Gupta S, Bhar A, Chatterjee M, Das S. Fusarium oxysporum f. sp. ciceri Race 1 induced redox state alterations are coupled to downstream defense signaling in root tissues of chickpea (Cicer arietinum L.). van Damme EJM, editor. PLoS ONE. 2013;8:e73163. https://doi.org/10.1371/journal.pone.0073163.
Hafrén A, Macia JL, Love AJ, Milner JJ, Drucker M, Hofius D. Selective autophagy limits cauliflower mosaic virus infection by NBR1-mediated targeting of viral capsid protein and particles. Proc Natl Acad Sci. 2017. https://doi.org/10.1073/pnas.1610687114.
Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N, et al. Autophagosomes form at ER–mitochondria contact sites. Nature. 2013;495:389–93. https://doi.org/10.1038/nature11910.
Han S, Wang Y, Zheng X, Jia Q, Zhao J, Bai F, et al. Cytoplastic glyceraldehyde-3-phosphate dehydrogenases interact with ATG3 to negatively regulate autophagy and immunity in Nicotiana benthamiana. Plant Cell. 2015;27:1316–31. https://doi.org/10.1105/tpc.114.134692.
Han S, Yu B, Wang Y, Liu Y. Role of plant autophagy in stress response. Protein Amp Cell. 2011;2:784–91. https://doi.org/10.1007/s13238-011-1104-4.
Hanaoka H, Noda T, Shirano Y, Kato T, Hayashi H, Shibata D, et al. Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an arabidopsis autophagy gene. Plant Physiol. 2002;129:1181–93. https://doi.org/10.1104/pp.011024.
Harding TM, Morano KA, Scott SV, Klionsky DJ. Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J Cell Biol. 1995;131:591–602. https://doi.org/10.1083/jcb.131.3.591.
Haxim Y, Ismayil A, Jia Q, Wang Y, Zheng X, Chen T, et al. Autophagy functions as an antiviral mechanism against geminiviruses in plants. Elife. 2017. https://doi.org/10.7554/elife.23897.
He C, Song H, Yorimitsu T, Monastyrska I, Yen WL, Legakis JE, et al. Recruitment of Atg9 to the preautophagosomal structure by Atg11 is essential for selective autophagy in budding yeast. J Cell Biol. 2006;175:925–35. https://doi.org/10.1083/jcb.200606084.
Henry E, Fung N, Liu J, Drakakaki G, Coaker G. Beyond Glycolysis: GAPDHs are multi-functional enzymes involved in regulation of ROS autophagy, and plant immune responses McDowell JM, editor. PLOS Genet. 2015;11:e1005199. https://doi.org/10.1371/journal.pgen.1005199.
Hofius D, Munch D, Bressendorff S, Mundy J, Petersen M. Role of autophagy in disease resistance and hypersensitive response-associated cell death. Cell Death Amp Differ. 2011;18:1257–62. https://doi.org/10.1038/cdd.2011.43.
Hofius D, Schultz-Larsen T, Joensen J, Tsitsigiannis DI, Petersen NHT, Mattsson O, et al. Autophagic components contribute to hypersensitive cell death in arabidopsis. Cell. 2009;137:773–83. https://doi.org/10.1016/j.cell.2009.02.036.
Ichimura Y, Kirisako T, Takao T, Satomi Y, Shimonishi Y, Ishihara N, et al. A ubiquitin-like system mediates protein lipidation. Nature. 2000;408:488–92. https://doi.org/10.1038/35044114.
Izumi M, Hidema J, Makino A, Ishida H. Autophagy contributes to nighttime energy availability for growth in arabidopsis. Plant Physiol. 2013;161:1682–93. https://doi.org/10.1104/pp.113.215632.
Izumi M, Ishida H, Nakamura S, Hidema J. Entire photodamaged chloroplasts are transported to the central vacuole by autophagy. Plant Cell. 2017;29:377–94. https://doi.org/10.1105/tpc.16.00637.
Jain A, Singh A, Singh S, Sarma BK, Singh HB. Biocontrol agents-mediated suppression of oxalic acid induced cell death during Sclerotinia sclerotiorum–pea interaction. J Basic Microbiol. 2014;55:601–6. https://doi.org/10.1002/jobm.201400156.
Jiao Y, Lei W, Xu W, Chen WL. Glucose signaling, AtRGS1 and plant autophagy. Plant Signal Amp Behav. 2019;14:1607465. https://doi.org/10.1080/15592324.2019.1607465.
Jones JDG, Dangl JL. The plant immune system. Nature. 2006;444:323–9. https://doi.org/10.1038/nature05286.
Kabbage M, Williams B, Dickman MB. Cell death control: the interplay of apoptosis and autophagy in the pathogenicity of Sclerotinia sclerotiorum Tyler B, editor. PLoS Pathog. 2013;9:e1003287. https://doi.org/10.1371/journal.ppat.1003287.
Katsiarimpa A, Kalinowska K, Anzenberger F, Weis C, Ostertag M, Tsutsumi C, et al. The deubiquitinating enzyme AMSH1 and the ESCRT-III subunit VPS2.1 are required for autophagic degradation inarabidopsis. Plant Cell. 2013;25:2236–52. https://doi.org/10.1105/tpc.113.113399.
Kim I, Rodriguez-Enriquez S, Lemasters JJ. Selective degradation of mitochondria by mitophagy. Arch Biochem Biophys. 2007;462:245–53. https://doi.org/10.1016/j.abb.2007.03.034.
Kirkin V. History of the selective autophagy research: how did it begin and where does it stand today? J Mol Biol. 2020;432:3–27. https://doi.org/10.1016/j.jmb.2019.05.010.
Kirkin V, McEwan DG, Novak I, Dikic I. A role for ubiquitin in selective autophagy. Mol Cell. 2009;34:259–69. https://doi.org/10.1016/j.molcel.2009.04.026.
Klionsky DJ. The molecular machinery of autophagy: unanswered questions. J Cell Sci. 2005;118:7–18.
Klionsky DJ. Autophagy revisited: a conversation with Christian de Duve. Autophagy. 2008;4:740–3. https://doi.org/10.4161/auto.6398.
Kloepper TH, Kienle CN, Fasshauer D. An elaborate classification of snare proteins sheds light on the conservation of the eukaryotic endomembrane system. Munro S, editor. Mol Biol Cell. 2007;18:3463–71. https://doi.org/10.1091/mbc.e07-03-0193.
Kushwaha NK, Hafrén A, Hofius D. Autophagy–virus interplay in plants: from antiviral recognition to proviral manipulation. Mol Plant Pathol. 2019;20:1211–6. https://doi.org/10.1111/mpp.12852.
Kweon Y, Rothe A, Conibear E, Stevens TH. Ykt6p is a multifunctional yeast R-SNARE that is required for multiple membrane transport pathways to the vacuole. Mol Biol Cell. 2003;14:1868–81. https://doi.org/10.1091/mbc.e02-10-0687.
Lai Z, Wang F, Zheng Z, Fan B, Chen Z. A critical role of autophagy in plant resistance to necrotrophic fungal pathogens. Plant J. 2011;66:953–68. https://doi.org/10.1111/j.1365-313X.2011.04553.x.
Laureano-Marín AM, Moreno I, Romero LC, Gotor C. Negative regulation of autophagy by sulfide in Arabidopsis thaliana is independent of reactive oxygen species. Plant Physiol. 2016. https://doi.org/10.1104/pp.16.00110.
Lenz HD, Haller E, Melzer E, Kober K, Wurster K, Stahl M, et al. Autophagy differentially controls plant basal immunity to biotrophic and necrotrophic pathogens. Plant J. 2011;66:818–30. https://doi.org/10.1111/j.1365-313X.2011.04546.x.
Li F, Chung T, Vierstra RD. Autophagy-related11 Plays a critical role in general autophagy- and senescence-induced mitophagy in arabidopsis. Plant Cell. 2014;26:788–807. https://doi.org/10.1105/tpc.113.120014.
Li F, Vierstra RD. Autophagy: a multifaceted intracellular system for bulk and selective recycling. Trends Plant Sci. 2012;17:526–37. https://doi.org/10.1016/j.tplants.2012.05.006.
Li F, Zhang C, Li Y, Wu G, Hou X, Zhou X, et al. Beclin1 restricts RNA virus infection in plants through suppression and degradation of the viral polymerase. Nat Commun. 2018. https://doi.org/10.1038/s41467-018-03658-2.
Li S, Zhang T, Zhu Y, Zhou G. Co-infection of two reoviruses increases both viruses accumulation in rice by up-regulating of viroplasm components and movement proteins bilaterally and RNA silencing suppressor unilaterally. Virol J. 2017;14:150. https://doi.org/10.1186/s12985-017-0819-0.
Li Y, Kabbage M, Liu W, Dickman MB. Aspartyl protease-mediated cleavage of BAG6 is necessary for autophagy and fungal resistance in plants. Plant Cell. 2016;28:233–47. https://doi.org/10.1105/tpc.15.00626.
Liang C, Feng P, Ku B, Dotan I, Canaani D, Oh BH, et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol. 2006;8:688–98. https://doi.org/10.1038/ncb1426.
Liao C, Wang P, Yin Y, Bassham DC. Interactions between autophagy and phytohormone signaling pathways in plants. FEBS Lett. 2022;596:2198–214. https://doi.org/10.1002/1873-3468.14355.
Liao CY, Bassham DC. Combating stress: the interplay between hormone signaling and autophagy in plants. Mittler R, editor. J Exp Bot. 2019;71:1723–33. https://doi.org/10.1093/jxb/erz515.
Lipatova Z, Belogortseva N, Zhang XQ, Kim J, Taussig D, Segev N. Regulation of selective autophagy onset by a Ypt/Rab GTPase module. Proc Natl Acad Sci. 2012;109:6981–6. https://doi.org/10.1073/pnas.1121299109.
Liu G, Tian D, Shi C, Wang DM. Autophagy is induced in haustorial mother cells of Puccinia triticina and is necessary for plant infection. Eur J Plant Pathol. 2016;147:833–43. https://doi.org/10.1007/s10658-016-1047-y.
Liu X, Chen S, Gao H, Ning G, Shi H, Wang Y, et al. The small <scp>GTP</scp>ase <scp>MoYpt</scp>7 is required for membrane fusion in autophagy and pathogenicity of <scp>M</scp>agnaporthe oryzae. Environ Microbiol. 2015;17:4495–510. https://doi.org/10.1111/1462-2920.12903.
Liu Y, Bassham DC. TOR Is a negative regulator of autophagy in Arabidopsis thaliana. Schumacher K, editor. PLoS ONE. 2010;5:e11883. https://doi.org/10.1371/journal.pone.0011883.
Liu Y, Burgos JS, Deng Y, Srivastava R, Howell SH, Bassham DC. Degradation of the endoplasmic reticulum by autophagy during endoplasmic reticulum stress in arabidopsis. Plant Cell. 2012;24:4635–51. https://doi.org/10.1105/tpc.112.101535.
Liu Y, Schiff M, Czymmek K, Tallóczy Z, Levine B, Dinesh-Kumar SP. Autophagy regulates programmed cell death during the plant innate immune response. Cell. 2005;121:567–77. https://doi.org/10.1016/j.cell.2005.03.007.
Liu Y, Xiong Y, Bassham DC. Autophagy is required for tolerance of drought and salt stress in plants. Autophagy. 2009;5:954–63. https://doi.org/10.4161/auto.5.7.9290.
Lu S, Yu J, Ma L, Dou D. Two phosphatidylinositol 3-kinase components are involved in interactions between Nicotiana benthamiana and Phytophthora by regulating pathogen effectors and host cell death. Funct Plant Biol. 2020;47:293. https://doi.org/10.1071/FP19155.
Mao K, Liu X, Feng Y, Klionsky DJ. The progression of peroxisomal degradation through autophagy requires peroxisomal division. Autophagy. 2014;10:652–61. https://doi.org/10.4161/auto.27852.
Mao K, Wang K, Liu X, Klionsky DJ. The scaffold protein atg11 recruits fission machinery to drive selective mitochondria degradation by autophagy. Dev Cell. 2013;26:9–18. https://doi.org/10.1016/j.devcel.2013.05.024.
Maria Fimia G, Stoykova A, Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R, et al. Ambra1 regulates autophagy and development of the nervous system. Nature. 2007;447:1121–5. https://doi.org/10.1038/nature05925.
Marshall RS, Li F, Gemperline DC, Book AJ, Vierstra RD. Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/Ubiquitin receptor RPN10 in arabidopsis. Mol Cell. 2015;58:1053–66. https://doi.org/10.1016/j.molcel.2015.04.023.
Marshall RS, Vierstra RD. Autophagy: the master of bulk and selective recycling. Annu Rev Plant Biol. 2018;69:173–208. https://doi.org/10.1146/annurev-arplant-042817-040606.
Marty F. Cytochemical studies on GERL, provacuoles, and vacuoles in root meristematic cells of Euphorbia. Proc Natl Acad Sci. 1978;75:852–6. https://doi.org/10.1073/pnas.75.2.852.
Marty F. The biogenesis of vacuoles insights from microscopy. In The Plant Vacuole. 1997. https://doi.org/10.1016/s0065-2296(08)60146-9.
Matile Ph, Moor H. Vacuolation: origin and development of the lysosomal apparatus in root-tip cells. Planta. 1968;80:159–75. https://doi.org/10.1007/BF00385592.
Matsunaga K, Saitoh T, Tabata K, Omori H, Satoh T, Kurotori N, et al. Two Beclin 1-binding proteins, Atg14L and Rubicon, reciprocally regulate autophagy at different stages. Nat Cell Biol. 2009;11:385–96. https://doi.org/10.1038/ncb1846.
McDowell JM, Dangl JL. Signal transduction in the plant immune response. Trends Biochem Sci. 2000;25:79–82. https://doi.org/10.1016/S0968-0004(99)01532-7.
Medeiros DB, Barros JAS, Fernie AR, Araújo WL. Eating away at ROS to regulate stomatal opening. Trends Plant Sci. 2020;25:220–3. https://doi.org/10.1016/j.tplants.2019.12.023.
Menand B, Desnos T, Nussaume L, Berger F, Bouchez D, Meyer C, et al. Expression and disruption of the Arabidopsis TOR (target of rapamycin) gene. Proc Natl Acad Sci. 2002;99:6422–7. https://doi.org/10.1073/pnas.092141899.
Michaeli S, Galili G. Degradation of organelles or specific organelle components via selective autophagy in plant cells. Int J Mol Sci. 2014;15:7624–38. https://doi.org/10.3390/ijms15057624.
Minibayeva F, Dmitrieva S, Ponomareva A, Ryabovol V. Oxidative stress-induced autophagy in plants: the role of mitochondria. Plant Physiol Biochem. 2012;59:11–9. https://doi.org/10.1016/j.plaphy.2012.02.013.
Minina EA, Filonova LH, Fukada K, Savenkov EI, Gogvadze V, Clapham D, et al. Autophagy and metacaspase determine the mode of cell death in plants. J Cell Biol. 2013;203:917–27. https://doi.org/10.1083/jcb.201307082.
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, et al. A protein conjugation system essential for autophagy. Nature. 1998;395:395–8. https://doi.org/10.1038/26506.
Mukaiyama H, Oku M, Baba M, Samizo T, Hammond AT, Glick BS, et al. Paz2 and 13 other PAZ gene products regulate vacuolar engulfment of peroxisomes during micropexophagy. Genes Cells. 2002;7:75–90. https://doi.org/10.1046/j.1356-9597.2001.00499.x.
Munch D, Rodriguez E, Bressendorff S, Park OK, Hofius D, Petersen M. Autophagy deficiency leads to accumulation of ubiquitinated proteins, ER stress, and cell death inArabidopsis. Autophagy. 2014;10(9):1579–87. https://doi.org/10.4161/auto.29406.
Nah J, Yuan J, Jung YK. Autophagy in neurodegenerative diseases from mechanism to therapeutic approach. Mol Cells. 2015;38:381–9. https://doi.org/10.14348/molcells.2015.0034.
Nakahara KS, Masuta C, Yamada S, Shimura H, Kashihara Y, Wada TS, et al. Tobacco calmodulin-like protein provides secondary defense by binding to and directing degradation of virus RNA silencing suppressors. Proc Natl Acad Sci. 2012;109:10113–8. https://doi.org/10.1073/pnas.1201628109.
Noda NN, Ohsumi Y, Inagaki F. Atg8-family interacting motif crucial for selective autophagy. FEBS Lett. 2010;584:1379–85. https://doi.org/10.1016/j.febslet.2010.01.018.
Patel S, Dinesh-Kumar SP. Arabidopsis ATG6 is required to limit the pathogen-associated cell death response. Autophagy. 2008;4:20–7. https://doi.org/10.4161/auto.5056.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, et al. Bcl-2 Antiapoptotic Proteins Inhibit Beclin 1-Dependent Autophagy. Cell. 2005. https://doi.org/10.1016/j.cell.2005.07.002.
Peoples MB, Beilharz VC, Waters SP, Simpson RJ, Dalling MJ. Nitrogen redistribution during grain growth in wheat (Triticum aestivum L.): II. Chloroplast senescence and the degradation of ribulose-1,5-bisphosphate carboxylase. Planta. 1980;149:241–51. https://doi.org/10.1007/BF00384560.
Phillips AR, Suttangkakul A, Vierstra RD. The ATG12-Conjugating enzyme ATG10 is essential for autophagic vesicle formation in arabidopsis thaliana. Genetics. 2008;178:1339–53. https://doi.org/10.1534/genetics.107.086199.
Popa C, Li L, Gil S, Tatjer L, Hashii K, Tabuchi M, et al. The effector AWR5 from the plant pathogen Ralstonia solanacearum is an inhibitor of the TOR signalling pathway. Sci Rep. 2016. https://doi.org/10.1038/srep27058.
Popelka H, Damasio A, Hinshaw JE, Klionsky DJ, Ragusa MJ. Structure and function of yeast Atg20, a sorting nexin that facilitates autophagy induction. Proc Natl Acad Sci. 2017. https://doi.org/10.1073/pnas.1708367114.
Ralston SH. Pathogenesis of paget’s disease of bone. Bone. 2008;43:819–25. https://doi.org/10.1016/j.bone.2008.06.015.
Ramirez Montero DF. An assembly model for the autophagy initiation complex. 2019; https://hdl.handle.net/1721.1/123711
Ramundo S, Casero D, Mühlhaus T, Hemme D, Sommer F, Crèvecoeur M, et al. Conditional depletion of the chlamydomonas chloroplast clpp protease activates nuclear genes involved in autophagy and plastid protein quality control. Plant Cell. 2014;26:2201–22. https://doi.org/10.1105/tpc.114.124842.
Ran J, Hashimi SM, Liu JZ. Emerging roles of the selective autophagy in plant immunity and stress tolerance. Int J Mol Sci. 2020;21:6321. https://doi.org/10.3390/ijms21176321.
Ren W, Liu N, Sang C, Shi D, Zhou M, Chen C, et al. The Autophagy gene BcATG8 regulates the vegetative differentiation and pathogenicity of botrytis cinerea schottel JL, editor. Appl Environ Microbiol. 2018. https://doi.org/10.1128/aem.02455-17.
Ren W, Zhang Z, Shao W, Yang Y, Zhou M, Chen C. The autophagy-related gene BcATG1 is involved in fungal development and pathogenesis in Botrytis cinerea. Mol Plant Pathol. 2016;18:238–48. https://doi.org/10.1111/mpp.12396.
Rodriguez M, Parola R, Andreola S, Pereyra C, Martínez-Noël G. TOR and SnRK1 signaling pathways in plant response to abiotic stresses: do they always act according to the “yin-yang” model? Plant Sci. 2019;288: 110220. https://doi.org/10.1016/j.plantsci.2019.110220.
Sakai Y, Koller A, Rangell LK, Keller GA, Subramani S. Peroxisome degradation by microautophagy in pichia pastoris: identification of specific steps and morphological intermediates. J Cell Biol. 1998;141:625–36. https://doi.org/10.1083/jcb.141.3.625.
Schepetilnikov M, Kobayashi K, Geldreich A, Caranta C, Robaglia C, Keller M, et al. Viral factor TAV recruits TOR/S6K1 signalling to activate reinitiation after long ORF translation: TAV activates the TOR pathway. EMBO J. 2011;30:1343–56. https://doi.org/10.1038/emboj.2011.39.
Schwarten M, Mohrlüder J, Ma P, Stoldt M, Thielmann Y, Stangler T, et al. Nix directly binds to GABARAP: a possible crosstalk between apoptosis and autophagy. Autophagy. 2009;5:690–8. https://doi.org/10.4161/auto.5.5.8494.
Shinozaki D, Merkulova EA, Naya L, Horie T, Kanno Y, Seo M, et al. Autophagy increases zinc bioavailability to avoid light-mediated reactive oxygen species production under zinc deficiency. Plant Physiol. 2020;182:1284–96. https://doi.org/10.1104/pp.19.01522.
Shoji-Kawata S, Levine B. Autophagy, antiviral immunity, and viral countermeasures. Biochimica et Biophysica Acta (BBA). Mol Cell Res. 2009;1793:1478–84. https://doi.org/10.1016/j.bbamcr.2009.02.008.
Signorelli S, Tarkowski ŁP, Van den Ende W, Bassham DC. Linking autophagy to abiotic and biotic stress responses. Trends Plant Sci. 2019;24:413–30. https://doi.org/10.1016/j.tplants.2019.02.001.
Sláviková S, Shy G, Yao Y, Glozman R, Levanony H, Pietrokovski S, et al. The autophagy-associated Atg8 gene family operates both under favourable growth conditions and under starvation stresses in Arabidopsis plants. J Exp Bot. 2005;56:2839–49. https://doi.org/10.1093/jxb/eri276.
Spitzer C, Li F, Buono R, Roschzttardtz H, Chung T, Zhang M, et al. The endosomal protein charged multivesicular body protein1 regulates the autophagic turnover of plastids in arabidopsis. Plant Cell. 2015;27:391–402. https://doi.org/10.1105/tpc.114.135939.
Stefaniak S, Wojtyla Ł, Pietrowska-Borek M, Borek S. Completing autophagy: formation and degradation of the autophagic body and metabolite salvage in plants. Int J Mol Sci. 2020;21:2205. https://doi.org/10.3390/ijms21062205.
Suzuki K, Akioka M, Kondo-Kakuta C, Yamamoto H, Ohsumi Y. Fine mapping of autophagy-related proteins during autophagosome formation in Saccharomyces cerevisiae. J Cell Sci. 2013. https://doi.org/10.1242/jcs.122960.
Svenning S, Lamark T, Krause K, Johansen T. Plant NBR1 is a selective autophagy substrate and a functional hybrid of the mammalian autophagic adapters NBR1 and p62/SQSTM1. Autophagy. 2011;7:993–1010. https://doi.org/10.4161/auto.7.9.16389.
Takatsuka C, Inoue Y, Higuchi T, Hillmer S, Robinson DG, Moriyasu Y. Autophagy in tobacco BY-2 cells cultured under sucrose starvation conditions: isolation of the autolysosome and its characterization. Plant Cell Physiol. 2011;52:2074–87. https://doi.org/10.1093/pcp/pcr137.
Thumm M, Egner R, Koch B, Schlumpberger M, Straub M, Veenhuis M, et al. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 1994;349:275–80. https://doi.org/10.1016/0014-5793(94)00672-5.
Titorenko VI, Keizer I, Harder W, Veenhuis M. Isolation and characterization of mutants impaired in the selective degradation of peroxisomes in the yeast Hansenula polymorpha. J Bacteriol. 1995;177:357–63. https://doi.org/10.1128/jb.177.2.357-363.1995.
Tsukada M, Ohsumi Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 1993;333:169–74. https://doi.org/10.1016/0014-5793(93)80398-E.
Uemura T, Ueda T. Plant vacuolar trafficking driven by RAB and SNARE proteins. Curr Opin Plant Biol. 2014;22:116–21. https://doi.org/10.1016/j.pbi.2014.10.002.
Üstün S, Hafrén A, Liu Q, Marshall RS, Minina EA, Bozhkov PV, et al. Bacteria exploit autophagy for proteasome degradation and enhanced virulence in plants. Plant Cell. 2018;30:668–85. https://doi.org/10.1105/tpc.17.00815.
van Janse Rensburg HC, Van den Ende W, Signorelli S. Autophagy in plants: both a puppet and a puppet master of sugars. Front Plant Sci. 2019. https://doi.org/10.3389/fpls.2019.00014.
VILLIERS TA. Cytolysomes in Long-dormant plant embryo cells. Nature. 1967;214:1356–7. https://doi.org/10.1038/2141356b0.
Wang P, Mugume Y, Bassham DC. New advances in autophagy in plants: regulation, selectivity and function. Semin Cell Amp Dev Biol. 2018;80:113–22. https://doi.org/10.1016/j.semcdb.2017.07.018.
Wei Y, Liu W, Hu W, Liu G, Wu C, Liu W, et al. Genome-wide analysis of autophagy-related genes in banana highlights MaATG8s in cell death and autophagy in immune response to Fusarium wilt. Plant Cell Rep. 2017;36:1237–50. https://doi.org/10.1007/s00299-017-2149-5.
Williams B, Njaci I, Moghaddam L, Long H, Dickman MB, Zhang X, et al. Trehalose accumulation triggers autophagy during plant desiccation Qu LJ, editor. PLOS Genet. 2015;11:e1005705. https://doi.org/10.1371/journal.pgen.1005705.
Wingler A, Masclaux-Daubresse C, Fischer AM. Sugars, senescence, and ageing in plants and heterotrophic organisms. J Exp Bot. 2009;60:1063–6. https://doi.org/10.1093/jxb/erp067.
Wittenbach VA, Lin W, Hebert RR. Vacuolar localization of proteases and degradation of chloroplasts in mesophyll protoplasts from senescing primary wheat leaves. Plant Physiol. 1982;69:98–102. https://doi.org/10.1104/pp.69.1.98.
Wleklik K, Borek S. Vacuolar processing enzymes in plant programmed cell death and autophagy. Int J Mol Sci. 2023;24:1198. https://doi.org/10.3390/ijms24021198.
Wu Y, Cheng S, Zhao H, Zou W, Yoshina S, Mitani S, et al. PI3P phosphatase activity is required for autophagosome maturation and autolysosome formation. EMBO Rep. 2014;15:973–81. https://doi.org/10.15252/embr.201438618.
Xie Z, Klionsky DJ. Autophagosome formation: core machinery and adaptations. Nat Cell Biol. 2007;9:1102–9. https://doi.org/10.1038/ncb1007-1102.
Xiong Y, Contento AL, Nguyen PQ, Bassham DC. Degradation of oxidized proteins by autophagy during oxidative stress in arabidopsis. Plant Physiol. 2006;143:291–9. https://doi.org/10.1104/pp.106.092106.
Xu G, Wang S, Han S, Xie K, Wang Y, Li J, et al. Plant bax inhibitor-1 interacts with ATG6 to regulate autophagy and programmed cell death. Autophagy. 2017;13:1161–75. https://doi.org/10.1080/15548627.2017.1320633.
Yan Q, Wang J, Fu ZQ, Chen W. Endocytosis of AtRGS1 Is regulated by the autophagy pathway after D-glucose stimulation. Front Plant Sci. 2017. https://doi.org/10.3389/fpls.2017.01229.
Yano K, Matsui S, Tsuchiya T, Maeshima M, Kutsuna N, Hasezawa S, et al. Contribution of the plasma membrane and central vacuole in the formation of autolysosomes in cultured tobacco cells. Plant Cell Physiol. 2004;45:951–7. https://doi.org/10.1093/pcp/pch105.
Yorimitsu T, Klionsky DJ. Atg11 links cargo to the vesicle-forming machinery in the cytoplasm to vacuole targeting pathway. Mol Biol Cell. 2005;16:1593–605. https://doi.org/10.1091/mbc.e04-11-1035.
Yoshimoto K, Jikumaru Y, Kamiya Y, Kusano M, Consonni C, Panstruga R, et al. Autophagy negatively regulates cell death by controlling NPR1-dependent salicylic acid signaling during senescence and the innate immune response inarabidopsis. Plant Cell. 2009;21:2914–27. https://doi.org/10.1105/tpc.109.068635.
Yoshimoto K, Ohsumi Y. Unveiling the molecular mechanisms of plant autophagy – from autophagosomes to vacuoles in plants. Plant Cell Physiol. 2018. https://doi.org/10.1093/pcp/pcy112.
Yoshimoto K, Takano Y, Sakai Y. Autophagy in plants and phytopathogens. FEBS Lett. 2010;584(7):1350–8. https://doi.org/10.1016/j.febslet.2010.01.007.
Yu SM. Cellular and genetic responses of plants to sugar starvation. Plant Physiol. 1999;121:687–93. https://doi.org/10.1104/pp.121.3.687.
Yuan W, Tuttle DL, Shi YJ, Ralph GS, Dunn WA. Glucose-induced microautophagy in Pichia pastoris requires the α-subunit of phosphofructokinase. J Cell Sci. 1997;110:1935–45. https://doi.org/10.1242/jcs.110.16.1935.
J yu Y, J lan J, W wen W, X rui J, H zhong W. Silencing of the calcium-dependent protein kinase TaCDPK27 improves wheat resistance to powdery mildew. BMC Plant Biol. 2023. https://doi.org/10.1186/s12870-023-04140-y.
Zhang R, Zhang C, Lyu S, Fang Z, Zhu H, Hou X. Functional analysis of BcSNX3 in regulating resistance to turnip mosaic virus (TuMV) by autophagy in Pak-choi (Brassica campestris ssp. chinensis). Agronomy. 2022;12:1757. https://doi.org/10.3390/agronomy12081757.
Zhang Z, Feechan A, Pedersen C, Newman M, Qiu J, Olesen KL, et al. A SNARE-protein has opposing functions in penetration resistance and defence signalling pathways. Plant J. 2006;49(2):302–12. https://doi.org/10.1111/j.1365-313X.2006.02961.x.
Zhang Z, Zhang Y, Wang Y, Fan J, Xie Z, Qi K, et al. Exogenous dopamine improves resistance to Botryosphaeria dothidea by increasing autophagy activity in pear. Plant Sci. 2023;329:111603. https://doi.org/10.1016/j.plantsci.2023.111603.
Zheng X, Wu M, Li X, Cao J, Li J, Wang J, et al. Actin filaments are dispensable for bulk autophagy in plants. Autophagy. 2019;15(12):2126–41. https://doi.org/10.1080/15548627.2019.1596496.
Zhou G, Xu D, Xu D, Zhang M. Southern rice black-streaked dwarf virus: a white-backed planthopper-transmitted fijivirus threatening rice production in Asia. Front Microbiol. 2013. https://doi.org/10.3389/fmicb.2013.00270/abstract.
Zhou J, Wang J, Cheng Y, Chi YJ, Fan B, Yu JQ, et al. NBR1-mediated selective autophagy targets insoluble ubiquitinated protein aggregates in plant stress responses gassmann W editor. PLoS Genet. 2013;9:e1003196. https://doi.org/10.1371/journal.pgen.1003196.
Zhou J, Zhang Y, Qi J, Chi Y, Fan B, Yu JQ, et al. E3 Ubiquitin Ligase CHIP and NBR1-mediated selective autophagy protect additively against proteotoxicity in plant stress responses gassmann W, editor. PLoS Genet. 2014;10:e1004116. https://doi.org/10.1371/journal.pgen.1004116.
Zhu XM, Li L, Wu M, Liang S, Shi HB, Liu XH, et al. Current opinions on autophagy in pathogenicity of fungi. Virulence. 2018;10:481–9. https://doi.org/10.1080/21505594.2018.
Zhuang X, Chung KP, Cui Y, Lin W, Gao C, Kang BH, et al. ATG9 regulates autophagosome progression from the endoplasmic reticulum in Arabidopsis. Proc Natl Acad Sci. 2017. https://doi.org/10.1073/pnas.1616299114.
Zhuang X, Wang H, Lam SK, Gao C, Wang X, Cai Y, et al. A BAR-domain protein SH3P2, which binds to phosphatidylinositol 3-phosphate and ATG8, regulates autophagosome formation in arabidopsis. Plant Cell. 2013;25:4596–615. https://doi.org/10.1105/tpc.113.118307.
Zientara-Rytter K, Łukomska J, Moniuszko G, Gwozdecki R, Surowiecki P, Lewandowska M, et al. Identification and functional analysis of Joka2, a tobacco member of the family of selective autophagy cargo receptors. Autophagy. 2011;7(10):1145–58. https://doi.org/10.4161/auto.7.10.16617.
Zientara-Rytter K, Sirko A. To deliver or to degrade – an interplay of the ubiquitin–proteasome system, autophagy and vesicular transport in plants. FEBS J. 2016;283:3534–55. https://doi.org/10.1111/febs.13712.
Zientara-Rytter K, Subramani S. Mechanistic insights into the role of Atg11 in selective autophagy. J Mol Biol. 2020;432:104–22. https://doi.org/10.1016/j.jmb.2019.06.017.
Zvereva AS, Golyaev V, Turco S, Gubaeva EG, Rajeswaran R, Schepetilnikov MV, et al. Viral protein suppresses oxidative burst and salicylic acid-dependent autophagy and facilitates bacterial growth on virus-infected plants. New Phytol. 2016;211:1020–34. https://doi.org/10.1111/nph.13967.
Acknowledgements
AB has not received any funding for this study. AJ is grateful to the Department of Science and Technology, India for grant under the DST Women Scientist scheme [SR/WOS-A/ LS-377/2018]. SD acknowledges Indian National Science Academy for her senior scientist fellowship.
Funding
No funding has received to perform this work.
Author information
Authors and Affiliations
Contributions
AB conceived the idea; AB, AJ wrote the manuscript; AB prepared the figures; DBB helps in final editing and referencing, SD read, modified and finally approved the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
This article is dedicated to Prof. Arun Kumar Sharma to commemorate his Birth Centenary.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Corresponding Editor: Umesh C. Lavania; Reviewers: Indranil Dasgupta, Sujit Roy.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bhar, A., Jain, A., Banerjee, D.B. et al. From cellular cleanup to defense: the stepwise process of plant autophagy with special reference to their crucial role in biotic stress tolerance. Nucleus (2024). https://doi.org/10.1007/s13237-024-00505-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13237-024-00505-2