Abstract
Microbes participate in a series of complex biogeochemical processes including nitrogen and sulfur circulation. Microbial species active in the nitrogen and sulfur cycles can be used for nitrogen and sulfur pollution remediation, preserving and keeping the ecosystem in balance. Hypolentic zones in brine lakes contain stable, elevated pH and total dissolved solids and provide a unique habitat for a rich diversity of haloalkaliphilic bacteria and archaea. Few studies have investigated the diversity and microbial community structure in the sedimentary environment of hypolentic zones in brine lakebeds located on desert plateaus. In this study, sedimentary environmental characteristics, species abundance and diversity were investigated, as well as the relationships between them. Analyses revealed important roles of different bacterial species in the microbially mediated nitrogen and sulfur biogeochemical cycles in the brine lakebed. Further, the study revealed details of the ecosystem dynamics of these extreme environments, under the action of microorganisms. The biogeochemical dynamic response of different hydraulic and hydrochemical characteristics was illuminated. It showed that the amount of deoxyribonucleic acid in the sediment was interrelated with the components of brine water and depth of the hypolentic zone and was higher in brine and reduction conditions, with the stronger hydraulic forces. In the hypolentic zone of the brine lake, microbial species composition was remarkably correlated with environmental factors, spatial environmental factors and hydrogeological conditions, such as hydraulic and hydrochemical characteristics.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Under the control of regional topography and climate, lakes have developed and are distributed across wide expanses of arid desert plateaus. These lakes are important regional groundwater drainage datum planes, for maintaining wetland and ecological vegetation (Mallast et al. 2014). In such regions, most lakes are saline or brine lakes, owing to the high levels of evaporation. Brine lakebeds have stable, elevated pH levels and total dissolved solids (TDS) concentrations. Indeed, the TDS concentration is much higher in such lakes than in the sea (Hallsworth et al. 2007; Wetzel 2001). Brine lakebeds provide a unique habitat to a rich diversity of haloalkaliphilic bacteria and archaea. Under the influence of human activities, interactions between the surface water and groundwater occur in a region known as the lake water–groundwater interaction zone or hypolentic zone (Teng et al. 2009). The hypolentic zone plays an important role in regional water cycles and the process of material circulation (Sophocleous 2002; Teng et al. 2009), because of the continuous transmission of water, solutes, nutrients and energy. The microbial community is a major component of a groundwater ecosystem and a key participant in biogeochemical evolution processes in the hypolentic zone. DNA is the carrier of biological information and can be used as an ideal marker for accurately identifying the microbial community structure and diversity in the hypolentic zone.
In brine lake ecosystems, the cycling of elemental sulfur and nitrogen in the sediment records environmental changes. Therefore, a wealth of information is stored, including vegetation succession, impacts of glaciers, links to global change and the ore-forming process in the Ordos basin (Feng et al. 2007; Zhai 2008; Jiang et al. 2007). However, the diversity and microbial community structure has several researches in deep-sea brine lakes (Hallsworth et al. 2007; Gugliandolo et al. 2015), but in sedimentary environments of brine lakebed on desert plateaus, characterized by high salt, total dissolved solids (TDSs) and alkalinity level, has not been properly investigated (Wang 2014).
In this study, bacteria in lakebed sedimentary environments that are subject to extreme conditions, such as high salt, TDSs and alkalinity levels were investigated, based on 16S rRNA. Lakebed sediments were collected to monitor and analyze the microbial communities and to identify and analyze microbial abundance in pore waters under various chemical conditions. Combining high-throughput sequencing technology with canonical correspondence analysis (CCA) has enabled detailed surveys of environmental gene banks and objective investigations of the potential relevance of biogeochemical functions between microbial communities (Huo et al. 2015; Johnson et al. 2015a, b). It has also allowed exploration of microbial community structure and relative abundance in complex environments (Sogin et al. 2006; Lauber et al. 2009; Ushio et al. 2014). This investigation also sought to reveal the response mechanisms based on the microbial structure and diversity, and the spatial–temporal distribution of environmental characteristics in the inland brine lakes of the desert plateau. These results should provide baseline measures for sediments and geochemical studies in desert plateau brine lakebeds, while supporting ecosystem protection in these lakebeds.
Sampling and methods
Study area
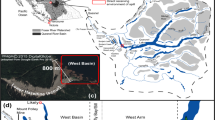
Ordos’ desert plateau is located in the northwest of China, to the west of Inner Mongolia’s autonomous region (Hou et al. 2008). It is an important center for the energy and chemical industry (Fig. 1). Dakebo Lake is located on the central part of the Ordos desert plateau within Etuokeqi County, Inner Mongolia Autonomous Region, which is topographically lower than the surrounding land. It has an approximately elliptic contour, with 4 km2 area and shallow water depth of 3–4 m. Desert plateaus are characterized by dry climates, which leads to strong evaporation, scarce and unevenly distributed precipitation, extremely scarce surface water and a very fragile ecosystem (Feng et al. 2007). The main groundwater discharge in the study area is precipitation infiltration and lateral runoff. The major mineral components of the lake sediments are Glauber’s salt, trona and rock salt (Guo 2012; Yang 2013; Wang 2014) (Table 1).
As shown in the hydrogeological profile (Fig. 2), the shore of the lakebed (HB2) and the central lakebed (HX) sediment hydrogeological structure has horizontal and vertical zonation. The lakebeds are covered with sandy loam, consisting of pores with good permeability. The lakes are easily infiltrated by atmospheric precipitation. Moreover, the lakebed bottom is covered with silty loam of different thicknesses. Groundwater mainly occurs in the Quaternary and Cretaceous aquifer, and shallow groundwater is primarily recharged by atmospheric precipitation. The hydraulic gradient allows groundwater to run off from around or the bottom of the lake controlled by lithology and topography, eventually the lake water is lost by evaporation. In HX, groundwater recharges the lake in a vertical direction. At the two sides of the lake, such as in HB2, groundwater is recharged from both vertical and lateral directions. The HX has different hydraulic and hydrochemical characteristics compared with HB2. Along with the differing lakebed sedimentary environment, biogeochemical reactions are very active in HB2. The microbial community responds to this, as it is the main component of life in groundwater ecosystems and an important part of biogeochemical cycles (Yang 2013; Wang 2014).
Sampling
The monitoring section was arranged approximately east–west across the lake (Fig. 2). No specific permissions were required for these locations, and the field studies did not involve endangered or protected species. Monitoring was conducted at different depth intervals across the lake. Factors considered included the mixing of different water bodies, the water chemical composition and the gradual change in environmental indicators, which tend to decrease gradually with depth in the hypolentic zone (Guo 2012). Major monitoring was conducted in two zones of the lake: HX and HB2. Groundwater hydrochemistry, sediments and microbial samples were collected in the study area.
Collection and preservation methods:
-
(a) Water sample collection
After flushing the wells, water samples were collected by a Proactive Mega-Typhoon-type pump (1767 Lakewood Ranch Blvd, Suite #113, Bradenton, FL |34211 USA). The pump was equipped with a professional controller that can achieve stable low-flow sampling (minimum sampling speed of 0.95 L/min). This low speed prevents changes in water oxidation–reduction-sensitive indicators (such as Eh and DO) during sampling, which can occur via disturbance caused by severe pumping (Yang 2013).
-
(b) Sediment sample collection
Undisturbed surface soil samples were collected from monitoring points in HX and HB2, each point within 1 m of the surface. Samples were collected by a Beeker portable lake sediment sampler (Eijkelkamp Agrisearch Equipment, Giesbeek, Netherlands) and a manual drill. Additional samples were collected from about 1–5 m using a manual drill.
-
(c) Microbial sample collection
Before sampling, the samplers were wiped with alcohol. Once the equipment had dried, microbial samples were taken at different depths, then sealed in sterile valve bags and quickly placed in a refrigerator at −20 °C for cryopreservation (Gurtner et al. 2000).
Test methods
Sensitive indices including water level, temperature, pH, Eh, DO and TDS were tested on-site using Hach portable electrodes, such as PHC, MTC and LDO. Iron and sulfide were also measured using a Hach D2800 spectrophotometer by test kits of 2105769 (total iron), 103769 (Fe2+), 2106769 (sulfate) and 2244500 (sulfide). Anion–cation ions such as K+, Na+, Ca2+, Mg2+, Cl−, CO3 2−, SO4 2−, HCO3 − and NO3 − were tested by ion chromatography. DNA was extracted directly from the soil after a purification treatment, using a suitable primer to amplify the 16S rRNA genes (Zhang et al. 2009; Liu et al. 2011; Wang 2014), through high-throughput sequencing technology, for detailed surveys of environmental gene banks and objective investigations of the potential relevance of biogeochemical functions between microbial communities, exploration of microbial community structure and relative abundance in complex environments.
Bacteria from lakebed sediments at different depths around HB2 and HX were analyzed. The Fast DNA®Spin Kit for Soil was used, combined with the MP Fast Prep Instruments (MP Biomedicals), to extract bacterial DNA from the sediments in the study area. The extracted genomic DNA was subject to agarose gel electrophoresis to check the integrity and concentration of genomic DNA, using a micro-ultraviolet spectrophotometer, NanoDrop®ND-1000UV-Vis (3411 Silverside Road Bancroft Building Wilmington, DE 19810 USA) (Ushio et al. 2014). The V4–V5 regions of 16S rRNA were amplified from the total DNA, using the prokaryotic universal primers 515F (GTGYCAGCMGCCGCGGTA) and 806R (GGACTACHVGGGTWTCTAAT), based on the Illumina sequencing methods (Caporaso et al. 2011, 2012; Wang and Qian 2009). The successfully amplified PCR products were preserved in a refrigerator at −20 °C (Liu et al. 2011) for use in subsequent experiments. Unweighted pair group method with arithmetic mean cluster was used to examine the similarity between the lanes. The sample of genomic PCR products was examined by the high-throughput sequencing technology. Identifying the different number of electrophoresis bands helps identify the diversity of different biological communities at different depths in the hypolentic zone.
The α biodiversity index (Shannon–Wiener index and Simpson index) and diversity indices like CHAO (Moura et al. 2009) were calculated to determine diversity and change trends of microbial communities, in different spaces and different depths of Dakebo Lake. The Shannon–Wiener index (H′) reflects the biological community structure in the environment and its degree of abundance diversity. The Simpson index (D) represents ecological dominance, reflecting the function and position of superior species that are more abundant in the community (Moura et al. 2009).
CCA was originally developed to relate community composition to environmental variables. CCA has enabled detailed investigations of the potential relevance of microbial community structure and spatial distribution of water chemistry, in complex environments. In this study, CCA was used to demonstrate relationships between microbial operational taxonomic unit composition and the characteristic indexes, using the software program CANOCO (version 4.5).
Results and discussion
Spatial distribution of DNA content in lakebed sediments
The spatial distribution of DNA content in lakebed sediments varied in the hypolentic zone (Fig. 2). The microbial DNA content in sediments differs in different hydrodynamic and hydrochemical locations of the hypolentic zone. The vertical profile of the hypolentic zone in the west of HB2 was divided into two different kinds of interaction region: the surrounding groundwater/brine lake interaction region (GBI) and the brine–hypolentic zone/deep pore water interaction region (BDI). In the lakebed sediments of HB2, DNA content was higher, and there were more microorganisms in the BDI, as well as in the GBI. In the lakebed sediments of HX, DNA content was lower in the BDI of the shallow lake-hypolentic zone. The microbial community structure responded to the unique hydraulic and hydrochemical characteristics of the lake, as indicated by significant differences in DNA content. These differences directly influenced spatial patterns in the environment. DNA content was higher in regions of the lakebed with stronger hydraulic conditions. The stronger hydraulic force in the hypolentic zone provides continuous transmission of water, solutes, nutrients and energy, needed for the activities of the bacteria.
Diversity analysis of microbial community
Within the study area, the microbial community structure of samples had high similarity to each other, but there were still some differences in the dominant bacteria and archaea at the genus level (Fig. 3). The abundance of microbes was significantly higher in HB2 than in HX. The content of acidophilic acidobacteria and halophilic archaea in the study area was high. In the HX and HB2, the dominant species, with a higher percentage of reads, included Marinobacter, Halomonas, Rhodobaca, Thioalkalivibrio, Alcanivorax, Candidatus Nitrososphaera, Anaerobacillus, Dehalobacter, Bacillus, Pseudomonas, Alkaliphilus and Portiera. The dominant species have developed various bioenergetic and structural adaptations to maintain pH and TDS homeostasis and intracellular osmotic pressure, and thus survive in the extreme environment (Spang et al. 2012; Sorokin et al. 2011, 2012a, b, 2014).
Denitrification in the brine lakebed is performed by heterotrophs, dominated by the denitrification ability of extremely salt-tolerant alkaliphiles, such as Marinobacter and Halomonas (Sorokin et al. 2012a, b; Duckworth et al. 1996; Grant and Sorokin et al. 2011). At the surrounding groundwater/lake–brine water interaction regions of HB2, Proteus mirabilis (Hauser) had become the dominant species. Desulfosporosinus, Desulfococcus and several lithotrophic sulfate-reducing bacteria, namely Desulfonatronospira and Desulfonatronovibrio (deltaproteobacterial genera) were found in the brine lakebeds. More Desulfonatronospira were found in HB2, which had higher TDS, than in HX (Fig. 4). It appears that microbial diversity plays an important role in the nitrogen and sulfur cycles in brine lakebed ecosystems. Previous studies have shown that the unique hydraulic exchanges often influenced nutrient or pollutants flux, biogeochemical reaction process, such as the reaction rate and degree of reactivity. And it directly altered the acid, alkali and redox conditions, which decided the water chemical reaction type and the chemical evolution of pore water, due to the different hydrochemical compositions between groundwater and lake water (Yang 2013; Wang 2014). The redox potentials have a certain response relationship with biogeochemical reaction process, hydrodynamic and hydrochemical exchange, aquifer medium structure and microbial community structure. These changed the survival environment for microorganisms, selecting the microbial species in the hypolentic zone (Hiscock and Grischek 2002; Tufenkji et al. 2002; Yang 2013; Wang 2014).
There were remarkable correlations in the vertical distribution of the Shannon–Wiener index and Simpson index of each location section (Figs. 5, 6). In the HX section, both the Shannon–Wiener index and Simpson index were higher in the lake-hypolentic zone with deep pore water interaction regions, than in the deep pore water. The microbial community structure and diversity was more abundant in the interaction regions. As aforementioned, the vertical profile of the hypolentic zone, at the west of HB2, was divided into two different kinds of interaction region: GBI and BDI. The microbial Simpson and Shannon–Wiener indices also showed well-zoned characteristics, consistent with the interaction regions. Indices were lower in the area surrounding the GBI.
In the both monitored cores (HX, HB2), the vertical distribution of the microbial Shannon–Wiener index and the Simpson index has two points in common:
-
(a) The higher indices were found in the interaction region;
-
(b) The region with the higher index was also a region of relatively low oxidation–reduction potential (HX: 0.2–3 m; HB2: 1–3 m).
Correlation analysis of environmental variables and microbial vertical distribution
Correlations between the environmental variables [TDS, pH, Eh, DO, sulfide, sulfate and total organic carbon (TOC)] and DNA content (Figs. 7, 8, 9, 10, 11) were analyzed. With the increase in dissolved oxygen content in the environment, there was decreased DNA content and microbial activity. Microbial DNA content and microbial community diversity tended to increase as Eh decreased (Fig. 8). The DO content in the environment was 0–5 mg/L. The low oxygen content led to a decrease in (obligate) aerobes. At the same time, when dissolved oxygen increased, the activity of the anaerobic prokaryotes was inhibited. Heterotrophic anaerobic fermentative haloalkaliphiles, such as the moderately salt-tolerant, Anaerobacillus diazotrophicus, actively fix nitrogen in brine lakebeds and water. Firmicutes were dominant in HX, with low DO/Eh content, high salt content and intense mixing with deep pore water. In the hypolentic zone, facultative anaerobic bacteria become dominant bacteria.
The study area was an alkaline environment, with pH ranging from 10.2 to 10.7 (Fig. 9). Species diversity in the environment was relatively limited. However, with an increase in basicity, alkalophilic microorganisms strengthened their microbial metabolic activity. These microorganisms have developed various bioenergetic and structural adaptations to maintain pH and TDS homeostasis and intracellular osmotic pressure, and thus survive in the extreme environment. The Alkaliphiles have been found in this area in environments with relatively high pH (Liu et al. 2016; Jie et al. 2016).
Indeed, the total dissolved solids (TDS) in such lakes are much higher than in the sea (Hallsworth et al. 2007; Wetzel 2001). With the increase in TDS, the DNA content had a decreasing trend (Fig. 10). Inhibiting the metabolic activity of microbes and increasing the water infiltration pressure led to a reduction in the biological degree of diversity. Some cells even exhibit plasmolysis and inactivation. With an increase in TDS, the secretion of microbial extracellular polymers was stimulated, inhibiting microbial activity (Wang 2014). Hypolentic zones in brine lakes contain stable elevated pH and TDS and provide a unique habitat to a rich diversity of haloalkaliphilic bacteria and archaea. Firmicutes were dominant in this area, which had a low DO content and high TDS content. Thioalkalivibrio can live in a wide range of salinities in this area. Carbon and nitrogen sources for microbial growth can be provided by TOC; as TOC concentration increased, microbial abundance increased (Fig. 11).
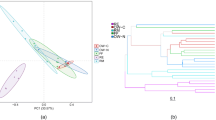
The diversity of the HB2 samples was significantly higher than that of the HX samples. Diversity index analysis revealed that the microbial diversity of HB2 and HX increased dramatically as the environment changed from oxidizing to reducing conditions. The diversity of the community structure of HB2 was highest at a depth of 1.4 m below the lakebed, which corresponded to the highest brine content (Table 2). The species compositions showed completely different trends and characteristics in HX and HB2, because of the impact of the differing hydraulic and hydrochemical characteristics. The principal component analysis (PCA) results revealed that the species composition and diversity of the HB2 sample were more abundant than those of HX (Fig. 12). The HX species composition had a higher similarity at the boundary to 0.9 m to the both monitored cores (HX and HB2) samples.
Correlations between environmental variables and species in the research area were analyzed (Figs. 13, 14). Detrended correspondence analysis (DCA), in which lengths of gradient have a maximum of 3.585, met the requirements of CCA. Microbial species composition and environmental factors in HB2 and HX were well correlated. From the CCA ordinations (Table 3), the distribution of the species was primarily correlated with S2− and TOC. However, in HB2, there was also a positive correlation between species and pH, TOC, sulfide content, DO content of pore water in the sediments. In the stronger hydraulic conditions (depth 0.9–1 m) of HB2 and HX, there was a positive correlation between NO3 − and the extremely salt-tolerant alkaliphilic genera, such as Marinobacter and Halomonas, due to their denitrification ability (Shapovalova et al. 2009). The maximum salt concentration for nitrification was 2.5 M of Na+ in HX and 6 M around HB2. The nitrogen cycle was inhibited at high pH (about 10.3) and in hypersaline environments, causing potential N loss from the ecosystem (Conrad et al. 1995; Sorokin et al. 2014) (Table 4).
The representatives of the genus Thioalkalivibrio, which were screened for sulfur-oxidizing bacteria, showed an inverse correlation with DO and a positive correlation with SO4 2−. Thioalkalivibrio were mainly distributed in reducing environments and high brine content areas (Hoeft et al. 2007; Sorokin et al. 2014). Several obligatory anaerobic and obligatory haloalkaliphilic bacteria, such as lithotrophic sulfate-reducing bacteria (SRB), can perform dissimilatory reductions of oxidized sulfur compounds in brine lakebeds (Sorokin et al. 2014). Heterotrophic SRB were oxidizers, using several volatile fatty acids (VFA) as e-donors or C-sources, with sulfates as e-acceptors. Several different lineages of obligatory anaerobic haloalkaliphiles were implicated in sulfur reduction. The Desulfonatronobacter can oxidize VFA completely with sulfate or others as e-acceptors, showing significant correlations with SO4 2−, in sulfate-reducing reactions (Sorokin et al. 2014). Additionally, there was a different correlation between dominant species and environmental variables (DO, NO3 −, TDS and SO4 2−), in HX and HB2, owing to different hydraulic and hydrochemical characteristics such as the redox conditions (Table 5).
Conclusions
The microbial community structure was closely associated with the unique hydraulic and hydrochemical characteristics of the lake, as indicated by significant differences in microbial abundance and community structure. These differences were directly influenced by spatial patterns in the environment and the mineralization process, such as Glauber’s salt with respect to sulfate-reducing bacteria.
The diversity of the microbial community structure in the HB2, which had stronger hydraulic conditions and higher TDS (25–150 g/L), was higher than in the HX. Additionally, the HB2 was higher in reducing environments than in oxidizing environments and more abundant in brine pore water.
The content of acidophilic acidobacteria and halophilic archaea in the study area was high. Dominant organisms included Proteobacteria (Marinobacter, Rhodobaca Halomonas and Thioalkalivibrio), Firmicutes (Bacillus), Clostridia (Alkaliphilus and Anaerobacillus), Actinomycetes and Acidobacteria.
There was a significant correlation between the dominant bacteria and the environmental factors in this extreme environment, on both a spatial and temporal scale. There was a dynamic response of the unique hydraulic and hydrochemical characteristics, and ecological function of dominant species, which are involved in nitrogen/sulfur cycles in the brine lakebed.
References
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108(S1):4516–4522
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6(8):1621–1624
Conrad R, Frenzel P, Cohen Y (1995) Methane emission from hypersaline microbial mats: lack of aerobic methane oxidation activity. FEMS Microbiol Ecol 16:297–306
Duckworth AW, Grant WD, Jones BE, van Steenburgen R (1996) Phylogenetic diversity of soda lake alkaliphiles. FEMS Microbiol Ecol 9:181–191
Feng ZD, Zhai XW, Wang WG, Zhang HC et al (2007) Eolian climatic variations during the past 30,000 years in the northern Mongolian Plateau, as indicated by geophysical, geochemical and geobotanical proxy date. Paleogeogr Paleoclimatology Paleoecol 245:505–517
Gugliandolo C, Lentini V, Bunk B, Overmann J, Italiano F, Maugeri T (2015) Changes in prokaryotic community composition accompanying a pronounced temperature shift of a shallow marine thermal brine pool (Panarea Island, Italy). Extremophiles 19:547–559
Guo JM (2012) Research on surfur biogeochemistry in hypolentic zone of lake in Kubuqi Desert Plateau Area. Jilin University
Gurtner C, Heyrman J, Piñar G, Lubitz W, Swings J, Rölleke S (2000) Comparative analyses of the bacterial diversity on two different biodeteriorated wall paintings by DGGE and 16S rDNA sequence analysis. Int Biodeterior Biodegradation 46:229–239
Hallsworth JE, Yakimov MM, Golyshin PN, Gillion JLM, D’Auria G, De Lima Alves F (2007) Limits of life in MgCl2-containing environments: chaotropicity defines the window. Environ Microbiol 9:801–813
Hiscock KM, Grischek T (2002) Attenuation of groundwater pollution by bank filtration. J Hydrol 266:139–144
Hoeft SE, Switzer Blum J, Stolz JF, Tabita FR, Witte B, King GM, Santini JM, Oremland RS (2007) Alkalilimnicola ehrlichii sp. nov., a novel arsenite-oxidizing, haloalkaliphilic gammaproteobacterium capable of chemoautotrophic or heterotrophic growth with nitrate or oxygen as the electron acceptor. Int J Syst Evol Microbiol 57:504–512
Hou GC, Liang YP, Su XS, Zhao ZH, Tao ZP, Yin LH, Yang YC, Wang XY (2008) Groundwater systems and resources in the Ordos Basin, China. Acta Geol Sin 5(6):801–811
Huo S, Ma C, Xi B, Su J, He Z, Li X (2015) Establishing water quality reference conditions for nutrients, chlorophyll a and Secchi depth for 7 typical lakes in arid and semiarid ecoregion, China. Environ Earth Sci 73:4739–4748
Jiang H, Dong H, Yu B, Liu X, Li Y, Ji S (2007) Microbial response to salinity change in Lake Chaka, a hypersaline lake on Tibetan plateau. Environ Microbiol 9:2603–2621
Jie T, Shuang L et al (2016) Effect of freeze–thaw cycles on carbon stocks of saline–alkali paddy soil. Arch Agron Soil Sci. doi:10.1080/03650340.2016.1159301
Johnson SS, Chevrette MG, Ehlmann BL, Benison KC (2015a) Insights from the metagenome of an acid salt lake: the role of biology in an extreme depositional environment. PLoS One 10:e0122869
Johnson WP, Swanson N, Black B, Rudd A, Carling G, Fernandez DP (2015b) Total- and methyl-mercury concentrations and methylation rates across the freshwater to hypersaline continuum of the Great Salt Lake, Utah, USA. Sci Total Environ 511:489–500
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-Based Assessment of Soil pH as a Predictor of Soil Bacterial Community Structure at the Continental Scale. Appl Environ Microbiol 75:5111–5120
Liu DS, Wang HY et al (2011) The review of DGGE technology application in the wastewater biological nitrogen removal system. Anhui Agric Sci 19:11695–11697
Liu W, Zhang W, Liu G, Zhang Y, Zhang G (2016) Microbial diversity in the saline-alkali soil of a coastal Tamarix chinensis woodland at Bohai Bay, China. J Arid Land 8(2):284–292
Mallast U, Gloaguen R, Friesen J, Rodiger T, Geyer S, Merz R, Siebert C (2014) How to identify groundwater-caused thermal anomalies in lakes based on multi-temporal satellite data in semi-arid regions. Hydrol Earth Syst Sci 18(7):2773–2787
Moura A, Tacao M, Henriques I (2009) Characterization of bacterial diversity in two aerated lagoons of a wastewater treatment plant using PCR-DGGE analysis. Microbiol Res 164(5):560–569
Shapovalova AA, Hizhniak TV, Tourova TP, Muyzer G, Sorokin DY (2009) Halomonas chromatireducens sp. nov., a novel haloalkaliphile from soda soil capable of aerobic chromate reduction. Microbiology (Moscow) 78:117–127
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR, et al (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci 103:12115–12120
Sophocleous M (2002) Interactions between groundwater and surface water: the state of the science. Hydrogeol J 10:52–67
Sorokin DY, Panteleeva AN, Tourova TP, Kaparullina EN, Muyzer G (2011) Natronoflexus pectinivorans gen. nov., sp. nov., an obligately anaerobic and alkaliphilic fermentative member of Bacteroidetes from soda lakes. Extremophiles 15:691–696
Sorokin DY, Tourova TP, Mordanov AV, Ravin NV (2012a) Microbial chitin utilization at extremely haloalkaline conditions. Extremophiles 16:883–894
Sorokin DY, Tourova TP, Panteleeva AN, Kaparullina EN, Muyzer G (2012b) Anaerobic utilization of pectinous substrates at extremely haloalkaline conditions by Natranaerovirga pectinivora gen. nov., sp. nov., and Natranaerovirga hydrolytica sp nov., isolated from hypersaline soda lakes. Extremophiles 16(2):307–315
Sorokin DY, Gumerov VM, Rakitin AL, Beletsky AV, Damsté JS, Mardanov AV, Ravin NV (2014) Genome analysis of Chitinivibrio alkaliphilus gen. nov., sp. nov., a novel extremely haloalkaliphilic anaerobic chitinolytic bacterium from the candidate phylum TG3. Environ Microbiol 16:1549–1565
Spang A, Poehlein A, Offre P et al (2012) The genome of the ammonia-oxidizing Candidatus Nitrososphaera gargensis: insights into metabolic versatility and environmental adaptations. Environ Microbiol 14(12):3122–3145
Teng YG, Zhang Zh, Feng D (2009) Pollutants in the river – groundwater interaction biological geochemical behavior and its detection technology. J Beijing Normal Univ (Nat Sci) Z1:515–519
Tufenkji N, Ryan JN, Elimelech M (2002) The promise of bankfiltration-a simple technologymay inexpensively clean up poorqualityraw surface water. Environ Sci Technol 36(21):422A–428A
Ushio M, Makoto K, Klaminder J, Takasu H, Nakano S-I (2014) High-throughput sequencing shows inconsistent results with a microscope-based analysis of the soil prokaryotic community. Soil Biol Biochem 76:53–56
Wang H (2014) Research on biogeochemical characteristics of sulfur and zonation in hypolentic zone in the Ordos Desert Plateau. Jilin University
Wang Y, Qian PY (2009) Conservative fragments in bacterial 16S rRNA genes and primer design for 16S ribosomal DNA amplicons in metagenomic studies. PLoS One 4(10):e7401
Wetzel RG (2001) Limnology, 3rd edn. Academic Press, San Diego
Yang GQ (2013) Research on the characteristics of hydraulic exchange and hydrochemical evolution in hypolentic zone of Dakebo Lake in Ordos Desert Plateau Area. Jilin University
Zhai XW (2008) The Holocene climate and environment changes in the Mongolian Plateau. Lanzhou University
Zhang HX, Tan ZJ et al (2009) The DGGE/TGGE technology progress in the study of soil microbial diversity. Nucl Agron 04:721–727
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (No. 41073054).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yuan, W., Su, X., Cui, G. et al. Microbial community structure in hypolentic zones of a brine lake in a desert plateau, China. Environ Earth Sci 75, 1132 (2016). https://doi.org/10.1007/s12665-016-5914-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12665-016-5914-9