Abstract
Gamma-aminobutyric acid (GABA) is a principal inhibitory neurotransmitter in the central nervous system and is produced by irreversible decarboxylation of glutamate. It possesses several physiological functions such as neurotransmission, diuretic, and tranquilizer effects and also regulates cardiovascular functions such as blood pressure and heart rate in addition to playing a role in the reduction of pain and anxiety. The objective of this study was to evaluate the GABA producing ability and probiotic capability of certain lactic acid bacteria strains isolated from dairy products. Around sixty-four bacterial isolates were collected and screened for their ability to produce GABA from monosodium glutamate, among which nine isolates were able to produce GABA. The most efficient GABA producer was Enterococcus faecium BS5. Further, assessment of several important and desirable probiotic properties showed that Ent. faecium BS5 was resistant to acid stress, bile salt, and antibiotics. Ent. faecium BS5 may potentially be used for large-scale industrial production of GABA and also for functional fermented product development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gamma-aminobutyric acid (GABA) is a widely distributed non-protein amino acid. It is produced from glutamate through irreversible alpha-decarboxylation by glutamate decarboxylase (GAD). GABA acts as a major neurotransmitter in the mammalian nervous system [1]. It has diuretic, anti-depression, and tranquilizing effects [2]. It induces hypotension and regulates growth hormone secretion. It has been used in the regulation of physiological disorders, including epilepsy, convulsions, Parkinson’s disease, Alzheimer’s disease, and Huntington’s chorea. GABA is widely distributed in the environment [3] and has notable physiological functions as an anti-stress molecule [4] inhibiting the spread of cancer [5] and preventing diabetes [6].
Microbial-derived GABA regulates visceral hypersensitivity [7], and also, in different parts of the GI tract, GABA and GABA receptors have functions, that focus on the regulation of GI motility and inflammation [8]. Supplementation of GABA regulates intestinal functions including intestinal immunity, intestinal amino acid profiles, gut microbiota [9], and the promotion of jejuna SIgA secretion. This could be linked to the T cell-dependent pathway and altered structure of gut microbiota as well as metabolism [10].
Furthermore, GABA is a bioactive component in pharmaceuticals as well [11]. These numerous value-added benefits of GABA on human health have recently gained increasing interest in the food industry. Many GABA-enriched functional foods are currently being developed such as tea leaves that are anaerobically treated, water-soaked rice germs, red mold rice, tempeh like fermented soybeans, and dairy products such as yogurt, fermented milk products, and cheese [12]. Therefore, the due acknowledgement should be given to the mass production of GABA and its subsequent use in the modern food industry as a bioactive food ingredient.
The screening of GABA-producing lactic acid bacteria (LAB) and the production of GABA-enriched food by LAB are currently being actively considered and investigated. Some researchers reported GABA-producing LAB strains from different sources, including Lactobacillus brevis from kimchi [13,14,15], L. brevis from pacoai [16], L. paracasei from Traditional fermented fish [17], L. plantarum from kimchi [18], L. sakei from jeot-gal, Korean fermented seafood [19], Lactococcus lactis from kimchi [20], and L. farciminis from fishery products [12]. The food industry needs to screen and find out whether various LAB have the ability to produce GABA because acid production, taste, and flavour formation ability, etc., are specific profiles of LAB. These profiles are regarded as essential factors in the use of LAB as starters in fermented food production [21].
Among the various GABA producing microbes, L. brevis, L. plantarum, and L. futsaii [18, 22, 23] have been assessed for their probiotic characteristics, making a good choice as a starter ingredient for functional GABA-enriched foods. These probiotic bacteria are thus widely used as starter cultures in the food industry [24]. They have to survive in diverse conditions such as the acidic condition in the stomach and in the presence of bile acids in the intestines to assimilate in the intestinal tract. These diverse conditions represent the essential parameters for selecting probiotics [25]. Also, different bacterial strains affect the host through different mechanisms of action [26, 27]. Therefore, the probiotic ability was shown to be species dependent. Selection of probiotic strains based their ability to modulate the very specific physiological properties such as GABA production is essential for demonstrating a probiotic health effects.
Numerous studies have reported LAB belonging to the genera Lactobacillus and Lactococcus as the most GABA producing organisms, while a few studies on the genus Enterococcus have also been published. Considering the above, the current study aimed to evaluate the capability of LAB isolated from dairy products to produce γ-aminobutyric acid (GABA), and also analyse the role of this LAB as a probiotic.
Materials and Methods
Ten samples of dairy products such as milk, commercial curd, and yogurt were collected from different places in Coimbatore, Tamil Nadu. The collected samples were stored in sterile bottles and transported in cooler boxes to the laboratory. The bacterial population was isolated by serial dilution of samples using sterile peptone physiological saline (1% peptone, and 0.85% w/v NaCl), and after incubation, plates with 30–100 colonies were enumerated and colonies with distinct morphological differences were selected and then purified using MRS agar media (Himedia, Mumbai, India). These isolates were stored in MRS broth (Himedia, Mumbai, India) containing 50% sterile glycerol at − 80 °C. The isolates were subcultured in MRS broth at 37 °C for 24 h and used as inoculum for the following experiments.
Screening of GABA Producing Isolates
Thin-Layer Chromatography:
GABA producing ability of the isolates and the positive control L. plantarum DM5 [28] were evaluated in bacterial supernatants following the method of Choi et al. [29] with some modifications of Kim and Kim [30]. The bacterial isolates were grown in MRS broth containing 2% monosodium glutamic acid (MSG) at 37 °C for 48 h. The cell-free supernatant (extracellular GABA) obtained by centrifugation (10,000 g, 4 C for 10 min) was filter-sterilized using a 0.2-µm syringe filter and stored for further analysis.
The collected bacterial cells (intracellular GABA) were washed three times with 0.9% NaCl and resuspended in 20 ml of phosphate buffer (pH 8.0). Then the cell suspension was treated in an ice bath with sonication (500 W, 20 min). The homogenate was centrifuged for 15 min at 8000 g at 4 °C, and the supernatant was collected for further analysis [31]. Four microlitres of extracellular GABA and intracellular GABA, along with 1 mg/ml of GABA standard (Sigma-Aldrich Co, St. Louis, MO, USA, cat No-03835) and MSG (Sigma Aldrich, Bengaluru, India) solution, was spotted on a silica TLC plate (Aluminium sheet silica gel 60 F254 Merck Germany). GABA separation by TLC was performed using a solvent mixture (1butanol:acetic acid:distilled water 3:2:1 v/v/v). When the solvent front had reached within about 1 cm of the top end of the adsorbent (after 20 to 30 min), the plate was taken out of the developing chamber, the location of the solvent front marked, and the solvent was evaporated from the plate. GABA spots were identified after spraying the plates with 2% (w/v) ninhydrin and exposing the plates to a heat source for a few minutes [12]. The conversion rate of MSG to GABA was quantified using ImageJ software and analysed with one-way ANOVA.
High-Performance Thin-Layer Chromatography:
Based on the TLC data, extracellular GABA (10 µl) of four isolates along with GABA standard, MSG, and culture of positive control were loaded with silica gel 60 F254 using a Hamilton syringe and run on a CAMAG Linomat 5 instrument. The mobile phase used was 1butanol:glacial acetic acid:water (3:2:1 v/v/v). The developed plate was viewed after spraying it with 2% ninhydrin in acetone and developing at 105 °C for 5 min. The scanned area of the samples was compared with the scanned area of the GABA standard. The GABA spots were viewed at 480 nm on HPTLC plates [32]. Quantification of the graph was performed using the area under curve analysis and analysed with one-way ANOVA.
Quantification of GABA by HPLC
GABA standard solution and extracellular GABA of the isolate (BS5) were derivatized with phenyl isothiocyanate (PITC) [33]. Aliquots of 1 ml GABA standard solution and culture supernatant were dried using a lyophilizer. About 100 μl of ethanol-water-triethylamine (2:2:1 v/v/v/) was added to the residue and was evaporated to dryness under vaccum. This residue was dissolved in 150 μl ethanol-water-triethylamine-PITC (7:1:1:1 v/v/v/v/) which was kept at room temperature for 20 min to form phenyl thiocarbamyl-GABA. Excess reagents were removed under vacuum. About 500 μl of mobile phase consisting of 80% solution A (1.4 mM sodium acetate, 0.1% triethylamine, and 6% acetonitrile) and 20% solution B (60% acetonitrile) was added to the dry residue, and the resultant solution was filtered using a 0.45-μm filter, and analysed with HPLC. The GABA analysis was performed using an HPLC system equipped with a gemini C18 column. The column was eluted at a flow rate of 1 mL/min for 50 min with a linear gradient of 0–100% solution B. A sample (20 μl) was injected and detected at a wavelength of 254 nm. A standard curve was created using known concentrations (250, 500, and 1000 ppm) of GABA standard and the quantity of GABA in the culture supernatant was measured from the standard curve and analysed with one-way ANOVA.
Morphological and Phenotypic Characterization
The selected isolate was grown overnight in MRS broth at 37 °C in a rotary shaker. Gram-staining was performed according to the method of Dussault [34]. FESEM was also carried out for morphological analysis. The genomic DNA of the isolate was extracted from the 12 h culture by the phenol-chloroform method [35]. The 16S rRNA gene of the selected organism was amplified by using the polymerase chain reaction (PCR) technique from a genomic DNA using the forward primer 5′-GAGTTTGATCCTGGCTCAG-3′ and the reverse primer 5′-ACGGCTACCTTGTTACGACTT-3′. PCR reaction was carried out by a TI thermocycler. The amplicon was obtained with a PCR cycling program of 96 °C for 5 min, 30 cycles 94 °C for 1 min, 48 °C for 1 min, and a final extension time of 72 °C for 2 min. The amplified product was visualized by electrophoresis separation in a 0.8% gel. The amplicon was eluted, purified, and sequenced commercially (Eurofins Genomics India Pvt. Ltd., Bengaluru). The partial sequence data of the 16S rRNA genes (956 bp) were deposited in the database of the National Center for Biotechnology Information (NCBI) with accession no MN394829. These sequence data were compared with the public database in Genbank using BLAST, closest known relatives were obtained and the phylogenetic tree was constructed based on the neighbour-joining method using MEGA version 7.0 [36].
Evaluation of High GABA Producing Isolate for Probiotic Attributes
Ent. faecium BS5 was selected based on quantification studies and subjected to a series of probiotic tests under in-vitro conditions to explore the probiotic potential of the isolate.
Tolerance to Low pH and Bile Salt
The selected GABA producing Ent. faecium BS5 was characterized to identify its major probiotic features, such as tolerance to low pH, and bile salt based on the method of Ahire et al. [37]. Ent. faecium BS5 was grown in MRS broth overnight at 37 °C and pelleted at 8000 g for 5 min at 4 °C. Cells were rinsed twice with sterile phosphate buffer saline (PBS, pH 7.3) and resuspended in 1 ml PBS. Ent. faecium BS5 was diluted in PBS (1:100) of varied pH (1, 2, 3, and 4) and then incubated at different time intervals (0, 1, 2, and 3 h) at 37 °C. The viability of the bacterial cells in terms of CFU ml−1 on the MRS agar plate was determined. After 3 h of incubation, the survival of the isolate in different pH was also reported in percentage.
Bile salt tolerance of Ent. faecium BS5 was determined by inoculating the bacterial isolate in MRS broth containing 2.5, 5.0, 7.5, and 10% of bile salt and incubated at 37 °C for 3 and 6 h. The growth medium with 0% bile salt was used as a control. The treated cells were then evaluated by an absorbance microplate reader by recording the absorbance at 595 nm. Data were subjected to two-way ANOVA for bile and time.
Hydrophobicity Assay
Cell surface hydrophobicity of Ent. faecium BS5 was evaluated based on the method of Thapa et al. [38]. Chloroform and ethyl acetate were used to detect the surface hydrophobicity of the isolates. The overnight grown cells were centrifuged, washed with PBS three times, and resuspended in 10-ml Ringer’s solution, and OD was measured as control (A0) at 600 nm. In the tested sample, the cell suspension was mixed with an equal volume of solvent by vortexing for 2 min and held at room temperature for 30 min. The aqueous phase was removed and absorbance measured at 600 nm (A1). The formula (1 − A1/A0) × 100 was used to measure the hydrophobicity of bacterial adhesion to the solvent.
Autoaggregation Assay
Autoaggregation assay of Ent. faecium BS5 was carried out with minor modifications to the method of Patel et al. [39]. Cells from the overnight grown culture were collected by centrifugation and washed three times with PBS (pH 7.3). The pellets were resuspended to obtain OD595 0.5, and 4 ml of the cell suspension was gently vortexed for 10 s and incubated for 2 h at 37 °C. The supernatant was removed after incubation, and absorbance was measured at 595 nm using a UV–Vis Spectrophotometer. Autoaggregation assay was expressed in percentage using the following formula: 1 − (At/A0) × 100, where At represents the cell suspension absorbance at time t = 2 h and A0 for absorbance at t = 0.
Hemolytic Activity
Hemolytic activity was evaluated by streaking Ent. faecium BS5 on MRS blood agar plates and incubating for 48 h at 37 °C. Partial hydrolysis and a greenish zone (α-hemolysis), a clear zone around the colonies (β-hemolysis), or no reaction (γ-hemolysis) were observed for a hemolytic reaction [40].
Antibiogram
Antibiotic susceptibility of Ent. faecium BS5 was determined using antibiotic diffusion discs. Ent. faecium BS5 was inoculated in MRS broth and incubated for 24 h at 37 °C. Sample culture (100 µl) was spread on nutrient agar plates, and a disc dispenser was used to apply antibiotic discs to plates. The plates were incubated at 37 °C and observed after 24 h of inoculation. The antibiotics (Himedia) used were tetracycline (30 μg), chloramphenicol (30 μg), ampicillin (10 μg), penicillin G (10 μg), streptomycin (300 μg), rifamycin (5 μg), kanamycin (30 μg), and erythromycin (15 μg). The diameter of the inhibition zone was measured (mm), and the antibiotic sensitivity was recorded based on their activity as different grades [41].
Antimicrobial Activity Assay
The antimicrobial activity of Ent. faecium BS5 was investigated against Escherichia coli, Bacillus subtilis, Pseudomonas aeruginosa, Staphylococcus aureus, and Candida albicans by the agar well diffusion method [42]. Isolate was grown overnight in MRS broth at 37 °C for 24 h, and the cell-free supernatant (CFC) pH was adjusted to 6.5 by adding 1 N NaOH. Briefly, aliquots of the supernatant in different volumes were loaded into wells in nutrient agar plates seeded with the indicator strain at a final concentration of 0.5% (v/v). The antibiotic disc was used as the positive control and water as the negative control. Then the plates were incubated at 37 °C for 24 h, and after incubation, the diameter of the inhibition zones around the well was measured.
Statistical Analysis
All experiments were conducted in triplicates with data expressed as mean ± standard deviation (SD). Results were tested for analysis of variance (ANOVA) using OriginPro 8.0 software to determine the statistical differences between the mean of the samples at the level of significance, p < 0.05.
Results and Discussion
Lactic Acid Bacteria from Dairy Products
The most interesting and practical group of bacteria for GABA production from dairy products are L. s paracasei, L. plantarum [43], and L. lactis [44], all of which possess special physiological activities and are generally regarded as safe. Therefore, using LAB as cell factories for GABA production offers a wide range of opportunities in the development of foods with potentially beneficial effects. In the present study, LAB was isolated from the dairy products and screened for GABA production and the highest GABA producing isolate was further assessed for its probiotic characteristics.
Screening of GABA Producing Isolates for Extracellular GABA Production Using TLC and HPTLC
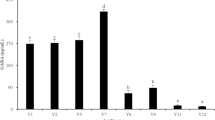
Screening of different microorganisms capable of producing GABA is important for the food industry because this type of GABA production offers natural GABA, a bioactive agent that modulates the health features and provides the consumer with new and attractive food products. The 64 isolates selected from MRS agar plates were screened for extracellular and intracellular GABA production with 2% MSG. Based on TLC spots, 9 isolates were screened for the final TLC run that produced the dark spot indicating extracellular GABA compared with intracellular GABA production. Earlier, Kim et al. [45], Ko et al. [46], Lee et al. [47], and Villegas et al. [48] have reported the highest GABA production when its culture supernatants were analysed by TLC. Figure 1 shows the TLC profile of the supernatants obtained in the present study. The isolates with a visible band of GABA were DM5 (positive control), BS5, BS1, BS2, BS3, and BS18. Their Rf values range from 0.51 to 0.53, similar to that (Rf = 0.52) of the GABA standard. Significant production of GABA was shown with BS1, BS2, BS3, and BS5 compared with other samples after spot quantification, and hence, these isolates were chosen for further experiments.
a TLC chromatogram showing GABA spots produced by the dairy isolates. The development solvent consists of 1butanol:acetic acid:water (3:2:1, v/v/v). The chromatogram was viewed after spraying the plates with a 2% ninhydrin solution and developing at 105 °C for 5 min. b Estimation of the amount of GABA produced from MSG by the isolates for screening and selection of the best isolate. c High-performance thin-layer chromatogram of GABA standard (1 mg/ml) and cell culture supernatants. The mobile phase used was 1butanol:glacial acetic acid:water (3:2:1 v/v/v). The developed plate was viewed after spraying with 2% ninhydrin in acetone and developing at 105 °C for 5 min. d Determination of quantity of GABA produced by the selected isolates using MSG as the substrate. e HPLC profile of GABA standard (1 mg/ml) and supernatant from BS5 in MRS medium. Mobile phase: 80% solution A (1.4 mM sodium acetate, 0.1% triethylamine, and 6% acetonitrile) and 20% solution B (60% acetonitrile). The column was eluted at a flow rate of 1 mL/min for 50 min and detected at a wavelength of 254 nm. Data are representative of three independent experiments. f Comparison of GABA concentration in terms of area under peak between GABA standard and the isolate BS5. The data represent the mean ± SD of three independent experiments and one-way ANOVA was performed with p < 0.05
In the previous research, Pradeep et al. [32] confirmed GABA from millets and legumes by HPTLC, and therefore, in our study, the culture supernatants of four isolates were subjected to HPTLC with the solvent system (1butanol:glacial acetic acid:water). The spectra of all the isolates and the standard are shown in Fig. 1. All the isolates showed a peak corresponding to the GABA standard, with an Rf value of 0.60. After analyzing the area under the curve, we found that the highest production of GABA was achieved with BS5 and there was no significant difference in GABA production with BS1, BS2, or BS3.
Quantification of GABA by HPLC
The GABA contents of the isolate BS5 were quantified using HPLC (Fig. 1). It was observed that the HPLC chromatogram of the culture supernatant of the presumptive GABA-producing isolate BS5 exhibits a peak that shows the same retention time (42 min) as that of GABA standard. The sample peak area was measured and compared with the calibration curve of the standard to quantify the GABA concentration of the isolate. In a similar study, Zmari et al. [49] confirmed GABA production in ethanolic extract (30 mg/ml) of germinated brown rice by HPLC whereby a retention time of 49 min was obtained. Tajabadi et al. [50] reported that L. plantarum Taj-apis 362 showed the highest GABA production (1.76 mM) as measured by HPLC.
General Phenotypic Characteristic of GABA Producing Bacteria
The highest GABA producing isolate BS5 was characterized based on morphological and biochemical analysis. The isolate BS5 formed creamy, circular, and opaque colonies on MRS agar plates and was seen as gram-positive cocci shape under the microscope. Further, the FESEM image of the isolate BS5 affirmed the cocci shaped morphology as shown in Fig. 2. Genetic identification by 16S rRNA sequencing of the isolate showed that BS5 belonged to the genus Ent. faecium. The partial sequences of Ent. faecium were submitted to the NCBI GenBank nucleotide sequence database, and the accession number MN394829 was obtained. The phylogenetic tree of the isolate BS5 (Accession no MN394829) with the most closely related enterococci was constructed using MEGA 7 software with neighbour-joining method (Fig. 2).
a Morphology of the GABA producing isolate, BS5 viewed under 60X magnification of the phase-contrast microscope. b Representative FESEM of the isolate BS5 showing the cocci shape of the bacterium. c Phylogenetic tree analysis of the 16S rRNA gene sequence of the isolate Ent. faecium BS5 (NCBI Accession no. MN394829) using the neighbour-joining method. The bacterial isolate was named based on its alignment to the homolog bacterial species. The bacterial isolate identified in this study has been represented in the box
Evaluation of the Highest GABA Producing Ent. Faecium BS5 for Probiotic Attributes
The prerequisite of a good probiotic is its ability to harbour and colonize the intestinal tract. Microorganisms are subjected to a variety of stresses during transit in the gastrointestinal tract, such as the action of enzymes in the oral cavity, the acidic environment of the stomach, and high bile concentration in the intestine to colonize, and thus, probiotics must survive these stresses and exert health benefits [51]. The isolates must be able to resist an acidic gastric environment with bile salt-resistance and epithelial cell adhesion. Before being used as a probiotics, their adherence to the appropriate surface and survival in the gastrointestinal tract has to be confirmed in vitro [52]. Furthermore, their safety and other functional features such as antibiotic resistance and antimicrobial activities should also be checked before they are commercially explored for human consumption. In this study, the highest GABA-producing isolate Ent. faecium BS5 was selected for further evaluation of probiotic attributes. All these experiments were performed under in vitro conditions, and predicting the designated isolate Ent. faecium BS5 may also mimic it`s in vivo activity when fed to the consumer.
Tolerance to Low pH and Bile Salt
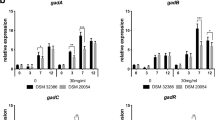
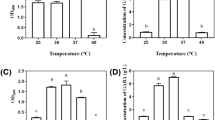
A probiotic must be tolerant to both bile and acid to benefit human health [53]. Stomach pH will rise to pH 3 if food is present, while in the stomach the pH of secreted HCl is 0.9 [54]. For entering into the small intestine through the stomach, probiotics need to survive at a pH value less than 3 [55]. Therefore, the ability of Ent. faecium BS5 to withstand acid stress at pH 1.0, 2.0, 3.0, and 4.0 was evaluated which showed the survival ability of Ent. faecium BS5 in the gastrointestinal tract. Ent. faecium BS5 showed the highest survivability of 76% and 78% in pH 3 and 4, respectively, after 3 h of incubation. The minimum survivability of 25% was observed in pH 2 and at pH 1; the isolate was not able to survive. Akalu et al. [56] have reported the survival rate (more than 80%) of Lactobacillus isolated from traditionally fermented Ethiopian beverage and food (Shamita and Kocho) at pH 2.5 and 3 for 3 and 6 h, respectively. Oh and Jung [57] reported that Pediococcus and Lactobacillus species isolated from Omegisool, a Korean traditional fermented millet alcoholic beverage can survive 3 h in pH 2 and pH 3.
Bile salt alters the structure of the cell membrane and is, therefore, harmful to living cells. Bile tolerance is a vital requirement for the survival and growth of LAB in the small intestine [58]. The viability percentage of cells with different concentrations of bile salt was carried out for 0, 3, and 6 h to determine the survival ability of Ent. faecium BS5 in the gastrointestinal tract. Two-way analysis of variance (ANOVA) revealed a significant difference in the survivability of Ent. faecium BS5 at 0, 3, and 6 h incubation at all concentrations of bile (Fig. 3). The survivability of Ent. faecium BS5 was significantly reduced at 10% bile compared with 2.5% bile. In conclusion, Ent. faecium BS5 was able to tolerate and survive in different concentrations of bile, in agreement with the earlier reports showing that the Lactobacilli and pediococcus possess a high tolerance to bile salt [59, 60] and the different rates of survival for L. plantarum strains indicate that survival of bacteria in the bile media is strain-dependent [61, 62].
a Ent. faecium BS5 showing bile salt tolerance grown in MRS broth supplemented with 0, 2.5, 5, 7.5, and 10% of bile. Growth was recorded by measuring the OD at 595 nm. b Cell surface hydrophobicity of Ent. faecium BS5 against ethyl acetate and chloroform. The data represent mean ± SD of three independent experiments. Two-way ANOVA for bile and one-way ANOVA for hydrophobicity were performed with p < 0.05
Cell Hydrophobicity Assay and Autoaggregation Assay
The adhesion property can provide information about the ability of probiotics to colonize and may modulate the immunological system of the host. One of the factors that contribute to the adhesion of bacterial cells to host tissues is cell hydrophobicity [63]. Many mechanisms about the colonization of the gastrointestinal tract involving microbial adhesions and surface hydrophobicity have been identified. In vitro analysis of microbial hydrophobicity which participates in microbial adhesion to ethyl acetate and chloroform was performed to investigate probiotic bacteria. The results revealed that GABA producing Ent. faecium BS5 is found to adhere to ethyl acetate at 45.09% and chloroform at 43.42% (Fig. 3). Cell hydrophobicity is involved in microbial-microbial interactions and may help isolate Ent. faecium BS5 to maintain its residence in the GI tract.
Autoaggregation is a significant bacterial feature in human and animal mucosa and bacteria with aggregation ability could adhere better onto the intestinal cells. Previous research has reported that some Lactobacillus [64, 65] and pathogens like Fusobacterium nucleatum [66] exhibit aggregation ability. Ent. faecium BS5 showed a noteworthy aggregation of 48%, and these findings are supported by previous data demonstrating a medium degree of aggregation of E. faecalis, and E. faecium [67]. However, aggregation varies among microbial species, and adhesive properties that lead to aggregation could potentially influence intestinal colonization.
Hemolytic Assay
One of the significant safety criteria often used to evaluate potential probiotic strains is the hemolytic activity. In this study, a partial hydrolysis and greening zone (α -hemolysis), the clear zone around the colonies (β-hemolysis), and no reaction (γ-hemolysis) have been observed for hemolytic activity. Ent. faecium BS5 had shown no clear zone or greenish zone around their colonies on the blood agar plate which illustrates the non-hemolytic activity, and these results were following the earlier studies in which lactobacilli strains were shown to be nonhemolytic [68].
Antibiogram
If the use of antibiotics in medicines and foods eliminates probiotics, probiotics will no longer function [69]. So, survival in the presence of antibiotics is essential for probiotic strain. So we have investigated the tolerance of E. faecium BS5 against eight antibiotics and the results are shown in Table 1. Ent. faecium BS5 found to be resistant to erythromycin, kanamycin, streptomycin, ampicillin, and penicillin and sensitive to tetracycline, chloramphenicol, and rifamycin. These findings concur with the report by Zhou et al. [70] who claimed that L. bulgaricus and S. thermophilus strains tend to be strongly resistant to kanamycin, streptomycin, and ampicillin. According to a review by Mathur and Singh [71], different strains exhibit different levels of antibiotic resistance. Hence, due to its antibiotic resistance, E. faecium BS5 can survive in the environment containing high levels of antibiotic concentrations.
Antimicrobial Activity
Another desirable property of the selection of a suitable starter culture is antimicrobial activity against potential pathogens and spoilage organisms to ensure the development of healthy fermented foods [72]. Hence, the selected Ent. faecium BS5 isolate was examined for its antimicrobial activity using the agar-well diffusion method. CFS of Ent. faecium BS5 inhibited the growth of E. coli, B. subtilis, P. aeruginosa, and S. aureus and Ent. faecium BS5 was inactive against C. albicans (Fig. 4). A similar result was reported by Bassyouni et al. [73] who showed that Lactobacillus species was able to inhibit E. coli, and Staphylococcus sp. Yuksekdag [74] found that LAB can inhibit the growth of pathogen and spoilage microorganisms through the production of antimicrobial substances such as organic acids, diacetyl, and hydrogen peroxide. Salminen et al. [75] also found that the ability to produce different antimicrobial compounds may be one of the important characteristics to prevent pathogen survival in the intestine and expression of a probiotic effect of a host.
a Antimicrobial activity of Ent. faecium BS5 against B. subtilis, E. coli, and S. aureus after 24 h of incubation at 37 °C. b Statistical analysis of zone of inhibition at different concentrations of the bacterial supernatant against test strains. The data represent mean ± SD of three independent experiments and one-way ANOVA was performed with p < 0.05
Conclusion
Identification of strains with biochemical characteristics and a mode of action that can mediate specific health effects in the host underpins the scientific selection of the next generation of probiotic strains. Production of GABA functional foods or enriched foods capable of providing GABA is a key objective of research and development in the food industry. Furthermore, the isolation of GABA-producing strains from various fermented foods is an important natural method for functional food design. Such isolates can represent next-generation probiotics with a specific mode of action based on the potential gut: brain axis modulation and GABA producing ability.
Based on the knowledge of the beneficial effects of GABA and as confirmed by many studies, this paper reports the high capacity of gamma-aminobutyric acid production of nine isolates from dairy products, and the selected isolate BS5 was identified as Ent. faecium through biochemical tests and 16S rRNA sequencing. Our results recommend that Ent. faecium BS5 could be a good candidate as a probiotic starter culture strain in the food industry. Although in vivo investigations are needed, these preliminary findings show that the Ent. faecium BS5 can substitute chemical GABA with natural GABA. Ent. faecium BS5 also represents a promising next-generation probiotic and a good starter culture for the production of GABA rich cultured dairy products that can be used as a functional food.
Data Availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Huang J, Mei L, Sheng Q, Yao S, Lin D (2007) Purification and characterization of glutamate decarboxylase of Lactobacillus brevis CGMCC 1306 isolated from fresh milk. Chin J Chem Eng 15(2):157–161. https://doi.org/10.1016/S1004-9541(07)60051-2
Hayakawa K, Kimura M, Yamori Y (2005) Role of the renal nerves in γ-aminobutyric acid-induced antihypertensive effect in spontaneously hypertensive rats. Eur J Pharmacol 524:120–125. https://doi.org/10.1016/j.ejphar.2005.09.020
Ueno H (2000) Enzymatic and structural aspects on glutamate decarboxylase. J Mol Catal B Enzym 10:67–79. https://doi.org/10.1016/S1381-1177(00)00114-4
Vaiva G, Thomas P, Ducrocq F et al (2004) Low posttrauma GABA plasma levels as a predictive factor in the development of acute posttraumatic stress disorder. Biol Psychiatry 55:250–254. https://doi.org/10.1016/j.biopsych.2003.08.009
Park KB, Oh SH (2007) Production of yogurt with enhanced levels of gamma-aminobutyric acid and valuable nutrients using lactic acid bacteria and germinated soybean extract. Bioresour Technol 98:1675–1679. https://doi.org/10.1016/j.biortech.2006.06.006
Soltani N, Qiu H, Aleksic M et al (2011) GABA exerts protective and regenerative effects on islet beta cells and reverses diabetes. Proc Natl Acad Sci U S A 108(28):11692–11697. https://doi.org/10.1073/pnas.1102715108
Pokusaeva K, Johnson C, Luk B et al (2016) GABA-producing Bifidobacterium dentium modulates visceral sensitivity in the intestine. Neurogastroenterol Motil 29: 1–14. https://doi.org/10.1111/nmo.12904
Auteri M, Zizzo MG, Serio R (2015) GABA and GABA receptors in the gastrointestinal tract: from motility to inflammation. Pharmacol Res 93:11–21. https://doi.org/10.1016/j.phrs.2014.12.001
Chen S, Tan B, Xia Y et al (2018) Effects of dietary gamma-aminobutyric acid supplementation on the intestinal functions in weaning piglets. Food Funct 10:366–378. https://doi.org/10.1039/c8fo02161a
Zhao Y, Wang J, Wang H et al (2020) Effects of GABA supplementation on intestinal SIgA secretion and gut microbiota in the healthy and ETEC-infected weanling piglets. Mediators Inflamm 2020:1–17. https://doi.org/10.1155/2020/7368483
Kim JY, Lee MY, Ji GE, Lee YS, Hwang KT (2009) Production of γ-aminobutyric acid in black raspberry juice during fermentation by Lactobacillus brevis GABA100. Int J Food Microbiol 130:12–16. https://doi.org/10.1016/j.ijfoodmicro.2008.12.028
Thwe SM, Kobayashi T, Luan T, Shirai T, Onodera M, Hamada-Sato N, Imada C (2011) Isolation, characterization, and utilization of γ-aminobutyric acid (GABA)-producing lactic acid bacteria from Myanmar fishery products fermented with boiled rice. Fish Sci 77:279–288. https://doi.org/10.1007/s12562-011-0328-9
Seo MJ, Lee JY, Nam YD et al (2013) Production of γ-aminobutyric acid by Lactobacillus brevis 340G isolated from kimchi and its application to skim milk. Food Eng Prog 17(4):418–423. https://doi.org/10.13050/foodengprog.2013.17.4.418
Binh TTT, Ju WT, Jung WJ, Park RD (2014) Optimization of γ-aminobutyric acid production in a newly isolated Lactobacillus brevis. Biotechnol Lett 36:93–98. https://doi.org/10.1007/s10529-013-1326-z
Lim HS, Cha IT, Roh SW, Shin HH, Seo MJ (2017) Enhanced production of gamma-aminobutyric acid by optimizing culture conditions of Lactobacillus brevis HYE1 isolated from kimchi, a Korean fermented food. J Microbiol Biotechnol 27(3):450–459. https://doi.org/10.4014/jmb.1610.10008
Huang J, Mei LH, Wu H, Lin DQ (2007) Biosynthesis of γ-aminobutyric acid (GABA) using immobilized whole cells of Lactobacillus brevis. World J Microbiol Biotechnol 23:865–871. https://doi.org/10.1007/s11274-006-9311-5
Komatsuzaki N, Shima J, Kawamoto S, Momose H, Kimura T (2005) Production of γ-aminobutyric acid by Lactobacillus paracaesai isolated from traditional fermented foods. Food Microbiol 22:497–504. https://doi.org/10.1016/j.fm.2005.01.002
Park SY, Lee JW, Lim SD (2014) The probiotic characteristics and GABA production of Lactobacillus plantarum K154 isolated from kimchi. Food Sci Biotechnol 23(6):1951–1957. https://doi.org/10.1007/s10068-014-0266-2
Sa HD, Park JY, Jeong SJ, Lee KW, Kim JH (2015) Characterization of glutamate decarboxylase (GAD) from Lactobacillus sakei A156 isolated from jeot-gal. J Microbiol Biotechnol 25(5):696–703. https://doi.org/10.4014/jmb.1412.12075
Lu XX, Xie C, Gu ZX (2009) Optimisation of fermentative parameters for GABA enrichment by Lactococcus lactis. Czech J Food Sci 27(6):433–442
Gran HM, Gadaga HT, Narvhus JA (2003) Utilization of various starter cultures in the production of amasi, a Zimbabwean naturally fermented raw milk product. Int J Food Microbiol 88:19–28. https://doi.org/10.1016/s0168-1605(03)00078-3
Mancini A, Carafa I, Franciosi E, Nardin T, Bottari B, Larcher R, Tuohy KM (2019) In vitro probiotic characterization of high GABA producing strain Lactobacillus brevis DSM 32386 isolated from traditional “wild” alpine cheese. Ann Microbiol 69:1435–1443. https://doi.org/10.1007/s13213-019-01527-x
Sanchart C, Rattanaporn O, Haltrich D, Phukpattaranont P, Maneerat S (2016) Technological and safety properties of newly isolated GABA-producing Lactobacillus futsaii strains. J Appl Microbiol 121(3):734–745. https://doi.org/10.1111/jam.13168
Foulquié Moreno MR, Sarantinopoulos P, Tsakalidou E, DeVuyst L (2006) The role and application of enterococci in food and health. Int J Food Microbiol 106:1–24. https://doi.org/10.1016/j.ijfoodmicro.2005.06.026
Kimoto-Nira H, Suzuki S, Suganuma H, Moriya N, Suzuki C (2015) Growth characteristics of Lactobacillus brevis KB290 in the presence of bile. Anaerobe 35:96–101. https://doi.org/10.1016/j.anaerobe.2015.08.001
Maassen CBM, Claassen E (2008) Strain-dependent effects of probiotic lactobacilli on EAE autoimmunity. Vaccine 26:2056–2057. https://doi.org/10.1016/j.vaccine.2008.02.035
Cani PD, Van Hul M (2015) Novel opportunities for next-generation probiotics targeting metabolic syndrome. Curr Opin Biotech 32:21–27. https://doi.org/10.1016/j.copbio.2014.10.006
Das D, Goyal A (2015) Antioxidant activity and γ-aminobutyric acid (GABA) producing ability of probiotic Lactobacillus plantarum DM5 isolated from Marcha of Sikkim. LWT - Food Sci Technol 61:263–268. https://doi.org/10.1016/j.lwt.2014.11.013
Choi SI, Lee JW, Park SM, Lee MY, JI GE, Park MS, Heo TR, (2006) Improvement of gamma-aminobutyric acid (GABA) production using cell entrapment of Lactobacillus brevis GABA 057. J Microbiol Biotechnol 16(4):562–568
Kim MJ, Kim KS (2012) Isolation and identification of γ-aminobutyric acid (GABA) producing lactic acid bacteria from kimchi. J Korean Soc Appl Biol Chem 55:777–785. https://doi.org/10.1007/s13765-012-2174-6
Yang SY, Lü FX, Lu ZX, Bie XM, Jiao Y, Sun LJ, Yu B (2008) Production of γ-aminobutyric acid by Streptococcus salivarius subsp. thermophilus Y2 under submerged fermentation. Amino Acids 34:473–478. https://doi.org/10.1007/s00726-007-0544-x
Pradeep SR, Malleshi NG, Guha M (2011) Germinated millets and legumes as a source of gamma-aminobutyric acid. World appl Sci 14:108–113
Rossetti V, Lombard A (1996) Determination of glutamate decarboxylase by high-performance liquid chromatography. J Chromatogr B 681: 63–67. https://doi.org/10.1016/0378-4347(96)88202-8
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70(4):484–485
Sambrook J, Russel DW (2001) Molecular cloning: A laboratory manual. Cold spring harbor laboratory press, New York
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5(2):150–163. https://doi.org/10.1093/bib/5.2.150
Ahire JJ, Patil KP, Chaudhari BL, Chincholkar SB (2011) A potential probiotic culture ST2 produces siderophore 2,3-dihydroxybenzoylserine under intestinal conditions. Food Chem 127:387–393. https://doi.org/10.1016/j.foodchem.2010.12.126
Thapa N, Pal J, Tamang JP (2004) Microbial diversity in ngari, hentak, and tungtap, fermented fish products of north-east India. World J Microbiol Biotechnol 20:599–607. https://doi.org/10.1023/B:WIBI.0000043171.91027.7e
Patel AK, Ahire JJ, Pawar SP, Chaudhari BL, Chincholkar SB (2009) Comparative accounts of probiotic characteristics of Bacillus spp. isolated from food wastes. Food Res Int 42:505–510. https://doi.org/10.1016/j.foodres.2009.01.013
Abedi J, Saatloo MV, Nejati V, Hobbenaghi R et al (2018) Selenium-enriched Saccharomyces cerevisiae reduces the progression of colorectal cancer. Biol Trace Elem Res 185:424–432. https://doi.org/10.1007/s12011-018-1270-9
Thankappan B, Ramesh D, Ramkumar S, Natarajaseenivasan K, Anbarasu K (2015) Characterization of Bacillus spp. from the gastrointestinal tract of labeo rohita—towards to identify novel probiotics against fish pathogens. Appl Biochem Biotechnol 175:340–353. https://doi.org/10.1007/s12010-014-1270-y
Cheikhyoussef A, Pogori N, Chen H et al (2009) Antimicrobial activity and partial characterization of bacteriocin-like inhibitory substances (BLIS) produced by Bifidobacterium infantis BCRC 14602. Food Control 20:553–559. https://doi.org/10.1016/j.foodcont.2008.08.003
Siragusa S, De Angelis M, Di Cagno R, Rizzello CG, Coda R, Gobbetti M (2007) Synthesis of γ -aminobutyric acid by lactic acid bacteria isolated from a variety of Italian cheeses. Appl Environ Microbiol 73(22):7283–7290. https://doi.org/10.1128/AEM.01064-07
Nomura M, Kimoto H, Someya Y, Suzuki I (1999) Novel characteristic for distinguishing Lactococcus lactis subsp. lactis from subsp. cremoris. Int J Syst Bacteriol 49:163–166. https://doi.org/10.1099/00207713-49-1-163
Kim SH, Shin BH, Kim YH, Nam SW, Jeon SJ (2007) Cloning and expression of a full-length glutamate decarboxylase gene from Lactobacillus brevis BH2. Biotechnol Bio Process Eng 12:707–712. https://doi.org/10.1007/BF02931089
Ko CY, Lin HTV, Tsai GJ (2013) Gamma-aminobutyric acid production in black soybean milk by Lactobacillus brevis FPA 3709 and the antidepressant effect of the fermented product on a forced swimming rat model. Process Biochem 48:559–568. https://doi.org/10.1016/j.procbio.2013.02.021
Lee BJ, Kim JS, Kang YM, Lim JH, Kim YM, Lee MS, Jeong MH, Ahn CB, Je JY (2010) Antioxidant activity and γ-aminobutyric acid (GABA) content in sea tangle fermented by Lactobacillus brevis BJ20 isolated from traditional fermented foods. Food Chem 122:271–276. https://doi.org/10.1016/j.foodchem.2010.02.071
Villegas JM, Brown L, Savoy de Giori G, Hebert EM (2016) Optimization of batch culture conditions for GABA production by Lactobacillus brevis CRL 1942, isolated from quinoa sourdough. Food Sci Technol 67:22–26. https://doi.org/10.1016/j.lwt.2015.11.027
Md Zamri ND, Imam MU, Abd Ghafar SA, Ismail M (2014) Antioxidative effects of germinated brown rice-derived extracts on H2O2-induced oxidative stress in HepG2 Cells. Evid based complement alternat medicine 2014:1–11. https://doi.org/10.1155/2014/371907
Tajabadi N, Ebrahimpour A, Baradaran A, Rahim RA, Mahyudin NA, Abdul Manap MY, Bakar FA, Saari N (2015) Optimization of γ-aminobutyric acid production by Lactobacillus plantarum Taj-Apis362 from honeybees. Molecules 20:6654–6669. https://doi.org/10.3390/molecules20046654
FAO/WHO (2006) Probiotics in food: health and nutritional properties and guidelines for evaluation. FAO Food Nutr Pap 85. https://www.fao.org/3/a-a0512e.pdf. Accessed 12 Sep 2019
Chiang SS, Pan TM (2012) Beneficial effects of Lactobacillus paracasei subsp. Paracasei NTU 101 and its fermented products. Appl Microbiol Biotechnol 93:903–916. https://doi.org/10.1007/s00253-011-3753-x
Ouwehand AC, Salminen S, Isolauri E (2002) Probiotics: an overview of beneficial effects. Anton Van Leeuw 82: 279–289. https://doi.org/10.1023/a:1020620607611
Erkkilä S, Petäjä E (2000) Screening of commercial meat starter cultures at low pH and in the presence of bile salts for potential probiotic use. Meat Sci 55:297–300. https://doi.org/10.1016/s0309-1740(99)00156-4
Mcdonald LC, Fleming HP, Hassan HM (1990) Acid tolerance of Leuconostoc mesenteroides and Lactobacillus plantarum. Appl Environ Microbiol 56(7):2120–2124
Akalu N, Assefa F, Dessalegn A (2017) In vitro evaluation of lactic acid bacteria isolated from traditional fermented Shamita and Kocho for their desirable characteristics as probiotics. Afr J Biotechnol 16(12):594–606. https://doi.org/10.5897/AJB2016.15307
Oh YJ, Jung DS (2015) Evaluation of probiotic properties of Lactobacillus and Pediococcus strains isolated from omegisool, a traditionally fermented millet alcoholic beverage in Korea. LWT—Food Sci Technol 63(1): 437–444. https://doi.org/10.1016/j.lwt.2015.03.005
Succi M, Tremonte P, Reale A, Sorrentino E, Grazia L, Pacifico S, Coppola R (2005) Bile salt and acid tolerance of Lactobacillus rhamnosus strains isolated from parmigiano reggiano cheese. FEMS Microbiol Lett 244:129–137. https://doi.org/10.1016/j.femsle.2005.01.037
Floros G, Hatzikamari M, Litopoulou-Tzanetaki E, Tzanetakis N (2012) Probiotic and technological properties of facultatively heterofermentative lactobacilli from Greek traditional cheeses. Food Biotechnol 26:85–105. https://doi.org/10.1080/08905436.2011.645941
Sukumar G, Ghosh AR (2010) Pediococcus spp.-a potential probiotic isolated from Khadi (an Indian fermented food) and identified by 16S rDNA sequence analysis. Afr J Food Sci 4(9):597–602
Lee H, Yoon H, Ji et al (2011) Functional properties of Lactobacillus strains isolated from kimchi. Int J Food Microbiol 145:155–161. https://doi.org/10.1016/j.ijfoodmicro.2010.12.003
Ramos CL, Thorsen L, Schwan RF, Jespersen L (2013) Strain-specific probiotics properties of Lactobacillus fermentum, Lactobacillus plantarum, and Lactobacillus brevis isolates from Brazilian food products. Food Microbiol 36:22–29. https://doi.org/10.1016/j.fm.2013.03.010
Ram C, Chander H (2003) Optimization of culture conditions of probiotic Bifidobacteria for maximal adhesion to hexadecane. World J Microbiol Biotechnol 19:407–410. https://doi.org/10.1023/a:1023946702949
Tuo Y, Yu H, Ai L, Wu Z, Guo B, Chen W (2013) Aggregation and adhesion properties of 22 Lactobacillus strains. J Dairy Sci 96:4252–4257. https://doi.org/10.3168/jds.2013-6547
De Souza BMS, Borgonovi TF, Casarotti SN et al (2018) Lactobacillus casei and Lactobacillus fermentum strains isolated from mozzarella cheese: Probiotic potential, safety, acidifying kinetic parameters and viability under gastrointestinal tract conditions. Probiotics & Antimicro Prot 11:382–396. https://doi.org/10.1007/s12602-018-9406-y
Merritt J, Niu G, Okinaga T, Qi F (2009) Autoaggregation response of Fusobacterium nucleatum. Appl Environ Microbiol 75(24):7725–7733. https://doi.org/10.1128/AEM.00916-09
Rajković J, Joković N (2015) Probiotic properties and safety assessment of lactic acid bacteria isolated from kajmak. Biol Nyssana 6(2):81–89
Vesterlund S, Vankerckhoven V, Saxelin M, Goossens H, Salminen S, Ouwehand AC (2007) Safety assessment of Lactobacillus strains: Presence of putative risk factors in faecal, blood and probiotic isolates. Int J Food Microbiol 116(3):325–331. https://doi.org/10.1016/j.ijfoodmicro.2007.02.002
Charteris WP, Kelly PM, Morelli L, Collins JK (2001) Gradient diffusion antibiotic susceptibility testing of potentially probiotic lactobacilli. J Food Prot 64(12): 2007–2014. https://doi.org/10.4315/0362-028x-64.12.2007
Zhou N, Zhang JX, Fan MT, Wang J, Guo G, Wei XY (2012) Antibiotic resistance of lactic acid bacteria isolated from Chinese yogurts. J Dairy Sci 95:4775–4783. https://doi.org/10.3168/jds.2011-5271
Mathur S, Singh R (2005) Antibiotic resistance in food lactic acid bacteria -a review. Int J Food Microbiol 105:281–295. https://doi.org/10.1016/j.ijfoodmicro.2005.03.008
Ratanaburee A, Kantachote D, Charernjiratrakul W, Sukhoom A (2013) Selection of γ-aminobutyric acid-producing lactic acid bacteria and their potential as probiotics for use as starter cultures in Thai fermented sausages (Nham). Int J Food Sci Tech 48:1371–1382. https://doi.org/10.1111/ijfs.12098
Bassyouni RH, Abdel-all WS, Abdel-all MGFS, Kamel Z (2012) Characterization of lactic acid bacteria isolated from dairy products in Egypt as a probiotic. Life Sci 9(4):2924–2933
Yuksekdag ZN, Aslim B (2010) Assessment of potential probiotic and starter properties of Pediococcus spp.isolated from turkish-type fermented sausages (sucuk). J Microbiol and Biotechnol 20:161–168 https://doi.org/10.4014/jmb.0904.04019
Salminen S, Wright AV, Morelli L et al (1998) Demonstration of safety of probiotics- a preview. Int J Food Microbiol 44:93–106. https://doi.org/10.1016/s0168-1605(98)00128-7
Funding
The second author received financial assistance from DST-WOS A(SR/WOS-A/LS-458/2018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research Involving Human and Animal Participants
The research does not contain any human or animal subjects for study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
BS, S., Thankappan, B., Mahendran, R. et al. Evaluation of GABA Production and Probiotic Activities of Enterococcus faecium BS5. Probiotics & Antimicro. Prot. 13, 993–1004 (2021). https://doi.org/10.1007/s12602-021-09759-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-021-09759-7