Abstract
Diverse pathogens, including Fusarium fujikuroi and Xanthomonas oryzae pv. oryzae (Xoo), cause significant yield losses in rice (Oryza sativa). The situation is expected to worsen due to rapid climate change. Thus, identifying novel genes conferring innate immunity against these pathogens is crucial for global food security. WRKY transcription factors are involved in various plant processes, including innate immunity. In rice, there are 125 OsWRKYs, with some functions reported. However, the roles of many OsWRKYs in rice immunity remain largely unknown. In this study, we investigate the role of OsWRKY9 in broad-spectrum disease resistance. OsWRKY9 transcripts increased in response to F. fujikuroi and Xoo. The promoter of OsWRKY9 was indirectly activated by OsWRKY65, which confers broad-spectrum resistance to F. fujikuroi and Xoo. Moreover, OsWRKY9-overexpressing transgenic plants exhibited enhanced resistance to both pathogens in a manner similar to transgenic plants overexpressing OsWRKY65. Additionally, OsWRKY9 modulated the expression of various defense-related genes regulated by OsWRKY65. These results indicate that the OsWRKY65-OsWRKY9 module enhances resistance to bakanae disease and bacterial blight.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa) is one of the main cereal crops that provides food nutrient globally. However, global rice yields can be seriously damaged by bakanae disease and bacterial blight (Son et al. 2024a). Fusarium fujikuroi is a primary agent of bakanae disease. Although F. fujikuroi is classified as a hemibiotrophic pathogen, it acts as a necrotrophic pathogen in susceptible rice varieties (Matic et al. 2016; Cheng et al. 2020). To increase plant susceptibility, F. fujikuroi promotes gibberellin (GA) biosynthesis and activates GA signaling (Matic et al. 2016). In contrast, the jasmonic acid (JA)-dependent defense response is critical for rice immunity against F. fujikuroi (Matic et al. 2016; Cheng et al. 2020). However, the positive or negative regulators controlling the rice-F. fujikuroi interaction are largely unknown. Xanthomonas oryzae pv. oryzae (Xoo) is a major agent causing bacterial blight disease. Although Xoo is mostly considered biotrophic, this bacterial pathogen is predominantly biotrophic hemibiotrophs (De Vleesschauwer et al. 2013). Xoo employs various effectors to induce host susceptibility (Deb et al. 2021). For example, transcription activator-like effectors (TALEs), including PthXo1, PthXo2, PthXo3, and AvrXa7, directly activate the promoters of SUGAR WILL EVENTUALLY BE EXPORTED TRANSPORTER 11 (OsSWEET11), OsSWEET13, and OsSWEET14, which are necessary for susceptibility to Xoo (Gupta et al. 2021). Given that mutagenesis of OsSWEET13 and OsSWEET14 led to enhanced resistance to various Xoo strains (Eom et al. 2019; Oliva et al. 2019), it has been suggested that transcription factors and signaling pathways regulating the expression of OsSWEETs in response to Xoo are crucial for bacterial blight resistance.
WRKY transcription factors play significant roles in plant immunity (Javed and Gao 2023). Transcriptome analyses suggest that various OsWRKYs are involved in rice immunity against F. fujikuroi and/or Xoo (Ji et al. 2016; Matic et al. 2016; Choi et al. 2017; Cheng et al. 2020). Indeed, OsWRKY114 enhances broad-spectrum disease resistance (BSDR) to F. fujikuroi and Xoo by regulating the expression of genes involved in defense responses and phytohormone signaling (Son et al. 2022; Song et al. 2023). Moreover, recent studies showed that OsWRKY65 induces BSDR to these pathogens by upregulating immune responses and downregulating susceptibility (S) genes, including OsSWEETs (Son et al. 2024b). OsWRKYs can regulate each other, and OsWRKY transcriptional regulatory cascades play key roles in disease responses. For instance, the OsWRKY45-OsWRKY13-OsWRKY42 module enhances defense responses to the fungal pathogen Magnaporthe oryzae (Cheng et al. 2015), and the OsWRKY80-OsWRKY4 module is involved in resistance to the fungal pathogen Rhizoctonia solani (Peng et al. 2016). Furthermore, transcriptional regulatory cascades comprising various OsWRKYs (e.g., OsWRKY6 and OsWRKY10) activate defense signaling against Xoo (Choi et al. 2015, 2020; Im et al. 2022). However, OsWRKY regulatory cascades are complex and remain largely unexplored.
Our previous study showed that OsWRKY65 induces the expression of various OsWRKYs, including OsWRKY9, OsWRKY13, and OsWRKY30 (Son et al. 2024b). OsWRKY13 and OsWRKY30 function as positive regulators of defense responses to Xoo by transcriptionally regulating various target genes (Qiu et al. 2008; Jalmi and Sinha 2016; Wang et al. 2022). However, the biological role of OsWRKY9 remains unknown. In this study, we explored whether OsWRKY9 confers innate immunity against F. fujikuroi and Xoo and its involvement in the OsWRKY transcriptional regulatory cascade.
Materials and Methods
Plant Material and Growth Conditions
The japonica rice variety Ilmi was utilized in this study. Kernels were surface-sterilized with 70% ethanol and 2% sodium hypochlorite, germinated, and seedlings were grown in soil or Murashige and Skoog (MS) medium under a 16-h light/8-h dark cycle at 28 ℃.
Generation of the Transgenic Plants
OsWRKY65-overexpressing plants (OsWRKY65OX) were previously generated and confirmed (Son et al. 2024b). OsWRKY9-overexpressing (OsWRKY9OX) and OsWRKY30-overexpressing (OsWRKY30OX) plants were generated previously described (Son et al. 2020). Briefly, the full-length cDNAs of OsWRKY9 (LOC_Os01g18584) and OsWRKY30 (LOC_Os08g38990) were amplified using specific primers (Table S1) and cloned into the pEarleygate104 vector using the Gateway system (Invitrogen, USA). The constructs were transformed into Agrobacterium tumefaciens strain LBA4404, which was then used for rice transformation. Transgenic plants were regenerated from calli and selected until T3 homozygous plants were obtained.
Analysis of Gene Expression Pattern
Total RNA was extracted from the leaves of wild-type and transgenic plants using TRIzol Reagent (QIAGEN, Germany). 3 µg of total RNA was reverse-transcribed into cDNA using SuperScript III reverse transcriptase (Invitrogen, USA) following the manufacturer’s protocol. Quantitative reverse-transcription polymerase chain reaction (RT-qPCR) was performed using specific primers (Table S1) as previously described (Son et al. 2024b). Gene expression levels were calculated using the comparative Ct method, with OsActin as the reference gene.
Subcellular Localization of OsWRKY9
Protoplasts were transiently transformed as previously described (Son et al. 2023). To determine the subcellular localization of OsWRKY9, the pEarleygate104/OsWRKY9 construct was co-transfected with a nuclear localization signal-red fluorescent protein (NLS-RFP) construct into protoplasts. After 10 h of incubation, fluorescence signals were observed using a Leica TCS SP8 confocal laser scanning microscope (Leica Microsystems, Germany). Yellow fluorescent protein signals were detected with excitation at 514 nm and emission at 530–570 nm, while RFP signals were detected with excitation at 543 nm and emission at 580–620 nm.
Promoter Activity Assays in Rice Protoplasts
For promoter assays, the promoter regions of OsWRKY9 were cloned into the DJ467 vector and co-introduced into protoplasts with the effector plasmid pEarleyGate104/OsWRKY65 and the reporter plasmids 35S:Renila luciferase (RLUC). Luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega, USA), and promoter activity was determined based on the ratio of firefly luciferase (FLUC) to RLUC activity.
Bakanae and Bacterial Blight Disease Assays
Bakanae and bacterial blight disease assays were conducted as previously described (Son et al. 2024a, 2024b). Bakanae disease indices ranged from 0 to 3, where 0 represented no symptoms, 1 indicated blight symptoms on leaf tips, 2 indicated wilted shoots and pale green leaf color, and 3 indicated complete leaf drying. Severity of bakanae disease (disease index 2 and 3) was used to assess rice susceptibility to F. fujikuroi in this study. Bacterial blight disease incidence (%) was calculated as follows: (average lesion length/leaf length) × 100.
Results
OsWRKY9 Localizes to the Nucleus and Responds to F. fujikuroi and Xoo
The cDNA of the WRKY Group IIb OsWRKY9 from the rice cultivar Ilmi comprises 1812 bp, encoding 603 amino acids with a WRKY domain containing the WRKYGQK motif and a C2H2 zinc finger motif (Fig. 1A). Subcellular localization was determined by transfecting pEarleygate104/OsWRKY9 constructs expressing YFP-OsWRKY9 into rice protoplasts and observing fluorescence signals. The results demonstrated specific nuclear targeting of OsWRKY9 (Fig. 1B).
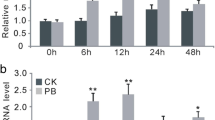
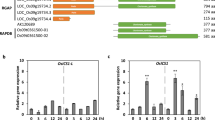
Subcellular localization and gene expression analysis of OsWRKY9. A Amino acid sequence and structural domains of OsWRKY9. The WRKY domain indicated by a solid line, the core WRKY motif by a dotted line, and the conserved zinc finger sequence by a triangle (▲). B Subcellular localization of OsWRKY9 in rice protoplasts. The N-terminal YFP conjugated OsWRKY9 was expressed in protoplasts with NLS-RFP and observed using confocal microscope. Scale bar: 8 μm. C, D Temporal expression analysis of OsWRKY9. Wild-type plants were inoculated with F. fujikuroi (C) and Xoo (D) for the indicated time. DAI, days after inoculation; HAI, hours after inoculation. Relative OsWRKY9 transcription levels were quantified using RT-qPCR, with OsActin serving as the reference gene. Data are presented as means ± SD, with asterisks indicating values significantly different from the controls (** P < 0.01)
To determine whether OsWRKY9 is involved in innate immunity against F. fujikuroi and Xoo, we assessed the abundance of OsWRKY9 transcripts following inoculation with each pathogen. Four-day-old wild-type seedlings were inoculated with F. fujikuroi CF283, and total RNA was extracted from shoots. The results revealed a significant upregulation of OsWRKY9 expression in response to F. fujikuroi infection (Fig. 1C). Additionally, 6-week-old wild-type plants were inoculated with the compatible Xoo strain KACC10859 using the clip method, and leaves were harvested. RT-qPCR analysis showed an increase in OsWRKY9 expression levels in response to Xoo (Fig. 1D). These findings suggest that OsWRKY9 likely plays a role in rice-pathogen interactions.
OsWRKY9 is Indirectly Induced by OsWRKY65 through Transcriptional Regulatory Mechanisms
Previous transcriptomic analysis indicated that OsWRKY9 transcriptional levels are elevated in OsWRKY65OX, similar to the upregulation observed for OsWRKY13 and OsWRKY30 (Son et al. 2024b). RT-qPCR analysis confirmed increased expression of OsWRKY9, OsWRKY13, and OsWRKY30 in OsWRKY65OX lines (Fig. 2A). Since OsWRKY13 and OsWRKY30 are known to play crucial roles in rice immunity against Xoo (Qiu et al. 2008; Jalmi and Sinha 2016; Wang et al. 2022), these findings suggest that OsWRKY65-induced expression of OsWRKY13 and OsWRKY30 contributes to bacterial blight resistance. Furthermore, we hypothesized that OsWRKY9 might also function as a regulator in this transcriptional regulatory cascade. To explore the correlation between OsWRKY9 and OsWRKY65, a promoter activity assay was conducted. Analysis of the 1-kb region upstream of the OsWRKY9 translation initiation site revealed the presence of four W-box motifs within the first 500 bp (Fig. 2B). Interestingly, OsWRKY65 induced promoter activity in the 1-kb upstream region of OsWRKY9, whereas it did not affect the 500 bp upstream region (Fig. 2B). Moreover, the promoter activity assay showed that the promoter region from -1000 to -500 of OsWRKY9, which lacks the W-box motif, is activated by OsWRKY65 (Fig. 2B). These results indicate that OsWRKY65 indirectly regulates the promoter activity of OsWRKY9.
Regulation of OsWRKY9 by OsWRKY65. A Expression levels of OsWRKY genes in OsWRKY65OX. Relative transcription levels of OsWRKY9, OsWRKY13, and OsWRKY30 in 2-week-old OsWRKY65OX and wild-type plants were quantified using RT-qPCR, with OsActin as the reference gene. Data are presented as means ± SD, with asterisks indicating values significantly different from the controls (** P < 0.01). B Promoter activity assays of truncated OsWRKY9 promoter regions. Schematic positions and directions of the W-box motifs located within 1 kb upstream of the translation initiation site in OsWRKY9 are shown. FLUC-conjugated truncated OsWRKY9 promoters were co-expressed with OsWRKY65 in rice protoplasts, and relative FLUC activity was measured. RLUC activity from 35S served as the control. Data are presented as means ± SD, with asterisks indicating values significantly different from the controls (** P < 0.01)
OsWRKY9 Confers Resistance to Both F. fujikuroi and Xoo Pathogens in Rice
To investigate the function of OsWRKY9 in rice immunity, transgenic rice cv. Ilmi lines overexpressing OsWRKY9 and OsWRKY30 were generated. Among the generated lines, two lines with strong expression were selected for OsWRKY9OX and OsWRKY30OX, confirmed by RT-qPCR in 2-week-old seedlings (Fig. 3A). To assess whether OsWRKY9 is involved in OsWRKY65-mediated immune responses, OsWRKY9OX, OsWRKY30OX, and OsWRKY65OX lines were infected with F. fujikuroi race CF283. Disease severity index scores ranged from 0 to 3, with a reduction in the sum of severe disease indices 2 and 3 observed in OsWRKY9OX and OsWRKY65OX compared to wild-type plants, whereas OsWRKY30OX showed symptoms similar to those of wild-type plants (Fig. 3B).
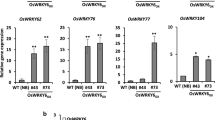
Disease phenotypes of OsWRKY9- and OsWRKY30-overexpressing plants. A Generation and verification of OsWRKY9OX and OsWRKY30OX. Relative expression levels of OsWRKY9 and OsWRKY30 were quantified using RT-qPCR, with OsActin as the reference gene. Data are presented as means ± SD, with asterisks indicating values significantly different from the controls (** P < 0.01). B Bakanae disease indices of the OsWRKYOX lines. Four-day-old OsWRKY9OX, OsWRKY30OX, OsWRKY65OX, and wild-type plants were inoculated with F. fujikuroi CF283 (n = 10). The disease index was evaluated at 6 days post-inoculation. Disease indices were categorized as follows: 0, no symptoms; 1, blight symptoms on the leaf tip; 2, wilted shoots and partial leaf drying; 3, entire leaf drying. The severity of bakanae disease (disease indices 2 and 3) was used to determine rice susceptibility to F. fujikuroi. Data are presented as means ± SD, with different letters indicating significant differences according to ANOVA (P < 0.05). C Bacterial blight disease resistance of the OsWRKYOX lines. Six-week-old OsWRKY9OX, OsWRKY30OX, OsWRKY65OX, and wild-type plants were inoculated with Xoo strain KACC10859 (n = 10). The incidence of bacterial blight disease (%) was determined at 14 days post-inoculation, calculated as (average lesion length/leaf length) × 100. Data are presented as means ± SD, with different letters indicating significant differences according to ANOVA (P < 0.05)
Furthermore, bacterial blight resistance assays revealed that OsWRKY9OX, OsWRKY30OX, and OsWRKY65OX lines exhibited significantly increased resistance to the compatible Xoo strain KACC10859 compared to wild-type plants (Fig. 3C). Overall, these findings collectively demonstrate that OsWRKY9 confers BSDR to F. fujikuroi and Xoo, akin to the resistance conferred by OsWRKY65.
OsWRKY9 Modulates the Expression of Genes Involved in the OsWRKY65 Transcriptional Regulatory Cascade
Transcriptome analysis and RT-qPCR revealed that the expression levels of key susceptibility-related genes to F. fujikuroi (e.g., GA 3-OXIDASE 2 [OsGA3OX2], OsGA20OX2, and JASMONATE ZIM-DOMAIN 10 [OsJAZ10]) and Xoo (e.g., OsSWEET13) are downregulated in OsWRKY65OX, whereas the expressions of defense-related genes, including 12-OXOPHYTODIENOIC ACID REDUCTASE 1 (OsOPR1) and PATHOGENESIS-RELATED GENE 10A (OsPR10a) are upregulated (Son et al. 2024b). Therefore, to further elucidate the function of OsWRKY9 and its relationship with OsWRKY65, we examined the expression levels of these genes in OsWRKY9OX plants. Transcription levels of GA biosynthesis-related genes, such as OsGA3OX2 and OsGA20OX2, along with the JA signaling repressor OsJAZ10, were down-regulated in 2-week-old OsWRKY9OX plants compared to wild-type plants, whereas transcripts of the JA biosynthesis-related gene OsOPR1 were up-regulated (Fig. 4A). Additionally, overexpression of OsWRKY9 decreased the expression of OsSWEET13, while increasing transcripts of OsPR10a (Fig. 4A). These findings suggest that OsWRKY9 is involved in OsWRKY65-mediated BSDR to both F. fujikuroi and Xoo. Moreover, to determine whether OsWRKY9 regulates the expression of OsWRKY65, we analyzed OsWRKY65 transcripts in 2-week-old OsWRKY9OX and wild-type plants. The result showed that the expression level of OsWRKY65 is not regulated by OsWRKY9 (Fig. 4B).
Expression levels of genes regulated by OsWRKY65 in OsWRKY9-overexpressing plants. A, B Expression levels of susceptibility and defense-related genes in OsWRKY9-overexpressing plants. The relative transcription levels of OsGA3OX2, OsGA20OX2, OsOPR1, OsJAZ10, OsSWEET13, and OsPR10a (A) and OsWRKY65 (B) were quantified using RT-qPCR in 2-week-old OsWRKY9OX and wild-type plants, with OsActin as the reference gene. Data are presented as means ± SD, with asterisks indicating values significantly different from the controls (** P < 0.01). C Conceptional model of the OsWRKY transcriptional regulatory cascade. OsWRKY65-induced expression of OsWRKY9 and OsWRKY30 enhances resistance to F. fujikuroi and/or Xoo
Discussion
Numerous studies have demonstrated that various WRKY transcription factors play critical roles in defense response-signaling pathways related to pathogen attack (Wani et al. 2021; Javed and Gao 2023). Although there are 125 OsWRKYs in the rice genome, many remain unexplored, presenting vast opportunities for future research. Previous RNA-sequencing result revealed that OsWRKY9 expression is increased in OsWRKY65OX, which exhibits enhanced resistance to F. fujikuroi and Xoo (Son et al. 2024b). Therefore, here, we focused on the role of OsWRKY9 in association with these pathogens.
OsWRKY9 belongs to WRKY group II and is specifically localized in the nucleus (Fig. 1A, B). In rice, eight OsWRKYs, including OsWRKY1, OsWRKY5, OsWRKY9, OsWRKY27, OsWRKY32, OsWRKY43, OsWRKY73, and OsWRKY97, belong to OsWRKYIIb (Yang et al. 2009). Among these, the expressions of OsWRKY1, OsWRKY9, and OsWRKY43 are induced by both compatible and incompatible Xoo strains (Choi et al. 2017), suggesting their involvement in rice immunity. In this study, we showed that the transcription levels of OsWRKY9 are significantly upregulated in response to not only Xoo but also F. fujikuroi (Fig. 1C, D). The initial interaction between rice and Xoo at the infection site determines the infection outcome, triggering rapid changes in gene expression even within minutes of Xoo inoculation (Kim et al. 2021). However, F. fujikuroi colonization progresses more slowly, with the earliest significant differential responses observed 7 days post-inoculation (Ji et al. 2016; Matic et al. 2016; Cheng et al. 2020). Indeed, the expression of OsWRKY9 was induced within 6 h following Xoo inoculation (Fig. 1D), whereas it was induced after 8 days in response to F. fujikuroi (Fig. 1C). These results suggest that OsWRKY9 plays a crucial role in the early defense response to both F. fujikuroi and Xoo. Additionally, we determined that the expression of OsWRKY9 is induced by OsWRKY65, and the 501–1,000 bp upstream promoter region excluding the W-box motif is necessary for its activation by OsWRKY65 (Fig. 2A, B). This suggests that OsWRKY65 indirectly activates the OsWRKY9 promoter through a non-WRKY transcription factor. Identifying this unknown transcription factor is essential for understanding the detailed regulatory mechanism related to OsWRKY9 and OsWRKY65.
To investigate whether OsWRKY9 is involved in the OsWRKY65 transcriptional cascade and plant immunity, we generated transgenic plants expressing OsWRKY9 and OsWRKY30, respectively, which are higher in OsWRKY65OX, and analyzed their resistance to F. fujikuroi and Xoo. OsWRKY9OX and OsWRKY65OX exhibited enhanced disease resistance to both pathogens, while OsWRKY30OX showed enhanced resistance only to Xoo (Fig. 3B, C). Previous studies have demonstrated that OsWRKY30 plays a crucial role in BSDR to M. oryzae, R. solani, Xanthomonas oryzae pv. oryzicola (Xoc), and Xoo (Peng et al. 2012; Han et al. 2013; Jalmi and Sinha 2016; Wang et al. 2022). Notably, the CCCH zinc finger transcription factor LEAF AND TILLER ANGLE INCREASED CONTROLLER (OsLIC) directly represses the expression of OsWRKY30, resulting in reduced resistance to Xoc and Xoo (Wang et al. 2022). However, MITOGEN-ACTIVATED PROTEIN KINASE 6 (OsMAPK6) negatively regulates OsLIC, thereby inducing OsWRKY30. Additionally, OsWRKY30 is implicated in the MAPK KINASE 3-OsMAPK6 signaling conferring resistance to Xoo (Jalmi and Sinha 2016). Therefore, further studies are needed to determine whether OsWRKY65 and/or OsWRKY9 are associated with disease resistance to other pathogens, such as M. oryzae, R. solani, and Xoc, as well as with the MAPK signaling pathways. The induction of OsSWEET genes, including OsSWEET13, by various TALEs is crucial for susceptibility to Xoo (Xu et al. 2019). The expression level of OsSWEET13 is significantly repressed in OsWRKY9OX (Fig. 4A), suggesting that Xoo must counteract the OsWRKY9-mediated suppression of OsSWEET13 to induce its expression. PthXo2 and PthXo2-like TALEs are major TALEs known to induce the expression of OsSWEET13 (Liu et al. 2024). Therefore, it is possible that the competition between the OsWRKY65-OsWRKY9 module and PthXo2 and PthXo2-like TALEs at the OsSWEET13 promoter may determine susceptibility to Xoo. This hypothesis also warrants further investigation in future study.
In this study, we demonstrated that OsWRKY9 contributes to BSDR to F. fujikuroi and Xoo and proposed a transcriptional regulatory cascade conferring rice immunity (Fig. 4C). Our findings provide information on novel signaling pathways controlling immune response, potentially helping to breed rice varieties with enhanced disease resistance.
Data Availability
The data and materials shown in this article are available from the corresponding author SRP upon a reasonable request.
Code Availability
N/A.
References
Cheng H, Liu H, Deng Y, Xiao J, Li X, Wang S (2015) The WRKY45-2 WRKY13 WRKY42 transcriptional regulatory cascade is required for rice resistance to fungal pathogen. Plant Physiol 167(3):1087–1099. https://doi.org/10.1104/pp.114.256016
Cheng AP, Chen SY, Lai MH, Wu DH, Lin SS, Chen CY, Chung CL (2020) Transcriptome analysis of early defenses in rice against Fusarium fujikuroi. Rice 13(1):65. https://doi.org/10.1186/s12284-020-00426-z
Choi C, Hwang SH, Fang IR, Kwon SI, Park SR, Ahn I, Kim JB, Hwang DJ (2015) Molecular characterization of Oryza sativa WRKY6, which binds to W-box-like element 1 of the Oryza sativa pathogenesis-related (PR) 10a promoter and confers reduced susceptibility to pathogens. New Phytol 208(3):846–859. https://doi.org/10.1111/nph.13516
Choi NY, Lee E, Lee SG, Choi CH, Park SR, Ahn I, Bae SC, Hwang CH, Hwang DJ (2017) Genome-wide expression profiling of OsWRKY superfamily genes during infection with Xanthomonas oryzae pv. oryzae using real-time PCR. Front Plant Sci 8:1628. https://doi.org/10.3389/fpls.2017.01628
Choi N, Im JH, Lee E, Lee J, Choi C, Park SR, Hwang DJ (2020) WRKY10 transcriptional regulatory cascades in rice are involved in basal defense and Xa1-mediated resistance. J Exp Bot 71(12):3735–3748. https://doi.org/10.1093/jxb/eraa135
De Vleesschauwer D, Gheysen G, Hofte M (2013) Hormone defense networking in rice: tales from a different world. Trends Plant Sci 18(10):555–565. https://doi.org/10.1016/j.tplants.2013.07.002
Deb S, Madhavan VN, Gokulan C, Patel HK, Sonti RV (2021) Arms and ammunitions: effectors at the interface of rice and it’s pathogens and pests. Rice 14(1):94
Eom JS, Luo D, Atienza-Grande G, Yang J, Ji C, Thi Luu V, Huguet-Tapia JC, Char SN, Liu B, Nguyen H, Schmidt SM, Szurek B, Vera Cruz C, White FF, Oliva R, Yang B, Frommer WB (2019) Diagnostic kit for rice blight resistance. Nat Biotechnol 37(11):1372–1379. https://doi.org/10.1038/s41587-019-0268-y
Gupta PK, Balyan HS, Gautam T (2021) SWEET genes and TAL effectors for disease resistance in plants: present status and future prospects. Mol Plant Pathol 22(8):1014–1026. https://doi.org/10.1111/mpp.13075
Han M, Ryu H-S, Kim C-Y, Park D-S, Ahn Y-K, Jeon J-S (2013) OsWRKY30 is a transcription activator that enhances rice resistance to the Xanthomonas oryzae pathovar oryzae. J Plant Biol 56:258–265. https://doi.org/10.1007/s12374-013-0160-0
Im JH, Choi C, Park SR, Hwang DJ (2022) The OsWRKY6 transcriptional cascade functions in basal defense and Xa1-mediated defense of rice against Xanthomonas oryzae pv. oryzae. Planta 255(2):47. https://doi.org/10.1007/s00425-022-03830-5
Jalmi SK, Sinha AK (2016) Functional involvement of a mitogen activated protein kinase module, OsMKK3-OsMPK7-OsWRK30 in mediating resistance against Xanthomonas oryzae in rice. Sci Rep 6:37974. https://doi.org/10.1038/srep37974
Javed T, Gao SJ (2023) WRKY transcription factors in plant defense. Trends Genet 39(10):787–801. https://doi.org/10.1016/j.tig.2023.07.001
Ji Z, Zeng Y, Liang Y, Qian Q, Yang C (2016) Transcriptomic dissection of the rice–Fusarium fujikuroi interaction by RNA-Seq. Euphytica 211(1):123–137
Kim S, Jang WE, Park J, Kim MS, Kim JG, Kang LW (2021) Combined analysis of the time-resolved transcriptome and proteome of plant pathogen Xanthomonas oryzae pv. oryzae. Front Microbiol 12:664857. https://doi.org/10.3389/fmicb.2021.664857
Liu L, Li Y, Wang Q, Xu X, Yan J, Wang Y, Wang Y, Shah SMA, Peng Y, Zhu Z, Xu Z, Chen G (2024) Constructed rice tracers identify the major virulent transcription activator-like effectors of the bacterial leaf blight pathogen. Rice 17(1):30. https://doi.org/10.1186/s12284-024-00704-0
Matic S, Bagnaresi P, Biselli C, Orru L, Amaral Carneiro G, Siciliano I, Vale G, Gullino ML, Spadaro D (2016) Comparative transcriptome profiling of resistant and susceptible rice genotypes in response to the seedborne pathogen Fusarium fujikuroi. BMC Genomics 17(1):608. https://doi.org/10.1186/s12864-016-2925-6
Oliva R, Ji C, Atienza-Grande G, Huguet-Tapia JC, Perez-Quintero A, Li T, Eom JS, Li C, Nguyen H, Liu B, Auguy F, Sciallano C, Luu VT, Dossa GS, Cunnac S, Schmidt SM, Slamet-Loedin IH, Vera Cruz C, Szurek B, Frommer WB, White FF, Yang B (2019) Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotechnol 37(11):1344–1350. https://doi.org/10.1038/s41587-019-0267-z
Peng X, Hu Y, Tang X, Zhou P, Deng X, Wang H, Guo Z (2012) Constitutive expression of rice WRKY30 gene increases the endogenous jasmonic acid accumulation, PR gene expression and resistance to fungal pathogens in rice. Planta 236(5):1485–1498. https://doi.org/10.1007/s00425-012-1698-7
Peng X, Wang H, Jang JC, Xiao T, He H, Jiang D, Tang X (2016) OsWRKY80-OsWRKY4 module as a positive regulatory circuit in rice resistance against Rhizoctonia solani. Rice 9(1):63. https://doi.org/10.1186/s12284-016-0137-y
Qiu D, Xiao J, Xie W, Liu H, Li X, Xiong L, Wang S (2008) Rice gene network inferred from expression profiling of plants overexpressing OsWRKY13, a positive regulator of disease resistance. Mol Plant 1(3):538–551. https://doi.org/10.1093/mp/ssn012
Son S, An HK, Seol YJ, Park SR, Im JH (2020) Rice transcription factor WRKY114 directly regulates the expression of OsPR1a and Chitinase to enhance resistance against Xanthomonas oryzae pv. oryzae. Biochem Biophys Res Commun 533(4):1262–1268. https://doi.org/10.1016/j.bbrc.2020.09.141
Son S, Im JH, Song G, Nam S, Park SR (2022) OsWRKY114 inhibits ABA-induced susceptibility to Xanthomonas oryzae pv. oryzae in rice. Int J Mol Sci 23(15):8825. https://doi.org/10.3390/ijms23158825
Son S, Im JH, Ko JH, Lee Y, Lee S, Han KH (2023) SNF1-related protein kinase 1 represses Arabidopsis growth through post-translational modification of E2Fa in response to energy stress. New Phytol 237(3):823–839. https://doi.org/10.1111/nph.18597
Son S, Song G, Nam S, Lee G, Im J, Lee KS, Park YJ, Suh EJ, Park SR (2024a) CRISPR/Cas9-mediated mutagenesis of rice NAC transcription factor genes results in altered innate immunity. Plant Physiol 195(2):1138–1142. https://doi.org/10.1093/plphys/kiae084
Son S, Song G, Nam S, Lee J, Hwang D-J, Suh E-J, Park SR (2024b) OsWRKY65 enhances immunity against fungal and bacterial pathogens in rice. Crop J 12(2):470–481. https://doi.org/10.1016/j.cj.2024.01.007
Song G, Son S, Nam S, Suh E-J, Lee SI, Park SR (2023) OsWRKY114 is a player in rice immunity against Fusarium fujikuroi. Int J Mol Sci 24(7):6604. https://doi.org/10.3390/ijms24076604
Wang L, Chen J, Zhao Y, Wang S, Yuan M (2022) OsMAPK6 phosphorylates a zinc finger protein OsLIC to promote downstream OsWRKY30 for rice resistance to bacterial blight and leaf streak. J Integr Plant Biol 64(5):1116–1130. https://doi.org/10.1111/jipb.13249
Wani SH, Anand S, Singh B, Bohra A, Joshi R (2021) WRKY transcription factors and plant defense responses: latest discoveries and future prospects. Plant Cell Rep 40(7):1071–1085
Xu Z, Xu X, Gong Q, Li Z, Li Y, Wang S, Yang Y, Ma W, Liu L, Zhu B, Zou L, Chen G (2019) Engineering broad-spectrum bacterial blight resistance by simultaneously disrupting variable TALE-binding elements of multiple susceptibility genes in rice. Mol Plant 12(11):1434–1446. https://doi.org/10.1016/j.molp.2019.08.006
Yang B, Jiang Y, Rahman MH, Deyholos MK, Kav NN (2009) Identification and expression analysis of WRKY transcription factor genes in canola (Brassica napus L.) in response to fungal pathogens and hormone treatments. BMC Plant Biol 9:68. https://doi.org/10.1186/1471-2229-9-68
Acknowledgements
This research was funded by Research Program for Agricultural Science and Technology Development (PJ01570601) and the Fellowship Program (PJ01661001) of the National Institute of Agricultural Sciences, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Contributions
SS conceptualized the study. GS, SRP, and YJ performed the experiments and data analysis. GS, JL, and D-JH generated the constructs and the transgenic plants. GS and SS wrote the manuscript. SRP, N-CP, and SS supervised the study.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no conflicts of interests.
Ethics Approval
N/A.
Consent to Participate
N/A.
Consent for Publication
N/A.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, G., Park, S.R., Jeong, Y. et al. OsWRKY9 is Involved in Transcriptional Regulatory Cascade Enhancing Broad-Spectrum Disease Resistance. J. Plant Biol. (2024). https://doi.org/10.1007/s12374-024-09439-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12374-024-09439-3