Abstract
In the breeding of intermediate clones of sugarcane crop, the quality of high sugar and early-mature sucrose are important factors in the breeding program. However, the genetic diversity relationship of high-sucrose clones between parents groups is not well understood. The present study was designed to assess the relationship between high-sucrose clones and parents groups and use their potential in the breeding program. Twenty-three high-sucrose sugarcane clones of GT (HSGT) series and forty-one commonly used parents (CP) including 7 CP series from the USA, 9 ROC from Taiwan China, 14 from mainland China and 11 from other countries were analyzed at Sugarcane Research Institute, Guangxi Academy of Agricultural Sciences, Nanning, Guangxi, China, for determining their genetic relationship using simple sequence repeats (SSR) molecular markers. A total of 308 bands were obtained, of which 293 were polymorphic (95.1%). We found that a total of 309 loci were scored with 23 primers using ISSR techniques and that 277 loci were polymorphic. The genetic diversity of these five populations was 99% of the total genetic variation present within populations, while about 1% of variation was found among populations. The Nei's genetic distance between populations was found to be relatively small (0.04), and the average (0.96) Nei's genetic similarity of the five populations was high. The results indicated that the genetic relationship between the high-sugar containing materials and the commonly used parents could be developed and high-sugar containing materials could be used as hybrid parents to supplement high-sugar parents and help to develop breeding programs for high-sugar containing sugarcane offspring.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane is a major agricultural cash crop grown in almost 110 countries worldwide. China is the third major sugarcane producer and plays a major role in sugar production, especially in Guangxi province, China (Li and Yang 2015; Verma et al. 2020; 2021). Sugarcane industries in Guangxi account for 58.9% of China's sugarcane production and produced around 6.34 million tonnes in the crushing season of 2018/19 (Wang 2018; Chen et al. 2020). Sugarcane has great potential as a major feedstock for biofuel production, globally. It is considered among the best options for producing biofuels due to its outstanding biomass production capacity, high carbohydrate content and good energy input/output ratio. To increase the production of biofuels, it is very important to produce improved sugarcane varieties with better biomass degradability (Hoang et al. 2015; Ali et al. 2019; Wu et al. 2019). The complexity and size of the sugarcane genome is a major drawback in genetic improvement. Genetic diversity can be determined based on various approaches, morphological traits, pedigree records and molecular markers.
Sugarcane breeding contributes to most of the sugarcane production, but it is a long process that generally takes about 12 years. Some intermediate materials that may have a significant value in research and development may be eliminated in the sugarcane breeding process. In addition, new parental materials are more important for the development of modern varieties (Deng et al. 2004; Wu et al. 2008, 2019; Medeiros et al. 2020). A high-sugar population's genetic inheritance has higher heritability than that of the sugarcane yield, and the former is far more stable to be expressed in the sugarcane breeding process (Jackson and McRae 2001; Todd et al. 2020).
Nowadays, the use of molecular markers for the evaluation of genetic diversity is drawing the attention of researchers (Rao et al. 2016; Wu et al. 2019). It is well known that intersimple sequence repeat (ISSR) and simple sequence repeat (SSR) markers were used to analyze the diversity and genetic background of the sugarcane population, such as parents of resource nurseries (Liu et al. 2015; Ali et al. 2019; Medeiros et al. 2020). Breeders have used morphological traits to identify the relationship among varieties in the traditional way; however, morphological traits are mainly affected by plant development and the environment. Identification, based on morphological traits, is not suitable (Wang et al. 2009; Tew and Pan 2010; Ahmad et al. 2018). Molecular markers are an accurate and suitable technique to determine the genetic diversity of sugarcane cultivars and species (Silva et al. 2012; You et al. 2013; Santos et al. 2014; Wu et al. 2019; Medeiros et al. 2020).
Presently, a large number of different molecular marker systems have been developed for use in sugarcane and simple sequence repeats (SSRs) have been shown to be more efficient markers for breeding program, due to their availability in large quantities, are required in low doses, are co-dominant, reliable and can be used for multi-allelic detecting (Powell et al. 1996; Pan 2016; Ali et al. 2019; Wu et al. 2019). SSRs are categorized into mono-, di-, tri-, tetra-, penta- or hexa-SSRs based on the number of repeated base pairs and into perfect, imperfect and compound SSRs, which display perfect repetitions, interruption with novel nucleotides and two or more tandem motifs. SSR markers can be sorted by genomic or expressed sequence tag (EST) levels. SSRs can be classified as nuclear (nuSSR), mitochondrial (mtSSR) or chloroplast SSRs (cpSSR) according to their location in the genome. Most genomic SSRs are nuclear SSRs (Soranzo et al. 1999; Weising and Gardner 1999; Selkoe and Toonen 2006; Ahmad et al. 2018).
SSR markers have been used mainly to study the structure of sugarcane genetic diversity and population (Nayak et al. 2014; Liu et al. 2018; Ali et al. 2019; Medeiros et al. 2020), varietal identification, genetic map (Marconi et al. 2011; Pan 2016) and genetic association (Banerjee et al. 2015; Ukoskit et al. 2019; Wu et al. 2019). However, fluorescence-labeled SSR markers combined with high-performance capillary electrophoresis (HPCE) have showed better performance in genotyping of polyploid sugarcane, due to higher accuracy and better detection power (Fu et al. 2016; Ali et al. 2017, 2019; Ahmad et al. 2018; Xu et al. 2018; Wu et al. 2019).
Simple sequence repeat markers with high stability, multiple quantity and high polymorphism are more efficient for evaluating sugarcane germplasm in China and other countries (Pan 2006; Chen 2009; Wu et al. 2019; Medeiros et al. 2020). Yu et al. (2018) concluded that hereditary base core parents in China were narrow because of the limited number of parents. Genetic diversity was analyzed on commonly used parents and genotypes by using SSR markers in various countries (Liu et al. 2015; Rao et al. 2016; Wu et al. 2019; Pocovi et al. 2020).
The objective of this study is to compare the Jaccard’ s genetic coefficient between the early maturing intermediate materials of GT series and the commonly used parents by SSR markers from the Guangxi Parental Resource Nursery, China. Understanding the genetic relationship between the high-sugar materials and the commonly used parents could help in developing these high-sugar materials as hybrid parents for the selection and development of novel high-sugar varieties of parents.
Materials and Methods
Plant Materials and DNA Extraction
In this study carried out in October 2014, a total of sixty-four sugarcane genotypes were tested. Twenty-three early maturity high-sucrose clones of GT series with Brix more than ROC22 (17.73%) were collected from Sugarcane Research Institute, Guangxi Academy of Agricultural Sciences (GxAAS), Nanning, Guangxi, China, and forty-one parents of commonly used in Sugarcane Cross Breeding Center germplasm were collected from the Chinese Academy of Agricultural Sciences, Hainan, China. Sixty-four sugarcane genotypes were divided into five groups which comprised: 23 high sugarcane sucrose clones of GT series (HSGT) and 41 commonly used parents, including 7 CP series from the USA, 9 ROC series from Taiwan Sugar Research Institute, Taiwan China, 14 mainland China (MLCH) series including some GT series and 11 others from other countries (Table 1).
DNA extraction was followed by the SDS method with minor modifications according to Huang et al. (2010) and Gao et al. (2012). Photosynthetically mature sugarcane leaves (200 mg) were collected from different clones and separately ground as fine powder in liquid nitrogen and transferred to 2-ml sterilized tubes containing 1 ml pre-warm SDS buffer (SDS 1.5%, Tris100 mM, EDTA 20 mM, NaCl 500 mM). The genomic DNA was extracted following the traditional method.
SSR Analysis
Twenty-three SSR primer pairs were used to determine the diversity among high-sucrose clones from the GT series and commonly used parents. Genomic primer sequences followed the International Federation of Sugarcane Biotechnology guidelines (Cordeiro et al. 2000), and EST-SSR primer sequences were obtained from the literature (Oliveira et al. 2009).
Polymerase chain reaction (PCR) amplification was performed in a total of 20 μL volume containing 2 μL 10 × Buffer (including 2 mmol/L MgCl2), 1 μmol/L of each forward and reverse primer, 0.4 μL dNTPs 0.2 mmol/L, 1 μL template 30 ng/L DNA and 0.2 μL Taq polymerase, 14.4 μL ddH2O. PCR was carried out by initial pre-denaturation at 95 °C (5 min), denaturation at 94 °C (30 s), annealing at 53 °C (30 s) and extension at 72 °C (1 min), followed by 35 cycles with final extension step at 72 °C for 5 min (PCR, T-gradient 96, Biometra, Germany). The PCR products were electrophoresed at 120 V in 7% polyacrylamide gel for 90 min and photographed under UV light using a gel documentation system.
The Genetic Diversity of the Five Populations
To assess the proportion of polymorphic loci (PPL), Shannon's diversity index (I), Nei's gene index (H), the observed number of alleles (Na) and the effective number of alleles (Ne) to evaluate levels of genetic diversity among the populations using the POPGENE program v. 1.32 were carried out (Francis et al. 1999). The principal coordinate analysis (PCoA) was carried out to determine the genetic relationships among populations. PCoA, analysis of molecular variance (AMOVA) and Nei’s genetic distance and similarity were carried out using the GenALEx6.5 program (Peakall and Smouse 2012). The estimated groups’ numbers were used by STRUCTURE program version 2.3.4 (Evanno et al. 2005), and the results were obtained by using STRUCTURE HARVESTE (Earl and VonHoldt 2012).
Data Analysis
The data on bands generated by the twenty-three primers on sixty-four genotypes were analyzed for genetic diversity. All segregating bands were scored manually as 1 for presence and 0 for absence. Based on Jaccard's coefficient (Jaccard 1908), the genetic similarity (GS) was carried out using NTSYS-pc 2.10d. Cluster trees under the unweighted pair-group method with arithmetic mean (UPGMA) were constructed by DPS software (Tang et al. 2013).
Results
Cluster Analysis
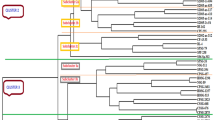
The cluster analyses of internodes in 64 genotypes revealed a general structure between HSGT (23) and commonly used parents (41) from the germplasm of Guangxi sugarcane 'parents' (Fig. 1; Table 1). Around the degree of 0.61, all genotypes could be divided into 10 clusters. In cluster I included most of HSGT by 21 in 24 clones and 8 MLCH, 4 ROC series and 3 others in the germplasm of Guangxi sugarcane 'parents.' It indicated that these HSGT were closely related to the parents in cluster I and were carefully crossed with each other when HSGT became parents in the future. Specially, ROC22 accounts for the largest area in terms of sugarcane acreage and the main hybrid parent in China, but it is high in usage rate and low in combining Brix's ability (Wu et al. 2019). Therefore, it is suggested that in future fewer high-sugar parents should be made. But GT08-509 (X), YC89-7 (IX), POJ2827 (VIII), NcO293 & CP84-1198 (VII) and Zanz74-141& CP81-2149 (VI) were far from other genotypes. Therefore, using YC89-7 (IX), POJ2827 (VIII), NcO293& CP84-1198 (VII) and Zanz74-141 & CP81-2149 (VI) as hybrid parents it might be difficult to get high-sucrose offspring, but GT08-509 as an HSGT should fully utilize its high sucrose and other genetic backgrounds for crossing with the parent in future.
SSR Analysis
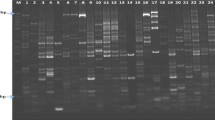
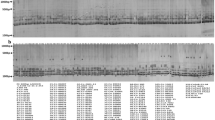
A total of 23 primer pairs were selected for SSR primer collection and to detect polymorphism in 64 sugarcane genotypes (Table 2). A total of 309 bands were amplified by 23 primers, with an average of about 13.48 polymorphic bands (ranking 7–22 bands). Two hundred ninety-four polymorphic bands were obtained with a polymorphic rate of 94.8% in 309 bands. Eleven primers reached 100% of polymorphic rate by mSSCIR1, mSSCIR21, mSSCIR3, mSSCIR43, mSSCIR66, mSSCIR9, mSSCIR9 and SMC851MS. This shows that the high polymorphic profiles by using these primers can assess genetic diversity in 64 genotypes.
Polymorphic analysis of 5 groups by 23 high-sucrose clones from GT series HSGT and 41 commonly used parents from CP series, ROC series, MLCH and others by 23 primers showed no more difference in the number and rate of occurrence of polymorphic bands. But HSGT showed the highest polymorphic band numbers (285) and polymorphic rate (95.3%) among the 5 groups (Table S1). This indicates that 23 primers worked excellently in all the 5 groups. The sharing status of amplification bands (Table S2) shows the same amplification bands in the correlation of two groups. HSGT was sharing 4 bands with the parent groups of MLCH, while the parent groups of CP series and others shared 6 bands.
Genetic Similarity Coefficient Analysis
According to the statistics of amplified bands from SSR locus, the genetic similarity coefficient of 64 genotypes ranged from 0.460 to 0.881, with an average of 0.613, and showed certain differences in 64 genotypes. The minimum genetic similarity coefficient was 0.460 and far from the genetic distance as seen in Q208 and GT008-509. The highest values of genetic similarity coefficients were found in GT07-548 and GT06-1857 (0.881), with the highest genetic correlation and close relationship of genetic similarity. In the collection of 23 clones in the HSGT group, the average GS was 0.633 indicating certain genetic differences in these clones. HSGT has great potential in the development of hybrid parents. GS of 23 clones in HSGT with 7 CP series by 0.604 and other parents by 0.608 was higher than CP series (0.581) and others-self (0.607), respectively (Table S3). It indicated that the relationship of HSGT was narrower than that of the parents of CP series and also with those of other countries in Guangxi germplasm. Since the 1950s, CP series characterized by high sugar has played an essential role in the breeding of sugarcane parents in China (Qi et al. 2012; Liu et al. 2015). Genetic similarity of 23 clones in HSGT with ROC series was the closest relation by 0.622. Presently, ROC22 and ROC16 account for the largest plantation area, more than 80% in China (Liu et al. 2018). Therefore, ROC varieties have been widely developed and utilized in sugarcane hybridization. The Guangxi province in China produces five varieties among the top 10 of the most widely commercial varieties in China including GT29, GT42, GT49, LC05-136 and ROC series offspring. Genetic similarity information between HSGT and parent groups should guide the breeder to make the program for high-sugar offspring and the utilization of HSGT as hybrid parents.
Genetic Diversity and Relationships Among Genotypes
Among the five populations, the mean values of the proportion of polymorphic loci (PPL), the observed number of alleles (Na), the effective number of alleles (Ne), Nei's gene index (H) and Shannon's diversity index (I) are presented in Table 3. At the species level, PPL, Na, Ne, I and H were 89.64%, 1.89 ± 0.03, 1.47 ± 0.01, 0.28 ± 0.01 and 0.43 ± 0.01, respectively.
Among the five populations, the total gene diversity index (Ht) and gene diversity within the population index (Hs) were 0.21 ± 0.03 and 0.15 ± 0.01, respectively. The genetic differentiation index among the five populations (Gst) was 0.28. Analysis of molecular variance (AMOVA) showed that a relatively large proportion of genetic variation (99%) occurred within the populations, whereas only 1% of genetic variation was observed among the eight populations (p < 0.001; Table 4). The average value of Nei's genetic distance and genetic similarity index was 0.07 and 0.93 (Table S4).
Principal Coordinate Analysis
The principal coordinate analysis helps to illustrate the genetic relationships of sugarcane parents as compared to individual units and is calculated based on the SSR data matrix of the 5 loci of all 64 sugarcane accessions available in the present study (Fig. 2). According to the principal coordinates (Fig. 2), the distribution of HSGT among others and CP series populations was far away and the GS similarity coefficient was relatively small by 0.604 and 0.068, respectively.
STRUCTURE Analysis
STRUCTURE analysis showed that the delta K displayed peaks at K = 3 (Fig. 3a and b). This indicates that the 64 individuals were clustered into 3 groups with 3 colors. Each individual is represented by a vertical colored line. The same color of different individuals indicates that they belong to the same cluster.
Discussion
Constant efforts are being made in China to improve sugarcane yield, particularly in cane yield and sugar content. Sugarcane breeding programs have enhanced these efforts to a great extent. Breeding programs based on genetic diversity are currently in the focus of agricultural research. In breeding programs, the hybrid varieties, when backcrossed with parents resulted in offspring with higher sugar content (Chen et al. 2009; Aitken et al. 2018; Medeiros et al. 2020). The molecular markers-based breeding program helps to overcome the limitations of conventional breeding techniques, as well as helps to understand the genetic susceptibility of the hybrids to various biotic and abiotic factors that affect plant growth and development (Govindaraj et al. 2011; Moore et al. 2013; Ahmad et al. 2018). For a successful breeding program, it is vital that the parents should be genetically divergent and the offspring tolerant to various stress factors (You et al. 2013; Neto et al. 2020). Microsatellite or SSRs markers are recent developments in research on breeding programs associated with crop improvement, and sugarcane breeding programs have identified SSR markers as more useful for identifying hybrid parents (Santos et al. 2014; Manechini et al. 2018; Ahmad et al. 2018). SSR-based breeding programs have demonstrated high variability, wide genomic distribution, co-dominant inheritance, high reproducibility, large multiallelic nature and specific chromosomal location and hence are gaining importance in breeding programs, especially in sugarcane (Neto et al. 2020; Medeiros et al. 2020).
In the present study, Jaccard's genetic coefficient was used to compare the genetic diversity of sugarcane hybrids. In this study, the sixty-four sugarcane genotypes when tested for genetic similarities using SSR markers, demonstrated high polymorphism. Based on the results obtained from the data on SSRs using the Jaccard similarity coefficient, HSGT showed the highest polymorphic band number of 285 bands and a polymorphic rate of 95.3% among the five groups. Among the 23 clones in the HSGT group, the average GS was 0.633 that indicated certain genetic differences in this clone. Studies on SSR-based breeding programs showed that those progenies which inherited parental traits, the possibility of contamination in the pedigree was evident. It was also observed that SSR markers can be used in differentiating true hybrids from those of contaminants (Santos et al. 2014; Parthiban et al. 2018). Under the breeding programs, Wu et al. (2019) applied the SSR markers for management of parental germplasm in sugarcane (Saccharum spp. hybrids). SSR markers are more suitable for the identification of parent clusters in breeding programs (Chen et al. 2009; You et al. 2013; Ahmad et al. 2018; Ali et al. 2019). Based on Jaccard's coefficient, it was observed that the GT07-548 and GT06-1857 were highly correlated, and therefore, together they can be regarded as poor parents together for the breeding program.
The present study also concluded that the 23 primers showed significant results in terms of measurement of genetic diversity and mapping with all the five groups, which was in line with the previous studies (Chen et al. 2009; Ahmad et al. 2018). These primers could be effectively used for sugarcane breeding programs to identify the genetically divergent parents. Researchers have shown that primer polymorphism enhances the efficacy of inter-specific hybrid identification (Yang et al. 2006; Saha et al. 2017; Manechini et al. 2018). The PCR-based DNA markers make it possible to analyze the degree of genetic variability that occurs among conventional progenitor species and commercial cultivars in sugarcane breeding trials (Singh et al. 2011; You et al. 2016; Ahmad et al. 2018; Medeiros et al. 2020).
Genetic similarity information between HSGT and parent groups could help the breeder to develop high-sugar offspring varieties using HSGT as hybridization parents. GT08-509 as an HSGT should fully utilize its high sucrose and other genetic background for crossing with the parent in the near future. The comparison with other populations as HSGT and ROC series found the largest similarity coefficient (0.622) by GS and small genetic distance value (0.048) by NEI. At the same time, according to structure (Fig. 3b), HSGT and ROC series showed that the rate of blue bars was more, indicating that the ROC group and HSGT are more or less similar. HSGT was at a far genetic distance from the two-parent populations. In the Guangxi breeding program, breeders should increase the rate of high sucrose in far genetic distance to obtain offspring with high sucrose and strong genetic diversity.
In conclusion, the analysis of variations in SSR fragments provides a useful tool for determining diversity to develop plant breeding strategies. In the coming years, the acceptance and use of SSR-based markers will increase significantly in the breeding of sugarcane. Identifying useful SSRs is critical, but in sugarcane, this can be a prolonged and complex process because the sugarcane genome is highly complex. These markers may be used for the construction of a genetic map in sugarcane. Further work on crosses between and within the groups identified in this study may provide useful strategies for identifying favorable genes and alleles in newly developed sugarcane varieties.
References
Ahmad, A., J.D. Wang, Y.B. Pan, R. Sharif, and S.J. Gao. 2018. Development and use of simple sequence repeats (SSRs) markers for sugarcane breeding and genetic studies. Agronomy 8: 260. https://doi.org/10.3390/agronomy8110260.
Aitken, K., J. Li, G. Piperidis, C. Qing, F. Yuanhong, and P. Jackson. 2018. Worldwide genetic diversity of the wild species Saccharum spontaneum and level of diversity captured within sugarcane breeding programs. Crop Science 58: 218–229.
Ali, A., J. Wang, Y. Pan, Z. Deng, Z. Chen, R. Chen, and S. Gao. 2017. Molecular identification and genetic diversity analysis of Chinese sugarcane (Saccharum spp. Hybrids) varieties using SSR markers. Tropical Plant Biology 10: 194–203.
Ali, A., Y. Pan, Q. Wang, J. Wang, J. Chen, and S. Gao. 2019. Genetic diversity and population structure analysis of Saccharum and Erianthus genera using microsatellite (SSR) markers. Scientific Reports 9 (1): 395. https://doi.org/10.1038/s41598-018-36630-7.
Banerjee, N., A. Siraree, S. Yadav, S. Kumar, J. Singh, S. Kumar, D.K. Pandey, and R.K. Singh. 2015. Marker-trait association study for sucrose and yield contributing traits in sugarcane (Saccharum spp. hybrid). Euphytica 205: 185–201.
Chen, G.L., F.J. Zheng, B. Lin, S.B. Lao, J. He, Z. Huang, Y. Zeng, J. Sun, and K.K. Verma. 2020. Phenolic and volatile compounds in the production of sugarcane vinegar. ACS Omega 5: 30587–30595.
Chen, P.H., Y.B. Pan, R.K. Chen, L.P. Xu, and Y.Q. Chen. 2009. SSR marker-based analysis of genetic relatedness among sugarcane cultivars (Saccharum spp. hybrids) from breeding programs in China and other countries. Sugar Tech 11: 347–354.
Cordeiro, G.M., G.O. Taylor, and R.J. Henry. 2000. Characterization of micro-satellite markers from sugarcane (Saccharum ssp.), a highly polyploid species. Plant Science 155: 161–168.
Deng, H.H., Q.W. Li, and Z.Y. Chen. 2004. Breeding and utilization of new sugarcane parents. Sugarcane 11 (3): 7–12.
Earl, D.A., and B.M. Vonholdt. 2012. Structure harvester: A website and program for visualizing structure output and implementing the evanno method. Conservation Genetics Resources 4: 359–361.
Evanno, G., S. Rengaut, and J. Goudet. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Molecular Ecology 14: 2611–2620.
Francis, C.Y., C.Y. Rong, and T. Boyle. 1999. POPGENE, Microsoft Window-based freeware for population genetic analysis. Version 1.31. Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton, Canada
Fu, Y., Y. Pan, C. Lei, M.P. Grisham, C. Yang, and Q. Meng. 2016. Genotype-specific microsatellite (SSR) markers for the sugarcane germplasm from the Karst Region of Guizhou, China. American Journal of Plant Science 7: 2209–2220.
Gao, Y.J., F.X. Fang, X.H. Liu, R.H. Zhang, H.Z. Song, R.Z. Yang, T. Luo, W.X. Duan, J.H. You, and G.M. Zhang. 2012. Identification of progeny from crosses between sugarcane (Saccharum spp.) and intergeneric hybrid complex (Erianthus arundinaceus × Saccharum spontaneum) with molecular markers. Journal of Plant Genetic Resources 13: 912–916.
Govindaraj, P., R. Sindhu, A. Balamurugan, and C. Appunu. 2011. Molecular diversity in sugarcane hybrids (Saccharum spp. complex) grown in peninsular and east coast zones of tropical India. Sugar Tech 13: 206–213.
Hoang, N.V., A. Furtado, F.C. Botha, B.A. Simmons, and R.J. Henry. 2015. Potential for genetic improvement of sugarcane as a source of biomass for biofuels. Frontiers in Bioengineering and Biotechnology 3: 182. https://doi.org/10.3389/fbioe.2015.00182.
Huang, D.L., X.L. Qin, Q. Liao, Y.J. Gao, and F.X. Fang. 2010. Simple and rapid procedure for isolation of high quality genomic DNA from sugarcane. Biotechnology Bulletin 5: 101–106.
Jaccard, P. 1908. Nouvelles recherches sur la distribution florale. Bulletin De La Sociéte Vaudoise Des Sciences Naturelles 44: 223–270.
Jackson, P.A., and T.A. McRae. 2001. Selection of sugarcane clones in small plots: Effects of plot size and selection criteria. Crop Science 41: 315–322.
Li, Y.R., and L.T. Yang. 2015. Sugarcane agriculture and sugar industry in China. Sugar Tech 17: 1–8.
Liu, H.L., X.P. Yang, Q. You, J. Song, L.P. Wang, J.S. Zhang, Z.H. Deng, R. Ming, and J.P. Wang. 2018. Pedigree, marker recruitment and genetic diversity of modern sugarcane cultivars in China and the United States. Euphytica 214: 48.
Liu, X.L., X.J. Li, H.B. Liu, L. Ma, C.H. Xu, and H.Y. Fan. 2015. Genetic diversity analysis of Yunnan commonly-used parents by using SSR marker. Journal of Plant Genetic Resources 16: 1214–1222.
Manechini, J.R.V., J.B. da Costa, B.T. Pereira, L.A. Carlini-Garcia, M.A. Xavier, M.G.dA. Landell, and L.R. Pinto. 2018. Unraveling the genetic structure of Brazilian commercial sugarcane cultivars through microsatellite markers. PLoS One 13: e0195623. https://doi.org/10.1371/journal.pone.0195623.
Marconi, T.G., E.A. Costa, H.R. Miranda, M.C. Mancini, and C.B. Cardososilva. 2011. Functional markers for gene mapping and genetic diversity studies in sugarcane. BMC Research Notes 4: 264.
Medeiros, C., T.W.A. Balsalobre, and M.S. Carneiro. 2020. Molecular diversity and genetic structure of Saccharum complex accessions. PLoS ONE 15: e0233211. https://doi.org/10.1371/journal.pone.0233211.
Moore, P.H., A.H. Paterson, and T. Tew. 2013. Sugarcane: The Crop, the Plant, and Domestication. In Sugarcane: Physiology, Biochemistry, and Functional Biology (eds P.H. Moore and F.C. Botha). https://doi.org/10.1002/9781118771280.ch1
Nayak, S.N., J. Song, A. Villa, B. Pathak, T. Ayala-Silva, X. Yang, J. Todd, N.C. Glynn, D.N. Kuhn, B. Glaz, R.A. Gilbert, J.C. Comstock, and J. Wang. 2014. Promoting utilization of Saccharum spp. genetic resources through genetic diversity analysis and core collection construction. PLoS ONE 9(10): e110856. https://doi.org/10.1371/journal.pone.0110856.
Neto, H.Z., L.G. da M. Borsuk, L.R.F. dos Santos, H.S. Angeli, G.S. Berton, and L.L. de Sousa. 2020. Genetic diversity and population structure of sugarcane (Saccharum spp.) accessions by means of microsatellites markers. Acta Scientiarum Agronomy 42: e45088. https://doi.org/10.4025/actasciagron.v42i1.45088.
Oliveira, K.M., L.R. Pinto, T.G. Marconi, M. Mollinari, E.C. Ulian, S.M. Chabregas, M.C. Falco, W. Burnquist, A.A.F. Garcia, and A.P. deSouza. 2009. Characterization of new polymorphic functional markers for sugarcane. Genome 52: 191–209.
Pan, Y.B. 2006. Highly polymorphic microsatellite DNA markers for sugarcane germplasm evaluation and variety identity testing. Sugar Tech 8: 246–256.
Pan, Y.B. 2016. Development and integration of an SSR-based molecular identity database into sugarcane breeding program. Agronomy 6: 28.
Parthiban, S., P. Govindaraj, and S. Senthilkumar. 2018. Comparison of relative efficiency of genomic SSR and EST-SSR markers in estimating genetic diversity in sugarcane. 3 Biotech 8: 144. https://doi.org/10.1007/s13205-018-1172-8
Peakall, R., and P.E. Smouse. 2012. Genalex 6.5: Genetic analysis in excel. Population genetic software for teaching and research–an update. Bioinformatics 28: 2537–2539.
Pocovi, M.L., N. Collavino, N. Gutirrez, G. Taboada, V. Castillo, R. Delgado, and J.A. Mariotti. 2020. Molecular versus morphological markers to describe variability in sugar cane (Saccharum officinarum) for germplasm management and conservation. Revista De La Facultad De Ciencias Agrarias 52: 40–60.
Powell, W., G.C. Machray, and J. Provan. 1996. Polymorphism revealed by simple sequence repeats. Trends in Plant Science 1: 215–222.
Qi, Y.W., H.H. Deng, and Q.W. Li. 2012. Advance in utilization of sugarcane germplasm in China Mainland. Corp Research 26: 443–446.
Rao, V.P., S. Singh, R. Chaudhary, M.K. Sharma, R.S. Sengar, U. Singh, and V. Sharma. 2016. Genetic variability in sugarcane (Saccharum spp. hybrid) genotypes through inter simple sequence repeats (ISSR) markers. Journal of Applied and Natural Science 8: 1404–1409.
Saha, D., R.S. Rana, S. Chakrabarty, S. Datta, A.A. Kumar, A.K. Chakrabarty, and P.G. Karmakar. 2017. Development of a set of SSR markers for genetic polymorphism detection and interspecific hybrid jute breeding. Crop Journal 5: 416–429.
Santos, J.M.D., and G.V.dS. Barbosa, C.E.R. Neto and C. Almeida. . 2014. Efficiency of biparental crossing in sugarcane analyzed by SSR markers. Crop Breeding and Applied Biotechnology 14: 102–107.
Selkoe, K.A., and R.J. Toonen. 2006. Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecology Letters 9: 615–629.
Silva, D.C., L.S.C. Duarte Filho, J.M. Santos, G.V. Barbosa, and C. Almeida. 2012. DNA fingerprinting based on simple sequence repeat (SSR) markers in sugarcane clones from the breeding program RIDESA. African Journal of Biotechnology 11: 4722–4728.
Singh, R.K., R.B. Singh, S.P. Singh, and M.L. Sharma. 2011. Identification of sugarcane microsatellites associated to sugar content in sugarcane and transferability to other cereal genomes. Euphytica 182: 335–354.
Soranzo, N., J. Provan, and W. Powell. 1999. An example of microsatellite length variation in the mitochondrial genome of conifers. Genome 42: 158–161.
Tang, Q.Y., and C.X. Zhang. 2013. Data Processing System (DPS) software with experimental design, statistical analysis and data mining developed for use in entomological research. Insect Science 20: 254–260.
Tew, T.L., and Y.B. Pan. 2010. Microsatellite (Simple Sequence Repeat) marker–based paternity analysis of a seven-parent sugarcane polycross. Crop Science 50: 1401–1408.
Todd, J., Y.B. Pan, and D. Boykin. 2020. Fidelity of sugarcane crosses assessed with SSR markers. Agronomy 386: 386.
Ukoskit, K., G. Posudsavang, N. Pongsiripat, P. Chatwachirawong, P. Klomsa-ard, P. Poomipant, and S. Tragoonrung. 2019. Detection and validation of EST-SSR markers associated with sugar-related traits in sugarcane using linkage and association mapping. Genomics 111: 1–9.
Verma, K.K., P. Singh, X.P. Song, M.K. Malviya, R.K. Singh, G.L. Chen, S. Solomon, and Y.R. Li. 2020. Mitigating climate change for sugarcane improvement: Role of silicon in alleviating abiotic stresses. Sugar Tech 22: 741–749.
Verma, K.K., X.P. Song, Y. Zeng, D.J. Guo, M. Singh, V.D. Rajput, M.K. Malviya, K.J. Wei, A. Sharma, D.P. Li, G.L. Chen, and Y.R. Li. 2021. Foliar application of silicon boosts growth, photosynthetic leaf gas exchange, antioxidative response and resistance to limited water irrigation in sugarcane (Saccharum officinarum L.). Plant Physiology and Biochemistry 166: 582–592. https://doi.org/10.1016/j.plaphy.2021.06.032
Wang, C.H. 2018. Development trend of sugar cane industry in Guangxi and counter measures for upgrading and transforma. Guangxi Sugar Industry 3: 45–48.
Wang, X.H., Q.H. Yang, F.S. Li, L.L. He, and S.C. He. 2009. Molecular identification of Saccharum spp × Erianthus fulvus hybrids using sequence-characterized amplified region markers. Crop Science 49: 864–870.
Weising, K., and R.C. Gardner. 1999. A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42: 9–19.
Wu, C.W., J.U. Liu, J. Zhao, P.F. Zhao, and C.X. Hou. 2008. Research on breeding potential and variety improvement of exotic parents in sugarcane. Southwest China Journal of Agricultural Sciences 21: 1671–1675.
Wu, J., Q. Wang, J. Xie, Y.-B. Pan, F. Zhou, Y. Guo, H. Chang, H. Xu, W. Zhang, C. Zhang, and Y. Qiu. 2019. SSR marker-assisted management of parental germplasm in sugarcane (Saccharum spp. hybrids) breeding programs. Agronomy 9: 449. https://doi.org/10.3390/agronomy9080449.
Xu, W.V., S.H. Long, C.S. Qiu, Y. Guo, H. Wang, D.M. Hao, and Y.F. Wang. 2018. Analysis of SSR primers and genetic diversity of 95 sugarcane culture materials. Southwest China Journal of Agricultural Sciences 31: 2498–2505.
Yang, X., B.E. Scheffler, and L.A. Weston. 2006. Recent developments in primer design for DNA polymorphism and mRNA profiling in higher plants. Plant Methods 2: 4. https://doi.org/10.1186/1746-4811-2-4.
You, Q., L. Xu, Y. Zheng, and Y. Que. 2013. Genetic Diversity analysis of sugarcane parents in Chinese Breeding Programmes using SSR markers. The Scientific World Journal 613062: 11.
You, Q., Y.B. Pan, S.W. Gao, Q.N. Wang, Y.C. Su, Y.Q. Yang, Q.B. Wu, D.G. Zhou, and Y.X. Que. 2016. Genetic diversity analysis of sugarcane germplasm based on fluorescence-labeled simple sequence repeat markers and a capillary electrophoresis-based genotyping platform. Sugar Tech 18: 380–390.
Yu, F., P. Wang, X. Li, Y. Huang, Q. Wang, L. Luo, Y. Jing, X. Liu, Z. Deng, J. Wu, Y. Yang, R. Chen, M. Zhang, and L. Xu. 2018. Characterization of chromosome composition of sugarcane in nobilization by using genomic in situ hybridization. Molecular Cytogenetics 11: 35.
Acknowledgements
The present study was supported by National Natural Science Foundation of China (32060468), Guangxi Natural Science Foundation Program (2020GXNSFBA159024), China Agriculture Research System (CARS-170105), Basic Research Program of Guangxi Academy of Agricultural Sciences (GNK2019M35), Government Booting Local Program (ZY20111001) and Guangxi Academy of Agricultural Sciences, Nanning, Guangxi, China, for providing the necessary facilities for this study.
Author information
Authors and Affiliations
Contributions
Hai-Rong Huang and Yi-Jing Gao designed, performed experiment and wrote original draft, Mukesh K. Malviya, Krishan K. Verma and Manoj K. Solanki helped in figure preparation, statistical analysis and writing the original draft, Yu-Xin Huang, Xiang Li, Yuchi Deng, Jing Yan, Shi-Yun Tang and Lun-Wang Wang helped in experimental analysis and software. Lin Xu critically revised the original draft. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for Publication
All authors included in this study consent for publication.
Availability of Data
None.
Human and Animal Rights
The present research did not involve human participants and/or animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huang, HR., Gao, YJ., Malviya, M.K. et al. Genetic Diversity Analysis of Sugarcane (Saccharum spp. hybrids) Among High-Sucrose Clones of GT Series and Commonly Used Parents by Using Microsatellite Markers in Guangxi, China. Sugar Tech 24, 397–407 (2022). https://doi.org/10.1007/s12355-021-01033-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-021-01033-y