Abstract
Epstein-Barr virus (EBV) is one of the most common viruses associated with multiple malignancies including hematopoietic, epithelial, and mesenchymal neoplasms. EBV is linked to B- and T-cell lymphomas, ranging from indolent to highly aggressive neoplasms. EBV-positive follicular lymphoma (FL) is not well characterized due to its low prevalence. In this report, we describe a case of EBV-positive FL and concurrent EBV-negative diffuse large B-cell lymphoma (DLBCL), and discuss their clonal relationship, and EBV status in the process of disease progression. Histology, immunohistochemistry, in situ hybridization, and next-generation sequencing studies were performed as previously described. The 58-year-old male presented with extensive axillary and subpectoral lymphadenopathy. The patient had a history of mixed connective tissue disease treated in the past with steroids and methotrexate, and at the time of current presentation with hydroxychloroquine. The excision of axillary lymph node showed coexistent EBV-positive FL (grade 3B) and EBV-negative DLBCL. There was no evidence of BCL2 gene rearrangement; however, both EBV-positive neoplastic follicles and diffuse component harbored MYC/IGH rearrangement. Next-generation sequencing suggested branching evolution with shared DDX3X mutation, a number of private mutations, and unique IGH usage in FL and DLBCL. The patient was treated with six cycles of R-CHOP (rituximab, cyclophosphamide, doxorubicin, vincristine, and prednisolone) with involved-field radiotherapy and remains in complete remission. To the best of our knowledge, this is the first report of BCL2 rearrangement negative, MYC/IGH-positive and EBV-positive FL, and concurrent EBV-negative DLBCL, which supports branched evolution model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Epstein-Barr (EBV)-positive follicular lymphoma (FL) is uncommon with a prevalence of 2.5 to 6% of follicular lymphomas [1] [2] [3]. While fewer than 30 cases have been previously reported, several common features have emerged. Typically, EBV-positive FL is of histologic grade 3 and/or commonly progresses to higher-grade FL or diffuse large B-cell lymphoma (DLBCL) [3] [4]. Secondly, in contrast to other EBV-associated lymphomas, many patients with EBV-positive FL are immunocompetent.
In this report, we describe a case of EBV-positive FL and concurrent EBV-negative DLBCL. Considering shared molecular genetic abnormalities, DLBCL is most likely clonally related to FL and this case illustrates branched evolution model which has been reported in the majority of FL [5]. In contrast to some other EBV-positive lymphoproliferations [6], this case was characterized by an indolent clinical course.
Clinical history
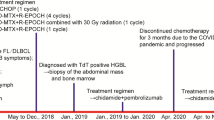
The patient is a 58 year-old Caucasian male with a history of mixed connective tissue disease diagnosed in 1992. In the past, the patient was treated with steroids and methotrexate, and currently has only been receiving hydroxychloroquine. The patient presented with extensive right axillary and subpectoral lymphadenopathy. He reported fevers but denied weight loss or night sweats. A PET/CT scan showed bulky, highly FDG-avid axillary lymphadenopathy. A core needle biopsy revealed focal proliferation of large atypical lymphoid cells positive for EBER and was considered not sufficient for a definitive diagnosis. Subsequently, a lymph node excision was performed and a diagnosis of EBV-positive FL (grade 3B) transforming to DLBCL was rendered. The patient was treated with six cycles of R-CHOP (rituximab, cyclophosphamide, doxorubicin, vincristine, and prednisolone) with involved-field radiotherapy and remains in complete remission since 2016.
Material and methods
Formalin-fixed and paraffin-embedded tissue (FFPE) was used for hematoxylin and eosin stain, immunohistochemistry (IHC), and in situ hybridization (ISH). IHC was performed using antibodies against CD20, PAX5, BCL6, CD10, HGAL, LMO2, BCL2, CD30, MYC, CD21, CD23, CD35, MUM1, kappa, and lambda light chain. EBV was detected by EBER 1 DNP Ventana ISH probe against EBV-encoded small RNAs (EBER). The IHC and EBV ISH were performed in the clinical diagnostic laboratory of Indiana University Health according to the previously described procedures. EBV nuclear antigen 1 (EBNA1) IHC was performed as previously described at the University of Wisconsin Madison [7]. Fluorescence in situ hybridization analysis for MYC, BCL2, and BCL6 rearrangements (Abbott Molecular, Abbott Park, IL) was performed at the Mayo Clinic (Rochester, MN) as previously described [8].
Next-generation sequencing (NGS) was performed separately for FL and DLBCL components at Indiana University Health. DNA and RNA sequencing included 453 genes involved in lymphomas, relevant viruses, and dominant immunoglobulin (IG) heavy chain usage. The average coverage was 334X and 240X for follicular and diffuse components, respectively. In addition, B-cell clonality was examined by PCR with BIOMED-2 immunoglobulin heavy chain (IgH) clonality assay using micro-dissected FFPE tissue from FL and DLBCL as previously described [9].
Results
Histologic sections showed lymph nodes with architecture almost entirely effaced by a nodular and diffuse proliferation of large, atypical lymphoid cells with moderately abundant cytoplasm, round to oval vesicular nuclei, and prominent multiple or single nucleoli consistent with centroblasts and immunoblasts (Fig. 1. A–D). Diffuse areas included prominent necrosis and extended into perinodal soft tissues. Neoplastic follicles were scattered about and were also present in areas with partially preserved nodal architecture including secondary follicles, paracortical expansion, and plasmacytosis. In addition, extensive granulomata composed of epithelioid histiocytes and giant cells were seen surrounding neoplastic follicles and were also scattered throughout lymph nodes.
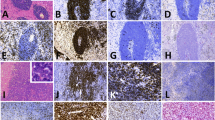
Histologic and cytomorphologic features of follicular and diffuse components. A. Neoplastic follicles, H&E 200x. B. Diffuse area with focal necrosis, H&E 200x. C. Neoplastic follicles were composed of large atypical lymphoid cells with abundant cytoplasm, round to oval vesicular nuclei and prominent nucleoli, H&E 400x. D. Similar cytomorphologic features were seen in diffuse areas, H&E 400x
The lymphoid cells in neoplastic follicles were positive for CD20, PAX5, BCL6, CD10, HGAL, and MYC (Fig. 2 A, B, E), and negative for LMO2 and MUM1. The BCL2 expression in neoplastic follicles was focal and weak. The reactive residual follicules were positive for CD20, PAX5, CD10, BCL6, LMO2, and HGAL. Immunostains for CD21, CD35, and CD23 showed prominent follicular dendritic cell meshworks in neoplastic follicles and were absent in areas transitioning to a diffuse pattern (Fig. 2 G). Neoplastic cells in diffuse areas were positive for CD20, PAX5, CD10, BCL6, BCL2, and MYC, weakly positive for HGAL and negative for LMO2 and MUM1 (Fig. 2 C, D, F). CD30 immunostain was negative in both follicular and diffuse areas. Kappa and lambda light chain immunostains were negative in neoplastic lymphoid cells and showed a polyclonal pattern of expression in plasma cells.
Immunohistochemical stains, EBER in situ hybridization and fluorescence in situ hybridization. A. CD20 immunostain in neoplastic follicles. B. CD10 immunostain in neoplastic follicles. C. BCL6 immunostain in diffuse area. D. BCL2 in diffuse component. MYC immunostain in follicular (E) and diffuse area (F). G. CD21 immunohistochemical stain highlights follicular dendritic cell meshwork in the neoplastic follicle. EBER in situ hybridization was positive in neoplastic follicles (H) and negative in diffuse areas (I). J. MYC break-apart FISH probe was positive (arrows) in follicular and diffuse areas. K. Fluorescence in situ hybridization using a MYC/IGH dual-color dual-fusion probe set demonstrated MYC/IGH rearrangement (arrows) in follicular and diffuse areas. L. BCL6 probe demonstrated a BCL6 rearrangement (arrow) in neoplastic lymphoid cells only in follicular areas
Neoplastic follicles were positive for EBER ISH in more than 80% of nuclei, whereas diffuse areas were negative for EBER (Fig. 2 H, I). Neoplastic follicles and diffuse areas were negative for EBNA1 and LMP1. EBV status was confirmed by NGS, which showed a presence of EBV in follicular component and DLBCL negative for EBV.
We also performed FISH studies for BCL2, BCL6, and MYC rearrangements. MYC break-apart probe (BAP) and MYC/IGH dual-color dual-fusion (D-FISH) probes sets identified a MYC/IGH rearrangement in 95–100% of interphase nuclei in both follicular and diffuse areas (Fig. 2 J, K). In addition, BCL6 (BAP) probe showed 58% of interphase nuclei with BCL6 rearrangement in follicular areas (Fig. 2 L), whereas it was negative in diffuse areas. Lastly, BCL2 BAP studies showed no rearrangement in follicular or diffuse areas, despite significant BCL2 expression as seen on immunohistochemistry in diffuse areas.
Three and 9 somatic mutations were identified in FL and DLBCL, respectively. The FL component showed the following mutations: ATM (V410A, variant allele frequency (VAF) 62%), DDX3X (Y243X, VAF 18%), and RAC1 (A159V, VAF 2%). The DDX3X mutation was designated as likely oncogenic due to the loss-of-function mechanism. The variants identified in DLBCL component included DDX3X (Y243X, VAF 62%), TP53 (R175H, VAF 66%), MYC (S21T, VAF 37%), KLHL6 (L58P, VAF 37%), RHOA (A61D, VAF 46%), APC (Q1444X, VAF 3%), KRAS (G60S, VAF 3%), NF1 (R2258X, VAF 3%), and NUP93 (Q15X, VAF 5%). DEAD-Box Helicase 3 X-Linked (DDX3X) Y243X was the only variant shared by FL and DLBCL and its VAF increased from 18 in FL to 62% in DLBCL component, respectively.
Heavy chain usage was determined separately for follicular and diffuse component and showed IGHV2-26, IGHD1-14, and IGHJ4 in FL, and IGHV3-43, IGHD6-13, and IGHJ4 in DLBCL. We also performed PCR with Biomed-2 primers on microdissected FL and DLBCL components. DLBCL showed 331 bp product in FR1 primer, and 248 bp and 266 bp products in FR2 primer. FL showed a polyclonal pattern in all three primer sets.
Discussion
EBV-associated lymphoproliferative disorders are a heterogeneous group of diseases including indolent conditions and highly aggressive lymphomas with variable presence of EBV and diverse latency types [6]. EBV-positive FL is not well characterized due to its low prevalence ranging from 2.5 to 6% of FL [1] [2] [3]. Nevertheless, it is known that the majority of cases are of histologic grade 3 and/or show progression to high-grade lymphoma. Mackrides et al. reported the two largest series (10 and 12 cases, respectively) of EBV-positive FL and identified several cases, which progressed to DLBCL or higher-grade FL within 1 to 6 years, although detailed pre- and post-transformation EBV data were not available in the majority of cases [3] (Table 1). EBV-positive FL is usually of latency type II [3] [2]. In the current case, the follicular areas were positive for EBER, and negative for EBNA1 and LMP1 by immunohistochemistry, indicating latency type 0. However, EBNA1 expression in type I or type II latency driven by viral promoter (qp) [10] [11] can be weak [12], and antibodies for EBNA1 may show poor sensitivity [13]. Therefore, we cannot exclude the possibility of type I latency in our case. This latency type has been rarely documented in EBV-positive FL [2] [3].
The IHC showed neoplastic follicles as well as diffuse areas to be positive for CD10, BCL6, and HGAL but negative for LMO2 and MUM1, which is consistent with germinal center cell phenotype [14] [15]. Of note, loss of LMO2 expression in germinal center cell-derived lymphomas was previously reported in lymphomas with MYC rearrangement [3, 16]. The BCL2 rearrangement was not detected in this case, despite significant BCL2 expression seen by IHC primarily in diffuse areas. The expression of BCL2 in neoplastic follicles was focal. Previously, Mackrides et al. reported a case of EBV-positive FL with BCL2 protein expression without a BCL2 rearrangement [3]. In BCL2/IGH-negative FL, mechanisms of BCL2 overexpression are largely unknown except for a few previously reported cases with BCL2 locus amplification and trisomy 18 [17, 18]. Although there are differences in gene expression and genetic alterations between BCL2 rearrangement-negative and -positive FL, prognosis and survival do not differ [17] [19] [20] [21].
Similar to the patient described in this report, Mackrides et al. reported a case of EBV-positive BCL2 rearrangement-negative FL with a history of hydroxychloroquine treatment (Table 1) [3]. Although the correlation between medication history and pathophysiology of BCL2 rearrangement-negative EBV-positive FL has not been explored, rare cases of iatrogenic immunodeficiency-associated lymphoproliferative disorders have been reported in association with hydroxychloroquine, and therefore, this case may be considered iatrogenic immunodeficiency-associated disorder [22] [23]. Hydroxychloroquine is an immunomodulatory drug used to treat malaria and autoimmune diseases [24]. It has been shown to reduce TNF alpha and IFN alpha production in plasmacytoid dendritic cells upon Toll-like receptor 9 stimulation and to decrease TNF alpha and IL-6 production in B-cells [25] [26]. Interestingly, chloroquine, which hydroxychloroquine is derived from, reportedly causes phosphorylation of KAP1 through ATM, leading to reactivation of EBV lytic replication [27]. Murata et al. suggested that EBV-infected cells in lytic cycle reactivation might experience genomic instability through activation of recombination or repair genes, and therefore, we speculate that hydroxychloroquine might introduce genomic instability and promote growth in EBV-positive germinal center cells [28] [29].
Constitutive MYC expression has also been shown to induce genomic instability in vitro and in vivo [30]. MYC gene rearrangement can be found in 8–38% of FL at the time of transformation [5, 31] [32]. When MYC rearrangement is observed in low-grade FL cases with BCL2 rearrangement, it may be associated with an aggressive disease course [32]. However, Yoshida et al. reported a case of FL without BCL2/IGH translocation with MYC rearrangement and with an indolent clinical course [33]. In the current report, BCL2 BAP FISH was negative in follicular and diffuse areas, and both follicular and diffuse areas showed MYC/IGH rearrangement. The patient responded well to chemotherapy and radiation, and remains in complete remission.
The prevalence of MYC/IGH rearrangement in EBV-associated B-cell lymphoproliferative disorders is variable. The highest frequency of MYC rearrangement is observed in Burkitt lymphoma and EBV-positive plasmablastic lymphoma, 80–90% and 74% of cases, respectively [6] [34] [35]. Both entities show EBV latency type I, similar to the reported case [6].
Next-generation sequencing provided further insights into the pathogenesis of the reported case. Mutations typically found in EBV-negative FL were not seen. Instead, the follicular component harbored mutations in ATM, DDX3X, and RAC1 genes. ATM is a master regulator of cell cycle checkpoint signaling pathways required for a response to DNA damage and maintaining genome stability. In small cell lymphoma, ATM mutations are frequent in mantle cell lymphomas and are not common in FL. In addition, even in mantle cell lymphoma cases, ATM mutation does not affect patients’ prognosis [36], suggesting that ATM mutation might not have impact on tumor progression and eventually ATM-positive clones may disappear during clonal evolution. DDX3X is the fourth most commonly mutated gene in Burkitt lymphoma and is considered to be a driver mutation in EBV-positive lymphomas [37, 38]. Moreover, a loss of function DDX3X mutations is enriched in MYC-rearranged DLBCL with functional cooperation between mutated DDX3X and MYC [39]. Not surprisingly, in addition to shared DDX3X mutation and MYC/IGH rearrangement, DLBCL show a number of private mutations affecting DNA damage response, cell cycle progression, B-cell receptor signaling, and apoptosis. Mutations of TP53, MYC, KLHL6, and RHOA genes were found to have the highest VAF, whereas other mutations represented minor variants. Such intratumoral heterogeneity has been previously described in transformed FL [40].
The presence of shared genetic abnormalities and private mutations supports a branched evolution model previously reported for the majority of transformed FL cases (Fig. 3) [5]. It is conceivable that EBV infection/reactivation was an initiating event and occurred in the context of hydroxychloroquine immunomodulation [41] [4] [42]. Since FL and DLBCL components harbored DDX3X mutation and MYC/IGH rearrangement, it is plausible that the EBV-infected common progenitor acquired DDX3X mutation and MYC/IGH as initial steps in lymphomagenesis [40]. In addition, DLBCL acquired numerous mutations promoting tumor growth and at this point EBV became dispensible, which would explain a loss of EBV in diffuse areas consistent with “hit-and-run” hypothesis [43] [44]. The unique IGH VDJ usage seen in FL and DLBCL is unusual and may represent two distinct amplified alleles. As an alternative, DDX3X mutations could have occurred later in the development of FL and DLBCL, and/or EBV infection/reactivation occurred independently as well, only in the FL branch.
Proposed branched evolution in the reported case of EBV-positive FL and concurrent EBV-negative DLBCL based on contributions of MYC, EBV, and shared and unique mutations to lymphomagenesis. EBV infection/reactivation and DDX3X mutation are shown in parenthesis at the level of common lymphoid progenitor as it is also plausible that these events occurred later in the evolution of the disease
Interestingly, recent phylogenetic analysis of B-cell receptor subclones in FL at single cell level demonstrated clonal heterogeneity in a subset of FL cases [45]. Such single-cell sequencing, if feasible, would have allowed for further clarification of clonal evolution and the role of individual mutations in our case.
In summary, we describe a case of EBV-positive FL and concurrent EBV-negative DLBCL. We speculate that hydroxychloroquine-induced immunomodulation may have led to EBV infection/reactivation as an initiating event. The presence of molecular genetic abnormalities shared by FL and DLBCL point to a common progenitor. The presence of mutations unique to FL and DLBCL and distinctive IGH usage support the branched clonal evolution model.
References
Mundo L et al (2019) EBV leaves its mark: new evidence of <<hit and run>> hypothesis in B-cell lymphomas from non-conventional methods. Hematol Oncol 37(S2):529
Mackrides N et al (2019) Prevalence, clinical characteristics and prognosis of EBV-positive follicular lymphoma. Am J Hematol 94(2):E62–E64. https://doi.org/10.1002/ajh.25357
Mackrides N et al (2017) Epstein-Barr virus-positive follicular lymphoma. Mod Pathol 30(4):519–529. https://doi.org/10.1038/modpathol.2016.214
Granai M et al (2019) Role of Epstein-Barr virus in transformation of follicular lymphoma to diffuse large B-cell lymphoma: a case report and review of the literature. Haematologica 104(6):e269–e273. https://doi.org/10.3324/haematol.2018.215053
Pasqualucci L et al (2014) Genetics of follicular lymphoma transformation. Cell Rep 6(1):130–140. https://doi.org/10.1016/j.celrep.2013.12.027
Rezk SA, Weiss LM (2019) EBV-associated lymphoproliferative disorders: update in classification. Surg Pathol Clin 12(3):745–770. https://doi.org/10.1016/j.path.2019.03.002
Romero-Masters JC et al (2018) An EBNA3C-deleted Epstein-Barr virus (EBV) mutant causes B-cell lymphomas with delayed onset in a cord blood-humanized mouse model. PLoS Pathog 14(8):e1007221. https://doi.org/10.1371/journal.ppat.1007221
Pophali PA et al (2020) High level MYC amplification in B-cell lymphomas: is it a marker of aggressive disease? Blood Cancer J 10(1):5. https://doi.org/10.1038/s41408-019-0271-z
van Dongen JJ et al (2003) Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17(12):2257–2317. https://doi.org/10.1038/sj.leu.2403202
Ok CY et al (2013) EBV-positive diffuse large B-cell lymphoma of the elderly. Blood 122(3):328–340. https://doi.org/10.1182/blood-2013-03-489708
Fields, B.N., D.M. Knipe, and P.M. Howley, Fields virology. 6th ed. 2013, Philadelphia: Wolters Kluwer Health/Lippincott Williams & Wilkins. 2 v. (xx, 2456, I-82 p.).
Grasser FA et al (1994) Monoclonal antibodies directed against the Epstein-Barr virus-encoded nuclear antigen 1 (EBNA1): immunohistologic detection of EBNA1 in the malignant cells of Hodgkin’s disease. Blood 84(11):3792–3798
Wilson, J.B., et al., EBNA1: oncogenic activity, immune evasion and biochemical functions provide targets for novel therapeutic strategies against Epstein-Barr virus- associated cancers. Cancers (Basel), 2018. 10(4) https://doi.org/10.3390/cancers10040109.
Natkunam Y et al (2005) Expression of the human germinal center-associated lymphoma (HGAL) protein, a new marker of germinal center B-cell derivation. Blood 105(10):3979–3986. https://doi.org/10.1182/blood-2004-08-3112
Natkunam Y et al (2001) Analysis of MUM1/IRF4 protein expression using tissue microarrays and immunohistochemistry. Mod Pathol 14(7):686–694. https://doi.org/10.1038/modpathol.3880373
Colomo L et al (2017) LMO2-negative expression predicts the presence of MYC translocations in aggressive B-cell lymphomas. Am J Surg Pathol 41(7):877–886. https://doi.org/10.1097/PAS.0000000000000839
Masir N et al (2012) Variation in BCL2 protein expression in follicular lymphomas without t(14;18) chromosomal translocations. Pathology 44(3):228–233. https://doi.org/10.1097/PAT.0b013e3283513fb2
Horsman DE et al (2003) Follicular lymphoma lacking the t(14;18)(q32;q21): identification of two disease subtypes. Br J Haematol 120(3):424–433. https://doi.org/10.1046/j.1365-2141.2003.04086.x
Leich E et al (2009) Follicular lymphomas with and without translocation t(14;18) differ in gene expression profiles and genetic alterations. Blood 114(4):826–834. https://doi.org/10.1182/blood-2009-01-198580
Leich E et al (2016) Similar clinical features in follicular lymphomas with and without breaks in the BCL2 locus. Leukemia 30(4):854–860. https://doi.org/10.1038/leu.2015.330
Zamo A et al (2018) Differences between BCL2-break positive and negative follicular lymphoma unraveled by whole-exome sequencing. Leukemia 32(3):685–693. https://doi.org/10.1038/leu.2017.270
Swerdlow, S., et al., WHO classification of tumours of haematopoietic and lymphoid tissues. 2017.
Marcelis L et al (2018) Other immunomodulatory agent-related lymphoproliferative diseases: a single-center series of 72 biopsy-confirmed cases. Mod Pathol 31(9):1457–1469. https://doi.org/10.1038/s41379-018-0054-2
Ben-Zvi I et al (2012) Hydroxychloroquine: from malaria to autoimmunity. Clin Rev Allergy Immunol 42(2):145–153. https://doi.org/10.1007/s12016-010-8243-x
Sacre K, Criswell LA, McCune JM (2012) Hydroxychloroquine is associated with impaired interferon-alpha and tumor necrosis factor-alpha production by plasmacytoid dendritic cells in systemic lupus erythematosus. Arthritis Res Ther 14(3):R155. https://doi.org/10.1186/ar3895
Torigoe M et al (2018) Hydroxychloroquine efficiently suppresses inflammatory responses of human class-switched memory B cells via Toll-like receptor 9 inhibition. Clin Immunol 195:1–7. https://doi.org/10.1016/j.clim.2018.07.003
Li X, Burton EM, Bhaduri-McIntosh S (2017) Chloroquine triggers Epstein-Barr virus replication through phosphorylation of KAP1/TRIM28 in Burkitt lymphoma cells. PLoS Pathog 13(3):e1006249. https://doi.org/10.1371/journal.ppat.1006249
Murata T (2014) Regulation of Epstein-Barr virus reactivation from latency. Microbiol Immunol 58(6):307–317. https://doi.org/10.1111/1348-0421.12155
Roughan JE, Thorley-Lawson DA (2009) The intersection of Epstein-Barr virus with the germinal center. J Virol 83(8):3968–3976. https://doi.org/10.1128/JVI.02609-08
Kuzyk, A. and S. Mai, c-MYC-induced genomic instability. Cold Spring Harbor Perspectives in Medicine, 2014. 4(4): ARTN a014373. https://doi.org/10.1101/cshperspect.a014373.
Aukema SM et al (2017) MYC expression and translocation analyses in low-grade and transformed follicular lymphoma. Histopathology 71(6):960–971. https://doi.org/10.1111/his.13316
Yano T et al (1992) MYC rearrangements in histologically progressed follicular lymphomas. Blood 80(3):758–767
Yoshida M et al (2015) Clinicopathological features of double-hit B-cell lymphomas with MYC and BCL2, BCL6 or CCND1 rearrangements. Pathol Int 65(10):519–527. https://doi.org/10.1111/pin.12335
Valera A et al (2010) IG/MYC rearrangements are the main cytogenetic alteration in plasmablastic lymphomas. Am J Surg Pathol 34(11):1686–1694. https://doi.org/10.1097/PAS.0b013e3181f3e29f
Allday MJ (2009) How does Epstein-Barr virus (EBV) complement the activation of Myc in the pathogenesis of Burkitt’s lymphoma? Semin Cancer Biol 19(6):366–376. https://doi.org/10.1016/j.semcancer.2009.07.007
Fang NY et al (2003) Oligonucleotide microarrays demonstrate the highest frequency of ATM mutations in the mantle cell subtype of lymphoma. Proc Natl Acad Sci U S A 100(9):5372–5377. https://doi.org/10.1073/pnas.0831102100
Schmitz R et al (2012) Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature 490(7418):116–120. https://doi.org/10.1038/nature11378
Okuno Y et al (2019) Defective Epstein-Barr virus in chronic active infection and haematological malignancy. Nat Microbiol 4(3):404–413. https://doi.org/10.1038/s41564-018-0334-0
Gong, C., et al., Sequential inverse dysregulation of the RNA helicases DDX3X and DDX3Y facilitates MYC-driven lymphomagenesis. Mol Cell, 2021. 81(19): p. 4059–4075 e11 https://doi.org/10.1016/j.molcel.2021.07.041.
Okosun J, Montoto S, Fitzgibbon J (2016) The routes for transformation of follicular lymphoma. Curr Opin Hematol 23(4):385–391. https://doi.org/10.1097/MOH.0000000000000255
Lieber MR (2016) Mechanisms of human lymphoid chromosomal translocations. Nat Rev Cancer 16(6):387–398. https://doi.org/10.1038/nrc.2016.40
Magrath I (2012) Epidemiology: clues to the pathogenesis of Burkitt lymphoma. Br J Haematol 156(6):744–756. https://doi.org/10.1111/j.1365-2141.2011.09013.x
Mundo L et al (2020) Frequent traces of EBV infection in Hodgkin and non-Hodgkin lymphomas classified as EBV-negative by routine methods: expanding the landscape of EBV-related lymphomas. Mod Pathol. https://doi.org/10.1038/s41379-020-0575-3
Niller HH, Wolf H, Minarovits J (2011) Viral hit and run-oncogenesis: genetic and epigenetic scenarios. Cancer Lett 305(2):200–217. https://doi.org/10.1016/j.canlet.2010.08.007
Haebe S et al (2021) Single-cell analysis can define distinct evolution of tumor sites in follicular lymphoma. Blood 137(21):2869–2880. https://doi.org/10.1182/blood.2020009855
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kitagawa, Y., Peterson, J.F., Kenney, S. et al. EBV-positive follicular lymphoma and concurrent EBV-negative diffuse large B-cell lymphoma illustrating branched evolution model and “Hit and Run” hypothesis. J Hematopathol 15, 157–167 (2022). https://doi.org/10.1007/s12308-022-00502-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12308-022-00502-x