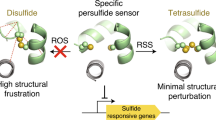
Abstract
Oxidative stresses, such as reactive oxygen species, reactive electrophilic species, reactive nitrogen species, and reactive chlorine species, can damage cellular components, leading to cellular malfunction and death. In response to oxidative stress, bacteria have evolved redox-responsive sensors that enable them to simultaneously monitor and eradicate potential oxidative stress. Specifically, redox-sensing transcription regulators react to oxidative stress by means of modifying the thiol groups of cysteine residues, functioning as part of an efficient survival mechanism for many bacteria. In general, oxidative molecules can induce changes in the three-dimensional structures of redox sensors, which, in turn, affects the transcription of specific genes in detoxification pathways and defense mechanisms. Moreover, pathogenic bacteria utilize these redox sensors for adaptation and to evade subsequent oxidative attacks from host immune defense. For this reason, the redox sensors of pathogenic bacteria are potential antibiotic targets. Understanding the regulatory mechanisms of thiol-based redox sensors in bacteria will provide insight and knowledge into the discovery of new antibiotics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oxidative stresses, such as reactive oxygen species (ROS), reactive electrophilic species (RES), reactive nitrogen species (RNS), and reactive chlorine species (RCS), detrimentally affect bacteria by influencing or depleting cellular thiol pools (Antelmann and Helmann 2011; Ortiz de Orué Lucana et al. 2012; Imlay 2013). As a defensive measure, bacteria have evolved a number of sensory and regulatory systems to eradicate oxidative stress or repair the damage caused to protect cellular components (Antelmann and Helmann 2011; Ortiz de Orué Lucana et al. 2012; Imlay 2013). Specifically, bacteria use redox-sensitive transcription factors to sense and respond to oxidative stress. These proteins sense oxidative stress via modifying the thiol groups of cysteine residues into reversible inter-/intra-molecular disulfides or into irreversible sulfinic acid, sulfonic acid, or thiol-(S)-alkylated forms (Chi et al. 2010; Barford 2004; Giles et al. 2003). Such thiol group modifications are observed in most life forms and ultimately result in changes in protein structure, leading to direct binding or dissociation of the transcription factors from the region of DNA to which they bind (Chi et al. 2010; Ji et al. 2013; Palm et al. 2012; Chen et al. 2010; Newberry et al. 2007; Antelmann and Helmann 2011; Lee et al. 2016).
Several representative bacterial thiol-based redox-sensing proteins that directly regulate gene expression are known, including OxyR, OhrR, Spx, YodB, CrtJ, and CprK. The OxyR protein, which belongs to the LysR family, senses ROS or RNS. The OxyR tetramer has been shown to increase the gene expression of antioxidant proteins in response to oxidative stress by interaction with RNA polymerase (Antelmann and Helmann 2011; Lee et al. 2004). The OhrR protein, which belongs to the MarR family, is a dimeric regulatory protein that senses organic peroxides and other ROS (Fuangthong et al. 2001; Sukchawalit et al. 2001). OhrR proteins are known to function as either one- or two-Cys OhrR proteins that vary in their redox-sensing mechanisms. For instance, the OhrR from Xanthomonas campestris has two redox-active cysteine residues that form an intermolecular disulfide bond, whereas the OhrR from B. subtilis has one redox-active cysteine residue that can be S-cysteinylated (Newberry et al. 2007; Fuangthong et al. 2001; Sukchawalit et al. 2001; Hong et al. 2005). The Spx protein behaves like a thiol-based activator in response to electrophiles, such as diamide and quinone (Leelakriangsak et al. 2007). The Spx protein controls several regulons that contain the genes needed to maintain the thiol-redox balance within cells and utilizes organosulfur compounds (Zuber 2004). The expression of Spx is induced when the peroxide regulon repressor PerR or the MarR/DUF24 family regulator YodB is oxidized (Antelmann and Helmann 2011; Leelakriangsak et al. 2007). The CrtJ protein from Rhodobacter capsulatus acts as a homotetramer that regulates the expression of photosynthesis and heme biosynthesis genes in response to oxygen and light (Elsen et al. 2005; Masuda et al. 2002). Finally, the CprK protein belongs to the CRP–FNR family and is involved in the regulation of halorespiration in Desulfitobacterium species (Pop et al. 2006). Redox regulation can also be achieved indirectly, including via RsrA anti-σ factor and RegB sensory histidine kinase (Antelmann and Helmann 2011).
Thiol-based redox-sensing proteins from pathogenic bacteria are known to play central roles during infection. Some representative examples are Salmonella typhimurium DksA (Crawford et al. 2016), Listeria monocytogenes SpxA1 (Whiteley et al. 2017), and Staphylococcus aureus QsrR (Ji et al. 2013), which are essential for bacterial pathogenesis and growth during infection. Here, we show representative examples of thiol-based redox-sensing proteins and explore their biological mechanisms at the molecular level. Because an oxidative signaling molecule can initiate specific structural changes in thiol-based redox-sensing proteins, structural comparisons of the various states of the related protein structures and related changes in transcriptional patterns can aid our understanding of the biological function of thiol-based redox-sensing proteins. Moreover, thiol-based redox-sensing proteins are also common in eukaryotes. Therefore, studies on prokaryotic redox-sensing proteins can provide greater insight into understanding human diseases related to redox regulation (Le Rossignol et al. 2017).
Modification of cysteine thiols by oxidative molecules
Because cysteine residues can be oxidized into different redox states and can also sense a range of oxidative signals, most redox signals are recognized by cysteine residues, whereas minor signals are recognized by iron cluster centers (Antelmann and Helmann 2011; Green and Paget 2004). In a highly reduced and low-oxygen environment, cysteine residues in cytoplasmic proteins are generally reduced (Sporer et al. 2017). When the levels of oxidative molecules, such as ROS, RES, RNS, or RCS, in the environment are high, cysteine thiolates can be oxidized and transitioned into more oxidized forms (Antelmann and Helmann 2011; Gray et al. 2013). Specifically, cysteine thiols can be oxidized into cysteine sulfenic acids (R-SOH) or can irreversibly be oxidized into other forms, including cysteine sulfinic acids (R-SO2H) and sulfonic acids (R-SO3H) (Antelmann and Helmann 2011; Hillion and Antelmann 2015). The sulfenic acids can form mixed disulfides with low-molecular weight (LMW) thiols, such as glutathione (GSH), mycothiol (MSH), and bacillithiol (BSH). More importantly, two adjacent cysteine sulfenic acids can reversibly form intramolecular/intermolecular disulfide bonds (Fig. 1a). These chemical modifications to cysteine residues allow redox sensor proteins to adopt specific conformations to modulate biological function by participating in transcriptional regulation of the expression of redox balance-related genes. ROS include superoxide anion (O ·−2 ), hydrogen peroxide (H2O2), and the highly reactive hydroxyl radical (OH·), which is produced by the one-electron reduction of molecular oxygen (O2) during aerobic respiration (Hillion and Antelmann 2015; Imlay 2008). RES include quinones, aldehydes, epoxides, diamide, and α,β-unsaturated carbonyl compounds (Hillion and Antelmann 2015). Generally, RES are secondary reactive intermediates from oxidation products of amino acids, lipids or carbohydrates (Hillion and Antelmann 2015). Because RES possesses electron-deficient centers, they can react with nucleophilic cysteine thiols, including thiol-containing proteins and LMW thiols, via thiol-S-alkylation chemistry (Antelmann and Helmann 2011) (Fig. 1b). For example, quinones are known to promote irreversible thiol-S-alkylation and protein aggregation, depleting protein thiols in the proteome in vivo (Loi et al. 2015). In addition, nitric oxide (·NO) is a main source of other RNS and is generated by the oxidation of l-arginine to l-citrulline via the inducible nitric oxide synthase (iNOS) in neutrophils (Forstermann and Sessa 2012). RNS include nitric oxide (·NO) and peroxynitrite (NO3−), which directly or indirectly form S-nitrosothiol (RS-NOs) and S-nitrothiol (R-SNO2), respectively (Antelmann and Helmann 2011) (Fig. 1c). In addition, the direct reaction of nitric oxide with LMW thiols leads to the formation of S-nitrothiol (Antelmann and Helmann 2011; Hillion and Antelmann 2015). Peroxynitrite, which is formed by the reaction of nitric oxide and superoxide anion, is a strong oxidative molecule that can cause cell damage (Nathan 2003; Nathan and Xie 1994). RCS, such as hypochlorous acid (HOCl) and chloramines, can lead to the oxidation of cysteine or methionine residues in proteins as well as to the breakage of nucleic acids and lipid peroxidation (Gray et al. 2013) (Fig. 1d). Therefore, in response to multiple RCS attacks from host cells, bacteria use various defense systems, such as detoxifying enzymes (catalases, peroxidases, and reductases), LMW thiols, and antioxidants, to survive. In addition, ROS and RNS are also used for antimicrobial defense, which involves activated neutrophils to remove pathogenic bacteria from the body (Hillion and Antelmann 2015; Loi et al. 2015; Ezraty et al. 2017).
Modification of cysteine thiols by oxidative species. When the levels of oxidative molecules, such as ROS, RES, RNS, or RCS, in the environment are high, cysteine thiolates are oxidized and transition into more oxidized forms. a Cysteine thiols can be reversibly oxidized into the cysteine sulfenic acids (R-SOH) or further oxidized into irreversible products, such as cysteine sulfinic acids (R-SO2H) and sulfonic acids (R-SO3H). Sulfenic acid can form a mixed disulfide with LMW thiols (R′-SH), such as glutathione (GSH), mycothiol (MSH), and bacillithiol (BSH). In the presence of adjacent sulfenic acids, cysteine thiols can reversibly form intramolecular/intermolecular disulfide. b RES with electrophilic carbon centers (δ +), such as benzoquinone and diamide, can react with the nucleophilic cysteine thiols via thiol-S-alkylation chemistry. Quinones promote irreversible thiol-S-alkylation, resulting in quinone-S-adducts, whereas the electrophilic diamide leads to the formation of disulfide bonds between cysteine residues. c RNS, including nitric oxide (·NO) and peroxynitrite (NO3−), modify thiols by forming reversible S-nitrosothiols (R-SNOs) and S-nitrothiols (R-SNO2), respectively. d RCS, such as hypochlorous acid (HOCl), lead to the chlorination of protein thiols, resulting in the production of sulfenylchloride intermediates (R-SCl) that form inter- or intramolecular disulfides via subsequent reactions. In the absence of proximal thiols, chlorinated thiols are overoxidized to cysteine sulfinic and sulfonic acids
Prokaryotic thiol-based redox sensors
Pathogenic bacteria have evolved redox-responsive sensors that enable them to monitor oxidative signals and survive in harsh environments even within a host species. Increases in specific oxidative signals are recognized by redox sensors, which trigger the appropriate cellular responses. Many redox sensors can recognize specific oxidative signals, binding to different oxidative signals and differentiating each of them. These dissimilar signals change the three-dimensional structures of redox sensors and affect the transcription of specific genes that encode corresponding detoxification pathways and defense mechanisms. In the following section, we will discuss the mechanisms of redox-responsive transcriptional factors from various bacteria in detail.
OxyR
OxyR is the LysR-type transcriptional regulator (LTTR) that senses cellular peroxide (H2O2) levels. OxyR is inactive under low levels of peroxide but is converted to an active form by an increase in peroxide. Large amounts of peroxide cause OxyR to form disulfide bonds and activate the transcription of antioxidant defense genes (Seaver and Imlay 2001; Aslund et al. 1999; Imlay 2008). OxyR, comprised of 305 amino acid residues, is composed of an N-terminal helix-loop-helix DNA binding motif and a C-terminal regulatory domain with two key cysteine residues (Cys199 and Cys208) (Choi et al. 2001) (Fig. 2). When peroxide is present, Cys199 in OxyR is oxidized to sulfenic acid by the bound peroxide in a specific manner, which is accompanied by the deprotonation of Cys199 and the donation of a proton to the peroxide molecule (Jo et al. 2015). The sulfenic acid formed on Cys199 forms an intramolecular disulfide bond with Cys208 (Jo et al. 2015; Choi et al. 2001) (Fig. 2b). The formation of a disulfide bond between Cys199 and Cys208 allows the following large structural rearrangements in the C-terminal domain: the loop spanning residues 199–206 moves, Cys199 flips out, and new secondary structural elements are formed (a beta strand appears along residues 212–217, and a pseudo-helical loop forms at residues 219–222). The structural changes in OxyR caused by peroxide lead to the OxyR protein activating the transcription of specific genes in the oxyR regulon (Zheng et al. 2001). These genes are related to the detoxification of hydrogen peroxide and include katG (hydroperoxidase), ahpCF (alkyl hydroperoxide reductase), oxyS (a regulatory RNA), dps (a non-specific DNA-binding protein), fur (ferric uptake regulation), gorA (glutathione reductase), and grxA (glutaredoxin) (Zheng et al. 2001). Furthermore, peroxide-activated OxyR induces the cooperative binding of RNA polymerase to activate and enhance transcription (Choi et al. 2001). In addition, oxidized OxyR binds to four DNA pairs, whereas the reduced OxyR binds only two pairs of DNA (Choi et al. 2001). The differential DNA-binding ability of the oxidized and reduced OxyR can be explained by the dissimilar dimeric interfaces of the two OxyR structures that result from different disulfide bond configurations. Ultimately, the different DNA-binding activities affects gene-regulation behaviors depending on the redox state of OxyR (Jo et al. 2015). After successful defense against oxidative stress, the disulfide bond between Cys199 and Cys208 in OxyR is reduced by glutaredoxin 1 (Grx1), and the related transcription is deactivated (Zheng et al. 1998). In addition to peroxide, OxyR is known to recognize S-nitrothiol to protect cells from various RNS attacks (Mukhopadhyay et al. 2004). To understand the exact mechanism by which OxyR recognizes and discriminates these two kinds of oxidative signals, further structural studies on OxyR are needed. Structural information on OxyR in various states has been published since the first structures from Escherichia coli were reported [PDB codes 1I69 (reduced form) and 1I6A (oxidized form)] (Choi et al. 2001). The structures available are Porphyromonas gingivalis OxyR [PDB codes 3HO7, 3T22, and 3UKI (reduced form)] (Svintradze et al. 2013), Neisseria meningitides OxyR [PDB code 3JV9 (reduced form)] (Sainsbury et al. 2010), Pseudomonas aeruginosa OxyR [PDB codes 4X6G, 4XWS, and 4Y0 M (reduced form)] (Jo et al. 2015), and the regulatory domains of Vibrio vulnificus OxyR2 [PDB codes 5X0Q (Cl-bound), 5B70, 5B7D (sulfate-bound), and 5X0V (reduced form)] (Jo et al. 2017).
Structures of the reduced and oxidized forms of OxyR, a sensor of peroxide stress. Two forms [reduced (gray) and oxidized (magenta)] are superimposed and depicted as ribbon diagrams. In all labeled residues, redox-sensing cysteines and mutants are represented as ball-and-stick models, with carbon, oxygen, and sulfur atoms colored green, red and yellow, respectively. The disulfide bridge is represented by yellow sticks. To crystallize the reduced OxyR, cysteine residue 199 was replaced with a serine residue. In the oxidized OxyR, Cys199 and Cys208 residues have been shown to form an intramolecular disulfide bond, leading to large structural changes in the loops [the corresponding residues (Cys199–Gly206, Gly212–Asp217, and Phe219–Thr222) are colored red]
OhrR, a MarR-family redox sensor
The organic hydroperoxide resistance protein regulator (OhrR) is a transcription factor that regulates the expression of the organic hydroperoxide resistance (ohr) gene in response to lipid hydroperoxides (Hong et al. 2005; Newberry et al. 2007). OhrR belongs to the multiple antibiotic resistance regulatory (MarR) family that controls the expression of toxin-resistance genes related to antibiotics, organic solvents, detergents, and oxidative stress signals (Hong et al. 2005; Newberry et al. 2007). OhrR proteins are classified into two groups based on the presence of additional cysteine residue(s) at the C-terminus. In general, one cysteine residue exists at the N-terminus (Newberry et al. 2007). Another cysteine residue is located at the C-terminus (two-Cys type), although some OhrR proteins do not have cysteines on their C-termini (one-Cys type). For instance, B. subtilis OhrR and X. campestris OhrR are homodimeric proteins comprising 147 and 153 amino acids, respectively (Fig. 3). The one-Cys type, including B. subtilis OhrR, senses hydroperoxides and forms a reversible sulphenic acid derivative with bacillithiol (Fuangthong and Helmann 2002). The oxidation of the conserved Cys15 of B. subtilis OhrR inhibits DNA-binding affinity, thereby inducing the expression of the organic hydroperoxidase gene ohrA (Fuangthong and Helmann 2002). In addition, a cyclic sulfenamide can also be formed by condensation of the sulfenate with a neighboring backbone amide nitrogen in B. subtilis OhrR (Antelmann and Helmann 2011). Unlike the one-Cys type, the two-Cys type, including X. campestris OhrR, forms intermolecular disulfide bonds between C22 and C127′ (or C22′ and C127) in two monomers in response to oxidative signals (Fig. 3c, d). Currently, a few OhrR structures have been published from two bacterial sources: B. subtilis [PDB codes 1Z91 (reduced form) and 1Z9C (DNA-bound form)] (Hong et al. 2005) and Xanthamonas campestris [PDB codes 2PFB (oxidized form) and 2PEX (reduced form)] (Newberry et al. 2007).
Structures of the reduced and oxidized forms of OhrR classified into two groups based on the presence of cysteine residues at the C-terminus. Dimeric structures of OhrR from Bacillus subtilis (a, b) and Xanthamonas campestris (c, d) are depicted based on their conformational changes. The monomeric and dimeric structures are presented in different colors (gray and blue). All labeled residues are represented as ball-and-stick models with carbon (green), oxygen (red) and sulfur (yellow) atoms. In the structure shown in (a), Cys15 (or Cys15′) was mutated to serine to facilitate crystallization of the reduced form of B. subtilis OhrR. In (b), the ohrA operator is represented by a wheat ribbon diagram. Cys15 was also replaced with serine to mimic the DNA-binding affinity of the reduced wild-type OhrR. In the reduced form (c) of X. campestris OhrR, Cys22 was substituted with serine to produce a conformation identical to the reduced wild-type form. In the oxidized form (d), Cys22 and Cys127′ (or Cys22′ and Cys127) form intermolecular disulfide bonds between the two subunits
YodB, QsrR, and HypR (redox sensors in the MarR/DUF24-family)
In the MarR/DUF24-family, three distinct redox sensors are recognized: YodB, QsrR, and HypR. Bacillus subtilis YodB is a prototypical MarR/DUF24 family transcriptional regulator composed of 112 amino acid residues. YodB senses and responds to RES, such as diamide, quinones, and aldehydes, in a specific manner (Antelmann and Helmann 2011). Unlike common redox sensors that sense only one kind of oxidative molecule, YodB possesses an elaborate molecular mechanism that discriminates between diamide and quinone by structural changes in the YodB dimers (Lee et al. 2016). More specifically, diamide induces intersubunit disulfide formation between Cys6 and Cys101′ (or Cys6′ and Cys101) in the YodB dimer (Fig. 4a). Disulfide formation between two YodB monomers induces large movements of secondary structural elements, including two DNA-recognition helices (α4 and α4′) and full dissociation of YodB from the operator DNA (Lee et al. 2016). In contrast to the diamide-mediated signaling pathway, YodB responds to the more toxic compound methyl-p-benzoquinone (MPBQ) via the S-alkylation of Cys6 (or Cys6′), which does not induce a large conformational change in the YodB dimer (not shown in the figure) (Lee et al. 2016). However, the MPBQ adducts on Cys6 (or Cys6′) also affect the DNA-recognition helices, causing YodB to dissociate from the operator DNA (Lee et al. 2016). As a result, detoxifying genes, including azoR1 and azoR2, are expressed to neutralize oxidative molecules in B. subtilis (Antelmann and Helmann 2011). Interestingly, MPBQ induces expression of the azoR1 gene at a 1000-fold lower concentration than diamide, indicating that more toxic compounds are efficiently treated in a specific manner (Lee et al. 2016). Second, S. aureus QsrR, termed the quinone-sensing and response repressor, has a redox-sensing Cys5 that covalently binds to menadione (Ji et al. 2013) (Fig. 4b). Binding between menadione and Cys5 causes rearrangement of the QsrR dimer and disruption of the QsrR–DNA interaction (Ji et al. 2013). In the case of B. subtilis HypR (hypochlorite responsive regulator; previously known as YybR), two cysteine residues (Cys14 and Cys49) are essential for sensing diamide and NaOCl and thus for the activation of hypO transcription (Palm et al. 2012) (Fig. 4c). With an increase in oxidative molecules, two disulfide bonds between HypR monomers are formed, causing structural rearrangement of the HypR dimer and altering the DNA-binding affinity of HypR (Palm et al. 2012). A comparison of the dimeric structures of S. aureus QsrR (112 amino acid residues) and B. subtilis HypR (125 amino acid residues) shows that the two are nearly identical to B. subtilis YodB (reduced and MPBQ-bound forms) (Fig. 4). However, the DNA-binding affinities of S. aureus QsrR and B. subtilis HypR are quite different depending on their redox states. Currently, several structures for the MarR/DUF24 family can be found: B. subtilis YodB [PDB codes 5HS7 (reduced form), 5HS8 (diamide-treated form), and 5HS9 (quinone-bound form)] (Lee et al. 2016), Staphylococcus aureus QsrR [PDB codes 4HQE (the DNA-complex form) and 4HQM (menadione-complex form)] (Ji et al. 2013), and B. subtilis HypR [PDB codes 4A5N (reduced form) and 4A5M (oxidized form)] (Palm et al. 2012).
Structures of the reduced and oxidized forms of YodB, QsrR, and HypR, redox sensors in the MarR/DUF24 family. In the structures above, dimers of the reduced and oxidized forms of YodB (a), QsrR (b) and HypR (c) are superimposed using sequence alignment. Dimeric structures of all reduced and oxidized forms are shown as orange/gray and yellow/gray, respectively. Carbon atoms of key residues in both forms are colored green, and the oxygen and sulfur atoms are colored red and yellow, respectively. In the focused view (reduced and oxidized forms are shown on the left and right side, respectively), the important residues and their interactions are highlighted with black lines. The oxidized form of YodB is induced by diamide, Cys6 and Cys101′ (or Cys6′ and Cys101) to form inter-subunit disulfide bonds (yellow) in the YodB dimer, causing a large conformational change. In the reduced form of QsrR, the QsrR dimer is shown as a complex, with palindromic DNA represented by a wheat ribbon diagram. In the oxidized form of QsrR, Cys5 covalently binds to menadione, colored dark gray, which inhibits the DNA interaction. In the reduced form of HypR, to mimic the reduced form of wild-type HypR, Cys14 was replaced with serine. In the oxidized form (right panel in c), Cys14 and Cys49′ form an intermolecular disulfide bond (yellow) between two HypR monomers
Spx
The Spx protein is composed of 131 amino acids, acts as a monomer and belongs to the arsenate reductase (ArsC) family (Newberry et al. 2005). Spx is a global transcription factor that regulates the transcription of multiple genes via directly interacting with the C-terminal domain (CTD) of RNA polymerase α subunit in response to disulfide stress (Lamour et al. 2009; Nakano et al. 2010; Newberry et al. 2005) (Fig. 5). Oxidative molecules promote disulfide formation between Cys10 and Cys13 in Spx, allowing direct interaction with the CTD of the RNA polymerase α subunit (Fig. 5b). The complex activates the transcription of genes involved in thiol homeostasis, including thioredoxin (trxA) and thioredoxin reductase (trxB) (Nakano et al. 2010; Newberry et al. 2005). The disulfide bond between Cys10 and Cys13 possibly affects the rearrangement of the side chains of several basic residues that are related to DNA binding affinity (Nakano et al. 2010). In addition to positive regulation, Spx may play a negative role in transcriptional regulation (Nakano et al. 2010). Structural information about Spx in complex with the C-terminal domain of the RNA Polymerase α subunit from B. subtilis is available (PDB codes 1Z3E, 3GFK, and 3IHQ) (Lamour et al. 2009; Nakano et al. 2010; Newberry et al. 2005).
Structures of the reduced and oxidized forms of Spx in complex with the c-terminal domain of the RNA polymerase α subunit (αCTD). In both structures of reduced (a) and oxidized (b) Spx, αCTD and Spx are colored gray and green, respectively. All labeled residues are shown as a ball-and-stick with carbon (green), oxygen (red), nitrogen (blue) and sulfur (yellow) atoms. In the oxidized Spx-αCTD complex, Cys10 and Cys13 form an intramolecular disulfide bond. A sulfate ion is present and interacts with Ser12, which is adjacent to the disulfide bond of Cys10/Cys13, and Arg92. In the reduced form, Cys10 was mutated to a serine to force the reduced conformation. No sulfate ion is present, and the amine group of Arg92 is thus oriented away from its position in the sulfate-containing oxidized form
PrrA/PrrB (RegA/RegB), a two-component system
The PrrA/PrrB system (RegA/RegB in Rhodobacter capsulatus) is a bacterial two-component regulatory system (TCS) that is involved in anaerobic synthesis of the photosystem in Rhodobacter capsulatus (Elsen et al. 2004; Laguri et al. 2003). For Mycobacterium tuberculosis, the PrrA/PrrB system is active in response to host-cell interactions after infection, suggesting a possible new antibacterial target (Nowak et al. 2006). Generally, TCS consist of two proteins: a membrane-bound histidine kinase that senses a specific signal and a response regulator that regulates a set of genes (Nowak et al. 2006; Capra and Laub 2012). The signal-sensor kinases have a histidine-containing dimerization domain and a catalytic domain, whereas the response regulators have an N-terminal receiver domain and a DNA-binding domain (Capra and Laub 2012). Structural studies have been performed on the response regulator PrrA from R. sphaeroides [PDB code 1UMQ (effector domain); residues 125–184] and M. tuberculosis (PDB code 1YS7; 233 amino acid residues) (Laguri et al. 2003; Nowak et al. 2006). The M. tuberculosis PrrA complexed with Mg2+ is shown in Fig. 6. In the active site, one Mg2+ is coordinated by Asp15, Asp58, and Lys108. Ser86 is an important residue involved in signal transduction via PrrB. The structure of PrrA is very similar to those of response regulators belonging to the OmpR/PhoB family (DrrB and DrrD), including NarL, CheB, and RsbQ (Nowak et al. 2006).
Structure of PrrA in complex with magnesium(II) ion. PrrA is shown in yellow, with the active site residues in a ball-and-stick diagram comprising carbon (green), oxygen (red) and nitrogen (blue). The magnesium(II) ion is represented as a gray sphere. The Mg2+ ion is coordinated by Asp15, Asp58, Asn60, and Lys108. Ser86, one of the important residues for signal transduction by PrrB, is also shown
MerR family and NmlR-like redox sensors
Mercury resistance regulators (MerR) control metal ion resistance determinants and efflux pumps in response to a diverse range of effectors, such as redox stress molecules (by SoxR and NmlR), xenobiotics (by BmrR), and metal ion overload (by MerR, ZntR, and CueR) (Counago et al. 2016; Antelmann and Helmann 2011). Generally, MerR family members sense metal ions by coordinating with cysteine residues (Fig. 7). As a representative example, Sox possesses a [2Fe–2S] cluster that senses superoxide anions or RNS molecules by oxidation or nitrosylation, allowing the transcriptional activation of soxS (Counago et al. 2016; Watanabe et al. 2008). NmlR-like redox sensors belonging to the MerR family have been identified in pathogenic bacteria, such as Neisseria species, Haemophilus influenza, and Streptococcus pneumoniae (Antelmann and Helmann 2011). Their virulence is highly related to recognition and response to RES and NO molecules, which triggers the expression of genes that protect cells against RNS (nitrosative stress). In the case of H. influenza NmlR (PDB codes 5D8C, 5D90, and 5E01), two cysteine residues (Cys54 and Cys71) sense reactive species generated by the host in response to formaldehyde. Via oxidation, NmlR can control the expression of an operon consisting of adhC and estD, which are involved in the glutathione-dependent detoxification of toxic formaldehyde to formate (Counago et al. 2016).
Structures of the apo- and mercury(II)-bound forms of the MerR dimer. The structures of apo (a) and Hg2+ (b) MerR are shown as ribbon diagrams, and each subunit is colored gray or salmon. All the important cysteine residues are represented as ball-and-stick models with carbon (green) and sulfur (yellow) atoms and Hg2+ ions shown as a red sphere. To clearly understand the active sites, the overall structures of MerR (left) are highlighted in the enlarged views (right)
The protein structures and mechanisms of MerR/NmlR family members have been extensively studied as follows: E. coli SoxR [PDB codes 2ZHG (DNA-bound form) and 2ZHH] (Watanabe et al. 2008); Pseudomonas aeruginosa, a mercury(II)-dependent Tn501 MerR (PDB code 5CRL) (Wang et al. 2016); Bacillus megaterium MerR [PDB codes 4UA1 (Hg2+-bound form) and 4UA2] (Chang et al. 2015); B. subtilis transcription activator MtaN (PDB code 1R8D) and multidrug-efflux transporter regulator BmrR (PDB codes 1R8E, 1EXI, and 1EXJ) (Newberry and Brennan 2004; Heldwein and Brennan 2001); E. coli copper efflux regulator CueR [PDB codes 1Q05 (Cu2+-bound form), 1Q06 (Ag2+-bound form), and 1Q07 (AU2+-bound form)] and zinc-sensing transcriptional regulator ZntR (PDB codes 1Q08, 1Q09, and 1Q0A) (Changela et al. 2003); and E. coli CueR [PDB codes 4WLS and 4WLW (DNA-bound form)] (Philips et al. 2015).
Redox sensors involved in RCS
Some redox sensors mentioned above (OxyR, OhrR, SoxR, and Spx) are also known to recognize and respond to RCS (Gray et al. 2013). In addition, ComR, MsrPQ, NemR, PerR, RclR, and YjiE have been identified as RCS-sensitive redox sensors. Among the RCS-responsive redox sensors, MsrPQ is a hypersensitive redox sensor of RCS and responds to HOCl (Ezraty et al. 2017). Escherichia coli RclR (previously known as YkgD) belongs to the AraC family and has highly conserved cysteine residues that are specifically sensitive to RCS (Parker et al. 2013). In NemR (PDB code 4YZE), Cys106 and Lys175 form a reversible sulfenamide bond in the presence of HOCl or reactive chloramines, such as N-chlorotaurine (Gray et al. 2015) (Fig. 8). This chemical modification leads to the expression of three proteins (NemR, NemA, and GloA) that block RCS attacks from neutrophils during inflammation (Gray et al. 2015). YjiE is a global regulator that upregulates the genes responsible for cysteine and methionine metabolism. YjiE also downregulates the genes responsible for iron acquisition after HOCl treatment (Gray et al. 2013).
Structure of NemR, a RCS-sensitive redox sensor. Two subunits of the NemR homodimer are shown as gray and pale cyan ribbons. The important residues in only one subunit are represented in a ball-and-stick model with carbon (green), nitrogen (blue) and sulfur (yellow) atoms. The overall structure of NemR (a) is enlarged in the view on the right (b) to clearly visualize the active site. Cys106 and Lys175, which can form a reversible sulfonamide bond with RCS, are in close proximity
Conclusions
Both prokaryotic and eukaryotic cells must maintain redox balance to survive harsh environments. If cells are exposed to oxidative stresses, such as ROS, RES, RCS, and RNS, cellular components, including lipids, DNA, and proteins, can be damaged or modified, leading to cellular malfunction and death. More specifically, the oxidation of polyunsaturated fatty acids in membranes can cause lipid peroxidation and decrease membrane fluidity, leading to the disruption of membrane-bound proteins (Humphries and Szweda 1998; Cabiscol et al. 2000). Additionally, oxidative molecules can induce single- and double-stranded breaks in the DNA backbone and modify chemical groups, which disrupts DNA replication (Cabiscol et al. 2000). Additionally, amino acids and protein structures can be modified by oxidative species, leading to the interruption or disturbance of specific cellular metabolic steps (Cabiscol et al. 2000). To defend against such oxidative attacks, cells constitutively produce antioxidants, such as NADPH/NADH pools, β-carotene, ascorbic acid, α-tocopherol, and glutathione (GSH), that maintain a reducing intracellular environment or remove oxidative molecules (Cabiscol et al. 2000; Antelmann and Helmann 2011). To treat specific oxidative molecules and restore redox balance, cysteine thiol groups in proteins are used to activate specific detoxification pathways (Lee et al. 2016). Specifically, transcription factors with redox sensing functions can directly sense and quickly respond to oxidative stress, which is regarded as an efficient way for bacteria to survive. Pathogenic bacteria use this strategy to adapt to their host immune defense system by evading oxidative attacks from the host. For this reason, the redox sensors of pathogenic bacteria are important potential antibiotic targets. Furthermore, low levels of ROS and RES also act as second messengers to modulate signaling transduction pathways in eukaryotes (Rudolph and Freeman 2009). Redox sensor-mediated regulation reflects the elaborate strategies bacteria use to finely tune the expression of relevant antioxidant genes by sensing specific arrays of oxidative molecules. The structures of redox sensors elucidated from many studies reveal how the presence of oxidative molecules initiate the recognition of an oxidation state and the subsequent conformational rearrangements that alter DNA-binding affinity at the molecular level.
References
Antelmann H, Helmann JD (2011) Thiol-based redox switches and gene regulation. Antioxid Redox Sign 14:1049–1063
Aslund F, Zheng M, Beckwith J, Storz G (1999) Regulation of the OxyR transcription factor by hydrogen peroxide and the cellular thiol-disulfide status. Proc Natl Acad Sci USA 96:6161–6165
Barford D (2004) The role of cysteine residues as redox-sensitive regulatory switches. Curr Opin Struct Biol 14:679–686
Cabiscol E, Tamarit J, Ros J (2000) Oxidative stress in bacteria and protein damage by reactive oxygen species. Int Microbiol 3:3–8
Capra EJ, Laub MT (2012) Evolution of two-component signal transduction systems. Annu Rev Microbiol 66:325–347
Chang CC, Lin LY, Zou XW, Huang CC, Chan NL (2015) Structural basis of the mercury(II)-mediated conformational switching of the dual-function transcriptional regulator MerR. Nucleic Acids Res 43:7612–7623
Changela A, Chen K, Xue Y, Holschen J, Outten CE, O’halloran TV, Mondragon A (2003) Molecular basis of metal-ion selectivity and zeptomolar sensitivity by CueR. Science 301:1383–1387
Chen H, Yi CQ, Zhang J, Zhang WR, Ge ZY, Yang CG, He CA (2010) Structural insight into the oxidation-sensing mechanism of the antibiotic resistance of regulator MexR. EMBO Rep 11:685–690
Chi BK, Albrecht D, Gronau K, Becher D, Hecker M, Antelmann H (2010) The redox-sensing regulator YodB senses quinones and diamide via a thiol-disulfide switch in Bacillus subtilis. Proteomics 10:3155–3164
Choi H, Kim S, Mukhopadhyay P, Cho S, Woo J, Storz G, Ryu SE (2001) Structural basis of the redox switch in the OxyR transcription factor. Cell 105:103–113
Counago RM, Chen NH, Chang CW, Djoko KY, Mcewan AG, Kobe B (2016) Structural basis of thiol-based regulation of formaldehyde detoxification in H. influenzae by a MerR regulator with no sensor region. Nucleic Acids Res 44:6981–6993
Crawford MA, Tapscott T, Fitzsimmons LF, Liu L, Reyes AM, Libby SJ, Trujillo M, Fang FC, Radi R, Vazquez-Torres A (2016) Redox-active sensing by bacterial DksA transcription factors is determined by cysteine and zinc content. Mbio 7:e02161-15
Elsen S, Swem LR, Swem DL, Bauer CE (2004) RegB/RegA, a highly conserved redox-responding global two-component regulatory system. Microbiol Mol Biol Rev 68:263–279
Elsen S, Jaubert M, Pignol D, Giraud E (2005) PpsR: a multifaceted regulator of photosynthesis gene expression in purple bacteria. Mol Microbiol 57:17–26
Ezraty B, Gennaris A, Barras F, Collet JF (2017) Oxidative stress, protein damage and repair in bacteria. Nat Rev Microbiol 15:385–396
Forstermann U, Sessa WC (2012) Nitric oxide synthases: regulation and function. Eur Heart J 33(7):829–837
Fuangthong M, Helmann JD (2002) The OhrR repressor senses organic hydroperoxides by reversible formation of a cysteine-sulfenic acid derivative. Proc Natl Acad Sci USA 99:6690–6995
Fuangthong M, Atichartpongkul S, Mongkolsuk S, Helmann JD (2001) OhrR is a repressor of ohrA, a key organic hydroperoxide resistance determinant in Bacillus subtilis. J Bacteriol 183:4134–4141
Giles NM, Giles GI, Jacob C (2003) Multiple roles of cysteine in biocatalysis. Biochem Biophys Res Commun 300:1–4
Gray MJ, Wholey WY, Jakob U (2013) Bacterial responses to reactive chlorine species. Annu Rev Microbiol 67:141–160
Gray MJ, Li Y, Leichert LIO, Xu ZH, Jakob U (2015) Does the transcription factor NemR use a regulatory sulfenamide bond to sense bleach? Antioxid Redox Sign 23:747–754
Green J, Paget MS (2004) Bacterial redox sensors. Nat Rev Microbiol 2:2954–2966
Heldwein EE, Brennan RG (2001) Crystal structure of the transcription activator BmrR bound to DNA and a drug. Nature 409:378–382
Hillion M, Antelmann H (2015) Thiol-based redox switches in prokaryotes. Biol Chem 396:415–444
Hong M, Fuangthong M, Helmann JD, Brennan RG (2005) Structure of an OhrR-ohrA operator complex reveals the DNA binding mechanism of the MarR family. Mol Cell 20:131–141
Humphries KM, Szweda LI (1998) Selective inactivation of alpha-ketoglutarate dehydrogenase and pyruvate dehydrogenase: reaction of lipoic acid with 4-hydroxy-2-nonenal. Biochemistry 37:15835–15841
Imlay JA (2008) Cellular defenses against superoxide and hydrogen peroxide. Annu Rev Biochem 77:755–776
Imlay JA (2013) The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat Rev Microbiol 11:443–454
Ji QJ, Zhang L, Jones MB, Sun F, Deng X, Liang HH, Cho H, Brugarolas P, Gao YHN, Peterson SN, Lan LF, Bae T, He C (2013) Molecular mechanism of quinone signaling mediated through S-quinonization of a YodB family repressor QsrR. Proc Natl Acad Sci USA 110:5010–5015
Jo I, Chung IY, Bae HW, Kim JS, Song S, Cho YH, Ha NC (2015) Structural details of the OxyR peroxide-sensing mechanism. Proc Natl Acad Sci USA 112:6443–6448
Jo I, Kim D, Bang YJ, Ahn J, Choi SH, Ha NC (2017) The hydrogen peroxide hypersensitivity of OxyR2 in Vibrio vulnificus depends on conformational constraints. J Biol Chem 292:7223–7232
Laguri C, Phillips-Jones MK, Williamson MP (2003) Solution structure and DNA binding of the effector domain from the global regulator PrrA (RegA) from Rhodobacter sphaeroides: insights into DNA binding specificity. Nucleic Acids Res 31:6778–6787
Lamour V, Westblade LF, Campbell EA, Darst SA (2009) Crystal structure of the in vivo-assembled Bacillus subtilis Spx/RNA polymerase alpha subunit C-terminal domain complex. J Struct Biol 168:352–356
Le Rossignol S, Ketheesan N, Haleagrahara N (2017) Redox-sensitive transcription factors play a significant role in the development of rheumatoid arthritis. Int Rev Immunol 12:1–15
Lee C, Lee SM, Mukhopadhyay P, Kim SJ, Lee SC, Ahn WS, Yu MH, Storz G, Ryu SE (2004) Redox regulation of OxyR requires specific disulfide bond formation involving a rapid kinetic reaction path. Nat Struct Mol Biol 11:1179–1185
Lee SJ, Lee IG, Lee KY, Kim DG, Eun HJ, Yoon HJ, Chae S, Song SH, Kang SO, Seo MD, Kim HS, Park SJ, Lee BJ (2016) Two distinct mechanisms of transcriptional regulation by the redox sensor YodB. Proc Natl Acad Sci USA 113:E5202–E5211
Leelakriangsak M, Kobayashi K, Zuber P (2007) Dual negative control of spx transcription initiation from the P-3 promoter by repressors PerR and YodB in Bacillus subtilis. J Bacteriol 189:1736–1744
Loi VV, Rossius M, Antelmann H (2015) Redox regulation by reversible protein S-thiolation in bacteria. Front Microbiol 6:187
Masuda S, Dong C, Swem D, Setterdahl AT, Knaff DB, Bauer CE (2002) Repression of photosynthesis gene expression by formation of a disulfide bond in CrtJ. Proc Natl Acad Sci USA 99:7078–7083
Mukhopadhyay P, Zheng M, Bedzyk LA, Larossa RA, Storz G (2004) Prominent roles of the NorR and Fur regulators in the Escherichia coli transcriptional response to reactive nitrogen species. Proc Natl Acad Sci USA 101:745–750
Nakano MM, Lin A, Zuber CS, Newberry KJ, Brennan RG, Zuber P (2010) Promoter recognition by a complex of Spx and the C-terminal domain of the RNA polymerase alpha subunit. PLoS ONE 5:e8664
Nathan C (2003) Specificity of a third kind: reactive oxygen and nitrogen intermediates in cell signaling. J Clin Investig 111:769–778
Nathan C, Xie QW (1994) Regulation of biosynthesis of nitric oxide. J Biol Chem 269:13725–13728
Newberry KJ, Brennan RG (2004) The structural mechanism for transcription activation by MerR family member multidrug transporter activation, N terminus. J Biol Chem 279:20356–20362
Newberry KJ, Nakano S, Zuber P, Brennan RG (2005) Crystal structure of the Bacillus subtilis anti-alpha, global transcriptional regulator, Spx, in complex with the alpha C-terminal domain of RNA polymerase. Proc Natl Acad Sci USA 102:15839–15844
Newberry KJ, Fuangthong M, Panmanee W, Mongkolsuk S, Brennan RG (2007) Structural mechanism of organic hydroperoxide induction of the transcription regulator OhrR. Mol Cell 28:652–664
Nowak E, Panjikar S, Konarev P, Svergun DI, Tucker PA (2006) The structural basis of signal transduction for the response regulator PrrA from Mycobacterium tuberculosis. J Biol Chem 281:9659–9666
Ortiz de Orué Lucana D, Wedderhoff I, Groves MR (2012) ROS-mediated signalling in bacteria: zinc-containing Cys-X-X-Cys redox centres and iron-based oxidative stress. J Signal Transduct 2012:605905
Palm GJ, Chi BK, Waack P, Gronau K, Becher D, Albrecht D, Hinrichs W, Read RJ, Antelmann H (2012) Structural insights into the redox-switch mechanism of the MarR/DUF24-type regulator HypR. Nucleic Acids Res 40:4178–4192
Parker BW, Schwessinger EA, Jakob U, Gray MJ (2013) The RclR protein is a reactive chlorine-specific transcription factor in Escherichia coli. J Biol Chem 288:32574–32584
Philips SJ, Canalizo-Hernandez M, Yildirim I, Schatz GC, Mondragon A, O’halloran TV (2015) Allosteric transcriptional regulation via changes in the overall topology of the core promoter. Science 349:877–881
Pop SM, Gupta N, Raza AS, Ragsdale SW (2006) Transcriptional activation of dehalorespiration—identification of redox-active cysteines regulating dimerization and DNA binding. J Biol Chem 281:26382–26390
Rudolph TK, Freeman BA (2009) Transduction of redox signaling by electrophile-protein reactions. Sci Signal 2:re7
Sainsbury S, Ren J, Nettleship JE, Saunders NJ, Stuart DI, Owens RJ (2010) The structure of a reduced form of OxyR from Neisseria meningitidis. BMC Struct Biol 10:10
Seaver LC, Imlay JA (2001) Hydrogen peroxide fluxes and compartmentalization inside growing Escherichia coli. J Bacteriol 183:7182–7189
Sporer AJ, Kahl LJ, Price-Whelan A, Dietrich LEP (2017) Redox-based regulation of bacterial development and behavior. Annu Rev Biochem 86:777–797
Sukchawalit R, Loprasert S, Atichartpongkul S, Mongkolsuk S (2001) Complex regulation of the organic hydroperoxide resistance gene (ohr) from Xanthomonas involves OhrR, a novel organic peroxide-inducible negative regulator, and posttranscriptional modifications. J Bacteriol 183:4405–4412
Svintradze DV, Peterson DL, Collazo-Santiago EA, Lewis JP, Wright HT (2013) Structures of the Porphyromonas gingivalis OxyR regulatory domain explain differences in expression of the OxyR regulon in Escherichia coli and P. gingivalis. Acta Crystallogr D 69:2091–2103
Wang D, Huang S, Liu P, Liu X, He Y, Chen W, Hu Q, Wei T, Gan J, Ma J, Chen H (2016) Structural analysis of the Hg(II)-regulatory protein Tn501 MerR from Pseudomonas aeruginosa. Sci Rep 6:33391
Watanabe S, Kita A, Kobayashi K, Miki K (2008) Crystal structure of the [2Fe-2S] oxidative-stress sensor SoxR bound to DNA. Proc Natl Acad Sci USA 105:4121–4126
Whiteley AT, Ruhland BR, Edrozo MB, Reniere ML (2017) A redox-responsive transcription factor is critical for pathogenesis and aerobic growth of Listeria monocytogenes. Infect Immun 85:e00978
Zheng M, Aslund F, Storz G (1998) Activation of the OxyR transcription factor by reversible disulfide bond formation. Science 279:1718–1721
Zheng M, Wang X, Doan B, Lewis KA, Schneider TD, Storz G (2001) Computation-directed identification of OxyR DNA binding sites in Escherichia coli. J Bacteriol 183:4571–4579
Zuber P (2004) Spx-RNA polymerase interaction and global transcriptional control during oxidative stress. J Bacteriol 186:1911–1918
Acknowledgements
This work was funded by the Korea Ministry of Science, Information, Communication, Technology, and Future Planning and National Research Foundation (NRF) of Korea (Grants NRF-2014K1A3A1A19067618 and NRF-2015R1A2A1A05001894 awarded to B.-J.L. and NRF-2016R1C1B2014609 awarded to S.J.L.). This work was also supported by the 2016 BK21 Plus Project for Medicine, Dentistry, and Pharmacy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors have no conflicts of interest to declare.
Rights and permissions
About this article
Cite this article
Lee, S.J., Kim, DG., Lee, KY. et al. Regulatory mechanisms of thiol-based redox sensors: lessons learned from structural studies on prokaryotic redox sensors. Arch. Pharm. Res. 41, 583–593 (2018). https://doi.org/10.1007/s12272-018-1036-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12272-018-1036-0