Abstract
Extracellular vesicles (EVs), including exosomes and microvesicles, emerge to be crucial mediators of cell-to-cell communication in multiple organs. Non-coding RNAs loaded inside EVs contribute as one major mechanism for remote information transfer among different cell types or organs. Increasing evidence suggests that EV-associated non-coding RNAs derived from cardiovascular or non-cardiac cells regulate cardiovascular pathophysiology in heart development and diseases. The functional relevance of the EV-associated ncRNAs in heart diseases provides an avenue to develop novel diagnostic tools and therapies for heart diseases. In this review, we summarize the recent advancement of EV-associated ncRNAs in different cardiovascular diseases, including myocardial infarction, arrhythmias, cardiac hypertrophy, and heart failure, with an emphasis on the underlying molecular mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Extracellular vesicles (EVs) are membrane-bound vesicles released by both normal and transformed cells, carrying bioactive molecules including proteins, lipids, and RNAs. EVs can be detected in all major bodily fluids, like blood, urine and saliva, among which the circulating EVs in the blood have been mostly studied [1]. Cumulative evidence highlights the important role of EVs in cell-to-cell communication by releasing their cargoes into the target cell [2]. This type of cell-to-cell communication is engaged in numerous pathophysiological processes, such as inflammation, immune modulation, neurological diseases, cancer, and cardiovascular diseases [3,4,5,6,7].
It has been well recognized that majority of the human transcriptome are non-coding RNAs (ncRNAs) with only 2% of them corresponding to protein-coding genes [8]. These ncRNAs are grouped into transfer RNAs (tRNAs), ribosomal RNAs (rRNAs), small nucleolar RNAs (snoRNAs), microRNAs (miRNAs), piwi-interacting RNAs (piRNAs), circular RNAs (circRNAs), and long non-coding RNAs (lncRNAs) [9]. The functionality of these ncRNAs in maintaining normal cell physiology has been repetitively verified by mounting studies [3,4,5,6,7]. Interestingly, ncRNAs also appear in plasma, where they are mostly packaged inside EVs rather than as free molecules, so that they are well protected from degradation by endogenous RNases [10, 11]. These circulating ncRNAs can then travel to neighboring or distant cells to transmit complex massages, particularly during the pathogenesis of diseases [12]. Thus, the EV-associated ncRNAs contribute as novel candidates for disease diagnostics or even therapeutics.
As the leading cause of death worldwide, heart diseases represent an important scenario whereby the EV-associated ncRNAs participate in the cross talk among different cell types. Here we summarize a recent finding about functional EV-associated ncRNAs in heart diseases, with an emphasis on the molecular mechanisms. We also discuss the potential application of EV-associated ncRNAs in diagnostics and therapeutics.
Function Mechanisms of EV-Associated ncRNAs
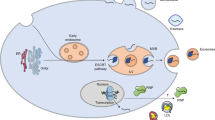
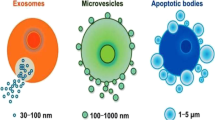
EVs are extracellular structures that are enclosed by a lipid bilayer and secreted by cells into their environment [13]. The two major classes of EVs are exosomes and microvesicles (MVs) [14]. Exosomes (30 ~ 150 nm diameter) are small vesicles released from cells when multivesicular bodies fuse with the plasma membrane. Micovesicles (MVs) are membranous vesicles with diverse sizes (100 ~ 1000 nm) compared to exosomes, and are released from the cell through blebbing of the plasma membrane. MVs are released into the extracellular space from plasma membrane instead of internal membranes by a calcium-dependent mechanism [14]. EV size is determined by its biosynthesis, associating with EV composition correlated with the cell line of origin [15].
Proteins, DNA, lipids, and RNAs are common EV cargoes that elicit diverse cellular responses in recipient cells [16]. Exosomes released from a certain tissue usually possesses a tissue-specific protein signature [17, 18]. Nevertheless, exosomes commonly carry different RNA species compared to their parental cells [19]. Accumulating evidence demonstrates that EVs actually offer a “protective shell” to protect the associated ncRNAs from degradation and maintain their functional integrity during circulation [20]. The EV content is affected by microenvironments, including aging, inflammatory process, oxygen content, and some other factors.
The identification of RNAs in EVs has progressed immensely due to highly sensitive methods of RNA detection (RNA-Seq) and reverse transcription and quantitative polymerase chain reaction (RT-qPCR) analysis in recent years. These RNA populations include various protein-coding transcript mRNAs and many types of non-coding RNAs, including miRNAs, lncRNAs, circRNAs, snoRNAs, snRNAs, tRNAs, rRNAs, and piRNAs [19]. These RNAs can be transferred from parent cells to recipient cells, where they can regulate or serve as templates for protein production [21, 22]. Here we focus on microRNAs, lncRNAs, and circRNAs identified in EVs, which could shed light on the source of diversity in the outcome of the EV-associated ncRNA mechanistic studies. The mechanisms of different RNAs entering the receptor cells are functionally summarized in the following (Fig. 1).
Regulatory mechanisms of EV-associated ncRNAs. (A) miRNAs participate in intercellular signaling, for example (nuclear factor-kappaB) NF-kB in TLR signaling pathways. (B) circRNAs act as microRNA sponges that regulate target gene expression. (C) miRNAs target mRNAs for degradation. (D) lncRNAs as decoy molecules regulating mRNA molecules or sponging miRNA molecules to block the inhibitory effect of miRNAs on mRNA molecules
miRNAs
miRNAs participate in multi-biological processes via the subtle and precise regulation of gene expression; miRNAs not only target the 3′ UTR of the mRNA but also interact with other parts of the mRNA, such as the 5′ UTR or coding sequence, and even the gene promoter [23]. Additionally, microRNAs can also participate in intercellular signaling. A report demonstrates that EV miRNAs can act as a Toll-like receptor (TLR) ligand and induce an immune response or inhibit the activation of macrophages by inhibiting TLR signals [24,25,26,27,28] (Fig. 1). Based on these complicated mechanisms, single miRNAs could impact on a series of pathways or axes to regulate one target gene expression.
circRNAs
CircRNAs mostly comprise protein-coding exons; some circRNAs contain both exons and introns, and others are derived from untranslated regions (5′ or 3′ UTRs) or introns of mRNAs or ncRNAs [29, 30]. One prominent function of circRNAs is to act as ceRNA (miRNA sponges), but they can also modulate RNA transcription, splicing, turnover, and translation, and may even have protein-coding potential [31]. CircRNAs have been shown to sponge some miRNAs and thereby control their function as regulators of mRNA stability and/or translation (Fig. 1). For instance, ciRS-7 or CRD1as is generated from the CDR1 gene, and there are over 70 conserved binding sites for miRNA-7 [32]. Moreover, CRD1as RNA binds to the protein Argonaut 2 (AGO2) and miRNA, forming an RNA-induced silencing complex in the cell [33]. Another study shows that iRS-7 sponges and regulates the function of miR-7. Interestingly, EIciEIF3J interacts with snRNP U1 and the promoter EIF3J to enhance EIF3J transcription [33]. It is important to note that not all circRNAs are enriched for miRNA-binding sites and repress miRNA [34].
lncRNAs
lncRNAs are operationally defined as transcripts of greater than 200 nucleotides that function by means other than coding for proteins. Numerous lncRNAs have been found in EVs that regulate gene transcription through transcriptional interference. For example, a finding indicates that lncRNAs, pseudogenes, and mRNAs cross talk by competing for a common pool of miRNAs and build a complex regulatory network [35] (Fig. 1). The lncRNA growth arrest-specific 5 (GAS5) was generally regarded as a tumor suppressor, acting as a miR-21 sponge, that could inhibit the proliferation and promote the apoptosis of various cancer cells [36]. Interestingly, the study showed that GAS5 contained a binding site for miR-23a and acted as a sponge of miR-23a. An upregulated GAS5 expression inhibited cardiomyocyte hypertrophy through negatively regulating miR-23a and its target forkhead box O3 (Foxo3a) [37, 38].
EV-Associated ncRNAs in Heart Diseases
In the heart, EVs are secreted from multiple cell types, such as cardiomyocytes, cardiac progenitor cells, endothelial cells, epithelial cells, macrophages, and fibroblasts [39, 40] (Fig. 2). A growing body of studies demonstrates that cardiac EVs, particularly the associated ncRNAs, play a crucial role in intercellular communication during the pathogenesis of heart diseases (Table 1).
Myocardial Infarction
Acute myocardial infarction with high mortality and morbidity rate is a life-threatening condition that occurs when blood flowing to the heart muscle is abruptly cut off, causing tissue damage [64].
Ong et al. [65] found that the exosome-associated miR126 and miR-210 mediated the cross talk between epithelial cells (ECs) and cardiac progenitor cells (CPCs). Hypoxia-inducible factor-1 (HIF1) is a transcription factor that mediates adaptive responses to ischemia [66]. Exosomes from ECs overexpressing HIF1 have higher contents of miR-126 and miR-210. These miRNAs regulate pro-survival kinases and induce glycolysis in CPCs, which eventually protect the CPCs in hypoxic stress conditions and improve the survival of CPCs, while depletion of miR-126 and miR-210 exosomes from the ECs abrogates the protective effects [65] (Fig. 2).
Given their cardiac developmental origins, endogenous CPCs have been proposed as candidates for heart repair. CPC-conditioned medium (CM) protects HL-1 cardiomyocytes in mice and promotes tube formation in human umbilical vein endothelial cells (HUVECs) [67]. Surprisingly, CPC-derived exosomes are enriched with several cardioprotective and pro-angiogenic miRNAs, such as miR-210, miR-132, and miR-146a-3p (Fig. 2), compared with cardiac fibroblasts. Particularly, the effect of miR-210 is associated with a functional downregulation of ephrin A3 and protein tyrosine phosphatase 1 (PTP1) [68]. miR-132 promotes tube formation in HUVECs, which is associated with a functional downregulation of the known miR-132 target, Ras GTPase-activating protein (RasGap-p120) [69]. Dosage- and time-dependent assays show an increase in the intracellular concentrations of miR-210 and miR-132 after exposure of the HL-1 cardiomyocyte line to EVs secreted by CPCs [70]. Functional miRNAs associated with CPC-derived exosomes appear to be a plausible mechanism in the recovery of cardiac function during myocardial infarction.
Increasing evidence shows that circulating miRNAs can be used as potential diagnostic biomarkers for myocardial infarction, including miR-133, miR-208b, and miR-499, which play important roles in cell differentiation and function [71]. This hypothesis is strengthened by a report that miRNAs derived from the heart release information into the circulation system upon myocardial injury. The absence of miR-133a expression leads to the ectopic expression of smooth cardiomyopathy and heart failure [72]. One research studying on 312 patients showed that the miR-133a level was higher in AMI patients compared with non-AMI patients [73]. miR-208a can be found in extracellular environments including blood, saliva, and urine [74]. miR-208a is reported as a gold standard marker of myocardial injury from increasing concentrations in myocardial injury mice [75], while in plasma, miR-499 shows a rapid increase in AMI patients, as well as unstable angina and non-ST elevation within 3 h of symptom onset. This provides evidence of miR-499 as a potentially novel biomarker to accelerate the diagnosis of MI or acute coronary syndrome patients [76, 77].
Cardiac Arrhythmias
Cardiac arrhythmias are associated with inflammation, metabolic disorder, and structural and physiological irregularity [78]. Cardiac electrophysiology dysfunction is one of the features in cardiac arrhythmias. It might be affected by ventricular tachycardia and atrial fibrillation (AF) [79]. Atrial fibrillation is a chronic and the most common form of arrhythmia. Variations of miRNA and its targeted genes are related to the initiation and progression of atrial fibrillation [80].
A recent study investigated the role of epicardial fat (eFat)-derived EVs in the pathogenesis of AF [81]. Several miRNAs, including miR-146b, miR-133a, and miR-29a (Fig. 2), released by EVs transfer the messages from cardiomyocytes or mesenchymal stem cells (MSCs) to cardiac fibroblasts to stimulate a cardiac fibroblast that forms the arrhythmogenic substrate for AF. eFat-EVs from patients with AF carried more profibrotic miRNAs (e.g., mmiR-146b) than patients without AF.
Another example from Ye et al. [82] suggests that exosomal miRNAs target mRNAs and lead to its deregulation. They find that miR-146-5p is released from CMs and represses the expression of the target protein TIMP metallopeptidase inhibitor 4 (TIMP4) in cardiac fibroblasts. Inhibiting miR-146b-5p in CMs of MI heart rescues TIMP4 expression, and consequently reduces fibrotic markers matrix metallopeptidase 9 (MMP9), transforming growth factor-beta (TGFB1), and collagen type I alpha 1 chain (COL1A1).
Yao et al. [83] find that miR-133a-3p is significantly reduced in atrium tissues of rats with AF induction. This provides a perspective that miRNA appears to be loaded selectively into exosomes and interacts with lncRNAs representing new shuttles of cell-to-cell communication. They found that miR-133a-3p from cardiomyocytes is identified as the target gene of lncRNA myocardial infarction-associated transcript (MIAT). Inhibition of MIAT reduces the mRNA expression of fibrosis-related gene TGF-β1, collagen I, and collagen III, while anti-miR-133a-3p administration significantly reversed the alleviation by knocking down MIAT. Thus, upregulating miR-133a-3p can decrease collagen content and inhibit atrial remodeling.
Similarly, miR-29a is transferred to cardiac fibroblasts to suppress collagen synthesis [81]. miR-29a-3p predominantly exists in mesenchymal stem cell (MSC)-derived EVs [84], serving as a mediator for long-distance cell-to-cell communications. A study validates calcium voltage-gated channel subunit alpha1 C (CACNA1C) to be the direct target gene of miR-29a-3p [85]. miR-29a-3p transfection in cardiomyocytes reduces the density of L-type calcium induced by electrical remodeling, which represents a novel approach to prevent atrial fibroblasts.
Cardiac Hypertrophy
Cardiac hypertrophy appears as an abnormal enlargement or thickening of the heart muscle, accompanied by increasing cardiomyocyte death and fibrotic remodeling, while the following reduction in systolic and diastolic function irreversibly develops into heart failure [86].
A recent report analyzed the potential paracrine miRNA cross talk between cardiac fibroblasts and cardiomyocytes in a mouse model of TAC [54]. It shows that miR-21-3p is abundant in fibroblast-derived exosomes. Once taken up by cardiomyocytes, miR-21-3p silences its target genes SH3 domain containing 2 (SORB2) and PDZ and LIM domain 5 (PDLIM5), and subsequently induces cardiomyocyte hypertrophy. Interestingly, administration of an antagonist of miR-21 in mice improves cardiac function in Ang II-induced cardiac hypertrophy.
Another two studies by Nie et al. [87] and Li et al. [88] also reveal interactions between cardiac fibroblasts and cardiomyocytes in the progression of cardiac remodeling. They show that miR-127 is highly expressed in the hearts of CHF patients and pressure overload–induced hypertrophic mice [87, 88]. Their results show that miR-217-enriched exosomes derived from cardiomyocytes induce cardiac hypertrophy, whereas the miR217-TUD-mediated downregulation of miR-217 reversed these effects. miR-217-containing exosomes targeting phosphatase and tension homolog (PTEN) enhances the proliferation of fibroblasts in vivo.
Exosomes are thought to play important roles in MSC-related cardioprotective effects [89, 90]. A study demonstrated the cross talk between bone marrow mesenchymal stem cells (BM-MSCs) and cardiomyocytes [91] (Fig. 2). They select the most relevant miRNA to cardiac hypertrophy from BM-MSC exosomes, and find miR-29 to be a key regulatory cargo contributing to the cardiac protective effects during pressure overload. It has been reported that the circulating miR-29a is upregulated in patients with hypertrophic cardiomyopathy (HCM) [92], representing a potential biomarker for cardiac remodeling assessment in HCM. In addition, miR-29a also plays a cardioprotective role in cardiac hypertrophy events by targeting proliferator-activated receptor δ (PPARδ) and downregulating ANF, which ameliorates the isoproterenol hydrochloride-induced cardiac hypertrophy response [93]. However, the mechanism of miR-29a on BM-MSC exosomes improving the function of the hypertrophic heart needs further investigation.
Heart Failure
Heart failure (HF) is a complex and progressive disease which may be caused by multi-pathological conditions including high blood pressure, coronary artery disease, faulty heart valves, and cardiomyopathy [94].
miR-21 is upregulated in the myocardium of HF murine models as well as in failing human myocardium [95]. It has been studied that miR-21 reveals a common way of communication between fibroblasts and cardiomyocytes (Fig. 2). Exosomes derived from cardiac fibroblasts selectively packaging miR-21 are taken up by cardiomyocytes to enhance cellular hypertrophy through repressing target genes SORB2 and PDLIM5 [96, 97]. miR-21 silencing attenuates the impairment of cardiac function and regression of cardiac hypertrophy fibrosis, which can prevent and even cure the structural and functional defeats in a mouse model of heart failure [98]. However, other research indicates that miR-21 inhibited by intravenous delivery of locked nucleic acid-modified (LNA) antimir oligonucleotides has no effect for remodeling response or preventing cardiac dysfunction in different mouse models of HF [99]. The results remain controversial since some microenvironment factors still need to be elucidated including individual microRNAs in vivo (Table 1).
Exosome-derived miR-92b-5p is found significantly increased in patients with acute HF caused by dilated cardiomyopathy [100]. Circulating miR-192 is a prognostic marker in ischemic heart failure [101]. It is characterized by accumulation of p53 causing the apoptosis of cardiomyocytes and resulting in the upregulation of miR-192 [102]. However, the exosome-derived miR-192 b-5p communication pathway in cardiac cells needs to be further clarified, since miR-192 molecular pathways still need to be further investigated (Fig. 2).
Other Heart Diseases
Peripartum cardiomyopathy (PPCM) is a life-threatening pregnancy-associated cardiomyopathy in previously healthy women. A finding suggests that miR-146a appears as a major mediator in the development of PPCM via a cross talk between endothelial cells and cardiomyocytes. The 16-kDa N-terminal prolactin fragment (16 K PRL) stimulates the release of miR-146a-loaded exosomes from ECs [103]. The miR-146a-containing exosomes lead to a subsequent decrease in metabolic activity and decrease in the expression of specific miR-146a-target mRNAs. In contrast, the pharmacological blockade of miR-146a or blockade of PRL with bromocriptine in postpartum CKO mice can attenuate miR-146a–target mRNA decrease and improve cardiac function.
Diabetic cardiomyopathy (DCM) is characterized by changes in myocardial structure and function, which increases the incidence of heart failure even after controlling for coronary artery disease and hypertension [104]. A report shows that exosomes derived from diabetic cardiomyocytes contain higher levels of miR-320. miR-320 is secreted by cardiomyocytes into exosomes which can be transferred to endothelial cells, and eventually downregulate the expression of its specific target genes such as insulin-like growth factor 1 (IGF-1), heat-shock protein 20 (HSP20), and transcription factor ETS2. The study reveals that cardiomyocytes could exert an anti-angiogenic effect through the release of miR-320-enriched exosomes in DCM.
Damages of cardiomyocytes and oxidative stress are stated as the main causes of Dox-induced cardiomyopathy [105]. A novel treatment of the human adipose–derived mesenchymal stem cells (MSCs) with hypoxia induced hypoxia-inducible factor lncRNA metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) accumulation in the secreted exosomes [106]. The exosome-mediated transfer of MALAT1 represses the expression of miR-92a-3p, which results in the destruction of cardiac homeostasis, for example inhibiting cardiomyocyte metabolism and autophagy via targeting ATG4a [107]. MALAT1 acts as a competing endogenous RNA (ceRNA) binding to miR-92a-3p, resulting in the activation of autophagy-related 4A cysteine peptidase (ATG4a), and improving mitochondrial metabolism. The MALAT1/miR-92a-3p/ATG4a axis partially mediates hypoxia in Dox-induced cardiac damage, which shows a new regulatory mechanism underlying the cross talk among lncRNAs, miRNAs, and mRNAs in response to stress stimulation.
Atherosclerosis is a pattern of the disease arteriosclerosis in which the wall of the artery develops abnormalities [108]. It is demonstrated that lncRNA growth-arrest specific transcript 5 (GAS5) plays a critical role in the progression of atherosclerosis [109]. THP-1 cells, when differentiating to macrophage-like cells, after taking up the exosomes derived from GAS5-overexpressing THP-1 cells, promote cell apoptosis [62]. However, exosomes shed by THP-1 cells with GAS5 knockdown inhibit the apoptosis of endothelial cells. This activation of exosomes provides an inspiration of an exosome mechanism for macrophage and endothelial cell communication mediated by lncRNA GAS5 in atherosclerosis [62] (Table 1).
Coronary artery disease (CAD) is caused by the buildup of the oxygen-rich blood plaque in the arteries [110]. A recent study identified that EVs incorporated with circulating lncRNA agap2-ANTISENSE RNA1[AS1] (PUNISHER) significantly increased in CAD [61]. In this case, EVs appear to be vehicles which convey information from circulating lncRNAs to ECs. EVs carrying PUNISHER reduce the expression vascular endothelial growth factor A (VEGFA) in recipient ECs through interacting with heterogeneous nuclear ribonucleoprotein K (hnRNPK). The inhibition of PUNISHER expression is accompanied by an impairment of the angiogenic response and a decrease in cell proliferation. This study reveals that EV-incorporated PUNISHER regulates the transcription and stability of VEGFA, thereby controlling the angiogenic function of ECs and promoting an angiogenic response (Table 1).
Translational Studies of EV-Associated ncRNAs
Cardiovascular disorders are still a major cause of morbidity and mortality worldwide, although much progress has been made in basic research [111]. Substantial studies are devoted to understanding the biology of EVs and their potential application in cardiovascular diseases. Advances in EV research provide a new avenue to develop EV-based therapeutic strategies. Below we summarized recent progresses in translational medicine leveraging EV-associated ncRNAs for diagnostics and personalized therapies of heart diseases.
Native EVs for Cardiovascular Biomarkers and Therapies
Specific miRNAs carried by EVs have been proposed as specific biomarkers for the early monitoring of CVD and other diseases [1]. For example, circRNA mitochondrial fission and apoptosis-related (MFACR) represents a valuable biomarker to predict cardiac cell death [112]. Increasing miR-126 or miR-199a expression in circulating EVs predicts the occurrence risk of CV events in stable CAD patients [113]. Exosomes from patients with MI show higher levels of miR-133a and miR-1 [114].
The therapeutic role of EVs in recipient cells is mostly studied in the delivery of ncRNAs, particularly miRNAs [115]. Some of the miRNAs are considered cell-based therapeutic targets, such as miRNA-34, miRNA-15, miRNA-24, and miRNA-208, which are relevant to cardiovascular repairation [116]. In addition, EV-associated ncRNAs trigger various cardioprotective effects, like improving cell survival in cardiomyocytes and endothelial cells [7, 117, 118] and activating pro-survival signaling pathways, including AKT, ERK, and Toll-like receptors [119]. For example, cardiosphere-derived EVs with miRNA-146 improve heart function after MI by increasing cardiomyocyte proliferation and angiogenesis [11]. Extracellular matrix-derived EVs carrying miRNA-199a-3p rescue electrical function in bioengineered atria by regulating the acetylation of transcription factor GATA4 [120]. EV-derived miRNA-486 plays a cardioprotective role via targeting PTEN and activating AKT signaling [121].
However, usage of EVs as diagnostic biomarkers or therapeutic media remains challenging due to the lack of standardization about sample collection, isolation, and quantification [122], which need to be further established in future. EVs are affected by the time of day when the sample is collected and the amount of physical activity will also influence for collection [122]. Moreover, physiological fluids contain EVs secreted from both diseased and non-diseased cells, which complicates the recovery of disease-specific EVs as biomarkers or therapeutic media [123].
Exosome Engineering
Beyond diagnostic and prognostic applications, many studies have focused on the use of engineered EVs in drug delivery to the injured heart. In one approach, the EV-secreting cell is genetically modified to express targeting peptides on the membrane of the secreted EVs to increase EV homing [124,125,126]. Wang et al. [124] modified exosomes with enriched membrane protein (Lamp2b) fused with ischemic myocardium-targeting peptide CSTSMLKAC (IMTP). Although no absolute quantification was provided for the accumulation of EVs in the heart, EVs isolated from engineered cardiosphere-derived cells (CDCs) expressed the cardiomyocyte-specific peptide (CMP) on their surface and retained their native physical properties. Peptide-modified EVs resulted in greater accumulation in fluorescence imaging and increased uptake by cardiomyocytes when compared to the EVs without the surface modification [124, 125].
In another approach reported by Gee et al. [127], an extracellular nanovesicle–based ribonucleoprotein delivery system named NanoMEDIC is developed by combining two distinct homing mechanisms to realize gene editing. Chemical-induced dimerization recruits Cas9 protein into extracellular nanovesicles, and then a viral RNA packaging signal and two self-cleaving riboswitches tether and release sgRNA into nanovesicles. This method successfully achieves exon skipping efficiencies in skeletal muscle cells derived from Duchenne muscular dystrophy (DMD) patient iPS cells. Similarly, Campbell et al. [128] develop a specialized extracellular vesicle termed “gesicle” using vesicular stomatitis virus glycoprotein to efficiently deliver Cas9 in a ribonucleoprotein form targeting the HIV long terminal repeat (LTR). This method bypasses the need for transgene delivery, and allows finer control of Cas9 expression.
In addition, EVs have been combined with virus-based gene therapies. A number of researches decipher how EVs influence viral and bacterial pathogenesis by spreading pathogen-derived factor [129]. Meliani et al. [130] utilize exosome-associated AAV (adeno-associated virus) to improve liver gene transfer and protect from pre-existing humoral immunity to the capsid in mice with hemophilia. Ju et al. [131] find that the endosomal sorting complex required for transport (ESCRT) system helps viruses to produce enveloped virions which avoid immune surveillance. These efforts might further accelerate the application of EV-associated ncRNAs in cardiovascular systems.
Perspectives
Before establishing EVs as an effective therapeutic tool, however, several challenges need to be resolved, including identification of specific EV-producing cells, replication of optimal vesicle production conditions, and manipulation of vesicle cargo contents. Further progress is also required to enhance the therapeutic effect of EVs by increasing their stability and targeting the EVs to a specific location, enriching their therapeutic contents, improving their internalization and intracellular trafficking, and controlling their spatial and temporal release from biomaterials. Above all, the molecular mechanisms how EV-associated ncRNAs impact cardiovascular pathophysiology need to be further explored.
Taken together, EV-associated ncRNAs provide a promising therapeutic strategy for CVD based on their advantageous properties as important factors involved in many physiological and pathological cardiovascular processes [132, 133], an essential component of the paracrine effect of stem cell-based therapies [134], low immunogenicity, low toxicity, and high capability to carry bioactive molecules to target cells. Future efforts to resolve the challenges in isolating, purifying, and manipulating EVs would help optimize the translation of EVs in a clinical setting.
Abbreviations
- EVs:
-
Extracellular vesicles
- MVs:
-
Microvesicles
- ncRNA:
-
Non-coding RNA
- lncRNA:
-
Long-non-coding RNA
- cicrRNA:
-
Circular RNA
- micRNA:
-
MicroRNA
- AMI:
-
Acute myocardial infarction
- AF:
-
Atrial fibrillation
- HF:
-
Heart failure
- CAD:
-
Coronary artery disease
- DCM:
-
Diabetic cardiomyopathy
- ECs:
-
Epithelial cells
- CPCs:
-
Cardiac progenitor cells
- HIF1:
-
Hypoxia-inducible factor-1
- CM:
-
CPC-conditioned medium
- HUVECs:
-
Human umbilical vein endothelial cells
- RISC:
-
RNA-induced silencing complexes
- RasGap:
-
Ras GTPase-activating protein
- TLR:
-
Toll-like receptor
- MIAT:
-
Myocardial infarction-associated transcript
- SORB2:
-
H3 domain-containing 2
- PTEN:
-
Phosphatase and tension homolog
- BM-MSCs:
-
Bone marrow mesenchymal stem cells
- PPARδ:
-
Proliferator-activated receptor δ
- GAS5:
-
Growth-arrest specific transcript 5
- PUNISHER:
-
Agap2-antisense RNA1[AS1]
- hnRNPK:
-
Heterogeneous nuclear ribonucleoprotein K
- PTP1:
-
Protein tyrosine phosphatase 1
- TIMP4:
-
TIMP metallopeptidase inhibitor 4
- MMP9:
-
Matrix metallopeptidase 9
- TGFB1:
-
Transforming growth factor-beta
- COL1A1:
-
Collagen type I alpha 1 chain
- CACNA1C:
-
Calcium voltage-gated channel subunit alpha1 C
- SORB2:
-
SH3 domain-containing 2
- PDLIM5:
-
PDZ and LIM domain 5
- PPCM:
-
Peripartum cardiomyopathy
- IGF-1:
-
Insulin-like growth factor 1
- HSP20:
-
Heat-shock protein 20
- ATG4a:
-
Autophagy-related 4A cysteine peptidase
- MALAT1:
-
Metastasis-associated lung adenocarcinoma transcript 1
- VEGFA:
-
Vascular endothelial growth factor A
- hnRNPK:
-
Heterogeneous nuclear ribonucleoprotein K
- MFACR:
-
Mitochondrial fission and apoptosis-related
References
Jansen, F., Nickenig, G., & Werner, N. (2017). Extracellular vesicles in cardiovascular disease: Potential applications in diagnosis, prognosis, and epidemiology. Circulation Research, 120, 1649–1657. https://doi.org/10.1161/CIRCRESAHA.117.310752
Lee, T. H., et al. (2011). Microvesicles as mediators of intercellular communication in cancer–The emerging science of cellular “debris.” Semin Immunopathol, 33, 455–467. https://doi.org/10.1007/s00281-011-0250-3
Cossetti, C., et al. (2014). Extracellular vesicles from neural stem cells transfer IFN-gamma via Ifngr1 to activate Stat1 signaling in target cells. Molecular Cell, 56, 193–204. https://doi.org/10.1016/j.molcel.2014.08.020
Hazan-Halevy, I., et al. (2015). Cell-specific uptake of mantle cell lymphoma-derived exosomes by malignant and non-malignant B-lymphocytes. Cancer Letters, 364, 59–69. https://doi.org/10.1016/j.canlet.2015.04.026
Lee, Y. S., & Dutta, A. (2009). MicroRNAs in cancer. Annual Review of Pathology: Mechanisms of Disease, 4, 199–227. https://doi.org/10.1146/annurev.pathol.4.110807.092222
Yanez-Mo, M., et al. (2015). Biological properties of extracellular vesicles and their physiological functions. The Journal of Extracellular Vesicles, 4, 27066. https://doi.org/10.3402/jev.v4.27066
Fasolo, F., Di Gregoli, K., Maegdefessel, L., & Johnson, J. L. (2019). Non-coding RNAs in cardiovascular cell biology and atherosclerosis. Cardiovascular Research, 115, 1732–1756. https://doi.org/10.1093/cvr/cvz203
International Human Genome Sequencing, C. (2004). Finishing the euchromatic sequence of the human genome. Nature, 431, 931–945. https://doi.org/10.1038/nature03001
Pennisi, E. (2012). ENCODE project writes eulogy for junk DNA. Science, 337, 1159–1161. https://doi.org/10.1126/science.337.6099.1159
Tsui, N. B., Ng, E. K., & Lo, Y. M. (2002). Stability of endogenous and added RNA in blood specimens, serum, and plasma. Clinical Chemistry, 48, 1647–1653.
Wang, G. K., et al. (2010). Circulating microRNA: A novel potential biomarker for early diagnosis of acute myocardial infarction in humans. European Heart Journal, 31, 659–666. https://doi.org/10.1093/eurheartj/ehq013
Kamm, R. C., & Smith, A. G. (1972). Nucleic acid concentrations in normal human plasma. Clinical Chemistry, 18, 519–522. https://doi.org/10.1093/clinchem/18.6.519
Gyorgy, B., et al. (2011). Membrane vesicles, current state-of-the-art: Emerging role of extracellular vesicles. Cellular and Molecular Life Sciences, 68, 2667–2688. https://doi.org/10.1007/s00018-011-0689-3
Zampetaki, A., Willeit, P., Drozdov, I., Kiechl, S., & Mayr, M. (2012). Profiling of circulating microRNAs: From single biomarkers to re-wired networks. Cardiovascular Research, 93, 555–562. https://doi.org/10.1093/cvr/cvr266
Zhang, H., et al. (2018). Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nature Cell Biology, 20, 332–343. https://doi.org/10.1038/s41556-018-0040-4
Toh, W. S., Lai, R. C., Hui, J. H. P., & Lim, S. K. (2017). MSC exosome as a cell-free MSC therapy for cartilage regeneration: Implications for osteoarthritis treatment. Seminars in Cell & Developmental Biology, 67, 56–64. https://doi.org/10.1016/j.semcdb.2016.11.008
Skog, J., et al. (2008). Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nature Cell Biology, 10, 1470–1476. https://doi.org/10.1038/ncb1800
Castellani, C., et al. (2020). Circulating extracellular vesicles as non-invasive biomarker of rejection in heart transplant. Journal of Heart and Lung Transplantation, 39, 1136–1148. https://doi.org/10.1016/j.healun.2020.06.011
Nolte-‘t Hoen, E. N., et al. (2012). Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nucleic Acids Res, 40, 9272–9285. https://doi.org/10.1093/nar/gks658
Yang, Q., Diamond, M. P., & Al-Hendy, A. (2016). The emerging role of extracellular vesicle-derived miRNAs: Implication in cancer progression and stem cell related diseases. Journal of Clinical Epigenetics, 2,
Montecalvo, A., et al. (2012). Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood, 119, 756–766. https://doi.org/10.1182/blood-2011-02-338004
Hergenreider, E., et al. (2012). Atheroprotective communication between endothelial cells and smooth muscle cells through miRNAs. Nature Cell Biology, 14, 249–256. https://doi.org/10.1038/ncb2441
Pu, M., et al. (2019). Regulatory network of miRNA on its target: Coordination between transcriptional and post-transcriptional regulation of gene expression. Cellular and Molecular Life Sciences, 76, 441–451. https://doi.org/10.1007/s00018-018-2940-7
Verweij, F. J., et al. (2019). Live tracking of inter-organ communication by endogenous exosomes In Vivo. Developmental Cell, 48, 573–589. https://doi.org/10.1016/j.devcel.2019.01.004
Alexander, M., et al. (2015). Exosome-delivered microRNAs modulate the inflammatory response to endotoxin. Nature Communications, 6, 7321. https://doi.org/10.1038/ncomms8321
Phinney, D. G., et al. (2015). Mesenchymal stem cells use extracellular vesicles to outsource mitophagy and shuttle microRNAs. Nature Communications, 6, 8472. https://doi.org/10.1038/ncomms9472
Chen, X., Liang, H., Zhang, J., Zen, K., & Zhang, C. Y. (2013). microRNAs are ligands of Toll-like receptors. RNA, 19, 737–739. https://doi.org/10.1261/rna.036319.112
Fabbri, M., et al. (2012). MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. The Proceedings of the National Academy of Sciences, 109, E2110-2116. https://doi.org/10.1073/pnas.1209414109
Zhang, Y., et al. (2013). Circular intronic long noncoding RNAs. Molecular Cell, 51, 792–806. https://doi.org/10.1016/j.molcel.2013.08.017
Li, Z., et al. (2015). Exon-intron circular RNAs regulate transcription in the nucleus. Nature Structural & Molecular Biology, 22, 256–264. https://doi.org/10.1038/nsmb.2959
Devaux, Y., et al. (2017). Circular RNAs in heart failure. European Journal of Heart Failure, 19, 701–709. https://doi.org/10.1002/ejhf.801
Hansen, T. B., et al. (2011). miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO Journal, 30, 4414–4422. https://doi.org/10.1038/emboj.2011.359
Memczak, S., et al. (2013). Circular RNAs are a large class of animal RNAs with regulatory potency. Nature, 495, 333–338. https://doi.org/10.1038/nature11928
Khan, M. A., et al. (2016). RBM20 regulates circular RNA production from the titin gene. Circulation Research, 119, 996–1003. https://doi.org/10.1161/circresaha.116.309568
Salmena, L., Poliseno, L., Tay, Y., Kats, L., & Pandolfi, P. P. (2011). A ceRNA hypothesis: The Rosetta Stone of a hidden RNA language? Cell, 146, 353–358. https://doi.org/10.1016/j.cell.2011.07.014
Filippova, E. A., et al. (2021). Long noncoding RNA GAS5 in breast cancer: Epigenetic mechanisms and biological functions. International Journal of Molecular Sciences, 22,. https://doi.org/10.3390/ijms22136810
Wei, W., Chen, Y., Gao, J., & Li, C. (2017). WITHDRAWN: The lncRNA-GAS5/miR-23a/Foxo3a axis regulates cardiac hypertrophy by Wnt/β-catenin signal pathway. Biochemical and Biophysical Research Communications, 494, 424. https://doi.org/10.1016/j.bbrc.2017.09.036
Zhang, Z., et al. (2013). Negative regulation of lncRNA GAS5 by miR-21. Cell Death and Differentiation, 20, 1558–1568. https://doi.org/10.1038/cdd.2013.110
Sahoo, S., & Losordo, D. W. (2014). Exosomes and cardiac repair after myocardial infarction. Circulation Research, 114, 333–344. https://doi.org/10.1161/CIRCRESAHA.114.300639
Sluijter, J. P., Verhage, V., Deddens, J. C., van den Akker, F., & Doevendans, P. A. (2014). Microvesicles and exosomes for intracardiac communication. Cardiovascular Research, 102, 302–311. https://doi.org/10.1093/cvr/cvu022
Feldman, A., et al. (2017). Analysis of circulating miR-1, miR-23a, and miR-26a in atrial fibrillation patients undergoing coronary bypass artery grafting surgery. Annals of Human Genetics, 81, 99–105. https://doi.org/10.1111/ahg.12188
Harling, L., et al. (2017). Elevated serum microRNA 483–5p levels may predict patients at risk of post-operative atrial fibrillation. European Journal of Cardio-Thoracic Surgery, 51, 73–78. https://doi.org/10.1093/ejcts/ezw245
Xiao, J., et al. (2011). MicroRNA expression signature in atrial fibrillation with mitral stenosis. Physiological Genomics, 43, 655–664. https://doi.org/10.1152/physiolgenomics.00139.2010
Bostjancic, E., Zidar, N., Stajner, D., & Glavac, D. (2010). MicroRNA miR-1 is up-regulated in remote myocardium in patients with myocardial infarction. Folia Biologica (Praha), 56, 27–31.
Liu, Y., Zou, J., Liu, X., & Zhang, Q. (2019). MicroRNA-138 attenuates myocardial ischemia reperfusion injury through inhibiting mitochondria-mediated apoptosis by targeting HIF1-alpha. Experimental and Therapeutic Medicine, 18, 3325–3332. https://doi.org/10.3892/etm.2019.7976
Wang, R., Li, N., Zhang, Y., Ran, Y., & Pu, J. (2011). Circulating microRNAs are promising novel biomarkers of acute myocardial infarction. Internal Medicine, 50, 1789–1795. https://doi.org/10.2169/internalmedicine.50.5129
Xue, Y., Fan, X., Yang, R., Jiao, Y., & Li, Y. (2020). miR-29b-3p inhibits post-infarct cardiac fibrosis by targeting FOS. Bioscience Reports, 40,. https://doi.org/10.1042/BSR20201227
Yuan, J., et al. (2017). Mir-21 promotes cardiac fibrosis after myocardial infarction via targeting Smad7. Cellular Physiology and Biochemistry, 42, 2207–2219. https://doi.org/10.1159/000479995
Eryilmaz, U., et al. (2016). Circulating microRNAs in patients with ST-elevation myocardial infarction. Anatolian Journal of Cardiology, 16, 392–396. https://doi.org/10.5152/AnatolJCardiol.2015.6603
Tang, C. M., et al. (2017). CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Science and Reports, 7, 40342. https://doi.org/10.1038/srep40342
Care, A., et al. (2007). MicroRNA-133 controls cardiac hypertrophy. Nature Medicine, 13, 613–618. https://doi.org/10.1038/nm1582
Li, D., et al. (2013). Transcriptome analysis reveals distinct patterns of long noncoding RNAs in heart and plasma of mice with heart failure. PLoS ONE, 8, e77938. https://doi.org/10.1371/journal.pone.0077938
Lim, T. B., et al. (2019). Targeting the highly abundant circular RNA circSlc8a1 in cardiomyocytes attenuates pressure overload induced hypertrophy. Cardiovascular Research, 115, 1998–2007. https://doi.org/10.1093/cvr/cvz130
Bang, C., et al. (2014). Cardiac fibroblast-derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. The Journal of Clinical Investigation, 124, 2136–2146. https://doi.org/10.1172/JCI70577
Heymans, S., et al. (2013). Macrophage microRNA-155 promotes cardiac hypertrophy and failure. Circulation, 128, 1420–1432. https://doi.org/10.1161/CIRCULATIONAHA.112.001357
Fang, X., et al. (2016). Adipocyte-specific loss of PPARgamma attenuates cardiac hypertrophy. JCI Insight, 1, e89908. https://doi.org/10.1172/jci.insight.89908
Goren, Y., et al. (2014). Relation of reduced expression of MiR-150 in platelets to atrial fibrillation in patients with chronic systolic heart failure. American Journal of Cardiology, 113, 976–981. https://doi.org/10.1016/j.amjcard.2013.11.060
Rooij, E. V., et al. (2007). Control of stress-dependent cardiac growth and gene expression by a microRNA. Science, 316, 575–579. https://doi.org/10.1126/science.1139089
Li, D., et al. (2010). MicroRNA-125a/b-5p inhibits endothelin-1 expression in vascular endothelial cells. Journal of Hypertension, 28, 1646–1654. https://doi.org/10.1097/HJH.0b013e32833a4922
Fukushima, Y., Nakanishi, M., Nonogi, H., Goto, Y., & Iwai, N. (2011). Assessment of plasma miRNAs in congestive heart failure. Circulation Journal, 75, 336–340. https://doi.org/10.1253/circj.cj-10-0457
Hosen, M. R., et al. (2021). CAD increases the long noncoding RNA PUNISHER in small extracellular vesicles and regulates endothelial cell function via vesicular shuttling. Molecular Therapy Nucleic Acids, 25, 388–405. https://doi.org/10.1016/j.omtn.2021.05.023
Chen, L., et al. (2017). Exosomal lncRNA GAS5 regulates the apoptosis of macrophages and vascular endothelial cells in atherosclerosis. PLoS ONE, 12, e0185406. https://doi.org/10.1371/journal.pone.0185406
Garcia, N. A., Ontoria-Oviedo, I., Gonzalez-King, H., Diez-Juan, A., & Sepulveda, P. (2015). Glucose starvation in cardiomyocytes enhances exosome secretion and promotes angiogenesis in endothelial cells. PLoS ONE, 10, e0138849. https://doi.org/10.1371/journal.pone.0138849
Roger, V. L., et al. (2012). Heart disease and stroke statistics–2012 update: A report from the American Heart Association. Circulation, 125, e2–e220. https://doi.org/10.1161/CIR.0b013e31823ac046
Ong, S. G., et al. (2014). Cross talk of combined gene and cell therapy in ischemic heart disease: Role of exosomal microRNA transfer. Circulation, 130, S60-69. https://doi.org/10.1161/CIRCULATIONAHA.113.007917
Ong, S. G., & Hausenloy, D. J. (2012). Hypoxia-inducible factor as a therapeutic target for cardioprotection. Pharmacology & Therapeutics, 136, 69–81. https://doi.org/10.1016/j.pharmthera.2012.07.005
Le, T., & Chong, J. (2016). Cardiac progenitor cells for heart repair. Cell Death Discov, 2, 16052. https://doi.org/10.1038/cddiscovery.2016.52
Chan, Y. C., Banerjee, J., Choi, S. Y., & Sen, C. K. (2012). miR-210: The master hypoxamir. Microcirculation, 19, 215–223. https://doi.org/10.1111/j.1549-8719.2011.00154.x
Katare, R., et al. (2011). Transplantation of human pericyte progenitor cells improves the repair of infarcted heart through activation of an angiogenic program involving micro-RNA-132. Circulation Research, 109, 894–906. https://doi.org/10.1161/CIRCRESAHA.111.251546
Barile, L., et al. (2014). Extracellular vesicles from human cardiac progenitor cells inhibit cardiomyocyte apoptosis and improve cardiac function after myocardial infarction. Cardiovascular Research, 103, 530–541. https://doi.org/10.1093/cvr/cvu167
Zhao, Y., et al. (2007). Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell, 129, 303–317. https://doi.org/10.1016/j.cell.2007.03.030
Liu, N., et al. (2008). microRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Genes & Development, 22, 3242–3254. https://doi.org/10.1101/gad.1738708
Ke-Gang, J., et al. (2016). Evaluating diagnostic and prognostic value of plasma miRNA133a in acute chest pain patients undergoing coronary angiography. Medicine (Baltimore), 95, e3412. https://doi.org/10.1097/MD.0000000000003412
Zhu, H., & Fan, G. C. (2011). Extracellular/circulating microRNAs and their potential role in cardiovascular disease. American Journal of Cardiovascular Diseases, 1, 138–149.
Jaffe, A. S., et al. (2000). It’s time for a change to a troponin standard. Circulation, 102, 1216–1220. https://doi.org/10.1161/01.cir.102.11.1216
Liu, X., et al. (2015). Plasma miR-1, miR-208, miR-499 as potential predictive biomarkers for acute myocardial infarction: An independent study of Han population. Experimental Gerontology, 72, 230–238. https://doi.org/10.1016/j.exger.2015.10.011
Corsten, M. F., et al. (2010). Circulating MicroRNA-208b and MicroRNA-499 reflect myocardial damage in cardiovascular disease. Circulation. Cardiovascular Genetics, 3, 499–506. https://doi.org/10.1161/CIRCGENETICS.110.957415
Hammerer-Lercher, A., Namdar, M., & Vuilleumier, N. (2020). Emerging biomarkers for cardiac arrhythmias. Clinical Biochemistry, 75, 1–6. https://doi.org/10.1016/j.clinbiochem.2019.11.012
Sustr, F., Starek, Z., Soucek, M., & Novak, J. (2020). Non-coding RNAs and cardiac arrhythmias. Advances in Experimental Medicine and Biology, 1229, 287–300. https://doi.org/10.1007/978-981-15-1671-9_17
Kleeberger, J. A., Tomsits, P. J., Kaab, S., & Clauss, S. (2020). Non-coding RNA and cardiac electrophysiological disorders. Advances in Experimental Medicine and Biology, 1229, 301–310. https://doi.org/10.1007/978-981-15-1671-9_18
Shaihov-Teper, O., et al. (2021). Extracellular vesicles from epicardial fat facilitate atrial fibrillation. Circulation, 143, 2475–2493. https://doi.org/10.1161/CIRCULATIONAHA.120.052009
Ye, Q., et al. (2021). MicroRNA-146b-5p promotes atrial fibrosis in atrial fibrillation by repressing TIMP4. Journal of Cellular and Molecular Medicine, 25, 10543–10553. https://doi.org/10.1111/jcmm.16985
Yao, L., Zhou, B., You, L., Hu, H., & Xie, R. (2020). LncRNA MIAT/miR-133a-3p axis regulates atrial fibrillation and atrial fibrillation-induced myocardial fibrosis. Molecular Biology Reports, 47, 2605–2617. https://doi.org/10.1007/s11033-020-05347-0
Asila, A., Yang, X., Kaisaer, Y., & Ma, L. (2021). SNHG16/miR-485-5p/BMP7 axis modulates osteogenic differentiation of human bone marrow-derived mesenchymal stem cells. The Journal of Gene Medicine, 23, e3296. https://doi.org/10.1002/jgm.3296
Zhao, Y., Yuan, Y., & Qiu, C. (2016). Underexpression of CACNA1C caused by overexpression of microRNA-29a underlies the pathogenesis of atrial fibrillation. Medical Science Monitor, 22, 2175–2181. https://doi.org/10.12659/msm.896191
Shimizu, I., & Minamino, T. (2016). Physiological and pathological cardiac hypertrophy. Journal of Molecular and Cellular Cardiology, 97, 245–262. https://doi.org/10.1016/j.yjmcc.2016.06.001
Nie, X., et al. (2018). miR-217 promotes cardiac hypertrophy and dysfunction by targeting PTEN. Mol Ther Nucleic Acids, 12, 254–266. https://doi.org/10.1016/j.omtn.2018.05.013
Li, H., et al. (2015). Identification of cardiac-related circulating microRNA profile in human chronic heart failure. Oncotarget, 7,
Cai, B., et al. (2015). Mesenchymal stem cells and cardiomyocytes interplay to prevent myocardial hypertrophy. Stem Cells Translational Medicine, 4, 1425–1435. https://doi.org/10.5966/sctm.2015-0032
Kelkar, A. A., et al. (2015). Mechanisms contributing to the progression of ischemic and nonischemic dilated cardiomyopathy: Possible modulating effects of paracrine activities of stem cells. Journal of the American College of Cardiology, 66, 2038–2047. https://doi.org/10.1016/j.jacc.2015.09.010
Chen, F., et al. (2020). Bone marrow mesenchymal stem cell-derived exosomes attenuate cardiac hypertrophy and fibrosis in pressure overload induced remodeling. In Vitro Cellular and Developmental Biology. Animal, 56, 567–576. https://doi.org/10.1007/s11626-020-00481-2
Zhang, S., et al. (2019). miR-29a attenuates cardiac hypertrophy through inhibition of PPARdelta expression. Journal of Cellular Physiology, 234, 13252–13262. https://doi.org/10.1002/jcp.27997
Zhang, S., et al. (2019). miR-29a attenuates cardiac hypertrophy through inhibition of PPARδ expression. Journal of Cellular Physiology, 234, 13252–13262. https://doi.org/10.1002/jcp.27997
Johnson, F. L. (2014). Pathophysiology and etiology of heart failure. Cardiol Clin, 32, 9–19. https://doi.org/10.1016/j.ccl.2013.09.015 vii.
Qiao, L., et al. (2019). microRNA-21-5p dysregulation in exosomes derived from heart failure patients impairs regenerative potential. The Journal of Clinical Investigation, 129, 2237–2250. https://doi.org/10.1172/JCI123135
Kioka, N., Ueda, K., & Amachi, T. (2002). Vinexin, CAP/ponsin, ArgBP2: A novel adaptor protein family regulating cytoskeletal organization and signal transduction. Cell Structure and Function, 27, 1–7. https://doi.org/10.1247/csf.27.1
Cheng, H., et al. (2010). Loss of enigma homolog protein results in dilated cardiomyopathy. Circulation Research, 107, 348–356. https://doi.org/10.1161/circresaha.110.218735
Duygu, B., de Windt, L. J., & de Costa Martins, P. A. (2016). Targeting microRNAs in heart failure. Trends in Cardiovascular Medicine, 26, 99–110. https://doi.org/10.1016/j.tcm.2015.05.008
Patrick, D. M., et al. (2010). Stress-dependent cardiac remodeling occurs in the absence of microRNA-21 in mice. The Journal of Clinical Investigation, 120, 3912–3916. https://doi.org/10.1172/JCI43604
Wu, T., et al. (2018). Serum exosomal MiR-92b-5p as a potential biomarker for acute heart failure caused by dilated cardiomyopathy. Cellular Physiology and Biochemistry, 46, 1939–1950. https://doi.org/10.1159/000489383
Klenke, S., et al. (2018). Circulating miR-192 is a prognostic marker in patients with ischemic cardiomyopathy. Future Cardiology, 14, 283–289. https://doi.org/10.2217/fca-2017-0108
Matsumoto, S., et al. (2013). Circulating p53-responsive microRNAs are predictive indicators of heart failure after acute myocardial infarction. Circulation Research, 113, 322–326. https://doi.org/10.1161/CIRCRESAHA.113.301209
Halkein, J., et al. (2013). MicroRNA-146a is a therapeutic target and biomarker for peripartum cardiomyopathy. The Journal of Clinical Investigation, 123, 2143–2154. https://doi.org/10.1172/JCI64365
Boudina, S., & Abel, E. D. (2010). Diabetic cardiomyopathy, causes and effects. Reviews in Endocrine & Metabolic Disorders, 11, 31–39. https://doi.org/10.1007/s11154-010-9131-7
Renu, K., V G, A., P B, T. P., & Arunachalam, S. (2018). Molecular mechanism of doxorubicin-induced cardiomyopathy – An update. European Journal of Pharmacology, 818, 241–253. https://doi.org/10.1016/j.ejphar.2017.10.043
Xia, W., Chen, H., Xie, C., & Hou, M. (2020). Long-noncoding RNA MALAT1 sponges microRNA-92a-3p to inhibit doxorubicin-induced cardiac senescence by targeting ATG4a. Aging (Albany NY), 12, 8241–8260. https://doi.org/10.18632/aging.103136
Rogg, E.-M., et al. (2018). Analysis of cell type-specific effects of MicroRNA-92a provides novel insights into target regulation and mechanism of action. Circulation, 138, 2545–2558. https://doi.org/10.1161/CIRCULATIONAHA.118.034598
Rafieian-Kopaei, M., Setorki, M., Doudi, M., Baradaran, A., & Nasri, H. (2014). Atherosclerosis: Process, indicators, risk factors and new hopes. International Journal of Preventive Medicine, 5, 927–946.
Chen, L., et al. (2016). Global transcriptomic study of atherosclerosis development in rats. Gene, 592, 43–48. https://doi.org/10.1016/j.gene.2016.07.023
Brill, A., Dashevsky, O., Rivo, J., Gozal, Y., & Varon, D. (2005). Platelet-derived microparticles induce angiogenesis and stimulate post-ischemic revascularization. Cardiovascular Research, 67, 30–38. https://doi.org/10.1016/j.cardiores.2005.04.007
Cohn, J. N. (2018). Cardiovascular disease progression: A target for therapy? The American Journal of Medicine, 131, 1170–1173. https://doi.org/10.1016/j.amjmed.2018.03.032
Wang, K., et al. (2017). Circular RNA mediates cardiomyocyte death via miRNA-dependent upregulation of MTP18 expression. Cell Death and Differentiation, 24, 1111–1120. https://doi.org/10.1038/cdd.2017.61
Jansen, F., et al. (2014). MicroRNA expression in circulating microvesicles predicts cardiovascular events in patients with coronary artery disease. Journal of the American Heart Association, 3, e001249. https://doi.org/10.1161/JAHA.114.001249
Kuwabara, Y., et al. (2011). Increased microRNA-1 and microRNA-133a levels in serum of patients with cardiovascular disease indicate myocardial damage. Circulation. Cardiovascular Genetics, 4, 446–454. https://doi.org/10.1161/CIRCGENETICS.110.958975
Cakmak, H. A., & Demir, M. (2020). MicroRNA and cardiovascular diseases. Balkan Medical Journal, 37, 60–71. https://doi.org/10.4274/balkanmedj.galenos.2020.2020.1.94
Boon, R. A., & Dimmeler, S. (2015). MicroRNAs in myocardial infarction. Nature Reviews. Cardiology, 12, 135–142. https://doi.org/10.1038/nrcardio.2014.207
Endo, K., et al. (2013). MicroRNA 210 as a biomarker for congestive heart failure. Biological &/and Pharmaceutical Bulletin, 36, 48–54. https://doi.org/10.1248/bpb.b12-00578
Lu, Y., et al. (2009). MicroRNA-1 downregulation by propranolol in a rat model of myocardial infarction: A new mechanism for ischaemic cardioprotection. Cardiovascular Research, 84, 434–441. https://doi.org/10.1093/cvr/cvp232
Poller, W., et al. (2018). Non-coding RNAs in cardiovascular diseases: Diagnostic and therapeutic perspectives. European Heart Journal, 39, 2704–2716. https://doi.org/10.1093/eurheartj/ehx165
Ventura, A., & Jacks, T. (2009). MicroRNAs and cancer: Short RNAs go a long way. Cell, 136, 586–591. https://doi.org/10.1016/j.cell.2009.02.005
Wang, H., et al. (2021). Percutaneous intracoronary delivery of plasma extracellular vesicles protects the myocardium against ischemia-reperfusion injury in Canis. Hypertension, 78, 1541–1554. https://doi.org/10.1161/HYPERTENSIONAHA.121.17574
Lane, R. E., Korbie, D., Hill, M. M., & Trau, M. (2018). Extracellular vesicles as circulating cancer biomarkers: Opportunities and challenges. Clinical and Translational Medicine, 7, 14. https://doi.org/10.1186/s40169-018-0192-7
Yekula, A., et al. (2020). From laboratory to clinic: Translation of extracellular vesicle based cancer biomarkers. Methods, 177, 58–66. https://doi.org/10.1016/j.ymeth.2020.02.003
Wang, X., et al. (2018). Engineered exosomes with ischemic myocardium‐targeting peptide for targeted therapy in myocardial infarction. Journal of the American Heart Association, 7, e008737. https://doi.org/10.1161/JAHA.118.008737
Mentkowski, K. I., & Lang, J. K. (2019). Exosomes engineered to express a cardiomyocyte binding peptide demonstrate improved cardiac retention in vivo. Science and Reports, 9, 10041. https://doi.org/10.1038/s41598-019-46407-1
Kim, H., et al. (2018). Cardiac-specific delivery by cardiac tissue-targeting peptide-expressing exosomes. Biochemical and Biophysical Research Communications, 499, 803–808. https://doi.org/10.1016/j.bbrc.2018.03.227
Gee, P., et al. (2020). Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping. Nature Communications, 11, 1334. https://doi.org/10.1038/s41467-020-14957-y
Campbell, L. A., et al. (2019). Gesicle-mediated delivery of CRISPR/Cas9 ribonucleoprotein complex for inactivating the HIV provirus. Molecular Therapy, 27, 151–163. https://doi.org/10.1016/j.ymthe.2018.10.002
Rodrigues, M., Fan, J., Lyon, C., Wan, M., & Hu, Y. (2018). Role of extracellular vesicles in viral and bacterial infections: Pathogenesis, diagnostics, and therapeutics. Theranostics, 8, 2709–2721. https://doi.org/10.7150/thno.20576
Meliani, A., et al. (2017). Enhanced liver gene transfer and evasion of preexisting humoral immunity with exosome-enveloped AAV vectors. Blood Advances, 1, 2019–2031. https://doi.org/10.1182/bloodadvances.2017010181
Ju, Y., Bai, H., Ren, L., & Zhang, L. (2021). The role of exosome and the ESCRT pathway on enveloped virus infection. International Journal of Molecular Sciences, 22, 9060. https://doi.org/10.3390/ijms22169060
Sahoo, S., et al. (2011). Exosomes from human CD34(+) stem cells mediate their proangiogenic paracrine activity. Circulation Research, 109, 724–728. https://doi.org/10.1161/CIRCRESAHA.111.253286
Beltrami, C., et al. (2017). Human pericardial fluid contains exosomes enriched with cardiovascular-expressed microRNAs and promotes therapeutic angiogenesis. Molecular Therapy, 25, 679–693. https://doi.org/10.1016/j.ymthe.2016.12.022
Lai, R. C., et al. (2010). Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell Research, 4, 214–222. https://doi.org/10.1016/j.scr.2009.12.003
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Associate Editor Junjie Xiao oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhao, Z., Guo, N., Chen, W. et al. Leveraging Extracellular Non-coding RNAs to Diagnose and Treat Heart Diseases. J. of Cardiovasc. Trans. Res. 15, 456–468 (2022). https://doi.org/10.1007/s12265-022-10252-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-022-10252-x