Abstract
Primates exhibit complex brain structures that augment cognitive function. The neocortex fulfills high-cognitive functions through billions of connected neurons. These neurons have distinct transcriptomic, morphological, and electrophysiological properties, and their connectivity principles vary. These features endow the primate brain atlas with a multimodal nature. The recent integration of next-generation sequencing with modified patch-clamp techniques is revolutionizing the way to census the primate neocortex, enabling a multimodal neuronal atlas to be established in great detail: (1) single-cell/single-nucleus RNA-seq technology establishes high-throughput transcriptomic references, covering all major transcriptomic cell types; (2) patch-seq links the morphological and electrophysiological features to the transcriptomic reference; (3) multicell patch-clamp delineates the principles of local connectivity. Here, we review the applications of these technologies in the primate neocortex and discuss the current advances and tentative gaps for a comprehensive understanding of the primate neocortex.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The mammalian neocortex is responsible for high-cognitive function and fine motor skills. The neocortex fulfills these functions through complicated networks of diverse neurons. Studies using rodents have established a basic framework of the neocortex, censused the transcriptomes of all the major cell types, and linked them to their physiological properties and principles of connection. Despite its augmented cognitive function, the primate neocortex shares the basic neuronal program with rodents. Accumulating evidence reveals the divergence between rodents and primates [1,2,3]. However, the distinctions are subtle. A multimodal census of the neurons of the primate neocortex is necessary to underpin evolutionary changes that augment the capacities of the primate neocortex.
Primates exhibit complex brain structures that augment cognitive function. However, the volume and number of neocortical neurons increased rapidly compared to subcortical structures during the evolutionary expansion of the neocortex [4,5,6,7,8]. While the general principles of cortical development and basic architecture are conserved, studies have shown differences in the cellular composition of the human cortex [9]. These differences include the expansion of superficial cortical layers during mammalian evolution, which may involve rare cell types and novel cellular interactions contributing to the complexity of primate brain function [10]. Notably, neurons such as von Economo [11] and rosehip neurons [12], which have unique morphological features, are primate-specific and do not exist in mice. These neurons may be involved in various cognitive processes, including facilitating rapid information transmission across different brain regions and promoting the integration of sensory, emotional, and mental information. In addition, there are transcriptional differences between mice, non-human primates, and humans, particularly in genes related to neuronal structure and function [2, 13,14,15]. For example, glutamatergic neuron transcriptome types are more diverse in the supragranular layer of the human neocortex [16]. The development of novel research technologies and massive high-throughput studies provide valuable resources for understanding the foundation of the augmented cognitive capacities of the primate brain.
First, it is crucial to systematically investigate the cell type composition within primate cortical areas. Since Frederick Sanger invented Sanger sequencing in 1977, when we could read the genetic code for the first time, it has taken several decades and significant efforts to promote the development of sequencing technology [17]. Due to its cost, sequencing became the conventional technology used in regular research until next-generation sequencing methods were announced in 2005, followed by the invention of the single-cell RNA-seq (scRNA-seq) method [18]. However, due to the limited application of scRNA-seq to frozen tissue with cell membranes ruptured during freezing, an essential complementary technology was developed that involved isolating a single nucleus sequencing the RNA (snRNA-seq) [19, 20]. This approach initiated a new chapter for investigating the cell type composition of primate cortical areas [21]. Previous neuronal classification was based on morphology, electrophysiology, or some specific molecules [22], while sc/snRNA-seq provides high-throughput analysis and single-cell resolution for the robust classification of cell types and generating a transcriptomic cellular atlas [23]. By comparing the cell type composition across species, primate-specific cell type composition and proportions might be evaluated to help explain the more complex brain functions of primates [24]. After generating a transcriptomic reference cellular atlas [25,26,27,28,29,30,31,32,33,34,35,36,37], thorough gene expression analysis might improve understanding of cell differentiation trajectories by identifying abundant related pathways and genes, which could serve as candidate targets for interventions targeting neurodevelopmental disorders [38,39,40,41,42,43,44,45]. When comparing physiological and pathological states, sc/snRNA-seq helps identify the modification of the composition and proportions of cell types with related pathogenic signaling pathways or genes, which can not only reveal the molecular mechanisms of pathological processes but also provide abundant promising targets for therapeutic protection and intervention [17, 46,47,48,49].
However, because of the multiple dimensions of the neuron, which is the basic unit of the nervous system, integrating the wealth of transcriptomic data with well-established morphological and electrophysiological data is still required [50]. Since Neher and Sakmann invented patch-clamp technology to study ionic currents at the single-cell level, this method has become the standard for investigating electrophysiology and morphology in single cells, especially neurons [51]. Genetically-labeled cells facilitate the study of the morphology and electrophysiology of neurons [52,53,54,55]. However, the application of these techniques is limited by the availability of known cell type-specific markers and the operational feasibility of primate experiments. Recently, several groups have developed and optimized Patch-seq, a multimodal method that describes electrophysiological, transcriptomic, and morphological profiles in single neurons of rodents [56,57,58,59,60] and adult human and non-human primate brain slices [16, 59, 61,62,63,64,65]. By combining this technology with innovative data analytical tools [66,67,68], neuroscientists can map the Patch-seq data to the transcriptomic reference atlas for assigning morphological and electrophysiologic annotations to enrich the transcriptomic cellular atlas to improve a comprehensive understanding of the primate cortical areas during physiological or pathological states [69,70,71,72,73].
After generating a transcriptomic cellular atlas with electrophysiological and morphological annotations, the next step is to analyze the connectivity principles within cortical areas. However, the currently popular technologies that identify neuronal types typically need more morphological data and heavily rely on genetic manipulation, which is challenging in primate experiments. To decipher the principles of local connectivity, a high-throughput technology and robust cell classification standards are critical [74,75,76,77,78]. Simultaneous multiple whole-cell patch-clamp recordings, such as dual, triple, and quadruple recordings, have proven invaluable in facilitating the study of connectivity between neurons [79, 80]. The number of test potential connections is significantly increased with simultaneous patch-clamp recording neurons. Multicell patch-clamp setups with up to 12 simultaneously recorded neurons were achieved before 2011 [81,82,83]. The stable simultaneous octuple patch-clamp recording technology achieved superior multicell patch-clamp recording results [84, 85]. Because it provided highly detailed morphological data from neurons for neuronal type identification and offered high-throughput evaluation of potential connections, it is optimally suited for primate research on connectivity and addresses concerns regarding the considerable workload in primate studies and the scarcity of primate tissues.
Here, we review the advances in primate neuroscience from the applications of the above three advanced technologies and discuss the potential of integrating the wealth of datasets obtained using those technologies to generate a primate brain atlas with multiple dimensions (describing transcriptomic, electrophysiological, and morphological profiles, as well as the principles of local connectivity) for comprehensively understanding the functional mechanisms of primate cortical areas. While sc/snRNA-seq can be used to generate a transcriptomic reference atlas of primate cortical areas with transcriptome-based classification, Patch-seq can provide morphological and electrophysiological annotations for the above transcriptomic reference atlas. In addition, the multicell patch-clamp measures the strength of monosynaptic connections between cells. These physiological properties can be mapped to the transcriptomic reference atlas, depicting a multimodal atlas of the primate brain, and facilitating the advancement of our knowledge in neuroscience.
Large-scale Transcriptomic Analysis Advances Primate Neuroscience
Single-cell or single-nucleus RNA-seq technologies, which have high throughput and robust resolution for cell type classification with gene expression analysis during both physiological and pathological states, have provided an excellent opportunity for the in-depth exploration of the primate brain [18, 19, 86,87,88] (Fig. 1B−E).
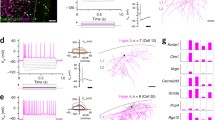
Schematic workflow of the three technologies for investigating the multimodal nature of the primate brain atlas. A Abridged general view of the organization pattern of the target area. B–E Schematic of a single-cell/single-nucleus RNA-seq experiment. B Preparation of single-cell/single-nucleus suspension. C Microfluidic device pairs individual cells or nuclei with barcoded beads that collect and barcode the cell's mRNAs in droplets. D The above droplets are then broken and reverse transcribed, amplified, and sequenced to generate a single-cell/single-nucleus RNA-seq dataset (upper). E This dataset establishes the transcriptomic reference atlas with transcriptome-based classification. D–G Schematic of Patch-seq experiment. F Patch-seq simultaneously performs electrophysiological recording on the acute slice and dye penetration for further staining and morphological reconstruction. G After electrophysiological recording, most of the cytoplasmic contents are aspirated using the patch pipette and transported into a lysis buffer. This pretreatment is followed by a standard single-cell RNA-seq protocol (D), including reverse transcription, amplification, and sequencing, and generates the Patch-seq sample (lower); the gene expression profile of the Patch-seq sample can be used to map the cell onto the above transcriptomic reference atlas (the red point in E), thus assigning electrophysiological and morphological annotations to the transcriptomically identified cell type. E, H–I Schematic of multicell patch-clamp experiments. H Connectivity diagram of the eight simultaneously-recorded neurons (triangles, excitatory neurons; circles, inhibitory neurons); In an octuple patch-clamp recording, 56 potential connections are tested (eight neurons are stimulated successively (left), and postsynaptic potentials of the other seven neurons are recorded for testing neuronal connections (right)). I Based on the morphological reconstruction and classification, connectivity principles within the selected cortical area can be delineated (black lines, excitatory connections; red lines, inhibitory connections; line thickness, connection probability). The dataset from multicell patch-clamp recording possesses the potential of mapping those connectivity principles into the above transcriptomic reference based on the shared morphological and electrophysiological message (dotted line with arrow in E).
The Taxonomy of Transcriptomic Cell Types
Accumulating research on the rodent and primate cortex at single-cell resolution suggested that the cell types are largely conserved across species. The neocortical brain cells of primates are commonly analyzed to generate a taxonomy of cell types according to transcriptomic similarity. Brain cells are grouped into neuronal cells and non-neuronal cells. Neuronal cells are usually divided into glutamatergic excitatory neurons and GABAergic inhibitory neurons [89]. Each class can be further divided into multiple subclasses (Table 1). In the primate neocortex, such as the human middle temporal gyrus and macaque primary visual cortex, the excitatory neurons have been categorized into different subclasses based on the laminar distribution, transcriptome type analysis, and the transcriptomic homology to well-established datasets. These subclasses include intratelencephalic cell subclasses (L2/3 IT, L4 IT/IT, L5 IT, L6 IT, and L6 IT Car3), the L5 extratelencephalic cell subclass (L5 ET), the L6 corticothalamic cell subclass (L6 CT), the near-projecting cell subclass (L5/6 NP), and the L6b cell subclass [10, 89,90,91,92,93]. Alternatively, in a different system, based on the expression of marker genes, excitatory neurons are also divided into four subclasses: CUX2-expressing cells (or LINC00507 [90], or HPCAL, combined with the application of the marker NXPH4 to distinguish upper neurons from L6b neurons[91]) that are mainly located in the upper layers, RORB-expressing cells (enriched in layer 4 but can be found across all layers), FEZF2-expressing cells (located in deep layers), and THEMIS-expressing cells (located in deep layers). The GABAergic inhibitory class contains four main subclasses (LAMP5, VIP, SST, and PVALB-expressing) [94, 95]), essentially correspond to their developmental origin in the medial ganglionic eminence (MGE: PVALB and SST subclasses) or caudal ganglionic eminence (CGE; LAMP5 and VIP subclasses) [15, 96, 97]. There are additional subclasses such as the PAX6 [91, 98], LAMP5 LHX6 [10, 91, 98], and PAX6 ADARB2 (SNCG) [90, 98] subclasses originating from the CGE, and the PVALB UNC5B (Chandelier) [90, 98] and SST CHODL [90, 98] subclasses originating from the MGE. Non-neuronal brain cells are grouped into six subclasses: astrocytes, oligodendrocyte precursor cells, oligodendrocytes, microglia, and perivascular macrophages, endothelial cells, and vascular leptomeningeal cells [90, 91, 93, 94, 96, 97]. These neuronal and non-neuronal subclasses are subdivided into cell types based on additional marker genes.
Cross-Species Transcriptomic Conservation and Divergence
Recent comprehensive cross-species transcriptomic studies that have generated transcriptomic cellular atlases with abundant molecular signatures have revealed surprisingly well-conserved neuronal and non-neuronal types across different cortical areas among primates and rodents [10, 15, 99, 100]. Humans share with rodents an evolutionarily-conserved regulatory program involved in the process of neuronal development, which controls the specification, migration, and differentiation of GABAergic interneurons [101]. However, the primate-specific cell types [12] and the differences in homologous cell types, including proportions, laminar distributions, gene expression, and morphological features, are inestimable [102, 103]. For example, homologous thalamocortical neurons in the primate dorsal lateral geniculate nucleus, which convey visual information from the retina to the primary visual cortex (V1), are distinct from those in rodents [99]. Detailed single-cell transcriptome analysis sampling from non-human primate V1, novel cell types (the NPY-expressing excitatory neuron type and the primate-specific activity-dependent OSTN+ neuron type), and different gene expression patterns have been revealed in the primary visual cortex in primates [91]. These findings may account for the high visual acuity and more complex color vision of primates.
Moreover, comparative RNA sequencing has revealed divergent expression patterns regulating cell morphogenesis, such as ZEB2 (zinc finger E-box binding homeobox 2) and human-specific NOTCH2NL (paralogs of the NOTCH2 receptor). The expression of ZEB2 promotes the neuroepithelial transition and manipulations of the related downstream signaling lead to the acquisition of non-human ape architecture in the human context and vice versa. On the other hand, NOTCH2NL expands cortical progenitors and enhances neuronal output, emphasizing the important role of neuroepithelial cell shape in human brain expansion [104, 105]. These findings suggest that cell type taxonomy is largely conserved from rodents to primates, yet differences exist.
The Transcriptomes During Brain Development and Neurogenesis
The primate brain, which has the largest volume relative to body size and ~1000 times more neurons than the rodent brain [106, 107], is much more complex. Single-cell and single-nucleus technologies have established cellular taxonomies of multiple cortical areas from developing primates using gene expression patterns. This considerably expands our knowledge of early neurogenesis, neuroplasticity, and cellular differentiation during the early developmental stage [108,109,110,111,112,113,114,115,116,117]. Transcriptomic data, which provides cell lineages, molecular signatures, and transcriptional regulatory networks that underlie the basis of physiological activities [118,119,120,121,122], can be used to assess human and non-human primate brain development and early neurogenesis [105, 107, 123, 124]. Moreover, synaptic gene expression patterns show considerable differences in human cortical areas during aging, accounting for the reduced functions of the aging brain [125]. Neurogenesis in adult primates, a recurring and crucial topic of primate neuroscience, has been comprehensively investigated through sc/snRNA-seq transcriptomic data accompanied by sufficient immunostaining evidence [126, 127]. Larger-scale transcriptomic studies have also focused on the diversity of glial cells, including oligodendrocytes and astrocytes [128], which exhibit developmental and metabolic regulation by neuronal activity in the developing human cerebral cortex [129, 130]. This result indicates that the balance of the interaction between glial cells and neurons is important for the normal development of the primate brain.
Transcriptome Changes of Neuropathological State
Single-cell and single-nucleus transcriptomic analyses enable the exploration of cell type composition, transcriptomic modifications, and their role in neurological diseases. These modifications, particularly in critical genes and pathways, regulate disease progression and offer therapeutic opportunities [131, 132]. Such analyses have been extensively used to study various primate neurological disorders, establishing cellular taxonomies, identifying vulnerable cell subpopulations with risk genes, and shedding light on pathogenesis mechanisms and potential therapeutics [133,134,135,136,137,138,139]. In this review, we dived into the detailed research on Alzheimer's disease (AD), autism spectrum disorder (ASD), and multiple sclerosis (MS).
AD is a progressive neurodegenerative disorder characterized by memory loss, cognitive decline, and executive dysfunction [140,141,142]. Analyses of single nuclei from the prefrontal cortex of individuals with AD have identified distinct neuronal and non-neuronal types with pathological gene expression associated with myelination, inflammation, and neuron survival. Disease-associated changes are highly cell-type specific, with some genes (HSP90AA1 and HSPA1A involved in protein folding) universally upregulated in late stages [143]. Transcriptional and pathological differences between sexes have also been reported [139]. Understanding cell-type-specific gene networks and transitions is crucial for unraveling AD pathogenesis. Integrated analysis of transcription factors and AD risk loci have revealed drivers of cell-type-specific transitions. It highlights the repression of AD risk genes in oligodendrocyte progenitor cells, astrocytes, and their upregulation in microglia [144,145,146]. Cell-type-specific vulnerability is a fundamental feature of neurodegenerative diseases in which different cellular populations show a gradient of susceptibility to degeneration. Abundant molecular signatures identified by snRNA-seq provide an unprecedented chance to characterize the specifically vulnerable neuronal subpopulations at the molecular level. In transcriptomic analysis of AD, RORB (RAR Related Orphan Receptor B) has been identified as a marker of selectively vulnerable excitatory neurons. At the same time, the downregulation of genes involved in homeostatic functions is used to characterize vulnerable astrocyte subpopulations [147, 148]. Recent multimodal methodology has identified potential AD treatments [149].
ASD is a neurodevelopmental condition impacting interaction and communication [150]. Recent research investigating 11 cortical areas in individuals with ASD and neurotypical controls has revealed widespread transcriptomic changes across the cortex in individuals with ASD. The findings exhibit an anterior-to-posterior gradient, with the most significant differences in the primary visual cortex. These differences coincide with reduced typical transcriptomic differences between cortical areas in neurotypical individuals [151]. In relation to the cell-type-specific molecular changes associated with ASD, Velmeshev et al. found that synaptic signaling in upper-layer excitatory neurons and the molecular state of microglia are preferentially affected in ASD. Moreover, the dysregulation of specific groups of genes in cortico-cortical projection neurons has been found to correlate with the clinical severity of ASD, as expected [48].
Oligodendrocytes are implicated in the pathogenesis of MS, a multifocal inflammatory disease affecting cortical areas [152]. SnRNA-seq has demonstrated different functional states of oligodendrocyte subpopulations in MS tissue and has identified selectively vulnerable neuronal subpopulations, stressed oligodendrocytes, reactive astrocytes, and activated microglia associated with the progression of MS lesions [153, 154]. Overlapping transcriptional profiles between MS and other neurodegenerative diseases suggest shared mechanisms and potential therapeutic approaches [155].
Linking Transcriptomes with Morphological and Electrophysiological Phenotypes
Single-cell or single-nucleus sequencing data provide valuable insights into the transcriptomic types (t-types) of homology across species [15, 90]. However, it cannot obtain the morphological and electrophysiological properties of the morpho-electrical-transcriptomic types [56, 59]. Patch-seq is a revolutionary technology that can simultaneously acquire the morphology, electrophysiology, and transcriptome of single cells, which are valuable resources to establish a multimodal atlas. Patch-seq is a modification of regular patch-clamp recording. It has been applied to record from cultured cells and acute brain sections in vitro [56,57,58, 60], and the recorded cells are labeled with dye for subsequent morphological reconstruction. After electrophysiological recording, most of the cytoplasmic contents are aspirated (generally including the nucleus) and transferred to an individual tube containing a lysis buffer followed by a standard single-cell or single-nucleus RNA-seq protocol (Fig. 1D−G). This powerful multimodal approach provides valuable resources for advancing our understanding of primate neuroscience.
Patch-seq techniques have been successfully implemented in the primate cortex [16, 59, 61, 62, 64, 90]. Patch-seq-sampled data from neurons across human cortical layers 1, 2, 3, and 5 have been mapped to a human transcriptomic cellular reference atlas and assigned electrophysiological and morphological features to the mapped t-types [16, 61, 64]. This multimodal analysis has identified human-specific double bouquet cells, mapping to two cortical GABAergic somatostatin (SST) t-types (SST CALB1 and SST ADGRG6) [62]. Studies on the human cortex have revealed higher divergence in the upper-layer neocortex compared to mice [16], whereas the cell density in layers 2/3 is lower in humans than in mice [16, 61, 156]. Among human L1 interneurons, subclasses defined by their transcriptomes exhibit similarly distinct morpho-electrical phenotypes. Two human cell types with specialized phenotypes have been identified (MC4R rosehip cells and the bursting PAX6 TNFAIP8L3 t-type) [61]. Indeed, observing a 'rosehip' cell type in human and not mouse neocortex emphasizes the importance of studying human L1 to uncover potential species-specific specializations [12].
In addition, the supragranular layer of the human neocortex exhibits increased diversity in glutamatergic neuron types, predominantly found in layers 2 and 3. Five human supragranular neuron t-types (LTK, GLP2R, FREM3, CARM1P1, and COL22A1) have corresponding morphology, physiology, and transcriptome phenotypes. The more superficially located LTK, GLP2R, and FREM3 types are homologous to the mouse supragranular IT types. The more deeply located CARM1P1 and COL22A1 types do not have direct counterparts in the mouse supragranular neocortex. Instead, they exhibit the closest transcriptomic similarity to infragranular mouse IT types [16]. These results suggest an increased diversity of deep L3 neurons in humans. The deep portion of layer 3 contains highly distinctive cell types, two pyramidal cell types (FREM3+ and CARM1P1+ transcriptomic cell types [15]) expressing neurofilament protein SMI-32 (encoded by the NEFH gene), which labels long-range projection neurons in primates that are selectively depleted in AD [157, 158], providing a promising entry to study the pathological mechanism and explore potential therapeutic options.
In contrast to the well-documented diversity of supragranular-layer excitatory neurons across regions and species, studies addressing the variability of deep-layer excitatory neurons, such as ET, IT, and CT neurons, are limited [90]. Axonal projections to lower brain regions predominantly originate from layer 5 (L5) ET neurons. L5 ET neurons exhibit unique morpho-electric properties, gene expression patterns, local synaptic connections, long-range afferents, and neuromodulatory responses, as primarily described in rodents. L5 ET neurons are traditionally characterized by thick apical dendritic tufts in layer 1, while L5 IT neurons have thinner or tuftless dendrites. In addition, L5 ET neurons strongly express hyperpolarization-activated cyclic nucleotide-gated (HCN) channels, likely contributing to their strong dendritic electrogenesis [64]. Still, in rodents, the HCN conductance tends to dampen dendritic electrogenesis. The differences in HCN expression between L5 ET and IT neurons may not be the primary reason for the differences in electrogenesis [159]. Further interrogations are necessary. These different properties of L5 ET and IT neurons contribute to distinct aspects of perception and behavior. Patch-seq analysis of L5 neurons in the primary motor cortex of both mice and macaques has revealed that macaque and human Betz cells are homologous to the thick-tufted L5 ET neurons in mice but exhibit species-specific differences in morphology, physiology, and gene expression. Macaque and human Betz ET neurons have specialized suprathreshold properties, such as biphasic firing patterns evoked by prolonged suprathreshold current injection [26]. Macaque and human L5 ET neurons are notably larger and possess long "taproot" basal dendrites, characteristic of the iconic Betz cells [160].
Gene expression patterns shape the electrophysiological and morphological phenotypes. Patch-seq provides a multimodal analysis strategy to identify potential molecular markers or pathways that predict the phenotype of single neurons [161, 162]. Using Patch-seq combined with Weighted Gene Co-expression Network Analyses of single human neurons in culture, certain gene clusters are correlated with neuronal maturation as determined by electrophysiological characteristics, and a list of candidate genes has been identified that have the potential to serve as biomarkers of neuronal maturation [163]. For example, Patch-seq recording from human induced pluripotent stem cell (iPSC)-derived astrocytes and neurons has revealed a continuum of low- to high-function electrophysiological states. Furthermore, a novel biomarker, GDAP1L1, effectively identifies the high-functioning neurons. These biomarkers facilitate the classification of neurons based on their functionality and enable the stratification of functional heterogeneity [161].
Subtle differences in the transcriptome may profoundly affect neuronal morphology and function [164]. Patch-seq links transcriptomes with phenotypes such as morphology and electrophysiology. This allows for targeted studies on specific neuronal populations based on factors like anatomical location, functional properties, and lineages. Patch-seq also facilitates studies on the molecular basis of morphological and functional diversity [165]. We can gain new insights into neurons by using Patch-seq to correlate transcriptomes with morphological characteristics, neuronal locations, and projection patterns. The acquisition and integration of transcriptome information is essential to neuronal classification.
Dissecting Local Connectivity Principles in Primate Cortical Areas
Understanding the functional mechanisms of primate cortical areas requires a comprehensive investigation of their cellular and synaptic organization. To enable a detailed understanding of microcircuits, multicell whole-cell patch-clamp recordings still represent the gold standard method [166, 167]. This method reliably detects unitary excitatory and inhibitory synaptic connectivity with its submillisecond and subthreshold resolution [168]. Increasing the number of simultaneously recorded neurons can considerably increase the number of probed synaptic connections and generate larger sample sizes from fewer experiments. For example, if the number of simultaneously patched neurons reaches eight in single slices, 56 potential connections will be tested. It is important to achieve relatively high throughput due to the limited availability of primate samples. By using multicell whole-cell patch-clamp recordings, simultaneous recording of multiple neurons becomes possible. This approach significantly enhances the number of monosynaptic connections tested per experiment, enabling us to explore the principles of connection between different cells (Fig. 1H, I) [83, 169,170,171,172,173].
The recent application of multicell patch-clamp technology to the study of the primate brain has considerably expanded our knowledge of its microcircuits. Recurrent excitatory connectivity is thought to be important in behavior [174] and disease [175], and has also been identified as a common feature in computational models of cortical working memory, receptive field shaping, attractor dynamics, and sequence storage [176,177,178,179,180]. Although there is a wide range of reported rates of recurrent connectivity among excitatory neurons in rodents [173, 181, 182], evidence has shown that the human cortex possesses a higher recurrent excitatory connectivity rate and mean amplitude, which might contribute to the related functional difference across species [183, 184]. The higher mean amplitude and other excitatory postsynaptic differences indicate stronger synaptic connectivity within the human cortex, which is explained by the larger presynaptic active zones and postsynaptic densities that may allow a higher release probability as well as more neurotransmitter release and binding [185, 186].
A more comprehensive survey of intralaminar connectivity has been applied to investigate all cortical layers detailing the connectivity atlas, and the analysis includes synaptic dynamics between layer-defined pyramidal neurons and inhibitory neurons, greatly increasing our understanding of primate neural microcircuits [187]. In this study of the connectivity between layer-defined neuronal types, the connectivity probability among layer 4 was nearly absent in the human cortex. At the same time, it was high in the mouse cortex. Moreover, disynaptic inhibition in the human cortex, which was not detected in the mouse cortex, was found between confirmed spiny pyramidal cells that were unidirectional, originating in layer 2 and targeting other layer 2 or layer 3 pyramidal cells. Consistent with previous results, the connectivity rates were estimated to decline with increasing distance but at a slower speed than in rodents. The unique characteristics of the human circuit findings regarding alterations in synaptic dynamics may help explain the complexity of information in the human cortex.
The morphologies of neurons are diverse. Their dendritic and axonal projections provide additional information for investigating the local connectivity. Recent advances in patch-clamp-based techniques and solution formulations have significantly improved the quality of neuron reconstruction [63], establishing a framework for the morphological classification of rodent neurons [60, 173]. In addition, high-throughput information can be obtained through the advances in imaging and cell labeling techniques, such as superresolution hopping probe ion conductance microscopy, a variant of scanning ion conductance microscopy [188,189,190] and viral tracers or transgenic animals (e.g., Cre-driven lines or tetracycline-controlled transcription factors for labeling specific neurons) [57, 191,192,193,194,195]. The morphological classification of inhibitory and excitatory neurons in rodents can serve as a preliminary reference for establishing the morphological classification of brain cells in primates. This reference can expedite the generation of a comprehensive connectivity atlas. By leveraging the knowledge and techniques developed in rodent studies, researchers can accelerate the mapping of neural circuits in primates, providing valuable insights into brain connectivity and function.
Leveraging the Core Advantages of the Above Technologies to Generate a Multimodal Atlas
The Advantages of these Three Technologies
The primate brain is organized into cortical areas responsible for different functions. Different types of cells within cortical areas with specific transcriptomic, morphologic, and electrophysiologic profiles establish synaptic connectivity following principles. During development, aging, and disease, the electrophysiology, morphology, and connectivity phenotypes of cell types are simultaneously affected and undergo different degrees of adaptation under the control of transcriptomic modifications. As in the mouse, the single-cell data on the morphology, electrophysiological properties, and gene expression patterns in the primate brain need to be integrated [196] for comprehensive knowledge of functional mechanisms in primate cortical areas. Single-cell and single-nucleus RNA-seq provide robust cell-type classification data to generate a transcriptomic reference atlas that can be used to illustrate the cellular composition of cortical areas and to perform bioinformatics analysis to explain the modifications that occur during physiological and pathological states (Fig. 2 A). Patch-seq results, which can be mapped to the transcriptomes provided in the above transcriptomic reference atlas, establish morphological and electrophysiological annotations for each cell type. Based on the advances in multicell patch-clamp recordings with detailed morphological reconstruction, especially the morphology of inhibitory neurons [60, 173], local connectivity principles among morphology-based neuronal types will be delineated in primates as in rodents (Fig. 2B, C). The strong correspondence between morphological and electrophysiological phenotypes of cells [16, 64] might be a theoretical basis for mapping the local connectivity principles to the above transcriptomic atlas with morphological and electrophysiological annotations. To this end, a primate brain atlas that integrates the datasets from the above three technologies can describe the transcriptomic, morphological, and electrophysiological profiles of each type and the local connectivity principles. This atlas will play a crucial role in investigating the cortical regions in primates and significantly contribute to research on degenerative diseases in primates.
Framework for depicting the primate brain atlas. Four modalities are illustrated. A Gene expression. Using single-cell/single-nucleus RNA-seq, single-cell ATAC-seq, spatial transcriptome, and Patch-seq, we can obtain the molecular characterization, gene expression dynamics, and gene expression regulation of each neuron. B Morphology. Patch-seq and multicell whole-cell patch clamps can provide laminar distributions, synaptic spines, input kinetics, and axo-dendritic projection patterns. C Physiology. Using Patch-seq and multicell whole-cell patch-clamp can provide the phenotypes of the neurons, such as the excitability of the membrane, firing properties, and ion channels. D Synaptic connectivity. Multicell whole-cell patch-clamp measures synaptic strength and kinetics, synaptic properties, synaptic latency, synaptic dynamics, and synaptic plasticity. Drawings of primate brains were created with BioRender.com.
Atlases with Multiple Dimensions Resolve Novel Neuroscience Questions
Extensive data on the applications of the three technologies are now being obtained from primate brains to generate a primate brain atlas with multiple dimensions within cortical areas. The atlas will describe the multiple profiles, including transcriptomes, electrophysiology, and morphology, as well as the local connectivity principles, in each cell type within primate cortical areas (Fig. 2). The organizational differences within homologous cortical areas could explain the higher complexity of functions in primates than in rodents. When establishing the above atlas, larger-scale transcriptome analysis might be used to identify species-specific cell types with distinct gene expression patterns [16, 64] and neuronal biology [90], and the local connectivity patterns of a selected cell type could then be comprehensively studied by the applications of Patch-seq and multicell patch-clamp (Fig. 2D).
Although studies show a surprising conservation of basic transcriptomic cell types across cortical areas between primates and rodents [10, 15, 99, 100], modifications of the primate neuronal profiles of transcriptomes, electrophysiology, morphology, and connectivity patterns might contribute to the more complex brain functions. In rodent studies, the integration of data from Patch-seq and sn/scRNA-seq has allowed for a comprehensive multimodal analysis. This approach successfully identified distinct morphology-electrophysiology-transcriptome types, showcasing unique neuronal properties. Moreover, these types can form continuous and correlated transcriptomic and morphological electrical landscapes within their respective families [72, 197]. This mutual predictability not only helps to predict the functional, morphological, or transcriptomic state based on one or two distinct neuronal properties but also provides promising potential for integration with experimental data from other technologies. In pathological studies, since large-scale transcriptome analysis has identified vulnerable cell types [139, 151, 153, 154], Patch-seq can be used to test the potential loss or gain changes in the transcriptomic, electrophysiological, and morphological aspects of these vulnerable cell types, and multicell patch-clamp can be used to detect the loss or gain changes of local connectivity involving the selected cell types. Gene expression pattern analysis not only explains the molecular mechanisms of functional changes but also supplies lists of marker genes of cell types to develop genetic manipulation tools, which might serve in the applications of Patch-seq and multicell patch-clamp in primates. Moreover, Patch-seq and multicell patch-clamp can be used to verify the therapeutic effectiveness of targeting drug candidates provided by the transcriptome analysis [48, 139, 198, 199].
Challenges and Future Efforts
These technologies have considerable potential to address crucial questions in primate neuroscience. Each of the above three technologies has already made important contributions to advances in primate neuroscience. Establishing a primate brain atlas with multiple dimensions by combining the advantages of the three technologies is extraordinarily promising. Still, several important challenges remain.
Due to the scarcity of primate tissues, especially those from primate models of neurological diseases, establishing a detailed primate brain atlas is challenging. In the future, integrating Patch-seq and multicell patch-clamp techniques with non-invasive imaging methods, such as magnetic resonance imaging and diffusion tensor imaging, offers a promising avenue for obtaining detailed structural and connectivity information about the primate brain in vivo [200].
Secondly, implementing Patch-seq and multicell patch-clamp techniques faces challenges due to their relatively low throughput. Future efforts should focus on developing automated and high-throughput methods for Patch-seq and multicell patch-clamp techniques. By streamlining the experimental workflow and optimizing protocols, more cells can be processed within a shorter time frame, allowing for a more comprehensive analysis of the diverse cell types in the primate brain.
Moreover, commonly used genetic manipulation tools are mainly used in rodents and organotypic section cultures in humans and non-human primates [201,202,203,204,205]. Future development of genetic manipulation tools, such as expressing fluorescent proteins to label specific neurons or using optogenetic approaches in vivo to dissect circuit connections, could reveal patterns of synaptic connectivity in different species, including non-human primates [193, 206,207,208,209,210].
Finally, to obtain transcriptome information from multicell patch experiments, future research should prioritize the development of techniques that enable the simultaneous patching of multiple cells, extraction of nuclei for RNA retrieval, and concurrent acquisition of morphological and electrophysiological information. For example, spatial single-cell analysis of multiple genes using multiplexed error-robust fluorescence in situ hybridization (MERFISH) has generated a molecularly-defined and spatially-resolved cell atlas [211]. Integrating Patch-seq with multicell patch-clamp and single-cell sequencing technologies is immensely important in establishing comparative connectivity maps across different states or cortical regions. By combining these techniques, researchers can establish comparative connectivity maps, enabling the study of neural circuitry across different conditions or regions of the cortex. This integration paves the way for a deeper understanding of brain function and connectivity.
In conclusion, a rodent transcriptomic cell atlas has become available with morphological and electrophysiological annotation [58, 72, 197, 212] and delineating the local connectivity principles [173, 213]. A primate neocortex transcriptomic atlas has been established, while the morphological, electrophysiological, and connective properties are mostly absent. Detailed interrogation of these neurons indicates a higher diversification of excitatory neurons in the supragranular layer. Further studies on a broader range of brain areas and cell types, together with a cellular atlas describing developmental, aging, and disease states, are necessary to fully understand the molecular and neuronal basis of the augmented cognitive and behavioral capabilities of higher primates.
References
Gidon A, Zolnik TA, Fidzinski P, Bolduan F, Papoutsi A, Poirazi P. Dendritic action potentials and computation in human layer 2/3 cortical neurons. Science 2020, 367: 83–87.
Bakken TE, Miller JA, Ding SL, Sunkin SM, Smith KA, Ng L, et al. A comprehensive transcriptional map of primate brain development. Nature 2016, 535: 367–375.
Beaulieu-Laroche L, Toloza EHS, van der Goes MS, Lafourcade M, Barnagian D, Williams ZM, et al. Enhanced dendritic compartmentalization in human cortical neurons. Cell 2018, 175: 643-651.e14.
Hill RS, Walsh CA. Molecular insights into human brain evolution. Nature 2005, 437: 64–67.
Lui JH, Hansen DV, Kriegstein AR. Development and evolution of the human neocortex. Cell 2011, 146: 18–36.
Rakic P. Evolution of the neocortex: A perspective from developmental biology. Nat Rev Neurosci 2009, 10: 724–735.
Sousa AMM, Meyer KA, Santpere G, Gulden FO, Sestan N. Evolution of the human nervous system function, structure, and development. Cell 2017, 170: 226–247.
Zilles K, Amunts K. Centenary of brodmann’s map—conception and fate. Nat Rev Neurosci 2010, 11: 139–145.
Defelipe J. The evolution of the brain, the human nature of cortical circuits, and intellectual creativity. Front Neuroanat 2011, 5: 29.
Ma S, Skarica M, Li Q, Xu C, Risgaard RD, Tebbenkamp ATN, et al. Molecular and cellular evolution of the primate dorsolateral prefrontal cortex. Science 2022, 377: eabo7257.
Hodge RD, Miller JA, Novotny M, Kalmbach BE, Ting JT, Bakken TE, et al. Transcriptomic evidence that von Economo neurons are regionally specialized extratelencephalic-projecting excitatory neurons. Nat Commun 2020, 11: 1172.
Boldog E, Bakken TE, Hodge RD, Novotny M, Aevermann BD, Baka J, et al. Transcriptomic and morphophysiological evidence for a specialized human cortical GABAergic cell type. Nat Neurosci 2018, 21: 1185–1195.
Zeng H, Shen EH, Hohmann JG, Oh SW, Bernard A, Royall JJ, et al. Large-scale cellular-resolution gene profiling in human neocortex reveals species-specific molecular signatures. Cell 2012, 149: 483–496.
Hawrylycz M, Miller JA, Menon V, Feng D, Dolbeare T, Guillozet-Bongaarts AL, et al. Canonical genetic signatures of the adult human brain. Nat Neurosci 2015, 18: 1832–1844.
Hodge RD, Bakken TE, Miller JA, Smith KA, Barkan ER, Graybuck LT, et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 2019, 573: 61–68.
Berg J, Sorensen SA, Ting JT, Miller JA, Chartrand T, Buchin A, et al. Human neocortical expansion involves glutamatergic neuron diversification. Nature 2021, 598: 151–158.
Hedlund E, Deng Q. Single-cell RNA sequencing: Technical advancements and biological applications. Mol Aspects Med 2018, 59: 36–46.
Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 2009, 6: 377–382.
Grindberg RV, Yee-Greenbaum JL, McConnell MJ, Novotny M, O’Shaughnessy AL, Lambert GM, et al. RNA-sequencing from single nuclei. Proc Natl Acad Sci U S A 2013, 110: 19802–19807.
Krishnaswami SR, Grindberg RV, Novotny M, Venepally P, Lacar B, Bhutani K, et al. Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat Protoc 2016, 11: 499–524.
Fishell G, Heintz N. The neuron identity problem: Form meets function. Neuron 2013, 80: 602–612.
Gouwens NW, Sorensen SA, Berg J, Lee C, Jarsky T, Ting J, et al. Classification of electrophysiological and morphological neuron types in the mouse visual cortex. Nat Neurosci 2019, 22: 1182–1195.
Andrews TS, Kiselev VY, McCarthy D, Hemberg M. Tutorial: Guidelines for the computational analysis of single-cell RNA sequencing data. Nat Protoc 2021, 16: 1–9.
Sousa AMM, Zhu Y, Raghanti MA, Kitchen RR, Onorati M, Tebbenkamp ATN, et al. Molecular and cellular reorganization of neural circuits in the human lineage. Science 2017, 358: 1027–1032.
Kozareva V, Martin C, Osorno T, Rudolph S, Guo C, Vanderburg C, et al. A transcriptomic atlas of mouse cerebellar cortex comprehensively defines cell types. Nature 2021, 598: 214–219.
Initiative Cell Census Network (BICCN) BRAIN. A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 2021, 598: 86–102
Tasic B, Menon V, Nguyen TN, Kim TK, Jarsky T, Yao Z, et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 2016, 19: 335–346.
Yao Z, Liu H, Xie F, Fischer S, Adkins RS, Aldridge AI, et al. A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex. Nature 2021, 598: 103–110.
Yao Z, van Velthoven CTJ, Nguyen TN, Goldy J, Sedeno-Cortes AE, Baftizadeh F, et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 2021, 184: 3222-3241.e26.
Zeisel A, Muñoz-Manchado AB, Codeluppi S, Lönnerberg P, La Manno G, Juréus A, et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 2015, 347: 1138–1142.
Tasic B, Yao Z, Graybuck LT, Smith KA, Nguyen TN, Bertagnolli D, et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 2018, 563: 72–78.
Bhaduri A, Sandoval-Espinosa C, Otero-Garcia M, Oh I, Yin R, Eze UC, et al. An atlas of cortical arealization identifies dynamic molecular signatures. Nature 2021, 598: 200–204.
Liao CH, Su B. Research proceedings on primate comparative genomics. Dongwuxue Yanjiu 2012, 33: 108–118.
Zeisel A, Hochgerner H, Lönnerberg P, Johnsson A, Memic F, van der Zwan J, et al. Molecular architecture of the mouse nervous system. Cell 2018, 174: 999-1014.e22.
Saunders A, Macosko EZ, Wysoker A, Goldman M, Krienen FM, de Rivera H, et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 2018, 174: 1015-1030.e16.
Welch JD, Kozareva V, Ferreira A, Vanderburg C, Martin C, Macosko EZ. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 2019, 177: 1873-1887.e17.
Wu DD, Qi XG, Yu L, Li M, Liu ZJ, Yoder AD, et al. Initiation of the primate genome project. Zool Res 2022, 43: 147–149.
Cheng S, Butrus S, Tan L, Xu R, Sagireddy S, Trachtenberg JT, et al. Vision-dependent specification of cell types and function in the developing cortex. Cell 2022, 185: 311-327.e24.
Di Bella DJ, Habibi E, Stickels RR, Scalia G, Brown J, Yadollahpour P, et al. Molecular logic of cellular diversification in the mouse cerebral cortex. Nature 2021, 595: 554–559.
Hrvatin S, Hochbaum DR, Nagy MA, Cicconet M, Robertson K, Cheadle L, et al. Single-cell analysis of experience-dependent transcriptomic states in the mouse visual cortex. Nat Neurosci 2018, 21: 120–129.
La Manno G, Siletti K, Furlan A, Gyllborg D, Vinsland E, Mossi Albiach A, et al. Molecular architecture of the developing mouse brain. Nature 2021, 596: 92–96.
Tosches MA, Yamawaki TM, Naumann RK, Jacobi AA, Tushev G, Laurent G. Evolution of pallium, hippocampus, and cortical cell types revealed by single-cell transcriptomics in reptiles. Science 2018, 360: 881–888.
Mayer C, Hafemeister C, Bandler RC, Machold R, Batista Brito R, Jaglin X, et al. Developmental diversification of cortical inhibitory interneurons. Nature 2018, 555: 457–462.
Li Y, Xu NN, Hao ZZ, Liu S. Adult neurogenesis in the primate hippocampus. Zool Res 2023, 44: 315–322.
Kim EJ, Zhang Z, Huang L, Ito-Cole T, Jacobs MW, Juavinett AL, et al. Extraction of distinct neuronal cell types from within a genetically continuous population. Neuron 2020, 107: 274-282.e6.
Kolodziejczyk AA, Kim JK, Svensson V, Marioni JC, Teichmann SA. The technology and biology of single-cell RNA sequencing. Mol Cell 2015, 58: 610–620.
Jovic D, Liang X, Zeng H, Lin L, Xu F, Luo Y. Single-cell RNA sequencing technologies and applications: A brief overview. Clin Transl Med 2022, 12: e694.
Velmeshev D, Schirmer L, Jung D, Haeussler M, Perez Y, Mayer S, et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 2019, 364: 685–689.
Ahmadi A, Gispert JD, Navarro A, Vilor-Tejedor N, Sadeghi I. Single-cell transcriptional changes in neurodegenerative diseases. Neuroscience 2021, 479: 192–205.
Saliba AE, Westermann AJ, Gorski SA, Vogel J. Single-cell RNA-seq: Advances and future challenges. Nucleic Acids Res 2014, 42: 8845–8860.
Neher E, Sakmann B. Single-channel Currents recorded from membrane of denervated frog muscle fibres. Nature 1976, 260: 799–802.
Fishell G, Rudy B. Mechanisms of inhibition within the telencephalon: Where the wild things are. Annu Rev Neurosci 2011, 34: 535–567.
Okaty BW, Miller MN, Sugino K, Hempel CM, Nelson SB. Transcriptional and electrophysiological maturation of neocortical fast-spiking GABAergic interneurons. J Neurosci 2009, 29: 7040–7052.
Okaty BW, Sugino K, Nelson SB. Cell type-specific transcriptomics in the brain. J Neurosci 2011, 31: 6939–6943.
Sugino K, Hempel CM, Miller MN, Hattox AM, Shapiro P, Wu C, et al. Molecular taxonomy of major neuronal classes in the adult mouse forebrain. Nat Neurosci 2006, 9: 99–107.
Cadwell CR, Palasantza A, Jiang X, Berens P, Deng Q, Yilmaz M, et al. Electrophysiological, transcriptomic and morphologic profiling of single neurons using Patch-seq. Nat Biotechnol 2016, 34: 199–203.
Cadwell CR, Scala F, Li S, Livrizzi G, Shen S, Sandberg R, et al. Multimodal profiling of single-cell morphology, electrophysiology, and gene expression using Patch-seq. Nat Protoc 2017, 12: 2531–2553.
Fuzik J, Zeisel A, Máté Z, Calvigioni D, Yanagawa Y, Szabó G, et al. Integration of electrophysiological recordings with single-cell RNA-seq data identifies neuronal subtypes. Nat Biotechnol 2016, 34: 175–183.
Lee BR, Budzillo A, Hadley K, Miller JA, Jarsky T, Baker K, Hill D, Kim L, Mann R, Ng L, Oldre A. Scaled, high fidelity electrophysiological, morphological, and transcriptomic cell characterization. Elife. 2021, 10: e65482.
Lipovsek M, Browne L, Grubb MS. Protocol for patch-seq of small interneurons. STAR Protoc 2020, 1: 100146.
Chartrand T, Dalley R, Close J, Goriounova NA, Lee BR, Mann R, et al. Morphoelectric and transcriptomic divergence of the layer 1 interneuron repertoire in human versus mouse neocortex. Science 2023, 382: eadf0805.
Lee BR, Dalley R, Miller JA, Chartrand T, Close J, Mann R, et al. Signature morphoelectric properties of diverse GABAergic interneurons in the human neocortex. Science 2023, 382: eadf6484.
Markram H, Toledo-Rodriguez M, Wang Y, Gupta A, Silberberg G, Wu C. Interneurons of the neocortical inhibitory system. Nat Rev Neurosci 2004, 5: 793–807.
Kalmbach BE, Hodge RD, Jorstad NL, Owen S, de Frates R, Yanny AM, et al. Signature morpho-electric, transcriptomic, and dendritic properties of human layer 5 neocortical pyramidal neurons. Neuron 2021, 109: 2914-2927.e5.
van den Hurk M, Erwin JA, Yeo GW, Gage FH, Bardy C. Patch-seq protocol to analyze the electrophysiology, morphology and transcriptome of whole single neurons derived from human pluripotent stem cells. Front Mol Neurosci 2018, 11: 261.
Bernaerts Y, Berens P, Kobak D. Sparse bottleneck neural networks for exploratory non-linear visualization of Patch-seq data. 2020: arXiv: 2006.10411. https://arxiv.org/abs/2006.10411.pdf"
Gala R, Gouwens N, Yao Z, Budzillo A, Penn O, Tasic B, et al. A coupled autoencoder approach for multi-modal analysis of cell types. 2019: arXiv: 1911.05663. https://arxiv.org/abs/1911.05663.
Kobak D, Bernaerts Y, Weis MA, Scala F, Tolias AS, Berens P. Sparse reduced-rank regression for exploratory visualisation of paired multivariate data. J Royal Stat Soc Ser C Appl Stat 2021, 70: 980–1000.
Kim JMH, Camarena A, Walker C, Lin MY, Wolseley V, Souaiaia T, et al. Robust RNA-Seq of aRNA-amplified single cell material collected by patch clamp. Sci Rep 1979, 2020: 10.
Paraskevopoulou F, Parvizi P, Senger G, Tuncbag N, Rosenmund C, Yildirim F. Impaired inhibitory GABAergic synaptic transmission and transcription studied in single neurons by Patch-seq in Huntington’s disease. Proc Natl Acad Sci U S A 2021, 118: e2020293118.
Parpaite T, Brosse L, Séjourné N, Laur A, Mechioukhi Y, Delmas P, et al. Patch-seq of mouse DRG neurons reveals candidate genes for specific mechanosensory functions. Cell Rep 2021, 37: 109914.
Scala F, Kobak D, Bernabucci M, Bernaerts Y, Cadwell CR, Castro JR, et al. Phenotypic variation of transcriptomic cell types in mouse motor cortex. Nature 2021, 598: 144–150.
Huang W, Xu Q, Su J, Tang L, Hao ZZ, Xu C, et al. Linking transcriptomes with morphological and functional phenotypes in mammalian retinal ganglion cells. Cell Rep 2022, 40: 111322.
Kantevari S, Matsuzaki M, Kanemoto Y, Kasai H, Ellis-Davies GCR. Two-color, two-photon uncaging of glutamate and GABA. Nat Methods 2010, 7: 123–125.
Bamann C, Nagel G, Bamberg E. Microbial rhodopsins in the spotlight. Curr Opin Neurobiol 2010, 20: 610–616.
Miesenböck G. The optogenetic catechism. Science 2009, 326: 395–399.
Yizhar O, Fenno LE, Davidson TJ, Mogri M, Deisseroth K. Optogenetics in neural systems. Neuron 2011, 71: 9–34.
Grienberger C, Konnerth A. Imaging calcium in neurons. Neuron 2012, 73: 862–885.
Markram H, Lübke J, Frotscher M, Roth A, Sakmann B. Physiology and anatomy of synaptic connections between thick tufted pyramidal neurones in the developing rat neocortex. J Physiol 1997, 500(Pt 2): 409–440.
Feldmeyer D, Egger V, Lubke J, Sakmann B. Reliable synaptic connections between pairs of excitatory layer 4 neurones within a single ‘barrel’ of developing rat somatosensory cortex. J Physiol 1999, 521: 169–190.
Le Bé JV, Markram H. Spontaneous and evoked synaptic rewiring in the neonatal neocortex. Proc Natl Acad Sci USA 2006, 103: 13214–13219.
Lefort S, Tomm C, Floyd Sarria JC, Petersen CCH. The excitatory neuronal network of the C2 barrel column in mouse primary somatosensory cortex. Neuron 2009, 61: 301–316.
Perin R, Berger TK, Markram H. A synaptic organizing principle for cortical neuronal groups. Proc Natl Acad Sci USA 2011, 108: 5419–5424.
Jiang X, Wang G, Lee AJ, Stornetta RL, Zhu JJ. The organization of two new cortical interneuronal circuits. Nat Neurosci 2013, 16: 210–218.
Lee AJ, Wang G, Jiang X, Johnson SM, Hoang ET, Lanté F, et al. Canonical organization of layer 1 neuron-led cortical inhibitory and disinhibitory interneuronal circuits. Cereb Cortex 2015, 25: 2114–2126.
Langseth CM, Gyllborg D, Miller JA, Close JL, Long B, Lein ES, et al. Comprehensive in situ mapping of human cortical transcriptomic cell types. Commun Biol 2021, 4: 998.
Zhang X, Lai GY, Volpe G, Han L, Maxwell PH, Liu LQ, et al. Towards a primate single-cell atlas. Zool Res 2022, 43: 691–694.
Wei JR, Xiao D, Tang L, Xu N, Liu R, Shen Y, et al. Neural cell isolation from adult macaques for high-throughput analyses and neurosphere cultures. Nat Protoc 2023, 18: 1930–1957.
Braun E, Danan-Gotthold M, Borm LE, Lee KW, Vinsland E, Lönnerberg P, et al. Comprehensive cell atlas of the first-trimester developing human brain. Science 2023, 382: eadf1226.
Bakken TE, Jorstad NL, Hu Q, Lake BB, Tian W, Kalmbach BE, et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 2021, 598: 111–119.
Wei JR, Hao ZZ, Xu C, Huang M, Tang L, Xu N, et al. Identification of visual cortex cell types and species differences using single-cell RNA sequencing. Nat Commun 2022, 13: 6902.
Siletti K, Hodge R, Albiach AM, Hu L, Lee KW, Lönnerberg P, et al. Transcriptomic diversity of cell types across the adult human brain. Science 2023, 382: eadd7046.
Jorstad NL, Close J, Johansen N, Yanny AM, Barkan ER, Travaglini KJ, et al. Transcriptomic cytoarchitecture reveals principles of human neocortex organization. Science 2023, 382: eadf6812.
Herring CA, Simmons RK, Freytag S, Poppe D, Moffet JJD, Pflueger J, et al. Human prefrontal cortex gene regulatory dynamics from gestation to adulthood at single-cell resolution. Cell 2022, 185: 4428-4447.e28.
Pfisterer U, Petukhov V, Demharter S, Meichsner J, Thompson JJ, Batiuk MY, et al. Identification of epilepsy-associated neuronal subtypes and gene expression underlying epileptogenesis. Nat Commun 2020, 11: 5038.
Franjic D, Skarica M, Ma S, Arellano JI, Tebbenkamp ATN, Choi J, et al. Transcriptomic taxonomy and neurogenic trajectories of adult human, macaque, and pig hippocampal and entorhinal cells. Neuron 2022, 110: 452-469.e14.
Krienen FM, Goldman M, Zhang Q, Del Rosario RCH, Florio M, Machold R, et al. Innovations present in the primate interneuron repertoire. Nature 2020, 586: 262–269.
Jorstad NL, Song JHT, Exposito-Alonso D, Suresh H, Castro-Pacheco N, Krienen FM, et al. Comparative transcriptomics reveals human-specific cortical features. Science 2023, 382: eade9516.
Bakken TE, van Velthoven CT, Menon V, Hodge RD, Yao Z, Nguyen TN, Graybuck LT, Horwitz GD, Bertagnolli D, Goldy J, Yanny AM. Single-cell and single-nucleus RNA-seq uncovers shared and distinct axes of variation in dorsal LGN neurons in mice, non-human Primates, and humans. Elife. 2021, 10: e64875.
Kebschull JM, Richman EB, Ringach N, Friedmann D, Albarran E, Kolluru SS, et al. Cerebellar nuclei evolved by repeatedly duplicating a conserved cell-type set. Science 2020, 370: eabd5059.
Shi Y, Wang M, Mi D, Lu T, Wang B, Dong H, et al. Mouse and human share conserved transcriptional programs for interneuron development. Science 2021, 374: eabj6641.
Xu X, Stoyanova EI, Lemiesz AE, Xing J, Mash DC, Heintz N. Species and cell-type properties of classically defined human and rodent neurons and glia. Elife. 2018, 7: e37551.
Khrameeva E, Kurochkin I, Han D, Guijarro P, Kanton S, Santel M, et al. Single-cell-resolution transcriptome map of human, chimpanzee, bonobo, and macaque brains. Genome Res 2020, 30: 776–789.
Benito-Kwiecinski S, Giandomenico SL, Sutcliffe M, Riis ES, Freire-Pritchett P, Kelava I, et al. An early cell shape transition drives evolutionary expansion of the human forebrain. Cell 2021, 184: 2084-2102.e19.
Suzuki IK, Gacquer D, Van Heurck R, Kumar D, Wojno M, Bilheu A, et al. Human-specific NOTCH2NL genes expand cortical neurogenesis through delta/Notch regulation. Cell 2018, 173: 1370-1384.e16.
Azevedo FAC, Carvalho LRB, Grinberg LT, Farfel JM, Ferretti REL, Leite REP, et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J Comp Neurol 2009, 513: 532–541.
Schmitz MT, Sandoval K, Chen CP, Mostajo-Radji MA, Seeley WW, Nowakowski TJ, et al. The development and evolution of inhibitory neurons in primate cerebrum. Nature 2022, 603: 871–877.
Bocchi VD, Conforti P, Vezzoli E, Besusso D, Cappadona C, Lischetti T, et al. The coding and long noncoding single-cell atlas of the developing human fetal striatum. Science 2021, 372: eabf5759.
Fan X, Dong J, Zhong S, Wei Y, Wu Q, Yan L, et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res 2018, 28: 730–745.
Ma H, Zhai J, Wan H, Jiang X, Wang X, Wang L, et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 2019, 366: eaax7890.
Nakajima R, Hagihara H, Miyakawa T. Similarities of developmental gene expression changes in the brain between human and experimental animals: Rhesus monkey, mouse, Zebrafish, and Drosophila. Mol Brain 2021, 14: 135.
Zhong S, Zhang S, Fan X, Wu Q, Yan L, Dong J, et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 2018, 555: 524–528.
Shang Z, Chen D, Wang Q, Wang S, Deng Q, Wu L, et al. Single-cell RNA-seq reveals dynamic transcriptome profiling in human early neural differentiation. GigaScience 2018, 7: giy117.
Trevino AE, Müller F, Andersen J, Sundaram L, Kathiria A, Shcherbina A, et al. Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 2021, 184: 5053-5069.e23.
Allman JM, Tetreault NA, Hakeem AY, Manaye KF, Semendeferi K, Erwin JM, et al. The von Economo neurons in frontoinsular and anterior cingulate cortex in great apes and humans. Brain Struct Funct 2010, 214: 495–517.
Polioudakis D, de la Torre-Ubieta L, Langerman J, Elkins AG, Shi X, Stein JL, et al. A single-cell transcriptomic atlas of human neocortical development during mid-gestation. Neuron 2019, 103: 785-801.e8.
La Manno G, Gyllborg D, Codeluppi S, Nishimura K, Salto C, Zeisel A, et al. Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 2016, 167: 566-580.e19.
Darmanis S, Sloan SA, Zhang Y, Enge M, Caneda C, Shuer LM, et al. A survey of human brain transcriptome diversity at the single cell level. Proc Natl Acad Sci USA 2015, 112: 7285–7290.
Twine NA, Janitz C, Wilkins MR, Janitz M. Sequencing of hippocampal and cerebellar transcriptomes provides new insights into the complexity of gene regulation in the human brain. Neurosci Lett 2013, 541: 263–268.
Yin S, Lu K, Tan T, Tang J, Wei J, Liu X, et al. Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain. Nat Commun 2020, 11: 474.
Zhong W, Barde S, Mitsios N, Adori C, Oksvold P, Feilitzen KV, et al. The neuropeptide landscape of human prefrontal cortex. Proc Natl Acad Sci U S A 2022, 119: e2123146119.
Boulting GL, Durresi E, Ataman B, Sherman MA, Mei K, Harmin DA, et al. Activity-dependent regulome of human GABAergic neurons reveals new patterns of gene regulation and neurological disease heritability. Nat Neurosci 2021, 24: 437–448.
Zhong S, Ding W, Sun L, Lu Y, Dong H, Fan X, et al. Decoding the development of the human hippocampus. Nature 2020, 577: 531–536.
Close JL, Yao Z, Levi BP, Miller JA, Bakken TE, Menon V, et al. Single-cell profiling of an in vitro model of human interneuron development reveals temporal dynamics of cell type production and maturation. Neuron 2017, 93: 1035-1048.e5.
Dillman AA, Majounie E, Ding J, Ding J, Hernandez D, Arepalli S, et al. Transcriptomic profiling of the human brain reveals that altered synaptic gene expression is associated with chronological aging. Sci Rep 2017, 7: 16890.
Hao ZZ, Wei JR, Xiao D, Liu R, Xu N, Tang L, et al. Single-cell transcriptomics of adult macaque hippocampus reveals neural precursor cell populations. Nat Neurosci 2022, 25: 805–817.
Wang W, Wang M, Yang M, Zeng B, Qiu W, Ma Q, et al. Transcriptome dynamics of hippocampal neurogenesis in macaques across the lifespan and aged humans. Cell Res 2022, 32: 729–743.
Fu Y, Yang M, Yu H, Wang Y, Wu X, Yong J, et al. Heterogeneity of glial progenitor cells during the neurogenesis-to-gliogenesis switch in the developing human cerebral cortex. Cell Rep 2021, 34: 108788.
Hasel P, Dando O, Jiwaji Z, Baxter P, Todd AC, Heron S, et al. Neurons and neuronal activity control gene expression in astrocytes to regulate their development and metabolism. Nat Commun 2017, 8: 15132.
Yang L, Li Z, Liu G, Li X, Yang Z. Developmental origins of human cortical oligodendrocytes and astrocytes. Neurosci Bull 2022, 38: 47–68.
Lake BB, Chen S, Sos BC, Fan J, Kaeser GE, Yung YC, et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat Biotechnol 2018, 36: 70–80.
Zhang HL, Long JW, Han W, Wang J, Song W, Lin GN, et al. Comparative analysis of cellular expression pattern of schizophrenia risk genes in human versus mouse cortex. Cell Biosci 2019, 9: 89.
Smajić S, Prada-Medina CA, Landoulsi Z, Ghelfi J, Delcambre S, Dietrich C, et al. Single-cell sequencing of human midbrain reveals glial activation and a Parkinson-specific neuronal state. Brain 2022, 145: 964–978.
Dols-Icardo O, Montal V, Sirisi S, López-Pernas G, Cervera-Carles L, Querol-Vilaseca M, et al. Motor cortex transcriptome reveals microglial key events in amyotrophic lateral sclerosis. Neurol Neuroimmunol Neuroinflamm 2020, 7: e829.
Bryois J, Calini D, MacNair W, Foo L, Urich E, Ortmann W, et al. Cell-type-specific cis-eQTLs in eight human brain cell types identify novel risk genes for psychiatric and neurological disorders. Nat Neurosci 2022, 25: 1104–1112.
Brenner E, Tiwari GR, Kapoor M, Liu Y, Brock A, Liu Y. Single cell transcriptome profiling of the human alcohol-dependent brain. Hum Mol Genet 2020, 29: 1144–1153.
Biermann J, Melms JC, Amin AD, Wang Y, Caprio LA, Karz A, et al. Dissecting the treatment-naive ecosystem of human melanoma brain metastasis. Cell 2022, 185: 2591-2608.e30.
Al-Dalahmah O, Sosunov AA, Shaik A, Ofori K, Liu Y, Vonsattel JP, et al. Single-nucleus RNA-seq identifies Huntington disease astrocyte states. Acta Neuropathol Commun 2020, 8: 19.
Luo ZG, Peng J, Li T. Single-cell RNA sequencing reveals cell-type-specific mechanisms of neurological diseases. Neurosci Bull 2020, 36: 821–824.
Braak H, Braak E. Neuropathological stageing of alzheimer-related changes. Acta Neuropathol 1991, 82: 239–259.
Chen ZY, Zhang Y. Animal models of Alzheimer’s disease: Applications, evaluation, and perspectives. Zool Res 2022, 43: 1026–1040.
Ying Y, Wang JZ. Illuminating neural circuits in alzheimer’s disease. Neurosci Bull 2021, 37: 1203–1217.
Mathys H, Davila-Velderrain J, Peng Z, Gao F, Mohammadi S, Young JZ, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 2019, 570: 332–337.
Nguyen AT, Wang K, Hu G, Wang X, Miao Z, Azevedo JA, et al. APOE and TREM2 regulate amyloid-responsive microglia in Alzheimer’s disease. Acta Neuropathol 2020, 140: 477–493.
Zhou Y, Song WM, Andhey PS, Swain A, Levy T, Miller KR, et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat Med 2020, 26: 131–142.
Olah M, Menon V, Habib N, Taga MF, Ma Y, Yung CJ, et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat Commun 2020, 11: 6129.
Galea E, Weinstock LD, Larramona-Arcas R, Pybus AF, Giménez-Llort L, Escartin C, et al. Multi-transcriptomic analysis points to early organelle dysfunction in human astrocytes in Alzheimer’s disease. Neurobiol Dis 2022, 166: 105655.
Leng K, Li E, Eser R, Piergies A, Sit R, Tan M, et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat Neurosci 2021, 24: 276–287.
Xu J, Zhang P, Huang Y, Zhou Y, Hou Y, Bekris LM, et al. Multimodal single-cell/nucleus RNA sequencing data analysis uncovers molecular networks between disease-associated microglia and astrocytes with implications for drug repurposing in Alzheimer’s disease. Genome Res 2021, 31: 1900–1912.
Brewer R, Biotti F, Catmur C, Press C, Happé F, Cook R, et al. Can neurotypical individuals read autistic facial expressions? atypical production of emotional facial expressions in autism spectrum disorders. Autism Res 2016, 9: 262–271.
Gandal MJ, Haney JR, Wamsley B, Yap CX, Parhami S, Emani PS, et al. Broad transcriptomic dysregulation occurs across the cerebral cortex in ASD. Nature 2022, 611: 532–539.
Guo MF, Ji N, Ma CG. Immunologic pathogenesis of multiple sclerosis. Neurosci Bull 2008, 24: 381–386.
Jäkel S, Agirre E, Mendanha Falcão A, van Bruggen D, Lee KW, Knuesel I, et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 2019, 566: 543–547.
Trobisch T, Zulji A, Stevens NA, Schwarz S, Wischnewski S, Öztürk M, et al. Cross-regional homeostatic and reactive glial signatures in multiple sclerosis. Acta Neuropathol 2022, 144: 987–1003.
Absinta M, Maric D, Gharagozloo M, Garton T, Smith MD, Jin J, et al. A lymphocyte-microglia-astrocyte axis in chronic active multiple sclerosis. Nature 2021, 597: 709–714.
DeFelipe J, Alonso-Nanclares L, Arellano JI. Microstructure of the neocortex: Comparative aspects. J Neurocytol 2002, 31: 299–316.
Hof PR, Cox K, Morrison JH. Quantitative analysis of a vulnerable subset of pyramidal neurons in Alzheimer’s disease: I. Superior frontal and inferior temporal cortex. J Comp Neurol 1990, 301: 44–54.
Bussière T, Giannakopoulos P, Bouras C, Perl DP, Morrison JH, Hof PR. Progressive degeneration of nonphosphorylated neurofilament protein-enriched pyramidal neurons predicts cognitive impairment in Alzheimer’s disease: Stereologic analysis of prefrontal cortex area 9. J Comp Neurol 2003, 463: 281–302.
Kole MHP, Bräuer AU, Stuart GJ. Inherited cortical HCN1 channel loss amplifies dendritic calcium electrogenesis and burst firing in a rat absence epilepsy model. J Physiol 2007, 578: 507–525.
Scheibel ME, Davies TL, Lindsay RD, Scheibel AB. Basilar dendrite bundles of giant pyramidal cells. Exp Neurol 1974, 42: 307–319.
Bardy C, van den Hurk M, Kakaradov B, Erwin JA, Jaeger BN, Hernandez RV, et al. Predicting the functional states of human iPSC-derived neurons with single-cell RNA-seq and electrophysiology. Mol Psychiatry 2016, 21: 1573–1588.
van den Hurk M, Lau S, Marchetto MC, Mertens J, Stern S, Corti O, et al. Druggable transcriptomic pathways revealed in Parkinson’s patient-derived midbrain neurons. NPJ Parkinsons Dis 2022, 8: 134.
Chen X, Zhang K, Zhou L, Gao X, Wang J, Yao Y, et al. Coupled electrophysiological recording and single cell transcriptome analyses revealed molecular mechanisms underlying neuronal maturation. Protein Cell 2016, 7: 175–186.
Cadwell CR, Sandberg R, Jiang X, Tolias AS. Q&A: Using Patch-seq to profile single cells. BMC Biol 2017, 15: 1–7.
Lipovsek M, Bardy C, Cadwell CR, Hadley K, Kobak D, Tripathy SJ. Patch-seq: Past, present, and future. J Neurosci 2021, 41: 937–946.
Hochbaum DR, Zhao Y, Farhi SL, Klapoetke N, Werley CA, Kapoor V, et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat Methods 2014, 11: 825–833.
Packer AM, Russell LE, Dalgleish HWP, Häusser M. Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat Methods 2015, 12: 140–146.
Geiger JR, Lübke J, Roth A, Frotscher M, Jonas P. Submillisecond AMPA receptor-mediated signaling at a principal neuron-interneuron synapse. Neuron 1997, 18: 1009–1023.
Perin R, Markram H. A computer-assisted multi-electrode patch-clamp system. JoVE J Vis Exper 2013, 18(80): e50630.
Peng Y, Barreda Tomás FJ, Klisch C, Vida I, Geiger JRP. Layer-specific organization of local excitatory and inhibitory synaptic connectivity in the rat presubiculum. Cereb Cortex 2017, 27: 2435–2452.
Wang G, Wyskiel DR, Yang W, Wang Y, Milbern LC, Lalanne T, et al. An optogenetics- and imaging-assisted simultaneous multiple patch-clamp recording system for decoding complex neural circuits. Nat Protoc 2015, 10: 397–412.
Guzman SJ, Schlögl A, Frotscher M, Jonas P. Synaptic mechanisms of pattern completion in the hippocampal CA3 network. Science 2016, 353: 1117–1123.
Jiang X, Shen S, Cadwell CR, Berens P, Sinz F, Ecker AS, et al. Principles of connectivity among morphologically defined cell types in adult neocortex. Science 2015, 350: aac9462.
Evans DA, Stempel AV, Vale R, Ruehle S, Lefler Y, Branco T. A synaptic threshold mechanism for computing escape decisions. Nature 2018, 558: 590–594.
Jin X, Prince DA, Huguenard JR. Enhanced excitatory synaptic connectivity in layer v pyramidal neurons of chronically injured epileptogenic neocortex in rats. J Neurosci 2006, 26: 4891–4900.
Olshausen BA, Field DJ. Emergence of simple-cell receptive field properties by learning a sparse code for natural images. Nature 1996, 381: 607–609.
Camperi M, Wang XJ. A model of visuospatial working memory in prefrontal cortex: Recurrent network and cellular bistability. J Comput Neurosci 1998, 5: 383–405.
Mongillo G, Barak O, Tsodyks M. Synaptic theory of working memory. Science 2008, 319: 1543–1546.
Brunel N. Is cortical connectivity optimized for storing information? Nat Neurosci 2016, 19: 749–755.
Pernice V, da Silveira RA. Interpretation of correlated neural variability from models of feed-forward and recurrent circuits. PLoS Comput Biol 2018, 14: e1005979.
Thomson AM, West DC, Wang Y, Wang Y. Synaptic connections and small circuits involving excitatory and inhibitory neurons in layers 2–5 of adult rat and cat neocortex: Triple intracellular recordings and biocytin labelling in vitro. Cereb Cortex 2002, 12: 936–953.
Hofer SB, Ko H, Pichler B, Vogelstein J, Ros H, Zeng H, et al. Differential connectivity and response dynamics of excitatory and inhibitory neurons in visual cortex. Nat Neurosci 2011, 14: 1045–1052.
Seeman SC, Campagnola L, Davoudian PA, Hoggarth A, Hage TA, Bosma-Moody A, Baker CA, Lee JH, Mihalas S, Teeter C, Ko AL. Sparse recurrent excitatory connectivity in the microcircuit of the adult mouse and human cortex. Elife. 2018, 7: e37349.
Hunt S, Leibner Y, Mertens EJ, Barros-Zulaica N, Kanari L, Heistek TS, et al. Strong and reliable synaptic communication between pyramidal neurons in adult human cerebral cortex. Cereb Cortex 2023, 33: 2857–2878.
Benavides-Piccione R, Ballesteros-Yáñez I, DeFelipe J, Yuste R. Cortical area and species differences in dendritic spine morphology. J Neurocytol 2002, 31: 337–346.
Yakoubi R, Rollenhagen A, von Lehe M, Miller D, Walkenfort B, Hasenberg M, Sätzler K, Lübke JH. Ultrastructural heterogeneity of layer 4 excitatory synaptic boutons in the adult human temporal lobe neocortex. Elife. 2019, 8: e48373.
Campagnola L, Seeman SC, Chartrand T, Kim L, Hoggarth A, Gamlin C, et al. Local connectivity and synaptic dynamics in mouse and human neocortex. Science 2022, 375: eabj5861.
Zhou C, Yang X, Wu S, Zhong Q, Luo T, Li A, et al. Continuous subcellular resolution three-dimensional imaging on intact macaque brain. Sci Bull Beijing 2022, 67: 85–96.
Novak P, Gorelik J, Vivekananda U, Shevchuk AI, Ermolyuk YS, Bailey RJ, et al. Nanoscale-targeted patch-clamp recordings of functional presynaptic ion channels. Neuron 2013, 79: 1067–1077.
Novak P, Li C, Shevchuk AI, Stepanyan R, Caldwell M, Hughes S, et al. Nanoscale live-cell imaging using hopping probe ion conductance microscopy. Nat Methods 2009, 6: 279–281.
Cadwell CR, Scala F, Fahey PG, Kobak D, Mulherkar S, Sinz FH, Papadopoulos S, Tan ZH, Johnsson P, Hartmanis L, Li S. Cell type composition and circuit organization of clonally related excitatory neurons in the juvenile mouse neocortex. Elife. 2020, 9: e52951.
Qiu L, Zhang B, Gao Z. Lighting up neural circuits by viral tracing. Neurosci Bull 2022, 38: 1383–1396.
Liu Q, Wu Y, Wang H, Jia F, Xu F. Viral tools for neural circuit tracing. Neurosci Bull 2022, 38: 1508–1518.
Shi L, Su B. A transgenic monkey model for the study of human brain evolution. Zool Res 2019, 40: 236–238.
Chen J, Li C, Lu Z, Zhan C. Optimal timing of a commonly-used rabies virus for neural recording and manipulation. Neurosci Bull 2022, 38: 548–552.
Poo MM. Transcriptome, connectome and neuromodulation of the primate brain. Cell 2022, 185: 2636–2639.
Gouwens NW, Sorensen SA, Baftizadeh F, Budzillo A, Lee BR, Jarsky T, et al. Integrated morphoelectric and transcriptomic classification of cortical GABAergic cells. Cell 2020, 183: 935-953.e19.
Grubman A, Chew G, Ouyang JF, Sun G, Choo XY, McLean C, et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 2019, 22: 2087–2097.
Schirmer L, Velmeshev D, Holmqvist S, Kaufmann M, Werneburg S, Jung D, et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 2019, 573: 75–82.
Sun S, Shi J, Wang Y, Cheng J, Huang Z. A temporal precision approach for deep transcranial optogenetics with non-invasive surgery. Neurosci Bull 2021, 37: 1260–1263.
Ting JT, Kalmbach B, Chong P, de Frates R, Keene CD, Gwinn RP, et al. A robust ex vivo experimental platform for molecular-genetic dissection of adult human neocortical cell types and circuits. Sci Rep 2018, 8: 8407.
Schwarz N, Hedrich UBS, Schwarz H, Harshad PA, Dammeier N, Auffenberg E, et al. Human Cerebrospinal fluid promotes long-term neuronal viability and network function in human neocortical organotypic brain slice cultures. Sci Rep 2017, 7: 12249.
Andersson R, Safari C, Dods R, Nango E, Tanaka R, Yamashita A, et al. Serial femtosecond crystallography structure of cytochrome c oxidase at room temperature. Sci Rep 2017, 7: 4518.
Lee C, Harkin EF, Yin X, Naud R, Chen S. Cell-type-specific responses to associative learning in the primary motor cortex. Elife. 2022, 11: e72549.
Kim MH, Radaelli C, Thomsen ER, Monet D, Chartrand T, Jorstad NL, Mahoney JT, Taormina MJ, Long B, Baker K, Bakken TE. Target cell-specific synaptic dynamics of excitatory to inhibitory neuron connections in supragranular layers of human neocortex. Elife. 2023, 12: e81863.
El-Shamayleh Y, Ni AM, Horwitz GD. Strategies for targeting primate neural circuits with viral vectors. J Neurophysiol 2016, 116: 122–134.
Lee C, Lavoie A, Liu J, Chen SX, Liu BH. Light up the brain: The application of optogenetics in cell-type specific dissection of mouse brain circuits. Front Neural Circuits 2020, 14: 18.
Lim DH, Ledue J, Mohajerani MH, Vanni MP, Murphy TH. Optogenetic approaches for functional mouse brain mapping. Front Neurosci 2013, 7: 54.
Han Y, Huang K, Chen K, Pan H, Ju F, Long Y, et al. MouseVenue3D: A markerless three-dimension behavioral tracking system for matching two-photon brain imaging in free-moving mice. Neurosci Bull 2022, 38: 303–317.
Shen Y, Ding LF, Yang CY, Xu F, Lau PM, Bi GQ. Mapping big brains at subcellular resolution in the era of big data in zoology. Zool Res 2022, 43: 597–599.
Fang R, Xia C, Close JL, Zhang M, He J, Huang Z, et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 2022, 377: 56–62.
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM III, et al. Comprehensive integration of single-cell data. Cell 2019, 177: 1888-1902.e21.
Zhao M, Ren M, Jiang T, Jia X, Wang X, Li A, et al. Whole-brain direct inputs to and axonal projections from excitatory and inhibitory neurons in the mouse primary auditory area. Neurosci Bull 2022, 38: 576–590.
Acknowledgements
This review was supported by the Natural Science Foundation of China (81961128021 and 82371095), the National Key R&D Program of China (2022YEF0203200), the Guangdong Provincial Key R&D Programs (2018B030335001), and the Science and Technology Program of Guangzhou (202007030011 and 202007030010).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors claim that there are no conflicts of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shen, Y., Shao, M., Hao, ZZ. et al. Multimodal Nature of the Single-cell Primate Brain Atlas: Morphology, Transcriptome, Electrophysiology, and Connectivity. Neurosci. Bull. 40, 517–532 (2024). https://doi.org/10.1007/s12264-023-01160-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-023-01160-4