Abstract
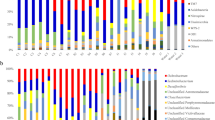
Bacterial clone libraries of the gut microbiota of nurtured and starved Cylindroiulus fulviceps specimens displayed the predominance of the phyla Bacteroidetes (55 and 37 %, respectively) and Proteobacteria (40 and 35 %, respectively) and a high similarity to bacteria previously detected in the intestinal tract of termites and beetles, which are known to harbor symbiotic bacteria essential for digestive activity. Bacterial isolates were dominated by Proteobacteria (74 %), followed by members of the phyla Actinobacteria, Firmicutes and Bacteroidetes. PCR-DGGE fingerprints of the gut samples showed that intestinal bacteria were affected by starvation, although the change was not significant.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- DGGE:
-
denaturing gradient gel electrophoresis
- ICD:
-
intestines of diplopods fed with cow dung
- ISA:
-
intestines of starved diplopods
- OTUs:
-
operational taxonomic units
- PCO:
-
principal coordinate analysis
- RDP:
-
ribosomal database project
References
Byzov B.A.: Intestinal microbiota of millipedes, pp. 89–114 in H. König, A. Varma (Eds): Intestinal Microorganisms of Soil Invertebrates. Springer, Berlin 2006.
Byzov B.A., Chernjakovskaya T.F., Zenova G.M., Dobrovolskaya T.G.: Bacterial communities associated with soil diplopods. Pedobiologia40, 67–79 (1996).
Cole J.R., Chai B., Farris R.J., Wang Q., Kulam S.A., Mcgarrell D.M., Garrity G.M., Tiedje J.M.: The Ribosomal Database Project (RDP-II): sequences and tools for high-throughput rRNA analysis. Nucl.Acids Res. 33, D294–D296 (2005).
Desantis T.Z., Hugenholtz P., Keller K., Brodie E., Larsen N., Piceno Y.M., Phan R., Andersen G.L.: NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucl.Acids Res. 34, W394–W399 (2006).
Dillon R.J., Webster G., Weightman A.J., Dillon V.M.: Composition of acridid gut bacterial communities as revealed by 16S rRNA gene analysis. J.Invertebr.Pathol. 97, 265–272 (2008).
Egert M., Wagner B., Lemke T., Brune A., Friedrich M.W.: Microbial community structure in midgut and hindgut of the humusfeeding larva of Pachnoda ephippiata (Coleoptera:Scarabaeidae). Appl.Environ.Microbiol. 69, 6659–6668 (2003).
Egert M., Stingl U., Bruun L.D., Pommerenke B., Brune A., Friedrich M.W.: Structure and topology of microbial communities in the major gut compartments of Melolontha melolontha larvae (Coleoptera:Scarabaeidae). Appl.Environ.Microbiol. 71, 4556–4566 (2005).
Feinstein L.M., Sul W.J., Blackwood C.B.: Assessment of bias associated with incomplete extraction of microbial DNA from soil. Appl.Environ.Microbiol. 75, 5428–5433 (2009).
Hongoh Y., Ohkuma M., Kudo T.: Molecular analysis of bacterial microbiota in the gut of the termite Reticulitermes speratus (Isoptera: Rhinotermitidae). FEMS Microb.Ecol. 44, 231–242 (2003).
Hongoh Y., Ekpornprasit L., Inoue T., Moriya S., Trakulnaleamsai S., Ohkuma M., Noparatnaraporn N., Kudo T.: Intracolony variation of bacterial gut microbiota among castes and ages in the fungus-growing termite Macrotermes gilvus. Mol. Ecol. 15, 505–516 (2006).
Hopkin S.P., Read H.J.: The Biology of Millipedes. Oxford University Press, New York 1992.
Hugenholtz P., Goebel B.M., Pace N.R.: Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J.Bacteriol. 180, 4765–4774 (1998).
Kemp P.F., Aller J.Y.: Estimating prokaryotic diversity: when are 16S rDNA libraries large enough? Limnol.Oceanogr.Methods2, 114–125 (2004).
Kersters K., de Vos P., Gillis M., Swings J., Vandamme P., Stackebrandt E.: Introduction to the Proteobacteria, pp. 3–37 in M. Dwarkin, S. Falkow, E. Rosenberg, K.-H. Schleifer, E. Stackebrandt (Eds): The Prokaryotes 5, 3rd ed. Springer, New York 2006.
Knapp B.A., Seeber J., Podmirseg S.M., Rief A., Meyer E., Insam H.: Molecular fingerprinting analysis of the gut microflora of Cylindroiulus fulviceps (Diplopoda). Pedobiologia52, 325–336 (2009).
Kumar S., Tamura T., Nei M.: MEGA3: integrated software for molecular evolutionary genetic analysis and sequence alignment. Brief Bioinform. 5, 150–163 (2004).
Lavelle P., Bignell D., Lepage M., Wolters V., Roger P., Ineson P., Heal O.W., Dhillion S.: Soil function in a changing world: the role of invertebrate ecosystem engineers. Eur.J.Soil Biol. 33, 159–193 (1997).
Lee A.H., Husseneder C., Hooper-Bui L.: Culture-independent identification of gut bacteria in fourth-instar red imported fire ant, Solenopsis invicta BUREN, larvae. J.Invertebr.Pathol. 98, 20–33 (2008).
Loy A., Lehner A., Lee N., Adamczyk J., Meier H., Ernst J., Schleifer K., Wagner M.: Oligonucleotide microarray for 16S rRNA gene-based detection of all recognized lineages of sulfate-reducing prokaryotes in the environment. Appl.Environ. Microbiol. 68, 5064–5081 (2002).
Marialigeti K., Contreras E., Barabas G., Heydrich M., Szabo I.M.: True intestinal actinomycetes of millipedes (Diplopoda). J.Invertebr.Pathol. 45, 120–121 (1985).
Mühling M., Woolven-Allen J., Murrell J.C., Joint I.: Improved group-specific PCR primers for denaturing gradient gel electrophoresis analysis of the genetic diversity of complex microbial communities. ISME J. 2, 379–392 (2008).
Muyzer G., de Waal E.C., Uitterlinden A.G.: Profiling of complex microbial populations by denaturing gradient gel electrophoresis analyses of polymerase chain reaction-amplified genes for 16S rRNA. Appl.Environ.Microbiol. 59, 695–700 (1993).
Nakajima H., Hongoh Y., Noda S., Yoshida Y., Usami R., Kudo T., Ohkuma M.: Phylogenetic and morphological diversity of Bacteroidales members associated with the gut wall of termites. Biosci.Biotechnol.Biochem. 70, 211–218 (2006).
Rawlins A.J., Bull I.A., Poirier N., Ineson P., Evershed R.P.: The biochemical transformation of oak (Quercus robur) leaf litter consumed by the pill millipede (Glomeris marginata). Soil Biol.Biochem. 38, 1063–1076 (2006).
Schmitt-Wagner D., Friedrich M.W., Wagner B., Brune A.: Phylogenetic diversity, abundance, and axial distribution of bacteria in the intestinal tract of two soil-feeding termites (Cubitermes spp.). Appl.Environ.Microbiol. 69, 6007–6017 (2003).
Schrott M., Knapp B.A., Wagner A.O., Seeber J., Meyer E., Insam H.: Molecular and cultivation-dependent analyses of the gut microbiota of Lumbricus rubellus and Cylindroiulus fulviceps, in K. Tajovský (Ed.): Proc. 10th Central European Workshop Soil Zoology, České Budějovice (Czechia) 2009.
Seeber J., Seeber G.U.H., Kässler W., Langel R., Scheu S., Meyer E.: Abundance and trophic structure of macrodecomposers on alpine pastureland (Central Alps, Tyrol): effects of abandonment of pasturing. Pedobiologia49, 221–228 (2005).
Vu A.T., Nguyen N.C., Leadbetter J.R.: Iron reduction in the metal-rich guts of wood-feeding termites. Geobiology2, 239–247 (2004).
Wang Q., Garrity G.M., Tiedje J.M., Cole J.R.: Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl.Environ.Microbiol. 73, 5261–5267 (2007).
Ward D.M., Weller R., Bateson M.M.: 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature345, 63–65 (1990).
Warnecke F., Luginbühl P., Ivanova N., Ghassemian M., Richardson T.H., Stege J.T., Cayouette M., Mchardy A.C., Djordjevic G., Aboushadi N., Sorek R., Tringe S.G., Podar M., Martin H.G., Kunin V., Dalevi D., Madejska J., Kirton E., Platt D., Szeto E., Salamov A., Barry K., Mikhailova N., Kyrpides N.C., Matson E.G., Ottesen E.A., Zhang X., Hernandez M., Murillo C., Acosta L.G., Rigoutsos I., Tamayo G., Green B.D., Chang C., Rubin E.M., Mathur E.J., Robertson D.E., Hugenholtz P., Leadbetter J.R.: Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature450, 560–569 (2007).
Yang H., Schmitt-wagner D., Stingl U., Brune A.: Niche heterogeneity determines bacterial community structure in the termite gut (Reticulitermes santonensis). Environ.Microbiol. 7, 916–932 (2005).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Knapp, B.A., Seeber, J., Rief, A. et al. Bacterial community composition of the gut microbiota of Cylindroiulus fulviceps (diplopoda) as revealed by molecular fingerprinting and cloning. Folia Microbiol 55, 489–496 (2010). https://doi.org/10.1007/s12223-010-0081-y
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-010-0081-y