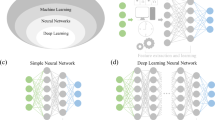
Abstract
Current imaging methods for diagnosing breast cancer (BC) are associated with limited sensitivity and specificity and modest positive predictive power. The recent progress in image analysis using artificial intelligence (AI) has created great promise to improve BC diagnosis and subtype differentiation. In this case, novel quantitative computational methods, such as radiomics, have been developed to enhance the sensitivity and specificity of early BC diagnosis and classification. The potential of radiomics in improving the diagnostic efficacy of imaging studies has been shown in several studies. In this review article, we discuss the radiomics workflow and current handcrafted radiomics methods in the diagnosis and classification of BC based on the most recent studies on different imaging modalities, e.g., MRI, mammography, contrast-enhanced spectral mammography (CESM), ultrasound imaging, and digital breast tumosynthesis (DBT). We also discuss current challenges and potential strategies to improve the specificity and sensitivity of radiomics in breast cancer to help achieve a higher level of BC classification and diagnosis in the clinical setting. The growing field of AI incorporation with imaging information has opened a great opportunity to provide a higher level of care for BC patients.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
The increasing number of breast cancer (BC) in recent years has made it the most common malignancy of women, with more than two million annual diagnoses worldwide [1]. Despite all the efforts to optimize the screening and diagnosis of BC, its survival rate is 50–60%, and its mortality is assumed to increase to 43% by 2030 [2]. Therefore, improving screening and diagnostic methods to achieve an early diagnosis of BC is mandatory to improve the treatment outcome and survival. Based on the American College of Radiology (ACR) and the Society of Breast Imaging (SBI) recommendations, all women must be assessed regarding BC by age 30. The annual mammographic screening in women older than 40 can reduce mortality by up to 40%. The current guidelines do not consider an upper age limit for BC screening [3]. Despite the considerable reduction of BC mortality via screening, late-stage diagnosis, the low specificity of current screening guidelines, and unnecessary biopsy sampling are still the main challenges [4].
Current guidelines for BC diagnosis are based on clinical evaluation of the patient and qualitative radiologic imaging using mammography, magnetic resonance imaging (MRI), ultrasound imaging, and digital breast tomosynthesis (DBT). These methods have multiple limitations, such as dependency on the radiologist’s skill and experience, suboptimal positive predictive value (PPV), and suboptimal sensitivity (approximately 70%) [5].
To obtain a definitive diagnosis of BC, histopathologic evaluation of the biopsy sample is required as the gold standard method of diagnosis. Biopsy obtaining is an invasive procedure, and its efficacy can be limited by the heterogeneity of BC, sampling errors, problematic sampling in small and necrotic tumors, beauty problems, and suboptimal accuracy of biopsy in large tumors [6]. Therefore, developing novel methods, specifically those based on artificial intelligence (AI)-based computational image analysis, could yield improved specificity and sensitivity and earlier diagnosis of BC, finally improving the outcome and quality of life.
It has been postulated that medical images include much more information than what can be seen by the naked human eye. Recently, a new field of computer-aided detection (CAD), named radiomics, has been developed, which works by extracting hand-crafted quantitative features from medical images using a high-throughput approach [7]. The extracted shape and textural features of radiomics are imaging biomarkers that reflect the mechanisms of cancer cells at the cellular and molecular level, which the human eye cannot qualitatively evaluate. Therefore, these data can be used as clues for the accurate diagnosis of malignancies, reduce the application of invasive diagnostic methods, improve the diagnostic accuracy and classification of the tumor, predict treatment response, and predict the survival and prognosis of the patients. Using radiomics methods could improve the early diagnosis of BC with higher specificity and sensitivity, discrimination of benign BC masses from malignant masses, tumor molecular subtype classification, prediction of the prognosis, and optimizing the treatment method [8].
This review differs from previously published reviews since it provides and updated review on the application of radiomics-based model for screening and diagnosis of BC. In this manuscript, we aim to comprehensively review the current screening and diagnostic protocols of BC, the standard radiomics workflow in BC, and the recent progress of radiomics in the screening and diagnosis of BC based on different radiologic methods with comparison to radiologist-based imaging. We also discuss the limitations and challenges confronted, and potential future directions to overcome these challenges. Previously, several reviews have been published in this area [9,10,11,12,13,14]. Although they discuss the role of radiomics in BC, there is a lack of specific addresses to the current advances in screening and diagnosis of BC using radiomics models. Thus, in this review, we have addressed and updated the current knowledge gaps on the screening and diagnosis of BC based on radiomics models.
2 Radiomics workflow
The application of radiomics in oncology imaging requires six main steps: image acquisition, segmentation and region-of-interest (ROI) selection, feature extraction, feature selection, model training, and validation. Figure 1 explains the radiomics workflow in BC. The application of radiomics in BC is a complex issue. In this regard, Lambin et al. have introduced a BC-specific radiomics quality score (RQS). Authors are highly encouraged to use this standardized method in future studies on BC radiomics (refer to [15] for more information).
2.1 Image acquisition
The first step is the acquisition of high-quality images. Standardized protocols, such as quantitative imaging biomarker alliance and quantitative imaging network [16], have been developed to improve the reproducibility and quality of the images. Moreover, the image biomarker standardization initiative by Zwanenburg et al. [17] is a reliable source for standardization of the workflow for image acquisition and quantization. This initiative includes standard steps for image processing, feature extraction, digital phantom texture matrices, and radiomics reporting guidelines and nomenclature. After acquisition, images commonly undergo pre-processing steps, e.g., re-sampling, re-segmentation, re-quantization, etc.
2.2 Segmentation and ROI selection
The second step is segmenting the ROI, the area from which the radiomic features will be extracted. To obtain reliable data, segmentation should be reproducible and accurate. ROI segmentation can be conducted automatically, semi-automatically, or manually. Manual segmentation of the ROI by expert radiologists is the gold standard method; however, some studies have used semi-automatic or automatic processes, which are superior if extensive data is being evaluated [18]. Considering semi-automatic methods, two main strategies can be used for ROI segmentation: region-based and threshold-based. Threshold-based methods segment the image based on the contrasts and intensities of the images, while region-based methods work based on the voxel similarity and connection [10]. Moreover, pre-processing is a necessary step that should be conducted before feature extraction to enhance the quality and consistency of medical images, which is vital for accurate and reliable radiomic feature extraction and model construction. The accuracy and reliability of the radiomics model heavily depend on the quality of the input data, making image preprocessing a crucial step. Appropriate pre-processing steps include image normalization (to adjust the pixel intensity values to a standard range), noise reduction by Gaussian and median filters (to reduce the noise in the acquired images), artifact correction (to reduce the artifacts caused by imaging equipment, patient movement, or implants), registration (to align images from different modalities or time points to a common coordinate system), and resampling (to adjust the image resolution or voxel size to ensure consistency across different images) [19].
2.3 Feature extraction
The third step is the computational extraction of features from the ROI, conducted through standard mathematical processes. These features can be divided into four different subgroups. The first group of features is shape (morphological) features, which are 3D-reconstructed images and include volume, shape, border heterogeneity, surface-to-volume ratio, sphericity, compactness, etc. An example of shape features is comparing the surface-to-volume ratio in two tumors with the same volume, of which one is speculated and the other is round. The surface-to-volume percentage is higher in speculated tumors and thus has a higher probability of malignancy. At the same time, it is lower in round tumors and thus has a higher likelihood of being benign. The second group of features is the histogram-based features commonly used for predicting patient prognosis. These features include mean, median, kurtosis, skewness, standard deviation, minimum, maximum, average volume, entropy, and uniformity [20]. The third group of features includes the textural features, which evaluate the voxels and the neighbors and commonly demonstrate intertumoral heterogeneity. Gray-level size zone matrix (GLSZM), Neighboring Gray Tone Difference Matrix (NGTDM), neighboring gray level dependence matrix (NGLDM), Grey Level Distance Zone Matrix (GLDZM), fractal self-similarity, gray-level run-length matrix, and gray-level co-occurrence matrix (GLCM) are the standard methods that can be used to extract textural features from images. Moreover, transform-based features, including wavelet or Laplacian of Gaussian, are histogram or texture features extracted from transformed images by various filtering convolutions. These features reflect the spatial relationships of the adjacent pixels [21]. Depeursinge et al. [22] have introduced standardized convolutional filtering for radiomics which can be used for standardization in further radiomics studies on BC. Several tools have been developed to extract the features from images, including the CGITA, 3DSlicer, Mazda, etc., which are open-access and available online [10].
2.4 Feature selection
The fourth step of radiomics is feature selection, which is a necessary step in radiomics workflow since among thousands of features, only a number are strongly related to the outcome. Through feature selection, the most reproducible and repeatable features are selected for training the model. In this context, statistical methods and machine learning (ML) strategies, such as random forest (RF), principal component analysis (PCA), and least absolute shrinkage and selection operator (LASSO), have been extensively applied in different radiomics studies [23]. The fifth step of the radiomics process is model construction. AI has been widely used to obtain predictive models for the extracted features. Several methods have been used in this context, such as RF, XG booster classifier, regularized linear regression (LR), support vector machine (SVM), and k-nearest neighborhood (KNN) [24, 25].
2.5 Validation
The last step is validating the model on the test and validation set. Validation is a critical step in radiomics analysis, ensuring that the predictive models developed are robust, reliable, and generalizable. [13]. Different methods could be used to validate the radiomics-based model. The main methods include Cross-Validation (either by K-fold cross-validation or Leave-One-Out Cross-Validation (LOOCV)), independent validation (on an external dataset from a different center or on a temporal dataset acquired at a different time point at the same center), bootstrapping, and hold-out validation (by splitting the primary dataset into training and test set) [26, 27]. The appropriate splitting and validation method should be selected according to the purpose of analysis, number of available data, etc.
The hold-out validation method splits data into training, validation, and test sets. The training set trains the model to learn the data’s underlying patterns, features, and relationships. The largest portion of the dataset is allocated to the training set to provide the model with sufficient information to learn effectively. The validation set tunes the model and selects the best hyperparameters, provides an unbiased evaluation of the model during the training phase, and helps prevent overfitting. Commonly, a smaller dataset portion is allocated to the validation set. The test set is used to evaluate the performance of the final model by providing an unbiased estimate of the model’s performance on new, unseen data. Usually, a portion of the dataset is allocated to the test set, typically around 20–30% of the total data, ensuring it is representative of the overall population [28]. Using one or a combination of these validation methods, radiomics models can be rigorously evaluated for accuracy, reliability, and generalizability, ensuring robust clinical applications in BC screening and diagnosis.
3 Current application of radiomics for breast cancer screening, diagnosis, and classification
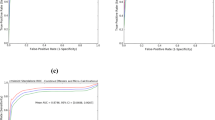
Radiomics, first used for head and neck cancer imaging [29], has been recently studied for screening, diagnosis, predicting treatment response, predicting lymph node metastasis, predicting recurrence chance, and prediction of prognosis in BC [13]. The most studied modality for applying radiomics approaches in BC is MRI. However, other modalities have also been studied, including contrast-enhanced spectral mammography (CEST), standard mammography, US, and DBT. Studies have shown that adding radiomics to the standard radiological processing of BC can increase its diagnostic accuracy [30]. Here, we aim to do a modality-based review of the current progress of radiomics for BC, focusing on BC screening and diagnosis, and compare AI-assisted BC screening/diagnosis with radiologist-based methods (Table 1). The percentage of each modality application based on Table 1 is presented in Fig 2. We also discuss the potential of radiomics as the developing modality for improving the accuracy of BC screening and diagnosis to decrease the bulk of invasive breast biopsies.
Percentage of each modality application for constructing radiomics models for breast-cancer diagnosis (based on data from Table 1). MRI: magnetic resonance imaging, CESM: contrast-enhanced mammography, DBT: digital breast tomosynthesis
3.1 Magnetic resonance imaging (MRI)
MRI is traditionally used to screen high-risk patients with positive BC gene-1, 2 (BRCA1, 2), specifically when US or mammography show suspicious findings. However, its role in screening intermediate and average-risk women is also being considered [31]. MRI has a high sensitivity for detecting malignant breast lesions; however, since malignant and benign breast lesions share similar characteristics, it has a low specificity for BC detection. Figure 2 is an example of a typical multiparametric breast image of a benign patient, including T1-weighted, T2-weighted, and DCE images [32, 33]. Based on the evidence, MRI can successfully discriminate between malignant and benign breast lesions in up to 72% of the cases [34]. Diffusion-weighted imaging (DWI) and dynamic contrast-enhancing (DCE)-MRI are the most studied MRI modality to discriminate benign from malignant breast lesions in radiomic studies [35]. Most studies focused on first- and second-order statistics. The primary target lesions were Breast Imaging-Reporting and Data System (BI-RADS) 4 and 5 for screening/diagnostic purposes. For example, in a 2018 study that included 222 patients, the authors aimed to develop a radiomic MRI model to predict malignancy in suspicious BI-RADS4 and five breast lesions. Results showed that a kurtosis-diffusion-weighted imaging model improved the sensitivity and specificity of MR-based discrimination of malignant breast lesions from benign breast lesions [36].
Typical multiparametric breast image of a benign patient. a Dynamic contract enhanced, b T2-weighted, c T1-weighted, d Pharmacokinetic-DCE (PK-DCE) overlay of K trans and EVF, where red indicates high K trans and blue demonstrates low K trans e ADC maps, and f whole breast entropy feature map (Licensed under CC BY 4.0, Source: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5686135/) [33]
More recently, multiparametric MRI, combining DCE and DWI-MRI with improved specificity, has been extensively used to extract features to reduce unnecessary biopsies [37]. In a study by Parekh et al. [33], multiparametric MRI was used for texture analysis of benign and malignant breast lesions. The results indicated that the radiomic feature map (RFM) related to tumor vascular and cellular heterogeneity significantly differed between benign and malignant lesions regarding different apparent diffusion coefficients and perfusion parameters. The multiview IsoSVM model consistently demonstrated a sensitivity of 93%, a specificity of 85%, and an area under the curve (AUC) of 0.91 in discriminating between benign and malignant breast lesions. In another study by Naranjo and colleagues, multiparametric MRI-based machine learning outperformed DWI and DCE-MRI in distinguishing benign and malignant breast lesions [38].
Although MRI is not routinely used for BC screening and diagnosis, its application could be more bright in discriminating suspicious lesions to avoid unnecessary biopsies and their cosmetic/financial burden to the patients. In a study, Pötsch et al. developed an AI classifier trained on 4D radiomic features of DCE-MRI of BI-RADS4 and five images. The AI classifier showed an AUC = 0.80 for predicting malignancy in testing and 0.85 in the validation set, respectively. More importantly, further analysis demonstrated that the classifier could reduce the rate of unnecessary biopsies up to 36.2% with a false negative score of only 4.5%. This study showed the great potential of a radiomic-based AI classifier in reducing unnecessary biopsies of suspicious BI-RADS-4 and five lesions [39].
Since different malignant breast lesions show other invasive behaviors, MRI-based radiomics models could also assist in identifying the phenotype/genotype of the BC, such as discrimination between triple-negative breast cancer (TNBC) and non-TNBC breast lesions [40]. In a study by Ma et al., the radiomic feature-based model showed an AUC = 0.741 for cross-validation and 0.867 for the testing dataset in differentiating TNBC from non-TNBC breast lesions [18].
Furthermore, the background parenchymal heterogeneity of the tumor can be identified by extracting its radiomic features, which can improve the potential to discriminate breast lesions and specifically define TNBC [41]. In a study, quantitative texture features of background parenchyma were extracted to identify the TNBC from other types of BC. The results indicated that radiomic features of the lesion had an AUC = 0.782 in discriminating TNBC from non-TNBC tumors. When the features of the background parenchymal features were added, the AUC was significantly improved to 0.878 [42]. Similar results were obtained in another study that combined breast-parenchymal enhancement (BPE) features with tumor-extracted features [41].
Considering BC screening and diagnosis, comparing the AI-based model’s diagnostic performance with radiologists' performance could be attractive. In a multicenter study, Naranjo et al. compared the performance of a radiomic-based ML decision-making model with the performance of two radiologists in discriminating between benign and malignant lesions. The performance of the multiparametric MRI radiomics with apparent diffusion coefficient (ADC) and BI-RADS was similar to the implementation of radiologists in discriminating benign from malignant lesions (p = 0.39). Thus, the authors concluded that this method could assist the decision-making of less experienced radiologists in differentiating between benign and malignant lesions [32].
Moreover, deep learning (DL) provides a more robust discriminative power. In a 2018 study by Truhn et al., a convolutional neural network (CNN)-based model outperformed radiomics in discriminating benign and malignant breast lesions. However, radiologists’ performance was superior compared to both of them [43]. Another study’s results demonstrated that combining the multiparametric MRI-based DL model improved the diagnostic performance of two junior radiologists, from an AUC = 0.823 to 0.876 and 0.833 to 0.885, respectively. However, the application of this model achieved comparable results to that of senior radiologists’ performance with an AUC = 0.944 in discriminating TNBC from fibroadenoma lesions [44].
3.2 Mammography
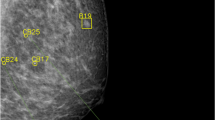
Mammography is the most used diagnostic modality for BC screening. Architectural distortions, microcalcifications, and tissue asymmetry are the most important biomarkers for BC diagnosis in mammography [45]. Figure 3 describes mammographic findings in a 45-year-old woman with infiltrating ductal carcinoma of the right breast [46]. Figure 4a, b describe mammography images of both breasts show heterogeneous dense breasts with left breast retroareolar, right breast retroareolar, and right breast lower inner quadrant dense lesions with obscured margins [47]. Based on different studies, the sensitivity of mammography for BC screening is 75–90% [48]. Its specificity for BC screening is consistently 80–90% [49, 50]. Although adjunctive ultrasonography and mammography can improve the accuracy of BC screening, specifically in small breast lesions under 2 cm [51], novel radiomics methods could improve its accuracy more efficiently.
A 45-year-old woman with infiltrating ductal carcinoma of the right breast. a a craniocaudal and mediolateral oblique projection b radiogram of the right breast with the presence of polymorphic microcalcifications extended to the upper sectors (arrow). (Licensed under CC BY 4.0, Source: https://www.mdpi.com/2227-9032/11/11/1596) [46]
Digital Mammography and Digital Breast Tomosynthesis imaging in a 40-year-old female presented with bilateral breast lumps. A Mediolateral oblique and B Craniocaudal Digital Mammography images of both breasts show heterogeneous dense breasts (ACR C) with left breast retroareolar, right breast retroareolar, and right breast lower inner quadrant dense lesions with obscured margins (arrows). No microcalcifications or speculated masses. C Mediolateral oblique and D Craniocaudal Digital Breast Tomosynthesis images show more margin characterizations of the lesions, which are medium-dense, well-defined rounded lesions with smooth margins and a characteristic halo sign (arrowheads). The lesions were classified as BI-RADS 3 according to Digital Mammography and BI-RADS 2 according to Digital Breast Tomosynthesis. Histopathological examination revealed simple cysts. (Licensed under CC BY 4.0, Source: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC10137942/#:~:text=The%20combination%20of%20DM%20and,for%20proper%20BI%2DRADS%20categorization) [168]
Mammography’s overall sensitivity and specificity for BC diagnosis is around 90%; however, a PPV of 15% has been reported for mammography [52]. Several studies have focused on improving the accuracy of BC diagnosis using AI-based radiomics approaches. AI-based radiomics models applied to mammography can reduce the burden on radiologists and improve diagnostic accuracy. In 2018, a retrospective study by Mao et al. aimed to develop radiomic features for diagnosing BC using mammography images. Among different regression models, the LR showed better results, having sensitivity, specificity, and diagnostic accuracy of 0.983, 0.975, and 0.978 for training data and 0.867, 0.900, and 0.886 for testing data, respectively. The authors concluded that quantitative mammography imaging could be an excellent diagnostic approach for BC diagnosis [53].
Combining trained AI-based DL tools with radiologist-based assessments could provide higher accuracy in identifying BC. In a study in 2020, Schaffter et al. aimed to evaluate whether combining radiomics with radiologist assessment can improve the accuracy of mammography-based BC screening. To do this, they collected data from 144,231 mammograms to train and validate a DL algorithm. The analysis revealed that the combination of this AI-based diagnosing modality with the radiologist assessments had an AUC = 0.942, higher than the AI-based and radiologist-based assessments. Moreover, the combination method also showed a higher specificity of 92%. The authors concluded that although the ML algorithms were inferior to radiologist assessments, combining these methods with a single-reader radiologist assessment improved the accuracy of mammography interpretations for BC screening [54]. Moreover, in another study, the AI-based system did better than 61.4% of the radiologists in diagnosing the malignancy, meaning that a high-performance AI system can do better than an average radiologist in diagnosing BC in the clinical setting [55].
Another paper by the same authors evaluated the feasibility of reducing the workload of mammographic studies using AI-based systems to exclude the lower malignancy likelihood results. The results of this study indicated that setting the AI threshold of malignancy probability at 2 and 5 reduces the mammography-reading workload up to 17% and 47%, respectively. However, choosing two and five points was also associated with excluding 1% and 7% of actual positive exams, respectively. This study showed that pre-selection of mammography exams using an AI-based system could reduce radiologists’ screening mammography reading tasks by excluding the low-likelihood exams. Nevertheless, further studies are still indicated to define the best likelihood threshold at which the lowest likelihood exams are excluded, but the true-positive studies are not excluded [56].
A recently published meta-analysis study aimed to evaluate the diagnostic accuracy of current ML methods on mammography. Thirty-six studies were included, of which the overall specificity, sensitivity, and AUC were 0.84, 0.83, and 0.90 [57]. Nevertheless, since the authors indicated that Deek’s LR model showed a bias, the data of this meta-analysis should be cautiously interpreted. Another systematic review and meta-analysis study was conducted, including 19 studies that had used radiomics as the preoperative diagnostic method for BC. They found that radiomic models had an AUC of 0.91, specificity of 0.83, and sensitivity of 0.84 in detecting BC in the preoperative setting [58]. However, since the number of primary studies included in the meta-analysis was relatively small, this study’s results must be investigated by large-scale trials.
One of the essential goals of BC screening by mammography is to reduce the number of unnecessary biopsies in high-risk patients. Mammography-based radiomics have shown promising results in this area. An example is a study conducted by Drukker et al. that combined the quantitative three-compartment breast (3CB) image analysis with mammography radiomics to assess its potential to reduce unnecessary biopsy sampling. The results indicated that compared to conventional mammography, the combined method could reduce 35.3% of total biopsies and had a sensitivity of 97% [59]. Microcalcifications, especially when they are present in dense breast parenchyma, could be challenging to distinguish and, therefore, may lead to unnecessary biopsies. Applying radiomics to mammography images could improve detection accuracy and reduce unnecessary biopsies. In a study, texture analysis of the surrounding tissue of the microcalcifications (excluding microcalcifications) had an AUC of 0.96 and reduced unnecessary biopsies [60]. Another study by Wang et al. has shown similar results [61].
A significant barrier to mammography-based BC screening is high densities, which can mask the tumor and delay diagnosis. Density is commonly evaluated based on the BI-RADS density score. A study aimed to develop a texture-based model for predicting the risk of tumor masking in high-density mammograms. The extracted model had an AUC of 0.75, while volumetric breast density, age-adjusted BI-RADS density, and BI-RADS density had an AUC of 0.72, 0.71, and 0.64, respectively. The authors concluded that the radiomic texture metrics could predict the masking of the breast lesion more accurately than other prediction methods [62]. The features of the contralateral breast parenchyma hold important histopathological data that could improve the radiomic-based diagnosis of BC. In a retrospective study, Li et al. included mammography data from 182 patients, comparing the radiomic features of the lesions with their contralateral ROI breast parenchyma. Six radiomic features were selected, including skewness, speculation, circularity, margin sharpness, size, and power-law beta. The results of this study indicated that the combination of parenchyma and lesion classifier (AUC = 0.84 ± 0.03) did better than the lesion classifier alone (AUC = 0.79 ± 0.03) [63].
CNN and other DL models have also been used for BC diagnosis based on mammographic images, and have shown superior results than radiomics-based models. An example is a large-scale study conducted by Kooi et al. on 45,000 mammography samples. This study evaluated two approaches: a CNN and the other performed by manual feature setting. The results demonstrated that the fully automated CNN outperformed conventional CAD methods in diagnosing BC at high and low sensitivity modes [64].
3.3 Contrast-enhanced spectral mammography (CESM)
CESM is a developing method for BC diagnosis with high sensitivity after intravenous (IV) injection of contrast material [65, 66]. Recently, CESM has shown similar accuracy to MRI in discriminating breast lesions [67]. Previous studies have focused on utilizing radiomics-based methods for analyzing CESM in differentiating malignant from benign breast lesions [68]. In a study, Massafra et al. studied 464 features from CESM exams of 53 patients. They trained three different classifiers, including LR, naïve Bayes, and RF, on each subset of principal components, of which the RF-based classifier showed the best potential to predict the malignancy of the ROIs with a sensitivity of 88.37% and a specificity of 100% [69]. Radiomic feature selection using CESM can also differentiate invasive forms from non-invasive forms of BC. In a study, six different radiomic features were analyzed in terms of their power to discriminate the invasiveness of the BC. It was shown that the co-occurrence matrix (COM), in combination with the first-order histogram (HIS) or mutual information (MI) coefficient, had an accuracy of 87.4% for differentiation of invasive from non-invasive BC [70]. In another study, the authors compared the radiomics analysis of DCE-MRI and CESM in evaluating BC invasiveness. This retrospective analysis included data from 48 women with 49 biopsy-confirmed BC. Analysis of DCE-MRI radiomics showed an accuracy of 90% in discriminating invasive from non-invasive BCs, while it was 92% for CEM [71]. One of the challenges of radiographic diagnosis of BC is the small size of lesions, especially < 1 cm. Radiomics could also help detect small malignant lesions. In 2020, Lin et al. conducted a retrospective study on 139 patients with lesions of sub-1 cm diameter. Their study showed that a radiologic nomogram combined with CESM-based radiomic features and predictive factors of age and BI-RADS score had an AUC = 0.94 in identifying benign from malignant breast lesions of < 1 cm [72].
The perilesional area of the tumor contains essential information that could help diagnose malignant lesions. A retrospective study by Wang et al. aimed to analyze the radiomic features of the perilesional regions of 190 women with 223 breast lesions. They surveyed 4098 radiomic characteristics from seven ROIs. Among the methods used, radiomic features of the annular perilesional region of 3 mm had the highest AUC = 0.93 for distinguishing benign from malignant lesions. Moreover, the authors reported that combining the radiomic analysis of the ROI with the annular perilesional region of 3 mm had a higher AUC = 0.94; thus, the combinatory method could do better in diagnosing BC [73].
It is already distinguished that some factors, such as lesion size, can influence the interpretation of data derived from CESM [74]. To determine the factors influencing the understanding of radiomic features extracted from CESM, Sun et al. aimed to identify the factors that influence the classification of CESM-based radiomic models in benign and malignant breast lesions. They collected the misclassified data using the RF algorithm and LASSO regression models to do this. Multivariate analysis demonstrated that the presence of air-trapping infarcts and the small size of the lesion were the two factors that led to misinterpretation of the malignant breast lesions. However, considering misclassifications of benign breast lesions, the presence of ripple and/or rim artifacts and larger lesion size were the two crucial influencing factors [75].
Combining CESM radiomic feature analysis with another diagnostic method could improve the accuracy of benign/malignant lesion discrimination. In a study by Fusco et al., 54 patients with 79 histopathologically-proven BCs underwent CESM and DCE-MRI and were analyzed for 48 textural radiomic features. Considering CESM, skewness (AUC = 0.71) and kurtosis (AUC = 0.71) were the best predictors of malignancy. Consistently, considering components extracted from DCE-MRI, gray-level run-length matrix (AUC = 0.72), GLN (gray-level non-uniformity) (AUC = 0.72), entropy (AUC = 0.70), energy (AUC = 0.72), and RANGE (AUC = 0.72) were the best predictors of malignancy [76].
3.4 Digital breast tomosynthesis (DBT)
Mammography has low sensitivity in detecting breast lesions in dense breasts due to the overlapping of the tissues. Figure 4 c, d describes mediolateral oblique and craniocaudal DBT images. The lesions were classified as BI-RADS 2 according to DBT. Histopathological examination revealed simple cysts [47]. DBT is the pseudo-3D reconstruction of mammographic images that enhances the discrimination of breast tissues [77]. Therefore, one of the applications of DBT is to diagnose malignancy in breasts with dense tissue. Recently, AI-based radiomics approaches have been used in DBT to improve diagnostic accuracy [78, 79]. In 2018, Tgliafico et al. included two groups of patients: 20 cancer-detected exams with dense breasts and negative mammography, with exams from 20 health-matched individuals. Three radiomic features correlated with tumor size, including 90 percentile, skewness, and entropy. Moreover, entropy was related to the estrogen receptor status of the BC [80]. In another study by Sakai et al., using ML and radiomic features, an automated classifier was trained to distinguish benign from malignant breast lesions on DBT. Among different classifiers, the accuracy of the SVM-based classifier was 55%for benign lesions and 84% for malignant lesions [81].
Data extracted from the peritumoral area are essential for distinguishing benign and malignant lesions in DBT. In a recently published study, Niu et al. aimed to extract the DBT-based peritumoral radiomic features for differentiating benign lesions from malignant ones. They extracted radiomic features from the lesion and the 2 mm diameter of the peritumoral area. The radiographic nomogram incorporated menstruation status, age, and radimoic features, which showed a specificity of 0.946, sensitivity of 0.970, and AUC of 0.980 in the training set. However, it had a specificity of 0.966, a sensitivity of 0.909, and an AUC of 0.985 in the validation cohort. The authors concluded that the radiomic nomogram that integrates the clinical data (menstruation status and age) with peritumoral radiomic features could help increase the accuracy of BC diagnosis [78].
Microcalcification clusters can disturb the diagnosis of BC in DBT images because of their spanning across slices. Therefore, differentiating malignant and benign microcalcification clusters in DBT is a critical issue radiomics studies address [82]. In a study, the authors developed a semi-automatic segmentation radiomic-based approach for differentiating benign microcalcifications clusters from malignant ones. Among different features, the view-based mode and case-based mode of both 2D and 3D-domain radiomics had the highest AUC of 0.834 and 0.868, respectively. Therefore, the authors concluded that radiomic-based models could perform well in differentiating microcalcification clusters that are challenging to diagnose on DBT [83]. In another study, to reduce the size of false positive calcifications in DBT, the authors developed a decision support system based on radiomic classifiers combined with the BI-RADS scoring system. The trained ML classifier reduced the false positive to half and improved the positive predictive value up to 50% [84].
3.5 Ultrasonography (US)
US is integral to the BC screening system, specifically in dense breasts. However, it has low specificity, increasing the rate of false-positive results and burdens unnecessary biopsies. Therefore, quantitative radiomics-based approaches have been widely studied to improve BC’s early diagnosis and classification and reduce unnecessary biopsies [85,86,87,88]. It has been illustrated that histogram, texture, and shape-oriented features are the essential gradients of US images to distinguish between benign and malignant lesions [89]. Figure 5 describes examples of ultrasound slices overlapped with intratumoral regions (green) and peritumoral regions (red) from two patients with positive axillary lymph node metastasis (ALN) and negative ALN [90]. In a recent study, Jabeen and colleagues proposed a new CNN model for predicting malignancy in US images. When validated on the Breast Ultrasound Images (BUSI) dataset, this model achieved the highest accuracy of 99.1% [91] (Fig. 6).
Examples of ultrasound slices overlapped with intratumoral regions (green) and peritumoral regions (red) from two patients. (Top) A patient with positive ALN. (Bottom) A patient with negative ALN. (Licensed under CC BY, no permission required, Source: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7006026/) [90]
Appropriate support medical decision-making models based in the US could improve the classification of breast lesions and reduce the rate of benign lesion biopsy. Interlenghi et al. proposed a ML model for predicting BI-RADS scores based on the US. The proposed model reduced the biopsy rate from 18 to 15% in benign lesions. The external validation of the model on two different datasets had a PPVPPV of 45.9% and 98% sensitivity, compared to the 41.5% PPV of a radiologist (p < 0.005) in the first dataset. Validation testing on the second dataset showed a PPV of 50.5% and sensitivity of 94%, compared to the PPV of 47.8% of a radiologist (p < 0.005). Moreover, compared to a board-certified breast radiologist, in six of nine images, the model outperformed the radiologist in giving a lower BI-RADS score to benign masses [92]. In another study, the proposed model reduced 67.86% of the unnecessary biopsies; however, a false-negative rate of 10.4% was observed [93].
DL models based on features extracted from morphological characteristics on US imaging can also distinguish the type of BC subtype. In a study, morphological characteristics of US images of 282 women with BC were included. While the TNBC subtype tended to have a parallel orientation and more round/oval morphology in the US, the luminal A subtype had a more distinctive feature but did not show a similar direction. Moreover, tumor size was associated with hypoechogenicity, a non-circumscribed margin, and irregular shape [94].
In another retrospective study, ten radiomic features were extracted from 206 percutaneously biopsied lesions (62 malignant and 144 benign). Different methods were used, of which the SVM had 76.9% specificity, 71.4% sensitivity, and the highest AUC of 0.840 [95]. Data from US imaging can also improve the differentiation of benign fibroadenoma from TNBC [79, 96,97,98,99]. A study by Du et al. developed a nomogram to differentiate TNBC from fibroadenoma based on radiomic features extracted from the US and patient clinical data. In the training cohort and validation, the nomogram had an AUC of 0.986 and 0.977, respectively, outperforming clinical models and radiomics signatures [100]. In a study to differentiate between phyllode tumors among fibroepithelial lesions in the US, the authors extracted 93 radiomic features from 182 fibroepithelial lesions biopsied and diagnosed via core needle biopsy. In the validation set, the radiomics-based classifier had an AUC of 0.765, an accuracy of 0.703, a specificity of 0.5, and a sensitivity of 0.857 [101].
Nomograms combining radiomic results with data from the BI-RADS scoring system could provide additional important information on differentiating malignant lesions from benign lesions. In a 2019 study, Luo et al. developed a nomogram based on US BI-RADS 4 or 5 lesions and nine radiomic features of the US images. Of the 315 pathologically-proven samples, 211 were included in the training and 104 in the validation groups. The nomogram combining radiomic features and BI-RADS scoring showed a better discrimination power with an AUC = 0.928 than radiomics and BI-RADS scoring alone (p value = 0.029 and 0.011, respectively). Therefore, the authors concluded that the nomogram incorporating radiomic features with the BI-RADS category is a potentially useful method for predicting malignancy in BI-RADS 4 and 5 US lesions [102].
Tumor vasculature consists of essential data for differentiating malignant and benign lesions. Color Doppler US is used to assess the vascularity of breast lesions. Higher vascularity lesions are more suspicious of being malignant. In 2020, Moustafa et al. extracted quantitative radiomic features from the color Doppler US images to develop a diagnostic model for BC diagnosis. To establish an AdaBoost ensemble classifier, two Doppler and seven grayscale features were extracted from 159 Doppler exams combined with the BIRADSUS category and age information. Training the model based on color Doppler and grayscale features improved the AUC from 0.925 to 0.958, reduced borderline diagnosis, and enhanced the diagnostic performance [103]. In another study, Ternifi et al. developed a quantitative, non-contrast high-definition microvasculature imaging (HDMI) for BC detection in highly suspicious breast masses. Four tumor microvessel features, including spatial vascularity pattern (SVP), bifurcation angle (BA), Murray’s deviation (MD), and microvessel fractal dimension (mvFD), were considered. For lesions greater than 20 mm, the HDMI model had a sensitivity of 95.6 and a specificity of 100%. Adding the BI-RADS score to the HDMI biomarkers produced a prediction model with 89.2% specificity, 93.8% sensitivity, and an AUC of 0.97 [104].
Similar to other modalities, peritumoral data from the US could also provide additional information that could improve the classification of breast lesions. Fuse the info of multiple tumoral regions (FMRNet) is an example of a CNN developed for breast tumor classification by combining the radiomic signature feature from multiple tumoral areas. When tested on the UDIAT dataset, FMRNet had an accuracy and specificity of 0.945 [105].
4 Limitations, challenges, and future directions
Radiomics has been extensively applied in BC screening and diagnosis. Despite progress in this area, clinical application is yet to be available for radiomics, and we still have a long way to bring radiomics to the bedside [106]. Based on a study, it has been shown that radiomic studies have an overall insufficient scientific quality and radiomics reporting, specifically in the fields of open science category, clinical utility, and reproducibility. Moreover, this retrospective meta-analysis study evaluated the quality of radiomic studies via radiomics quality scoring (RQS). The results indicated a low rate of the 17 included studies [11]. This section discusses the limitations and challenges of radiomics application in BC screening and diagnosis and possible solutions.
The first and significant challenge is the reproducibility of the features [107]. The variability of radiomic features, the heterogeneity of imaging protocols (especially in MRI), the heterogeneity of processing methods, and the retrospective nature of the studies are the significant factors that dampen the reproducibility of radiomics studies [108]. It has been shown that voxel shape and slice thickness are the two features that have the most effect on the reproducibility of elements [109]. Another major factor that affects the reproducibility of radiomics is the method used for ROI segmentation. Although automated methods are suitable for extensive data, they can only explore their best function if the lesions have a well-defined margin [110].
Moreover, although the manual method is more accurate, it is time-consuming, and the intra and inter-interpreter variability reduces the reproducibility of the radiomics method [111]. The application of novel CNN-based methods, such as the Ensemble Deep Neural Network model, has contributed significantly to automating the segmentation task, as shown in a recently published study [112]. Another critical factor affecting the reproducibility of high-level features is the image quality, which has affected US-based radiomics studies more than other modalities [113]. Augmentation of the images using DL models can help gain better results. One of the methods used for image augmentation is the generative adversarial network (GAN) [114]. A study used GAN to generate high-quality US images of breast lesions. Using a CNN model for interpretation, synthetic GAN-augmented images resulted in better classification of malignant from benign breast lesions compared to the baseline method, with an accuracy of 90.41% [115].
Developing novel technologies and algorithms for high-quality image acquisition could yield better results and higher accuracy in radiomic studies shortly. Examples are novel MRI-based developing technologies, such as blood oxygen level-dependent (BOLD) MRI, ultrafast MRI, abbreviated MRI, and arterial-spin labeling (ASL)-MRI, which produce high-quality images for feature extraction [116,117,118]. The scanner model, imaging parameters, and imaging protocol heterogeneity are among other factors that may influence the result and reproducibility of the radiomics methods, specifically regarding MRI, which includes many more parameters than US or mammography [106]. An example of the scanner effect is a study that showed that compared to breast tumors, the radiomic features of the fibroglandular tissue are more dependent on the MRI scanner model [119,120,121].
Moreover, the same algorithm may show different results in two sample sizes. Therefore, the sample size is another issue that can influence the results of a study. From a statistics view, predictive models built on a more extensive study size have more accuracy ref. Thus, since most radiomic studies use a small size, their results must be externally validated on larger datasets.
In addition, variations in the imaging protocols and image post-processing for image normalization are two critical factors that influence the quality of the images and can thus reduce the reproducibility of BC radiomic studies. Many studies have investigated harmonization as a novel method for enhancing repeatability and reproducibility in different observers, scanners, and imaging protocols. For example, Whitney et al. [122] considered investigating the effects of harmonization on a database of radiomic features extracted from DCE-MRI. The authors evaluated their method on 3150 benign and malignant lesions from international datasets using the ComBat method, the current trend in harmonizing radiomic features. Results demonstrated that classification based on postharmonization features were higher than preharmonization features (p < 0.001). The authors concluded that radiomic feature harmonization enables the combination of different databases and achieves better results. Similar results of the harmonization technique were also observed in the study by Prinzi et al. [123].
Proper normalization protocols are required to minimize the effects of different factors on database quality. Nevertheless, according to the established normalization techniques in the texture analysis of MR images, authors must apply the normalization process in future studies. For example, applying intensity normalization processes [124], such as z-score normalization [125], linear histogram match [126], and N4 bias field correction [127], enhances the robustness of radiomics features. For more information on image intensity normalization, refer to [128,129,130].
The hardware used, dataset size and type, dataset quality, and data architecture are essential factors affecting the results when training the algorithms. Parallel computation strategies on graphics processing units (GPUs) can be used to reduce the time required for training the algorithms [131].
In addition, overfitting and validation are essential barriers in radiomics studies due to the extensive application of ML algorithms. Multicenter data could be a possible solution to this problem. Since multicentric studies might only be available for some, the best solution might be the application of open databases. Dataset accessibility is another solution that has an essential effect on the external validation and reproducibility of radiomics studies. Limited dataset sharing and the availability of the algorithms used in different studies are critical challenges. The Cancer Imaging Archive (TCIA) is the most used open database for radiomics studies; however, regarding BC, it only includes 1000 mammography and 500 breast MRI images, which is not sufficient [132]. Regarding mammography, several other public datasets have recently become available. These datasets include the BGH, DDSM, MIAS, INBreast, OPTIMAM, VICTRE, BCDR, and LAPIMO [133].
Several factors can affect the results of radiomics studies that discriminate benign and malignant lesions. The first issue is the modality used for imaging. When applied to distinct modalities, the same algorithm could demonstrate different performances [134]. Intra and inter-observer segmentation variability is another issue that can affect feature extraction results. Automatic segmentation of images could reduce the effects of this variability. However, a study showed that 32.8% of Pyradiomics and 41.6% of radiomiX features were robust, meaning that inter-observer variability did not affect them [111].
The progressing role of DL in radiomics has resulted in the development of fully automated models that can conduct all phases of radiomics workflow, from image segmentation to classification [135]. An example is a recent work performed by Vigil et al., which developed a dual-intended DL model that showed an accuracy of 78.5% in the classification of US breast lesions [136]. Several other models have been developed that focus on classifying BC, and most of them are surveyed on currently available online databases.
To improve the accuracy of screening BC, pre-screening risk assessment with thermal radiomics and AI can be used to generate a personalized risk score for BC. Thermalytix Risk Score (TRS) is an AI-based, radiation-free, non-invasive method for identifying high-risk populations based on the metabolic activity of breast tissue. Compared to age-based risk assessment, this method has shown a 21% improvement in AUC [2]. With the development of AI and DL strategies, novel methods incorporating the potential of AI may be used for radiation-free pre-screening personalized risk assessment of patients. This method could finally lead to higher accuracy in BC screening [137]. Another topic of significant interest is developing automatic predicting models based on transfer learning from neural network (NN) systems, which has gained much attention recently [138]. It has been shown that NN-based models are more accurate than conventional ML tools such as SVR/KNN; however, NNs are more expensive and have more complexities [139]. Currently, several CNN-based models have been developed and tested on different modalities, mostly mammogram databases, which have shown the high accuracy of these models in discriminating benign from malignant breast lesions [140, 141].
Another approach is developing nomograms for a more accurate diagnosis in BC studies. The concept is that different factors, such as age and menstrual cycle, can affect the density of the breast. Therefore, to obtain more reliable and accurate results, recent studies have incorporated radiomic studies with patient clinical information, such as age, menstrual cycle, and BI-RADS score, to develop more accurate nomograms. Nomograms have shown better accuracy and higher AUCs in predicting BC in different studies [142, 143] and should be considered in further studies.
One of the main challenges would be the identification of small-size lesions less than 1 cm by radiomic features. Despite the progress in improving the diagnostic accuracy of radiomics for subcentimeter lesions, future studies are still indicated to achieve more accurate models and methods [144]. Another significant issue is the undetermined zone, where benign and malignant tissues share the same radiomic features, accounting for 15% of lesions. The most common lesions of the undetermined zone are Fibroadenoma, phyllodes tumor, etc. An undetermined radiomics zone is a dilemma in the interpretation of radiomic studies. Despite studies showing the potential of computer-aided feature extraction to replicate the human BI-RADS score definition [145], currently, the radiomic features cannot discriminate the undetermined radiomic lesion. Therefore, expert radiologists’ interpretation of the data is still required to discriminate these lesions by defining the BI-RADS score [146].
DBT acquisition angle is another factor that can impact the quality of breast parenchymal feature findings in DBT. The effects of acquisition angles of 15° and 40° on radiomic-based BC diagnosis were analyzed in a study. GLCM features significantly differed between 15° and 40° images, while there was no difference in histogram-based features. The texture analysis revealed that the DBT acquisition angle affects the results of radiomics studies, especially GLCM [147].
Besides screening and diagnosis, recent studies have also focused on extracting radiomic features for predicting other aspects of BC, including classification of its molecular subtypes [148], axillary lymph node metastasis [149], treatment response [150], prediction of prognosis [151], prediction of survival [152], and prediction of its risk of recurrence [153]. The growing role of radiomics and AI in this area may assist patient management and contribute to the personalized treatment of patients in the future [154]. The promising results of current studies indicate the potential of radiomics to become part of the routine decision-making for clinical practice regarding BC [155].
5 Conclusion and discussion
In recent years, radiomics, combined with the expanding role of ML and AI, has shown great potential in BC research, specifically in improving BC screening and diagnostic accuracy. This review article presents the current progress of radiomic-based studies for BC screening and diagnosis using MRI, US, DBT, mammography, and CESM modalities. MRI is getting more attention in BC screening, especially after recent studies showing its potential to reduce unnecessary biopsies [39]. However, MRI-based radiomics cannot replace a biopsy’s role in diagnosing BC, at least in the near future [156]. Moreover, based on evidence, combining radiomic features of different modalities can improve the accuracy of diagnosis and reduce false-positive results. The big question is whether radiomic-based AI models can replicate the role of radiologists in diagnosing BC. Despite promising results of radiomic-based models’ performance, in most cases, these models did not perform better than expert radiologists [44]. However, radiomic-based ML models did improve the performance of less-experienced radiologists. Therefore, the development of radiomic-based models could improve the decision-making capability of radiologists in BC diagnosis. Moreover, the growing role of AI, ML models could improve the accuracy of radiomics and enhance its application in the screening and diagnosis of BC in the clinical setting.
Data availability
Data will be available upon request by reviewers.
References
Piechocki M, Koziołek W, Sroka D, Matrejek A, Miziołek P, Saiuk N, et al. Trends in incidence and mortality of gynecological and breast cancers in Poland (1980–2018). Clin Epidemiol. 2022;14:95–114.
Kakileti ST, Madhu HJ, Manjunath G, Wee L, Dekker A, Sampangi S. Personalized risk prediction for breast cancer pre-screening using artificial intelligence and thermal radiomics. Artif Intell Med. 2020;105: 101854.
Monticciolo DL, Malak SF, Friedewald SM, Eby PR, Newell MS, Moy L, et al. Breast cancer screening recommendations inclusive of all women at average risk: update from the ACR and society of breast imaging. J Am Coll Radiol. 2021;18(9):1280–8.
Meenalochini G, Ramkumar S. Survey of machine learning algorithms for breast cancer detection using mammogram images. Mater Today Proc. 2021;37:2738–43.
Pesapane F, Rotili A, Agazzi GM, Botta F, Raimondi S, Penco S, et al. Recent radiomics advancements in breast cancer: lessons and pitfalls for the next future. Curr Oncol. 2021;28(4):2351–72.
Tzeng YT, Liu SI, Wang BW, Chen YC, Chang PM, Chen IS, et al. The efficacy of thrombin-gelatin matrix in hemostasis for large breast tumor after vacuum-assisted breast biopsy. J Pers Med. 2022. https://doi.org/10.3390/jpm12020301.
Gillies RJ, Schabath MB. Radiomics improves cancer screening and early detection. Cancer Epidemiol Biomarkers Prev. 2020;29(12):2556–67.
Ghelich Oghli M, Shabanzadeh A, Moradi S, Sirjani N, Gerami R, Ghaderi P, et al. Automatic fetal biometry prediction using a novel deep convolutional network architecture. Phys Med. 2021;88:127–37.
Pesapane F, De Marco P, Rapino A, Lombardo E, Nicosia L, Tantrige P, et al. How radiomics can improve breast cancer diagnosis and treatment. J Clin Med. 2023;12(4):1372.
Conti A, Duggento A, Indovina I, Guerrisi M, Toschi N. Radiomics in breast cancer classification and prediction. Semin Cancer Biol. 2021;72:238–50.
Valdora F, Houssami N, Rossi F, Calabrese M, Tagliafico AS. Rapid review: radiomics and breast cancer. Breast Cancer Res Treat. 2018;169(2):217–29.
Crivelli P, Ledda RE, Parascandolo N, Fara A, Soro D, Conti M. A new challenge for radiologists: radiomics in breast cancer. Biomed Res Int. 2018;2018(1):6120703.
Lee SH, Park H, Ko ES. Radiomics in breast imaging from techniques to clinical applications: a review. Korean J Radiol. 2020;21(7):779–92.
Tagliafico AS, Piana M, Schenone D, Lai R, Massone AM, Houssami N. Overview of radiomics in breast cancer diagnosis and prognostication. The Breast. 2020;49:74–80.
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017;14(12):749–62.
Press RH, Shu HG, Shim H, Mountz JM, Kurland BF, Wahl RL, et al. The use of quantitative imaging in radiation oncology: a quantitative imaging network (QIN) perspective. Int J Radiat Oncol Biol Phys. 2018;102(4):1219–35.
Zwanenburg A, Leger S, Vallières M, Löck S. Image biomarker standardisation initiative. 2016. arXiv preprint arXiv:161207003.
Ma M, Gan L, Jiang Y, Qin N, Li C, Zhang Y, et al. Radiomics analysis based on automatic image segmentation of DCE-MRI for predicting triple-negative and nontriple-negative breast cancer. Comput Math Methods Med. 2021;2021:2140465.
Demircioğlu A. The effect of preprocessing filters on predictive performance in radiomics. Eur Radiol Exp. 2022;6(1):40.
Debbi K, Habert P, Grob A, Loundou A, Siles P, Bartoli A, et al. Radiomics model to classify mammary masses using breast DCE-MRI compared to the BI-RADS classification performance. Insights Imaging. 2023;14(1):64.
Demircioğlu A. Benchmarking feature selection methods in radiomics. Investigat Radiol. 2022;57(7):433–43.
Depeursinge A, Andrearczyk V, Whybra P, van Griethuysen J, Müller H, Schaer R, et al. Standardised convolutional filtering for radiomics. 2020. arXiv preprint arXiv:200605470.
Kim SM, Kim Y, Jeong K, Jeong H, Kim J. Logistic LASSO regression for the diagnosis of breast cancer using clinical demographic data and the BI-RADS lexicon for ultrasonography. Ultrasonography. 2018;37(1):36–42.
Guiot J, Vaidyanathan A, Deprez L, Zerka F, Danthine D, Frix A-N, et al. A review in radiomics: making personalized medicine a reality via routine imaging. Med Res Rev. 2022;42(1):426–40.
Teng X, Zhang J, Zwanenburg A, Sun J, Huang Y, Lam S, et al. Building reliable radiomic models using image perturbation. Sci Rep. 2022;12(1):10035.
Kim M, Lee JH, Joo L, Jeong B, Kim S, Ham S, et al. Development and validation of a model using radiomics features from an apparent diffusion coefficient map to diagnose local tumor recurrence in patients treated for head and neck squamous cell carcinoma. Korean J Radiol. 2022;23(11):1078–88.
Zhang J, Cao G, Pang H, Li J, Yao X. Development and validation of radiomics machine learning model based on contrast-enhanced computed tomography to predict axillary lymph node metastasis in breast cancer. Biomol Biomed. 2023;23(2):317–26.
Mezzadri G, Laloë T, Mathy F, Reynaud-Bouret P. Hold-out strategy for selecting learning models: application to categorization subjected to presentation orders. J Math Psychol. 2022;109: 102691.
Aerts HJWL, Velazquez ER, Leijenaar RTH, Parmar C, Grossmann P, Carvalho S, et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun. 2014;5(1):4006.
Rashid HU, Ibrikci T, Paydaş S, Binokay F, Çevik U. Analysis of breast cancer classification robustness with radiomics feature extraction and deep learning techniques. Expert Syst. 2022;39(8): e13018.
Reig B. Radiomics and deep learning methods in expanding the use of screening breast MRI. Eur Radiol. 2021;31(8):5863–5.
Daimiel Naranjo I, Gibbs P, Reiner JS, Lo Gullo R, Thakur SB, Jochelson MS, et al. Breast lesion classification with multiparametric breast MRI using radiomics and machine learning: a comparison with radiologists’ performance. Cancers (Basel). 2022;14(7):1743.
Parekh VS, Jacobs MA. Integrated radiomic framework for breast cancer and tumor biology using advanced machine learning and multiparametric MRI. NPJ Breast Cancer. 2017;3:43.
Tagliafico AS, Mariscotti G, Valdora F, Durando M, Nori J, La Forgia D, et al. A prospective comparative trial of adjunct screening with tomosynthesis or ultrasound in women with mammography-negative dense breasts (ASTOUND-2). Eur J Cancer. 2018;104:39–46.
Tsougos I, Vamvakas A, Kappas C, Fezoulidis I, Vassiou K. Application of radiomics and decision support systems for breast mr differential diagnosis. Comput Math Methods Med. 2018;2018(1):7417126.
Bickelhaupt S, Jaeger PF, Laun FB, Lederer W, Daniel H, Kuder TA, et al. Radiomics based on adapted diffusion kurtosis imaging helps to clarify most mammographic findings suspicious for cancer. Radiology. 2018;287(3):761–70.
Parekh VS, Jacobs MA. Multiparametric radiomics methods for breast cancer tissue characterization using radiological imaging. Breast Cancer Res Treat. 2020;180(2):407–21.
Daimiel Naranjo I, Gibbs P, Reiner JS, Lo Gullo R, Sooknanan C, Thakur SB, et al. Radiomics and machine learning with multiparametric breast MRI for improved diagnostic accuracy in breast cancer diagnosis. Diagnostics. 2021;11(6):919.
Pötsch N, Dietzel M, Kapetas P, Clauser P, Pinker K, Ellmann S, et al. An A.I. classifier derived from 4D radiomics of dynamic contrast-enhanced breast MRI data: potential to avoid unnecessary breast biopsies. Eur Radiol. 2021;31(8):5866–76.
Agner SC, Rosen MA, Englander S, Tomaszewski JE, Feldman MD, Zhang P, et al. Computerized image analysis for identifying triple-negative breast cancers and differentiating them from other molecular subtypes of breast cancer on dynamic contrast-enhanced MR images: a feasibility study. Radiology. 2014;272(1):91–9.
Yang Q, Li L, Zhang J, Shao G, Zheng B. A new quantitative image analysis method for improving breast cancer diagnosis using DCE-MRI examinations. Med Phys. 2015;42(1):103–9.
Wang J, Kato F, Oyama-Manabe N, Li R, Cui Y, Tha KK, et al. Identifying triple-negative breast cancer using background parenchymal enhancement heterogeneity on dynamic contrast-enhanced MRI: a pilot radiomics study. PLoS ONE. 2015;10(11): e0143308.
Truhn D, Schrading S, Haarburger C, Schneider H, Merhof D, Kuhl C. Radiomic versus convolutional neural networks analysis for classification of contrast-enhancing lesions at multiparametric breast MRI. Radiology. 2019;290(2):290–7.
Yin HL, Jiang Y, Xu Z, Jia HH, Lin GW. Combined diagnosis of multiparametric MRI-based deep learning models facilitates differentiating triple-negative breast cancer from fibroadenoma magnetic resonance BI-RADS 4 lesions. J Cancer Res Clin Oncol. 2023;149(6):2575–84.
Zhang J, Wu J, Zhou XS, Shi F, Shen D. Recent advancements in artificial intelligence for breast cancer: Image augmentation, segmentation, diagnosis, and prognosis approaches. Semin Cancer Biol. 2023;96:11–25.
Nicosia L, Gnocchi G, Gorini I, Venturini M, Fontana F, Pesapane F, et al. History of mammography: analysis of breast imaging diagnostic achievements over the last century. Healthcare. 2023;11(11):1596.
Hassan RM, Almalki YE, Basha MAA, Alduraibi SK, Aboualkheir M, Almushayti ZA, et al. The impact of adding digital breast tomosynthesis to BI-RADS categorization of mammographically equivocal breast lesions. Diagnostics (Basel). 2023;13(8):1423.
Grimm LJ, Avery CS, Hendrick E, Baker JA. Benefits and risks of mammography screening in women ages 40 to 49 years. J Prim Care Community Health. 2022. https://doi.org/10.1177/21501327211058322.
Domingo L, Hofvind S, Hubbard RA, Román M, Benkeser D, Sala M, et al. Cross-national comparison of screening mammography accuracy measures in U.S., Norway, and Spain. Eur Radiol. 2016;26(8):2520–8.
Institute of M, National Research Council Committee on New Approaches to Early D, Diagnosis of Breast C. The National Academies Collection: Reports funded by National Institutes of Health. In: Joy JE, Penhoet EE, Petitti DB, editors. Saving Women's Lives: Strategies for Improving Breast Cancer Detection and Diagnosis. Washington (DC): National Academies Press (US) Copyright © 2005, National Academy of Sciences.; 2005.
Chen H-l, Zhou J-q, Chen Q, Deng Y-c. Comparison of the sensitivity of mammography, ultrasound, magnetic resonance imaging and combinations of these imaging modalities for the detection of small (≤2 cm) breast cancer. Medicine. 2021;100(26): e26531.
Head JF, Lipari CA, Elliott RL, editors. Comparison of mammography and breast infrared imaging: sensitivity, specificity, false negatives, false positives, positive predictive value and negative predictive value. Proceedings of the First Joint BMES/EMBS Conference 1999 IEEE Engineering in Medicine and Biology 21st Annual Conference and the 1999 Annual Fall Meeting of the Biomedical Engineering Society (Cat N; 1999 13–16 Oct. 1999.
Mao N, Yin P, Wang Q, Liu M, Dong J, Zhang X, et al. Added value of radiomics on mammography for breast cancer diagnosis: a feasibility study. J Am Coll Radiol. 2019;16(4 Pt A):485–91.
Schaffter T, Buist DSM, Lee CI, Nikulin Y, Ribli D, Guan Y, et al. Evaluation of combined artificial intelligence and radiologist assessment to interpret screening mammograms. JAMA Netw Open. 2020;3(3): e200265.
Rodriguez-Ruiz A, Lång K, Gubern-Merida A, Broeders M, Gennaro G, Clauser P, et al. Stand-alone artificial intelligence for breast cancer detection in mammography: comparison with 101 radiologists. J Natl Cancer Inst. 2019;111(9):916–22.
Rodriguez-Ruiz A, Lång K, Gubern-Merida A, Teuwen J, Broeders M, Gennaro G, et al. Can we reduce the workload of mammographic screening by automatic identification of normal exams with artificial intelligence? A feasibility study. Eur Radiol. 2019;29(9):4825–32.
Hanis TM, Islam MA, Musa KI. Diagnostic accuracy of machine learning models on mammography in breast cancer classification: a meta-analysis. Diagnostics (Basel). 2022;12(7):1643.
Li Z, Ye J, Du H, Cao Y, Wang Y, Liu D, et al. Preoperative prediction power of radiomics for breast cancer: a systemic review and meta-analysis. Front Oncol. 2022. https://doi.org/10.3389/fonc.2022.837257.
Drukker K, Giger ML, Joe BN, Kerlikowske K, Greenwood H, Drukteinis JS, et al. Combined benefit of quantitative three-compartment breast image analysis and mammography radiomics in the classification of breast masses in a clinical data set. Radiology. 2019;290(3):621–8.
Karahaliou A, Skiadopoulos S, Boniatis I, Sakellaropoulos P, Likaki E, Panayiotakis G, et al. Texture analysis of tissue surrounding microcalcifications on mammograms for breast cancer diagnosis. Br J Radiol. 2014;80(956):648–56.
Wang G, Shi D, Guo Q, Zhang H, Wang S, Ren K. Radiomics based on digital mammography helps to identify mammographic masses suspicious for cancer. Front Oncol. 2022;12: 843436.
Mainprize JG, Alonzo-Proulx O, Alshafeiy TI, Patrie JT, Harvey JA, Yaffe MJ. Prediction of cancer masking in screening mammography using density and textural features. Acad Radiol. 2019;26(5):608–19.
Li H, Mendel KR, Lan L, Sheth D, Giger ML. Digital mammography in breast cancer: additive value of radiomics of breast parenchyma. Radiology. 2019;291(1):15–20.
Kooi T, Litjens G, van Ginneken B, Gubern-Mérida A, Sánchez CI, Mann R, et al. Large scale deep learning for computer aided detection of mammographic lesions. Med Image Anal. 2017;35:303–12.
Cozzi A, Magni V, Zanardo M, Schiaffino S, Sardanelli F. Contrast-enhanced mammography: a systematic review and meta-analysis of diagnostic performance. Radiology. 2022;302(3):568–81.
Ferranti FR, Vasselli F, Barba M, Sperati F, Terrenato I, Graziano F, et al. Diagnostic accuracy of contrast-enhanced, spectral mammography (CESM) and 3T magnetic resonance compared to full-field digital mammography plus ultrasound in breast lesions: results of a (pilot) open-label, single-centre prospective study. Cancers (Basel). 2022;14(5):1351.
Fallenberg EM, Dromain C, Diekmann F, Renz DM, Amer H, Ingold-Heppner B, et al. Contrast-enhanced spectral mammography: does mammography provide additional clinical benefits or can some radiation exposure be avoided? Breast Cancer Res Treat. 2014;146(2):371–81.
Fusco R, Piccirillo A, Sansone M, Granata V, Rubulotta MR, Petrosino T, et al. Radiomics and artificial intelligence analysis with textural metrics extracted by contrast-enhanced mammography in the breast lesions classification. Diagnostics (Basel). 2021;11(5):815.
Massafra R, Bove S, Lorusso V, Biafora A, Comes MC, Didonna V, et al. Radiomic feature reduction approach to predict breast cancer by contrast-enhanced spectral mammography images. Diagnostics. 2021;11(4):684.
Marino MA, Pinker K, Leithner D, Sung J, Avendano D, Morris EA, et al. Contrast-enhanced mammography and radiomics analysis for noninvasive breast cancer characterization: initial results. Mol Imag Biol. 2020;22(3):780–7.
Marino MA, Leithner D, Sung J, Avendano D, Morris EA, Pinker K, et al. Radiomics for tumor characterization in breast cancer patients: a feasibility study comparing contrast-enhanced mammography and magnetic resonance imaging. Diagnostics (Basel). 2020;10(7):492.
Lin F, Wang Z, Zhang K, Yang P, Ma H, Shi Y, et al. Contrast-enhanced spectral mammography-based radiomics nomogram for identifying benign and malignant breast lesions of sub-1 cm. Front Oncol. 2020;10: 573630.
Wang S, Sun Y, Li R, Mao N, Li Q, Jiang T, et al. Diagnostic performance of perilesional radiomics analysis of contrast-enhanced mammography for the differentiation of benign and malignant breast lesions. Eur Radiol. 2022;32(1):639–49.
Yu L, Wang Y, Xing D, Gong P, Chen Q, Lv Y. Background parenchymal enhancement on contrast-enhanced spectral mammography does not represent an influencing factor for breast cancer: a preliminary study. Medicine (Baltimore). 2020;99(52): e23857.
Sun Y, Wang S, Liu Z, You C, Li R, Mao N, et al. Identifying factors that may influence the classification performance of radiomics models using contrast-enhanced mammography (CEM) images. Cancer Imaging. 2022;22(1):22.
Fusco R, Di Bernardo E, Piccirillo A, Rubulotta MR, Petrosino T, Barretta ML, et al. Radiomic and artificial intelligence analysis with textural metrics extracted by contrast-enhanced mammography and dynamic contrast magnetic resonance imaging to detect breast malignant lesions. Curr Oncol. 2022;29(3):1947–66.
Bai J, Posner R, Wang T, Yang C, Nabavi S. Applying deep learning in digital breast tomosynthesis for automatic breast cancer detection: a review. Med Image Anal. 2021;71: 102049.
Niu S, Yu T, Cao Y, Dong Y, Luo Y, Jiang X. Digital breast tomosynthesis-based peritumoral radiomics approaches in the differentiation of benign and malignant breast lesions. Diagn Interv Radiol. 2022;28(3):217–25.
Sechopoulos I, Teuwen J, Mann R. Artificial intelligence for breast cancer detection in mammography and digital breast tomosynthesis: State of the art. Semin Cancer Biol. 2021;72:214–25.
Tagliafico AS, Valdora F, Mariscotti G, Durando M, Nori J, La Forgia D, et al. An exploratory radiomics analysis on digital breast tomosynthesis in women with mammographically negative dense breasts. Breast. 2018;40:92–6.
Sakai A, Onishi Y, Matsui M, Adachi H, Teramoto A, Saito K, et al. A method for the automated classification of benign and malignant masses on digital breast tomosynthesis images using machine learning and radiomic features. Radiol Phys Technol. 2020;13(1):27–36.
Zhang F, Wu S, Zhang C, Chen Q, Yang X, Jiang K, et al. Multi-domain features for reducing false positives in automated detection of clustered microcalcifications in digital breast tomosynthesis. Med Phys. 2019;46(3):1300–8.
Peng Y, Wu S, Yuan G, Wu Z, Du Q, Sun H, et al. A radiomics method to classify microcalcification clusters in digital breast tomosynthesis. Med Phys. 2020;47(8):3435–46.
Alì M, D’Amico NC, Interlenghi M, Maniglio M, Fazzini D, Schiaffino S, et al. A decision support system based on BI-RADS and radiomic classifiers to reduce false positive breast calcifications at digital breast tomosynthesis: a preliminary study. Appl Sci. 2021;11(6):2503.
Youk JH, Kwak JY, Lee E, Son EJ, Kim JA. Grayscale ultrasound radiomic features and shear-wave elastography radiomic features in benign and malignant breast masses. Ultraschall Med. 2020;41(4):390–6.
Michael E, Ma H, Li H, Qi S. An optimized framework for breast cancer classification using machine learning. Biomed Res Int. 2022;2022(1):8482022.
Zhang X, Li H, Wang C, Cheng W, Zhu Y, Li D, et al. Evaluating the accuracy of breast cancer and molecular subtype diagnosis by ultrasound image deep learning model. Front Oncol. 2021;11: 623506.
Magnuska ZA, Theek B, Darguzyte M, Palmowski M, Stickeler E, Schulz V, et al. Influence of the computer-aided decision support system design on ultrasound-based breast cancer classification. Cancers. 2022;14(2):277.
Mishra AK, Roy P, Bandyopadhyay S, Das SK. Breast ultrasound tumour classification: a machine learning—radiomics based approach. Expert Syst. 2021;38(7): e12713.
Sun Q, Lin X, Zhao Y, Li L, Yan K, Liang D, et al. Deep learning vs. radiomics for predicting axillary lymph node metastasis of breast cancer using ultrasound images: don’t forget the peritumoral region. Front Oncol. 2020. https://doi.org/10.3389/fonc.2020.00053.
Jabeen K, Khan MA, Alhaisoni M, Tariq U, Zhang Y-D, Hamza A, et al. Breast cancer classification from ultrasound images using probability-based optimal deep learning feature fusion. Sensors. 2022;22(3):807.
Interlenghi M, Salvatore C, Magni V, Caldara G, Schiavon E, Cozzi A, et al. A machine learning ensemble based on radiomics to predict BI-RADS category and reduce the biopsy rate of ultrasound-detected suspicious breast masses. Diagnostics (Basel). 2022;12(1):187.
Zhang X, Li H, Wang C, Cheng W, Zhu Y, Li D, et al. Evaluating the accuracy of breast cancer and molecular subtype diagnosis by ultrasound image deep learning model. Front Oncol. 2021. https://doi.org/10.3389/fonc.2021.623506.
Kim YS, Lee SE, Chang JM, Kim SY, Bae YK. Ultrasonographic morphological characteristics determined using a deep learning-based computer-aided diagnostic system of breast cancer. Medicine (Baltimore). 2022;101(3): e28621.
Fleury E, Marcomini K. Performance of machine learning software to classify breast lesions using BI-RADS radiomic features on ultrasound images. Eur Radiol Exper. 2019;3(1):34.
Lee SE, Han K, Kwak JY, Lee E, Kim E-K. Radiomics of US texture features in differential diagnosis between triple-negative breast cancer and fibroadenoma. Sci Rep. 2018;8(1):13546.
Moon WK, Huang YS, Lo CM, Huang CS, Bae MS, Kim WH, et al. Computer-aided diagnosis for distinguishing between triple-negative breast cancer and fibroadenomas based on ultrasound texture features. Med Phys. 2015;42(6):3024–35.
Xu M, Li F, Yu S, Zeng S, Weng G, Teng P, et al. Value of histogram of gray-scale ultrasound image in differential diagnosis of small triple negative breast invasive ductal carcinoma and fibroadenoma. Cancer Manag Res. 2022;14:1515–24.
Chen Q, Xia J, Zhang J. Identify the triple-negative and non-triple-negative breast cancer by using texture features of medicale ultrasonic image: a STROBE-compliant study. Medicine (Baltimore). 2021;100(22): e25878.
Du Y, Zha HL, Wang H, Liu XP, Pan JZ, Du LW, et al. Ultrasound-based radiomics nomogram for differentiation of triple-negative breast cancer from fibroadenoma. Br J Radiol. 2022;95(1133):20210598.
Sim Y, Lee SE, Kim EK, Kim S. A Radiomics approach for the classification of fibroepithelial lesions on breast ultrasonography. Ultrasound Med Biol. 2020;46(5):1133–41.
Luo WQ, Huang QX, Huang XW, Hu HT, Zeng FQ, Wang W. Predicting breast cancer in breast imaging reporting and data system (BI-RADS) ultrasound category 4 or 5 lesions: a nomogram combining radiomics and BI-RADS. Sci Rep. 2019;9(1):11921.
Moustafa AF, Cary TW, Sultan LR, Schultz SM, Conant EF, Venkatesh SS, et al. Color Doppler ultrasound improves machine learning diagnosis of breast cancer. Diagnostics (Basel). 2020;10(9):631.
Ternifi R, Wang Y, Gu J, Polley EC, Carter JM, Pruthi S, et al. Ultrasound high-definition microvasculature imaging with novel quantitative biomarkers improves breast cancer detection accuracy. Eur Radiol. 2022;32(11):7448–62.
Cui W, Peng Y, Yuan G, Cao W, Cao Y, Lu Z, et al. FMRNet: a fused network of multiple tumoral regions for breast tumor classification with ultrasound images. Med Phys. 2022;49(1):144–57.
Pang T, Wong JHD, Ng WL, Chan CS. Deep learning radiomics in breast cancer with different modalities: overview and future. Expert Syst Appl. 2020;158: 113501.
Fusco R, Granata V, Grazzini G, Pradella S, Borgheresi A, Bruno A, et al. Radiomics in medical imaging: pitfalls and challenges in clinical management. Jpn J Radiol. 2022;40(9):919–29.
Traverso A, Wee L, Dekker A, Gillies R. Repeatability and reproducibility of radiomic features: a systematic review. Int J Radiat Oncol Biol Phys. 2018;102(4):1143–58.
Shafiq-Ul-Hassan M, Zhang GG, Latifi K, Ullah G, Hunt DC, Balagurunathan Y, et al. Intrinsic dependencies of CT radiomic features on voxel size and number of gray levels. Med Phys. 2017;44(3):1050–62.
Frid-Adar M, Diamant I, Klang E, Amitai M, Goldberger J, Greenspan H. GAN-based synthetic medical image augmentation for increased CNN performance in liver lesion classification. Neurocomputing. 2018;321:321–31.
Granzier RWY, Verbakel NMH, Ibrahim A, van Timmeren JE, van Nijnatten TJA, Leijenaar RTH, et al. MRI-based radiomics in breast cancer: feature robustness with respect to inter-observer segmentation variability. Sci Rep. 2020;10(1):14163.
Nagalakshmi T. Breast cancer semantic segmentation for accurate breast cancer detection with an ensemble deep neural network. Neural Process Lett. 2022;54(6):5185–98.
Geras KJ, Mann RM, Moy L. Artificial intelligence for mammography and digital breast tomosynthesis: current concepts and future perspectives. Radiology. 2019;293(2):246–59.
Wu E, Wu K, Lotter W. Synthesizing lesions using contextual GANs improves breast cancer classification on mammograms. 2020. arXiv preprint arXiv:200600086.
Pang T, Wong JHD, Ng WL, Chan CS. Semi-supervised GAN-based radiomics model for data augmentation in breast ultrasound mass classification. Comput Methods Programs Biomed. 2021;203: 106018.
Zhang L, Xie D, Li Y, Camargo A, Song D, Lu T, et al. Improving sensitivity of arterial spin labeling perfusion MRI in Alzheimer’s disease using transfer learning of deep learning-based ASL denoising. J Magn Reson Imaging. 2022;55(6):1710–22.
Abdel Razek AAK, Alksas A, Shehata M, AbdelKhalek A, Abdel Baky K, El-Baz A, et al. Clinical applications of artificial intelligence and radiomics in neuro-oncology imaging. Insights Imaging. 2021;12(1):152.
Kalantari S, Soltani M, Maghbooli M, Khoshe Mehr FS, Kalantari Z, Borji S, et al. Alteraciones del flujo sanguíneo cerebral medidas con RM-ASL como predictor de demencia vascular en la enfermedad isquémica de pequeño vaso. Radiología. 2024. https://doi.org/10.1016/j.rx.2024.03.006.
Baeßler B, Weiss K, dos Pinto Santos D. Robustness and reproducibility of radiomics in magnetic resonance imaging: a phantom study. Invest Radiol. 2019;54(4):221–8.
Huang SY, Franc BL, Harnish RJ, Liu G, Mitra D, Copeland TP, et al. Exploration of PET and MRI radiomic features for decoding breast cancer phenotypes and prognosis. NPJ Breast Cancer. 2018;4:24.
Saha A, Yu X, Sahoo D, Mazurowski MA. Effects of MRI scanner parameters on breast cancer radiomics. Expert Syst Appl. 2017;87:384–91.
Whitney HM, Li H, Ji Y, Liu P, Giger ML. Harmonization of radiomic features of breast lesions across international DCE-MRI datasets. J Med Imaging (Bellingham). 2020;7(1): 012707.
Prinzi F, Orlando A, Gaglio S, Vitabile S. Breast cancer classification through multivariate radiomic time series analysis in DCE-MRI sequences. Expert Syst Appl. 2024;249: 123557.
Shinohara RT, Sweeney EM, Goldsmith J, Shiee N, Mateen FJ, Calabresi PA, et al. Statistical normalization techniques for magnetic resonance imaging. NeuroImage Clin. 2014;6:9–19.
Hatamikia S, George G, Schwarzhans F, Mahbod A, Woitek R. Breast MRI radiomics and machine learning-based predictions of response to neoadjuvant chemotherapy—how are they affected by variations in tumor delineation? Comput Struct Biotechnol J. 2024;23:52–63.
Nyul LG, Udupa JK, Xuan Z. New variants of a method of MRI scale standardization. IEEE Trans Med Imaging. 2000;19(2):143–50.
Dovrou A, Nikiforaki K, Zaridis D, Manikis GC, Mylona E, Tachos N, et al. A segmentation-based method improving the performance of N4 bias field correction on T2weighted MR imaging data of the prostate. Magn Reson Imaging. 2023;101:1–12.
Loizou CP, Pantziaris M, Seimenis I, Pattichis CS, editors. Brain MR image normalization in texture analysis of multiple sclerosis. 2009 9th International Conference on Information Technology and Applications in Biomedicine; 2009 4–7 Nov. 2009.
Carré A, Klausner G, Edjlali M, Lerousseau M, Briend-Diop J, Sun R, et al. Standardization of brain MR images across machines and protocols: bridging the gap for MRI-based radiomics. Sci Rep. 2020;10(1):12340.
Schwarzhans F, George G, Sanchez LE, Zaric O, Abraham JE, Woitek R, et al. Intensity Normalization Techniques and Their Effect on the Robustness and Predictive Power of Breast MRI Radiomics. 2024. arXiv preprint arXiv:240601736.
Cano A. A survey on graphic processing unit computing for large-scale data mining. Wiley Interdiscip Rev Data Mining and Knowl Discov. 2018;8: e1232.
Li H, Zhu Y, Burnside ES, Huang E, Drukker K, Hoadley KA, et al. Quantitative MRI radiomics in the prediction of molecular classifications of breast cancer subtypes in the TCGA/TCIA data set. NPJ Breast Cancer. 2016;2:16012.
Abdelrahman L, Al Ghamdi M, Collado-Mesa F, Abdel-Mottaleb M. Convolutional neural networks for breast cancer detection in mammography: a survey. Comput Biol Med. 2021;131: 104248.
Antropova N, Huynh BQ, Giger ML. A deep feature fusion methodology for breast cancer diagnosis demonstrated on three imaging modality datasets. Med Phys. 2017;44(10):5162–71.
Saber A, Sakr M, Abo-Seida O, Keshk A, Chen H. A novel deep-learning model for automatic detection and classification of breast cancer using the transfer-learning technique. IEEE Access. 2021. https://doi.org/10.1109/ACCESS.2021.3079204.
Vigil N, Barry M, Amini A, Akhloufi M, Maldague XPV, Ma L, et al. Dual-intended deep learning model for breast cancer diagnosis in ultrasound imaging. Cancers (Basel). 2022;14(11):2663.
Prusty S, Dash S, Patnaik S. A novel transfer learning technique for detecting breast cancer mammograms using VGG16 bottleneck feature. ECS Trans. 2021. https://doi.org/10.1149/10701.0733ecst.
Whitney HM, Li H, Ji Y, Liu P, Giger ML. Comparison of breast MRI tumor classification using human-engineered radiomics, transfer learning from deep convolutional neural networks, and fusion methods. Proc IEEE Inst Electr Electron Eng. 2020;108(1):163–77.
Chen LD, Li W, Xian MF, Zheng X, Lin Y, Liu BX, et al. Preoperative prediction of tumour deposits in rectal cancer by an artificial neural network-based US radiomics model. Eur Radiol. 2020;30(4):1969–79.
Ghoushchi S, Ranjbarzadeh R, Najafabadi SA, Osgooei E, Tirkolaee EB. An extended approach to the diagnosis of tumour location in breast cancer using deep learning. J Ambient Intell Human Comput. 2021. https://doi.org/10.1007/s12652-021-03613-y.
Rajendran R, Balasubramaniam S, Ravi V, Sennan S. Hybrid optimization algorithm based feature selection for mammogram images and detecting the breast mass using multilayer perceptron classifier. Comput Intell. 2022;38(4):1559–93.
Lin F, Wang Z, Zhang K, Yang P, Ma H, Shi Y, et al. Contrast-enhanced spectral mammography-based radiomics nomogram for identifying benign and malignant breast lesions of sub-1 cm. Front Oncol. 2020. https://doi.org/10.3389/fonc.2020.573630.
Liu H, Chen Y, Zhang Y, Wang L, Luo R, Wu H, et al. A deep learning model integrating mammography and clinical factors facilitates the malignancy prediction of BI-RADS 4 microcalcifications in breast cancer screening. Eur Radiol. 2021;31(8):5902–12.
Lo Gullo R, Daimiel I, Rossi Saccarelli C, Bitencourt A, Gibbs P, Fox MJ, et al. Improved characterization of sub-centimeter enhancing breast masses on MRI with radiomics and machine learning in BRCA mutation carriers. Eur Radiol. 2020;30(12):6721–31.
Whitney HM, Taylor NS, Drukker K, Edwards AV, Papaioannou J, Schacht D, et al. Additive benefit of radiomics over size alone in the distinction between benign lesions and luminal A cancers on a large clinical breast MRI dataset. Acad Radiol. 2019;26(2):202–9.
Fleury EFC, Marcomini K. Impact of radiomics on the breast ultrasound radiologist’s clinical practice: from lumpologist to data wrangler. Eur J Radiol. 2020;131: 109197.
Savini A, Feliciani G, Amadori M, Rivetti S, Cremonesi M, Cesarini F, et al. The role of acquisition angle in digital breast tomosynthesis: a texture analysis study. Appl Sci. 2020;10(17):6047.
Ma W, Zhao Y, Ji Y, Guo X, Jian X, Liu P, et al. Breast cancer molecular subtype prediction by mammographic radiomic features. Acad Radiol. 2019;26(2):196–201.
Han L, Zhu Y, Liu Z, Yu T, He C, Jiang W, et al. Radiomic nomogram for prediction of axillary lymph node metastasis in breast cancer. Eur Radiol. 2019;29(7):3820–9.
Fusco R, Granata V, Maio F, Sansone M, Petrillo A. Textural radiomic features and time-intensity curve data analysis by dynamic contrast-enhanced MRI for early prediction of breast cancer therapy response: preliminary data. Eur Radiol Exp. 2020;4(1):8.
Jiang X, Zou X, Sun J, Zheng A, Su C. A nomogram based on radiomics with mammography texture analysis for the prognostic prediction in patients with triple-negative breast cancer. Contrast Media Mol Imaging. 2020;2020(1):5418364.
Park H, Lim Y, Ko ES, Cho HH, Lee JE, Han BK, et al. Radiomics signature on magnetic resonance imaging: association with disease-free survival in patients with invasive breast cancer. Clin Cancer Res. 2018;24(19):4705–14.
Li H, Zhu Y, Burnside ES, Drukker K, Hoadley KA, Fan C, et al. MR imaging radiomics signatures for predicting the risk of breast cancer recurrence as given by research versions of mammaprint, oncotype DX, and PAM50 gene assays. Radiology. 2016;281(2):382–91.
Tran WT, Jerzak K, Lu FI, Klein J, Tabbarah S, Lagree A, et al. Personalized breast cancer treatments using artificial intelligence in radiomics and pathomics. J Med Imaging Radiat Sci. 2019;50(4 Suppl 2):S32-s41.
Ibrahim A, Primakov S, Beuque M, Woodruff HC, Halilaj I, Wu G, et al. Radiomics for precision medicine: current challenges, future prospects, and the proposal of a new framework. Methods. 2021;188:20–9.
Pesapane F, Suter MB, Rotili A, Penco S, Nigro O, Cremonesi M, et al. Will traditional biopsy be substituted by radiomics and liquid biopsy for breast cancer diagnosis and characterisation? Med Oncol. 2020;37(4):29.
Lyu S, Zhang M, Zhang B, Zhu J, Gao L, Qiu Y, et al. The value of radiomics model based on ultrasound image features in the differentiation between minimal breast cancer and small benign breast masses. J Clin Ultrasound. 2023;51(9):1536–43.
Yang X, Fan X, Lin S, Zhou Y, Liu H, Wang X, et al. Assessment of lymphovascular invasion in breast cancer using a combined MRI morphological features, radiomics, and deep learning approach based on dynamic contrast-enhanced MRI. J Magn Reson Imaging. 2024;59(6):2238–49.
Hao W, Gong J, Wang S, Zhu H, Zhao B, Peng W. Application of MRI radiomics-based machine learning model to improve contralateral BI-RADS 4 lesion assessment. Front Oncol. 2020;10: 531476.
Liu Y, Jia X, Zhao J, Peng Y, Yao X, Hu X, et al. A machine learning-based unenhanced radiomics approach to distinguishing between benign and malignant breast lesions using T2-weighted and diffusion-weighted MRI. J Magn Reson Imaging. 2024;60(2):600–12.
Qi X, Wang W, Pan S, Liu G, Xia L, Duan S, et al. Predictive value of triple negative breast cancer based on DCE-MRI multi-phase full-volume ROI clinical radiomics model. Acta Radiol. 2024;65(2):173–84.
Militello C, Rundo L, Dimarco M, Orlando A, Woitek R, D’Angelo I, et al. 3D DCE-MRI radiomic analysis for malignant lesion prediction in breast cancer patients. Acad Radiol. 2022;29(6):830–40.
Wang L, Ding Y, Yang W, Wang H, Shen J, Liu W, et al. A radiomics nomogram for distinguishing benign from malignant round-like breast tumors. Front Oncol. 2022;12: 677803.
Lei C, Wei W, Liu Z, Xiong Q, Yang C, Yang M, et al. Mammography-based radiomic analysis for predicting benign BI-RADS category 4 calcifications. Eur J Radiol. 2019;121: 108711.
Debelee T, Gebreselasie A, Schwenker F, Amirian M, Ashenafi DY. Classification of mammograms using texture and CNN based extracted features. J Biomim Biomater Biomed Eng. 2019;42:79–97.
Kayode AA, Akande NO, Adegun AA, Adebiyi MO. An automated mammogram classification system using modified support vector machine. Med Devices (Auckl). 2019;12:275–84.
Romeo V, Cuocolo R, Apolito R, Stanzione A, Ventimiglia A, Vitale A, et al. Clinical value of radiomics and machine learning in breast ultrasound: a multicenter study for differential diagnosis of benign and malignant lesions. Eur Radiol. 2021;31(12):9511–9.
Hassan RM, Almalki YE, Basha MAA, Alduraibi SK, Aboualkheir M, Almushayti ZA, et al. The impact of adding digital breast tomosynthesis to BI-RADS categorization of mammographically equivocal breast lesions. Diagnostics. 2023;13(8):1423.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted without commercial or financial relationships, which could be considered a potential conflict of interest.
Ethical approval
Since this article is a review manuscript and no specific patient or animal material has been used, the ethics approval is not applicable to this study.
Informed consent
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Elahi, R., Nazari, M. An updated overview of radiomics-based artificial intelligence (AI) methods in breast cancer screening and diagnosis. Radiol Phys Technol (2024). https://doi.org/10.1007/s12194-024-00842-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12194-024-00842-6