Abstract
Purpose
This study aims to develop radiomics models and a nomogram based on machine learning techniques, preoperative dual-energy computed tomography (DECT) images, clinical and pathological characteristics, to explore the tumor microenvironment (TME) of clear cell renal cell carcinoma (ccRCC).
Methods
We retrospectively recruited of 87 patients diagnosed with ccRCC through pathological confirmation from Center I (training set, n = 69; validation set, n = 18), and collected their DECT images and clinical information. Feature selection was conducted using variance threshold, SelectKBest, and the least absolute shrinkage and selection operator (LASSO). Radiomics models were then established using 14 classifiers to predict TME cells. Subsequently, we selected the most predictive radiomics features to calculate the radiomics score (Radscore). A combined model was constructed through multivariate logistic regression analysis combining the Radscore and relevant clinical characteristics, and presented in the form of a nomogram. Additionally, 17 patients were recruited from Center II as an external validation cohort for the nomogram. The performance of the models was assessed using methods such as the area under the receiver operating characteristic curve (AUC), calibration curve, and decision curve analysis (DCA).
Results
The validation set AUC values for the radiomics models assessing CD8+, CD163+, and αSMA+ cells were 0.875, 0.889, and 0.864, respectively. Additionally, the external validation cohort AUC value for the nomogram reaches 0.849 and shows good calibration.
Conclusion
Radiomics models could allow for non-invasive assessment of TME cells from DECT images in ccRCC patients, promising to enhance our understanding and management of the tumor.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Renal cell carcinoma (RCC), the most prevalent form of malignant kidney tumor, has exhibited a rising incidence rate. Current projections estimate that, by 2023, approximately 14,890 individuals in the United States will die from RCC [1]. A significant proportion of these fatalities are expected to due to clear cell renal cell carcinoma (ccRCC), the most common and aggressive subtype of RCC. With a 5 year survival rate ranging from 50 to 60% [2], ccRCC underscores the critical need for advancements in diagnosis and treatment to improve patient outcomes. In a pan-cancer comparison, RCC was identified as one of the most immune-infiltrated cancers [3], highlighting the profound impact the tumor microenvironment (TME) on the disease’s biological behavior.
Non-malignant cells within the TME contribute to cancer's oncogenic potential by stimulating uncontrolled cell proliferation affecting every stage of cancer development from initiation to metastasis [4], significantly influencing disease progression and treatment responses. In tumor immunology research, lymphocytes and myeloid cells are significant components of the tumor immune microenvironment (TIME). Among these, tumor-infiltrating lymphocytes (TILs) are one of the most extensively studied immune cell populations, CD8+ T lymphocytes are recognized as the primary effector lymphocytes in most tumors [5]. Tumor-associated macrophages (TAMs) are an important and extensively studied immune cell population within the TIME. TAMs often differentiate towards the immunosuppressive M2 phenotype, with CD163 serving as a definitive marker [6]. Furthermore, cancer associated fibroblasts (CAFs) are one of the most major components of the TME, they promote tumor proliferation and induce immune evasion in cancer cells, making them an attractive therapeutic target [7]. α-smooth muscle actin (αSMA), a marker for fibroblasts, is closely associated with the degree of fibrosis in the tumor’s surrounding tissue, with its high expression [8]. Therefore, this study selected CD8+ T lymphocytes, CD163+ TMAs and αSMA+ CAFs as representative immune and stromal cells to assess the TME in ccRCC.
Currently, the landscape of the TME evaluation is largely anchored to invasive practices such as biopsies or surgeries. The inherent heterogeneity of tumors, however, raise concerns about the ability of biopsy samples to fully capture the TME's complexity. Surgery, aside from being invasive, is subject to procedural delays and associated risks, posing significant challenges to timely and accurate cancer assessment. Therefore, there is an urgent need for a non-invasive preoperative assessment method. The development and introduction of various advanced imaging technologies and the increased utilization of radiology have significantly improved the detection rate of renal masses [9]. As a novel technology, dual-energy computed tomography (DECT) surpasses traditional CT single-parameter imaging by generating multiple-parameter images in a single scan, revealing more subtle differences in tissue composition [10]. However, identifying these complex data solely through visual inspection is a challenge. Therefore, the introduction of radiomics, an emerging technology, has become a central strategy to address this issue. Radiomics not only provides structural information about the tumor but also detects information on different cell types, vascular distribution, metabolic activity, and more [11], thus mapping the composition and behavior of the TME.
This study aims to construct radiomics models for quantitative analysis of CD8+ T lymphocytes, CD163+ TAMs, and αSMA+ CAFs in the TME of ccRCC and illustrate the potential of radiomics features as imaging biomarkers for TME cells. At the same time, this may seek to unveil the DECT assessment models’ additional clinical value.
Materials and methods
Cohort
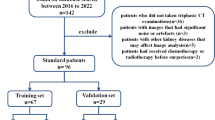
The Institutional Ethics Committee of our hospital approved this retrospective study and waived the requirement for patient consent. Data were obtained from two hospital districts. We collected patients with confirmed ccRCC between January 2019 and June 2023 based on inclusion and exclusion criteria. Patients must meet the following inclusion criteria: 1. Patients with pathologically confirmed ccRCC following surgery; 2. Complete clinical data available; 3. Contrast-enhanced DECT imaging was conducted within the 2 weeks preceding surgery, with no other treatments. The exclusion criteria are as follows: 1. Patients with other concurrent malignancies; 2. Patients with non-standard DECT scan patterns or insufficiently clear images, affecting the qualitative assessment of tumor; 3. Patients with predominantly cystic RCC; 4. Patients with severe underlying conditions affecting the heart, liver, lungs, or other organs. The criteria for measuring complete clinical data are provided in Appendix S1, and the classification thresholds for cystic-dominant RCC cases are provided in Appendix S2.
Ultimately, a total of 104 patients with ccRCC were included in the study. 87 patients from Centre I (January 2019 to January 2022) were randomized in an 8:2 ratio into a training cohort (n = 69) and a validation cohort (n = 18) in order to construct the radiomics models. For the construction of nomogram, patients from Centre I were used as the training set and patients (n = 17) from Centre II (February 2022 to June 2023) were treated as an independent external validation cohort.
DECT image data acquisition
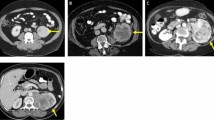
Following a standard scanning protocol (Table S1), all patients underwent DECT (Somatom Force; Siemens Healthcare, Forchheim, Germany) scans of the kidneys. The non-ionic contrast agent iopatol (a concentration of 300 mg/mL, a dose approximately 60–80 mL) was injected via the antecubital vein at a rate of 3.5 mL/s. Scanning was carried out using automatic exposure mode and DE mode, with the delay times for the corticomedullary phase (CMP) and renal parenchymal phase (NP) set at 30 and 80 s after contrast agent injection. The acquired images were post-processed using the workstation (syngo.via) to obtain the virtual monoenergetic 70 keV images, single-energy 100 keV and 150 keV images, virtual non-contrast (VNC) images, mixed energy imaging (MEI), and iodine overlay maps (IOM).
Pathological analysis
Tumor slides were stained with Hematoxylin and Eosin (HE), evaluated by a pathologist with 8 years of work experience, for WHO/ISUP grade, and T-stage of the tumor was performed according to the pathological Tumor-Node-Metastasis (pTNM) stage system [12]. Specific antibodies, including CD8 (ZSGB-Bio rabbit# ZA-0508), CD163 (ZSGB-Bio mouse# ZM-0428), and αSMA (ZSGB-Bio mouse# ZM-0003), were utilized for immunohistochemical staining. The above antibodies were supplied by Zhongshan Jinqiao Biotechnology Co., Ltd. China. Using an optical microscope (Olympus BX53, Japan), the slides were initially examined at 40 × magnification to identify regions with higher staining density, and then 5 micrographs were captured for each sample at 200 × magnification. Immunohistochemical stained areas were quantified using Image-Pro Plus 6.0 (Media Cybernetics, Silver Spring, MD, USA). The number of CD8+ T lymphocytes and CD163+ TAMs, as well as the percentage area of αSMA+ CAFs, were recorded in 5 micrographs for each case. The average of these values was calculated as the final immune quantitation for each case. Subsequently, the median final immune quantitation of all cases was calculated and used as a threshold for binary classification.
Radiomics features segmentation and extraction
All images and clinical data were uploaded to the Radcloud platform (Huiying Medical Technology Co., Ltd, https://www.huiyihuiying.com) for data processing, radiomics feature extraction and models construction. The regions of interest (ROI) were defined as the tumor regions. In preliminary experiments, we observed that the area under the Receiver Operating Characteristic Curve (AUC) values obtained from the model based on NP images were slightly lower than those obtained from the CMP images. Therefore, we chose CMP as the base phase for ROI outlining. A radiologist with 3 years of experience in diagnostic imaging, blinded to the patient’s clinical information, manually outlined the tumor boundaries and measured its maximum diameter. Subsequently, a senior radiologist with 16 years of diagnostic experience reviewed the results.
In order to reduce redundant features, several feature selection methods were used, including the variance threshold (0.8), SelectKBest and the least absolute shrinkage and selection operator (LASSO). The preprocessing steps, radiomics feature extraction and screening criteria are described in Appendix S3. After feature reduction and selection, radiomics features with high sensitivity and specificity for immunohistochemical quantitative parameters were ultimately obtained. Radscore (Radiomics score) is calculated as follows:
This formula includes the selected n radiomics features, where α represents the intercept, βi represents the value of the radiomics features, and Xi represents the corresponding coefficient.
Radiomics models construction
The Radcloud platform has 14 built-in supervised learning classifiers for machine learning. The classifiers were screened based on efficacy metrics such as Precision, Sensitivity, F1-Score, and Support. The relevant values are presented in Table S2. We used the AUC, 95% confidence intervals (CI), Sensitivity, and Specificity to evaluate the performance of radiomics models. Decision curve analysis (DCA) was obtained using the “rmda” package, which reflects the clinical utility of models by quantifying the trade-offs between the true positive rate and the false positive rate across various decision thresholds. The workflow schema is displayed in Fig. 1.
Nomogram construction
Univariate logistic regression analysis was performed on factors including clinical data, laboratory variables, pathological characteristics, and image feature to identify factors significantly influencing the probability of an event occurrence. Based on the Radscore and the aforementioned characteristic parameters, a multivariate logistic regression model was constructed for ccRCC to generate a nomogram. We considered WHO/ISUP grade as a potential prognostic indicator. The performance of the nomogram was assessed using AUC. The “rms” package was utilized to obtain calibration curves, evaluating the model’s net benefit performance across different probability thresholds. Its clinical utility was further assessed using DCA. The workflow schema is displayed in Fig. 1.
Statistical analysis
When comparing differences between different groups of continuous variables, the independent samples t-test method was chosen. For comparing differences between categorical variables, a chi-square test was employed. P-value < 0.05 is considered statistically significant. All data analyses were conducted using R Software (version 4.2.1, https://www.r-project.org/) and SPSS version 26 (IBM, Armonk, NY).
Results
Patient demographics
A total of 87 eligible ccRCC patients (60 male, 27 female) with a mean age of 57 years ± 10.94 were finally enrolled in Center I. 65 and 22 patients were in Grade I-II and Grade III-IV, respectively. Immunohistochemical analysis demonstrated the expression levels of CD8+ (Fig. 2A), CD163+ (Fig. 3A) and αSMA+ (Fig. 4A) cells in ccRCC TME. Using a threshold of 187 CD8+ cells per high-power field (HPF), 48 (55%) cases were classified as high-infiltration and 39 (45%) as nonhigh-infiltration. Using a threshold of 284 CD163+ cells per HPF, 46 (53%) cases were classified as high-infiltration and 41 (47%) as nonhigh-infiltration. Similarly, using a threshold of 0.014 αSMA+ cells area percentage per HPF, 44 (51%) high-infiltration cases and 43 (49%) nonhigh-infiltration. There were no significant differences in clinical, pathological features between the training and validation groups (Table 1).
Radiomics prediction model of CD8+ T lymphocytes in clear cell renal cell carcinoma. A Immunohistochemical expression. B Radiomics features obtained by variance thresholding, SelectKBest, and LASSO methods. C The boxplot. 0 indicating CD8-nonHigh and 1 indicating CD8-High. D Receiver operating characteristic curves based on the GaussianNB classifier, with the legend next to the curves showing the AUCs for the training and validation sets. E Decision curve analysis. The Y-axis represents the net benefit, calculated by adding true positives and subtracting false positives. The X-axis corresponds to the probability threshold. The higher the curve, the higher the clinical practicality of the model at this threshold
Radiomics prediction model of CD163+ TAMs in clear cell renal cell carcinoma. A Immunohistochemical expression. B Radiomics features. C The boxplot. 0 indicating CD163-nonHigh and 1 indicating CD163-High. D Receiver operating characteristic curves based on the LDA classifier, with the legend next to the curves showing the AUCs for the training and validation sets. E Decision curve analysis
Radiomics prediction model of SMA+ CAFs in clear cell renal cell carcinoma. A Immunohistochemical expression. B Radiomics features. C The boxplot. 0 indicating SMA-nonHigh and 1 indicating SMA-High. D Receiver operating characteristic curves based on the LR classifier, with the legend next to the curves showing the AUCs for the training and validation sets. E Decision curve analysis
Center II included 17 eligible ccRCC patients (11 male, 6 female) with a mean age of 56 years ± 10.31 years as an external validation cohort for the nomogram. 11 and 6 patients were in Grades I-II and III-IV, respectively. According to the immunohistochemical classification thresholds, there were 9 and 8 CD8+ high- and nonhigh-infiltrating cases, 11 and 6 CD163+ high- and nonhigh-infiltrating cases, and 11 and 6 αSMA+ high- and nonhigh-infiltrating cases, respectively.
Feature selection in radiomics models
A total of 10,128 quantitative features were extracted from DECT images. These features were categorized into three distinct classes. The first class encompassed 756 first-order statistical descriptors that provided a quantitative analysis of the CT image's voxel intensity distribution. The second class included 84 three-dimensional features representing the shape and size of ROI. The third category was the 3150 texture features derived from the Grey-Level Run-Length Matrix (GLRLM) and the Grey-Level Co-occurrence texture Matrix (GLCM). Through features selection and dimensionality reduction, the classification performance of GLRLM improved for CD8+ T lymphocytes and CD163+ TAMs, whereas the classification performance of αSMA+ CAFs was mainly influenced by first-order radiomics features. Figure 2, 3, 4B and Table S3 depict the radiomics features that were ultimately selected.
The performance of radiomics models
5 classifiers with stable differentiation performance in constructing the radiomics assessment models were selected, and their performance is shown in Table 2. The GaussianNB classifiers could effectively distinguish high and nonhigh CD8+ T lymphocytes infiltration, with the training and validation set AUC values were 0.930 (95% CI 0.843–1.000) and 0.875 (95% CI 0.699–1.000), respectively. The LDA classifiers could effectively distinguish high and nonhigh CD163+ TAMs infiltration, with the training and validation set AUC values were 0.978 (95% CI 0.924–1.000) and 0.889 (95% CI 0.677–1.000), respectively. The LR classifiers could effectively distinguish high and nonhigh αSMA+ CAFs infiltration, with the training and validation set AUC values were 0.947 (95% CI 0.865–1.000) and 0.864 (95% CI 0.721–1.000), respectively. Figure 2, 3, 4D illustrates the ROCs for different classifiers in the three radiomics models. DCA demonstrates that the evaluation models achieved improved clinical utility within a reasonable range of probability thresholds (Fig. 2, 3, 4E).
Construction and validation of the nomogram
As shown in Table 3, blood lipids and urea nitrogen were higher in high-grade patients (25.3%) compared with low-grade patients (P < 0.05). After multivariate logistic regression analysis, we created an integrated nomogram combining statistically significant clinical factors and Radscore to predict the pathological grade of ccRCC (Fig. 5A). The AUC for the nomogram was 0.820 (95% CI 0.709–0.930) (Fig. 5B) and the external validation cohort was 0.849 (95% CI 0.639–1.000) (Fig. 5E). Figure 5C, F show the calibration curves for the nomogram, indicating that the model has good consistency between the predictive model and the actual observed results. DCA showed that the nomogram had good clinical utility (Fig. 5D, G).
A The nomogram. In the visualization nomogram, first draw a vertical line based on the values of the most influential factors to determine the corresponding scores, and add these scores to obtain the total score. Then, draw another vertical line based on the total score value to determine the probability of ccRCC grade. B, E Receiver operating characteristic curves. C, F The calibration curves. The 45° straight line indicates the ideal prediction. The closer the two curves are, the higher the accuracy. D, G Decision curve analysis. (B, C, D Center I set; E, F, G Center II set.)
Discussion
The precise characterization of the TME components is crucial for the development and application of immunotherapeutic drugs, as well as for individualized cancer diagnosis, treatment, and prognosis prediction. The distribution and composition of immune cell infiltration within tumors often exhibit heterogeneity, with variations observed between different types of tumors. Even within different subtypes of the same cancer type, the composition of infiltrating immune cells in tumors may vary [13]. Although the abundant presence of intratumoral CD8+ T lymphocytes is usually regarded as a sign of anti-tumor immunity [14, 15], it has been shown that their increased infiltration may portend a poorer prognosis in ccRCC [16, 17], reflecting their complex role in the TME. This not only reveals their multiple functions in TME, but is also expected to inform future therapeutic strategies and drug development. TAMs are another key cellular component of the TIME and play an important role in anti-tumor therapy. It has been found that exhibit significant plasticity in both phenotype and function [18]. In the TIME, a significant portion of macrophages differentiates into M2 Phenotype TAMs, which possess immunosuppressive characteristics [19]. Additionally, tumor growth and infiltration are often associated with fibrosis in the surrounding tissue, characterized primarily by the activation and proliferation of CAFs. This, in turn, leads to the remodeling and fibrosis of the TME [20]. The components of the TME display dynamic heterogeneity influenced by a variety of internal and external regulatory factors, such as the dynamic development of tumors, immune evasion mechanisms, and therapeutic strategies. In this study, we used CD8+, CD163+, and αSMA+ cells as the entry point and found that radiomics features can be used as non-invasive imaging biomarkers for TME evaluation.
A growing number of studies have shown that radiomics analysis can successfully predict the TME and a range of predictive models have been developed. Wang et al. [21] utilized MRI radiomics analysis to assess TILs in locally advanced breast cancer (AUC = 0.891). Li et al. [22] and Chen et al. [23] constructed MRI-based radiomics models to predict TAMs infiltration in gliomas, with the latter reaching a maximum AUC value of 0.849. In related studies of ccRCC TME, CT radiomics models have been used to predict ccRCC CD8+ T cells [24], CD68+ TAMs cells [25] and immune checkpoint cells expression, including PD-L1 [24] and CTLA-4 [26]. All of the above radiomics assessment models achieved good performance, which is consistent with our findings. However, most relevant studies exploring TME tend to selectively focus on one to two types of TME cells or focus their studies only on immune cells. In addition, there have been few studies using radiomic models to evaluate the infiltration of αSMA + stromal cell tumors.
In the selection of tumor types, although the potential of radiomics in revealing the internal complexity of tumors has been recognized, current studies have mainly focused on breast cancers and gliomas, etc., and in the selection of imaging modalities, monoenergetic CT and MRI have been mainly applied, with relatively few relevant DECT radiomics studies targeting the TME features of ccRCC [27]. In this study, we developed radiomics evaluation models based on DECT imaging, which could well distinguish key immune and stromal cells in ccRCC TME, enabling the analysis of multiple cellular components in TME. The correlation between Radscore, TME cells, and tumor grade is important for the prognostic assessment of tumor patients and the formulation of personalized treatment plans, which of course needs to be confirmed by further experiments and more extensive cases.
Radiomics is a promising approach that allows for the quantitative analysis of the inherent heterogeneity within tumor lesions. High-order texture features have the capability to quantitatively assess tumor heterogeneity from various angles and scales [28]. Relevant research has found a negative correlation between heterogeneity indices (e.g., GLRLM non-uniformity) and several immune cell signal features in head and neck tumors [29]. This finding aligns with our study's results, where GLRLM features exhibited a negative correlation with quantitative features of CD8+ and CD163+ cells. The classification performance of αSMA+ CAFs is mainly influenced by first-order radiomics features, this suggests that the delineation of ROI may have a significant impact on the model performance for CAFs. In comparison to texture features, first-order radiomics features extracted from the entire tumor volume are robust and have demonstrated potential robustness for longitudinally monitoring tumor responses in a wide range of clinical settings, as indicated in a study [30]. This suggests that radiomics features can reveal features such as shape, margins, texture, and internal structure of ccRCC and translate the features into quantifiable data metrics, which are often closely related to pathological information of ccRCC.
The present study, however, has several limitations. First, as a retrospective study, selection bias in patient enrolment cannot be completely avoided. Future studies will be conducted prospectively to reduce bias in retrospective studies. Secondly, there are currently three types of DECT scanning technologies, and we only chose to use the technology provided by Siemens equipment. In the future, we plan to perform larger-scale validation on more types of equipment to improve the generalizability of the model. Third, given that RCC is a highly heterogeneous disease, large multi-center cohort studies are needed in the future to explore more biomarkers and cell types. Additionally, immunohistochemistry-positive cells were only quantified in the tumors without assessing staining intensity or location. Neglect of these factors may affect the comprehensiveness and accuracy of the study results.
In summary, this study developed ccRCC TME assessment models based on DECT radiomics, focusing on evaluating TME components such as CD8+, CD163+, and αSMA+ cells. The findings suggest that DECT radiomics features can serve as imaging biomarkers for quantitative analysis of ccRCC TME. This has the potential to provide more comprehensive and accurate tumor information to help enable more precise treatment strategies.
Data availability
The original contributions presented in the study are included in the article/Supplementary Material. You can always obtain further data from the corresponding author, [Ouyang A.], upon reasonable request.
References
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17–48. https://doi.org/10.3322/caac.21763.
Xing J, Liu Y, Wang Z, et al. Incremental value of radiomics with machine learning to the existing prognostic models for predicting outcome in renal cell carcinoma. Front Oncol. 2023;13:1036734. https://doi.org/10.3389/fonc.2023.1036734.
Vuong L, Kotecha RR, Voss MH, Hakimi AA. Tumor microenvironment dynamics in clear-cell renal cell carcinoma. Cancer Discov. 2019;9:1349–57. https://doi.org/10.1158/2159-8290.CD-19-0499.
Mier JW. The tumor microenvironment in renal cell cancer. Curr Opin Oncol. 2019;31:194–9. https://doi.org/10.1097/CCO.0000000000000512.
Sikic D, Weyerer V, Geppert CI, et al. Utility of stromal tumor infiltrating lymphocyte scoring (sTILs) for risk stratification of patients with muscle-invasive urothelial bladder cancer after radical cystectomy. Urol Oncol Semin Orig Investig. 2022;40:63.e19-63.e26. https://doi.org/10.1016/j.urolonc.2021.07.025.
Kinoshita J, Fushida S, Yamaguchi T, et al. Prognostic value of tumor-infiltrating CD163+macrophage in patients with metastatic gastric cancer undergoing multidisciplinary treatment. BMC Cancer. 2022;22:608. https://doi.org/10.1186/s12885-022-09713-y.
Chen Y, McAndrews KM, Kalluri R. Clinical and therapeutic relevance of cancer-associated fibroblasts. Nat Rev Clin Oncol. 2021;18:792–804. https://doi.org/10.1038/s41571-021-00546-5.
Muchlińska A, Nagel A, Popęda M, et al. Alpha-smooth muscle actin-positive cancer-associated fibroblasts secreting osteopontin promote growth of luminal breast cancer. Cell Mol Biol Lett. 2022;27:45. https://doi.org/10.1186/s11658-022-00351-7.
Krishna S, Leckie A, Kielar A, et al. Imaging of renal cancer. Semin Ultrasound CT MRI. 2020;41:152–69. https://doi.org/10.1053/j.sult.2019.12.004.
Mileto A, Marin D. Dual-energy computed tomography in genitourinary imaging. Radiol Clin North Am. 2017;55:373–91. https://doi.org/10.1016/j.rcl.2016.10.006.
Xu Y, Su G-H, Ma D, et al. Technological advances in cancer immunity: from immunogenomics to single-cell analysis and artificial intelligence. Signal Transduct Target Ther. 2021;6:312. https://doi.org/10.1038/s41392-021-00729-7.
Amin MB, Greene FL, Edge SB, et al. The 18th edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin. 2017;67:93–9. https://doi.org/10.3322/caac.21388.
Stanton SE, Adams S, Disis ML. Variation in the incidence and magnitude of tumor-infiltrating lymphocytes in breast cancer subtypes: a systematic review. JAMA Oncol. 2016;2:1354. https://doi.org/10.1001/jamaoncol.2016.1061.
Paijens ST, Vledder A, De Bruyn M, Nijman HW. Tumor-infiltrating lymphocytes in the immunotherapy era. Cell Mol Immunol. 2021;18:842–59. https://doi.org/10.1038/s41423-020-00565-9.
Fortis SP, Sofopoulos M, Goulielmaki M, et al. Association between intratumoral CD8+ T cells with FoxP3+ and CD163+ cells: a potential immune intrinsic negative feedback mechanism for acquired immune resistance. Cancers. 2022;14:6208. https://doi.org/10.3390/cancers14246208.
Murakami T, Tanaka N, Takamatsu K, et al. Multiplexed single-cell pathology reveals the association of CD8 T-cell heterogeneity with prognostic outcomes in renal cell carcinoma. Cancer Immunol Immunother. 2021;70:3001–13. https://doi.org/10.1007/s00262-021-03006-2.
Schraml P, Athelogou M, Hermanns T, et al. Specific immune cell and lymphatic vessel signatures identified by image analysis in renal cancer. Mod Pathol. 2019;32:1042–52. https://doi.org/10.1038/s41379-019-0214-z.
Murray PJ, Allen JE, Biswas SK, et al. Macrophage activation and polarization: nomenclature and experimental guidelines. Immunity. 2014;41:14–20. https://doi.org/10.1016/j.immuni.2014.06.008.
Pittet MJ, Michielin O, Migliorini D. Clinical relevance of tumour-associated macrophages. Nat Rev Clin Oncol. 2022;19:402–21. https://doi.org/10.1038/s41571-022-00620-6.
Chen X, Chen W, Zhao Y, et al. Interplay of Helicobacter pylori, fibroblasts, and cancer cells induces fibroblast activation and serpin E1 expression by cancer cells to promote gastric tumorigenesis. J Transl Med. 2022;20:322. https://doi.org/10.1186/s12967-022-03537-x.
Wang X, Xie T, Luo J, et al. Radiomics predicts the prognosis of patients with locally advanced breast cancer by reflecting the heterogeneity of tumor cells and the tumor microenvironment. Breast Cancer Res. 2022;24:20. https://doi.org/10.1186/s13058-022-01516-0.
Li G, Li L, Li Y, et al. An MRI radiomics approach to predict survival and tumour-infiltrating macrophages in gliomas. Brain. 2022;145:1151–61. https://doi.org/10.1093/brain/awab340.
Chen D, Zhang R, Huang X, et al. MRI-derived radiomics assessing tumor-infiltrating macrophages enable prediction of immune-phenotype, immunotherapy response and survival in glioma. Biomark Res. 2024;12:14. https://doi.org/10.1186/s40364-024-00560-6.
Varghese B, Cen S, Zahoor H, et al. Feasibility of using CT radiomic signatures for predicting CD8-T cell infiltration and PD-L1 expression in renal cell carcinoma. Eur J Radiol Open. 2022;9: 100440. https://doi.org/10.1016/j.ejro.2022.100440.
Shieh A, Cen SY, Varghese BA, et al. Radiomics correlation to CD68+ tumor-associated macrophages in clear cell renal cell carcinoma. Oncology. 2023. https://doi.org/10.1159/000534078.
He H, Jin Z, Dai J, et al. Computed tomography-based radiomics prediction of CTLA4 expression and prognosis in clear cell renal cell carcinoma. Cancer Med. 2022. https://doi.org/10.1002/cam4.5449.
Frankowska K, Zarobkiewicz M, Dąbrowska I, Bojarska-Junak A. Tumor infiltrating lymphocytes and radiological picture of the tumor. Med Oncol. 2023;40:176. https://doi.org/10.1007/s12032-023-02036-3.
Zhou Z, Qian X, Hu J, et al. CT-based peritumoral radiomics signatures for malignancy grading of clear cell renal cell carcinoma. Abdom Radiol. 2021;46:2690–8. https://doi.org/10.1007/s00261-020-02890-z.
Wang JH, Wahid KA, Van Dijk LV, et al. Radiomic biomarkers of tumor immune biology and immunotherapy response. Clin Transl Radiat Oncol. 2021;28:97–115. https://doi.org/10.1016/j.ctro.2021.03.006.
Lin Y-C, Lin C-H, Lu H-Y, et al. Deep learning for fully automated tumor segmentation and extraction of magnetic resonance radiomics features in cervical cancer. Eur Radiol. 2020;30:1297–305. https://doi.org/10.1007/s00330-019-06467-3.
Funding
The study received funding from the Jinan clinical medical science and technology innovation program (Grant: 202019036) and central guidance on local science and technology development fund project of Shandong Province (Grant: YDZX2021012).
Author information
Authors and Affiliations
Contributions
Ruobing Li: data curation, formal analysis, validation, visualization, writing—original draft, writing—review & editing. Xue Bing: data curation, formal analysis, writing—review & editing, supervision. Xinyou Su: formal analysis, validation, writing—review & editing, Supervision. Chunling Zhang: investigation, methodology, conceptualization, writing—review & editing. Haitao Sun: methodology, conceptualization, writing—review & editing, funding provided. Aimei Ouyang: methodology, supervision, project administration, writing—review & editing, funding provided. Co-author Zhengjun Dai are from a medical technical corporation provided technical support for model development, Writing—review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The data from this study was analyzed and controlled by authors who are from Central Hospital Affiliated to Shandong First Medical University. The authors declare that they have no conflict of interest.
Ethical approval
This study protocol was reviewed and approved by the Medical Ethics Committee of Central Hospital Affiliated to Shandong First Medical University, approval number 2023-012-01. The study has been granted an exemption from requiring written informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, R., Bing, X., Su, X. et al. The potential value of dual-energy CT radiomics in evaluating CD8+, CD163+ and αSMA+ cells in the tumor microenvironment of clear cell renal cell carcinoma. Clin Transl Oncol (2024). https://doi.org/10.1007/s12094-024-03637-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12094-024-03637-8