Abstract
Purpose
This study aimed to investigate the relationship between the interferon-gamma (IFN-γ) pathway in different tumor microenvironments (TME) and patients' prognosis, as well as the regulatory mechanisms of this pathway in tumor cells.
Methods
Using RNA-seq data from the TCGA database, we analyzed the predictive value of the IFN-γ pathway across various tumors. We employed a univariate Cox regression model to assess the prognostic significance of IFN-γ signaling in different tumor types. Additionally, we analyzed single-cell RNA sequencing (scRNA-seq) data from the Gene Expression Omnibus (GEO) database to examine the distribution characteristics of the IFN-γ pathway and explore its regulatory mechanisms, highlighting how IFN-γ influenced cellular interactions within the TME.
Results
Our analysis revealed a significant association between the IFN-γ pathway and adverse prognosis in pan-cancer tissues (P < 0.001). Interestingly, this correlation varied regarding positive and negative regulation across different tumor types. Through a detailed examination of scRNA-seq data, we found that the IFN-γ pathway exerted substantial regulatory effects on stromal and immune cells. In contrast, its expression and regulatory patterns in tumor cells exhibited diversity and heterogeneity. Further analysis indicated that the IFN-γ pathway not only enhanced the immunogenicity of tumor cells but also inhibited their proliferation. Cell–cell interaction analysis confirmed the pivotal role of the IFN-γ pathway within the overall regulatory network. Moreover, we identified HMGB2 (high mobility group box 2) in T cells as a potential key regulator of tumor cell proliferation.
Conclusions
The IFN-γ pathway exhibited a dual function by both suppressing tumor cell proliferation and enhancing their immunogenicity, positioning it as a pivotal target for refined cancer diagnosis and cancer strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The advent of tumor immunotherapy marks a pivotal evolution in oncology, introducing innovative strategies to harness the body's immune system in the fight against cancer [1]. Despite its promising therapeutic potential, the practical deployment of immunotherapy faces significant challenges. These include limitations in specificity and effectiveness, tumor immune evasion, undesirable side effects, drug resistance, and the absence of reliable predictive biomarkers [2, 3].
Recent advancements in understanding the tumor microenvironment (TME) and identifying immune-centric therapeutic targets have paved the way for innovative interventions. These include strategies to alter the tumor milieu with mesenchymal stem cells and their derivatives, as well as techniques to amplify immune responses by activating the cGAS/STING pathway [4,5,6]. The advent of combination therapies has broadened the spectrum of available treatments, leading to improved outcomes for a wider range of patients. However, the complexity of these therapies and the potential for adverse side effects require a measured and cautious approach [7]. Ongoing research and technological innovation are essential to refine these treatments, enhancing their effectiveness and making them more accessible to a wider patient population.
Interferon-gamma (IFN-γ) plays a crucial and multifaceted role within the TME, acting as a critical catalyst for the immune system's response to cancer through its complex regulatory functions [8]. It initiates the immune response by activating natural killer (NK) cells and enhancing the effectiveness of CD8+T cells, thereby significantly bolstering both humoral and cellular immunity to facilitate the direct elimination of tumor cells [9, 10]. Additionally, IFN-γ finely the activity of immune cells, notably by inhibiting the functions of regulatory T (Treg) cells [11] and suppressive macrophages [12]. This action helps dismantle the immunosuppressive environment, fostering conditions that boost immune system vigor and enhance tumor immune surveillance and attack.
Moreover, IFN-γ induces tumor cells to upregulate the expression of major histocompatibility complex (MHC)-I molecules, thereby enhancing their antigen presentation capabilities and rendering them more detectable and susceptible to the immune system's attack [13, 14]. Ultimately, IFN-γ transforms the TME from a state of immunosuppression to one of immunostimulation, curbing tumor growth and spread while promoting the involvement of additional immune cells. This establishes a synergistic immune network that effectively combats the tumor. However, research also indicates that IFN-γ may contribute to tumor cells' immune evasion tactics [15, 16]. Thus, the intricate regulatory role of IFN-γ in cancer immunotherapy underscores its significance as a therapeutic target. A deep understanding of its mechanisms holds the potential for developing novel and more effective treatment approaches.
This investigation aimed to uncover the intricate roles of IFN-γ signaling pathways within the TME, evaluating their implications for patients' outcomes and elucidating their regulatory functions within tumor cells. By leveraging RNA-seq and single-cell RNA sequencing (scRNA-seq) data from public databases, our study highlighted the critical role of IFN-γ within the TME, emphasizing its pivotal importance in modulating the microenvironmental dynamics during tumor immunotherapy processes. This research significantly advanced our understanding of the complex interactions within the TME, providing essential insights for the development of novel therapeutic strategies.
Materials and methods
Public data sources
The Cancer Genome Atlas (TCGA) pan-cancer RNA-seq data were obtained from the UCSC Xena website (https://xena.ucsc.edu/), encompassing 11,768 samples with pan-cancer gene expression matrices and survival information. During data curation, samples adjacent to tumors and those lacking survival information were excluded. This refinement led to the selection of 11,506 samples, which were used for further survival analyses. RNA-seq data for mRNA expressions in pre-treatment melanomas undergoing anti-PD-1 checkpoint inhibition therapy were acquired from the GEO database (GEO: GSE78220), which contained 14 responsive and 14 non-responsive melanomas.
scRNA-seq data for patients with low-grade glioma (LGG) and melanoma were obtained from the Gene Expression Omnibus (GEO) database (GEO: GSE182109 and GSE115978) [17, 18]. The dataset for patients with LGG included four untreated tumor samples, while the melanoma dataset contained 33 tumor samples, with 15 of these being from melanomas unresponsive to treatment.
RNA-seq data analysis
Gene sets corresponding to the IFN-γ signaling pathway and cell cycle pathways were acquired from the MSigDB database (https://www.gsea-msigdb.org/gsea/msigdb). Enrichment scores of pathways were computed using the gene set variation analysis (GSVA) method within the GSVA package. According to the median value, the proportions of scores were divided into two groups: “low” and “high”. Subsequently, Kaplan–Meier survival curves and univariate Cox regression analyses were performed. Cox regression analysis was conducted using the coxph function from the R package survival. The results were visualized using the R package forestplot.
scRNA-seq data analysis
Utilizing the officially recommended analysis workflow by Seurat, we conducted a standardized analysis on the downloaded scRNA-seq data. Each cell cluster was meticulously annotated in accordance with cell population marker genes identified in the literature. The IFN-γ signaling pathway's score was determined by the AddModuleScore function, while the scores for the S phase and G2M phase were calculated with the CellCycleScoring feature, leveraging Seurat's integrated gene sets. Differential expressed genes (DEGs) analysis was performed using the FindMarkers function. Subsequently, the fgsea package was employed for the enrichment analysis and visualization of selected DEGs.
Immune infiltration analysis
The FindAllMarkers function was applied to identify signature genes for each cluster within scRNA-seq datasets. Subsequently, scores for each cluster in melanoma and LGG RNA-seq datasets are scored using the gsva function within the GSVA package. The Pearson correlation coefficient among clusters and IFN-γ signaling pathway's score was computed using the cor function, and the results are visualized by pheatmap package.
Cell–cell interactions analysis
The scRNA-seq data from and melanoma samples were employed to infer cell–cell interactions. To identify potential ligand/receptor interactions, we utilized the NicheNetr package with default parameters. The identified interactions identified were subsequently visualized using the p_ligand_target_network function from NicheNetr.
Statistical analysis
All statistical analyses were performed using R-4.2.3 software, with the relevant R packages sourced from the official websites of CRAN and Bioconductor. Specific details about the significance testing are provided in the corresponding figure legends.
Results
Survival analysis of the IFN-γ pathway in pan-cancer
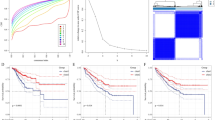
Utilizing the TCGA database, we embarked on an extensive survival analysis of the IFN-γ pathway across a broad spectrum of cancers. Our results strikingly revealed that elevated IFN-γ pathway score were markedly associated with poorer overall survival rates in patients (P < 0.001) (Fig. 1A). This underscored the pivotal role of the IFN-γ pathway in contributing to adverse outcomes across multiple cancer types. Delving deeper, univariate Cox regression model analysis across 33 cancer types demonstrated a nuanced picture. In four distinct cancers, adrenocortical carcinoma (ACC: P = 0.018), ovarian cancer (OV: P = 0.022), sarcoma (SARC: P = 0.044), and melanoma (SKCM: P < 0.001), higher IFN-γ pathway scores were associated with a favorable prognosis. Conversely, in three other cancers, acute myeloid leukemia (LAML: P = 0.012), LGG (P < 0.001), and uveal melanoma (UVM: P = 0.003), higher scores were linked to a poorer prognosis (Fig. 1B). Further analysis illuminated the diverse impact of the IFN-γ pathway on patient survival across different cancer types. Notably, melanoma patients with higher IFN-γ scores showed significantly improved outcomes (P < 0.001) (Fig. 1C), whereas LGG patients exhibited a significant correlation with adverse outcomes (P < 0.001) (due to insufficient numbers, UVM patients were not included in further analysis) (Fig. 1D). These insights underscore the critical role of the IFN-γ pathway in pan-cancer survival, highlighting its significant biological influence and its variable impact across different cancer types.
Survival analysis based on IFN-γ pathway's score based on the TCGA database. A Kaplan–Meier analysis of the IFN-γ pathway score among pan-cancer, based on the TCGA database. B Univariate Cox's regression model analysis of the IFN-γ pathway score across pan-cancer. C Kaplan–Meier analysis of the IFN-γ pathway score in cancer types with better prognosis. D Kaplan–Meier analysis of the IFN-γ pathway score in cancer types with worse prognosis. The high and low expression groups are defined by the median value of the IFN-γ pathway's score, the Kaplan–Meier analysis was used to calculate the P value, and univariate Cox's regression model analysis was used to calculate the hazard ratio (HR) and 95% confidence interval (95% CI)
scRNA-seq data analysis of IFN-γ pathway in melanoma and LGG
To further elucidate the differential activity of the IFN-γ pathway across various tumors, we extensively analyzed scRNA-seq data from the GEO database. This approach allowed us to investigate the regulatory effects of the IFN-γ pathway on diverse cellular components within TME. Our analysis of scRNA-seq data from LGG patients revealed that the IFN-γ pathway exhibited lower activity in glioma cells and oligodendrocytes compared to stromal and immune cells (Fig. 2A–C, Supplemental Fig. 1A). Extending this investigation to melanoma patients, we observed a similar pattern: the activity of IFN-γ pathway in melanoma cells was diminished relative to that in stromal and immune cells (Fig. 2D–F, Supplemental Fig. 1B). Based on the clustering results of scRNA-seq data, we quantified the infiltration scores for each cell type within the TCGA database and conducted a correlation analysis among various cells and IFN-γ pathway. Our findings revealed a significant negative correlation between glioma tumor cells and the IFN-γ pathway (Fig. 2G). Conversely, melanoma tumor cells exhibited no notable correlation with the IFN-γ pathway (Fig. 2H).
scRNA-seq data analysis reveals diverse expression patterns of the IFN-γ pathway's score among the TME. A–C LGG. D–F melanoma. A and C UMAP analysis showing the clustering results of different cell populations. B and E UMAP analysis showing the s IFN-γ pathway scores in different cell populations. C and F Violin plots show the IFN-γ pathway scores in different cell populations. G and H Heatmaps show scores of cell populations and IFN-γ pathway on the TCGA database, calculated by clustering results in scRNA-seq analysis. G Represents low-grade glioma and H represents melanoma
While the IFN-γ pathway showed lower expression levels in melanoma tumor cells compared to other immune and stromal cells, it nonetheless displayed an upward trend when contrasted with glioma tumor cells. This finding indicated its dynamic regulatory role within the TME.
The role of the IFN-γ pathway in melanoma tumor cells
Through sophisticated differential analysis, we successfully isolated genes exhibiting distinct expression profiles between melanoma tumor cells characterized by high versus low IFN-γ pathway activity, revealing significant disparities between these cohorts (Fig. 3A). Subsequent gene set enrichment analysis (GSEA) of these DEGs provided a detailed mapping of the regulatory landscape of the IFN-γ pathway within melanoma cells. The GSEA results revealed that melanoma tumor cells with high IFN-γ pathway activity showed a significant upregulation in key immune-related pathways, including those related to IFN, tumor necrosis factor-α (TNF-α), interleukin-6 (IL-6), and the complement system. This highlighted the crucial role of the IFN-γ pathway in immune regulation and the activation of numerous immune response genes. In contrast, melanoma tumor cells with low IFN-γ pathway activity demonstrated notable enhancement in pathways linked to cell cycle progression, such as MYC, E2F, and G2M. This finding suggested that the IFN-γ pathway might play a role in inhibiting melanoma cell proliferation (Fig. 3B).
The impact of the IFN-γ pathway on melanoma tumor cells. A Heatmap showing the differential expression of the IFN-γ pathway score in melanoma tumor cells. B Enrichment analysis results of the IFN-γ pathway in melanoma cells with high and low expression. C Violin plots showing the scores of cell cycle S phase and G2M phase in high and low expression of the IFN-γ pathway. D Heatmap showing the ratio of o/e values for cell cycle phases G1, S, and G2M under high and low expression conditions of the IFN-γ pathway.The differences in scores for the S phase and G2M phase of the cell cycle between high and low expression of the IFN-γ pathway were calculated using the Wilcoxon test, ***P < 0.001
Upon further investigation, we evaluated the cell cycle dynamics by calculating cell cycle scores in melanoma tumor cells. In groups with elevated IFN-γ pathway activity, there was a notable rise in the scores for both the S phase and G2M phase (P < 0.001). This suggested a possible link between IFN-γ pathway activation and precise control of cell cycle mechanisms in melanoma cells (Fig. 3C). By examining the observed versus expected (o/e) ratio for cell cycle scores, our analysis shed light on nuanced variations. Specifically, the dominance of the S phase and G2M phase was significantly higher in melanoma cells with reduced IFN-γ pathway activity, while a tendency toward an increased G2M phase was evident in cells with heightened IFN-γ pathway activity (Fig. 3D). This detailed analysis further clarified the varied impact of the IFN-γ pathway on cell cycle regulation within melanoma tumor cells, emphasizing its intricate and multifaceted role in cancer biology.
Survival analysis of cell cycle-related pathways in pan-cancer
Our examination of melanoma tumor cells revealed a profound association between cell cycle-related signaling pathways and the degree of tumor malignancy. To deepen our understanding of their potential implications for patient outcomes, we meticulously explored four pathways related to the cell cycle: MYC targets V1 and V2, E2F, and G2M. By calculating the scores of these pathways and employing the median score as a threshold, we categorized patients into groups with high or low expression levels and generated Kaplan–Meier survival curves. The survival analysis results revealed a notable negative correlation between pan-cancer cell cycle-related pathway scores and patient prognosis (Fig. 4A–D). This pattern was consistently evident in both LGG (Fig. 4E–H) and melanomas (Fig. 4I–L), strongly indicating the widespread importance of cell cycle-related pathways in determining cancer survival rates. These findings highlighted the critical role of cell cycle-related pathways in determining patient outcomes across various cancer types, emphasizing the need for further research into their potential as therapeutic targets.
Survival analysis based on cell cycle-related pathway scores from the TCGA database. A–D Pan-cancer. E–H LGG. I–L Melanoma. A, E and I E2F pathway. B, F, and J G2M pathway. C, G, and K MYC targets V1 pathway. D, H, and L MYC targets V2 pathway. The high expression group and the low expression group are defined by the median value of percentage, using the Kaplan–Meier test to calculate the P value for the difference between the high and low expression groups
Differences in the IFN-γ pathway in melanoma tumor cells of non-responding patients
IFN-γ stimulation resulted in increased expression of major histocompatibility complex-I (MHC-I) molecules in tumor cells, enhancing the processing and presentation of internal antigens and making tumor cells more susceptible to recognition by CD8+T cells. Simultaneously, IFN-γ could also trigger the expression of MHC-II molecules, augmenting the recognition of tumor cells by CD4+T cells and further strengthening immunogenicity. By modulating the expression of MHC molecules, IFN-γ could influence the interactions between tumor and immune cells, thereby enhancing the efficacy of immunotherapy. Given the pivotal role of the IFN-γ pathway in cancer immunotherapy, we examined its specific mechanisms within melanoma. Analysis of scRNA-seq data from melanomas resistant to immunotherapy revealed that the IFN-γ pathway scores were either elevated or remained constant in nearly all cells after treatment (Fig. 5A and B). However, a decrease in IFN-γ pathway scores in melanoma tumor cells following treatment indicated a critical role of the IFN-γ pathway within these cells. By examining DEG expression pre- and post-treatment, we identified a widespread downregulation of genes related to the IFN-γ pathway, including numerous HLA molecules associated with antigen presentation (Fig. 5C). Enrichment analysis further revealed a significant upregulation of cell cycle-related pathways after treatment (Fig. 5D). These findings supported the idea that IFN-γ not only boosted the immunogenicity of tumor cells but also enhanced the efficacy of immunotherapy by inhibiting cell proliferation capabilities.
The impact of the IFN-γ pathway on melanoma tumor cells in non-responsive patients. A UMAP showing the IFN-γ pathway scores in melanoma patients with immunotherapy. B Violin plot showing the IFN-γ pathway scores in melanoma patients with immunotherapy. C UMAP showing the differential genes related to the IFN-γ pathway's score in melanoma tumor cells before and after immunotherapy. D UMAP showing the enriched signaling pathways in melanoma tumor cells before and after immunotherapy. E Box plots showing scores of IFN-γ pathway and cell cycle-related pathway, based on RNA-seq from melanoma patients treated with immunotherapy. The differences in IFN-γ pathway's scores among cell populations before and after treatment were calculated using the Wilcoxon test, *P < 0.05 and ***P < 0.001
To further elucidate the relationship between IFN-γ and the immune response in melanoma patients, we analyzed RNA-seq data from melanoma patients categorized by their response status in the GEO database. We calculated scores for both IFN-γ and cell cycle-related pathways. Our analysis revealed that the IFN-γ pathway score was higher in complete responsive patients, while it was lower in those with partial or progressive disease. Conversely, scores for cell cycle-related pathways were lower in complete responsive patients and higher in those with partial or progressive disease (Fig. 5E). This discovery provided a new perspective on understanding the mechanistic role of the IFN-γ pathway in cancer treatment, highlighting its potential impact on patient outcomes and its significance in modulating the immune response and cell cycle dynamics.
Cell–cell communication analysis between immune cells and melanoma tumor cells on the IFN-γ pathway
To examine the impact of immune cells within the TME on the IFN-γ pathway, we employed the NicheNetr package for our analysis. Initially, we identified highly expressed ligands in T cells and myeloid cells, then constructed a ligand–target gene regulatory network to investigate the potential effects of these ligands on the IFN-γ pathway in tumor cells (Fig. 6A and C). The analysis of T cells revealed that the regulatory modes of many ligands aligned with the IFN-γ pathway, including TNF, Fas ligand (FASLG), CD40 ligand (CD40LG). These ligands not only increased the expression of genes related to MHC molecules in tumor cells but also enhanced the expression of the cell cycle inhibitory gene CDKN1A (p21). Conversely, the target genes of HMGB2 exhibited a different pattern, involving the expression of cell cycle-related genes such as CDK1 (Fig. 6A and B). Analysis of myeloid cells showed that the regulatory modes of most ligands were consistent, promoting the immunogenicity of tumor cells and inhibiting their proliferative capabilities (Fig. 6C and D). This finding suggested that immune cells within the tumor microenvironment positively influence the IFN-γ pathway in tumor cells by modulating the ligand–target gene network. This study provides valuable insights into understanding the complex regulatory mechanisms within the TME, highlighting the pivotal role of immune cell–tumor cell interactions in modulating the IFN-γ pathway.
Cell–cell communication between immune cells and melanoma cells on the IFN-γ pathway. A, B Showing the effect of T cells on the IFN-γ pathway in tumor cells, and C, D showing the effect of myeloid cells on the IFN-γ pathway in tumor cells. A Dot plot showing potential T cell ligands driving the IFN-γ pathway in melanoma tumor cells. B Heatmap showing potential T cell ligands driving the IFN-γ pathway in melanoma tumor cells. C Dot plot showing potential myeloid cell ligands driving the IFN-γ pathway in melanoma tumor cells. D Heatmap showing potential myeloid cell ligands driving the IFN-γ pathway in melanoma tumor cells
Discussion
When leveraging the IFN-γ pathway for tumor treatment, a pronounced anti-proliferative effect is observed, demonstrating a complex modulation of tumor cell behavior through various mechanisms. Initially, IFN-γ markedly restricts the proliferation rate of tumor cells by altering cell cycle dynamics, particularly slowing the G1/S phase transition and reducing DNA synthesis [19, 20]. Moreover, IFN-γ orchestrates the suppression of tumor cell growth by finely regulating gene expression, including the downregulation of genes associated with proliferation and the activation of oncogene suppressors [21, 22]. Additionally, IFN-γ induces apoptosis in tumor cells, further inhibiting their proliferation [23]. Overall, the robust anti-proliferative efficacy of IFN-γ underscores its potent therapeutic potential in tumor treatment, highlighting its role in the intricate modulation of cellular proliferation processes.
IFN-γ plays a pivotal role in the modulation of MHC molecule expression. It induces the upregulation of MHC class I molecules on the surface of tumor cells, enhancing the processing and presentation of endogenous antigens, thereby rendering tumor cells more readily identifiable by CD8+T cells [13]. Additionally, IFN-γ may trigger the expression of MHC class II molecules, enhancing tumor cell susceptibility to CD4+T cells [24,25,26]. This regulatory effect not only amplifies the immunogenicity of tumor cells but also facilitates the immune system's targeted response against the tumor. Through the strategic modulation of MHC molecule expression, IFN-γ intricately influences the interactions between tumor cells and immune cells, thereby enhancing the impact of immunotherapeutic strategies. The regulation of MHC molecule expression by IFN-γ is critical for the recognition and clearance of tumor cells by the immune system [26, 27]. This improvement in antigen presentation efficacy on the tumor cell surface significantly strengthens the immune system's ability to detect and eliminate tumor cells, underscoring the indispensable role of IFN-γ in immune regulation and the orchestration of anti-tumor immune responses.
In certain scenarios, IFN-γ exhibits a tendency to enhance the immunosuppressive capabilities of tumor cells [16, 28]. This can be evidenced by its ability to suppress the expression of MHC class I molecules, impairing the immune system's recognition of tumor cells. Additionally, IFN-γ can induce the production of immunosuppressive agents such as PD-L1, leading to diminished activity of immune cells [29, 30]. Furthermore, IFN-γ may influence the functionality of immune cells, thereby reducing their impact on tumor cells [31]. The intricate and multifaceted role of IFN-γ involvement in the regulation of tumor immunity is influenced by the influence of various factors, including the type of tumor, its microenvironment, and the prevailing immune state.
The IFN-γ signaling pathway has emerged as a promising therapeutic target, poised to play a pivotal role within the TME by intricately modulating immune responses and cell cycle regulation. This research not only establishes a novel theoretical framework but also significantly bolsters the foundation for the personalized design of future clinical interventions. Delving into the multifaceted effects of the IFN-γ pathway holds the potential to drive groundbreaking advancements in cancer therapy, heralding a new era of innovation and development.
Data availability
All sequencing data in this study were obtained from publicly available articles, and no additional data were generated during the course of this research.
References
Zhang Y, Zhang Z. The history and advances in cancer immunotherapy: understanding the characteristics of tumor-infiltrating immune cells and their therapeutic implications. Cell Mol Immunol. 2020;17(8):807–21.
Lei Y, Li X, Huang Q, Zheng X, Liu M. Progress and challenges of predictive biomarkers for immune checkpoint blockade. Front Oncol. 2021;11: 617335.
Lorentzen CL, Haanen JB, Met O, Svane IM. Clinical advances and ongoing trials on mRNA vaccines for cancer treatment. Lancet Oncol. 2022;23(10):e450–8.
Chehelgerdi M, Chehelgerdi M. The use of RNA-based treatments in the field of cancer immunotherapy. Mol Cancer. 2023;22(1):106.
Han Y, Yang J, Fang J, Zhou Y, Candi E, Wang J, et al. The secretion profile of mesenchymal stem cells and potential applications in treating human diseases. Signal Transduct Target Ther. 2022;7(1):92.
Ou L, Zhang A, Cheng Y, Chen Y. The cGAS-STING pathway: a promising immunotherapy target. Front Immunol. 2021;12: 795048.
Shyr ZA, Cheng YS, Lo DC, Zheng W. Drug combination therapy for emerging viral diseases. Drug Discov Today. 2021;26(10):2367–76.
Ivashkiv LB. IFNgamma: signalling, epigenetics and roles in immunity, metabolism, disease and cancer immunotherapy. Nat Rev Immunol. 2018;18(9):545–58.
Sakatani T, Kita Y, Fujimoto M, Sano T, Hamada A, Nakamura K, et al. IFN-gamma expression in the tumor microenvironment and CD8-positive tumor-infiltrating lymphocytes as prognostic markers in urothelial cancer patients receiving pembrolizumab. Cancers (Basel). 2022;14(2):263.
Kim H, Abbasi A, Sharrock J, Santosa EK, Sheppard S, Lau CM, et al. Cutting edge: STAT4 promotes Bhlhe40 induction to drive protective IFN-gamma from NK cells during viral infection. J Immunol. 2023;211(10):1469–74.
Overacre-Delgoffe AE, Chikina M, Dadey RE, Yano H, Brunazzi EA, Shayan G, et al. Interferon-gamma drives T(reg) fragility to promote anti-tumor immunity. Cell. 2017;169(6):1130–4111.
Zhang M, Huang L, Ding G, Huang H, Cao G, Sun X, et al. Interferon gamma inhibits CXCL8-CXCR2 axis mediated tumor-associated macrophages tumor trafficking and enhances anti-PD1 efficacy in pancreatic cancer. J Immunother Cancer. 2020;8(1): e000308.
Zhang S, Kohli K, Black RG, Yao L, Spadinger SM, He Q, et al. Systemic interferon-gamma increases MHC class I expression and T-cell Infiltration in cold tumors: results of a phase 0 clinical trial. Cancer Immunol Res. 2019;7(8):1237–43.
Rodig SJ, Gusenleitner D, Jackson DG, Gjini E, Giobbie-Hurder A, Jin C, et al. MHC proteins confer differential sensitivity to CTLA-4 and PD-1 blockade in untreated metastatic melanoma. Sci Transl Med. 2018;10(450):eaar342.
Chen Y, Niu N, Xue J. Double-edged roles of IFNγ in tumor elimination and immune escape. J Pancreatol. 2023;06(01):8–17.
Gocher AM, Workman CJ, Vignali DAA. Interferon-gamma: teammate or opponent in the tumour microenvironment? Nat Rev Immunol. 2022;22(3):158–72.
Abdelfattah N, Kumar P, Wang C, Leu JS, Flynn WF, Gao R, et al. Single-cell analysis of human glioma and immune cells identifies S100A4 as an immunotherapy target. Nat Commun. 2022;13(1):767.
Jerby-Arnon L, Shah P, Cuoco MS, Rodman C, Su MJ, Melms JC, et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell. 2018;175(4):984-97 e24.
Guinn ZP, Petro TM. IFN-gamma synergism with poly I: C reduces growth of murine and human cancer cells with simultaneous changes in cell cycle and immune checkpoint proteins. Cancer Lett. 2018;438:1–9.
Gao AH, Hu YR, Zhu WP. IFN-gamma inhibits ovarian cancer progression via SOCS1/JAK/STAT signaling pathway. Clin Transl Oncol. 2022;24(1):57–65.
Tate DJ, Patterson JR, Finkel-Jimenez B, Zea AH. Interferon (IFNγ) induces cell cycle arrest in RCC cell lines. J Immunother Cancer. 2013;1(Suppl 1):P125.
Zhao YH, Wang T, Yu GF, Zhuang DM, Zhang Z, Zhang HX, et al. Anti-proliferation effects of interferon-gamma on gastric cancer cells. Asian Pac J Cancer Prev. 2013;14(9):5513–8.
Song M, Ping Y, Zhang K, Yang L, Li F, Zhang C, et al. Low-dose IFNgamma induces tumor cell stemness in tumor microenvironment of non-small cell lung cancer. Cancer Res. 2019;79(14):3737–48.
Neuwelt AJ, Kimball AK, Johnson AM, Arnold BW, Bullock BL, Kaspar RE, et al. Cancer cell-intrinsic expression of MHC II in lung cancer cell lines is actively restricted by MEK/ERK signaling and epigenetic mechanisms. J Immunother Cancer. 2020;8(1): e000441.
Axelrod ML, Cook RS, Johnson DB, Balko JM. Biological consequences of MHC-II expression by tumor cells in cancer. Clin Cancer Res. 2019;25(8):2392–402.
Baleeiro RB, Bouwens CJ, Liu P, Di Gioia C, Dunmall LSC, Nagano A, et al. MHC class II molecules on pancreatic cancer cells indicate a potential for neo-antigen-based immunotherapy. Oncoimmunology. 2022;11(1):2080329.
Accolla RS, Ramia E, Tedeschi A, Forlani G. CIITA-driven MHC class II expressing tumor cells as antigen presenting cell performers: toward the construction of an optimal anti-tumor vaccine. Front Immunol. 2019;10:1806.
Dubrot J, Du PP, Lane-Reticker SK, Kessler EA, Muscato AJ, Mehta A, et al. In vivo CRISPR screens reveal the landscape of immune evasion pathways across cancer. Nat Immunol. 2022;23(10):1495–506.
Dong E, Yue XZ, Shui L, Liu BR, Li QQ, Yang Y, et al. IFN-gamma surmounts PD-L1/PD1 inhibition to CAR-T cell therapy by upregulating ICAM-1 on tumor cells. Signal Transduct Target Ther. 2021;6(1):20.
Abiko K, Hamanishi J, Matsumura N, Mandai M. Dynamic host immunity and PD-L1/PD-1 blockade efficacy: developments after “IFN-gamma from lymphocytes induces PD-L1 expression and promotes progression of ovarian cancer.” Br J Cancer. 2023;128(3):461–7.
Lukhele S, Rabbo DA, Guo M, Shen J, Elsaesser HJ, Quevedo R, et al. The transcription factor IRF2 drives interferon-mediated CD8(+) T cell exhaustion to restrict anti-tumor immunity. Immunity. 2022;55(12):2369-85 e10.
Funding
The present study was supported by the Jiangsu Provincial Medical Association Laboratory Medicine Research Special Fund Project (SYH-3201160-0068) and Science and Technology Project of Yixing (2023SF10).
Author information
Authors and Affiliations
Contributions
(I) Conception and design: G Qiao, L Zhou and X Lu. (II) Administrative support: G Qiao and L Zhou. (III) Collection and assembly of data: L Zhou and X Lu; (V) Data analysis and interpretation: L Zhou and X Lu; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper. The authors declare no conflict of interest related to this publication.
Ethical approval and consent to participate
The authors are accountable for all aspects of the work, including ensuring that any questions related to the accuracy or integrity of any part of the work have been appropriately investigated and resolved.
Informed consent
All participants provided informed consent prior to their participation.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12094_2024_3574_MOESM1_ESM.png
Supplementary file1 Marker genes of low-grade glioma and melanoma scRNA-seq clustering analysis. (A) LGG. (B) Melanoma (PNG 915 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, L., Lu, X. & Qiao, G. Single-cell transcriptomic sequencing analysis of mechanistic insights into the IFN-γ signaling pathway in different tumor cells. Clin Transl Oncol (2024). https://doi.org/10.1007/s12094-024-03574-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12094-024-03574-6